Abstract
Nicotinamide adenine dinucleotide phosphate (NADPH) is a vital electron donor essential for macromolecular biosynthesis and protection against oxidative stress. Although NADPH is compartmentalized within the cytosol and mitochondria, the specific functions of mitochondrial NADPH remain largely unexplored. Here we demonstrate that NAD+ kinase 2 (NADK2), the principal enzyme responsible for mitochondrial NADPH production, is critical for maintaining protein lipoylation, a conserved lipid modification necessary for the optimal activity of multiple mitochondrial enzyme complexes, including the pyruvate dehydrogenase complex. The mitochondrial fatty acid synthesis (mtFAS) pathway utilizes NADPH for generating protein-bound acyl groups, including lipoic acid. By developing a mass-spectrometry-based method to assess mammalian mtFAS, we reveal that NADK2 is crucial for mtFAS activity. NADK2 deficiency impairs mtFAS-associated processes, leading to reduced cellular respiration and mitochondrial translation. Our findings support a model in which mitochondrial NADPH fuels the mtFAS pathway, thereby sustaining protein lipoylation and mitochondrial oxidative metabolism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The raw metabolomics data are deposited in the Mendeley Repository at https://data.mendeley.com/datasets/x4tkj57mb5/1. The proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE60 partner repository with the dataset identifier PXD060981. All other data are available from the corresponding author upon request. Source data are provided with this paper.
Code availability
The code used to generate the gene coessentiality networks is available via Zenodo at https://doi.org/10.5281/zenodo.14902791 (ref. 61).
Change history
22 May 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41556-025-01695-w
References
Pollak, N., Dolle, C. & Ziegler, M. The power to reduce: pyridine nucleotides—small molecules with a multitude of functions. Biochem. J. 402, 205–218 (2007).
Chandel, N. S. NADPH—the forgotten reducing equivalent. Cold Spring Harb. Perspect. Biol. 13, a040550 (2021).
Ohashi, K., Kawai, S. & Murata, K. Identification and characterization of a human mitochondrial NAD kinase. Nat. Commun. 3, 1248 (2012).
Bartman, C. R., Faubert, B., Rabinowitz, J. D. & DeBerardinis, R. J. Metabolic pathway analysis using stable isotopes in patients with cancer. Nat. Rev. Cancer 23, 863–878 (2023).
Praharaj, P. P. et al. Cryo-EM structure and regulation of human NAD kinase. Sci. Adv. 11, eads2664 (2025).
Murphy, M. P. How mitochondria produce reactive oxygen species. Biochem. J. 417, 1–13 (2009).
Tran, D. H. et al. Mitochondrial NADP+is essential for proline biosynthesis during cell growth. Nat. Metab. 3, 571–585 (2021).
Zhu, J. et al. Mitochondrial NADP(H) generation is essential for proline biosynthesis. Science 372, 968–972 (2021).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Hoxhaj, G. & Manning, B. D. The PI3K-AKT network at the interface of oncogenic signalling and cancer metabolism. Nat. Rev. Cancer 20, 74–88 (2020).
Wedan, R. J., Longenecker, J. Z. & Nowinski, S. M. Mitochondrial fatty acid synthesis is an emergent central regulator of mammalian oxidative metabolism. Cell Metab. 36, 36–47 (2024).
Nowinski, S. M. et al. Mitochondrial fatty acid synthesis coordinates oxidative metabolism in mammalian mitochondria. eLife 9, e58041 (2020).
Nowinski, S. M., Van Vranken, J. G., Dove, K. K. & Rutter, J. Impact of mitochondrial fatty acid synthesis on mitochondrial biogenesis. Curr. Biol. 28, R1212–R1219 (2018).
Chuman, L. & Brody, S. Acyl carrier protein is present in the mitochondria of plants and eucaryotic micro-organisms. Eur. J. Biochem. 184, 643–649 (1989).
Goldman, P., Alberts, A. W. & Vagelos, P. R. The condensation reaction of fatty acid synthesis III. Identification of the protein-bound product of the reaction and its conversion to long chain fatty acids. J. Biol. Chem. 238, 3579–3583 (1963).
Solmonson, A. & DeBerardinis, R. J. Lipoic acid metabolism and mitochondrial redox regulation. J. Biol. Chem. 293, 7522–7530 (2018).
Rowland, E. A., Snowden, C. K. & Cristea, I. M. Protein lipoylation: an evolutionarily conserved metabolic regulator of health and disease. Curr. Opin. Chem. Biol. 42, 76–85 (2018).
Brody, S., Oh, C., Hoja, U. & Schweizer, E. Mitochondrial acyl carrier protein is involved in lipoic acid synthesis in Saccharomyces cerevisiae. FEBS Lett. 408, 217–220 (1997).
Rahman, M. T. et al. An engineered variant of MECR reductase reveals indispensability of long-chain acyl-ACPs for mitochondrial respiration. Nat. Commun. 14, 619 (2023).
Chen, Z. J. et al. Structural enzymological studies of 2-enoyl thioester reductase of the human mitochondrial FAS II pathway: new insights into its substrate recognition properties. J. Mol. Biol. 379, 830–844 (2008).
Heimer, G. et al. MECR mutations cause childhood-onset dystonia and optic atrophy, a mitochondrial fatty acid synthesis disorder. Am. J. Hum. Genet. 99, 1229–1244 (2016).
Nair, R. R. et al. Genetic modifications of MECR reveal a role for mitochondrial 2-enoyl-CoA/ACP reductase in placental development in mice. Hum. Mol. Genet. 26, 2104–2117 (2017).
Houten, S. M. et al. Mitochondrial NADP(H) deficiency due to a mutation in NADK2 causes dienoyl-CoA reductase deficiency with hyperlysinemia. Hum. Mol. Genet. 23, 5009–5016 (2014).
Arnold, P. K. et al. A non-canonical tricarboxylic acid cycle underlies cellular identity. Nature 603, 477–481 (2022).
Martínez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Sahu, U. et al. Pyrimidines maintain mitochondrial pyruvate oxidation to support de novo lipogenesis. Science 383, 1484–1492 (2024).
Patel, M. S., Nemeria, N. S., Furey, W. & Jordan, F. The pyruvate dehydrogenase complexes: structure-based function and regulation. J. Biol. Chem. 289, 16615–16623 (2014).
Arnold, P. K. & Finley, L. W. S. Regulation and function of the mammalian tricarboxylic acid cycle. J. Biol. Chem. 299, 102838 (2023).
Luengo, A. et al. Increased demand for NAD+relative to ATP drives aerobic glycolysis. Mol. Cell 81, 691–707 e696 (2021).
Mathias, R. A. et al. Sirtuin 4 is a lipoamidase regulating pyruvate dehydrogenase complex activity. Cell 159, 1615–1625 (2014).
Tsvetkov, P. et al. Copper induces cell death by targeting lipoylated TCA cycle proteins. Science 375, 1254–1261 (2022).
Ni, M. et al. Functional assessment of lipoyltransferase-1 deficiency in cells, mice, and humans. Cell Rep. 27, 1376–1386 e1376 (2019).
Mary, C. et al. Crystal structure of human NADK2 reveals a dimeric organization and active site occlusion by lysine acetylation. Mol. Cell 82, 3299–3311.e8 (2022).
Tao, R. et al. Genetically encoded fluorescent sensors reveal dynamic regulation of NADPH metabolism. Nat. Methods 14, 720–728 (2017).
Solmonson, A. et al. Compartmentalized metabolism supports midgestation mammalian development. Nature 604, 349–353 (2022).
Endo, S. et al. Human carbonyl reductase 4 is a mitochondrial NADPH-dependent quinone reductase. Biochem. Biophys. Res. Commun. 377, 1326–1330 (2008).
Nam, J. W. et al. A general method for quantification and discovery of acyl groups attached to acyl carrier proteins in fatty acid metabolism using LC–MS/MS. Plant Cell 32, 820–832 (2020).
Wainberg, M. et al. A genome-wide atlas of co-essential modules assigns function to uncharacterized genes. Nat. Genet. 53, 638–649 (2021).
Van Vranken, J. G. et al. ACP acylation is an acetyl-CoA-dependent modification required for electron transport chain assembly. Mol. Cell 71, 567–580 e564 (2018).
Kursu, V. A. et al. Defects in mitochondrial fatty acid synthesis result in failure of multiple aspects of mitochondrial biogenesis in Saccharomyces cerevisiae. Mol. Microbiol. 90, 824–840 (2013).
Schonauer, M. S., Kastaniotis, A. J., Hiltunen, J. K. & Dieckmann, C. L. Intersection of RNA processing and the type II fatty acid synthesis pathway in yeast mitochondria. Mol. Cell. Biol. 28, 6646–6657 (2008).
Brown, A. et al. Structures of the human mitochondrial ribosome in native states of assembly. Nat. Struct. Mol. Biol. 24, 866–869 (2017).
Dibley, M. G. et al. The mitochondrial acyl-carrier protein interaction network highlights important roles for LYRM family members in complex I and mitoribosome assembly. Mol. Cell Proteom. 19, 65–77 (2020).
Yousefi, R. et al. Monitoring mitochondrial translation in living cells. EMBO Rep. 22, e51635 (2021).
Wada, H., Shintani, D. & Ohlrogge, J. Why do mitochondria synthesize fatty acids? Evidence for involvement in lipoic acid production. Proc. Natl Acad. Sci. USA 94, 1591–1596 (1997).
Tajima, K. et al. Mitochondrial lipoylation integrates age-associated decline in brown fat thermogenesis. Nat. Metab. 1, 886–898 (2019).
Goodman, R. P., Calvo, S. E. & Mootha, V. K. Spatiotemporal compartmentalization of hepatic NADH and NADPH metabolism. J. Biol. Chem. 293, 7508–7516 (2018).
Masud, A. J., Kastaniotis, A. J., Rahman, M. T., Autio, K. J. & Hiltunen, J. K. Mitochondrial acyl carrier protein (ACP) at the interface of metabolic state sensing and mitochondrial function. Biochim. Biophys. Acta Mol. Cell. Res. 1866, 118540 (2019).
Hoxhaj, G. et al. Direct stimulation of NADP+synthesis through Akt-mediated phosphorylation of NAD kinase. Science 363, 1088–1092 (2019).
Soflaee, M. H. et al. Purine nucleotide depletion prompts cell migration by stimulating the serine synthesis pathway. Nat. Commun. 13, 2698 (2022).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Brautigam, C. A., Zhao, H., Vargas, C., Keller, S. & Schuck, P. Integration and global analysis of isothermal titration calorimetry data for studying macromolecular interactions. Nat. Protoc. 11, 882–894 (2016).
Keller, S. et al. High-precision isothermal titration calorimetry with automated peak-shape analysis. Anal. Chem. 84, 5066–5073 (2012).
Scheuermann, T. H. & Brautigam, C. A. High-precision, automated integration of multiple isothermal titration calorimetric thermograms: new features of NITPIC. Methods 76, 87–98 (2015).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT—a platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Brautigam, C. A. Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol. 562, 109–133 (2015).
Jenkins, L. M., Nam, J. W., Evans, B. S. & Allen, D. K. Quantification of acyl–acyl carrier proteins for fatty acid synthesis using LC-MS/MS. Methods Mol. Biol. 2295, 219–247 (2021).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576 e516 (2017).
Tran, D. H. et al. De novo and salvage purine synthesis pathways across tissues and tumors. Cell 187, 3602–3618.e20 (2024).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–d553 (2025).
Marckx, A. gene-coessentiality-map. Zenodo https://doi.org/10.5281/zenodo.14902791(2025).
Acknowledgements
We thank Y. Yang (East China University of Science and Technology) for providing iNAP plasmids34 and Y. Zhang for insightful discussions. We also thank the UTSW Quantitative Light Microscopy Core, a shared resource of the Harold C. Simmons Cancer Center, supported in part by NCI (1P30 CA142543-01), the UT Southwestern Proteomics Core and the US Department of Agriculture, Agriculture Research Service that supports D.K.A. The research is supported by the NIH (grant no. R01GM143236 to G.H., grant no. 3R01GM143236-02S1 (equipment grant) to G.H.), Welch foundation (grant no. I-2067-20240404 to G.H), CPRIT (grant no. RP240035 to G.H.) and USDA National Institute of Food and Agriculture Award (grant no. 2021-67013-33778 to D.K.A.). G.H. is a recipient of the Pew-Stewart Scholar, CPRIT Scholar (CPRIT; RR190087), ACS Scholar awards (RSG-22-177-01-TBE) and V Scholar (V2021-019). D.K. is supported by a CPRIT training grant (grant no. RP210041 to D.K.) and C.M. by an NIH F30 fellowship (grant no. DK137407 to C.M.).
Author information
Authors and Affiliations
Contributions
D.K. performed and analysed most experiments. R.K., T.D., C.M., P.P.P., M.H.S., K.R., H.B., H.J.T. and P.M. conducted experiments and analysed and discussed data. S.M. and D.K.A. contributed to the mtFAS analysis, A.M. to the computational work, A.L. to the post-translational modification analysis, R.K. and M.M. to the microscopy, S.T. and C.A.B. to the ITC and B.K., F.C. and H.S.V. to mass spectrometry. D.K. and G.H. conceptualized the study and wrote the manuscript, and G.H. directed the study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Yihui Shen, Huiyong Yin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 NADK2 supports pyruvate oxidation and tumor growth.
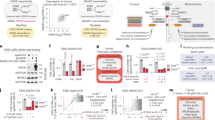
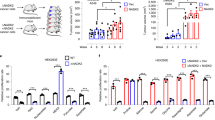
(a) Fractional abundance (%) of the indicated TCA cycle metabolites from isogenic ∆NADK2 A375 cells expressing either empty vector or NADK2 and labeled with [13C3]-pyruvate for 30 min in 0.2 mM proline. (n = 4 biological replicates). (b) As in (a), but the experiments were performed in isogenic ∆NADK2 A549 cells expressing either empty vector or NADK2 and labeled with [13C3]-pyruvate in the presence of 0.2 mM proline for 15, 30, or 60 min. (n = 4 biological replicates). Related to Fig. 1d. (c) As in (b), but the experiments were performed in isogenic ∆NADK2 HeLa cells expressing either empty vector or NADK2. (n = 4 biological replicates). Related to Fig. 1e. (d) As in (b), but the cells were labeled with [13C3]-pyruvate for 1 h. The peak area was normalized to protein abundance. (n = 4 biological replicates). (e) As in (d), but the experiments were performed in isogenic ∆NADK2 HeLa cells expressing either empty vector or NADK2. (n = 4 biological replicates). (f) Relative citrate (M + 2)/ pyruvate (M + 2) ratio from (d). (n = 4 biological replicates). (g) Relative citrate (M + 2)/ pyruvate (M + 2) ratio from (e). (n = 4 biological replicates). (h) Schematic of A375 tumor xenograft. Related to Fig. 1l, m. (i) Tumor growth from isogenic ∆NADK2 A375 cells expressing either empty vector or NADK2. (n = 8 biologically independent animals). Related to Fig. 1l, m. (j) The size of tumors derived from isogenic ∆NADK2 A375 cells expressing either empty vector or NADK2 at the end of the experiment. (n = 8 biologically independent tumors).Related to Fig. 1l, m. (k) Normalized proline abundance from tumors derived from isogenic ∆NADK2 A375 cells expressing either empty vector or NADK2. (n = 4 biologically independent tumors). Data are presented as the mean ± standard deviation (a-g, j, and k) or mean ± s.e.m. (i). *P < 0.05, **P < 0.01, and ***P < 0.001 were calculated using a two-tailed Student’s t-test (a-c, and i-k) or one-way ANOVA with Tukey’s post hoc test (d-g).
Extended Data Fig. 2 Impact of NADK2 on redox ratios and the TCA cycle.
(a) The relative levels of NAD+ and NADH and NAD+/NADH ratio in mitochondria isolated from isogenic ∆NADK2 HeLa cells expressing either empty vector or NADK2. The cells were cultured in the presence of 0.2 mM proline. (n = 4 biologically independent replicates). (b) The relative levels of NAD+ and NADH and NAD+/NADH ratio in mitochondria isolated from isogenic ∆NADK2 A549 cells expressing either empty vector or NADK2. The cells were cultured in the presence of 0.2 mM proline. (n = 4 biologically independent replicates). (c) Coomassie staining of immunoprecipitated DLAT-HA from isogenic ∆NADK2 HeLa cells expressing either empty vector or NADK2 and transfected with DLAT-HA. The cells were cultured in the presence of 0.2 mM proline. The bands were cut out and subjected to mass spectrometry for detecting lipoylated peptides. (d) The abundance of lipoylated DLAT as described in (c). (e) Schematic of [13C5]-glutamine tracing. (f) The normalized peak area (labeled and total pools) of glutamine in isogenic ΔNADK2 HeLa cells expressing either empty vector or NADK2 labeled with [13C5]-glutamine for 1 h. The peak areas were normalized to the protein amount assessed with a BCA assay. (n = 4 biological replicates). (g) As in (f), but the normalized peak area (labeled and total pools) of glutamate are shown. (n = 4 biological replicates). (h) As in (f), but the normalized peak area (labeled and total pools) of alpha-ketoglutarate are shown. (n = 4 biological replicates). (i) As in (f), but the normalized peak area (labeled and total pools) of succinyl-CoA are shown. (n = 4 biological replicates). (j) The relative OGDH activity from mitochondrial extracts of isogenic ΔNADK2 HeLa cells expressing either empty vector or NADK2, which were grown in DMEM containing 10% FBS and 0.2 mM proline. (n = 3 biological replicates). Data are presented as the mean ± standard deviation (a,b,f-j). *P < 0.05, **P < 0.01, and ***P < 0.001 were calculated using a two-tailed Student’s t-test (a,b,f-j).
Extended Data Fig. 3 Impact of ROS, proline, and lipids on protein lipoylation.
(a) Immunoblots showing DLAT lipoylation, DLAT, NADK2, and vinculin levels from isogenic ΔNADK2 A549 cells expressing either empty vector or NADK2. Cells were grown in the presence of 0.2 mM proline and treated with the indicated antioxidants for 48 h. GSHee; glutathione ethyl ester (5 mM), NAC; N-acetyl cysteine (1 mM), and Trolox (5 μM). (b) Immunoblotting was done as in (a), but cells were grown in the presence or absence of 0.2 mM proline. (c) Immunoblotting was done as in (a), but cells were grown in the presence of 0.2 mM proline and treated with octanoic acid (50 μM), lipoic acid (50 μM), palmitic acid (100 μM), oleic acid (100 μM), low-density lipoprotein (LDL, 50 μg/ml), or high-density lipoprotein (HDL, 50 μg/ml) for 48 h. (d) Immunoblotting was done as in (a) but in ΔNADK2 A549 cells expressing either empty vector or NADK2. (e) Immunoblotting was done as in (a), but from WT or isogenic ΔNADK2 A549 cells expressing empty vector, NADK2, D161A NADK2, MTS-NADK, or cytosolic NADK. D161A NADK2: kinase-dead NADK2; MTS: Mitochondrial Targeting Sequence. (f) The abundance of intracellular lipoic acid from isogenic ∆NADK2 HeLa or A549 cells expressing either empty vector or NADK2 treated with either vehicle or lipoic acid (50 μM) for 48 h assessed via LC/MS. The peak areas were normalized to the protein amount. (n = 4 biologically independent replicates). Data are presented as the mean ± standard deviation. **P < 0.01 was calculated using a two-tailed Student’s t-test. (g) Immunoblots showing ACP, NADK2, and vinculin levels from WT or ∆NADK2 HeLa cells.
Extended Data Fig. 4 The synthesis and detection of acyl-ACP standards via LC/MS.
(a) Schematic illustrating workflow for synthesizing acyl-DSL standard for acyl-ACP measurement. (b) The mass spectrometry parameters of various acyl-DSLs. (c-j) The chromatogram of synthesized standards (black) and biological HeLa WT (blue) for Apo-ACP (c), Holo-ACP (d), C2-ACP (e), C4-ACP (f), C6-ACP (g), C8-ACP (h), C10-ACP (i), C12-ACP (j) run on LC/MS.
Extended Data Fig. 5 Impact of NADK2 and mtFAS on mitochondrial translation.
(a) Coomassie staining of purified ACP-FLAG from WT and ΔMECR HeLa cells, which was then subjected to Asp-N peptidase treatment for assessment of acyl-ACP species via LC/MS. (b) Representative immunofluorescence images for ACP-FLAG (green) detected with anti-FLAG antibody MitoTracker Red (red), and nuclei (blue) stained with Hoechst for WT and ΔNADK2 HeLa cells cultured in the presence of 0.2 mM proline. Scale bars, 10 µm. (c) The NADK2 coessentiality network showing the top 6 enrichment gene clusters. (d) The MECR coessentiality network showing the top 6 enrichment gene clusters. (e) Volcano plot illustrating the log2 fold change of mitoribosome-related proteins identified in ACP immunoprecipitates from wild-type (WT), ΔNADK2, and ΔMECR HeLa cells. Proteins belonging to the mitochondrial ribosomal protein large subunit (MRPL) or small subunit (MRPS), as well as other interactors, are indicated and color-coded. (f) Schematic illustrating the workflow of click chemistry-based mitochondrial translation assay.
Supplementary information
Supplementary Table (download XLSX )
Oligonucleotide sequences.
Source data
Source Data (download PDF )
Uncropped gels and blots.
Source Data (download XLSX )
Source numerical data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, D., Kesavan, R., Ryu, K. et al. Mitochondrial NADPH fuels mitochondrial fatty acid synthesis and lipoylation to power oxidative metabolism. Nat Cell Biol 27, 790–800 (2025). https://doi.org/10.1038/s41556-025-01655-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01655-4
This article is cited by
-
SOX8/CPT2 axis regulates lipid metabolism to support enzalutamide resistance in prostate cancer
Cancer Cell International (2026)
-
Remodeling and retooling metabolism
Nature Chemical Biology (2025)