Abstract
Epstein–Barr virus (EBV) infects >95% of adults and contributes to several human cancers. EBV can remain latent where viral lytic genes are silenced, precluding the use of antiviral agents such as ganciclovir. Little is known about the host factors involved in EBV latency. Here we performed a human genome-wide CRISPR–Cas9 screen in Burkitt lymphoma B cells, which identified lysine-specific histone demethylase 1 (LSD1) and its corepressors REST corepressor 1 (CoREST) and zinc finger protein 217 (ZNF217) as critical for EBV latency. Gene knockout or LSD1 inhibition triggered EBV reactivation, and the latter sensitized cells to ganciclovir cytotoxicity, including in murine tumour xenografts. Mechanistically, ZNF217 recruits LSD1 and CoREST to form a complex that binds a specific DNA motif associated with regions implicated in EBV reactivation. It removes histone 3 lysine 4 (H3K4) methylation marks and restricts host DNA looping. Alternatively, the H3K4 lysine methyltransferase 2D supports EBV lytic reactivation. Our results highlight H3K4 methylation as a major EBV lytic switch regulator and therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data supporting the findings of this study are available within the Article and its Supplementary Information. The ChIP-seq data has been deposited into the NIH GEO omnibus (accession number GSE284859). The CRISPR–Cas9 screen hit results are presented in Supplementary Tables 1 and 2. Source data are provided with this paper.
Code availability
The ChIP-seq analysis in this study was conducted using publicly available software and code, and no specialized code was generated.
References
Farrell, P. J. Epstein–Barr virus and cancer. Annu. Rev. Pathol. 14, 29–53 (2019).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Gewurz, B., Longnecker, R. & Cohen, J. Epstein–Barr virus. Fields Virol. 2, 7 (2021).
Bjornevik, K. et al. Longitudinal analysis reveals high prevalence of Epstein–Barr virus associated with multiple sclerosis. Science 375, 296–301 (2022).
Thorley-Lawson, D. A. EBV persistence–introducing the virus. Curr. Top. Microbiol. Immunol. 390, 151–209 (2015).
Reusch, J. A., Nawandar, D. M., Wright, K. L., Kenney, S. C. & Mertz, J. E. Cellular differentiation regulator BLIMP1 induces Epstein–Barr virus lytic reactivation in epithelial and B cells by activating transcription from both the R and Z promoters. J. Virol. 89, 1731–1743 (2015).
Feederle, R. et al. The Epstein–Barr virus lytic program is controlled by the co-operative functions of two transactivators. EMBO J. 19, 3080–3089 (2000).
El-Guindy, A., Ghiassi-Nejad, M., Golden, S., Delecluse, H. J. & Miller, G. Essential role of Rta in lytic DNA replication of Epstein–Barr virus. J. Virol. 87, 208–223 (2013).
Frost, T. C. & Gewurz, B. E. Epigenetic crossroads of the Epstein–Barr virus B-cell relationship. Curr. Opin. Virol. 32, 15–23 (2018).
Murata, T. & Tsurumi, T. Switching of EBV cycles between latent and lytic states. Rev. Med. Virol. 24, 142–153 (2014).
Kenney, S. C. & Mertz, J. E. Regulation of the latent-lytic switch in Epstein–Barr virus. Semin Cancer Biol. 26, 60–68 (2014).
Li, D., Fu, W. & Swaminathan, S. Continuous DNA replication is required for late gene transcription and maintenance of replication compartments in gammaherpesviruses. PLoS Pathog. 14, e1007070 (2018).
Djavadian, R., Hayes, M. & Johannsen, E. CAGE-seq analysis of Epstein–Barr virus lytic gene transcription: 3 kinetic classes from 2 mechanisms. PLoS Pathog. 14, e1007114 (2018).
Woellmer, A. & Hammerschmidt, W. Epstein–Barr virus and host cell methylation: regulation of latency, replication and virus reactivation. Curr. Opin. Virol. 3, 260–265 (2013).
Hammerschmidt, W. & Sugden, B. Identification and characterization of oriLyt, a lytic origin of DNA replication of Epstein–Barr virus. Cell 55, 427–433 (1988).
Djavadian, R., Chiu, Y. F. & Johannsen, E. An Epstein–Barr virus-encoded protein complex requires an origin of lytic replication in cis to mediate late gene transcription. PLoS Pathog. 12, e1005718 (2016).
Guo, R. et al. MYC controls the Epstein–Barr virus lytic switch. Mol. Cell. 78, 653–69 e8 (2020).
Tanner, J., Weis, J., Fearon, D., Whang, Y. & Kieff, E. Epstein–Barr virus gp350/220 binding to the B lymphocyte C3d receptor mediates adsorption, capping, and endocytosis. Cell 50, 203–213 (1987).
Meng, Q. et al. The Epstein–Barr virus (EBV)-encoded protein kinase, EBV-PK, but not the thymidine kinase (EBV-TK), is required for ganciclovir and acyclovir inhibition of lytic viral production. J. Virol. 84, 4534–4542 (2010).
Feng, W. H., Hong, G., Delecluse, H. J. & Kenney, S. C. Lytic induction therapy for Epstein–Barr virus-positive B-cell lymphomas. J. Virol. 78, 1893–1902 (2004).
Li, H. et al. Therapies based on targeting Epstein–Barr virus lytic replication for EBV-associated malignancies. Cancer Sci. 109, 2101–2108 (2018).
Tikhmyanova, N., Schultz, D. C., Lee, T., Salvino, J. M. & Lieberman, P. M. Identification of a new class of small molecules that efficiently reactivate latent Epstein–Barr virus. ACS Chem. Biol. 9, 785–795 (2014).
Tikhmyanova, N. et al. Development of a novel inducer for EBV lytic therapy. Bioorg. Med. Chem. Lett. 29, 2259–2264 (2019).
Chan, A. T. et al. Azacitidine induces demethylation of the Epstein–Barr virus genome in tumors. J. Clin. Oncol. 22, 1373–1381 (2004).
Dalton, T. et al. Epigenetic reprogramming sensitizes immunologically silent EBV+ lymphomas to virus-directed immunotherapy. Blood 135, 1870–1881 (2020).
Ben-Sasson, S. A. & Klein, G. Activation of the Epstein–Barr virus genome by 5-aza-cytidine in latently infected human lymphoid lines. Int. J. Cancer 28, 131–135 (1981).
Zhang, Y. et al. Histone loaders CAF1 and HIRA restrict Epstein–Barr virus B-cell lytic reactivation. mBio 11, e01063–20 (2020).
Murray-Nerger, L. A. et al. The nucleic acid binding protein SFPQ represses EBV lytic reactivation by promoting histone H1 expression. Nat. Commun. 15, 4156 (2024).
Sanson, K. R. et al. Optimized libraries for CRISPR–Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Shi, Y. J. et al. Regulation of LSD1 histone demethylase activity by its associated factors. Mol. Cell. 19, 857–864 (2005).
Lee, M. G., Wynder, C., Cooch, N. & Shiekhattar, R. An essential role for CoREST in nucleosomal histone 3 lysine 4 demethylation. Nature 437, 432–435 (2005).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Quinlan, K. G. et al. Specific recognition of ZNF217 and other zinc finger proteins at a surface groove of C-terminal binding proteins. Mol. Cell. Biol. 26, 8159–8172 (2006).
Mendez, L. M. et al. CtBP is an essential corepressor for BCL6 autoregulation. Mol. Cell. Biol. 28, 2175–2186 (2008).
Hatzi, K. et al. Histone demethylase LSD1 is required for germinal center formation and BCL6-driven lymphomagenesis. Nat. Immunol. 20, 86–96 (2019).
Graveline, R. et al. The chromatin-associated Phf12 protein maintains nucleolar integrity and prevents premature cellular senescence. Mol. Cell. Biol. 37, e00522–16 (2017).
Song, Y. et al. Mechanism of Crosstalk between the LSD1 Demethylase and HDAC1 Deacetylase in the CoREST Complex. Cell Rep. 30, 2699–711 e8 (2020).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Ersing, I. et al. A temporal proteomic map of Epstein–Barr virus lytic replication in B cells. Cell Rep. 19, 1479–1493 (2017).
Zeng, C. et al. Demethylase-independent roles of LSD1 in regulating enhancers and cell fate transition. Nat. Commun. 14, 4944 (2023).
Myoung, J. & Ganem, D. Generation of a doxycycline-inducible KSHV producer cell line of endothelial origin: maintenance of tight latency with efficient reactivation upon induction. J. Virol. Methods 174, 12–21 (2011).
Yiu, S. P. T., Dorothea, M., Hui, K. F. & Chiang, A. K. S. Lytic induction therapy against Epstein–Barr virus-associated malignancies: past, present, and future. Cancers 12, 2142 (2020).
Tikhmyanova, N. et al. Small molecule perturbation of the CAND1-Cullin1-ubiquitin cycle stabilizes p53 and triggers Epstein–Barr virus reactivation. PLoS Pathog. 13, e1006517 (2017).
Cowger, J. J., Zhao, Q., Isovic, M. & Torchia, J. Biochemical characterization of the zinc-finger protein 217 transcriptional repressor complex: identification of a ZNF217 consensus recognition sequence. Oncogene 26, 3378–3386 (2007).
Collins, C. et al. Positional cloning of ZNF217 and NABC1: genes amplified at 20q13.2 and overexpressed in breast carcinoma. Proc. Natl Acad. Sci. USA 95, 8703–8708 (1998).
Nunez, N. et al. The multi-zinc finger protein ZNF217 contacts DNA through a two-finger domain. J. Biol. Chem. 286, 38190–38201 (2011).
Calderwood, M. A., Holthaus, A. M. & Johannsen, E. The Epstein–Barr virus LF2 protein inhibits viral replication. J. Virol. 82, 8509–8519 (2008).
Froimchuk, E., Jang, Y. & Ge, K. Histone H3 lysine 4 methyltransferase KMT2D. Gene 627, 337–342 (2017).
Ma, Y. et al. CRISPR/Cas9 screens reveal Epstein–Barr virus-transformed B cell host dependency factors. Cell Host Microbe 21, 580–591 e7 (2017).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Tsai, K., Thikmyanova, N., Wojcechowskyj, J. A., Delecluse, H. J. & Lieberman, P. M. EBV tegument protein BNRF1 disrupts DAXX-ATRX to activate viral early gene transcription. PLoS Pathog. 7, e1002376 (2011).
Niller, H. H., Szenthe, K. & Minarovits, J. Epstein–Barr virus-host cell interactions: an epigenetic dialog?. Front. Genet. 5, 367 (2014).
Xu, H. et al. IFI16 partners with KAP1 to maintain Epstein–Barr virus latency. J. Virol. 96, e0102822 (2022).
Ichikawa, T. et al. Regulation of Epstein–Barr virus life cycle and cell proliferation by histone H3K27 methyltransferase EZH2 in Akata cells. mSphere 3, e00478–18 (2018).
Imai, K. et al. Role of the histone H3 lysine 9 methyltransferase Suv39 h1 in maintaining Epsteinn-Barr virus latency in B95-8 cells. FEBS J. 281, 2148–2158 (2014).
Krig, S. R. et al. Identification of genes directly regulated by the oncogene ZNF217 using chromatin immunoprecipitation (ChIP)-chip assays. J. Biol. Chem. 282, 9703–9712 (2007).
Su, S. T. et al. Involvement of histone demethylase LSD1 in Blimp-1-mediated gene repression during plasma cell differentiation. Mol. Cell. Biol. 29, 1421–1431 (2009).
Laichalk, L. L. & Thorley-Lawson, D. A. Terminal differentiation into plasma cells initiates the replicative cycle of Epstein–Barr virus in vivo. J. Virol. 79, 1296–1307 (2005).
Zhang, J. et al. Disruption of KMT2D perturbs germinal center B cell development and promotes lymphomagenesis. Nat. Med. 21, 1190–1198 (2015).
Ortega-Molina, A. et al. The histone lysine methyltransferase KMT2D sustains a gene expression program that represses B cell lymphoma development. Nat. Med. 21, 1199–1208 (2015).
Pasqualucci, L. et al. Analysis of the coding genome of diffuse large B-cell lymphoma. Nat. Genet. 43, 830–837 (2011).
Okosun, J. et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat. Genet. 46, 176–181 (2014).
Beguelin, W. et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell. 23, 677–692 (2013).
Gao, L. M. et al. Somatic mutations in KMT2D and TET2 associated with worse prognosis in Epstein–Barr virus-associated T or natural killer-cell lymphoproliferative disorders. Cancer Biol. Ther. 20, 1319–1327 (2019).
Canzio, D. et al. Antisense lncRNA transcription mediates DNA demethylation to drive stochastic protocadherin α promoter choice. Cell 177, 639–53 e15 (2019).
Mansouri, S., Wang, S. & Frappier, L. A role for the nucleosome assembly proteins TAF-Iβ and NAP1 in the activation of BZLF1 expression and Epstein–Barr virus reactivation. PLoS ONE 8, e63802 (2013).
Kristie, T. M. Dynamic modulation of HSV chromatin drives initiation of infection and provides targets for epigenetic therapies. Virology 479-480, 555–561 (2015).
Zhou, G., Du, T. & Roizman, B. The role of the CoREST/REST repressor complex in herpes simplex virus 1 productive infection and in latency. Viruses 5, 1208–1218 (2013).
Dembowski, J. A., Dremel, S. E. & DeLuca, N. A. Replication-coupled recruitment of viral and cellular factors to herpes simplex virus type 1 replication forks for the maintenance and expression of viral genomes. PLoS Pathog. 13, e1006166 (2017).
Liang, Y. et al. A novel selective LSD1/KDM1A inhibitor epigenetically blocks herpes simplex virus lytic replication and reactivation from latency. mBio 4, e00558-12 (2013).
Liang, Y., Vogel, J. L., Narayanan, A., Peng, H. & Kristie, T. M. Inhibition of the histone demethylase LSD1 blocks alpha-herpesvirus lytic replication and reactivation from latency. Nat. Med. 15, 1312–1317 (2009).
Cliffe, A. R. et al. Neuronal stress pathway mediating a histone methyl/phospho switch is required for herpes simplex virus reactivation. Cell Host Microbe 18, 649–658 (2015).
Gu, H., Liang, Y., Mandel, G. & Roizman, B. Components of the REST/CoREST/histone deacetylase repressor complex are disrupted, modified, and translocated in HSV-1-infected cells. Proc. Natl Acad. Sci. USA 102, 7571–7576 (2005).
Gu, H. & Roizman, B. Herpes simplex virus-infected cell protein 0 blocks the silencing of viral DNA by dissociating histone deacetylases from the CoREST–REST complex. Proc. Natl Acad. Sci. USA 104, 17134–17139 (2007).
Zhou, G., Te, D. & Roizman, B. The CoREST/REST repressor is both necessary and inimical for expression of herpes simplex virus genes. mBio 2, e00313-10 (2010).
Cliffe, A. R. & Knipe, D. M. Herpes simplex virus ICP0 promotes both histone removal and acetylation on viral DNA during lytic infection. J. Virol. 82, 12030–12038 (2008).
Wang, L. W. et al. Epstein–Barr-virus-induced one-carbon metabolism drives B cell transformation. Cell Metab. 30, 539–55 e11 (2019).
Guo, R. et al. Methionine metabolism controls the B cell EBV epigenome and viral latency. Cell Metab. 34, 1280–97 e9 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Picard Toolkit. GitHub http://broadinstitute.github.io/picard/ (Broad Institute, 2019).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Acknowledgements
This work was supported by NIH grant nos. R01AI164709, R01CA228700, R01DE033907 and P01CA269043 to B.E.G.; by a Lymphoma Research Foundation Postdoctoral Fellowship to Y.L.; by grant no. U01CA275301 to B.E.G and L.G.-R.; and by a Broad Institute grant to B.E.G and Z.L. The funders had no role in study design; data collection and analysis; decision to publish; or preparation of the manuscript. We thank J. M. Middeldorp (Amsterdam University Medical Centre, Amsterdam, the Netherlands) for sharing the anti-EBV gp350/220 antibody (OT6).
Author information
Authors and Affiliations
Contributions
Y.L., J.Y., Z.L., T.Y. and D.M. performed the experiments and data analysis; W.D. performed the ChIP-seq bioinformatic analysis; I.Y.K., S.C. and L.G.-R. performed the xenograft experiments; and Y.L. and B.E.G. supervised the study. Y.L., J.Y. and B.E.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Lori Frappier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 MYC knockout rapidly triggers EBV lytic reactivation.
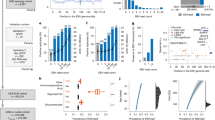
(A) Cross-comparison of Avana sgRNA library Day 6 versus Brunello library Day 9 CRISPR screens for host factors that repress EBV reactivation. MYC scored at Day 6 but not Day 9 in the Akata library screen. (B) Immunoblot analysis of WCL from P3HR-1 (left) or Akata (right) cells that expressed control vs Brunello MYC targeting sgRNAs at the indicated days post-transduction. (C) Mean ± SD PM gp350 levels from n = 3 replicates of P3HR-1 (left) or Akata (right) cells expressing control vs MYC sgRNAs as in (B). NS: not significant, *** p < 0.001 by unpaired two-sided Student’s t-test. (D) FACS analysis of Cas9+ EBV+ vs EBV− Akata cells that expressed control or independent screen hit UROD targeting sgRNAs. UROD depletion strongly increase FL4 channel autofluorescence signal. (E) Immunoblot analysis of Akata cells as in (D) that expressed control or UROD targeting sgRNAs. WCL from EBV+ Akata cells ± anti-immunoglobulin (αIgG) cross-linking were used as positive and negative controls. All blots shown are representative of n = 3 replicates.
Extended Data Fig. 2 LSD1–ZNF217–CoREST complex co-repressors restrict Burkitt B-cell EBV reactivation.
(A) STRING interaction network analysis94 of LSD1–ZNF217–CoREST complex and its associated factors. Shown are edges, which depict protein-protein associations, and confidence scores, which depict the estimated likelihood, based on supporting evidence, that the predicted interaction is biologically meaningful, specific and reproducible. (B) Immunoblot analysis of WCL from Akata cells expressing control or CTBP1 targeting sgRNA. (C) FACS analysis of PM gp350 abundance in Akata cells expressing control or independent Brunello library CTBP1 targeting sgRNAs. (D) Immunoblot analysis of WCL from Akata cells expressing control or BCL6 targeting sgRNA. (E) FACS analysis of PM gp350 levels in Akata cells expressing control or BCL6 targeting sgRNA. (F) Immunoblot analysis of WCL from Akata cells expressing control or independent Brunello PHF12 sgRNAs. (G) FACS analysis of PM gp350 levels in Akata cells expressing control or independent Brunello PHF12 targeting sgRNAs. (H) Mean ± SD intracellular EBV genome copy number from n = 3 replicates of Akata cells expressing control, CTBP1, BCL6, or PHF12 sgRNAs. ** p < 0.01, *** p < 0.001 by unpaired two-sided Student’s t-test. Blots shown are representative of n = 3 replicates.
Extended Data Fig. 3 LSD1, ZNF217 and CoREST maintain Burkitt B-cell EBV latency.
(A) WCL of P3HR-1 cells were subjected to immunoprecipitation using control IgG or anti-LSD1 antibody, followed by immunoblot analysis with 1% input for the indicated proteins. (B) (Left) LSD1, ZNF217 and CoREST relative protein abundances in P3HR-1 ZHT/RHT cells uninduced or induced for lytic replication by 4-HT for the indicated times, or (Right) in EBV-negative (EBV−) versus EBV-positive (EBV+) Akata mock induced or induced for lytic replication by anti-IgG crosslinking for 48 h, using data from Ersing et al.47. Mean ± SD from n = 2 replicates were presented. (C-E) FACS analysis of PM gp350 abundances in P3HR-1 cells expressing control, LSD1 (C), ZNF217 (D) or CoREST (E) targeting sgRNAs. (F) Immunoblot analysis of WCL from EBV super-infected AGS or EBV+ SNU719 gastric carcinoma cells expressing control or ZNF217 sgRNAs. (G) Immunoblot analysis of WCL from EBV+ or EBV- Akata cells expressing control or CoREST sgRNA. Blots are representative of n = 3 independent replicates.
Extended Data Fig. 4 LSD1 inhibition by the small molecule antagonist C12 induces EBV reactivation in B and epithelial cells.
(A) Immunoblot analysis of WCL from latency I EB3 or Rael Burkitt cells, latency III GM15892 LCLs or KEM III LCLs, EBV+ AGS gastric carcinoma and C666-1 nasopharyngeal carcinoma cells treated with C12 (0,1,2 or 5 μM) for 48 h. (B) Immunoblot analysis of WCL from MUTU I that were treated with the indicated concentration of C12 for 6 or 24 h and then grown in C12-free RPMI for the remainder of 48 h. (C) Immunoblot analysis of WCL from KEM I cells treated with C12 (0, 1, 2, 5, or, 10 μM) alone or in combination with NaB (0.5 mM) for 48 h. (D-E) Immunoblot analysis of WCL (D) and FACS analysis of PM gp350 (E) from P3HR-1 cells treated with C12 (0, 1, 2, 5, or, 10 μM) alone or in combination with NaB (0.5 mM) for 48 h. (F-G) Immunoblot analysis of WCL (F) and FACS analysis of PM gp350 levels (G) from MUTU I cells treated with C12 (0, 1, 2, 5, or, 10 μM) alone or in combination with NaB (0.5 mM) for 48 h. Shown in the top right of the panels in (E) and (G) are %gp350+ cells. Blots are representative of n = 3 independent replicates.
Extended Data Fig. 5 Corin induces Burkitt B-cell EBV lytic reactivation.
(A) Schematic diagram depicting bifunctional small molecule antagonist corin inhibition of LSD1 and HDAC activity. (B) Immunoblot analysis of WCL from P3HR-1, Akata or MUTU I cells that were treated with the indicated concentration of corin for 6 h and then grown in corin-free RPMI for the remainder of 48 h. (C) Mean ± SD intracellular EBV genome copy number from n = 3 replicates of P3HR-1 or MUTU I cells treated with corin for 48 h. **p < 0.01, ***p < 0.001 by unpaired two-sided Student’s t-test. (D) Immunoblot analysis of WCL from Akata cells with stable GFP versus MYC cDNA overexpression that were treated with corin for 48 h, as indicated. (E) Immunoblot analysis of WCL from control or BZLF1 knockout (KO) Akata cells treated with corin for 48 h, suggesting that corin requires BZLF1 to induce early and late protein expression. (F) Immunoblot analysis of WCL from Akata, KEM I, EB3, Rael Burkitt cells, GM15892 LCLs or SNU719 gastric carcinoma cells treated with corin for 48 h. (G) Immunoblot analysis of WCL from EBV+ /KSHV+ BC-1 or KSHV+ BCBL-1 primary effusion lymphoma cells treated with corin for 48 h. (H) KSHV+ iSLK.219 epithelial cells with conditional doxycycline (Dox) inducible KSHV immediate early RTA expression was treated with DMSO, Dox (0.5 ug/ml) or corin (5 μM) for 24 h, and then maintained in growth media without Dox or corin. 48 h later, immunofluorescence analysis was performed for GFP (stably expressed from the KSHV genome) versus for red fluorescence protein (RFP, controlled by lytic PAN RNA promoter, which reports KSHV lytic gene expression). Scale bar = 650 μm. Blots and immunofluorescence images are representative of n = 3 independent replicates. Created with BioRender.com.
Extended Data Fig. 6 Corin induces EBV lytic reactivation in EBV+ transformed Burkitt and lymphoblastoid B-cells and in Burkitt xenograft tumors in vivo.
(A) Corin and ganciclovir co-treatment effects on EBV-negative Burkitt and EBV+ LCL cells. Shown are mean ± SD fold changes of live cell numbers, relative to DMSO-treated controls, from n = 3 replicates on day 6 of treatment of EBV-negative MUTU I (left) and of EBV+ GM15892 LCLs (right). Cells were treated as described in Fig. 3e with DMSO or corin (0.5 μM). (B) Corin and ganciclovir co-treatment effects on KEM III LCLs. KEM III LCLs were treated with a single dose on day 1 of or with multiple drug doses as described in Fig. 3e. Shown are mean ± SD fold changes of live cell numbers, relative to DMSO-treated controls, from n = 3 replicates on day 6 of treatment. (C) P3HR-1 Burkitt cells were treated with a single dose on day 1 or with multiple doses of DMSO vehicle, LSD1 inhibitor C12 (0.5 μM), the HDAC inhibitor NaB (0.1 mM), C12 + NaB or with the dual LSD1/HDAC inhibitor corin (0.5 μM), using the regimen described in Fig. 3e. Shown are mean ± SD fold changes of live cell numbers, relative to DMSO-treated controls, from n = 3 replicates on day 6 of treatment. (D) Immunoblot analysis of WCL prepared from MUTU I xenografts harvested from mice treated with DMSO vehicle vs corin as described in Fig. 3g. (E) Immunohistochemical analysis of BZLF1 expression in MUTU I xenografts harvested from mice treated with DMSO vs corin, as described in Fig. 3g. Scale bar = 100 μm. Representative images from n = 4 randomly selected fields are presented. (F) Quantification of BZLF1+ (left panel) versus BMRF1+ (right panel) cell numbers in MUTUI xenograft tumors as in Fig. 3h and Extended Data Fig. 6E. Shown are mean ± SEM numbers of BZLF1+ or BMRF1+ cells from four randomly selected fields per mouse, quantitated by Image J using the Cell Counter plugin. (G) qPCR analysis of BZLF1, early BMRF1 and late BLLF1 (encodes gp350) mRNA abundances in xenograft tumors from tumors shown in Fig. 3h and Extended Data 6E. Shown are mean of n = 3 or 4 qPCR values. (H) qPCR defined intracellular EBV genome copy number from the xenograft tumors shown in Fig. 3h. Box heights indicate mean of n = 3 or 4 qPCR values. (I) MUTU I xenograft tumor sizes from the vehicle control, corin, GCV or corin and GCV treated mice (n = 6 or 7) shown in Fig. 3j. Data are presented as mean ± standard error of the mean (SEM). Significance was determined by cross-comparison of mean tumor sizes from dual drug treated versus vehicle or single drug treated mice. Blots shown are representative images of n = 3 replicates. A-C, statistical significance was determined using unpaired two-sided t-test; F, Statistical significance was determined by two-tail nested t-test; I, Statistical significance was determined using multiple t-test. *p < 0.05, **p < 0.01, ***p < 0.001, NS, not significant.
Extended Data Fig. 7 ZNF217 recruits LSD1 and CoREST to restrict EBV genomic H3K4 methylation.
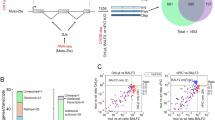
(A) ChIP-qPCR analysis of CoREST occupancy at oriLyt versus BZLF1p and BXLF1 regions in Akata control versus ZNF217 KO cells. Mean ± SD percentages of input values from n = 3 replicates are shown. (B) ZNF217 mutant validation. Immunoblot analysis of WCL vs α-V5 epitope tagged control GFP, wildtype or deletion mutant ZNF217 complexes immunopurified from Cas9+ Akata cells that expressed sgRNA against endogenous ZNF217. Red asterisks indicate V5-tagged ZNF217 bands. (C-D) ChIP-qPCR analysis of LSD1 (C) or CoREST (D) occupancies at the BZLF1 promoter or oriLytL in Cas9+ Akata cells with stable wildtype or DNA binding domain deletion (Δ471-526) ZNF217 cDNAs and that also expressed negative control or ZNF217 targeting sgRNA. Mean ± SD percentages of input values from n = 3 replicates are shown. (E) Immunoblot analysis of WCL from Akata cells expressing V5-tagged GFP or ZNF217 cDNAs, with or without α-IgG (10 μg/ml) stimulation for 24 h (F) ChIP-qPCR analysis of BZLF1 promoter or oriLyt LSD1 or CoREST occupancy in Akata cells with stable expression of control GFP or ZNF217 cDNAs. Mean ± SD percentages of input values from n = 3 replicates are shown. (G) Immunoblot analysis of WCL from Cas9+ Akata cells that expressed negative control sgRNA, an independent sgRNA targeting the oriLyt R or L regions with the highest ChIP-seq defined ZNF217 occupancy (sgRNA #1-4 against oriLyt R and #6-9 against oriLyt L), or positive control sgRNAs targeting oriLyt R vs oriLyt L region MYC-occupied E-Box sites critical for EBV latency19 (sgRNA #5 and #10, respectively). (H) ChIP-qPCR analysis of ZNF217 (left) or H3K4me1 (right) levels in negative control vs oriLyt region expressing Akata cells. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. All blots shown are representative images of n = 3 replicates. A, C, D, F, and H, *p < 0.05, **p < 0.01, ***p < 0.001; NS: not significant as defined by unpaired two-tail Student’s t-test.
Extended Data Fig. 8 Analysis of LSD1–ZNF217–CoREST depletion effects on epigenetic marks on EBV genome.
(A-B) ChIP-qPCR analysis of H3K4me1, H3K4me2, and H3K4me3 levels in control vs LSD1, ZNF217 or CoREST depleted Akata cells. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates of Akata cells expressing control, LSD1, ZNF217, or CoREST sgRNAs, using primers specific for oriLytR (A) or the BZLF1 promoter (B). (C) ChIP-qPCR analysis of H3K4me1 levels in control vs LSD1 depleted Akata cells in the presence of acyclovir (100 μg/ml). Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates of Akata cells expressing control or LSD1 sgRNAs, using primers specific for BZLF1p, BXLF1, or oriLyt regions. (D) ChIP-qPCR analysis of H3K4me1 levels in control vs LSD1, ZNF217, or CoREST depleted Akata cells. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates of Akata cells expressing control or LSD1, ZNF217, or CoREST sgRNAs, using primers specific for the BXLF1 region. (E) Mean ± SD ChIP-qPCR % input values from n = 3 replicates of Akata cells expressing control or LSD1 sgRNA, using primers specific for the BZLF1 promoter or oriLyt regions. A-E, *p < 0.05; **p < 0.01; ***p < 0.001, NS: not significant as defined by unpaired two-tail Student’s t-test. Significance values in panels A, B, and E refer to comparisons between cells with the indicated vs control sgRNAs.
Extended Data Fig. 9 Characterization of LSD1 KO, C12 and corin effects on BZLF1 and oriLyt H3K9 and H3K27 epigenetic marks.
(A) ChIP-qPCR analysis of LSD1 and ZNF217 occupancy in Akata cells 24 h after treatment with vehicle, C12 (2.5 μM) or corin (2.5 μM), in the presence of acyclovir (100 μg/ml) to block EBV lytic DNA synthesis. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (B) ChIP-qPCR analysis of H3K4me1, H3K4me2 and H3K4me3 abundances at the BZLF1 promoter or oriLyt regions in Akata cells 24 h after treatment with vehicle, C12 (2.5 μM) or corin (2.5 μM). Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (C) ChIP-qPCR analysis of H3K9Ac and H3K27Ac abundances at the BZLF1 promoter or oriLyt regions in Akata cells 24 h after treatment with vehicle, C12 (2.5 μM) or corin (2.5 μM). Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (D) ChIP-qPCR analysis of H3K9 and H3K27 acetylation levels in Akata cells expressing control versus independent LSD1 targeting sgRNAs. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (E) ChIP-qPCR analysis of H3K9me2 and H3K9me3 abundances at the BZLF1 promoter or oriLyt regions in Akata cells treated with vehicle control, C12 (2.5 μM) or corin (2.5 μM) for 24 h. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. A-E, unpaired two-tail Student’s t-test was performed to cross-compare ChIP-qPCR values from C12 or corin treated cells or LSD1 KO cells with corresponding values from control cells: *p < 0.05, **p < 0.01, ***p < 0.001; NS: not significant.
Extended Data Fig. 10 KMT2D supports Burkitt B-cell EBV lytic reactivation.
(A) Immunoblot analysis of WCL from Akata cells expressing control or KMT2D sgRNAs and that were mock-stimulated or stimulated for lytic reactivation by α-human IgG (10 μg/ml) cross-linking for 24 h. (B) FACS analysis of PM gp350 levels on live Akata cells (as defined by exclusion of the vital dye PI and by forward and side scatter analysis) that expressed control or KMT2D sgRNA following 24 h of mock-stimulation or αIgG crosslinking as in (A). (C) Mean ± SD MFI PM gp350 values from n = 3 replicates, as in (B). (D) qPCR analysis of intracellular EBV genome copy number in Akata cells expressing control or KMT2D sgRNA, treated with or without α-human IgG (10 μg/ml) for 24 h. (E) Immunoblot analysis of WCL from Akata cells expressing control or KMT2D sgRNA and treated with vehicle or corin (2.5 μM) for 24 h. (F) Immunoblot analysis of WCL from KEM III LCLs expressing control or KMT2D sgRNA and treated with vehicle or corin (2.5 μM) for 24 h. (G) ChIP-qPCR analysis of H3K4me1 abundances at the BZLF1 promoter or oriLyt regions in Akata cells expressing control or KMT2D sgRNA 24 h after treatment with vehicle or corin (2.5 μM). Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (H) ChIP-qPCR analysis of H3K4me1 abundances at the BZLF1 promoter or oriLyt regions in Akata cells expressing GFP or KMT2D SET cDNA. Shown are mean ± SD ChIP-qPCR % input values from n = 3 replicates. (I-J) Immunoblot analysis of WCL from Akata cells expressing GFP or KMT2D SET cDNA, treated with or without α-human IgG (10 μg/ml) (I) or corin (2.5 μM) (J) for 24 h. Blots are representative images of n = 3 replicates. C-D and G-H, **p < 0.01; ***p < 0.001 as defined by unpaired two-tail Student’s t-test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Table 1 (download XLSX )
Supplementary Tables 1–5.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, Y., Yan, J., Kong, I.Y. et al. Lysine-specific histone demethylase complex restricts Epstein–Barr virus lytic reactivation. Nat Microbiol 10, 3290–3304 (2025). https://doi.org/10.1038/s41564-025-02165-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02165-7