Abstract
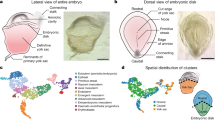
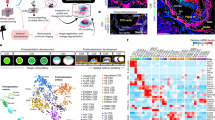
Gastrulation marks a pivotal stage in mammalian embryonic development, establishing the three germ layers and body axis through lineage diversification and morphogenetic movements. However, studying human gastrulating embryos is challenging due to limited access to early tissues. Here we show the use of spatial transcriptomics to analyse a fully intact Carnegie stage 7 human embryo at single-cell resolution, along with immunofluorescence validations in a second embryo. Employing 82 serial cryosections and Stereo-seq technology, we reconstructed a three-dimensional model of the embryo. Our findings reveal early specification of distinct mesoderm subtypes and the presence of the anterior visceral endoderm. Notably, primordial germ cells were located in the connecting stalk, and haematopoietic stem cell-independent haematopoiesis was observed in the yolk sac. This study advances our understanding of human gastrulation and provides a valuable dataset for future research in early human development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the Genome Sequence Archive63 in National Genomics Data Center, China National Center for Bioinformation64, under accession code HRA006197. Previously published human CS7 embryo data that were re-analysed here are available under accession code E-MTAB-9388. Previously published cynomolgus monkey embryo data that were re-analysed here are available under accession code GSE193007. Previously published marmoset CS5–CS7 embryo data that were re-analysed here are available under accession code E-MTAB-9367. Previously published mouse E5.25–E6.5 embryo data that were re-analysed here are available under accession code CRA008972. Previously published human CS8 embryo data that were re-analysed here are available under accession code HRA005567. Previously published human primordial germ cell-like cell data that were re-analysed here are available under accession code GSE205611. Previously published PGC data that were re-analysed here are available under accession code GSE223036. Previously published human primordial germ cell-like cell data that were re-analysed here are available under accession code GSE231812. Human reference genome (hg38) and gene annotations (Ensembl Genome Browser GRCh38.107) were used for alignment and gene count matrix generation. The CellchatDB.human was obtained through https://github.com/sqjin/CellChat. Data supporting the findings of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Bergmann, S. et al. Spatial profiling of early primate gastrulation in utero. Nature 609, 136–143 (2022).
Zhai, J. et al. Primate gastrulation and early organogenesis at single-cell resolution. Nature 612, 732–738 (2022).
Tyser, R. C. V. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Xiao, Z. et al. 3D reconstruction of a gastrulating human embryo. Cell 187, 2855–2874 e19 (2024).
de Bakker, B. S. et al. An interactive three-dimensional digital atlas and quantitative database of human development. Science 354, aag0053 (2016).
Wang, M. et al. High-resolution 3D spatiotemporal transcriptomic maps of developing Drosophila embryos and larvae. Dev. Cell 57, 1271–1283 e1274 (2022).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 e1721 (2022).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
DeTomaso, D. & Yosef, N. Hotspot identifies informative gene modules across modalities of single-cell genomics. Cell Syst. 12, 446–456 e449 (2021).
Liu, G. et al. Spatial transcriptomic profiling to identify mesoderm progenitors with precision genomic screening and functional confirmation. Cell Prolif. 55, e13298 (2022).
Tani, S., Chung, U. I., Ohba, S. & Hojo, H. Understanding paraxial mesoderm development and sclerotome specification for skeletal repair. Exp. Mol. Med. 52, 1166–1177 (2020).
Newton, A. H., Williams, S. M., Major, A. T. & Smith, C. A. Cell lineage specification and signalling pathway use during development of the lateral plate mesoderm and forelimb mesenchyme. Development 149, dev200702 (2022).
Sambasivan, R. & Steventon, B. Neuromesodermal progenitors: a basis for robust axial patterning in development and evolution. Front. Cell Dev. Biol. 8, 607516 (2020).
Yang, R. et al. Amnion signals are essential for mesoderm formation in primates. Nat. Commun. 12, 5126 (2021).
Pham, T. X. A. et al. Modeling human extraembryonic mesoderm cells using naive pluripotent stem cells. Cell Stem Cell 29, 1346–1365 e1310 (2022).
Beddington, R. S. & Robertson, E. J. Axis development and early asymmetry in mammals. Cell 96, 195–209 (1999).
Rivera-Perez, J. A. & Magnuson, T. Primitive streak formation in mice is preceded by localized activation of Brachyury and Wnt3. Dev. Biol. 288, 363–371 (2005).
Madabhushi, M. & Lacy, E. Anterior visceral endoderm directs ventral morphogenesis and placement of head and heart via BMP2 expression. Dev. Cell 21, 907–919 (2011).
Soares, M. L., Torres-Padilla, M. E. & Zernicka-Goetz, M. Bone morphogenetic protein 4 signaling regulates development of the anterior visceral endoderm in the mouse embryo. Dev. Growth Differ. 50, 615–621 (2008).
Sun, X., Meyers, E. N., Lewandoski, M. & Martin, G. R. Targeted disruption of Fgf8 causes failure of cell migration in the gastrulating mouse embryo. Genes Dev. 13, 1834–1846 (1999).
Li, C., Li, Y. P., Fu, X. Y. & Deng, C. X. Anterior visceral endoderm SMAD4 signaling specifies anterior embryonic patterning and head induction in mice. Int J. Biol. Sci. 6, 569–583 (2010).
Andersson, O., Bertolino, P. & Ibanez, C. F. Distinct and cooperative roles of mammalian Vg1 homologs GDF1 and GDF3 during early embryonic development. Dev. Biol. 311, 500–511 (2007).
Kimura, C. et al. Visceral endoderm mediates forebrain development by suppressing posteriorizing signals. Dev. Biol. 225, 304–321 (2000).
Kinder, S. J. et al. The organizer of the mouse gastrula is composed of a dynamic population of progenitor cells for the axial mesoderm. Development 128, 3623–3634 (2001).
Zhu, Q. et al. Decoding anterior-posterior axis emergence among mouse, monkey, and human embryos. Dev. Cell 58, 63–79 e64 (2023).
Finley, K. R., Tennessen, J. & Shawlot, W. The mouse secreted frizzled-related protein 5 gene is expressed in the anterior visceral endoderm and foregut endoderm during early post-implantation development. Gene Expr. Patterns 3, 681–684 (2003).
Kimura-Yoshida, C. et al. Canonical Wnt signaling and its antagonist regulate anterior-posterior axis polarization by guiding cell migration in mouse visceral endoderm. Dev. Cell 9, 639–650 (2005).
Matsuda, K. & Kondoh, H. Dkk1-dependent inhibition of Wnt signaling activates Hesx1 expression through its 5’ enhancer and directs forebrain precursor development. Genes Cells 19, 374–385 (2014).
Kemp, C. R. et al. Expression of Frizzled5, Frizzled7, and Frizzled10 during early mouse development and interactions with canonical Wnt signaling. Dev. Dyn. 236, 2011–2019 (2007).
Richardson, B. E. & Lehmann, R. Mechanisms guiding primordial germ cell migration: strategies from different organisms. Nat. Rev. Mol. Cell Biol. 11, 37–49 (2010).
Guo, F. et al. The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell 161, 1437–1452 (2015).
Garcia-Alonso, L. et al. Single-cell roadmap of human gonadal development. Nature 607, 540–547 (2022).
Ai, Z. et al. Dissecting peri-implantation development using cultured human embryos and embryo-like assembloids. Cell Res 33, 661–678 (2023).
Tang, W. W. C. et al. Sequential enhancer state remodelling defines human germline competence and specification. Nat. Cell Biol. 24, 448–460 (2022).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Yu, L. et al. Derivation of intermediate pluripotent stem cells amenable to primordial germ cell specification. Cell Stem Cell 28, 550–567 e512 (2021).
Irie, N. et al. DMRT1 regulates human germline commitment. Nat. Cell Biol. 25, 1439–1452 (2023).
Zheng, Y. et al. Single-cell analysis of embryoids reveals lineage diversification roadmaps of early human development. Cell Stem Cell 29, 1402–1419 e1408 (2022).
Harrison, S. E., Sozen, B., Christodoulou, N., Kyprianou, C. & Zernicka-Goetz, M. Assembly of embryonic and extraembryonic stem cells to mimic embryogenesis in vitro. Science 356, eaal1810 (2017).
Zhang, J. et al. OTX2 restricts entry to the mouse germline. Nature 562, 595–599 (2018).
Yu, S. et al. BMP4 drives primed to naive transition through PGC-like state. Nat. Commun. 13, 2756 (2022).
Zheng, Y. et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature 573, 421–425 (2019).
Murase, Y. et al. In vitro reconstitution of epigenetic reprogramming in the human germ line. Nature 631, 170–178 (2024).
Esfahani, S. N. et al. Derivation of human primordial germ cell-like cells in an embryonic-like culture. Nat. Commun. 15, 167 (2024).
Vijayakumar, S. et al. Monolayer platform to generate and purify primordial germ-like cells in vitro provides insights into human germline specification. Nat. Commun. 14, 5690 (2023).
Bian, Z. et al. Deciphering human macrophage development at single-cell resolution. Nature 582, 571–576 (2020).
Dzierzak, E. & Bigas, A. Blood development: hematopoietic stem cell dependence and independence. Cell Stem Cell 22, 639–651 (2018).
Wang, H. et al. Decoding human megakaryocyte development. Cell Stem Cell 28, 535–549 e538 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Kumar, N., Mishra, B., Athar, M. & Mukhtar, S. Correction to: inference of gene regulatory network from single-cell transcriptomic data using pySCENIC. Methods Mol. Biol. 2328, C1 (2021).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Jin, S. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. 20, 180–219 (2025).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat. Methods 19, 567–575 (2022).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Chen, T. et al. The genome sequence archive family: toward explosive data growth and diverse data types. Genomics Proteom. Bioinform. 19, 578–583 (2021).
CNCB-NGDC Members and Partners. Database resources of the National Genomics Data Center, China National Center for Bioinformation in 2022. Nucleic Acids Res. 50, D27–D38 (2022).
Acknowledgements
We thank Q. Zhou for his invaluable support and guidance. This work was supported by the National Key Research and Development Program of China (2022YFA1104100 and 2022YFC2702600 to J.G. and 2022YFA1103100 and 2022YFA1104300 to L.Y.), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0820000 to J.G. and L.Y.), the start-up funding support from the Institute of Zoology and Chinese Academy of Sciences to J.G. and L.Y., Initiative Scientific Research program supported by the Institute of Zoology, Chinese Academy of Sciences (202310Z0102 to J.G.), the Chinese Universities Scientific Fund to Y.W., the Biological Breeding-National Science and Technology Major Project 2023ZD0407504 to Y.W., The National Natural Science Foundation of China (32400663 to Y.W., 2022CX11015 and 82371685 to Z.X., and 82101764 to N.H.) and the Beijing Nova Program to Z.X. We acknowledge assistance with the access of analytic instruments from the Translational Medicine Center at The First Affiliated Hospital of Zhengzhou University. We thank all the members from Guo Laboratory, Wei Laboratory, Xiao Laboratory and Yu Laboratory for their fruitful discussions.
Author information
Authors and Affiliations
Contributions
J.G. and L.Y. conceptualized the idea. J.G., Y.W., Z.X. and L.Y. designed, interpreted and supervised the experiments. N.H. performed the human embryo sample collection. Z.X. and Y.W. performed Stereo-seq library construction and sequencing. L.C., S.L., X. Yang, X.W., X.X., J.Y., X.Z., X.L., X. Yan, Y.G., R.L., R.Z., H.M. and B.Z. performed bioinformatics analyses. L.C. and X.W. constructed the online website. N.H. performed immunostaining and confocal photography with assistance from M.J. J.G., Y.W., Z.X. and L.Y. performed paper writing, review and editing with input and feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control and clusters annotation.
a. Immunofluorescent staining results showing the localization of AFP, GABBR2, and DAPI in the human CS7 embryo. Biologically independent experiments were repeated three times with similar results. b. Histogram graph of UMIs/genes/percent.mito per slice. The dashed lines represent the threshold value of quality control. c. Bar graph showing the number of filtered spots in each slice. d. UMAP plots as in Fig. 1b, showing the log expression of UCHL1, TBXT, NOTO, LEFTY2, BMP4, GABRP, AFP, IGF2 and HBG1. e. Bar plot of cell cycle phase by analysing the expression levels of S- and M-phase genes. f. Spatial visualization of clusters with coloured spots in slice 1 (S1), slice 5 (S5), slice 10 (S10), slice 15 (S15), slice 20 (S20), slice 25 (S25), slice 30 (S30), slice 35 (S35), slice 40 (S40), slice 45 (S45), slice 50 (S50), slice 55 (S55), slice 60 (S60), slice 65 (S65), slice 70 (S70), slice 75 (S75), slice 80 (S80).
Extended Data Fig. 2 Spatial characteristics in human CS7 embryo.
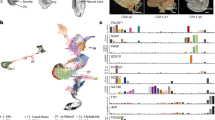
a. Bar plot showing the fractions of clusters in each slice. b. Scatter-plot showing the spatial distribution of the clusters including Epi, Em/EXE.Meso, DE/VE, PS, Noto, Connecting stalk, AM, YS.EXE.Meso, YS.Endo, HEP, Ery. c. Scatter-plot showing the normalized expression of CHRD, CER1, SOX17, BMP4, NANOS3, AFP, CDX1, IGF2, SOX2, TBXT, CDX2, SHH and GABRP in human CS7 reconstructed 3D model.
Extended Data Fig. 3 Spatial characteristics in human CS7 embryo.
a. Bubble plot showing the regulon specificity score (RSS) of significant top 8 regulons by SCENIC analysis in clusters shown in Fig. 1b. Significant, p < 0.05. The hypergeometric P values were adjusted for multiple testing using the Benjamini–Hochberg method. b. Spatial visualization of functional regulon modules based on hotspot analysis in CS7 embryo. Colours represents module scores in each spot. c. Heatmap plot showing the dynamic patterns of regulon modules along anterior-posterior axes. d. Heatmap plot showing the dynamic patterns of regulon modules across cell types. e. Bar plot of the enriched significantly pathways in each regulon module. The hypergeometric P values were adjusted for multiple testing using the Benjamini–Hochberg method.
Extended Data Fig. 4 Spatial distribution of embryonic clusters.
a. Bar graph of the number of spots shown in Fig. 2a in per slice. b. UMAP plots showing the log-normalized expression of MT1G, SOX2, CDX1, TBXT, CHRD, LEFTY2, MSX1, CCKBR, CYB5A, TFAP2A, RGS4, and MYL7. c. Spatial plots showing the embryonic clusters in each slice. The spots coloured in grey indicate extra-embryonic tissues.
Extended Data Fig. 5 Comparison and trajectory analysis in human CS7 embryo.
a. UMAP plot overlaid with PAGA graph showing the developmental trajectory in human CS7 embryonic clusters. b. The scatter-plots showing the expression levels of the changed genes along the velocity pseudotime in lateral plate mesoderm (LP.Meso), axial mesoderm (Axial.Meso) paraxial mesoderm (Para.Meso) and extra-embryonic mesoderm progenitor (EXE.Meso.Prog) lineages. c. UMAP plot showing the cell trajectory in Gast-derived cells along the pseudotime by Monocle3 analysis. d. Heatmap of the percentage cells of embryonic subclusters from Stereo-seq mapped to human cell types at CS7, which suggests our Gast closely resembled their Primitive Steak, our LP.Meso and Axial.Meso closely resembled their Nascent Mesoderm, our Para.Meso closely resembled Emergent Mesoderm, and our EXE.Meso.Prog closely resembled Advanced Mesoderm. e. UMAP visualization of the integrated datasets based on 2000 HVGs of human CS7 embryonic clusters from Stereo-seq and Smart-seq. f. UMAP plot showing the cell trajectory of embryonic subclusters from Stereo-seq and Smart-seq at CS7 along the pseudotime by Monocle3 analysis. g. Ridge plot showing the cell distribution along the pseudotime by Monocle3 analysis.
Extended Data Fig. 6 Comparison analysis with the published dataset.
a. Heatmaps, and bar plots with enrichment pathways showing the expression patterns of the genes that change along the pseudotime trajectory in our data and Tyser data by Monocle3 analysis. The hypergeometric P values were adjusted for multiple testing using the Benjamini–Hochberg method. b. Heatmap showing the scaled expression pattern of genes in human CS7 gastrula. c. Bar plot showing the average expression of CDH2, TBXT, NOTO, MIXL1, TBX6, NTS and FGF4 in human CS7 embryonic clusters from Stereo-seq (blue) and Smart-seq (sky blue) datasets.
Extended Data Fig. 7 Comparison analysis of AVE from human, monkey and mouse.
a. Heatmap plot showing the expression of top 20 markers in endoderm cells. b. Left: PCA plot of visceral endoderm subclusters in mouse E5.25-E6.5 stages. Right: PCA plot of visceral endoderm subclusters in monkey CS8 gastrula. ExVE, embryonic visceral endoderm; EmVE, embryonic visceral endoderm; AVE, anterior visceral endoderm; VE, visceral endoderm. c. PCA plot of the visceral endoderm (VE) cells from human CS7 and monkey CS8 stage. Left: colours indicate clusters; shape indicates monkey. Right: colours indicate cell clusters of monkeys. d. PCA plot of the visceral endoderm (VE) cells from human CS7 and mouse E5.25-E6.5 stages. Left: colours indicate clusters; shape indicate mouse. Right: colours indicate cell clusters of mice. e. Bar plot showing the percentage of cells based on cell cycle-related genes in endodermal subclusters. f. Heatmap showing the percentage cells of endoderm cells allocated to human cells at CS7, monkey cells at CS8, and mouse cells at E5.25-E6.5. DE, definitive endoderm; VE, visceral endoderm; AVE, anterior visceral endoderm; EmVE, embryonic visceral endoderm.
Extended Data Fig. 8 Spatial analysis of key signalling pathways in AVE.
a. Bar plots showing the communication counts of source cells (left) and target cells (right) by ligand–receptor analysis. b. Bubble plots showing the communications between anterior visceral endoderm (AVE) and other clusters in human CS7 embryo. Red indicates target/receptor; Blue indicates source/ligand. The p values obtained from the permutation test were adjusted for multiple testing using the Benjamini–Hochberg method. c. Bubble plots showing the communications between anterior visceral endoderm (AVE) or definitive endoderm (DE2) (left) and other clusters in human CS7 embryo. Red indicates target/receptor; Blue indicates source/ligand. The p values obtained from the permutation test were adjusted for multiple testing using the Benjamini–Hochberg method.
Extended Data Fig. 9 Molecular characterization of AVE and PGCs.
a. Heatmaps showing the expression of genes in BMP, NOTCH, FGF, TGFb, VEGF, and WNT pathways. Blue/red/green represent ligands/receptors/inhibitors. b. Heatmap of top 20 markers of epiblast-derived cells and Connecting stalk. Gast, gastrulating cells; AM.Ecto, amniotic ectoderm; PGCs, primordial germ cells; Epi(P), posterior epiblast. c. PAGA graph showing the two lineage cells by partition-based graph abstraction analysis. d. Left: 3D scatter-plot showing the spatial location of primordial germ cells (PGCs), posterior epiblast (Epi(P)), amniotic ectoderm (AM.Ecto) and amnion (AM). Right: 3D Scatter-plot showing the spatial latent time of primordial germ cells (PGCs), posterior epiblast (Epi(P)), amniotic ectoderm (AM.Ecto) and amnion (AM) by RNA velocity analysis. e. Deconvolution of the Monocle2 pseudotime plot according to clusters. AM.Ecto, amniotic ectoderm; PGC, primordial germ cells; Epi(P), posterior epiblast; AM, amnion. f. Ridge plots showing the distribution of primordial germ cells (PGCs), posterior epiblast (Epi(P)), amniotic ectoderm (AM.Ecto) and amnion (AM) along the pseudotime in CS7 by Monocle2 analysis. g. Bubble plot of ligand–receptor interactions among PGCs, Gast, connecting stalk, and AM.Ecto. Colours indicate communication probability, size indicate significance. The p values obtained from the permutation test were adjusted for multiple testing using the Benjamini–Hochberg method.
Extended Data Fig. 10 Molecular characterization of subtypes of hematopoietic precursors.
a. Bubble plot showing the expression of markers in hematopoietic progenitors. Mesoderm.Prog, mesoderm progenitor; Primitive.HSPCs, primitive hematopoietic stem/progenitor cells; Definitive.HSPCs, definitive hematopoietic stem/progenitor cells; HEC, hemogenic endothelial cells; MP, myeloid progenitors; MEP, megakaryocyte/erythroid progenitors; YS.Endo.Mega, yolk sac endoderm megakaryocyte; YS.Endo.MP, yolk sac endoderm myeloid progenitors; YS.Endo.Ery, yolk sac endoderm erythroblast. b. 3D scatter-plot showing the spatial distribution of hematopoietic precursors in human 3D reconstructed model. c. Pearson correlations among clusters. The numbers indicate Pearson correlations. d. UMAP plot overlayed with RNA velocity showing the trajectory among hematopoietic precursors. e. Immunofluorescent staining results showing the localization of CD45(PTPRC), TFRC(CD71), CD14, AFP and RUNX1 in the human CS7 embryo. Biologically independent experiments were repeated three times with similar results. f. Immunofluorescent staining results showing the localization of DAPI, AFP and CDH1 in the human CS7 embryo. Biologically independent experiments were repeated three times with similar results. g. Enrichment pathways in yolk sac endoderm myeloid progenitors (YS.Endo.MP). The size of the dots represents the counts of genes. The hypergeometric P values were adjusted for multiple testing using the Benjamini–Hochberg method. h. The chord diagram showing the cell-cell communication networks in myeloid progenitors (MP) and hemogenic endothelial cells (HEC). The thickness of the line represents the weight of ligand–receptor pairs. The p values obtained from the permutation test were adjusted for multiple testing using the Benjamini–Hochberg method. i. Bubble plot showing the significant cell-cell communications between MP and other hematopoietic precursors in human CS7. j. Enrichment pathways in yolk sac endoderm megakaryocyte (YS.Endo.Mega). The size of the dots represents the counts of genes. The hypergeometric P values were adjusted for multiple testing using the Benjamini–Hochberg method.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1.
Supplementary Tables 1–8 (download XLSX )
Supplementary Table 1. Top 20 marker genes in each cluster in human CS7 embryo. Supplementary Table 2. Abbreviations in this study. Supplementary Table 3. The representative markers in mesodermal subtypes in the embryonic clusters. Supplementary Table 4. The top 20 markers in subclusters in this study. Supplementary Table 5. The results of ligand–receptor pairs analysis among embryo clusters in this study. Supplementary Table 6. The results of ligand–receptor pairs analysis among epiblast-derived cells in this study. Supplementary Table 7. Markers in haematopoietic progenitors. Supplementary Table 8. Top 20 marker genes in haematopoietic progenitors.
Supplementary Video 1 (download MP4 )
Video of the process of constructing the 3D model of the human CS7 embryo.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig.10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cui, L., Lin, S., Yang, X. et al. Spatial transcriptomic characterization of a Carnegie stage 7 human embryo. Nat Cell Biol 27, 360–369 (2025). https://doi.org/10.1038/s41556-024-01597-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-024-01597-3
This article is cited by
-
Spatial architecture of development and disease
Nature Reviews Genetics (2026)
-
Modelling late gastrulation in stem cell-derived monkey embryo models
Nature (2026)
-
Progress in stem cell-based embryo models and their applications in developmental biology and biomedicine
Nature Reviews Molecular Cell Biology (2026)
-
Mechanisms of human germ cell development
Nature Reviews Molecular Cell Biology (2026)
-
Deciphering signaling mechanisms and developmental dynamics in extraembryonic mesoderm specification from hESCs
Nature Communications (2025)