Abstract
Dietary nutrients are inextricably linked to antitumour immune responses. However, the effect of diet-derived galactose on antitumour immunity remains unclear. Here we show that dietary galactose augments CD8+ T cell immunity to suppress tumour progression. High-galactose feeding drives hepatocyte-derived insulin-like growth factor binding protein 1 (IGFBP-1) production, thus restraining IGF-1 signalling-dependent T cell exhaustion. IGF-1 receptor (IGF-1R) deficiency in T cells potentiates antitumour CD8+ T cell responses and phenocopies high-galactose feeding by preventing T cell exhaustion. Circulating galactose reprograms hepatocyte metabolism to inactivate mTORC1, thereby inducing the production of IGFBP-1 to boost CD8+ T cell function. Furthermore, patients with cancer who have high plasma IGFBP-1 levels exhibit blocked T cell exhaustion and enhanced T cell responses in tumour tissues. These findings reveal that dietary galactose specifically elicits potent antitumour CD8+ T cell responses by facilitating hepatocyte-derived IGFBP-1 production, providing insights into the development of more effective immunotherapies against cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw scRNA-seq, ATAC–seq and RNA-seq data reported in this study have been deposited in the Genome Sequence Archive (Genomics, Proteomics and Bioinformatics 2021) in National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA015185, CRA015187, CRA015188, CRA015190 and CRA015203) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa. The processed public human liver scRNA-seq datasets were collected from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE124395. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
References
McIntyre, C. L., Temesgen, A. & Lynch, L. Diet, nutrient supply, and tumor immune responses. Trends Cancer 9, 752–763 (2023).
Ji, M., Xu, Q. & Li, X. Dietary methionine restriction in cancer development and antitumor immunity. Trends Endocrinol. Metab. 35, 400–412 (2024).
Wu, Y. et al. Neutrophil profiling illuminates anti-tumor antigen-presenting potency. Cell 187, 1422–1439 (2024).
Wu, Y., Zou, Q., Jiang, P. & Gao, Q. Tumor–host cometabolism collaborates to shape cancer immunity. Cancer Discov. 14, 653–657 (2024).
Fan, H. et al. Trans-vaccenic acid reprograms CD8+ T cells and anti-tumour immunity. Nature 623, 1034–1043 (2023).
Bender, M. J. et al. Dietary tryptophan metabolite released by intratumoral Lactobacillus reuteri facilitates immune checkpoint inhibitor treatment. Cell 186, 1846–1862 (2023).
Zhang, Y. et al. Dietary fructose-mediated adipocyte metabolism drives antitumor CD8+ T cell responses. Cell Metab. 35, 2107–2118 (2023).
Newens, K. J. & Walton, J. A review of sugar consumption from nationally representative dietary surveys across the world. J. Hum. Nutr. Diet. 29, 225–240 (2016).
Conte, F., van Buuringen, N., Voermans, N. C. & Lefeber, D. J. Galactose in human metabolism, glycosylation and congenital metabolic diseases: time for a closer look. Biochim. Biophys. Acta, Gen. Subj. 1865, 129898 (2021).
Drozdowski, L. A. & Thomson, A. B. Intestinal sugar transport. World J. Gastroenterol. 12, 1657–1670 (2006).
Dall’Olio, F. Glycobiology of aging. Subcell. Biochem. 90, 505–526 (2018).
Chang, C. H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Rossignol, R. et al. Energy substrate modulates mitochondrial structure and oxidative capacity in cancer cells. Cancer Res. 64, 985–993 (2004).
Lee, J. W. et al. Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature 567, 249–252 (2019).
Li, X. et al. The immunological and metabolic landscape in primary and metastatic liver cancer. Nat. Rev. Cancer 21, 541–557 (2021).
Stone, M. L. et al. Hepatocytes coordinate immune evasion in cancer via release of serum amyloid A proteins. Nat. Immunol. 25, 755–763 (2024).
Coelho, A. I., Berry, G. T. & Rubio-Gozalbo, M. E. Galactose metabolism and health. Curr. Opin. Clin. Nutr. Metab. Care 18, 422–427 (2015).
Baxter, R. C. IGF binding proteins in cancer: mechanistic and clinical insights. Nat. Rev. Cancer 14, 329–341 (2014).
Allard, J. B. & Duan, C. IGF-binding proteins: why do they exist and why are there so many? Front. Endocrinol. 9, 117 (2018).
Lin, Y. W. et al. IGFBP-1 in cancer: expression, molecular mechanisms, and potential clinical implications. Am. J. Transl. Res. 13, 813–832 (2021).
Tong, L. et al. Transketolase promotes MAFLD by limiting inosine-induced mitochondrial activity. Cell Metab. 36, 1013–1029 (2024).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Orozco, J. M. et al. Dihydroxyacetone phosphate signals glucose availability to mTORC1. Nat. Metab. 2, 893–901 (2020).
Li, M. et al. Aldolase is a sensor for both low and high glucose, linking to AMPK and mTORC1. Cell Res. 31, 478–481 (2021).
Rao, R. R., Li, Q., Gubbels Bupp, M. R. & Shrikant, P. A. Transcription factor Foxo1 represses T-bet-mediated effector functions and promotes memory CD8+ T cell differentiation. Immunity 36, 374–387 (2012).
Jerome, L., Shiry, L. & Leyland-Jones, B. Deregulation of the IGF axis in cancer: epidemiological evidence and potential therapeutic interventions. Endocr. Relat. Cancer 10, 561–578 (2003).
Clark, R. The somatogenic hormones and insulin-like growth factor-1: stimulators of lymphopoiesis and immune function. Endocr. Rev. 18, 157–179 (1997).
Smith, T. J. Insulin-like growth factor-I regulation of immune function: a potential therapeutic target in autoimmune diseases? Pharm. Rev. 62, 199–236 (2010).
DiToro, D. et al. Insulin-like growth factors are key regulators of T helper 17 regulatory T cell balance in autoimmunity. Immunity 52, 650–667 (2020).
Ajona, D. et al. Short-term starvation reduces IGF-1 levels to sensitize lung tumors to PD-1 immune checkpoint blockade. Nat. Cancer 1, 75–85 (2020).
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (grant numbers 82225020 and 82430054 to Q.Z., and 82373072 to Z.-Y.N.), National Key Research and Development Program of China (grant numbers 2020YFA0803603 and 2021YFA1301402 to Q.Z.), Natural Science Foundation of Hebei Province (grant number H2022201067 to Z.-Y.N.), and Central Government Guides Local Science and Technology Development Special Project (grant number 236Z7744G to Z.-Y.N.).
Author information
Authors and Affiliations
Contributions
X.D., W.L., G.L. and C.G. designed and performed the experiments, prepared the figures and wrote the manuscript. X.T., R.B., R.L., H.H., S.X., X.Y., Q.H., J.W., S.L., J.S., R.Z. and Y.Y. contributed to the performance of the experiments. Q.G., Z.-Y.N., X.C. and Q.Z. supervised the work and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Ping-Chih Ho, Cliona O’Farrelly and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Diet-derived galactose augments antitumour immunity to control tumour growth.
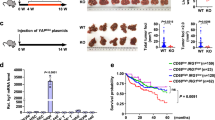
a, Pearson correlation analysis of plasma central carbon-associated metabolites versus the level of plasma galactose of patients with RCC (n = 27 patients), LC (n = 28 patients) and CRC (n = 25 patients). Points represent individual metabolites (positive correlated metabolites highlight in red). b, Survival curve (n = 10 mice) of C57BL/6 WT mice fed with a high-galactose diet or normal diet and injected with MC38 colon cancer cells at day 14 after high-galactose treatment. c, The orthotopic tumours on day 15 after orthotopic implantation of MC38 cells (n = 6 mice per group). WT mice fed with a high-galactose diet or normal diet and orthotopically implanted with MC38 cancer cells at day 14 after high-galactose treatment. Scale bar, 1 cm. d, The weight of orthotopic tumours was measured on day 18 after implantation of MC38 cells (n = 6 mice). WT mice fed with a high-galactose diet or normal diet and orthotopically implanted with MC38 cancer cells at day 14 after high-galactose treatment. The experiments were independently repeated two (c,d) or three (b) times. The data (d) are presented as mean ± SEM. P values were calculated by two-sided Pearson’s correlation coefficient assay (a), log-rank (Mantel–Cox) test (b) and two-tailed Student’s t-test (d).
Extended Data Fig. 2 scRNA-seq analysis of tumour-infiltrating T cells.
a, Dot Plot illustrates the relative average expression levels of selected marker genes across all identified clusters from Fig. 2a. Marker genes for each cluster were determined based on their log-fold change relative to the other clusters. b, Dot plot illustrates the relative average expression levels of selected marker genes across all identified clusters from Fig. 2c. Marker genes for each cluster were determined based on their log-fold change relative to the other clusters.
Extended Data Fig. 3 Dietary galactose prevents tumour-infiltrating CD8+ T cell exhaustion.
a, Cell numbers of tumour-infiltrating CD8+ T cells from B16-F10 or LLC tumour-bearing C57BL/6 WT mice fed a high-galactose diet or normal diet. Tumour-infiltrating CD8+ T cells were isolated at day 14 after tumour injection. Cell number was normalized to 1 g tumour tissue (n = 7 mice). b, Flow cytometric analysis of the proportion and numbers of tumour-infiltrating IFN-γ+CD8+ T cells and Ki-67+CD8+ T cells from B16-F10 tumour-bearing C57BL/6 WT mice on a high-galactose or normal diet (n = 7 mice). c, Proportion of tumour-infiltrating Ki-67+ (n = 7 mice) and IFN-γ+ (n = 5 mice) CD4+ T cells from B16-F10 tumour-bearing C57BL/6 mice on a high-galactose or normal diet. d, Flow cytometric analysis of proportion and numbers of tumour-infiltrating PD-1+TIM-3+, TCF-1+TIM-3− CD8+ T cells from LLC tumour-bearing C57BL/6 WT mice fed a high-galactose diet or normal diet. Cell number was normalized to 1 g tumour tissue (n = 7 mice). e,f, Flow cytometric analysis of the proportion of tumour-infiltrating IFN-γ+CD8+ T cells (e, n = 8 mice), Ki-67+CD8+ T cells (n = 7 mice), IFN-γ+CD4+ T cells and Ki-67+CD4+ T cells (f, n = 6 mice) from LLC tumour-bearing C57BL/6 WT mice on a high-galactose or normal diet. g, Tumour size of B16-F10 tumour-bearing C57BL/6 WT mice. C57BL/6 WT mice were injected with B16-F10 tumour cells and then fed with normal control diet (n = 12 mice) or high-galactose diet (n = 13 mice). h, Flow cytometric analysis of proportion of PD-1+TIM-3+, TCF-1+TIM-3− CD8+ T cells from tumours of B16-F10 tumour-bearing mice from g (Ctrl, n = 7 mice; Gal, n = 8 mice). i, Flow cytometric analysis of proportion of IFN-γ+CD8+, Ki-67+CD8+, Ki-67+CD4+ T cells from tumours of B16-F10 tumour-bearing mice from g (n = 8 mice). The experiments were independently repeated two times. Data are presented as mean ± SEM. P values were calculated by two-tailed Student’s t-test (a-f,h,i) and two-way ANOVA with Geisser–Greenhouse correction (g). ns, not statistically significant.
Extended Data Fig. 4 The effect of dietary galactose on systemic T cell phenotypes.
a, Flow cytometric analysis of proportion of CD4+, CD4+Ki-67+, CD4+IFN-γ+, CD8+, CD8+Ki-67+, CD8+IFN-γ+ T cells from liver (n = 6 mice), spleen (n = 6 mice) and lymph node (n = 6 mice) of B16-F10 tumour-bearing C57BL/6 WT mice fed a high-galactose diet or normal diet. b, Tumour size of female C57BL/6 WT mice fed with normal (Ctrl, n = 12 mice) diet or high-galactose diet (Gal, n = 13 mice) and injected with B16-F10 melanoma cells at day 14 after high-galactose treatment. c, Flow cytometric analysis of proportion of PD-1+TIM-3+, TCF-1+TIM-3− and IFN-γ+ CD8+ T cells (n = 9 mice) in tumours from mice of b. The experiments were independently repeated two times. Data are presented as mean ± SEM. P values were calculated by two-tailed Student’s t-test (a,c) and two-way ANOVA with Geisser–Greenhouse correction (b). ns, not statistically significant.
Extended Data Fig. 5 Dietary galactose promotes antitumour CD8+ T cell immunity via IGFBP-1.
a, Flow cytometric analysis of proportion of IFN-γ production in CD4+ T cells and CD8+ T cells and exhausted CD8+ T cells treated with 10 mM glucose, 10 mM galactose or 10 mM glucose plus 100 μM galactose (n = 4 biological replicates). CD4+ T cells and CD8+ T cells were isolated from C57BL/6 WT mice. b, Flow cytometric analysis of exhausted CD8+ T cells treated with or without IGF-1 or/and IGFBP-1. c, Flow cytometric analysis of exhausted Igf1r+/+Cd4-Cre and Igf1rfl/flCd4-Cre CD8+ T cells treated with or without IGF-1. d, ELISA analysis of serum IGF-1 from C57BL/6 WT mice fed a high-galactose diet or normal diet at the indicated time points after tumour challenge (n = 5 mice). d, day. e, Immunoblot analysis of the IGFBP-1 proteins in the liver tissue from C57BL/6 WT mice treated with either GalNAc–siIGFBP1 or GalNAc–NC. Liver samples were collected at day 3 after GalNAc–siIGFBP1 treatment. f, Flow cytometric analysis of exhausted CD8+ T cells cultured with medium of hepatocyte from mice with or without GalNAc–siIGFBP1 treatment supplied with or without 10 mM galactose (n = 5 biological replicates). g, Tumour size in C57BL/6 WT mice fed a high-galactose or normal diet and injected s.c. with B16-F10 cells at day 14 after high-galactose treatment. These mice were treated intraperitoneally with 0.03 mg/kg anti-IGFBP-1 or control antibodies (Ctrl Ab) every two days (n = 10 mice) at day 7 after high-galactose treatment. h, Flow cytometric analysis of PD-1 and TIM-3 expression (left, n = 10 mice for Ctrl, n = 8 mice for galactose plus Ctrl Ab and n = 9 mice for galactose plus anti-IGFBP-1) or IFN-γ expression (right, n = 9 mice) in CD8+ T cells from tumours of B16-F10 tumour-bearing C57BL/6 WT mice from g. Tumour-infiltrating CD8+ T cells were isolated at day 14 after tumour injection. The experiments were independently repeated two (a,d–h) or three (b,c) times. Data are presented as mean ± SEM. P values were calculated by one-way ANOVA with Tukey’s multiple comparisons (a,f,h), two-tailed Student’s t-test (d) and two-way ANOVA with Geisser–Greenhouse correction (g). ns, not statistically significant.
Extended Data Fig. 6 IGF-1 signaling transcriptionally facilitates exhausted CD8 + T cell differentiation.
a, Principal component analysis (PCA) of ATAC–seq (upper panel) and RNA-seq (lower panel) for exhausted CD8+ T cells with no treatment (Ctrl) or treated with IGF-1 or IGF-1 and IGFBP-1. b, Gene Ontology (GO) enrichment for pathways upregulated (red) and downregulated (blue) according to co-upregulated or co-downregulated genes illustrated in Fig. 4a. The gradient of colour indicates the significance of enrichment, while the size of the bubbles corresponds to the number of differentially expressed genes. c, ChromVAR motif accessibility heatmap for each ATAC–seq sample. Selected transcription factors are indicated on the right. d,e, Relative mRNA levels of terminal exhausted-associated genes (d) and progenitor exhausted-associated genes (e) in exhausted CD8+ T cells with no treatment (Ctrl) or treated with IGF-1 or IGF-1 and IGFBP-1 (n = 3 biological replicates). The experiments were independently repeated two (d,e) times. Data are presented as mean ± SEM. P values were calculated by hypergeometric test, with FDR correction (b) and one-way ANOVA with Tukey’s multiple comparisons (d,e).
Extended Data Fig. 7 Galactose enhances hepatocyte-derived IGFBP-1 production.
a, Igfbp1 mRNA in various tissues from WT mice fed a high-galactose or normal diet (n = 3 mice). b, Igfbp1 mRNA (left, n = 5 mice) and Igf1 mRNA (right, n = 6 mice) in liver tissue from WT mice fed a high-galactose diet. c, Immunoblotting analysis of IGFBP-1 protein expression (n = 6 mice) in liver tissue from WT mice fed a high-galactose diet, with band intensities normalized to β-actin. d, Igfbp1 mRNA (left) and Igf1 mRNA (right) in primary hepatocytes cultured with or without 10 mM galactose treatment for 24 h (n = 3 biological replicates). e, Heatmap of central carbon metabolism-related metabolites in primary hepatocytes treated with or without 10 mM galactose (n = 5 biological replicates). Hepatocytes were undergone a starvation for 30 min and treated with galactose for 60 min, followed by collection and analysis. f, Heatmap of mTORC1 signaling pathway-related genes in the liver tissue from WT mice fed a high-galactose diet (n = 3 mice). Liver samples were collected at day 14 after galactose treatment. g, Representative immunoblot analysis of the indicated proteins in the liver tissue from WT mice fed a high-galactose diet. Liver samples were collected at day 14 after galactose treatment. h,i, Immunoblot analysis of the indicated proteins (h) and Igfbp1 mRNA (i, n = 6 mice) in the liver tissue from WT mice fed a high-fructose diet. Liver samples were collected at day 14 after fructose treatment. j, Serum IGFBP-1 from C57BL/6 WT mice fed a high-fructose diet (n = 6 mice) at day 14 after fructose treatment. k,l, Immunoblot analysis of the indicated proteins (k) and Igfbp1 mRNA (l, n = 7 mice) in the liver tissue from WT mice fed a high-glucose diet. Liver samples were collected at day 14 after glucose treatment. m, Serum IGFBP-1 from C57BL/6 WT mice fed a high-glucose diet (n = 5 mice) at day 14 after glucose treatment. The experiments were independently repeated two (g–m) or three (a–d) times. Data are presented as mean ± SEM. P values were calculated by two-tailed Student’s t-test (a–d,i,j,l,m). ns, not statistically significant.
Extended Data Fig. 8 Galactose triggers mTORC1 inhibition to promote Foxo1-induced IGFBP-1 expression in hepatocytes.
a,b, Representative immunoblot analysis of phosphorylated S6 (p-S6) and phosphorylated S6K1 (p-S6K1) in the liver tissue from tamoxifen-treated Rptor+/+Alb-CreER and Rptorfl/flAlb-CreER mice (a) and Pten+/+Alb-CreER, Ptenfl/flAlb-CreER mice (b) fed on a high-galactose (Gal) diet or normal (Ctrl) diet. c,d, Representative immunoblot analysis of phosphorylated Foxo1 (p-Foxo1) and total Foxo1 in liver tissue from tamoxifen-treated Rptor+/+Alb-CreER, Rptorfl/flAlb-CreER mice (c) and Pten+/+Alb-CreER, Ptenfl/flAlb-CreER mice (d) fed on a high-galactose (Gal) diet or normal control (Ctrl) diet. Liver samples in a–d were collected at day 14 after galactose treatment. e,f, Igfbp1 mRNA level (e, n = 3 biological replicates per group) and protein level in supernatant (f, n = 6 biological replicates per group) from primary hepatocytes treated with DMSO or 1 μM AS184256 for 24 h supplied with 10 mM galactose or not. g, Schematic diagram of the Igfbp1 promoter constructed using a luciferase reporter vector. “WT” and “Mut” represent the wild-type and mutant versions of the Foxo1-binding site, respectively. Luc denotes luciferase. h, Luciferase assay of HEK293T cells transfected with a Foxo1 overexpression plasmid and a luciferase reporter plasmid containing the IGFBP-1 promoter region (n = 4 biological replicates per group), as constructed in (g). “NC” refers to the negative control. The experiments were independently repeated two times. Data are presented as mean ± SEM (e,f,h). P values were calculated by one-way ANOVA with Tukey’s multiple comparisons (e,f) and two-tailed Student’s t-test (h). ns, not statistically significant.
Supplementary information
Supplementary Table 1 (download XLSX )
Primer sequences used in this paper.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Unprocessed blots (download PDF )
Unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Du, X., Li, W., Li, G. et al. Diet-derived galactose reprograms hepatocytes to prevent T cell exhaustion and elicit antitumour immunity. Nat Cell Biol 27, 1357–1366 (2025). https://doi.org/10.1038/s41556-025-01716-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01716-8