Abstract
Identifying centres of neo- and palaeo-endemism is crucial for defining global conservation priorities, yet data gaps hinder prioritization in megadiverse regions. Using a dated phylogeny of 16,585 native vascular plant species, we unravel spatio-temporal dynamics of China’s extant flora. Temporally, most genera originated in the Oligocene–Miocene, whereas the Pleistocene was key for speciation, tying floristic assembly to Pleistocene climatic oscillations. Spatially, mismatched phylogenetic and taxonomic endemism centres highlight complementary conservation priorities. We identify three major taxonomic endemism centres across China, with Central China, embedded within the East Asian subtropical evergreen broad-leaved forest ecoregion, emerging as a critical and overlooked conservation gap. This region harbours over 14,431 vascular plant species, including at least 2,024 endemics, but has experienced severe loss of its original natural vegetation. We therefore advocate its designation as a global biodiversity hotspot. Our multi-dimensional assessment challenges frameworks that neglect evolutionary history and underscores the global conservation importance of Central China.
Similar content being viewed by others
Main
Global biodiversity is currently collapsing at an unprecedented rate, marked by extinction rates exceeding historical baselines, widespread erosion of ecosystem services and rapid declines in genetic diversity1,2,3. In response, the Kunming–Montreal Global Biodiversity Framework commits its signatories to protecting at least 30% of Earth’s land and seas by 2030 through ecologically representative, well-connected conservation networks4,5. Achieving this goal requires filling critical gaps in biodiversity knowledge, particularly in megadiverse regions where endemism and anthropogenic threats intersect.
Identification of global biodiversity hotspots (for regions hosting ≥1,500 endemic vascular plants with ≤30% of their original natural vegetation)6 is pivotal for determining conservation priorities7,8 to guide protected area networks and ecological corridor planning9,10. Modern conservation also encompasses biodiversity dimensions beyond taxonomic distinction, including the phylogenetic (evolutionary) diversity of species11,12,13. Furthermore, distinguishing between neo-endemics (recently diverged taxa) and palaeo-endemics (ancient relicts) is crucial for interpreting regional biodiversity but remains challenging due to inconsistent temporal thresholds14,15 and limited fossil or molecular data16. Phylogenetic endemism (PE) and the Categorical Analysis of Neo- and Palaeo-endemism (CANAPE) offer quantitative solutions by incorporating evolutionary history11,17. Yet, comprehensive assessments combining taxonomic and phylogenetic endemism remain rare in megadiverse regions with persistent data gaps18,19.
China’s extensive landmass and exceptional climatic and topographic heterogeneity (Extended Data Fig. 1) support over 30,000 vascular plant species, approximately 50% of which are endemic to the country20,21. The country spans four currently recognized global biodiversity hotspots: the Himalaya, Indo–Burma, Mountains of Central Asia and Mountains of Southwest China22 (Extended Data Fig. 2). Among these, the Mountains of Southwest China hotspot exhibits the highest levels of species richness and endemism10, a pattern attributed to the combined influences of tectonic uplift and the intensification of the Asian monsoon23,24,25. These geological and palaeoclimatic processes also fostered the evolution of unique evergreen broad-leaved forests26, which starkly contrast with the deserts and savannahs prevalent in analogous subtropical latitudes elsewhere27. Despite this richness, historical gaps in taxonomy, distribution data and phylogenetic resolution have obscured accurate assessments of China’s diversity and endemism patterns. Compounding these challenges, China’s flora is threatened by spatially heterogeneous extinction drivers, from changes in land-use and vegetation structure to extreme weather events and climate change28. An evaluation of plant diversity and endemism to identify conservation gaps and align spatial priorities with global biodiversity targets is therefore urgent4.
Here we provide a comprehensive spatio-temporal assessment of China’s endemic vascular plants by integrating taxonomic and phylogenetic endemism. Using 49,488 sequences from 18,259 taxa, we reconstruct the most comprehensive time-calibrated phylogeny for China’s vascular plants to date (99% of genera, 53% species coverage). By integrating >1.4 million distribution records with this phylogeny, we identify centres of neo- and palaeo-endemism at genus and species levels and evaluate their protection under national and global frameworks. Our analysis uncovers a critically overlooked biodiversity hotspot in Central China, a 1.54-million-km2 region characterized by moderate topographic relief and centred on Hubei, Hunan and Jiangxi provinces. Under the pressure of dense human populations, this region demands urgent conservation attention to safeguard its ecological integrity and connectivity under accelerating anthropogenic changes5,28.
Results and discussion
The origin and diversification of endemic plants in China
According to our newly compiled checklist, China harbours 124 endemic genera (4.1%) and 15,942 endemic species (51.4%) of vascular plants (Supplementary Table 1; Supplementary Data 1 and 2 provide more details). Our dated phylogeny includes 3,029 genera (99.0%) and 16,585 species (53.4%) of native Chinese vascular plants, of which 117 genera and 6,265 species are endemic (Supplementary Table 2). In this Article, we generated 2,996 sequences (atpB: 608, matK: 593, matR: 620, ndhF: 580 and rbcL: 595) for 620 species and 353 genera, of which 436 species and 35 genera lacked target sequences in public databases before this work. Our updated phylogeny now leaves only 32 accepted genera native to China unsampled, often due to low representation in herbarium collections, which warrant prioritized sampling in future research (Supplementary Table 3). Our geospatial analysis reveals gaps in available molecular data from species predominantly clustered in Hengduan Mountains and adjacent areas (Supplementary Fig. 1a,b), while the Qinghai–Tibet Plateau and Xinjiang possess the highest proportion of unsampled species (Supplementary Fig. 1c), with Xinjiang showing particularly acute under-sampling of taxa endemic to China (Supplementary Fig. 1d).
Our dated phylogeny reveals consistent relationships and divergence times for major clades in comparison with previous large-scale phylogenies29,30,31 (Fig. 1 and Extended Data Figs. 3 and 4; Supplementary Discussion provides details). Notably, we find that the endemic plants are unevenly distributed across the phylogeny (Fig. 1). The top five orders with the most endemic genera are Lamiales, Asterales, Apiales, Ranunculales and Gentianales, whereas those with the highest number of endemic species are Lamiales, Ericales, Asterales, Poales and Ranunculales.
The phylogeny includes 16,585 species and 3,029 genera of vascular plants native to China (outgroups not shown). Major clades and bootstrap support values for major nodes are indicated with different colours. Bar charts illustrate the percentage of China’s endemic genera (blue bars) and species (red bars) in each order. Representative images illustrate the ten orders with the highest proportion of species endemic to China, with the ranks (1–10) corresponding to numbers on the tree.
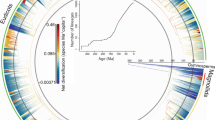
Our lineage accumulation rate (LAR) analysis, which quantifies speciation events per unit time32, reveals a prominent peak in diversification for all genera at ~19.3 million years ago (Ma) during the Early Miocene, consistent with previous studies33,34, underscoring the Miocene as a pivotal epoch for the divergence of Chinese plant genera. For endemic genera, the LAR exhibits an initial lower peak at ~46.7 Ma (Middle Eocene), followed by a more pronounced peak at ~18.9 Ma (Early Miocene) (Fig. 2a). This pattern is supported by the largest proportions of endemic genera diversified during the Miocene (37.6%) and Eocene (18.0%) (Supplementary Table 4). Endemic genera originating in the Eocene and Miocene reach their highest richness in the Hengduan Mountains (Supplementary Fig. 2a,c), highlighting the region’s pivotal role in China’s endemic flora. The earlier Eocene peak coincides with the emergence of palaeo-endemics such as Acanthochlamys, probably facilitated by geographic isolation during the India–Asia soft collision and the early uplift of the Hengduan Mountains25. The subsequent Miocene peak aligns with the rapid in situ diversification of alpine flora in the Hengduan Mountains, probably driven by intensified tectonic uplift and monsoon dynamics that increased environmental heterogeneity and climatic gradients23,24.
a,b, Number of genera (a) and species (b) that originated during specific geological timespans, with red columns showing the number of China’s endemic taxa and grey columns for non-endemic taxa. The LARs are shown with blue lines for all taxa and red lines for endemic taxa. Asterisks on the lines indicate peaks in LAR. c,d, The inset violin plots illustrate comparisons in age distribution between all, endemic and non-endemic taxa. Horizontal lines in the violin plots represent medians, bottom and top edges of boxes represent first and third quartiles and whiskers extend to 1.5 times the interquartile range. Sample sizes (n): all genera = 2,725, endemic genera = 101, non-endemic genera = 2,624, all species = 14,925, endemic species = 5,748, non-endemic species = 9,177. Plc., Palaeocene; Olig., Oligocene; Plio., Pliocene; Plt., Pleistocene.
The LAR reveals a congruent trend for all species and endemic species, each exhibiting a pronounced Pleistocene diversification peak (~2.3 Ma and ~2.1 Ma, respectively; Fig. 2b), highlighting the Pleistocene as a critical epoch for speciation of China’s extant plant diversity. The Pleistocene climate was marked by repeated glacial–interglacial cycles35, with glacial phases driving plant populations southward or to lower altitudes on mountains, while interglacial intervals enabled northward or upslope range expansions35,36. Species originating during the Pleistocene are primarily concentrated in southern China, including the Hengduan Mountains in the southwest and the Daba, Dalou and Nanling Mountains in the southeast (Supplementary Fig. 2g,h). The exceptionally high species endemism in the Hengduan Mountains is closely linked to continuous and rapid in situ diversification since the mid-Miocene24, exemplified by spectacular alpine radiations in major clades such as Rhododendron37 and Corydalis38. Moreover, growing evidence suggests that the low- to mid-elevation mountains in southeastern China acted as crucial Pleistocene refugia36,39,40 and evolutionary ‘pumps’. During glacial–interglacial cycles, population contractions and expansions facilitated isolation, secondary contact and hybridization among refugial populations, promoting lineage divergence and speciation41,42,43,44,45.
Pleistocene climatic dynamics are inferred to have spurred rapid diversification of young, geographically restricted species, as median divergence age of endemic species is significantly younger than widespread species (4.8 Ma vs 7.2 Ma, P < 0.001; Fig. 2d). This pattern highlights the profound and lasting influence of Pleistocene climatic oscillations on the assembly of China’s modern floristic diversity, in the aftermath of geological and climatic events at global and regional levels that set the stage for diversification46. Crucially, these insights are uniquely accessible through a comprehensive, species-level reconstruction of the Tree of Life, underscoring the value of phylogenetic frameworks in deciphering evolutionary drivers31.
Conservation implications of endemism centres
China’s endemic vascular plant richness shows a strong positive correlation with total richness at both genus and species levels (Supplementary Fig. 3). Spatially, China’s endemic species share a similar diversity pattern with all species that peaks in southwestern China, particularly the Hengduan Mountains and adjacent regions (Extended Data Fig. 5 and Supplementary Fig. 4b). In contrast, China’s endemic genera exhibit a markedly distinct distribution of diversity, concentrated in the mountainous areas surrounding the Sichuan Basin, including the Hengduan and Daba mountains (Extended Data Fig. 6 and Supplementary Fig. 4a). This discrepancy is probably attributed to the limited proportion of endemic genera, approximately 4% of all vascular plant genera native to China (Supplementary Table 1).
We identify taxonomic centres of neo- and palaeo-endemism as the top 5% richest grid cells within the oldest and youngest quartiles of taxa (Extended Data Fig. 7), following a threshold widely applied in large-scale biodiversity studies16,33,47. Both genus- and species-level analyses consistently reveal three major endemism centres: (1) the Hengduan Mountains, (2) Central China and (3) Yunnan–Guizhou–Guangxi boundary region (Fig. 3a,b; Supplementary Table 5 provides area, richness and endemism data). At the genus level, the two eastern centres (Central China and Yunnan–Guizhou–Guangxi boundary region) are dominated by palaeo-endemic genera, whereas the Hengduan Mountains centre predominantly harbours neo-endemic genera (Fig. 3a). In contrast, species-level endemism centred in the Hengduan Mountains exhibits a mixed distribution of both neo- and palaeo-endemism (Fig. 3b).
a,b, Taxonomic endemism centres based on the top 5% criterion at the genus (a) and species (b) levels: (1) the Hengduan Mountains, (2) Central China and (3) Yunnan–Guizhou–Guangxi boundary region. Grid cells in pink represent centres of the youngest quartile, blue the oldest quartile and purple mixed centres of the two types. c,d, Phylogenetic endemism centres identified by the CANAPE analysis at the genus (c) and species (d) levels. Grid cells in pink represent centres of neo-endemism, blue centres of palaeo-endemism and purple centres of mixed-endemism. The donut charts show the proportion of grid cells occupied by different types of endemism centres. Current national nature reserves are highlighted in dark green, while four global biodiversity hotspots are highlighted in brown. Numbers above the maps denote taxa involved in the analyses. The national boundary layer was downloaded from the Standard Map Service website (approval number GS(2019)1823; http://bzdt.ch.mnr.gov.cn/browse.html?picId=%224o28b0625501ad13015501ad2bfc0256%22; accessed October 2022).
To validate the robustness of our quartile-based classification, we applied an alternative approach by categorizing neo- and palaeo-endemic taxa using fixed temporal thresholds (23 Ma for genera and 5 Ma for species; Extended Data Fig. 8a–d), according to previous studies16. This method yields comparable patterns of endemic richness and endemism centres (Extended Data Fig. 8e,f) to those derived from the quartile approach (Fig. 3a,b). Moreover, centres based on the top 5% richness (and top 5% proportions) of all China’s endemic species with distribution data available (13,724 species) also cluster in the Hengduan Mountains, Central China and Yunnan–Guizhou–Guangxi boundary region (Supplementary Fig. 5). This congruence reinforces the robustness of taxonomic endemism centres identified based on 39.2% of China’s endemic species (6,248 species) with both divergence age estimates and distribution data available (Fig. 3b).
Our genus-level CANAPE analyses of Chinese vascular plants identify centres of phylogenetic endemism across southern China (including the coastal regions of South China, Yunnan, Hainan, Taiwan and the Himalayas), northwestern China (Tianshan–Altai Mountains) and northeastern China (Changbai Mountains) (Fig. 3c), consistent with a previous angiosperm analysis12. Phylogenetic endemism patterns at the species level align broadly with the genus-level results, except for a notable expansion of neo-endemism centres in the Hengduan Mountains (Fig. 3d). This discrepancy is consistent with previous studies that genus-level CANAPE analyses tend to underestimate neo-endemism18,48. Such underestimation arises because large genera, particularly those with rapid evolutionary radiations, are treated as single operational units in genus-level analyses, obscuring recent species-level diversification. Notably, the phylogenetic approach often reveals endemism centres influenced by ‘border effects’12, particularly in countries such as China where geopolitical boundaries ‘fragment’ natural distributions. To minimize this bias, we applied the CANAPE analysis exclusively to China’s endemic taxa, which confirms the robustness of species-level phylogenetic endemism centres (Fig. 3d and Extended Data Fig. 9b). The observed discrepancies between endemic and all genera analyses largely arise from the limited number of endemic vascular plant genera in China (Fig. 3c and Extended Data Fig. 9a).
The disparities between centres of taxonomic and phylogenetic endemism underscore their complementary conservation implications. Taxonomic endemism centres highlight regions rich in taxa restricted to China, many of which face heightened vulnerability from habitat degradation and climate change49. Protecting these areas is critical for both national and global conservation strategies, as the loss of these taxa would represent an irreplaceable reduction of planetary biodiversity50,51. In contrast, phylogenetic endemism centres reflect over-representation of lineages with restricted ranges in China, even if some taxa extend beyond the national boundaries. While geopolitical boundaries may obscure geographic endemism patterns, they remain critical for conservation, as efforts to protect such taxa rarely receive equal attention across political boundaries, particularly in regions with stark socio-economic disparities52,53. Moreover, the substantial spatial incongruence between taxonomic and phylogenetic endemism centres demonstrates that neither metric alone captures the full breadth of biodiversity, limiting their effectiveness in isolation. Integrating both dimensions is therefore essential for comprehensive conservation planning and ensuring biologically unique areas in megadiverse regions are not overlooked.
Several methodological limitations in the identification of endemism centres should be acknowledged. First, while our species sampling is broadly representative, it remains incomplete, especially in the Qinghai–Tibet Plateau and Xinjiang of western China (Supplementary Fig. 1), which may hinder precise delineation of neo- and palaeo-endemism centres in these regions. Second, geographic distribution data, primarily derived from public databases and herbarium collections, are often spatially biased towards more accessible or well-surveyed areas. Third, phylogenetic reconstruction and dating, based on a limited number of molecular loci, may introduce uncertainties, particularly for deep divergences and rapid radiations. Despite these limitations, our core conclusions regarding spatio-temporal patterns and conservation priorities remain robust. This robustness is further supported by the consistent patterns of both taxonomic and phylogenetic endemism centres across ten randomly selected bootstrap trees (Supplementary Figs. 6 and 7). These patterns suggest that our findings are not artefacts of phylogenetic uncertainty. Future studies that incorporate broader species sampling, more geographically balanced distribution data and high-resolution genomic datasets will further refine these findings.
An ignored global biodiversity hotspot in Central China
We identify conservation gaps across China by overlaying endemism centres with national nature reserves and previously recognized global biodiversity hotspots. A pronounced spatial imbalance in conservation efforts emerges along the Hu Huanyong Line, a demarcation of China’s east–west population divide54,55. West of this line, national nature reserves are fewer in number but larger and more contiguous (n = 78, total area: 889,410 km2), whereas the densely populated east hosts a higher density of smaller, fragmented reserves (n = 346, total area: 103,417 km2). This fragmentation, reflecting intense anthropogenic pressures, including land-use conversion for urban, agricultural and infrastructural development10,47, directly challenges implementation of the Kunming–Montreal Global Biodiversity Framework’s target for maintaining ecosystem connectivity4,5. Globally, western China’s endemism centres intersect with three global biodiversity hotspots, Mountains of Central Asia, Himalaya and Mountains of Southwest China, whereas eastern China’s tropical–subtropical zones show only limited overlap with the Indo–Burma hotspot (Fig. 3 and Extended Data Fig. 2). Strikingly, Central China emerges as a critical conservation gap in both national and global frameworks (Figs. 3 and 4). To address this gap, we present updated, comprehensive taxonomic checklists (Supplementary Data 3 and 4) and advocate the formal recognition of Central China as a global biodiversity hotspot.
The newly proposed Central China hotspot (light green) encompasses taxonomic endemism centres (yellow grid cells) at genus and species levels using quartile and temporal threshold criteria. Brown shading shows partial overlap with four recognized global biodiversity hotspots in China, and dark green indicates current national nature reserves. The Hu Huanyong Line (black dashed) demarcates China’s east–west population divide, extending from Heihe City (Heilongjiang Province) to Tengchong City (Yunnan Province). Orange curves indicate major mountain ranges in Central China: (1) Qinling, (2) Daba, (3) Huaying, (4) Dalou, (5) Wuling, (6) Miaoling, (7) Xuefeng, (8) Luoxiao, (9) Nanling, (10) Wuyi, (11) Tianmu and (12) Dabie. Representative vascular plant genera endemic to Central China are illustrated on the right. The national boundary layer was downloaded from the Standard Map Service website (approval number GS(2019)1823; http://bzdt.ch.mnr.gov.cn/browse.html?picId=%224o28b0625501ad13015501ad2bfc0256%22; accessed October 2022).
Myers et al. have previously noted several regions of exceptional endemism under threat, including southeastern China, whose hotspot status remains unresolved due to data limitations6,22. Robust data foundations now exist for revising China’s hotspot system, built over two decades through compilation of national/regional checklists and floras20, digitization of >8.53 million specimens (Chinese Virtual Herbarium, http://www.cvh.ac.cn/) and updated regional phylogenies29,56. Building on this major increase in data availability and the results from previous analyses, we propose Central China (24–34° N and 103–122° E; encompassing much of southeastern China) as a biodiversity hotspot spanning 1.54 million km2, broadly corresponding to Takhtajan’s Central China Floristic Province57 (Supplementary Note provides details of geographic delineation).
Comprehensive data analyses confirm that Central China meets the strict criteria for hotspot designation6 (Supplementary Tables 6 and 7). Comparisons with the four recognized subtropical hotspots further highlight its exceptional plant diversity and extensive vegetation loss (Table 1). This region harbours extraordinary biodiversity and endemism, hosting at least 14,431 vascular plant species, 2,024 of which are endemic to this region (Supplementary Table 6 and Supplementary Data 4). A parallel assessment based on coordinate data from the Global Biodiversity Information Facility suggests that the number of endemic species may be as high as 2,158 (Supplementary Data 5). Whereas the exact figures require further validation, current data place Central China sixth in total vascular plant species richness and 27th in endemics among the 36 recognized global biodiversity hotspots22,58,59. However, as one of China’s most densely populated areas (east of the Hu Huanyong Line), it faces intense land-use pressures and has experienced extensive habitat degradation55, with over 93% of its original vegetation lost (Extended Data Fig. 10 and Supplementary Table 7). Protected areas currently cover just 7% of the region, with national nature reserves and national parks accounting for only 3% (Supplementary Table 8).
Situated within the unique East Asian subtropical evergreen broad-leaved forest ecoregion (Fig. 4 and Supplementary Fig. 8), Central China serves as both a museum and a cradle of biodiversity16,33,40. This dual role is exemplified by its unparalleled concentration of palaeo-endemic genera (Fig. 3a and Extended Data Fig. 8e), including ‘living fossils’ such as Ginkgo, Metasequoia and Davidia34,60,61, alongside a mixture of neo- and palaeo-endemic plant species (Fig. 3b and Extended Data Fig. 8f). The region is also a critical hub for insect and vertebrate diversity62,63, providing refuge for flagship conservation species including the giant panda and snub-nosed monkey64,65. Amphibians, the most threatened vertebrate class (40.7% of species globally threatened)66, further exemplify the region’s richness and endemism, with over 100 regionally endemic species documented (Supplementary Table 6). However, six key amphibian diversity centres in this region remain inadequately protected47, reflecting systemic conservation gaps spanning both flora and fauna.
Alarmingly, accelerating climate change now represents the greatest extinction risk to the region’s flowering plants28, compounding the existing pressures from dense human populations and fragmented landscapes. Addressing protection gaps in this complex socio-ecological region requires a multi-path strategy that integrates spatial, institutional and socio-economic approaches. First, remaining intact patches of native vegetation should be promptly incorporated into high-level conservation instruments, such as the national park system, in line with China’s commitment to deliver on the ‘30 × 30’ protection target of the Kunming–Montreal Global Biodiversity Framework4,67, to ensure strict land-use regulation and enhance cross-jurisdictional coordination68. Priority should be given to formally establishing national park pilots in key areas, such as the Daba Mountains (Shennongjia pilot) in the north and the Nanling Mountains (Nanling pilot) in the south (Extended Data Fig. 10). Second, in heavily human-dominated landscapes, fine-scale conservation interventions led by local governments, such as plant micro-reserves69, stepping-stone habitats70 and a range of Other Effective Area-Based Conservation Measures71 (for example, community-managed forests, traditional sacred natural sites72 and biodiversity-friendly agricultural mosaics), can enhance functional connectivity and safeguard range-restricted endemic species67. Third, socio-economic mechanisms, including ecological compensation, community co-management and livelihood diversification, are vital for reducing conservation-development conflicts and fostering long-term stewardship73,74. These should be considered by both the public and private sectors, such as through the establishment of a science-based market for high-integrity biodiversity credits to support conservation and restoration efforts75. Finally, recognizing Central China as a global biodiversity hotspot during the upcoming revision of the framework announced by the International Union for Conservation of Nature76 would elevate the region’s international conservation priority and guide targeted, context-appropriate planning77, advancing conservation action and supporting China’s sustainability ambitions.
Methods
Taxon sampling and phylogenetic reconstruction
We used sequences of five loci, including four chloroplast genes (atpB, matK, ndhF and rbcL) and one mitochondrial gene (matR) for phylogenetic reconstruction. Our phylogeny was built upon two early versions of Chinese vascular plant tree of life. The first version included 93% genera and 19% species56 and the second version encompassed 96% genera and 44% species29. Compared with the second version phylogeny by Hu et al.29, we newly added 10,023 sequences representing 3,267 species and 996 genera, of which 2,996 sequences (representing 620 species and 353 genera) were newly sequenced in this study and 7,027 (representing 2,727 species and 858 genera) were newly downloaded from GenBank.
Initially, we downloaded all target sequences for approximately 2,727 species from GenBank uploaded after Hu et al.29 (accessed September 2022). Low-quality sequences were identified and replaced or removed through phylogenetic tree reconstruction. For species with multiple sequences of the same locus, we retained the longest one. To address gaps in public databases, we conducted targeted field surveys to collect taxa lacking available sequences. Voucher specimens for newly sampled taxa were deposited at the Herbarium of Institute of Botany, Chinese Academy of Sciences (PE). We newly generated a total of 2,996 sequences for 620 species (92.6% are species endemic to China), most of which were either absent from GenBank or of low quality (for example, shorter than 200 bp). Of these, 436 species had no molecular data previously available in GenBank and the remaining 184 species were replaced with newly generated, higher-quality sequences.
Our phylogeny has sampled all orders and families of vascular plants native to China, with the sole exception of Corsiaceae, represented by the presumably extinct genus Corsiopsis78. At the genus level, previous studies lacked molecular data for 133 genera29, many of which are monotypic, oligotypic or narrowly distributed and have been historically underrepresented in herbarium collections. We newly incorporated 70 of these genera into our phylogeny (Supplementary Table 9) and evaluated taxonomic statuses of the remaining unsampled genera. Only 32 accepted genera native to China remain absent from the phylogeny (Supplementary Table 3), highlighting critical targets for future research. Additionally, we examined the spatial distribution of molecular data coverage by mapping the richness and proportion of taxa lacking target molecular loci within grid cells to locate regional data gaps.
Total genomic DNA was extracted from silica-gel dried leaf materials and sequenced on the Illumina HiSeq and DNBSEQ-T7 platforms by Beijing Novogene Bioinformatics Technology Co. Ltd. The raw sequencing data yielded ~10 Gb of 150 bp paired-end reads per sample. For all newly sampled species, full-length reference sequences for each target gene from the same family were downloaded from GenBank as the target file. Coding sequences of the five genes were extracted from the whole-genome sequencing reads using the hybpiper assemble command of the Hybpiper79 pipeline v.1.3.1. Voucher information for newly sampled species and corresponding GenBank accession numbers are provided in Supplementary Data 6. To validate identification for species with newly generated sequences, we used the BLAST tool provided by the National Center for Biotechnology Information (NCBI) and checked morphology of specimens. The newly obtained sequences were automatically aligned using MAFFT80 v7.508 and then manually checked in BioEdit81.
Incorporating newly generated sequences with alignments from Hu et al.29, our final concatenated matrix included 18,259 taxa and 49,488 sequences (7,844 atpB, 15,666 matK, 1,843 matR, 8,326 ndhF and 15,809 rbcL). One hornwort (Anthoceros angustus), two liverworts (Pellia endiviifolia and Aneura mirabilis) and two mosses (Syntrichia ruralis and Physcomitrium patens) were selected as outgroup taxa following Chen et al.56. Each gene in the concatenated matrix was treated as a separate partition. To ensure taxonomic accuracy, we conducted preliminary maximum likelihood analyses and iteratively validated species placements against established classifications such as the Angiosperm Phylogeny Group classification (APG) IV, the Pteridophyte Phylogeny Group classification (PPG) I and Flora of China20,82,83. Sequences yielding phylogenetically incongruent positions were either replaced or removed until all placements were resolved as taxonomically consistent. Final phylogenetic reconstruction was performed in RAxML84 8.2.12 under the GTRGAMMA model, with 100 bootstrap replicates. The resulting tree included 18,259 tips, representing 17,853 species and 3,288 genera, of which 16,585 species (53%) and 3,029 genera (99%) are native to China. Monophyly assessments revealed six non-monophyletic families and 692 non-monophyletic genera (Supplementary Data 7; Supplementary Discussion provides details). The phylogenetic tree was visualized using iTOL85 (https://itol.embl.de/).
Divergence time estimation
Divergence times were estimated using the penalized likelihood (PL) approach implemented in treePL86, a computationally efficient and widely adopted method for dating large phylogenies31,33. We incorporated a total of 220 calibration points covering major clades (Extended Data Fig. 3; Supplementary Data 8 provides calibration sources). The maximum and minimum crown ages were constrained to 454–416 Ma for vascular plants, 366.8–318 Ma for seed plants and 245–136 Ma for flowering plants, following previous studies87,88. We dated the best maximum likelihood (ML) tree with the optimal parameters (opt = 2, optad = 2, optcvad = 0 and smooth = 1 × 10−10). To quantify uncertainty, confidence intervals for node ages were calibrated based on 100 bootstrap trees, which were constrained using topology of the best ML tree allowing branch lengths to vary, following Magallón et al.89. Divergence times for the 100 bootstrap trees were calculated using the same treePL parameters. Statistical distributions of ages for all nodes (for example, median, maximum and minimum) were summarized in TreeAnnotator v.2.7.590.
To cross validate our divergence time estimates, we compared node ages at family and genus levels with those from two recent global macroevolutionary studies30,31. Whereas these studies employed different maximum age constraints for angiosperms, we focused on their estimates derived under constraints comparable to our own calibration (crown age constraint for angiosperms: max = 256 Ma or 247 Ma) to ensure comparability. We used Spearman’s rank correlation coefficient to evaluate concordance between estimates. The results revealed broad temporal concordance across clades, particularly at the genus level, suggesting robustness of our divergence time estimates.
Compilation and revision of vascular plants endemic to China
The list of China’s endemic vascular plant genera was compiled primarily from ref. 91, applying a strict definition of taxonomic endemism that excluded genera with updated distribution records beyond China. Scientific names for these genera were standardized according to Catalogue of Life China (Species 2000 China Node, http://www.sp2000.org.cn/) and Plants of the World Online (https://powo.science.kew.org). Discrepancies between the two databases were resolved through expert consultation to ensure taxonomic consistency. After excluding synonyms, 124 genera (4.1% of China’s native vascular plant genera) were retained (Supplementary Data 1).
For endemic species, we integrated datasets from Huang et al.92 for seed plants and Zhou et al.93 for lycophytes and ferns, resulting in 15,942 endemic species (Supplementary Data 2). These represent 51.4% of China’s vascular plant diversity. Our phylogenetic analysis included 117 endemic genera (94.4% of total endemic genera) and 6,265 endemic species (39.3% of total endemic species), covering all major clades of vascular plants (Extended Data Fig. 3). To evaluate endemism patterns across lineages of the phylogenetic tree, we quantified endemic genera and species per plant order and calculated their proportional representation within the total endemic taxa.
Distribution data assemblage
Geographic distribution records of vascular plants in China were assembled from two primary sources: the gridded distribution data for angiosperms from ref. 28 and county-level records for gymnosperms, lycophytes and ferns from the Chinese Virtual Herbarium (accessed December 2021). Circumscription of families followed PPG I for lycophytes and ferns83, Christenhusz et al. for gymnosperms94 and APG IV for angiosperms82. To ensure data quality, newly acquired records were cleaned and validated based on protocols from refs. 28,33, including (1) standardizing taxonomic names according to Flora of China20, (2) removing cultivation/non-native occurrences, (3) correcting county names, (4) combining infraspecific taxa records to corresponding species and (5) excluding duplicate records in each county. The angiosperm and non-angiosperm datasets were merged and mapped onto a standard map of China (review drawing number: GS(2019)1823), which was divided into 1,155 grid cells of 100 km × 100 km under the Albers equal-area projection. Grid cells <50% land area (that is, <5,000 km2) were excluded, resulting in a final dataset of 1,421,390 records across 941 grid cells, which encompassed 26,604 species (13,725 endemic) and 2,873 genera (116 endemic) native to China. After matching phylogeny with the distribution database, we obtained a total of 1,112,348 records for 14,906 species for downstream phylogenetically based analyses.
Temporal diversity pattern analyses
Divergence times for endemic and non-endemic vascular plant genera and species in China were extracted from a time-calibrated phylogeny (chronogram). To mitigate biases from incomplete sampling, we used the stem ages (rather than crown ages) for genera. For monophyletic genera, the stem node age was directly extracted. For non-monophyletic genera, the stem age of the largest clade within the genus was used95. For species, divergence times were derived from the terminal tip ages of the chronogram96. To account for phylogenetic uncertainty, ages for each genus and species were calculated as the median values across 100 time-calibrated trees with a fixed topology.
We quantified the number of endemic and non-endemic genera that originated during each five-million-year period since the Jurassic, whereas species-level analysis was conducted in two-million-year intervals during the Cenozoic. We also documented the number of genera, species, endemic genera and endemic species originated in each geological epoch (Supplementary Table 4). Age distributions for all, endemic and non-endemic taxa were log10-transformed and compared using the Kruskal–Wallis and Wilcoxon rank sum test. To reduce influence of outliers, we only included data within the 5th–95th percentiles range of ages for statistical analyses.
To reconstruct the diversification dynamics of endemic vascular plants in China, we estimated the lineage accumulation rates (LARs) at genus and species levels since the Cenozoic using the Julia package Biohistoria.jl97 following a previous study32. LAR quantifies speciation events per unit time by summing the probability density functions of the age distributions pointwise over all lineages. Compared to traditional indices, LAR carries additional statistical information beyond means, medians and confidence intervals32.
Spatial diversity pattern analyses
We quantified taxonomic richness for both endemic and non-endemic taxa within each grid cell as the total number of genera or species, using the specnumber function in the R package vegan98. We also calculated richness patterns separately for the youngest 25% and oldest 25% of genera and species. Because regions with high endemism do not necessarily coincide spatially with those of high overall diversity99, we assessed their spatial congruence by calculating the Pearson correlation coefficient in R100 between endemic richness and total richness at both genus and species levels across grid cells.
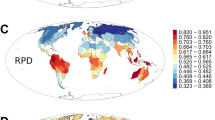
We used the R package canaper101 to calculate phylogenetic metrics, conduct randomization tests and perform statistical analyses. These analyses were performed separately for all genera/species and endemic genera/species. We calculated three phylogeny-based diversity metrics, including phylogenetic diversity (PD), phylogenetic endemism (PE) and relative phylogenetic endemism (RPE). PD was calculated as Faith’s PD by summing the total branch length connecting all species within each grid cell102. PE was calculated as the sum of branch lengths connecting species present in a grid cell, weighted by the inverse of range size of their descendant species, to quantify the geographic concentration of evolutionary history17. RPE was defined as the ratio of PE calculated on the original phylogenetic tree to PE calculated on a comparison tree with identical topology but equal lengths for all branches11. This ratio distinguishes areas dominated by palaeo-endemism (rare, long branches) from those dominated by neo-endemism (rare, short branches). To ensure comparability, branch lengths were scaled as proportions of the total tree length before calculating PD and PE, resulting in values between 0 and 1.
To identify grid cells with statistically significant patterns of RPE, we conducted randomization tests using 999 iterations of the ‘curveball’ algorithm implemented in the canaper package101. This algorithm generates null matrices by randomizing species occurrences within each grid cell while maintaining both the observed species richness per cell and the geographic range size of each species103. We then performed a two-tailed test (α = 0.05) against this null model. Grid cells were classified as significantly higher or lower than expected if the observed value fell above 97.5% or below 2.5% of the randomized values. Grid cells not meeting these thresholds were deemed consistent with patterns expected under random conditions11.
Identification of taxonomic endemism centres
We applied the quartile method to identify centres of taxonomic endemism and distinguish between young and old endemism33. We first ranked all endemic taxa (genera and species) by their estimated evolutionary age, from youngest to oldest. We then divided this ranked list into quartiles. For each grid cell, we calculated the richness (number of taxa) within the youngest 25% and oldest 25% of endemic taxa separately. We defined centres of taxonomic endemism as grid cells falling within the top 5% richness values for either the youngest quartile or the oldest quartile. Cells ranking in the top 5% for both quartiles were classified as mixed centres, while those only in the top 5% for the youngest quartile were young centres, and those only in the top 5% for the oldest quartile were old centres. Adjacent grid cells identified as centres were subsequently aggregated into major endemism centres based on geographic proximity and prior biogeographic knowledge104. Finally, we calculated the proportional representation of young, old and mixed centres at both the genus and species levels. To evaluate the impact of phylogenetic uncertainty on our results, we repeated the identification of quartile-based endemism centres using ten trees randomly selected from a set of 100 bootstrap trees.
To assess the robustness of our quartile-based classification of young and old endemism centres, we implemented an alternative temporal threshold method following established approaches16,91. Selecting appropriate thresholds to distinguish palaeo-endemics from neo-endemics is critical. At the genus level, we adopted 23 Ma, corresponding to the Palaeogene–Neogene boundary, as the threshold for palaeo-endemics and neo-endemics. Nearly all Chinese endemic gymnosperm genera originated in the Cretaceous or earlier (palaeo-endemics). For angiosperms, palaeo-endemic genera mainly arose in the Late Cretaceous–Palaeogene, consistent with evidence that many relict lineages (for example, Davidia, Eucommia) arose during this interval and later contracted to China91. In contrast, neo-endemic genera mostly emerged during the Neogene, matching recent large-scale phylogenetic studies that identify the Miocene as the primary diversification epoch for China’s plant genera33,34,95.
At the species level, the threshold distinguishing palaeo- from neo-endemics remains debated16. Nonetheless, given that the majority of China’s endemic species are concentrated in the Hengduan Mountains, a region that experienced rapid uplift and intensified monsoon influence during the late Miocene to Pliocene105,106, this interval represents a critical phase of accelerated in situ diversification23. Moreover, recent studies of the Sino-Himalayan flora highlight ~5 Ma as a key turning point, marked by rapid radiations that generated a substantial number of young, range-restricted species96. Therefore, we set the species-level boundary at 5 Ma.
Using these thresholds, we calculated richness patterns for old (>23 Ma) and young (<23 Ma) genera and old (> 5 Ma) and young (< 5 Ma) species, identifying endemism centres based on the top 5% richness criterion. The resulting richness patterns and endemic centres were broadly consistent with those derived from the quartile method. Consequently, we present the quartile-based results in the main text, with the temporal threshold method results provided in Extended Data.
While our quartile method used estimated lineage ages to classify species as old and young, only 6,248 species endemic to China (39.2%) have both divergence age estimates and distribution data available. To address potential impacts of this incomplete sampling on identifying endemism centres, we utilized distribution records for a larger subset: 13,724 endemic species (86.1%). We calculated the species richness and endemic proportion for each grid cell. Grid cells within the top 5% for each metric were identified as endemism centres. Comparing these results with those derived from the phylogenetically incomplete dataset confirmed the robustness of our analysis, specifically validating Central China’s status as a taxonomic endemism centre.
Identification of phylogenetic endemism centres
We identified phylogenetic endemism centres using the Categorical Analysis of Neo- and Palaeo-endemism (CANAPE)11. For species-level analysis, we directly utilized the species tree and corresponding distribution data. For genera exhibiting non-monophyly, we treated each monophyletic lineage as an independent operational unit in genus-level analysis12. The CANAPE workflow comprised two steps. First, we identified grid cells with significantly high PE on the actual tree (numerator of RPE), PE on a comparison tree with equal branch lengths (denominator of RPE) or both significantly higher than expected (observed value > 95% of randomized values; one-tailed test, α = 0.05). Second, grid cells meeting the above criteria were categorized into three types: if a grid cell has a significantly high or low RPE ratio (two-tailed test, α = 0.05), it is classified as a centre of palaeo-endemism or neo-endemism, respectively. If it is significantly high in both the numerator and the denominator (taken alone) but not significant for RPE, it is classified as a centre of mixed endemism11. Thus, neo-endemic centres cluster recent radiations (short branches), palaeo-endemic centres cluster narrow-range relictual lineages (long branches) and mixed centres contain both. Additionally, we repeated the CANAPE workflow using ten trees randomly selected from the 100 bootstrap trees to assess the impact of phylogenetic uncertainty on identifying phylogenetic endemism centres.
As CANAPE relies on range-weighted phylogenetic endemism17, restricted sampling to China may inflate PE values near borders where taxa extend beyond our dataset (‘border effect’)12. To evaluate this bias, we replicated analyses using strictly Chinese endemic taxa. We extracted the phylogenetic subtrees and distribution data subsets (116–genus; 6,248–species) for analyses based on the list of Chinese endemic vascular plants. The subtrees were extracted using the drop.tip function in the ape package107 in R100, whereas the distribution data subsets were filtered using a series of functions from the dplyr package108. These endemic datasets, unaffected by cross-border ranges, provide more accurate range-size estimates for PE calculations. The CANAPE workflow was reapplied identically to these subsets.
Mapping biodiversity hotspots and gaps in protection
To assess protection status of taxonomic and phylogenetic endemism centres, we overlaid these centres with current protected areas to identify conservation gaps. We used spatial data of protected areas primarily from ref. 33, focusing exclusively on national nature reserves. After excluding non-terrestrial reserves, a total of 424 nature reserves were included for downstream analysis. Furthermore, recognizing the growing emphasis on China’s national park system10,109, we incorporated both officially established national parks and pilots specifically within the Central China region when calculating protected area coverage. The list of national parks was sourced from the Chinese National Forestry and Grassland Administration (http://www.forestry.gov.cn/main/5960/index.html). We further assessed the east–west distribution disparity of protected areas using the Hu Huanyong Line, a seminal demographic geographic boundary dividing China into densely populated southeastern and sparsely populated northwestern regions54. The number and total area of national nature reserves on each side were calculated. For reserves intersecting the line, classification was based on the side containing the larger area.
Globally recognized biodiversity hotspots represent critical conservation priorities6. To identify conservation gaps of global importance, particularly potential hidden hotspots, we overlaid four global biodiversity hotspots involving China (that is, Mountains of Central Asia, Indo–Burma, Mountains of Southwest China and Himalaya)58 with our identified endemic centres. Spatial data for global biodiversity hotspots were downloaded from ref. 110, which provides vector layers for all 36 global hotspots. Our gap analysis revealed that Central China, a distinct floristic unit, receives inadequate global conservation attention. We subsequently delineated its geographic boundaries and quantified its vascular plant endemism and primary vegetation loss (Supplementary Note provides details) to evaluate whether Central China meets the formal criteria for designation as a global biodiversity hotspot6.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequences newly generated for this study are available in GenBank and all accession numbers are provided in Supplementary Data 6. The phylogeny and species distribution data generated in this study are available via the ScienceDB repository at https://doi.org/10.57760/sciencedb.27130 (ref. 111).
Code availability
The code used to analyse the data is available via the ScienceDB repository at https://doi.org/10.57760/sciencedb.27130 (ref. 111).
References
Barnosky, A. D. et al. Has the Earth’s sixth mass extinction already arrived?. Nature 471, 51–57 (2011).
Andermann, T., Faurby, S., Turvey, S. T., Antonelli, A. & Silvestro, D. The past and future human impact on mammalian diversity. Sci. Adv. 6, eabb2313 (2020).
Shaw, R. E. et al. Global meta-analysis shows action is needed to halt genetic diversity loss. Nature 638, 704–710 (2025).
Kunming–Montreal Global Biodiversity Framework (UN-CBD, 2022).
Brodie, J. F. et al. A well-connected earth: the science and conservation of organismal movement. Science 388, eadn2225 (2025).
Myers, N., Mittermeier, R. A., Mittermeier, C. G., Da Fonseca, G. A. B. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858 (2000).
Antonelli, A. et al. Madagascar’s extraordinary biodiversity: evolution, distribution, and use. Science 378, eabf0869 (2022).
Kier, G. et al. A global assessment of endemism and species richness across island and mainland regions. Proc. Natl Acad. Sci. USA 106, 9322–9327 (2009).
Brooks, T. M. et al. Global biodiversity conservation priorities. Science 313, 58–61 (2006).
Lu, L. M. et al. A comprehensive evaluation of flowering plant diversity and conservation priority for national park planning in China. Fundam. Res. 3, 939–950 (2023).
Mishler, B. D. et al. Phylogenetic measures of biodiversity and neo- and paleo-endemism in Australian Acacia. Nat. Commun. 5, 4473 (2014).
Zhang, X. X. et al. Spatial phylogenetics of the Chinese angiosperm flora provides insights into endemism and conservation. J. Integr. Plant Biol. 64, 105–117 (2022).
Cai, L. R. et al. Climatic stability and geological history shape global centers of neo- and paleoendemism in seed plants. Proc. Natl Acad. Sci. USA 120, e2300981120 (2023).
Fjeldså, J. & Lovett, J. C. Geographical patterns of old and young species in African forest biota: the significance of specific montane areas as evolutionary centres. Biodivers. Conserv. 6, 325–346 (1997).
Cardoso Da Silva, J. M. & Bates, J. M. Biogeographic patterns and conservation in the South American Cerrado: a tropical savanna hotspot. Bioscience 52, 225–234 (2002).
López-Pujol, J., Zhang, F. M., Sun, H. Q., Ying, T. S. & Ge, S. Centres of plant endemism in China: places for survival or for speciation?. J. Biogeogr. 38, 1267–1280 (2011).
Rosauer, D., Laffan, S. W., Crisp, M. D., Donnellan, S. C. & Cook, L. G. Phylogenetic endemism: a new approach for identifying geographical concentrations of evolutionary history. Mol. Ecol. 18, 4061–4072 (2009).
Qian, H., Mishler, B. D., Zhang, J. & Qian, S. Global patterns and ecological drivers of taxonomic and phylogenetic endemism in angiosperm genera. Plant Divers. 46, 149–157 (2024).
Omollo, W. O. et al. Spatial heterogeneity of neo- and paleo-endemism for plants in Madagascar. Curr. Biol. 34, 1271–1283 (2024).
Wu, Z. Y., Raven, P. H. & Hong, D. Y. (eds) Flora of China (Science Press & Missouri Botanical Garden Press, 1994–2013).
Huang, J. H. et al. Diversity hotspots and conservation gaps for the Chinese endemic seed flora. Biol. Conserv. 198, 104–112 (2016).
Mittermeier, R. et al. Hotspots Revisited. Earth’s Biologically Richest and Most Endangered Terrestrial Ecoregions (CEMEX, 2004).
Xing, Y. W. & Ree, R. H. Uplift-driven diversification in the Hengduan Mountains, a temperate biodiversity hotspot. Proc. Natl Acad. Sci. USA 114, E3444–E3451 (2017).
Ding, W. N., Ree, R. H., Spicer, R. A. & Xing, Y. W. Ancient orogenic and monsoon-driven assembly of the world’s richest temperate alpine flora. Science 369, 578–581 (2020).
Cao, G. L. et al. Cenozoic geoclimatic changes drove the evolutionary dynamics of floristic endemism on the Qinghai–Tibet Plateau. Proc. Natl Acad. Sci. USA 122, e2426017122 (2025).
Meng, H. H. et al. Evolution of East Asian subtropical evergreen broad-leaved forests: when and how?. J. Syst. Evol. 63, 1045–1060 (2025).
Dinerstein, E. et al. An ecoregion-based approach to protecting half the terrestrial realm. Bioscience 67, 534–545 (2017).
Zhao, L. N. et al. Spatial heterogeneity of extinction risk for flowering plants in China. Nat. Commun. 15, 6352 (2024).
Hu, H. H. et al. An updated Chinese vascular plant tree of life: phylogenetic diversity hotspots revisited. J. Syst. Evol. 58, 663–672 (2020).
Dimitrov, D. et al. Diversification of flowering plants in space and time. Nat. Commun. 14, 7609 (2023).
Zuntini, A. R. et al. Phylogenomics and the rise of the angiosperms. Nature 629, 843–850 (2024).
Zhang, Q. et al. Meta-analysis provides insights into the origin and evolution of East Asian evergreen broad-leaved forests. New Phytol. 242, 2369–2379 (2024).
Lu, L. M. et al. Evolutionary history of the angiosperm flora of China. Nature 554, 234–238 (2018).
Chen, Y. S., Deng, T., Zhou, Z. & Sun, H. Is the East Asian flora ancient or not?. Natl Sci. Rev. 5, 920–932 (2018).
Hewitt, G. The genetic legacy of the Quaternary ice ages. Nature 405, 907–913 (2000).
Qian, H. & Ricklefs, R. E. Large-scale processes and the Asian bias in species diversity of temperate plants. Nature 407, 180–182 (2000).
Ma, Y. Z. et al. Pervasive hybridization during evolutionary radiation of Rhododendron subgenus Hymenanthes in mountains of southwest China. Natl Sci. Rev. 9, nwac276 (2022).
Peng, H. W. et al. Repeated evolution of storage root and invasions of alpine biome drove replicated radiations of the megadiverse Corydalis (Papaveraceae) in the Qinghai–Tibet Plateau. Syst. Biol. 74, 359–372 (2025).
Qiu, Y. X., Fu, C. X. & Comes, H. P. Plant molecular phylogeography in China and adjacent regions: tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol. Phylogenet. Evol. 59, 225–244 (2011).
Tang, C. Q. et al. Identifying long-term stable refugia for relict plant species in East Asia. Nat. Commun. 9, 4488 (2018).
Comes, H. P. & Kadereit, J. W. The effect of Quaternary climatic changes on plant distribution and evolution. Trends Plant Sci. 3, 432–438 (1998).
Stewart, J. R., Lister, A. M., Barnes, I. & Dalén, L. Refugia revisited: individualistic responses of species in space and time. Proc. R. Soc. Ser. B Biol. Sci. 277, 661–671 (2010).
Kadereit, J. W. & Abbott, R. J. Plant speciation in the Quaternary. Plant Ecolog. Divers. 14, 105–142 (2021).
Xu, X. X. et al. Late Pleistocene speciation of three closely related tree peonies endemic to the Qinling–Daba Mountains, a major glacial refugium in Central China. Ecology and Evolution 9, 7528–7548 (2019).
Nevado, B., Contreras-Ortiz, N., Hughes, C. & Filatov, D. A. Pleistocene glacial cycles drive isolation, gene flow and speciation in the high-elevation Andes. New Phytol. 219, 779–793 (2018).
Perrigo, A., Hoorn, C. & Antonelli, A. Why mountains matter for biodiversity. J. Biogeogr. 47, 315–325 (2020).
Xu, W. et al. Hidden hotspots of amphibian biodiversity in China. Proc. Natl Acad. Sci. USA 121, e2320674121 (2024).
Thornhill, A. H. et al. Continental-scale spatial phylogenetics of Australian angiosperms provides insights into ecology, evolution and conservation. J. Biogeogr. 43, 2085–2098 (2016).
Harrison, S. & Noss, R. Endemism hotspots are linked to stable climatic refugia. Ann. Bot. 119, 207–214 (2017).
Shipley, B. R. & McGuire, J. L. Interpreting and integrating multiple endemism metrics to identify hotspots for conservation priorities. Biol. Conserv. 265, 109403 (2022).
Lamoreux, J. F. et al. Global tests of biodiversity concordance and the importance of endemism. Nature 440, 212–214 (2006).
Waldron, A. et al. Reductions in global biodiversity loss predicted from conservation spending. Nature 551, 364–367 (2017).
Seidl, A., Mulungu, K., Arlaud, M., van den Heuvel, O. & Riva, M. The effectiveness of national biodiversity investments to protect the wealth of nature. Nat. Ecol. Evol. 5, 530–539 (2021).
Chen, M. X., Gong, Y. H., Li, Y., Lu, D. D. & Zhang, H. Population distribution and urbanization on both sides of the Hu Huanyong line: answering the Premier’s question. J. Geogr. Sci. 26, 1593–1610 (2016).
Xu, W. B. et al. Human activities have opposing effects on distributions of narrow-ranged and widespread plant species in China. Proc. Natl Acad. Sci. USA 116, 26674–26681 (2019).
Chen, Z. D. et al. Tree of life for the genera of Chinese vascular plants. J. Syst. Evol. 54, 277–306 (2016).
Takhtajan, A. Floristic Regions of the World (Univ. of California Press, 1986).
Zachos, F. E., & Habel, J. C. (eds) Biodiversity Hotspots: Distribution and Protection of Conservation Priority Areas (Springer Science & Business Media, 2011).
Noss, R. F. et al. How global biodiversity hotspots may go unrecognized: lessons from the North American Coastal Plain. Divers. Distrib. 21, 236–244 (2015).
Zhao, Y. P. et al. Resequencing 545 ginkgo genomes across the world reveals the evolutionary history of the living fossil. Nat. Commun. 10, 4201 (2019).
Zhu, S. S. et al. Genomic insights on the contribution of balancing selection and local adaptation to the long-term survival of a widespread living fossil tree, Cercidiphyllum japonicum. New Phytol. 228, 1674–1689 (2020).
Fan, H. Z. et al. Geographical patterns and determinants of insect biodiversity in China. Sci. China Life Sci. 67, 1255–1265 (2024).
Hu, Y. B. et al. Spatial patterns and conservation of genetic and phylogenetic diversity of wildlife in China. Sci. Adv. 7, eabd5725 (2021).
Guang, X. M. et al. Chromosome-scale genomes provide new insights into subspecies divergence and evolutionary characteristics of the giant panda. Sci. Bull. 66, 2002–2013 (2021).
Wu, H. et al. Hybrid origin of a primate, the gray snub-nosed monkey. Science 380, eabl4997 (2023).
Luedtke, J. A. et al. Ongoing declines for the world’s amphibians in the face of emerging threats. Nature 622, 308–314 (2023).
Ren, H., Corlett, R. T., Ouyang, Z. Y. & Blackmore, S. How can China protect 30% of its land?. Trends Ecol. Evol. 40, 824–826 (2025).
Zhao, W. J. Beginning: China’s national park system. Natl Sci. Rev. 9, nwac150 (2022).
Fos, S., Laguna, E., Jiménez, J. & Gómez-Serrano, M. Á Plant micro-reserves in Valencia (E. Spain): a model to preserve threatened flora in China?. Plant Divers. 39, 383–389 (2017).
Saura, S., Bodin, Ö & Fortin, M.-J. Stepping stones are crucial for species’ long-distance dispersal and range expansion through habitat networks. J. Appl. Ecol. 51, 171–182 (2014).
Brodie, J. F. et al. The contribution of other effective area-based conservation measures (OECMs) to protecting global biodiversity. Nat. Commun. 16, 7886 (2025).
Liang, L. Z. et al. A review of fengshui forests: ecological functions, humanistic values, and potential applications to enhance biodiversity in urban green landscapes and achieve sustainable development goals. Sustainability 17, 3314 (2025).
Pan, X. L., Xu, L. Y., Yang, Z. F. & Yu, B. Payments for ecosystem services in China: policy, practice, and progress. J. Cleaner Prod. 158, 200–208 (2017).
Shang, W. X., Gong, Y. C., Wang, Z. J. & Stewardson, M. J. Eco-compensation in China: theory, practices and suggestions for the future. J. Environ. Manage. 210, 162–170 (2018).
Croci, E., Lucchitta, B. & Cusa, M. Biodiversity credits schemes: a comparative analysis. J. Cleaner Prod. 523, 146382 (2025).
Comprehensive update to the world’s biodiversity hotspots project begins. IUCN https://iucn.org/news/202510/comprehensive-update-worlds-biodiversity-hotspots-project-begins (2025).
Smith, R. J. & Matimele, H. Twenty-five years of misinterpreting the biodiversity hotspot approach. Nat. Ecol. Evol. 9, 2175–2177 (2025).
Qin, H. N. et al. Threatened species list of China’s higher plants. Biodivers. Sci. 25, 696 (2017).
Johnson, M. G. et al. HybPiper: extracting coding sequence and introns for phylogenetics from high-throughput sequencing reads using target enrichment. Appl. Plant Sci. 4, 1600016 (2016).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Hall, T. A. in Nucleic Acids Symposium Series 95–98 (Oxford, 1999).
The Angiosperm Phylogeny Group. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181, 1–20 (2016).
PPG I. A community-derived classification for extant lycophytes and ferns. J. Syst. Evol. 54, 563–603 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Smith, S. A. & O’Meara, B. C. treePL: divergence time estimation using penalized likelihood for large phylogenies. Bioinformatics 28, 2689–2690 (2012).
Nie, Y. et al. Accounting for uncertainty in the evolutionary timescale of green plants through clock-partitioning and fossil calibration strategies. Syst. Biol. 69, 1–16 (2020).
Ran, J. H., Shen, T. T., Wang, M. M. & Wang, X. Q. Phylogenomics resolves the deep phylogeny of seed plants and indicates partial convergent or homoplastic evolution between Gnetales and angiosperms. Proc. R. Soc. Ser. B Biol. Sci. 285, 20181012 (2018).
Magallón, S., Gómez-Acevedo, S., Sánchez-Reyes, L. L. & Hernández-Hernández, T. A metacalibrated time-tree documents the early rise of flowering plant phylogenetic diversity. New Phytol. 207, 437–453 (2015).
Bouckaert, R. et al. BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Wu, Z. Y., Sun, H., Zhou, Z. K., Peng, H. & Li, D. Z. Origin and differentiation of endemism in the flora of China. Front. Biol. China 2, 125–143 (2007).
Huang, J. H., Ma, K. P. & Chen, B. Diversity and Geographic Distribution of Endemic Species of Seed Plants in China (Science Press, 2014).
Zhou, X. L., Zhang, X. C., Sun, J. Q. & Yan, Y. H. Diversity and distribution of lycophytes and ferns in China. Biodivers. Sci. 24, 102–107 (2016).
Christenhusz, M. J. M. et al. A new classification and linear sequence of extant gymnosperms. Phytotaxa 19, 55–70 (2011).
Hu, H. H. et al. Temporal and spatial comparisons of angiosperm diversity between eastern Asia and North America. Natl Sci. Rev. 9, nwab199 (2022).
Liu, Y. et al. The Sino-Himalayan flora evolved from lowland biomes dominated by tropical floristic elements. BMC Biol. 21, 239 (2023).
Yang, Y. C. Biohistoria.jl: lineage accumulation rate calculator (v0.0.1). Github https://github.com/Mikumikunisiteageru/Biohistoria.jl (2025).
Oksanen, J. et al. vegan: community ecology package. R package version 2.6–6.1 https://CRAN.R-project.org/package=vegan (2025).
Orme, C. D. L. et al. Global hotspots of species richness are not congruent with endemism or threat. Nature 436, 1016–1019 (2005).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2024).
Nitta, J. H., Laffan, S. W., Mishler, B. D. & Iwasaki, W. canaper: categorical analysis of neo- and paleo-endemism in R. Ecography 2023, e06638 (2023).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Strona, G., Nappo, D., Boccacci, F., Fattorini, S. & San-Miguel-Ayanz, J. A fast and unbiased procedure to randomize ecological binary matrices with fixed row and column totals. Nat. Commun. 5, 4114 (2014).
Ying, T. S. & Zhang, Z. S. Endemism in the flora of China-studies on the endemic genera. J. Syst. Evol. 22, 259–268 (1984).
Clark, M. K. et al. Late Cenozoic uplift of southeastern Tibet. Geology 33, 525 (2005).
Wang, E. et al. Two-phase growth of high topography in eastern Tibet during the Cenozoic. Nat. Geosci. 5, 640–645 (2012).
Paradis, E. Analysis of Phylogenetics and Evolution with R (Springer New York, 2012).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: a grammar of data manipulation. R package version 1.1.4 https://CRAN.R-project.org/package=dplyr (2023).
Pan, X. et al. Evaluating conservation gaps of China’s national key protected wild plants: insights from county-level distribution data. BMC Biol. 23, 156 (2025).
Hoffman M. Koenig K. Bunting G. Costanza J. Williams K. J. Biodiversity hotspots (version 2016.1) Zenodo https://doi.org/10.5281/zenodo.3261807 (2016).
Feng, Y. L. et al. A comprehensive dated phylogeny of China’s vascular plants reveals a hidden global biodiversity hotspot. Science Data Bank https://doi.org/10.57760/sciencedb.27130 (2025).
Nan, X., Li, A. & Deng, W. Data set of ‘digital mountain map of China’ (2015). National Tibetan Plateau / Third Pole Environment Data Center https://doi.org/10.11888/Terre.tpdc.272523 (2022).
Acknowledgements
We are grateful to L.-X. Yuan, X.-X. Zhou, J.-F. Ye, B.-H. Dao, F.-L. Li, Q.-B. Gong, J.-B. Pan, Z.-Z. Yang, J.-C. Chi, J. Wang, X.-W. Li and H.-L. Zheng for field assistance and/or sample collection, Q.-H. Guo, Y.-J. Su and T.-Y. Hu for sharing the updated vegetation map of China (1:1000000) and Y.-F. Li, L.-N. Zhao and Q. Zhang for suggestions and assisting with data analysis. This study was supported by National Natural Science Foundation of China (32122009 to L.-M.L. and 32221001 to Z.-D.C.), National Key Research Development Program of China (2022YFF0802300 to L.-M.L. and 2023YFF0805800 to Z.-D.C.), the International Partnership Program of Chinese Academy of Sciences (063GJHZ2024053FN to L.-M.L. and 151853KYSB20190027 to Z.-D.C.), The Youth Innovation Promotion Association CAS (2020080 to L.-M.L.), Biological Resources Program of CAS (KFJBRP-017-087 to B.L.), Sino-Africa Joint Research Center, CAS (SAJC202527ZD01 to Z.-D.C.) and the Chinese Academy of Sciences President’s International Fellowship Initiative (to R.L.B. and A.A.).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.-M.L. and Z.-D.C. Methodology: Y.-L.F., H.-H.H., D.-X.P., Y.-C.Y. and L.-M.L. Investigation: Y.-L.F., B.L., H.-H.H. and D.-X.P. Visualization: Y.-L.F. and Y.-C.Y. Funding acquisition: L.-M.L. and Z.-D.C. Supervision: Z.-D.C., L.-M.L. and B.L. Writing–original draft: Y.-L.F. and L.-M.L. Writing–review and editing: Y.-L.F., L.-M.L., Z.-D.C., R.L.B and A.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Thomas Givnish, Sarah Mathews and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 China’s physical geography and administrative divisions.
a, Topography and major mountain ranges in China. Black lines depict major mountain ranges in China, with labeled ranges (in black text) indicating mountains within or adjacent to endemism centres identified in this study. Geographic data of mountains were obtained from the digital mountain map of China (National Tibetan Plateau Data Center)112. b, China’s administrative divisions. Blue lines show major rivers: (1) Amur River; (2) Yellow River; (3) Yangtze River; (4) Pearl River; (5) Yarlung Tsangpo River; (6) Tarim River. The river’s geospatial data were downloaded from the Resource and Environmental Science Data Platform (https://www.resdc.cn/, accessed May 2025). Review drawing number for map: GS(2019)1823. Mts: Mountains.
Extended Data Fig. 2 Geographic extent of the four recognized global biodiversity hotspots overlapping China.
The map shows the spatial distribution of four recognized global biodiversity hotspots intersecting China: (1) Mountains of Central Asia, (2) Himalaya, (3) Mountains of Southwest China, and (4) Indo-Burma. The black dashed line represents the Hu Huanyong Line, which delineates China’s east-west population divide, extending from Heihe City (Heilongjiang Province) to Tengchong City (Yunnan Province). Review drawing number for map: GS(2019)1823.
Extended Data Fig. 3 Dated phylogeny of Chinese vascular plants estimated in treePL with 220 calibrations.
The phylogeny includes 16,585 species and 3,029 genera of vascular plants native to China. The sampled 6,265 species endemic to China are highlighted with red branches. Major clades, including ferns, lycophytes, gymnosperms, basal angiosperms, monocots, magnoliids, basal eudicots, superrosids and superasterids, are indicated with different coloured stripes encircling the tree. Calibration points are shown with black dots. Ma: million years ago.
Extended Data Fig. 4 Comparison of stem ages of angiosperm families and genera inferred in this study and two recent publications.
a-c, Stem age comparison at the family level between our tree and those of Dimitrov et al.30 (a) and Zuntini et al.31 (b). The kernel density plot shows the distribution of family age differences, calculated as the age in this study minus the age in Dimitrov et al.30 and Zuntini et al.31 (c), respectively. d-f, Stem age comparison at the genus level between our tree and those of Dimitrov et al.30 (d) and Zuntini et al.31 (e). The kernel density plot shows the distribution of genus age differences (f). The association between variables was evaluated using Spearman’s rank correlation (ρ), and statistical significance was assessed using two-sided tests. Panel a: n = 226, ρ = 0.47, P = 1.3 × 10⁻¹³; panel b: n = 226, ρ = 0.61, P < 2.2 × 10⁻¹⁶; panel d: n = 2,613, ρ = 0.46, P < 2.2 × 10⁻¹⁶; panel e: n = 1,912, ρ = 0.54, P < 2.2 × 10⁻¹⁶. In each panel, “n” denotes the number of paired observations. P values were not adjusted for multiple comparisons. Ma: million years ago.
Extended Data Fig. 5 Diversity and endemism patterns for Chinese vascular plants at the species level.
a, b, Taxonomic richness for all species (a) and endemic species (b). c, d, Phylogenetic diversity (PD) for all species (c) and endemic species (d). e, f, Phylogenetic endemism (PE) for all species (e) and endemic species (f). Review drawing number for maps: GS(2019)1823.
Extended Data Fig. 6 Diversity patterns for Chinese vascular plants at the genus level.
a, b, Taxonomic richness for all genera (a) and endemic genera (b). c, d, Phylogenetic diversity (PD) for all genera (c) and endemic genera (d). e, f, Phylogenetic endemism (PE) for all genera (e) and endemic genera (f). Gray cells indicate the absence of endemic genera. Review drawing number for maps: GS(2019)1823.
Extended Data Fig. 7 Geographic distribution of endemic genera and endemic species of the youngest and oldest quartiles.
a, b, Richness of the youngest 25% of endemic genera (a) and endemic species (b). c, d, Richness of the oldest 25% of endemic genera (c) and endemic species (d). Gray cells indicate the absence of endemic genera or species. Review drawing number for maps: GS(2019)1823. Ma: million years ago.
Extended Data Fig. 8 Geographic patterns and taxonomic endemism centres of genera and species of vascular plants endemic to China, divided by 23 Ma and 5 Ma as boundaries between young and old components, respectively.
a-d, Richness of genera younger than 23 Ma (a), species younger than 5 Ma (b), genera older than 23 Ma (c), species older than 5 Ma (d). e, f, Taxonomic endemism centres identified by the top 5% criterion at the genus (e) and species (f) levels: (1) the Hengduan Mountains, (2) Central China, and (3) Yunnan-Guizhou-Guangxi boundary region. China’s national nature reserves are highlighted in dark green, while four global biodiversity hotspots are highlighted in brown. Review drawing number for maps: GS(2019)1823. Ma: million years ago.
Extended Data Fig. 9 Geographic distribution of phylogenetic endemism centres based on taxa endemic to China.
a, Centres derived from endemic genera. b, Centres derived from endemic species. Centres of neo-endemism, palaeo-endemism, and mixed-endemism are represented by pink, blue, and purple grid cells, respectively. China’s national nature reserves are highlighted in dark green, and four global biodiversity hotspots are highlighted in brown. Numbers above the maps denote taxa involved in the analyses. Review drawing number for maps: GS(2019)1823.
Extended Data Fig. 10 The remaining native vegetation in Central China.
The black line outlines the proposed new biodiversity hotspot (Central China) in this study, while green area highlights the distribution of remaining native vegetation. Yellow dots represent established national parks, while blue dots indicate national park pilots. National parks (or pilots) in Central China: 1. Giant Panda National Park; 2. Shennongjia National Park (pilot); 3. Nanshan National Park (pilot); 4. Nanling National Park (pilot); 5. Wuyi Mountains National Park; 6. Qianjiangyuan National Park (pilot). Review drawing number for maps: GS(2019)1823.
Supplementary information
Supplementary Information (download PDF )
Supplementary discussion, note, Figs. 1–8, Tables 1–9 and references.
Supplementary Data (download XLSX )
Supplementary Data 1–8.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, YL., Hu, HH., Liu, B. et al. A comprehensive dated phylogeny of China’s vascular plants reveals a hidden global biodiversity hotspot. Nat Ecol Evol (2026). https://doi.org/10.1038/s41559-026-03025-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41559-026-03025-1