Abstract
Tumour-associated macrophages (TAMs) play a pivotal role in shaping the immune microenvironment of hepatocellular carcinoma (HCC), influencing tumour progression and immunotherapy response. WDR4, a tRNA-binding cofactor of the N7-methylguanosine (m7G) methyltransferase complex, remains poorly understood in its independent functions. Here we show that WDR4 is significantly upregulated in HCC-associated TAMs and correlates with poor prognosis. Loss of WDR4 in monocyte-derived macrophages, but not in resident Kupffer cells, reprogrammes TAMs towards an antitumoral phenotype and suppresses HCC progression. Mechanistically, cytoplasmic WDR4 acts independently of m7G modification by directly interacting with eIF4E2 to enhance eIF4E-mediated selective translation of ABCA1, thereby promoting membrane cholesterol efflux and maintaining pro-tumoral polarization. Targeted silencing of WDR4 in TAMs using a CpG-small interfering RNA delivery system enhances antitumour immunity, inhibits tumour progression and improves the efficacy of anti-PD-1 therapy. Our findings identify WDR4 as a key regulator of TAM polarization and a promising therapeutic target to enhance immunotherapeutic outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Single-cell RNA-sequencing data supporting the findings of this study have been deposited in the NCBI Gene Expression Omnibus under the accession number GSE301487. Previously published data from GSE162698 (M2 and MDM dataset) were also re-analysed in this study. Ribo-seq and bulk RNA-sequencing data have been deposited in the NCBI Gene Expression Omnibus under the accession number GSE302134. Proteomics data (LC–MS/MS) have been deposited in the ProteomeXchange Consortium via the PRIDE repository under the accession number PXD058385. Untargeted metabolomics data are available at MetaboLights under the accession number MTBLS12764. Source data are provided with this paper. Additional information relevant to data reanalysis is available from the corresponding author on reasonable request.
Code availability
No custom code was developed or reported in this study.
References
Rumgay, H. et al. Global burden of primary liver cancer in 2020 and predictions to 2040. J. Hepatol. 77, 1598–1606 (2022).
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48 (2023).
Llovet, J. M. et al. Immunotherapies for hepatocellular carcinoma. Nat. Rev. Clin. Oncol. 19, 151–172 (2022).
Cheng, K. et al. Tumor-associated macrophages in liver cancer: from mechanisms to therapy. Cancer Commun. 42, 1112–1140 (2022).
Yeung, O. W. et al. Alternatively activated (M2) macrophages promote tumour growth and invasiveness in hepatocellular carcinoma. J. Hepatol. 62, 607–616 (2015).
Chen, F. et al. Extracellular vesicle-packaged HIF-1α-stabilizing lncRNA from tumour-associated macrophages regulates aerobic glycolysis of breast cancer cells. Nat. Cell Biol. 21, 498–510 (2019).
Liu, G. et al. The miR-92a-2-5p in exosomes from macrophages increases liver cancer cells invasion via altering the AR/PHLPP/p-AKT/β-catenin signaling. Cell Death Differ. 27, 3258–3272 (2020).
Song, G. et al. Global immune characterization of HBV/HCV-related hepatocellular carcinoma identifies macrophage and T-cell subsets associated with disease progression. Cell Discov. 6, 90 (2020).
Cassetta, L. & Pollard, J. W. Targeting macrophages: therapeutic approaches in cancer. Nat. Rev. Drug Discov. 17, 887–904 (2018).
Mantovani, A., Allavena, P., Marchesi, F. & Garlanda, C. Macrophages as tools and targets in cancer therapy. Nat. Rev. Drug Discov. 21, 799–820 (2022).
Zhai, K. et al. Pharmacological inhibition of BACE1 suppresses glioblastoma growth by stimulating macrophage phagocytosis of tumor cells. Nat. Cancer 2, 1136–1151 (2021).
Cassetta, L. & Pollard, J. W. A timeline of tumour-associated macrophage biology. Nat. Rev. Cancer 23, 238–257 (2023).
Lin, S. et al. Mettl1/Wdr4-mediated m7G tRNA methylome is required for normal mRNA translation and embryonic stem cell self-renewal and differentiation. Mol. Cell 71, 244–255 (2018).
Li, J. et al. Structural basis of regulated m7G tRNA modification by METTL1–WDR4. Nature 613, 391–397 (2023).
Katsara, O. & Schneider, R. J. m7G tRNA modification reveals new secrets in the translational regulation of cancer development. Mol. Cell 81, 3243–3245 (2021).
Xia, P. et al. MYC-targeted WDR4 promotes proliferation, metastasis, and sorafenib resistance by inducing CCNB1 translation in hepatocellular carcinoma. Cell Death Dis. 12, 691 (2021).
Dai, Z. et al. N7-methylguanosine tRNA modification enhances oncogenic mRNA translation and promotes intrahepatic cholangiocarcinoma progression. Mol. Cell 81, 3339–3355 (2021).
Ma, J. et al. METTL1/WDR4-mediated m7G tRNA modifications and m7G codon usage promote mRNA translation and lung cancer progression. Mol. Ther. 29, 3422–3435 (2021).
Han, H. et al. N7-methylguanosine tRNA modification promotes esophageal squamous cell carcinoma tumorigenesis via the RPTOR/ULK1/autophagy axis. Nat. Commun. 13, 1478 (2022).
Chen, J. et al. Aberrant translation regulated by METTL1/WDR4-mediated tRNA N7-methylguanosine modification drives head and neck squamous cell carcinoma progression. Cancer Commun.42, 223–244 (2022).
Orellana, E. A. et al. METTL1-mediated m7G modification of Arg-TCT tRNA drives oncogenic transformation. Mol. Cell 81, 3323–3338 (2021).
Huang, M. et al. METTL1-mediated m7G tRNA modification promotes lenvatinib resistance in hepatocellular carcinoma. Cancer Res. 83, 89–102 (2023).
Ma, S. et al. YTHDF2 orchestrates tumor-associated macrophage reprogramming and controls antitumor immunity through CD8+ T cells. Nat. Immunol. 24, 255–266 (2023).
Sakai, M. et al. Liver-derived signals sequentially reprogram myeloid enhancers to initiate and maintain Kupffer cell identity. Immunity 51, 655–670 (2019).
Shao, N. et al. Targeting lipid metabolism of macrophages: a new strategy for tumor therapy. J. Adv. Res. 68, 99–114 (2024).
Goossens, P. et al. Membrane cholesterol efflux drives tumor-associated macrophage reprogramming and tumor progression. Cell Metab. 29, 1376–1389 (2019).
Sag, D., Cekic, C., Wu, R., Linden, J. & Hedrick, C. C. The cholesterol transporter ABCG1 links cholesterol homeostasis and tumour immunity. Nat. Commun. 6, 6354 (2015).
Zhao, Y., Van Berkel, T. J. & Van Eck, M. Relative roles of various efflux pathways in net cholesterol efflux from macrophage foam cells in atherosclerotic lesions. Curr. Opin. Lipido. 21, 441–453 (2010).
Rosettani, P., Knapp, S., Vismara, M. G., Rusconi, L. & Cameron, A. D. Structures of the human eIF4E homologous protein, h4EHP, in its m7GTP-bound and unliganded forms. J. Mol. Biol. 368, 691–705 (2007).
Wiebe, S. et al. The eIF4E homolog 4EHP (eIF4E2) regulates hippocampal long-term depression and impacts social behavior. Mol. Autism 11, 92 (2020).
Wang, F. et al. METTL16 promotes translation and lung tumorigenesis by sequestering cytoplasmic eIF4E2. Cell Rep. 42, 112150 (2023).
Pickering, B. M. & Willis, A. E. The implications of structured 5′ untranslated regions on translation and disease. Semin. Cell Dev. Biol. 16, 39–47 (2005).
Forester, C. M. et al. Regulation of eIF4E guides a unique translational program to control erythroid maturation. Sci. Adv. 8, eadd3942 (2022).
Feoktistova, K., Tuvshintogs, E., Do, A. & Fraser, C. S. Human eIF4E promotes mRNA restructuring by stimulating eIF4A helicase activity. Proc. Natl Acad. Sci. USA 110, 13339–13344 (2013).
Roberts, T. C., Langer, R. & Wood, M. J. A. Advances in oligonucleotide drug delivery. Nat. Rev. Drug Discov. 19, 673–694 (2020).
Kortylewski, M. et al. In vivo delivery of siRNA to immune cells by conjugation to a TLR9 agonist enhances antitumor immune responses. Nat. Biotechnol. 27, 925–932 (2009).
Wen, J. et al. Targeting squalene epoxidase restores anti-PD-1 efficacy in metabolic dysfunction-associated steatohepatitis-induced hepatocellular carcinoma. Gut 73, 2023–2036 (2024).
Zhu, S. et al. Combination strategies to maximize the benefits of cancer immunotherapy. J. Hematol. Oncol. 14, 156 (2021).
Murciano-Goroff, Y. R., Warner, A. B. & Wolchok, J. D. The future of cancer immunotherapy: microenvironment-targeting combinations. Cell Res. 30, 507–519 (2020).
Schoenfeld, A. J. & Hellmann, M. D. Acquired resistance to immune checkpoint inhibitors. Cancer Cell 37, 443–455 (2020).
Xie, Q., Zeng, Y., Zhang, X. & Yu, F. The significance of lipid metabolism reprogramming of tumor-associated macrophages in hepatocellular carcinoma. Cancer Immunol. Immunother. 73, 171 (2024).
Sharma, A. et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183, 377–394 (2020).
Franklin, R. A. & Li, M. O. Ontogeny of tumour-associated macrophages and its implication in cancer regulation. Trends Cancer 2, 20–34 (2016).
Agirre-Lizaso, A. et al. Targeting the heterogeneous tumour-associated macrophages in hepatocellular carcinoma. Cancers 15, 4977 (2023).
Guillot, A. & Tacke, F. Liver macrophages: old dogmas and new insights. Hepatol. Commun. 3, 730–743 (2019).
El-Kenawi, A. et al. Macrophage-derived cholesterol contributes to therapeutic resistance in prostate cancer. Cancer Res. 81, 5477–5490 (2021).
Hoppstadter, J. et al. Dysregulation of cholesterol homeostasis in human lung cancer tissue and tumour-associated macrophages. eBioMedicine 72, 103578 (2021).
Esobi, I., Olanrewaju, O., Echesabal-Chen, J. & Stamatikos, A. Utilizing the LoxP–Stop–LoxP system to control transgenic ABC-transporter expression in vitro. Biomolecules 12, 679 (2022).
Kloosterman, D. J. et al. Macrophage-mediated myelin recycling fuels brain cancer malignancy. Cell 187, 5336–5356 (2024).
Ho, C. et al. AKT (v-akt murine thymoma viral oncogene homolog 1) and N-Ras (neuroblastoma ras viral oncogene homolog) coactivation in the mouse liver promotes rapid carcinogenesis by way of mTOR (mammalian target of rapamycin complex 1), FOXM1 (forkhead box M1)/SKP2, and c-Myc pathways. Hepatology 55, 833–845 (2012).
Heideveld, E. et al. Methods for macrophage differentiation and in vitro generation of human tumor associated-like macrophages. Methods Enzymol. 632, 113–131 (2020).
Cassetta, L. et al. Human tumor-associated macrophage and monocyte transcriptional landscapes reveal cancer-specific reprogramming, biomarkers, and therapeutic targets. Cancer Cell 35, 588–602 (2019).
Xia, P. et al. METTL5 stabilizes c-Myc by facilitating USP5 translation to reprogram glucose metabolism and promote hepatocellular carcinoma progression. Cancer Commun. 43, 338–364 (2023).
Schmidt, E. K., Clavarino, G., Ceppi, M. & Pierre, P. SUnSET, a nonradioactive method to monitor protein synthesis. Nat. Methods 6, 275–277 (2009).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Charoentong, P. et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 18, 248–262 (2017).
Su, R. et al. METTL16 exerts an m(6)A-independent function to facilitate translation and tumorigenesis. Nat. Cell Biol. 24, 205–216 (2022).
Acknowledgements
This work was funded by research funds from the Natural Science Foundation of China (grant numbers 82403904 to P.X. and 82300675 to X.C.), the Key Research and Development Program of Hubei Province (grant number 2021BCA114 to Y.Y.), the Cancer Research and Translational Platform Project of Zhongnan Hospital of Wuhan University (grant number ZLYNXM202004 to Y.Y.) and the medical Sci-Tech innovation platform of Zhongnan Hospital of Wuhan University (grant number PTXM2023008 to Y.Y.). P.X. was supported by the Chicago Tumour Institute, endowments from the Ludwig Cancer Research Foundation, Barry and Kimberly Fields. The authors thank the Center for Research Informatics of the University of Chicago for the use of the Gardner High-Performance Computing cluster, and the Cancer Center Support Grant (P30 CA014599) for support of the Genomics Core Facility.
Author information
Authors and Affiliations
Contributions
J.L., P.X. and Y.Y. conceived and designed the project. J.L., Y.L., P.X., T.M., X.G., C.Q. and X.C. performed the experiments. J.L., M.Z. and C.Q. conducted the database searches and analysed the data. Y.L., H.W. and P.X. were responsible for data statistics. J.L. and P.X. wrote the paper. J.L., P.X., W.H., C.Q., Y.L., X.C. and Y.Y. participated in the revision of the draft. P.X., X.C. and Y.Y. contributed with grant support. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Huilin Huang, Brian Ruffell and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
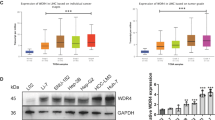
Extended Data Fig. 1 Gating strategies for TAM isolation and validation of WDR4 upregulation in vivo and in vitro.
a, Immune cell type fractions in BRCA, CRC, KIRC, LIHC, LUAD, OV, and STAD samples analysed using CIBERSORT with the LM22 signature in the The Cancer Immunome Atlas (TCIA) database. Absolute scores are shown; TAM populations are highlighted in red. b, Schematic of the flow cytometry gating strategy for the isolation and identification of human CD14+CD11b+CD68+ macrophages. Zombie Aqua was used to exclude dead cells. c, Schematic of the flow cytometry strategy for analysing the expression of WDR4, CD45.1, and CD45.2 in mouse CD45+CD11b+F4/80+ macrophages. d, Workflow illustrating the isolation of mouse bone marrow monocytes and their differentiation into BMDMs using 20 ng/mL macrophage colony-stimulating factor (M-CSF). Created with BioRender.com. e, qPCR analysis of pro-tumour macrophage markers (Cd163, Mrc1, and Arg1) in TAMs and BMDMs (n = 3 biological replicates). f, ELISA measurement of IL-10 levels in culture supernatants of TAMs and BMDMs (n = 3 biological replicates). g,h, qPCR (g) and immunoblotting (h) analysis of WDR4 expression in BMDMs and F4/80+ TAMs (n = 5 biological replicates). ns, not significant. i, Representative mIHC images showing WDR4 expression in macrophages from orthotopic Hepa1-6 tumours and adjacent tissues. j,k, THP-1 (j) and iBMDMs (k) cells were stimulated with tumour-conditioned medium to induce a TAM-like phenotype in vitro, followed by WDR4 expression analysis by flow cytometry (n = 3 biological replicates). All data are presented as mean ± s.d. Statistical significance was determined by two-tailed paired (g) or two-tailed unpaired (e,f,j,k) Student’s t-test.
Extended Data Fig. 2 WDR4 knockout in TAMs suppresses HCC progression.
a, Immunoblotting analysis of WDR4 protein levels in THP-1- and iBMDM-derived TAMs with or without WDR4 knockout (KO), with band intensity quantification shown at right (n = 3 biological replicates). β-Actin served as a loading control. b, Schematic illustrating the co-culture of Wdr4-KO iBMDM-derived TAMs with Hepa1-6 cells using 0.4 μm Transwell inserts. c, CCK-8 assay assessing the proliferation of untreated (UN) Hepa1-6 cells or those co-cultured with Wdr4-KO TAMs (n = 3 biological replicates). d, Representative images from colony formation assays showing the effect of Wdr4-KO TAMs on Hepa1-6 cell proliferation (left), with quantification of colony numbers (right; n = 3 biological replicates). e, EdU assay of Hepa1-6 cells cultured alone or co-cultured with Wdr4-KO TAMs (n = 3 biological replicates). Scale bar, 50 μm. f, Schematic illustrating the co-culture of Huh-7 cells with THP-1-derived TAMs. g–i, CCK-8 assay (g), colony formation assay (h), and EdU assay (i) evaluating the impact of WDR4-KO TAMs on the proliferation of Huh-7 cells (n = 3 biological replicates). Scale bar for EdU assay, 50 μm. j,k, qPCR (j) and immunoblotting (k) analyses of WDR4 expression in CD11b+ myeloid cells isolated from the spleens of WDR4f/f and WDR4cKO mice (n = 5 biological replicates). l,m, qPCR (l) and immunoblotting (m) of WDR4 expression in BMDMs isolated from Wdr4f/f and Wdr4cKO mice (n = 5 biological replicates). n, Flow cytometry analysis of tumour-infiltrating macrophage percentages in tumour-bearing Wdr4f/f and Wdr4cKO mice treated with clodronate or PBS liposomes (n = 5 biological replicates). o, Schematic illustrating the subcutaneous co-injection of BMDMs from Wdr4f/f and Wdr4cKO mice together with Hepa1-6 cells into wild-type C57BL/6 mice. p, Tumour growth curves and final tumour weights (n = 5 biological replicates). All data are presented as mean ± s.d. Statistical significance was determined by two-tailed unpaired Student’s t-test (j,l; quantifications in k,m, and p), one-way ANOVA (n), or two-way ANOVA (tumour growth curves in p) with Tukey’s post-hoc test. Panels b,f,o created with BioRender.com.
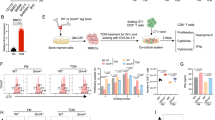
Extended Data Fig. 3 Immune cell profiling highlights TAM subset heterogeneity and WDR4-associated functional states.
a, UMAP plot showing the clustering of CD45+ immune cells in orthotopic Hepa1-6 tumours from Wdr4f/f and Wdr4cKO mice. b, Dot plots showing the expression of canonical marker genes used to identify immune cell types. c,d, UMAP plots showing the major immune cell populations (c) and the distribution of immune cell subsets (d) in orthotopic Hepa1-6 tumours from Wdr4f/f and Wdr4cKO mice. e, Heatmap displaying differentially expressed genes (rows) across five TAM subsets (columns). Representative marker genes are indicated. f, scRNA-seq analysis showing the distribution of Wdr4-positive cells among TAM subsets. g, Flow cytometry analysis of WDR4 expression in TAM subsets from orthotopic Hepa1-6 tumours (n = 5 biological replicates). h, GO enrichment analysis of differentially expressed genes among TAM subsets. i, Violin plots showing the expression levels of Nos2 and Arg1 across TAM subsets, as determined by scRNA-seq analysis. Data are presented as mean ± s.d. Statistical significance was calculated by Hypergeometric test (h).
Extended Data Fig. 4 WDR4 modulates macrophage polarization in HCC models.
a,b, Flow cytometry analysis of CD11b+F4/80+iNOS+ antitumoral macrophages and CD11b+F4/80+Arg1+ pro-tumoral macrophages in hydrodynamically induced primary liver tumours (a) and subcutaneous Hepa1-6 tumours (b) from Wdr4f/f and Wdr4cKO mice (n = 5 biological replicates). c,d, qPCR analysis of antitumoral markers (Nos2, Il1b, Il12a, Csf3r, Cxcr2, Tnf, Fpr1, and Cd80) and pro-tumoral markers (Arg1, Apoe, Pf4, Mrc1, Spp1, Ccl9, C1qa, and Cd163) in F4/80+ TAMs sorted from hydrodynamically induced primary liver tumours (c) and subcutaneous Hepa1-6 tumours (d) (n = 3 biological replicates). e, Flow cytometry analysis of iNOS and Arg1 expression in iBMDM-derived and THP-1-derived TAMs with or without WDR4 knockout (KO) (n = 3 biological replicates). All data are presented as mean ± s.d. Statistical significance was determined by two-tailed unpaired Student’s t-test (c,d; quantifications in a,b) or one-way ANOVA with Dunnett’s post-hoc test (quantifications in e).
Extended Data Fig. 5 WDR4 knockout in Kupffer cells has limited impact on HCC progression.
a, Flow cytometry gating strategy used to isolate F4/80+ Tim4⁻ monocyte-derived macrophages (MDMs) and F4/80+ Tim4+ Kupffer cells from orthotopic Hepa1-6 tumours. b, Flow cytometry analysis of WDR4 expression in MDMs and Kupffer cells (n = 5 mice per group). c, Representative images and quantification of tumour weights in orthotopic Hepa1-6 tumours from Wdr4f/f and Kupffer cell-specific Wdr4 knockout (Wdr4f/fClec4fcre) mice (n = 5 mice per group). d, Representative images and quantification of tumour burden in hydrodynamically induced primary liver tumours from Wdr4f/f and Wdr4f/fClec4fcre mice (n = 5 mice per group). e,f, Flow cytometry analysis of CD11b+F4/80+iNOS+ antitumoral and CD11b+F4/80+Arg1+ pro-tumoral macrophages in orthotopic Hepa1-6 tumours (e) and hydrodynamically induced primary liver tumours (f) from Wdr4f/f and Wdr4f/fClec4fcre mice (n = 5 mice per group). All data are presented as mean ± s.d. Statistical significance was determined by two-tailed unpaired Student’s t-test (quantifications in b–d,e,f).
Extended Data Fig. 6 Loss of WDR4 reduces translational efficiency and alters cholesterol efflux in TAMs.
a, Representative polysome profiles of WT and WDR4-KO THP-1-derived TAMs obtained by sucrose density gradient fractionation. b, Representative immunoblotting image from SUnSET assays quantifying nascent (puromycin-labelled) peptides in THP-1-derived TAMs with or without WDR4 KO. c, Cell numbers of wild-type (WT) and WDR4-KO THP-1-derived TAMs at 0 h and 24 h (n = 3 biological replicates). d, Scatter plot showing the correlation of average read counts from input RNA between Wdr4-KO and control TAMs, based on RNA sequencing (RNA-seq). e, Venn diagram showing overlap of transcripts with reduced translational efficiency (TE) upon Wdr4 deletion across experimental replicates. f, KEGG pathway enrichment analysis of genes with decreased TE following Wdr4 KO. g, Bubble plot of KEGG enrichment analysis for differentially abundant metabolites in TAMs from Wdr4f/f and Wdr4cKO mice bearing orthotopic Hepa1-6 tumours. Bubble size indicates the number of metabolites associated with each pathway. h, Total cholesterol levels in BMDMs from tumour-bearing Wdr4f/f and Wdr4cKO mice. i, Membrane cholesterol in TAMs from tumour-bearing Wdr4f/f and Wdr4cKO mice as detected by cholera toxin B (CTB) staining (n = 3 biological replicates). j, qPCR analysis of key cholesterol efflux-related genes in WDR4-KO TAMs derived from THP-1 or iBMDMs (n = 3 biological replicates). k, qPCR analysis of polysomal mRNA distribution showing the localization of Abca1 and β-actin transcripts in TAMs from tumour-bearing Wdr4f/f and Wdr4cKO mice (n = 3 biological replicates). Ribo, Ribosome; Mono, Monosomes; low MW poly, low molecular weight polysomes; high MW poly, high molecular weight polysomes. For a,b, data shown represent 3 independent experiments with similar results. All data are presented as mean ± s.d. Statistical significance was determined by two-tailed Pearson correlation analysis (d), two-tailed unpaired Student’s t-test (h–j) or two-way ANOVA with Tukey’s post-hoc test (c).
Extended Data Fig. 7 ABCA1 mediates the functional impact of WDR4 on cholesterol efflux and tumour-promoting activity in TAMs.
a, Total cholesterol levels in WT and Abca1-KO iBMDM-derived TAMs with or without Wdr4 KO (n = 3 biological replicates), and in THP-1-derived TAMs with knockout of ABCG1, APOA1, APOA2, or APOA4, with or without Wdr4 KO (n = 3). b, Membrane cholesterol levels in TAMs derived from THP-1 cells or iBMDMs as detected by cholera toxin B (CTB) staining (n = 3 biological replicates). CTCF, corrected total cell fluorescence. c, NBD-cholesterol efflux percentage in TAMs derived from THP-1 cells or iBMDMs (n = 3 biological replicates), quantified by fluorescence intensity in cells and medium. d, Immunoblotting analysis of ABCA1 expression in ABCA1-overexpression (OE) TAMs (n = 3 biological replicates). e,f, Flow cytometry analysis of iNOS (e) and Arg1 (f) expression in WDR4-KO TAMs transfected with vector or ABCA1-OE plasmid (n = 3 biological replicates). g, Schematic diagram of the co-culture of Huh-7 cells with THP-1-derived TAMs. Created with BioRender.com. h–j, CCK8 assay (h), colony formation assay (i), and EdU assay (j) were performed to assess whether ABCA1 OE could rescue the WDR4-KO TAM-induced inhibition of Huh-7 cell proliferation (n = 3 biological replicates). k, Tumour growth curves, and final tumour weights in WT C57BL/6 mice subcutaneously co-injected with Hepa1-6 cells and Wdr4-KO iBMDMs transfected with ABCA1-OE or vector (n = 5 mice per group). All values are presented as mean ± s.d. Statistical significance was calculated using two-tailed unpaired Student’s t-test (quantifications in d), two-way ANOVA (a–c,h, tumour growth curves in k), or one-way ANOVA (i,j; quantifications in e,f,k) with Tukey’s post-hoc test.
Extended Data Fig. 8 Cytoplasmic WDR4 maintains stable localization under stress in TAMs and promotes tumour progression via METTL1-independent regulation of ABCA1 translation.
a, Immunoblotting analysis showing the distribution of WDR4 in the cytoplasmic (C) and nuclear (N) fractions of THP-1-derived TAMs treated with 1% O2, low glucose (1 g/L), LPS (100 ng/mL), or IFN-γ (20 ng/mL). Data shown represent three independent experiments. b, LC–MS/MS analysis of total RNA m7G methylation levels in WDR4 KO TAMs derived from THP-1 (n = 3 biological replicates). c, Dot blot assay showing total RNA m7G methylation levels in THP-1-derived TAMs transfected with sgWDR4-1 plasmid (n = 3 biological replicates), with quantification shown on the right. d, Immunoblotting analysis of METTL1 expression in THP-1-derived TAMs transfected with siMETTL1 (n = 3 biological replicates). e–h, WDR4-KO TAMs transfected with WDR4mut, WDR4NLS, or Vector were co-cultured with Hepa1-6 or Huh-7 cells (e). CCK8 assay (f), colony formation assay (g), and EdU assay (h) were used to assess the proliferation of Hepa1-6 and Huh-7 cells (n = 3 biological replicates). i,j, Tumour growth curves (i) and final tumour weight (j) in WT C57BL/6 mice subcutaneously co-injected with Hepa1-6 cells and Wdr4-KO iBMDMs transfected with Vector, Wdr4, or Wdr4NLS (n = 5 mice per group). k, Tumour weights of subcutaneous tumours in mice co-injected with Hepa1-6 cells and Wdr4-KO iBMDMs transfected with Vector, Wdr4, or Wdr4mut (n = 5 mice per group). l, Total cholesterol levels and membrane cholesterol in Wdr4-KO TAMs transfected with Vector, Wdr4NLS, or Wdr4mut (n = 3 biological replicates). Membrane cholesterol was detected by CTB staining. CTCF, corrected total cell fluorescence. m, qPCR analysis of polysomal mRNA distribution showing the localization of Abca1 and β-actin mRNAs across sucrose gradient fractions in Wdr4-KO TAMs transfected with Vector, Wdr4NLS, or Wdr4mut (n = 3 biological replicates). Ribo, ribosome; Mono, monosomes; low MW poly, low molecular weight polysomes; high MW poly, high molecular weight polysomes. n, Immunoblotting analysis of ABCA1 expression in THP-1-derived TAMs transfected with siMETTL1 (n = 3 biological replicates). o, qPCR analysis of polysomal mRNA distribution showing the effect of METTL1 knockdown on ABCA1 translation (n = 3 biological replicates). p, Immunoblotting analysis of ABCA1 protein levels in THP-1-derived TAMs treated with a METTL1-WDR4 inhibitor (n = 3 biological replicates). q, qPCR analysis of polysomal mRNA distribution assessing the impact of METTL1-WDR4 inhibitor on ABCA1 translation (n = 3 biological replicates). All values are presented as mean ± s.d. Statistical significance was calculated using two-tailed unpaired Student’s t-test (quantifications in c,d,n,p) or one-way ANOVA (b,g,h,j,k) or two-way ANOVA (f,i) with Tukey’s post-hoc test. Panel e created with BioRender.com.
Extended Data Fig. 9 WDR4 interacts with eIF4E2 to regulate cholesterol efflux gene translation and drive HCC progression.
a, GO enrichment analysis of the cellular component (CC) category for WDR4-interacting proteins in TAMs derived from THP-1 cells. b, Translation initiation-related genes identified by WDR4 immunoprecipitation and mass spectrometry. c,d, Co-immunoprecipitation (Co-IP) (c) and reciprocal Co-IP (d) assays examining the interaction between WDR4 and eIF4E2 in THP-1-derived TAMs, with or without RNase treatment. Data shown represent 3 independent experiments. e, Coomassie blue-stained gel showing purified GST-tagged WDR4 fusion proteins (GST–WDR4). f, Proximity ligation assay (PLA) showing the in situ interaction of WDR4 with eIF4E2 and eIF4E in TAMs isolated from tumour-bearing WT C57BL/6 mice (n = 8 biological replicates). Scale bar, 10 μm. g, Immunoblotting analysis of eIF4E2 and ABCA1 protein expression in eIF4E2-OE TAMs derived from THP-1 (n = 3 biological replicates). h, GC content, minimum free energy (MFE), and length of the 5′ UTRs of translationally downregulated genes in WT and WDR4- KO TAMs. i, Luciferase reporter assay measuring 5′ UTR-mediated translation of target mRNAs (ABCA1, ABCG1, APOA1, APOA2, and APOA4) and the housekeeping gene β-actin in control and WDR4-KO TAMs derived from THP-1. pGL3 refers to the empty luciferase vector lacking a 5′ UTR insert (n = 3 biological replicates). j, Luciferase reporter assay showing the effect of transversion (TV) mutations in the C-rich motif of the APOA1 5′ UTR (nucleotides 147–155) on reporter activity in WT and WDR4-KO TAMs derived from THP-1 (n = 3 biological replicates). ns, not significant. k, Immunoblotting analysis of eIF4E expression in TAMs with or without eIF4E knockdown (KD) (n = 3 biological replicates). l–o, eIF4E KD and control THP-1-derived TAMs with or without WDR4 KO, were co-cultured with Huh-7 cells (l). CCK8 assay (m), colony formation assay (n), and EdU assay (o) were used to evaluate the proliferation of Huh-7 cells (n = 3 biological replicates). All values are presented as mean ± s.d. Statistical significance was calculated using a two-tailed unpaired Student’s t-test (h,i; quantifications in f,g,k) or two-way ANOVA with Tukey’s post-hoc test (j,m–o). Panel l created with BioRender.com.
Extended Data Fig. 10 Assessment of WDR4 expression and immune cell profiling in tumour models treated with CpG-siRNA.
a, Immunoblotting analysis of WDR4 protein expression in TAMs after intratumoral injection of CpG-siRNAWdr4 or CpG-siRNANC (n = 5 mice per group). b, Flow cytometry analysis of WDR4 expression in major cell populations isolated from subcutaneous Hepa1-6 tumours of mice treated with CpG-siRNAWdr4 or CpG-siRNANC (n = 5 mice per group). c, Flow cytometry gating strategy for CD62L−CD44+ effector memory T cells, CD3+CD4+ and CD3+CD8+ T cells, CD11c+MHCII+ dendritic cells (DCs), and NKp46+ natural killer (NK) cells. All values are presented as mean ± s.d. Statistical significance was calculated using two-tailed unpaired Student’s t-test (b; quantifications in a).
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1.
Supplementary Table 1 (download XLSX )
Find Individual Motif Occurrences-based analysis of C-rich motif enrichment in 5′ UTRs of cholesterol efflux-related genes.
Supplementary Table 2 (download XLSX )
The clinicopathological characteristics of the 40 patients with HCC.
Supplementary Table 3 (download XLSX )
Reagents or resources.
Supplementary Table 4 (download XLSX )
Summary of oligonucleotide sequences and plasmids used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Unprocessed western blots and gels (download PDF )
Unprocessed western blots and gels for Figs. 1 and 3–6 and Extended Data Figs. 1, 2 and 6–10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Qu, C., Liu, Y. et al. WDR4 drives tumour-associated macrophage reprogramming and tumour progression via selective translation and membrane cholesterol efflux. Nat Cell Biol 27, 2152–2166 (2025). https://doi.org/10.1038/s41556-025-01815-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01815-6
This article is cited by
-
Engineered extracellular vesicles displaying bi-specific T-cell engagers for targeted therapy of B-cell malignancies
Experimental Hematology & Oncology (2026)