Abstract
Although peritumoural visceral adipose tissue (tVAT) is anatomically close to tumours such as colorectal cancer, the immune landscape of this tissue and its functional contribution to tumour immunity remain poorly defined. Here, we performed single-cell RNA analysis on the tVAT from patients with colorectal cancer to map its immune landscape and observed that tVAT exhibited a highly immune-infiltrated microenvironment enriched with lymphocytes, especially tumour-specific CD8⁺ T cells. Mechanistically, tVAT competes with the tumour for these immunocytes by activating the CXCL12–CXCR4 axis to promote tumour immune escape. Moreover, tumour-derived factors induce an adipose–mesenchymal transformation process where the adipose stromal cells trans-differentiated into adipose-derived cancer-associated fibroblasts, which secrete large amounts of CXCL12 in tVAT. Clinically, targeting adipose–tumour interaction substantially enhances diagnostic and therapeutic efficacy of anti-PD-1 therapy. These findings offer an understanding of the dynamic crosstalk between tVAT and tumour immune escape, highlighting the tVAT as a potential target for cancer immunotherapy.
Similar content being viewed by others
Main
The tumour microenvironment (TME) is a complex and dynamic ecosystem1. Currently, there is increasing evidence suggesting that TME plays pivotal roles in tumourigenesis, metastasis and immune evasion, and numerous therapeutic targets within the TME have been developed2. While these studies have mostly focused on tumour-intrinsic mechanisms, research on the peritumoural microenvironment remains limited3. Many solid tumours, particularly those in visceral organs such as colorectal cancer (CRC), have been shown to be closely associated with various types of adipose tissues, including omental and mesenteric4,5, which are increasingly recognized as integral components of the broader TME6. Therefore, it is of great theoretical and clinical significance to study the cellular heterogeneity, spatial organization and tumour-related functional states of peritumoural visceral adipose tissue (tVAT).
Adipose tissue is a highly dynamic, plastic and heterogeneous organ that functions both as an endocrine organ and an energy-storage depot7,8. Structurally, adipose tissue consists of mature adipocytes and a stromal-vascular fraction, which includes fibroblasts, adipose-derived stromal cells, immune cells, vasculature and nerve fibres9. Emerging evidence has shown that adipocytes or adipose tissue may promote tumour progression by supplying cancer cells with metabolites, adipokines and chemokines6,10,11. Additionally, adipose components have been shown to contribute to immune evasion by suppressing cytotoxic immune cell activity and promoting the survival of immunosuppressive cells12,13. Conversely, tumour cells can induce the reprogramming of neighbouring adipocytes into cancer-associated adipocytes to support tumour expansion by supplying essential nutrients14. One study also reported that implanting engineered browning adipocytes into tumour-bearing models could suppress tumour growth15. Thus, elucidating the crosstalk between tumour and tVAT will further expand our understanding of the TME and antitumour immunity.
In recent years, the development of single-cell RNA sequencing (scRNA-seq) technologies has greatly expanded our understanding of how TME heterogeneity contributes to cancer progression. Recently, scRNA-seq analyses have unveiled the characteristics and dynamic evolution of T cells, macrophages and dendritic cells (DCs) in CRC16,17,18,19. Similarly, scRNA-seq has been increasingly applied to investigate adipose tissue in endocrine research. For instance, a recent study provided comprehensive cellular atlases of subcutaneous and visceral white fat at single-cell resolution across a range of body weights8. Additionally, another study identified the subsets and developmental trajectories of adipose-resident immune cell populations that accumulated in obese WAT20. Although CRC is surrounded by abundant adipose tissue, the functional roles of tVAT in tumour immune escape remain largely unknown. Notably, the immune landscape of tVAT has not yet been reported.
In this study, our findings not only reveal the dynamic crosstalk between the tumour and peritumoural adipose tissue (PAT), but also support tVAT as a potential target for improving the diagnostic and therapeutic efficacy of immunotherapy for patients with visceral tumours.
Results
Single-cell RNA profiling unveils tissue-specific immune landscape of tVAT
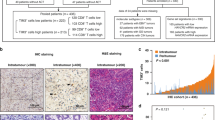
To analyse systematically the characteristics of the peritumoural microenvironment, we collected 36 paired CRC tissue specimens (tumour, tVAT and distal visceral adipose tissue (dVAT)) from 12 patients as well as two tumour samples obtained from the remaining two patients and performed scRNA-seq analysis (Fig. 1a). We also integrated our dataset with a previously published CRC scRNA-seq atlas containing 56 tumours and 12 peritumoural normal tissues21. All CRC patients are in stage T4a, the tumour is close to the mesenteric side and has broken through the serosal layer as shown (Fig. 1a). After rigorous quality control and filtering, we obtained 371,381 high-quality single-cell transcriptomes across four tissue types, comprising 70 tumour, 12 normal, 12 tVAT and 12 dVAT samples (Fig. 1b). To define the major cell populations in the TME, graph-based clustering was performed and 13 distinct cell subsets were identified based on canonical marker expression (Fig. 1b and Extended Data Fig. 1a,b). As expected, we observed that epithelial cells were predominantly found in tumour tissues, whereas plasma cells were enriched in healthy tissues. In contrast, neutrophils, stromal, monocytes and natural killer (NK) cells were more abundant in both tVAT and dVAT (Fig. 1c).
a, Study design illustrating the data analysis (left), and representative images of surgical specimens from patients with CRC (Stage, T4a; right). b, Uniform Manifold Approximation and Projection (UMAP) visualization showing major cell clusters from tumour, normal, tVAT and dVAT samples derived from patients with CRC. The cell types shown NK cells, DC, Mono (monocytes), Mφ (macrophages), Neutro (neutrophils), Mast (mast cells), Endo (endothelial cells) and Epi (epithelial cells). c, Heatmap showing the tissue preferences of 13 cell types by the Ratio of observed to expected (Ro/e) index, illustrating preferential cell type enrichment across different tissue regions. d, UMAP visualization of neighbourhoods (Nhoods) identified by Milo, highlighting the differentially abundant neighbourhoods between tVAT and dVAT. Each Nhood is represented as a node, coloured according to log2 fold change (FC) between tVAT (n = 12) and dVAT (n = 12). Non-differentially abundant neighbourhoods (false discovery rate ≥ 0.1) are displayed in white. Node sizes are proportional to the number of cells in each Nhood, with graph edges representing shared cell quantities between adjacent neighbourhoods. e, Beeswarm plot illustrating the distribution of adjusted log2 FC in abundance of Nhoods between tVAT (n = 12) and dVAT (n = 12) across all cell types. The cell types shown include major immune cell and stromal cell lineages. f, Boxplot comparing the relative abundance of CD4+ T cells, CD8+ T cells, B cells and stromal cells between tVAT (n = 12) and dVAT (n = 12), analysed using a two-sided paired Wilcoxon test. The box hinges denote the first and third quartiles, the median is represented by the centre line and the whiskers encompass the full data range. Individual data points are shown as dots. g, Representative images of hematoxylin and eosin staining (23 samples) and mIHC staining (four samples) for TLSs in dVAT and tVAT sections from patients with CRC. Scale bar, 200 μm. Panel a created in BioRender: Huaiqiang, J. https://biorender.com/vnv4y0b (2026).
To compare the cellular composition between tVAT and dVAT, we conducted a differential abundance analysis using the Milo method22, identifying 17,739 cell neighbourhoods, among which, 3,714 showed marked differences in abundance (Fig. 1d). Notably, immune cells such as B/plasma cells, CD4⁺ T cells and CD8⁺ T cells were significantly enriched in tVAT, whereas stromal cells were more prominent in dVAT (Fig. 1e,f). Next, a Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was conducted and we observed tVAT was enriched in inflammatory signalling pathways, consistent with its increased immune cell content (Extended Data Fig. 1c). Also, multiplex immunohistochemistry (mIHC) was performed, which confirmed a higher presence of scattered T cells and B cells in tVAT (Extended Data Fig. 1d). In addition, we unexpectedly identified the presence of tertiary lymphoid structures (TLSs) within tVAT (Fig. 1g). Based on established classification criteria23,24, the early-stage TLSs (E-TLSs), primary follicle-like TLSs (PFL-TLSs) and secondary follicle-like TLSs (SFL-TLSs) were all detected in the tVAT samples (Extended Data Fig. 1e), which indicates the potential immunoregulatory ability of tVAT. Together, these results demonstrate that tVAT forms a unique tumour-associated microenvironment characterized by extensive immune cell infiltration and immune activity.
Characteristic of lymphocytes in tVAT compared with dVAT
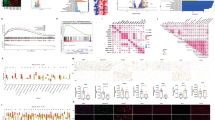
To investigate the transcriptional heterogeneity of lymphocytes, we performed unsupervised clustering on 65,148 CD4⁺ T cells, 63,276 cytotoxic lymphocytes and 62,816 B/plasma cells (Fig. 2a). Based on differentially expressed genes and canonical immune markers (Extended Data Fig. 2a), we identified seven transcriptional states of CD8⁺ T cells including naive-like (TN, C1), central memory (TCM, C2–C3), effector memory (TEM, C4–C5), terminal effector memory (TEMRA, C6), tissue-resident memory (TRM, C7–C8), exhausted (TEX, C9) and stress-response (TSTR, C10) subsets (Fig. 2a). In addition, we identified three types of innate cytotoxic lymphocytes: γδT cells, NK cells and mucosal-associated invariant T (MAIT) cells. For CD4+ T and B/plasma cells, six transcriptional states of CD4⁺ T cells and eight transcriptional clusters of B/plasma cells were defined based on top-ranked markers (Fig. 2a). Tissue distribution analysis revealed preferential infiltration patterns across tissues, whereby LAG3⁺CD8⁺ TEX, ZNF683⁺CD8⁺ TRM, TNFRSF18⁺CD4⁺ Treg and FOXP3⁺CD4⁺ Treg cells were found to be predominantly enriched in tumour tissues, whereas in healthy tissues, IGHG2⁺, IGHV5⁺ and IGLV2⁺ plasma cells, as well as KLRC2⁺CD8⁺ T cells, were more abundant. In both dVAT and tVAT, we observed increased infiltration of LTB⁺CD8⁺ TN, GPR183⁺CD8⁺ TCM, GZMH⁺CD8⁺ TEMRA, TCF7⁺CD4⁺ TN, CD16⁺ NK cells, XCL1⁺ NK cells and IgM⁻ memory B cells (Fig. 2b).
a, UMAP embedding showing subsets of CD4+ T cells, cytotoxic lymphocytes and B cells across all tissue samples from patients with CRC. b, Heatmap displaying the abundance of each lymphocyte subset across various tissues, illustrating their distribution patterns in the TME. c, Boxplot comparing the abundance of different subclusters of CD4+ T cells, cytotoxic lymphocytes and B cells between tVAT (n = 12) and dVAT (n = 12), analysed using the two-sided paired Wilcoxon test. d, Heatmap showing the fraction of TCR clonotype overlap among tissue samples and their respective tumours (left). A boxplot comparing TCR overlap between tVAT (n = 3) and dVAT (n = 3), analysed with a paired two-sided Student’s t-test (right). e, UMAP visualization of 13 tumour-specific CD8+ T cell subsets in dVAT and tVAT from patients with CRC (left). Stack plot showing the relative abundance of these 13 subsets in dVAT and tVAT (right). The data are presented as a box-and-whisker graph (bounds of box show the first to third quartile, bottom and top line show the minimum to maximum and central line shows the median) for c,d.
To further examine the differences between tVAT and dVAT, differential abundance analysis was performed and revealed significant enrichment of lymphocytes in tVAT, including TCF7⁺CD4⁺ TN, CXCL13⁺CD4⁺ TFH, GPR183⁺CD8⁺ TCM, TNFSF9⁺CD8⁺ TEM, GZMK⁺CD8⁺ TEM, mature naive B cells, and both IgM⁺ and IgM⁻ memory B cells (Fig. 2c). Single-cell T cell receptor (scTCR) analysis revealed a higher degree of T cell receptor (TCR) clonotype overlap between tVAT and tumour than between dVAT and tumour (Fig. 2d and Extended Data Fig. 2b,c). In contrast, single-cell B cell receptor (scBCR) analysis showed no significant difference in B cell receptor (BCR) overlap between tVAT and dVAT with tumour (Extended Data Fig. 2d–g). To further investigate the presence of tumour-reactive T cells, we identified tumour-specific CD8⁺ T cells based on the reported approach25,26. Notably, GZMK⁺ TEM cells represented a major subset of the tumour-specific CD8⁺ T cells enriched in tVAT (Fig. 2e). The above results revealed the unique immune landscape of tVAT, suggesting its potential functional contribution to tumour immunity.
tVAT competes with tumour tissue for the infiltration lymphocytes
tVAT is located adjacent to CRC boundary and may act as an initial immunological barrier against tumour progression27,28,29. Based on the observation that tVAT shows increased infiltration of cytotoxic CD8⁺ T cells, we hypothesized that tVAT might possess antitumour potential. To investigate this hypothesis, we established a tumour-associated adipose tissue model by implanting tumour cells adjacent to the inguinal adipose tissue in mice, as previously reported30. One week after tumour implantation, we surgically removed approximately 0.1 g of PAT to simulate tVAT depletion (Fig. 3a). Contrary to our expectations, the removal of this adipose tissue resulted in a substantial reduction in tumour growth and volume (Fig. 3b,c and Extended Data Fig. 3a,b). Also, the depletion of tVAT led to increased infiltration of CD4⁺ and CD8⁺ T cells into the tumour tissue, as well as a higher proportion of tumour antigen-specific CD8⁺ T cells (Fig. 3d,e), suggesting that tVAT removal enhances the antitumour immune response.
a, Experimental design of the PAT immune-competition model using C57BL/6J or BALB/c mice with the removal of either PAT or contralateral inguinal adipose tissue (control). b,c, Tumour weights of MC38-OVA (b) or CT26 (c) tumours at day 16 of the experiment in C57BL/6J or BALB/c mice (n = 5). d,e, Flow cytometry analysis of the infiltration of T cells, CD4+ T cells, CD8+ T cells, and tumour-specific CD8+ T cells in MC38-OVA (d) or CT26 (e) tumours, with and without removal of PAT (n = 5). To identify OVA-specific CD8+ T cells, H-2Kb tetramers were employed in the MC38-OVA tumour model that endogenously expresses ovalbumin. f, Representative images of MC38 tumours (left) and tumour weights (right) in C57BL/6J mice following removal of PAT or control, and treatment with ATS-GNP or saline (n = 5). g, Experimental design of the PAT immune-deficient model (BALB/c-Nude or NSG) with removal of PAT. h,i, Representative images of MC38 tumours (left) and tumour weights (right) at day 16 in C57BL/6 J, BALB/c-Nude (h), or NSG mice (i) (n = 6). j, Schematic representation of the strategy to detect CD45.1+ cells in MC38-OVA tumours, with and without removal of PAT or contralateral inguinal adipose tissue (control), using the CD45.2 receptor mouse model transferred with CD45.1 splenocytes from CD45.1 donor mice. i.v., intravenous. k, Quantitative analysis of various types of CD45.1+ cells in MC38-OVA tumours with and without removal of PAT or control, as determined by flow cytometry (n = 5). Data represent ≥3 independent experiments. Statistical significance was assessed using a two-sided, unpaired Student’s t-test (d,e,k), one-way analysis of variance (ANOVA) with Tukey’s correction for multiple comparisons (b,c,f right), a two-way ANOVA with Tukey’s correction for multiple comparisons (h right) or a two-way ANOVA with Tukey’s correction for multiple comparisons (i right). Graphs display mean ± s.d. (b–f,h i,k). Panels created with BioRender: a, g and j, Huaiqiang, J. https://biorender.com/f9uacgq (2026).
To exclude confounding effects related to altered blood supply caused by surgical manipulation, we employed an adipocytolytic nanomaterial, ATS-GNP, to chemically eliminate PAT31. Consistent with the surgical model, removal of PAT using ATS-GNP also resulted in reduced tumour growth (Fig. 3f and Extended Data Fig. 3c). Also, PAT removal led to a weak tumour suppression effect in immunodeficient BALB/c-Nude and NSG mice (Fig. 3g–i and Extended Data Fig. 3d), suggesting that the observed effect requires functional immune cells. Similar results were also observed using the E0771 cell model (Extended Data Fig. 3e,f). Based on these results, we hypothesized that tVAT might act as a competing ‘reservoir’ for lymphocytes, which limits their infiltration into the tumour. To test this, we performed a competitive study and found that mice in the PAT removal group had significantly higher numbers of CD45.1⁺ cells within the tumour, especially CD45.1⁺CD8⁺ tumour-specific T cells showed an increase trend (Fig. 3j,k and Extended Data Fig. 3g). These findings support the hypothesis that tVAT competes with the tumour for immune cells, and its removal relieves this competition, allowing more immune cells to enter the tumour and exert antitumour effects.
tVAT recruits CXCR4⁺ immune cells through CXCL12 secretion
Here, we hypothesized that differences in the stromal microenvironment between tVAT and tumour tissue contribute to the competition for immune cell recruitment. scRNA-seq and intercellular communication analysis between stromal and immune cells revealed that the CXCL12–CXCR4 signalling axis was more active in tVAT than in tumour and dVAT (Fig. 4a,b). Further analysis showed that stromal cells in tVAT were the primary source of CXCL12, whereas CXCR4 was broadly expressed on immune cells, particularly lymphocytes (Fig. 4c). Moreover, the overall expression of CXCL12 was significantly higher in tVAT than in tumour tissue and dVAT (Fig. 4d), indicating that tVAT may recruit CXCR4⁺ immune cells through increased CXCL12 secretion. In addition, the low expression of CXCR7, the alternative receptor for CXCL12, in immune cells suggests a limited role for the CXCL12–CXCR7 axis (Extended Data Fig. 3h).
a, Cell–cell communication analysis based on ligand–receptor interactions (top six) between stromal cells and lymphocytes in tVAT (left) and a comparison between tVAT, dVAT and tumour (right). b, Marked CXCL12–CXCR4 interactions among CD8+ T cells, CD4+ T cells, B cells, plasma cells and stromal cell populations in tVAT, dVAT and tumour. The width of the lines represents the probability of communication. c, Violin plots showing the expression of CXCL12 (top) and CXCR4 (bottom) across all cell types in patients with CRC. d, Violin plots comparing the expression of CXCL12 in dVAT versus tVAT (top) and tumour versus tVAT (bottom) in patients with CRC, analysed using a two-sided Wilcoxon test. e, Experimental design for the PAT C57BL/6J mouse model treated with IgG or anti-CXCL12 antibody (left), and representative MC38 tumour images at day 16 of the experiment (right) (n = 5). f, Tumour growth (left) and tumour weights (right) of MC38 tumours at day 16 of the experiment in C57BL/6J mice (n = 5). g, Representative MC38 tumour images (left) and tumour growth (right) of experiments in control and Cxcl12fl/fl cKO mice (n = 6). h, Tumour weights of MC38 tumours in control and Cxcl12fl/fl cKO mice at day 16 of the experiment (n = 6). i, Flow cytometry analysis of the infiltration of various CXCR4+ immune cells in MC38 tumours in Control and Cxcl12fl/fl cKO mice (n = 6). j, Schematic diagram of the chemotaxis assay using T cells as ‘sensors’ and conditional medium as a ‘sink’ (left), and the aggregated trajectories of control or CXCL12-induced T cells migrating for 1 h (right). k, Quantitative analysis of CXCR4+ CD45.1+ T cells in MC38 tumours with and without removal of PAT or contralateral inguinal adipose tissue (control) by flow cytometry (n = 5). Data represent ≥3 independent experiments. Statistical significance was assessed by a two-sided permutation test (a), two-sided unpaired Student’s t-test (f right, h and i), one-way analysis of variance (ANOVA) with Tukey’s correction for multiple comparisons (k right) or two-way ANOVA (f left and g right). Graphs display mean ± s.d. (f–i,k). Panels created with BioRender: e and k, Huaiqiang, J. https://biorender.com/ovq2e39 (2026).
To validate the functional role of the CXCL12–CXCR4 axis in immune cell recruitment, we injected a CXCL12-neutralizing antibody (anti-CXCL12) into the PAT of tumour-bearing mice (Fig. 4e). The results demonstrated that CXCL12 neutralization substantially suppressed tumour growth (Fig. 4e, f). Next, we generated Cxcl12 conditional knockout mice (cKO) by injecting AAV2/9-CAG-Cre into the PAT of Cxcl12fl/fl mice (Extended Data Fig. 3i–l). The results showed that tumours in the Cxcl12 cKO were significantly smaller and showed increased infiltration of CXCR4⁺ immune cells than those in wild-type controls (Fig. 4g–i). A Transwell chemotaxis assay further confirmed the strong chemotactic activity of CXCL12 toward T cells (Fig. 4j). In addition, the competitive study showed that removal of PAT resulted in more CXCR4⁺ CD45.1⁺ immune cells infiltrating the tumour (Fig. 4k). Together, these findings demonstrate that tVAT increases CXCL12 levels, which in turn recruits CXCR4⁺ immune cells, thereby limiting their infiltration into tumour tissue and acting as a competitive immune sink that reduces the availability of cytotoxic T cells in the TME.
Tumour induces adipose mesenchymal transformation in tVAT
To identify the cellular origin of elevated CXCL12 expression in tVAT, we further analysed the stromal cell compartment (Fig. 5a and Extended Data Fig. 4a). Three major cancer-associated fibroblast (CAF) lineages were identified, including matrix CAFs (mCAFs), inflammatory CAFs (iCAFs) and antigen-presenting CAFs (apCAFs). Besides, we also discovered VAT-associated stromal cells, mainly adipose stromal cells (ASCs; PI16 and CD55) and preadipocytes (pACs; APOD and APOE)32,33 (Fig. 5a and Extended Data Fig. 4a). Tissue distribution analysis showed notably differences in stromal components across diverse tissues, with VAT-associated stromal populations, particularly accumulated ASCs and pACs, enriched in VATs (Fig. 5b and Extended Data Fig. 4b,c). We then characterized the precise clustering of VAT-associated stromal cells in VATs and identified two ASC and five pAC subclusters (Fig. 5c and Extended Data Fig. 5a). Notably, we identified a unique fibroblastic-like subcluster characterized by high expression of COL1A1 and MDK (Fig. 5c and Extended Data Fig. 5b). This subcluster was markedly expanded in tVAT, but was rarely detected in dVAT (Fig. 5d,e). Given its unique transcriptional profile and preferential enrichment in tVAT, we designated this subpopulation as adipose-derived cancer-associated fibroblasts (adCAFs). Further analysis revealed that adCAFs displayed fibroblastic-like phenotypes (ACTA2, COLA1 and LUM), CAF-like features (FAP and PDGFRB) (might be precursor cells of CAF), as well as strong secretory activity with multiple cytokines such as CXCL12, IGF1 and MDK, etc. (Fig. 5f and Extended Data Fig. 4d).
a, UMAP of all stromal cells in tVAT, dVAT, tumour and normal from patients with CRC, with ten clusters labelled by inferred cell types. Major lineages included ASCs, pACs, CAFs, pericytes (PCs) and mesothelial cells (Mesos). b, Heatmap displaying the distribution of eight stromal cell subtypes across different tissue types. c, UMAP of eight subsets of VAT-associated stromal cells in tVAT and dVAT from patients with CRC, including ASCs, pACs and adCAFs. d, Beeswarm plot showing the distribution and abundance of VAT-associated stromal cell types in Nhoods between tVAT and dVAT. e, Stack plot displaying the abundance of the eight VAT-associated stromal cell subsets in dVAT and tVAT. f, Heatmap showing the RNA expression of various marker genes in VAT-associated stromal cell types, including ASC/pAC markers, CAF markers, cytokines and stromal markers. g, Representative multiplex immunofluorescence images showing the presence of adCAFs in tVAT samples from patients with CRC. Scale bar, 10 μm. DAPI, 4,6-diamidino-2-phenylindole. h, Relative expression levels of multiple cytokines and protumoural factors in adCAF-enriched stromal cells (n = 4) and non-adCAF stromal cells (n = 4) derived from the PAT of mice xenograft models. The data are presented as a box-and-whisker graph (bounds of box show first to third quartile, bottom and top line show minimum to maximum and the central line shows the median). i, Cell–cell communication analysis based on ligand–receptor pairs (top six) between adCAFs and lymphocytes in tVAT (left) and a comparison between tVAT and dVAT (right). j, RT–qPCR (left) and ELISA (right) detecting the RNA expression and protein secretion of CXCL12 in sorted adCAF-enriched stromal cells and non-adCAF stromal cells from PAT of mice. Data represent ≥3 independent experiments. All data are shown as mean ± s.d. and statistical significance was assessed by a two-sided, unpaired Wilcoxon test (h), two-sided permutation test (i) and Student’s t-test (j).
To confirm the presence of adCAFs within tVAT in patients with CRC, we performed mIHC staining and confirmed the localization of adCAFs within tVAT (Fig. 5g). We further isolated MDK⁺FAP⁺PDGFRB⁺ stromal cells, enriched for adCAF, from the PAT in tumour-bearing mice (Extended Data Fig. 5c). The results showed that adCAF-enriched stromal cells exhibited a spindle-shaped, fibroblastic morphology and showed transcriptional similarity to the adCAF cluster (Extended Data Fig. 5d–f), as well as strong secretory activity with multiple cytokines such as Cxcl12, Igf1, Mdk, etc. (Fig. 5h). The intercellular communication analysis revealed a stronger activation of the CXCL12–CXCR4 axis in adCAF–immune cell interactions in tVAT compared with dVAT (Fig. 5i), and robust protumoural effects through the adCAF–tumour interactions of multiple growth factors (Extended Data Fig. 5g). Consistently, adCAFs exhibited significantly elevated Cxcl12 expression at both the messenger RNA and protein levels (Fig. 5j and Extended Data Fig. 5h). Collectively, these findings demonstrate that adCAFs represent a major source of CXCL12 in tVAT and plays a central role in establishing a local immune cell reservoir through activation of the CXCL12–CXCR4 signalling axis.
To validate our findings, we performed single-nucleus RNA sequencing (snRNA-seq) on paired tissues from five patients with CRC to overcome the technical limitations of scRNA-seq in capturing mature adipocytes (Extended Data Fig. 6a,b). Subclustering of mature adipocytes identified six major subpopulations and revealed a notably high abundance of the SCD + AC subgroup in tVAT (Extended Data Fig. 6c–e). However, our snRNA-seq data revealed that mature adipocytes themselves show minimal CXCL12 expression and do not significantly rely on CXCL12–CXCR4 axis for immune cell recruitment (Extended Data Fig. 6f,g), indicating that the dominant source of CXCL12-mediated immune recruitment originates from adCAFs rather than mature adipocytes. Further subclustering of VAT-associated stromal cells confirmed that adCAFs are specifically and highly enriched in tVAT and exhibit co-expression patterns characteristic of both CAFs and ASCs/pACs (Extended Data Fig. 6h–k), consistent with our initial scRNA-seq findings.
Tumour-derived factors induce ASC to differentiate into adCAF
The high abundance of adCAFs in tVAT suggests that tumour-derived factors may play a role in their formation. To explore this, we performed cell trajectory analyses via Monocle 3 (Fig. 6a,b). The results showed that ASCs predominantly differentiated into pACs34, with only a small fraction transitioning into adCAFs in dVAT. In contrast, ASCs in tVAT predominantly differentiated into adCAFs rather than pACs (Fig. 6a), indicating that the tumour environment drives a shift in stromal cell fate near the tumour site. We also validated the developmental trajectories using another model via Monocle2, which also revealed a predominant differentiating potential of ASCs into adCAFs rather than pACs in tVAT (Fig. 6c–e). This shift in differentiation was accompanied by altered transcriptional regulation, with increased activity of transcription factors such as STAT2, TWIST1, CREB3L1, MXD4 and SNAI1, which are likely involved in promoting the adCAF phenotype (Fig. 6f,g). To further evaluate whether tumour-derived factors can induce the differentiation of ASCs into adCAF, we established the tumour-derived conditioned medium (CM) assay and performed in vitro validations (Extended Data Fig. 7a). Compared with the control group, mouse ASCs exposed to CM underwent a distinct morphological transition toward a fibroblast-like phenotype, followed by upregulation of adCAF-specific markers, transcription factors and signalling pathways associated with adCAFs (Fig. 6h–j and Extended Data Fig. 7b). In addition, adCAF-specific markers and transcription factors were elevated in peritumoural versus contralateral adipose tissue in MC38 tumour-bearing mice (Extended Data Fig. 7c,d). These results indicate that soluble factors secreted by tumour cells may induce adCAF differentiation in a paracrine manner.
a, Cell trajectories (top) and cell abundance (bottom) of ASC-derived pAC and ASC-derived adCAFs inferred by Monocle3, with cell trajectories in dVAT (left) and tVAT (right). The pseudotime colour code is shown in the right box. b, Dynamic changes in the expression of several marker genes over time in dVAT (left) and tVAT (right) by monocle3 pseudotime. c, Pseudotime (top) and cell trajectories (bottom) of ASC-derived pAC and ASC-derived adCAFs inferred by Monocle2. The pseudotime colour code is shown in the right box. d, Cell trajectories in dVAT (top) and tVAT (bottom) of ASC-derived pAC and ASC-derived adCAFs inferred by Monocle2. e, Dynamic changes in the expression of several marker genes over time in pAC markers (left) and adCAF markers (right) by monocle2 pseudotime. f, Dot plot showing the inferred differential transcription factor (TF) activities in ASC, pAC and adCAF. g, Dynamic changes in inferred activities and RNA expression levels of TFs in ASC, pAC, and adCAF. h, Representative images of the morphology of mASCs treated with MC38 tumour-derived CM from MC38-derived tumour tissues for 72 h. Scale bar, 100 μm. i, RT–qPCR detecting the RNA expression of adCAF-associated genes (top) and TFs (bottom) in mASCs treated with CM. j, Bar chart of KEGG pathway enrichment analysis showing differentially enriched signalling pathways in mASCs treated with tumour CM. (k) Bar chart of KEGG pathway enrichment analysis showing differentially enriched signalling pathways between adCAF-enriched stromal cells and non-adCAF stromal cells. Data represent ≥3 independent experiments. All data are shown as mean ± s.d. and statistical significance was assessed by a two-sided, unpaired Student’s t-test (i).
To identify the key tumour-derived signalling pathways involved, we performed KEGG pathway analysis on sorted adCAF and non-adCAF populations and observed significant enrichment of the TGF-β signalling pathway in adCAF, suggesting its potential role in driving their formation (Fig. 6k). ELISA further confirmed that TGF-β1 levels were significantly elevated in supernatants from MC38-derived tumours compared with those from normal intestinal tissue (Extended Data Fig. 7e). To validate the role of TGF-β1 in inducing adCAF, we stimulated mASCs in vitro with recombinant murine TGF-β1 and found that this treatment resulted in a pronounced fibroblast-like morphological change and a significant increase in the expression of adCAF markers (Extended Data Fig. 7f,g). Taken together, these findings indicate that tumour-derived TGF-β1 is a key driver of ASC differentiation into adCAFs within the tumour-associated adipose microenvironment.
Clinical significance of the tumour-adipose tissue crosstalk
Given that we had established the role of adCAFs in competitively recruiting immune cells to tVAT, we hypothesized that targeting adCAFs could reduce immune cell sequestration in tVAT, enhancing the efficacy of immune-checkpoint blockade (ICB). To test this, we constructed MdkDTR mice to specifically eliminate the majority of adCAFs in the PAT (Fig. 7a and Extended Data Fig. 8a,b), based on the high expression of the MDK in adCAFs. The results demonstrated that Mdk+ adCAF depletion substantially enhanced the therapeutic effects of anti-PD-1, as evidenced by the reduced tumour burden and increased infiltration of immune cells, including CXCR4⁺ or tumour antigen-specific immune subsets within the tumour (Fig. 7a–d). These findings further support the role of adCAFs in diverting immune cells away from the tumour, thereby highlight the therapeutic potential of targeting adCAFs to enhance the efficacy of immunotherapy in tVAT-associated tumours.
a, Experimental design for constructing Control and MdkDTR cKO mice bearing MC38 tumours near PAT, followed by αPD-1 therapy. b,c, Representative MC38 tumour images (b), tumour weights (c left) and tumour growth (c right) in control and MdkDTR cKO mice treated with IgG or anti-PD-1 (n = 6). d, Flow cytometry analysis of the infiltration of immunocytes, including T cells, CD4+ T cells, CD8+ T cells, and tumour-specific CD8+ T cells, CXCR4+ immunocytes, CXCR4+ T cell, CXCR4+ CD4+ T cell, CXCR4+ CD8+ T cell and CXCR4+ tumour-specific T cell in MC38-OVA tumours from the four treatment groups (n = 6). e,f, Representative MC38 tumour images (e) and tumour weights (f) of the experiment in mice treated with anti-CXCL12 and/or anti-PD-1 (n = 5). g, Representative MRI image of CRC tumour and corresponding tVAT area region of CR and non-CR patients pre- and post-immuno-chemoradiotherapy. The yellow area represents the tVAT area, whereas the red area denotes the tumour region. Note that the mass visible in the intestinal lumen (top right) is faecal material. h, Pre-treatment tVAT area difference based on 3D Slicer between CR (n = 30) and non-CR (n = 37) patients. The data are presented as a box-and-whisker graph (bounds of box show first to third quartile, bottom and top line show minimum to maximum and the central line shows the median). i, ROC plot of response predicting ability of pre-treatment PAT area in immuno-chemoradiotherapy of proficient mismatch repair patients with CRC, compared with conventional indexes, including CPS, TPS, CEA and CA199 (n = 67) with optimal cutoff. j, Comparison of pCR ratio in tVAT high and low group according to the optimal cutoff. k, Graphical abstract depicting how tumours reshape the stromal environment in tVAT and how tVAT competes for immunocytes from the tumour to promote immune escape. Data represent ≥3 independent experiments. Statistical significance was assessed using a two-sided, unpaired Student’s t-test (d,h), one-way ANOVA with Tukey’s correction for multiple comparisons (c left, f) or two-way ANOVA with Tukey’s correction for multiple comparisons (c right). Graphs display mean ± s.d. (c,d,f,h). Panels created with BioRender: a and k, Huaiqiang, J. https://biorender.com/e5jwcye (2026).
To further explore the therapeutic and diagnostic significance of adipose–tumour crosstalk, we evaluated the therapeutic effects of anti-CXCL12 in combination with anti-PD-1 in MC38 tumours (Extended Data Fig. 8c). We blocked PAT-derived CXCL12 by local injection and observed the combination therapy exhibited a synergistic antitumour effect as evidenced by reduced tumour growth and increased immune cell infiltration (Fig. 7e,f and Extended Data Fig. 8d–f). Additionally, we observed that combining AMD3100 (the small-molecule CXCR4 antagonist) with anti-PD-1 substantially enhanced tumour suppression compared with either monotherapy (Extended Data Fig. 9a–d)35. To fully simulate the impact of visceral fat on immune infiltration in tumours, we established a caecal orthotopic CRC model. In this setting, both AMD3100 (Extended Data Fig. 9e–g) and anti-CXCL12 (Extended Data Fig. 9h–l) potently synergized with anti-PD-1 therapy, leading to significantly enhanced antitumour immunity compared with monotherapy regimens.
Furthermore, we analysed a cohort of 67 patients with locally advanced CRC (a well-established tVAT-associated malignancy) treated with neoadjuvant immuno-chemoradiotherapy at our institute (Supplementary Table 1). Notably, patients who achieved complete response (CR) exhibited significantly smaller tVAT area compared with non-CR patients (Fig. 7g,h). Imaging assessment of tVAT area may serve as a predictive biomarker for immune response in CRC achieving an area under the curve of 0.887 in all patients, as well as competitive predictive power in both T3 and T4 stage patients. It demonstrates superior predictive performance compared with conventional histopathological metrics such as combined positive score (CPS) and tumour proportion score (TPS) (Fig. 7i and Extended Data Fig. 8g,h). In patients with greater tVAT area, the ICB response rate decreased sharply from 88.5% to 17.1%, further supporting tVAT elimination as a promising strategy to improve ICB efficiency (Fig. 7j). As illustrated in Fig. 7k, we proposed a working model of how PAT drives immune evasion in CRC via adipose–mesenchymal transformation.
Discussion
Recent research by our team focuses on how TME remodelling promotes immune escape of gastrointestinal tumours36,37,38. Furthermore, this study provides a comprehensive analysis of tVAT in CRC and uncovers a high-resolution landscape and a previously unrecognized role of tVAT in shaping the immune macroenvironment. The spatial proximity between tumours and tVAT suggests that adipose tissue may play roles in tumour progression through paracrine signalling or metabolic crosstalk10,39. Indeed, most studies have focused on how adipose tissue or adipocytes promote tumour growth and metastasis40,41. Despite the growing interest in the interactions between tumours and tVAT, only a limited number of studies have explored their bidirectional regulatory relationship. One such study showed that renal carcinoma induces the browning remodelling of tVAT and further promoted tumours by releasing lactate into the TME10. Our study revealed a previously unrecognized mechanism of immune diversion, expanding the concept of effector immune cell deployment42. This mechanism also partially explains why CRCs tend to form an immune-excluded TME and generally exhibit poor responses to ICB43.
VAT-associated factors promote tumour immune evasion through both physical and biochemical mechanisms44,45,46, and we identified a role of PAT in this process via CXCL12 secretion. The CXCL12–CXCR4 axis serves as a central mediator of tumour progression by contributing to metastasis, angiogenesis and immunosuppression within the TME47,48. Tumours, as the epicentre of inflammatory responses, recruit large numbers of immune cells through systemic circulation and surrounding tissues49. In this study, we found that tVAT competitively recruits functionally diverse CD8⁺ T cells through the CXCL12–CXCR4 axis. This observation expands our understanding of immune cell exclusion mechanisms in tumours50,51. Moreover, the reported expression of CXCR4 on tumour cells may explain why tumours often invade the adipose tissue boundaries or exhibit skip metastases in peritumoural regions52. While previous studies and trials have validated the therapeutic relevance of CXCL12 in CRC53,54, our work demonstrates that its pivotal role is specifically mediated by adipose–tumour crosstalk, thus elucidating the underlying mechanism.
The previously uncharacterized adCAFs arise in response to tumour-derived TGF-β1, aligning with established views on the role of TGF-β1 in modulating stromal cells within the TME55,56. Furthermore, this aligns with previous findings showing that adipose tissue adjacent to tumour lesions exhibits greater fibrosis compared with regions distant from invasive tumour cells57. Building upon previous evidence for adipose-derived CXCL12 (refs. 58,59), our study defines adCAFs as the key cellular mediator within the peritumoural niche, providing its precise cellular identity. The coexistence of intratumoural CAFs and peritumoural adCAFs suggests a dynamic, multilayered process of T cell exclusion: CAFs may initially expel T cells from the tumour core, whereas adCAFs further sequester them within adjacent adipose tissue, reinforcing immune evasion60,61. Collectively, these findings establish the adCAF–CXCL12–CXCR4 axis as a key regulator of immune cell distribution and substantially advance our understanding of the tumour and PAT interaction network.
Several studies have demonstrated that the density of PAT is strongly associated with tumour progression in breast, colorectal and renal cancer62,63,64, with its radiological features serving as clinically valuable indicators for assessment and prediction. Consistently, our study indicated that imaging assessment of PAT may serve as a predictive biomarker for immune response in patients with visceral tumours. Targeting the CXCL12–CXCR4 axis has been proven to enhance the efficacy of tumour immunity in multiple types of cancer65,66,67. Building on this, our study also indicated that targeting the adCAF–CXCL12 axis further enhanced responses to PD-1 therapy. The proximity of visceral tumours to adipose tissues further highlights the therapeutic potential of targeting the adCAF–CXCL12 axis. Our team has also previously explored clinical intervention strategies targeting different stages of the immune cycle to enhance immune efficacy of CRC68,69. Clinically, these results further emphasize the importance of considering the TME beyond the tumour core, incorporating adjacent non-tumoural structures such as adipose tissue, to evaluate the immunoefficacy and explore the strategy of immunotherapies.
In conclusion, this study offers a major breakthrough in our understanding of the dynamic crosstalk between tumour and adipose tissues. Moreover, targeting adipose–tumour interaction holds promise for enhancing diagnostic and therapeutic efficacy of immunotherapy for patients with visceral tumours.
Methods
Ethical approval
All samples were collected from patients with informed consent, and informed consent for publication was obtained from all participants. All related procedures were performed with the approval of the Medical Ethics Committee of Sun Yat-sen University Cancer Center (SYSUCC; (B2024-759-01). Animal experiments were approved by the Institutional Animal Care and Use Committee of Sun Yat-Sen University (L025504202205013). This study was performed according to the Declaration of Helsinki. The maximum permitted tumour volume was set at 2,000 mm3 or 20% of the animal’s body weight, whichever threshold was reached earlier. Throughout the study, all observed tumours remained below these approved limits. In compliance with institutional protocols, any animal that met predefined humane end points was promptly killed, and all surviving mice were humanely killed at the study conclusion.
Tissue specimens
The CRC tissue specimens (n = 38 paired samples), including primary tumour, adjacent visceral adipose tissue (tVAT, ≤2 cm) and dVAT (>10 cm), were collected from SYSUCC for omics analysis or experimental verification. All samples were collected from patients with informed consent, and all related procedures were performed with the approval of the Medical Ethics Committee of SYSUCC (B2024-759-01). Patients were recruited with no perceived bias, and were not treated with adjuvant radiotherapy or chemotherapy before their operation.
scRNA-seq data generation
Fresh tissue specimens were dissected into approximately 1–2-mm3 pieces and enzymatically digested with adipose or a Tumour Tissue Dissociation kit, respectively (Miltenyi) on a rotor at 37 °C for 40 min. Dissociated cells were subsequently filtered through a 40-μm cell strainer and then centrifuged at 450g for 5 min. After removal of supernatant, the pelleted cells were suspended in red blood cell lysis buffer Hybri-Max (Sigma-Aldrich) and incubated on ice for 3–5 min to lyse red blood cells. After washing twice with 1× phosphate-buffered saline (PBS), the cells were resuspended in FACS buffer (PBS supplemented with 2% fetal bovine serum (FBS)). Given the sensitivity of primary cells to prolonged processing, we prioritized rapid library preparation without live/dead cell separation to preserve transcriptomic integrity. The concentration of single-cell suspensions was adjusted to approximately 1,000 cells per μl as determined by hemacytometer. Cells were captured in droplets and then underwent reverse transcription and cell barcoding in droplets. Emulsions were disrupted and complementary DNA purified followed by PCR amplification. According to the manufacturer’s instructions, the amplified cDNA was utilized for constructing the 10x Chromium Single-cell 3′ library. For the samples used for TCR/BCR enrichment, the amplified cDNA was utilized to construct both the 10x Chromium Single-cell 5′ library and the human variable, diversity and joining (VDJ) library (10x Genomics) for gene expression library construction and TCR/BCR enrichment. The purified libraries were analysed by an Illumina Novaseq 6000 sequencer with 150-bp paired-end reads (Beijing CapitalBio Technology).
scRNA-seq data aggregation, processing and clustering
The scRNA-seq data derived from four distinct tissue types (tVAT, dVAT, normal and tumour) were aligned and quantified with the Cell Ranger (v.4.0) program using the GRCh38 human reference genome. Preliminary filtered data generated from Cell Ranger were used for downstream analysis. Quality control was applied to filter out low-quality cells with either (1) <500 or >8,000 expressed genes or (2) >20% mitochondrial genes. Besides, droplets containing two or more cells were filtered using the DoubletFinder program. For the integrated analysis of single-cell samples with Seurat tool (v.4.1.1), the IntegrateData function was used to combine all individual sample datasets into a single integrated dataset. Each dataset underwent normalization, feature selection and data correction, and robust principal-component analysis (PCA)-based integration was performed to remove batch effects. Then, the single-cell data were processed following the Seurat standard protocol. The FindVariableFeatures method was employed to identify the top 3,000 differentially expressed genes, which were utilized as hypervariable genes for subsequent studies. This dataset was dimension-reduced using PCA with highly variable features, and the first 15 principal components were selected for subsequent analysis. Unsupervised clustering was performed using the FindNeighbors and FindClusters functions based on the edge weights between any two cells and the shared nearest neighbour graph generated by the Louvain algorithm. The detected cell clusters were visualized using t-distributed stochastic neighbour embedding (t-SNE) and Uniform Manifold Approximation and Projection. Highly differentially expressed genes and canonical markers were used for the annotation of unsupervised clusters. After rough clustering and annotation, more refined clustering and annotation were performed for each major cell lineage, such as lymphocytes, myeloid cells and stromal cells in four CRC tissue types, including tVAT. Finally, the refined annotation information was further annotated onto the whole dataset to generate a curated annotation.
scTCR/BCR-seq data processing
The TCR and BCR sequence data corresponding to CRC tissues (tVAT, dVAT and tumour) from 10x Genomics were processed using Cell Ranger software (v.4.0) with the manufacturer-supplied human VDJ reference genome. For each sample, the output file filtered_contig_annotations.csv, containing TCR α/β-chain CDR3 nucleotide sequences and BCR light/heavy chain CDR3 nucleotide sequences, was applied to downstream analysis via the scRepertoire package (v.1.8.0)70. Only the assembled chains that were productive, highly confident, full length, with a valid cell barcode and an unambiguous chain type (for example, α) assignment were retained. For each patient, cells with an identical TCR α/β-chain pair or BCR light/heavy chain were considered as having originated from the same clonotype and were therefore identified as clonal cells. Subsequently, TCR data were matched to the single-cell atlas of CD4+ and CD8+ T cells to explore the distribution of TCR across tVAT, dVAT and tumour. T cells containing TCRs shared with those within tumours were defined as tumour-antigen-specific T cells.
Cell–cell communication interactions analysis based on ligand–receptor pairs
To evaluate the potential interaction relationship between immune cells and various cellular components in the stroma of tVAT, we used the CellChat program (v.2.1) to analyse cell–cell interactions. The raw count matrix and the corresponding cell-type annotations exported from the Seurat object were used as input. The underlying interaction strength between ligand and receptor was predicted according to their average expression. Subsequently, crucial ligand–receptor pairs were extracted for presentation and analysis.
Single-cell trajectory analysis
We used the Monocle2 algorithm (v.2.18.0) and Monocle3 algorithm (v.1.3.1) with default parameters to infer the interconversion and evolutionary path between individual cells in the adipocyte-associated stromal cell subpopulations to study the plasticity and dynamic differentiation process of each cell subpopulation of the same origin. Specifically, to reflect the intrinsic physiological properties of cells, we chose to use an unsupervised approach and employed the raw count matrix of single-cell data.
RNA-seq
Cells were digested with trypsin and washed with PBS, and RNA was subsequently extracted using the TRIzol method. Transcriptome sequencing was performed using the Illumina platform. Gene expression (counts and fragments per kilobase million) was quantified using the GRCh38 (hg38) p13 reference genome with Gencode v.32 annotation. Read alignment was performed with STAR, followed by transcript abundance estimation via RSEM. Functional enrichment analysis was performed using clusterProfiler (v.4.0) with default parameters, referencing Gene Ontology (GO) and KEGG gene sets. Input genes consisted of differentially expressed genes between cell subsets or experimental groups. For signature evaluation in both single-cell and bulk transcriptome datasets, we applied the single sample gene set variation analysis (ssGSVA) algorithm (v.1.14.1), which is a nonparametric, unsupervised method.
Cell culture
MC38, CT26 and E0771 cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco) supplemented with 10% FBS and 1% penicillin–streptomycin. The MC38 cell line was kindly provided by W. Yang (Guangdong Provincial People’s Hospital). CT26 and E0771 cells were purchased from the American Type Culture Collection. The mouse ASCs were purchased from IMMOCELL and cultured in specific medium specified by the supplier. All cells were incubated in 5% CO2 at 37 °C in a humidified incubator. To prepare conditional medium (CM) from primary tumour and healthy intestine tissue, the tissues were cultured in DMEM supplemented with 1% penicillin–streptomycin. After 72 h, the supernatants were collected and centrifuged, and then were passed through 0.22-μm filters and stored at −80 °C for use. All cells were negatively tested for mycoplasma contamination and authenticated based on short tandem repeat fingerprinting before use.
Animal studies
Six-week-old C57BL/6J mice (strain no. 219), BALB/c mice (strain no. 211), BALB/c-Nude mice (strain no. 401) or NOD.SCID γc-deficient (NSG) mice (strain no. 408) were purchased from Beijing Vital River Laboratories. CD45.1 (C57BL/6JGpt-Ptprcem1Cin(p.K302E)/Gpt) mice (strain no. T054816), Mdk-GFP (Mdk-P2A-EGFP Cas9-KI) mice, MdkDTR (Mdk-LSL-P2A-DTR) mice and Cxcl12fl/fl (C57BL/6JGpt-Cxcl12em1Cflox/Gpt) mice (strain no. T008662) were generated by Gempharmatech. Genetically engineered mice were from in-house matings. All mice were housed under specific-pathogen-free conditions in a temperature-controlled facility with a 12-h light–dark cycle. Standard rodent chow and water were provided ad libitum. All related protocols were approved by the Institutional Animal Care and Use Committee of Sun Yat-Sen University.
Antibodies
The following antibodies were used: anti-human CD3 (ZM-0417, ZSGB-BIO, 1:500), CD20 (ab-78237, Abcam, 1:200), CD21 (ZM-0040, ZSGB-BIO, 1:50), CD23 (ZA-0516, ZSGB-BIO, 1:200), Ki67 (ZM-0166, ZSGB-BIO, 1:100), PANCK (ZM-0069, ZSGB-BIO, 1:100); anti-FAP (66562, Cell Signalling Technology, 1:200), anti-PDGFRB (3169S, Cell Signalling Technology, 1:200), anti-MDK (11009-1-AP, Proteintech, 1:200); anti-mouse CD45-APC-Fire750 (103154, BioLegend, 1:100), CD31-PC5.5 (102420, BioLegend, 1:100), FAP-AF647 (FAB9727R, R&D Systems, 1:100), PDGFRB-PE (12-1402-81, eBioscience, 1:50), MDK-PE-Cy7 (Sanyou Bio, 1:200), CD45-APC-Cy7 (103116, BioLegend, 1:100), CD3-FITC (100204, BioLegend, 1:100), CD4-PE-Dazzle594 (100456, BioLegend, 1:100), CD8-PC5.5 (100734, BioLegend, 1:50), CXCR4-APC (146508, BioLegend, 1:50), CD45.1-BV510 (110741, BioLegend, 1:100); anti-mouse β-tubulin (TA-10, ZSGB-BIO, 1:1,000), CXCL12 (3530S, Cell Signalling Technology, 1:1,000); and H-2Kb OVA Tetramer-PE (TS-5001-1C, MBL, 1:20). Secondary antibodies were Alexa Fluor 488-goat anti-mouse (A0428, Beyotime, 1:100), Alexa Fluor 647-goat anti-rabbit (A0468, Beyotime, 1:100), HRP-goat anti-mouse (ZB-2305, ZSGB-BIO, 1:100) and HRP-goat anti-rabbit (ZB-2301, ZSGB-BIO, 1:100). Functional antibodies were CXCL12-neutralizing antibody (MABC184, Merck Millipore), anti-mouse PD-1 (BE0146, Bio X Cell), anti-mouse CD3ε (100340, BioLegend, 2.5 μg ml−1) and anti-mouse CD28 (102116, BioLegend, 1 μg ml−1).
T cell co-stimulation assay
A T cell co-stimulation assay was conducted as previously described71. T cells were isolated from the spleens of C57BL/6J mice and then co-stimulated. Splenocytes were cultured in RPMI-1640 supplemented with 10% FBS, 1% penicillin–streptomycin, 10 ng ml−1 recombinant mouse IL-2 (PeproTech) and 100 mM β-mercaptoethanol (Sigma-Aldrich). For T cell co-stimulation, splenocytes were treated with 2.5 μg ml−1 anti-CD3ε monoclonal antibody (BioLegend) and 1 μg ml−1 anti-CD28 monoclonal antibody (BioLegend) for 2–3 days.
IB, IF and mIHC staining
Immunoblot (IB) and immunofluorescent (IF) staining were performed according to the standard procedure as previously reported37,72. To validate the existence of TLSs or adCAFs in tVAT, mIHC staining was performed on the remaining biopsy sections of patients with CRC. Tyramide signal amplification (TSA) was applied to enhance the signal, and seven-colour or four-colour AlphaTSA Multiplex IHC kit (Alpha X Biotech) was used as reported previously according to the manufacturer’s protocol73,74. In brief, endogenous peroxidase in the rehydrated sections was inactivated by incubation in 3% H2O2 in methanol for 10 min at room temperature. Later, antigen retrieval was performed on them with a microwave heating method following routine methods. After being blocked with 10% normal goat serum (Absin) to block nonspecific sites, the sections were incubated with one of the primary antibodies for 1 h at 37 °C and incubated with the secondary antibody for 10 min at 37 °C. Later, the sections were labelled with corresponding TSA dye for 5 min at room temperature with full light protection. After completing the above-mentioned round of staining, the sections re-enter the cycle of antigen retrieval-blocking-staining until all the primary antibodies’ staining was finished. Finally, the samples were mounted in anti-fluorescence quenching mounting medium that contains 4,6-diamidino-2-phenylindole (DAPI).
For the validation of TLS, a panel of anti-human antibodies (Panel 1) was used, including anti-CD3, anti-CD20, anti-CD21, anti-CD23, anti-Ki67 and anti-PANCK. The staining procedure was performed by AlphaPainter X30 Multi-target Pathological Staining Machine (Alpha X Biotech), using human tonsil sections as positive controls. Multispectral images were scanned with a Zeiss Axioscan7 whole-slide imaging system (Carl Zeiss), and visualized using ZEN v.3.3 software (Carl Zeiss) to distinguish TLS, B cells, T cells, germinal centre (GC) B cells and follicular DCs (FDCs). For the validation of adCAF, a panel of anti-human antibodies (Panel 2), including anti-FAP, anti-PDGFRB and anti-MDK was used. The staining procedure was performed manually and the multispectral images were acquired by a digital pathology section scanner (KFBIO, KF-PRO-020).
ATS-GNP synthesis
For reducing peritumoural fat specifically and avoiding damaging blood vessels, ATS-GNP was used to replace the surgical elimination of PAT. As adipocytolytic polymer nanoparticles, ATS-GNP can be selectively absorbed by adipocytes and generate carbon dioxide gas within late endosomes/lysosomes to induce adipocytolysis, leading to the decrease in the number of adipocytes. ATS-GNP were synthesized as reported previously according to the custom protocol31. In brief, nanoparticles were prepared with a calcium carbonate (CaCO3) (Sigma-Aldrich) core and a poly (D,L-lactide-co-glycolide) (PLG) (Lactel Absorbable Polymers) shell via a certain method, thus producing gas-generating nanoparticles (GNPs). Later, adipocyte-targeting sequence (ATS; CKGGRAKDC) peptide (Peptron) was covalently conjugated to the surface of these nanoparticles using a carbodiimide reaction. This reaction occurred between the carboxyl group of ATS and the amino group of the ethylenediamine-modified PLG nanoparticles, thus obtaining ATS-GNP. GNP was regarded as a control to evaluate the efficacy of ATS-GNP.
RNA isolation and qPCR analysis
TRIzol Reagent (Thermo Fisher Scientific) or RNA Quick Purification Kit (ESscience) was used to extract RNA from tissues and cells according to the manufacturer’s instructions. Next, Prime Script RT Master Mix kit (Takara) was used to synthesize complementary DNA, which then served as a template for real-time PCR using the GoTaq qPCR Master Mix kit (Promega) according to the manufacturer’s instructions. β-Actin or β-tubulin was used as the internal control genes, and relative changes in gene expression were analysed using the 2−ΔCt or 2−ΔΔCt formula.
CXCL12 detection by ELISA
The supernatant of mouse adCAFs or non-adCAFs was collected and concentrated 72 h after cell culture. The CXCL12 concentrations were determined using the Mouse CXCL12 ELISA kit (R&D system). Standard curve calibrators and supernatant samples were assayed according to the manufacturer’s instructions.
Tumour-derived conditioned medium preparation
For preparation of MC38 tumour-derived CM (adipose-CM), freshly resected MC38 tumour tissue were promptly transferred to sterile tubes containing 5 ml of serum-free RPMI 1640 medium. The tissues were washed five times with PBS to remove residual blood. The tissue was carefully dissected using sterile surgical blades, followed by three additional PBS washes. Isolated adipose samples were incubated in T25 flasks containing serum-free RPMI 1640 medium supplemented with 1% (v/v) penicillin–streptomycin (500 mg 8 ml−1) for 24 h at 37 °C in a humidified incubator with 5% CO2. Following incubation, the supernatants were collected, centrifuged at 500g for 10 min at 4 °C, and filtered through 0.22-μm membranes to remove cellular debris and potential microbial contaminants. CM was prepared under identical conditions but without adipose tissue and served as a baseline reference.
Chemotaxis assay in vitro
To observe the chemotactic behaviour of T cells, a three-dimensional chemotaxis assay with μ-Slide Chemotaxis (ibidi) was used as reported previously according to the manufacturer’s protocol75,76,77. T cells were isolated from the spleens of C57BL/6J mice, co-stimulated, and then suspended at a density of approximately 3 × 106 cells per ml in the culture medium containing 1–3 mg ml−1 Matrigel Matrix (Corning). The cells with gel matrix were seeded in one filling port of the central channel and incubated for 1–2 h at 37 °C. After the gel matrix was solidified, a chemoattractant or chemoattractant-free solution was added into the two reservoirs of the chamber respectively according to the determined loading scheme.
To obtain the detailed documentation of the T cell migration over time, live cell images were acquired every minute for 2 h by Nikon ECLIPSE Ti2 (Nikon), a fully motorized fluorescence microscope with live cell imaging incubator. Specifically, the experiment included tracking data from 20–40 single cells. Then, for quantification of their movement, single cells were tracked over time by determining their position (xand y) on each frame of the image using the ImageJ Manual Tracking Plugin (Fabrice Cordelières). After tracking, the paths of the cells were visualized in trajectory plots providing information about chemotactic effects using Chemotaxis and Migration Tool Software (Ibidi).
Isolation of adCAF and non-adCAF stromal cells
MDK–GFP mice were inoculated with 1 × 106 tumour cells (suspended in 50 μl saline) near the inguinal adipose tissue. After 2 weeks, mice were killed and PAT was dissected to be digested in single-cell suspension with full light protection. MDK-coupled GFP fluorescence (from MDK–GFP mice) and a panel of anti-mouse antibodies (Panel 6), including anti-mouse CD45, CD31, FAP and PDGFRB, were used for sorting the adCAF-enriched stromal cells. Stromal cells were characterized and defined as Live/Dead−, CD45− and CD31−. In the gate of stromal cells, cells that were triple-positive for FAP+, PDGFRB+, MDK+ were considered to be adCAF-enriched stromal cells, while the remaining cells were regarded as non-adCAF stromal cells. These cells were cultured in DMEM supplemented with 10% FBS and 1% penicillin–streptomycin. The adCAF-enriched stromal cells and non-adCAF stromal cells were cultured and detected by morphological examination under the microscope. Next, the CXCL12 protein and RNA expression levels were detected by western blot, RT–qPCR and ELISA. Then RNA sequencing was carried out for both of them.
To explore the differentiation from ASCs to adCAFs, mouse ASCs were treated with tumour supernatant for 72 h. Moreover, mouse ASCs were treated with TGFβ1 protein (Novoprotein) at concentrations of 5 ng ml−1 and 10 ng ml−1, respectively for 5–7 days, and then the characteristics of adCAFs were detected.
Tumour-bearing mouse models
For mouse models with tumour adjacent to inguinal adipose tissue, C57BL/6J, BALB/c, BALB/c-Nude mice, NSG mice, Cxcl12fl/fl mice, MdkDTR mice and MDK-GFP mice were used. A surgical procedure was employed to establish tumours adjacent to adipose tissue. Following anaesthesia, a small skin incision (~1 cm) was made in the inguinal region using sterile surgical scissors. This approach allowed for direct visualization of the well-defined inguinal fat pad, a substantial subcutaneous adipose depot consistently present at this location. Under direct observation, a 50 μl suspension containing 1 × 106 MC38-OVA or CT26 CRC cells was slowly injected into the adjacent loose connective tissue space using a 29-gauge insulin syringe. While the initial access was subcutaneous, the final cell delivery was periadipose. Tumour size was routinely monitored every 3 days using a caliper, and tumour volume was calculated using the standard formula V = length × width2/2. After the tumours grew to the indicated sizes (50–100 mm3), mice were randomized into different groups (n = 5 per group). Mice were killed when they met the institutional euthanasia criteria for tumour size and overall health condition.
For the surgical elimination of PAT, mice were anaesthetized with 1% pentobarbital sodium, positioned in lateral recumbency with the tumour facing upward, and a skin incision was made around the tumour. The PAT (weighing approximately 0.1 g post-resection) located immediately adjacent to the tumour mass was surgically removed. Control animals underwent either a sham operation or resection of the contralateral inguinal adipose tissue. Haemostasis was achieved by applying gentle pressure to bleeding points with cotton swabs, followed by wound suturing and disinfection with iodophor. The tissue resected in the experimental group specifically corresponded to the adipose pad directly surrounding the tumour, rather than representing distant or systemic fat depots.
For the CD45.1 transfer model, CD45.1+ T cells were isolated from the spleen of CD45.1 donor mice and co-stimulated in vitro. Thereafter, CD45.1+ T cells (5 × 106) were intravenously transferred for three consecutive days into CD45.2 recipient mice that were bearing tumours near the inguinal adipose tissue and underwent PAT resection78. After 5 days, the infiltration of CD45.1+ cells in tumour and PAT was detected by flow cytometry analysis.
For drug treatment, CXCL12 neutralizing antibody (Merck Millipore) was administered at a dose of 15 μg per mouse in PBS by injection into the inguinal adipose tissue every 3 days for a total of four times. The anti-mouse PD1 antibody (Bio X Cell) was given at 100 μg per mouse in PBS by intraperitoneal injection every 3 days for a total of four times.
Orthotopic caecal models
To establish an orthotopic caecal model for CRC, the following procedures were performed. First, MC38-luc cells were grown subcutaneously in donor mice to produce cell-derived xenograft tumour tissue. After 2 weeks, the resulting tumours were excised and cut into uniform 1-mm slices using a metal tissue slicer, and the slices were kept in PBS until use. Recipient mice were anaesthetized with 1% sodium pentobarbital and placed in a supine position. A 1-cm midline abdominal incision was made to expose the caecum. At a well-vascularized site on the caecal surface, the vasculature was gently scratched with a needle tip to create a mild injury. A tumour slice was then attached at this site using a biocompatible tissue adhesive. After returning the caecum into the abdominal cavity, the wound was sutured and disinfected with iodophor. At 5–7 days after surgery, mice were intraperitoneally injected with D-luciferin potassium salt (15 mg ml−1, 200 µl per mouse). Tumour development was assessed using a small animal in vivo imaging system (IVIS Spectrum, PerkinElmer), allowing subsequent processing and data recording.
For drug treatment, a CXCL12 neutralizing antibody (Merck Millipore) and an anti-mouse PD-1 antibody (Bio X Cell) were administered via intraperitoneal injection. AMD3100 (Selleck) was dissolved in saline and given daily at a dose of 10 mg kg−1 until the experimental end point.
Tissue dissociation and flow cytometric analysis
Freshly isolated tumour and PAT samples from mice were dissociated as previously described79. The samples were minced into small pieces by a scalpel, digested with an Adipose/Tumour Tissue Dissociation kit (Miltenyi Biotec) and incubated at 37 °C with rotation for 40 min. Thereafter, the samples were filtered through 40-μm nylon cell strainers, washed with FACS buffer, and centrifuged at 450g for 5 min. After removal of the supernatant, the pelleted cells were gently suspended in 40% Percoll (Cytiva) and subjected to gradient centrifugation at 900g for 20 min (Inc Spd 9, Dec Spd 3). The pelleted cells were resuspended in FACS buffer, stained with Live/Dead Fix Blue-400 (Thermo Fisher Scientific) for 15 min at room temperature. The cells were washed and stained with several antibody panels for 30 min at room temperature to identify different subsets of immune cells or stromal cells. Then the cells were fixed and permeabilized overnight at 4 °C and were washed the next day by using Foxp3/Transcription Factor Staining Buffer Set (Thermo Fisher Scientific). Next, the fixed cells were suspended in PBS with Countbright Absolute Counting Beads (Thermo Fisher Scientific). Then the cells were evaluated by CytoFLEX LX Flow Cytometer (Beckman Coulter) and the data were analysed using CytoExpert software (Beckman Coulter).
Panel 3 included anti-mouse CD45, CD3, CD4, CD8, CXCR4, T-Select H-2Kb OVA Tetramer (OVA 257–264) for the detection of CXCR4+ T cells. Panel 4 included anti-mouse CD45.1, CD3, CD4, CD8, CXCR4, T-Select H-2Kb OVA Tetramer (OVA 257-264) for the detection of CD45.1+ CXCR4+ T cells. Panel 5 included anti-mouse CD45, CD31, FAP, PDGFRB, MDK for confirming the sorting efficiency of adCAF.
Cxcl12 fl/fl mice and Mdk DTR mice models
To specifically knock out Cxcl12 in the PAT, Cxcl12fl/fl mice were treated with 5 × 1010 V.G. AAV2/9-CAG-Cre or its control AAV by injection into the inguinal adipose tissue to obtain PAT-specific Cxcl12 homozygous knockout mice. All AAVs were purchased from Taitool Bioscience. After 4 weeks, the efficiency of viral infection was determined by detecting GFP fluorescence intensity in OCT-embedded frozen sections, and the knockout efficiency of Cxcl12 in PAT was verified by qPCR and western blot.
To specifically eliminate the vast majority of adCAFs in the PAT, MdkDTR mice were injected daily with 100 ng diphtheria toxin (DT; List Biological Laboratories) into PAT for at least 7 days. For evaluating adCAF elimination efficiency, PAT was dissected and digested into a single-cell suspension and then the quantity of adCAF was detected using flow cytometry.
Patient enrolment
We enrolled 67 patients with clinically staged T3-4 or N + M0 rectal adenocarcinoma from SYSUCC, all of whom underwent neoadjuvant immuno-chemoradiotherapy. The treatment regimen consisted of (1) pre-radiotherapy: one cycle of Capeox (oxaliplatin 130 mg m−² intravenously on day 1, capecitabine 1,000 mg m−² orally twice daily on days 1–14, every 3 weeks) combined with PD-1 inhibitor (sintilimab, 200 mg intravenously on day 1, every 3 weeks). (2) Concurrent radiotherapy: two cycles of Capeox plus sintilimab (oxaliplatin dose reduced to 100 mg m−², with other parameters unchanged). (3) Post-radiotherapy: one additional cycle of Capeox with sintilimab (reverting to the initial oxaliplatin dose of 130 mg m−²). Following immuno-chemoradiotherapy, treatment response was assessed using radiological imaging, colonoscopy and clinical evaluation. Patients were closely monitored every 3–4 months to determine whether a clinical CR (cCR) was achieved. Radical surgery was performed for those without cCR. Postoperative pathological evaluation was independently conducted by two senior pathologists using the AJCC Seventh Edition Tumour Regression Grade system. PD-L1 expression in tumour samples was assessed using an IHC staining kit (containing the 22C3 antibody). Two experienced pathologists, blinded to clinical outcomes, interpreted the results following the manufacturer’s guidelines.
tVAT area analysis
MRI was performed using a 3.0-T whole-body MRI system (Signa Pioneer, GE Healthcare) with a 28-channel phased-array coil. Patients fasted and refrained from water intake for 4 h before the scan. They were positioned head-first in the supine position, with the coil centre aligned at the pubic symphysis level and adjusted based on tumour location. High-resolution MRI was acquired using a two-dimensional fast spin-echo T2-weighted sequence in three orientations: oblique axial (perpendicular to the tumour’s long axis), sagittal (parallel to the tumour’s long axis) and oblique coronal (parallel to the anal canal). The MRI parameters of oblique axial included an echo time (TE) of 88 ms, repetition time (TR) of 3,693 ms, slice thickness of 3 mm, slice spacing of 0.5 mm, field of view of 200 × 200 mm, matrix size of 256 × 320, and two excitations. Two experienced radiologists, blinded to pathological results, independently reviewed and reinterpreted the MRI findings. Using 3D Slicer software (https://www.slicer.org), they manually delineated the mesorectal fat area on the largest cross-sectional oblique axial T2-weighted image, tracing along the mesorectal fascia, while excluding the tumour and lumen, which referred to tVAT area. The final tVAT area value was calculated as the average of their measurements.
Statistics and reproducibility
No statistical method was used to predetermine sample size; however, our sample sizes are comparable with those reported in previous relevant studies80. The investigators were not blinded to group allocation during experiments and outcome assessment, except for the MRI analyses, which were performed independently by two radiologists who were blinded to the experimental groups. In animal experiments, mice were randomly allocated to experimental groups unless otherwise specified. For studies involving genetically engineered mice, group assignment was determined by genotype, with all comparison groups matched at baseline for key covariates including age and sex. No data were excluded from the analyses. Before statistical testing, data were formally assessed for normality and homogeneity of variance, and all met the assumptions of the applied tests. The statistical methods used for each analysis are described in the figure legends. In brief, a paired Student’s t-test and paired Wilcoxon test were used to determine significant differences for the comparison between two paired groups. An unpaired Student’s t-test was used to detect whether there were significant differences between two independent groups. A one-way analysis of variance (ANOVA) combined with the Tukey’s multiple comparison method was used to conduct pairwise comparisons among multiple independent groups (≥3). A two-way ANOVA combined with Tukey’s or Bonferroni’s multiple comparison method was used to perform pairwise comparisons among multiple groups with two factors. A two-way ANOVA combined with the Tukey’s multiple comparison method was used to conduct pairwise comparisons between groups of tumour growth curves. All statistical tests were two-sided and the statistical results are presented as mean ± s.d. All experiments were repeated at least three times yielding consistent results. Representative outcomes and images are presented. Statistical analyses were performed with GraphPad Prism (v.8.0.1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data of scRNA-seq, scTCR/BCR-seq and snRNA-seq data of this study are available in Genome Sequence Archive database under accession code (HRA009867). Processed bulk transcriptome data are available on Mendeley at https://doi.org/10.17632/gnmygsz5zr. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
This study has not generated any custom codes or programs. All the software and packages used in this study have been mentioned in detail in Methods.
References
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell 41, 374–403 (2023).
Bejarano, L., Jordāo, M. J. C. & Joyce, J. A. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 11, 933–959 (2021).
Huang, C. X. et al. Pericancerous cross-presentation to cytotoxic T lymphocytes impairs immunotherapeutic efficacy in hepatocellular carcinoma. Cancer Cell 42, 2082–2097.e2010 (2024).
Nieman, K. M. et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat. Med. 17, 1498–1503 (2011).
Lee, M. J., Wu, Y. & Fried, S. K. Adipose tissue heterogeneity: implication of depot differences in adipose tissue for obesity complications. Mol. Aspects Med. 34, 1–11 (2013).
Lengyel, E., Makowski, L., DiGiovanni, J. & Kolonin, M. G. Cancer as a matter of fat: the crosstalk between adipose tissue and tumors. Trends Cancer 4, 374–384 (2018).
Maniyadath, B., Zhang, Q., Gupta, R. K. & Mandrup, S. Adipose tissue at single-cell resolution. Cell Metab. 35, 386–413 (2023).
Emont, M. P. et al. A single-cell atlas of human and mouse white adipose tissue. Nature 603, 926–933 (2022).
Cypess, A. M. Reassessing human adipose tissue. N. Engl. J. Med. 386, 768–779 (2022).
Wei, G. et al. The thermogenic activity of adjacent adipocytes fuels the progression of ccRCC and compromises anti-tumor therapeutic efficacy. Cell Metab. 33, 2021–2039.e2028 (2021).
Maguire, O. A. et al. Creatine-mediated crosstalk between adipocytes and cancer cells regulates obesity-driven breast cancer. Cell Metab. 33, 499–512.e496 (2021).
Pacella, I. et al. Fatty acid metabolism complements glycolysis in the selective regulatory T cell expansion during tumor growth. Proc. Natl Acad. Sci. USA 115, E6546–e6555 (2018).
Ringel, A. E. et al. Obesity shapes metabolism in the tumor microenvironment to suppress anti-tumor immunity. Cell 183, 1848–1866.e1826 (2020).
Lyu, X. et al. Contribution of adipocytes in the tumor microenvironment to breast cancer metabolism. Cancer Lett. 534, 215616 (2022).
Nguyen, H. P. et al. Implantation of engineered adipocytes suppresses tumor progression in cancer models. Nat. Biotechnol. 43, 1979–1995 (2025).
Pelka, K. et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 184, 4734–4752.e4720 (2021).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459.e429 (2020).
Liu, Y. et al. Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell 40, 424–437.e425 (2022).
Hildreth, A. D. et al. Single-cell sequencing of human white adipose tissue identifies new cell states in health and obesity. Nat. Immunol. 22, 639–653 (2021).
Joanito, I. et al. Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer. Nat. Genet. 54, 963–975 (2022).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2022).
Helmink, B. A. et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577, 549–555 (2020).
Lynch, K. T. et al. Heterogeneity in tertiary lymphoid structure B-cells correlates with patient survival in metastatic melanoma. J. Immunother. Cancer 9, e002273 (2021).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Huang, Q. et al. The primordial differentiation of tumor-specific memory CD8(+) T cells as bona fide responders to PD-1/PD-L1 blockade in draining lymph nodes. Cell 185, 4049–4066.e4025 (2022).
Nieman, K. M., Romero, I. L., Van Houten, B. & Lengyel, E. Adipose tissue and adipocytes support tumorigenesis and metastasis. Biochim. Biophys. Acta 1831, 1533–1541 (2013).
Pond, C. M. & Mattacks, C. A. The activation of the adipose tissue associated with lymph nodes during the early stages of an immune response. Cytokine 17, 131–139 (2002).
MacQueen, H. A. & Pond, C. M. Immunofluorescent localisation of tumour necrosis factor-alpha receptors on the popliteal lymph node and the surrounding adipose tissue following a simulated immune challenge. J. Anat. 192, 223–231 (1998).
Iwamoto, H. et al. Cancer lipid metabolism confers antiangiogenic drug resistance. Cell Metab. 28, 104–117.e105 (2018).
Han, Y. K. et al. Adipocytolytic polymer nanoparticles for localized fat reduction. ACS Nano. 17, 70–83 (2023).
Cai, L., Kolonin, M. G. & Anastassiou, D. The fibro-adipogenic progenitor APOD+DCN+LUM+ cell population in aggressive carcinomas. Cancer Metastasis Rev. 43, 977–980 (2024).
Zhu, K., Cai, L., Cui, C., de Los Toyos, J. R. & Anastassiou, D. Single-cell analysis reveals the pan-cancer invasiveness-associated transition of adipose-derived stromal cells into COL11A1-expressing cancer-associated fibroblasts. PLoS Comput. Biol. 17, e1009228 (2021).
Merrick, D. et al. Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science 364, eaav2501 (2019).
Daniel, S. K., Seo, Y. D. & Pillarisetty, V. G. The CXCL12-CXCR4/CXCR7 axis as a mechanism of immune resistance in gastrointestinal malignancies. Semin. Cancer Biol. 65, 176–188 (2020).
Han, Y. et al. IL-1β-associated NNT acetylation orchestrates iron-sulfur cluster maintenance and cancer immunotherapy resistance. Mol. Cell 83, 1887–1902.e1888 (2023).
Li, J. J. et al. The liver microenvironment orchestrates FGL1-mediated immune escape and progression of metastatic colorectal cancer. Nat. Commun. 14, 6690 (2023).
Li, Y. et al. CAF-macrophage crosstalk in tumour microenvironments governs the response to immune checkpoint blockade in gastric cancer peritoneal metastases. Gut 74, 350–363 (2025).
Xiang, W. et al. Dietary fats suppress the peritoneal seeding of colorectal cancer cells through the TLR4/Cxcl10 axis in adipose tissue macrophages. Signal Transduct. Target. Ther. 5, 239 (2020).
Mukherjee, A., Bilecz, A. J. & Lengyel, E. The adipocyte microenvironment and cancer. Cancer Metastasis Rev. 41, 575–587 (2022).
Mukherjee, A. et al. Adipocytes reprogram cancer cell metabolism by diverting glucose towards glycerol-3-phosphate thereby promoting metastasis. Nat. Metab. 5, 1563–1577 (2023).
Zhang, J., Huang, D., Saw, P. E. & Song, E. Turning cold tumors hot: from molecular mechanisms to clinical applications. Trends Immunol. 43, 523–545 (2022).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550 (2018).
Bochet, L. et al. Adipocyte-derived fibroblasts promote tumor progression and contribute to the desmoplastic reaction in breast cancer. Cancer Res. 73, 5657–5668 (2013).
Wang, Z. et al. Paradoxical effects of obesity on T cell function during tumor progression and PD-1 checkpoint blockade. Nat. Med. 25, 141–151 (2019).
Wu, B. et al. Genetic ablation of adipocyte PD-L1 reduces tumor growth but accentuates obesity-associated inflammation. J. Immunother. Cancer 8, e000964 (2020).
Janssens, R., Struyf, S. & Proost, P. The unique structural and functional features of CXCL12. Cell Mol. Immunol. 15, 299–311 (2018).
Smit, M. J. et al. The CXCL12/CXCR4/ACKR3 axis in the tumor microenvironment: signaling, crosstalk, and therapeutic targeting. Annu. Rev. Pharmacol. Toxicol. 61, 541–563 (2021).
Pernot, S., Evrard, S. & Khatib, A. M. The give-and-take interaction between the tumor microenvironment and immune cells regulating tumor progression and repression. Front. Immunol. 13, 850856 (2022).
Steele, M. M. et al. T cell egress via lymphatic vessels is tuned by antigen encounter and limits tumor control. Nat. Immunol. 24, 664–675 (2023).
Choi, Y. W. et al. Senescent tumor cells build a cytokine shield in colorectal cancer. Adv. Sci. 8, 2002497 (2021).
Dong, L. et al. SPP1(+) TAM regulates the metastatic colonization of CXCR4(+) metastasis-associated tumor cells by remodeling the lymph node microenvironment. Adv. Sci. 11, e2400524 (2024).
Biasci, D. et al. CXCR4 inhibition in human pancreatic and colorectal cancers induces an integrated immune response. PNAS 117, 28960–28970 (2020).
Suarez-Carmona, M. et al. Combined inhibition of CXCL12 and PD-1 in MSS colorectal and pancreatic cancer: modulation of the microenvironment and clinical effects. J. Immunother. Cancer 9, e002505 (2021).
Katsuno, Y. et al. Chronic TGF-β exposure drives stabilized EMT, tumor stemness, and cancer drug resistance with vulnerability to bitopic mTOR inhibition. Sci. Signal. 12, eaau8544 (2019).
Ren, L. L. et al. TGF-β as a master regulator of aging-associated tissue fibrosis. Aging Dis. 14, 1633–1650 (2023).
Zhu, Q. et al. Adipocyte mesenchymal transition contributes to mammary tumor progression. Cell Rep. 40, 111362 (2022).
Saha, A. et al. Proinflammatory CXCL12-CXCR4/CXCR7 signaling axis drives myc-induced prostate cancer in obese mice. Cancer Res. 77, 5158–5168 (2017).
Su, F. et al. Progression of prostate carcinoma is promoted by adipose stromal cell-secreted CXCL12 signaling in prostate epithelium. NPJ Precis. Oncol. 5, 26 (2021).
Feig, C. et al. Targeting CXCL12 from FAP-expressing carcinoma-associated fibroblasts synergizes with anti-PD-L1 immunotherapy in pancreatic cancer. PNAS 110, 20212–20217 (2013).
Yang, K. D. et al. CAF-derived midkine promotes EMT and cisplatin resistance by upregulating lncRNA ST7-AS1 in gastric cancer. Mol. Cell. Biochem. 477, 2493–2505 (2022).
Lee, J. W. et al. Peritumoral adipose tissue features derived from [(18)F]fluoro-2-deoxy-2-d-glucose positron emission tomography/computed tomography as predictors for response to neoadjuvant chemotherapy in breast cancer patients. J. Pers. Med. 14, 952 (2024).
Ahn, H. et al. Prognostic significance of imaging features of peritumoral adipose tissue in FDG PET/CT of patients with colorectal cancer. Eur. J. Radiol. 145, 110047 (2021).
Sahin, Y. et al. Peritumoral adipose tissue density predicts the malignancy in cT1 renal masses. Urol. Res. Pract. 49, 191–197 (2023).
Chen, I. X. et al. Blocking CXCR4 alleviates desmoplasia, increases T-lymphocyte infiltration, and improves immunotherapy in metastatic breast cancer. Proc. Natl Acad. Sci. USA 116, 4558–4566 (2019).
Lu, Y. et al. CXCL12(+) tumor-associated endothelial cells promote immune resistance in hepatocellular carcinoma. J. Hepatol. 82, 634–648 (2025).
Yang, D. et al. Genetic deletion of galectin-3 inhibits pancreatic cancer progression and enhances the efficacy of immunotherapy. Gastroenterology 167, 298–314 (2024).
Wang, F. et al. Combined anti-PD-1, HDAC inhibitor and anti-VEGF for MSS/pMMR colorectal cancer: a randomized phase 2 trial. Nat. Med. 30, 1035–1043 (2024).
Xiao, W. W. et al. Effect of neoadjuvant chemoradiotherapy with or without PD-1 antibody sintilimab in pMMR locally advanced rectal cancer: a randomized clinical trial. Cancer Cell 42, 1570–1581.e1574 (2024).
Borcherding, N., Bormann, N. L. & Kraus, G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res 9, 47 (2020).
Wang, J. et al. Fibrinogen-like protein 1 is a major immune inhibitory ligand of LAG-3. Cell 176, 334–347.e312 (2019).
Liu, J. et al. The macrophage-associated LncRNA MALR facilitates ILF3 liquid-liquid phase separation to promote HIF1α signaling in esophageal cancer. Cancer Res. 83, 1476–1489 (2023).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Liu, H. et al. Integrative molecular and spatial analysis reveals evolutionary dynamics and tumor-immune interplay of in situ and invasive acral melanoma. Cancer Cell 42, 1067–1085.e1011 (2024).
Laganenka, L., Colin, R. & Sourjik, V. Chemotaxis towards autoinducer 2 mediates autoaggregation in Escherichia coli. Nat. Commun. 7, 12984 (2016).
Biswenger, V. et al. Characterization of EGF-guided MDA-MB-231 cell chemotaxis in vitro using a physiological and highly sensitive assay system. PLoS ONE 13, e0203040 (2018).
Suraneni, P. et al. The Arp2/3 complex is required for lamellipodia extension and directional fibroblast cell migration. J. Cell Biol. 197, 239–251 (2012).
Zhao, Y. et al. cis-B7:CD28 interactions at invaginated synaptic membranes provide CD28 co-stimulation and promote CD8(+) T cell function and anti-tumor immunity. Immunity 56, 1187–1203.e1112 (2023).
Li, T. et al. Methionine deficiency facilitates antitumour immunity by altering m(6)A methylation of immune checkpoint transcripts. Gut 72, 501–511 (2023).
Maudet, C. et al. Bacterial inhibition of Fas-mediated killing promotes neuroinvasion and persistence. Nature 603, 900–906 (2022).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (82372920 and 82341010), Changping Laboratory (2025C-12-02), Open Funds of State Key Laboratory of Oncology in South China (HN2023-03), Young Talents Program of Sun Yat-sen University Cancer Center (YTP-SYSUCC-0019) and the Chih Kuang Scholarship for Outstanding Young Physician-Scientists of Sun Yat-sen University Cancer Center (2023001, 2024001 and 2024009). Figures created with BioRender: Fig. 1a, Huaiqiang, J. https://biorender.com/vnv4y0b (2026); Fig. 3a,g,j, Huaiqiang, J. https://biorender.com/f9uacgq (2026); Fig. 4e,k, Huaiqiang, J. https://biorender.com/ovq2e39 (2026); Fig. 7a,k, Huaiqiang, J. https://biorender.com/e5jwcye (2026); Extended Data Fig. 3c,i,j, Huaiqiang, J. https://biorender.com/cy7rgjm (2026); Extended Data Fig. 7a, Huaiqiang, J. https://biorender.com/y65a209 (2026); Extended Data Fig. 8a,c, Huaiqiang, J. https://biorender.com/33r2gmm (2026); Extended Data Fig. 9a, Huaiqiang, J. https://biorender.com/wbcolts (2026).
Author information
Authors and Affiliations
Contributions
Q.J., R.H.X. and J.Z.L. designed the study. J.H.W., Y.Q.Z., T.T., J.Z. L., L.Y.W., R.B.W., J.H.P., T.L. and H.Y.M. collected the data. J.H.W., Y.Q.Z., Z.Y.Q., X.T.D., Y.Q.P. and J.H.P. analysed and interpreted the data. J.H.W., Y.Q.Z. and Z.Y.Q., performed the statistical analysis. J.H.W., Z.Y.Q., Y.Q.P. and H.Q.J. wrote the manuscript. W.Y., Y.Y.Z., Y.H., J.W., K.L., Z.X.L. and G.J.S. contributed to discussion and data interpretation. Y.Q.Z., Z.Y.Q. and Y.Q.P. revised the manuscript. All authors reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-cell RNA profiling unveils tissue-specific immune landscape of tVAT.
a. UMAP plots showing the expression patterns of marker genes used to define all cell types within the TME. b. Expression of representative signature genes across major clusters, highlighting key molecular markers for each cell type. c.Enriched pathways based on differentially expressed genes between tVAT and dVAT, providing insight into the functional differences between these tissue types. A two-sided hypergeometric test was used to calculate p values and Benjamini-Hochberg method was used to adjusted the p values. d. Representative images of H&E staining and immunofluorescence (IF) staining for lymphocytes in tVAT and dVAT sections from CRC patients (n = 3 patient samples per group). Scale bar = 200 μm. e. Representative images of multiplex immunohistochemistry (mIHC) staining showing various maturation states of tertiary lymphoid structures (TLSs) in tVAT sections from CRC patients (n = 4 patient samples). Scale bars = 500 μm (overall image) or 200 μm (insets).
Extended Data Fig. 2 Characteristic of lymphocytes in tVAT compared with dVAT.
a. Dot plot showing RNA expression of marker genes used to define clusters of CD4+ T cells, CD8+ T/NK cells, and B cells. b. Longitudinal tracking of the top 20 most expanded TCR clones in dVAT, Tumour, and tVAT for each patient. Each stratum represents a unique TCR clonotype, highlighted by a different colour. Coloured bands between columns represent shared clones across time points. c. Boxplot comparing TCR clonal diversity between tVAT (n = 3) and dVAT (n = 3), analysed using a two-sided paired Student’s t-test. d. Longitudinal tracking of the top 20 most expanded BCR clones in dVAT, Tumour, and tVAT for each patient. e. Boxplot comparing BCR clonal diversity between tVAT (n = 3) and dVAT (n = 3), analysed using a two-sided paired Student’s t-test. f. Heatmap showing the fraction of BCR clonotype overlap among tissue samples and their respective tumours. g. Boxplot comparing BCR overlap between tVAT (n = 3) and dVAT (n = 3) with their respective tumours, analysed using a two-sided paired Student’s t-test. The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median) for (c, e, g).
Extended Data Fig. 3 tVAT competes with tumour tissue for the infiltration lymphocytes.
a. Representative images of MC38-OVA (left) and tumour growth (right) in C57BL/6J mice (n = 5). b. Representative images of CT26 tumours (left) and tumour growth (right) in BALB/c mice (n = 5). c. Schematic diagram illustrating the working principle of ATS-GNP adipocytolysis. ATS-GNP, containing a CaCO3 core, is receptor-mediated for endocytosis. Upon reaching the acidic environment, ATS-GNP releases CO2 gas, disrupting the adipose cell membrane. d. Tumour growth of MC38 tumours in C57BL/6J, BALB/c-Nude (left), and NSG (right) mice (n = 5). e. Representative images of E0771 tumours at day 16 of the experiment in C57BL/6J mice following removal of PAT (n = 5). f. Tumour weights of E0771 tumours at day 16 in C57BL/6J mice with the removal of PAT (n = 5). g. Representative flow cytometry plots of CD45+ cells gated on live cells in MC38-OVA tumours. h. Violin plots showing the expression of CXCR7 across all cell types in CRC patients, analysed using the Kruskal-Wallis test. i. Editing strategy for constructing Cxcl12fl/fl cKO mouse. j. Experimental design for constructing Control and Cxcl12fl/fl cKO mice bearing MC38-OVA tumours near PAT. k. Western blot analysis for verifying the knockout efficiency of Cxcl12 in PAT of Control mice and Cxcl12fl/fl cKO mice. l. RT-qPCR analysis for verifying the Cxcl12 knockout efficiency in liver, spleen, uterus, and tumour tissues from Control mice and Cxcl12fl/fl cKO mice (n = 6). Data represent ≥ 3 independent experiments. P-values were calculated using two-way ANOVA with Tukey’s correction for multiple comparisons (a-right, b-right, and d), two-way ANOVA with Bonferroni’s correction for multiple comparisons (f), or a two-sided, unpaired Student’s t-test (l). Graphs display mean ± SD (a, b, d, f, l). Panels created with BioRender: c, i and j, Huaiqiang, J. https://biorender.com/cy7rgjm (2026).
Extended Data Fig. 4 Cross-tissue diversity of stromal cells in VATs, PT and PN.
a. Top-ranked markers to define the stromal cell subclusters. b. Cross-tissue diversity (top) and abundance (bottom) of stromal cells in VATs, Tumour and Normal. c. Box plot to compare the abundance of stromal cells in VATs (n = 24), Tumour (n = 70) and Normal (n = 12). P-values were calculated using a two-side Wilcoxon test. The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median). d. Relative expression levels of CAF-associated markers (FAP and COL1A1) and adipose markers (APOE and APOD) in CAFs versus adCAFs from peritumoral adipose tissue of CRC patients, analysed using the two-sided Wilcoxon test.
Extended Data Fig. 5 Tumour induces adipose mesenchymal transformation in tVAT.
a. UMAP plot of VAT-associated stromal cells in tVAT and dVAT from CRC patients. The 8 clusters, labelled by inferred cell types, are denoted by colour. b. Dot plot showing RNA expression of marker genes used to define pAC, adCAF, and APC subclusters. Circle size represents the log-normalized P-value, while colour intensity indicates the log-transformed mean expression of marker genes. c. Sorting strategy for isolating adCAFs from mouse PAT by flow cytometry. d. RNA sequencing analysis of adCAF markers in sorted adCAF-enriched stromal cells and non-adCAF stromal cells (n = 4). Statistical significance was assessed using a two-sided, unpaired Student’s t-test. e. RNA sequencing analysis of adipogenesis markers in sorted adCAF-enriched stromal cells and non-adCAF stromal cells (n = 4). Statistical significance was assessed using a two-sided, unpaired Student’s t-test. f. Representative images of the morphology of sorted adCAF-enriched stromal cells and non-adCAF stromal cells. Scale bar = 500 μm. g. Cross-tissue interactions analysis based on ligand-receptor pairs between adCAFs and tumour cells (left) and a comparison between tVAT and dVAT (right). A two-sided permutation test was used to determine the significance of pathways. h. Western blot analysis of CXCL12 protein expression in sorted adCAF-enriched stromal cells and non-adCAF stromal cells (n = 3). The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median) for (d-e).
Extended Data Fig. 6 Single-nucleus RNA profiling characterizes the stromal microenvironment and mature adipocytes of tVAT.
a. UMAP of all cells in tVAT and dVAT from 5 CRC patients, with 16 clusters labelled by inferred cell types. Major lineages included various immunocytes, VAT-associated stromal cells, adipocytes (ACs), endothelial cells (ECs), pericytes (PCs) and mesothelial cells (Mesos). b. Heatmap of representative marker genes across all cell populations. c. UMAP of 6 subsets of ACs in tVAT and dVAT from 5 CRC patients. d. Beeswarm plot showing the distribution and abundance of ACs in Nhoods between tVAT and dVAT. e. Box plot to compare the abundance of ACs in dVAT (n = 5) and tVAT (n = 5). P-values were calculated using a two-side Wilcoxon test. f. Cell-cell communication analysis based on ligand-receptor interactions (top 3) between CD8+ T cells and adipocytes, and between CD8+ T cells and VAT-associated stromal cells in tVAT. A two-sided permutation test was used to determine the significance of pathways. Only significant ligand-receptor pairs (P < 0.05) are shown. Dot size encodes the interaction score, and colour scale indicates the communication probability. g. Violin plots showing the expression of CXCL12, VAT-associated markers, and CAF-associated markers in adipocytes and VAT-associated stromal cells from CRC patients. h. UMAP of 8 subsets of VAT-associated stromal cells in tVAT and dVAT from 5 CRC patients, including APCs, pACs and adipocyte-derived cancer-associated fibroasts (adCAFs). The 8 clusters, labelled by inferred cell types, are denoted by colour. i. Beeswarm plot showing the distribution and abundance of VAT-associated stromal cell types in Nhoods between tVAT (n = 5) and dVAT (n = 5). j. Box plot to compare the abundance of VAT-associated stromal cells in dVAT and tVAT. P-values were calculated using a two-side Wilcoxon test. k. Heatmap showing the RNA expression patterns of representative marker genes across all cell populations, including CAF markers, stromal markers, APC/pAC markers, white/brown/beige fat markers, and cytokines and growth markers. The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median) for (e, j).
Extended Data Fig. 7 Tumour-derived factors induce APC to obtain adCAF characteristics.
a. Experimental flow chart for dissecting MC38 tumours from tumour-bearing mice and preparing tumour conditioned media. b. RNA sequencing detecting the RNA expression of 4 adCAF-associated transcript factors in mAPCs treated with tumour conditioned media (n = 10). The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median). Statistical significance was assessed using a two-sided, unpaired Student’s t-test. c. RT-qPCR detecting the RNA expression of adCAF-associated genes in peritumoral adipose tissue (PAT) and contralateral adipose tissue (Control) from MC38 tumour-bearing mice (n = 6). Statistical significance was assessed by a two-sided, unpaired Student’s t-test. d. RT-qPCR detecting the RNA expression of adCAF-associated TFs in PAT and contralateral adipose tissue from MC38 tumour-bearing mice (n = 6). Statistical significance was assessed by a two-sided, unpaired Student’s t-test. e. ELISA detecting the concentration of TGF-β1 in the supernatant of MC38-derived tumours (n = 3) or normal intestinal tissue (n = 3). Statistical significance was assessed using a two-sided, unpaired Student’s t-test. f. Representative images of the morphology of mouse adipose progenitor cells (mAPCs) treated with mouse TGF-β1 protein. g. Western blot analysis detecting the expression of adCAF marker genes (MDK) in mAPCs treated with mouse TGF-β1 protein at concentrations of 5 or 10 ng/mL. Panel created with BioRender: a, Huaiqiang, J. https://biorender.com/y65a209 (2026).
Extended Data Fig. 8 Clinical significance of the tumour-adipose tissue crosstalk.
a. Editing strategy for constructing MdkDTR mouse. b. Flow cytometry representative plots and bar graphs for verifying the elimination efficiency of adCAFs (FAP+PDGFRB+MDK+ stromal cells) in PAT of Control mice and MdkDTR cKO mice. c. Experimental design for combination therapy with αCXCL12 and αPD-1 in mice bearing MC38-OVA tumours near PAT. d. MC38 tumour weights of the experiment in mice treated with αCXCL12 and/or αPD-1 (n = 6). e. Flow cytometry analysis of the infiltration of immunocytes, including T cells, CD4+ T cells, CD8+ T cells, and tumour-specific CD8+ T cells in MC38-OVA tumours from the 4 treatment groups (n = 6). f. Flow cytometry analysis of the infiltration of CXCR4+ immunocytes, including CXCR4+ T cell, CXCR4+ CD4+ T cell and CXCR4+ CD8+ T cell in MC38-OVA tumours from the 4 treatment groups (n = 6). g. Boxplot of difference of tVAT area in CR and non-CR patients with T3 or T4 stage separately. The data are presented as a box-and-whisker graph (bounds of box: first to third quartile, bottom and top line: minimum to maximum, central line: median). h. ROC plot of prediction ability of tVAT area in patients with T3 or T4 stage separately. Data represent ≥ 3 independent experiments. P-values were calculated using a two-sided, unpaired Student’s t-test (b, e-g) and two-way ANOVA with Tukey’s correction for multiple comparisons (d). Graphs display mean ± SD (b, d, e-g). Panels created with BioRender: a and c, Huaiqiang, J. https://biorender.com/33r2gmm (2026).
Extended Data Fig. 9 Clinical significance of the tumour-adipose tissue crosstalk.
a. Experimental design for combination therapy with AMD3100 and αPD-1 in mice bearing MC38-OVA tumours near subcutaneous PAT. b. Representative MC38 tumour images of the mice treated with AMD3100 and/or αPD-1 (n = 5). c. Representative MC38 tumour weights of the mice treated with AMD3100 and/or αPD-1 (n = 5). d. Representative MC38 tumour growth of the mice treated with AMD3100 and/or αPD-1 (n = 5). e. Representative MC38 tumour images in MC38-OVA caecal orthotopic tumour-bearing mice treated with AMD3100 and/or αPD-1 (n = 5). f. Representative MC38 tumour weights in MC38-OVA caecal orthotopic tumour-bearing mice treated with AMD3100 and/or αPD-1 (n = 5). g. Representative MC38 Bioluminescence Images in MC38-OVA caecal orthotopic tumour-bearing mice treated with AMD3100 and/or αPD-1 (n = 5). h. Representative MC38 tumour images in MC38-OVA caecal orthotopic tumour-bearing mice treated with αCXCL12 and/or αPD-1 (n = 5). i. Representative MC38 tumour weights in MC38-OVA caecal orthotopic tumour-bearing mice treated with αCXCL12 and/or αPD-1 (n = 5). j. Representative MC38 Bioluminescence Images in MC38-OVA caecal orthotopic tumour-bearing mice treated with αCXCL12 and/or αPD-1 (n = 5). k. Flow cytometry analysis of the infiltration of immunocytes, including T cells, CD4+ T cells, CD8+ T cells, and tumour-specific CD8+ T cells in MC38-OVA tumours from 4 treatment groups (n = 5). l. Flow cytometry analysis of the infiltration of CXCR4+ immunocytes, including CXCR4+ T cell, CXCR4+ CD4+ T cell and CXCR4+ CD8+ T cell in MC38-OVA tumours from 4 treatment groups (n = 5). Data represent ≥ 3 independent experiments. P-values were calculated using a two-sided, unpaired Student’s t-test (c, f, i, k, l) and two-way ANOVA with Tukey’s correction for multiple comparisons (d). Graphs display mean ± SD (c-d, f, i, k-l). Panel created with BioRender: a, Huaiqiang, J. https://biorender.com/wbcolts (2026).
Supplementary information
Supplementary Table 1 (download XLSX )
Clinical characteristics of patients with CRC treated with neoadjuvant immuno-chemoradiotherapy at SYSUCC.
Supplementary Table 2 (download XLSX )
Marker genes (top-ranked) for the identified cell clusters in scRNA-seq analysis, associated with Fig. 5.
Supplementary Table 3 (download XLSX )
Marker genes (top-ranked) for the identified cell clusters in snRNA-seq analysis, associated with Extended Data Fig. 6.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 6.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 7.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 5 (download PDF )
Uncropped blots for Extended Data Figs. 3k, 5h and 7g.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, JH., Zheng, YQ., Qian, ZY. et al. Peritumoural adipose tissue drives immune evasion in colorectal cancer via adipose–mesenchymal transformation. Nat Cell Biol 28, 581–595 (2026). https://doi.org/10.1038/s41556-026-01885-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-026-01885-0