Abstract
Probiotics are promising alternatives to antibiotics for the treatment of intestinal infections, but the effects of probiotics alone are often insufficient. Here we uncovered synergism between antibacterial iron–sulfur nanozymes (nFeS) and tryptophan derivatives that protects mice and pigs against bacterial gut infections. nFeS selectively inhibited potential intestinal pathogens while sparing commensal Lactobacillus vaginalis, whose presence enhanced the protective activity of nFeS against Salmonella enterica subsp. enterica serovar Typhimurium in vivo. Metabolomics and mutational analysis revealed that L. vaginalis synthesized 2-indolecarboxylic acid from a tryptophan derivative, indole-3-carboxaldehyde, a reaction that was catalysed by nFeS. The cytoplasmic pH of L. vaginalis (pH 7.5) allowed 2-indolecarboxylic acid to chelate free ferrous ions released by nFeS, thereby protecting it from antibacterial effects, whereas pathogens such as S. Typhimurium with a lower cytoplasmic pH were susceptible (pH 6.5). Pretreatment of pigs and mice with L. vaginalis and nFeS protected them against Salmonella infection. Our findings provide a foundation for improving probiotic bacteria-based therapies against intestinal infections.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions are presented in the article and its Extended Data Figs. 1–10 and Supplementary Information. Raw data can be found in the source data files for each figure item. Source data are provided with this paper and are available via figshare at https://doi.org/10.6084/m9.figshare.29652707 (ref. 110). The 16S rRNA gene sequencing data of gut microbiota have been deposited in the NCBI Sequence Read Archive under PRJNA1314424 and PRJNA1314429. Metabolomic (weaned piglets, mice and L. vaginalis) sequencing data have been deposited in CNGBdb under CNP0006014, CNP0006015 and CNP0006017. Transcriptomic sequencing of L. vaginalis has been deposited in NCBI Sequence Read Archive under PRJNA1310755.
References
Lin, Z., Feng, Y., Wang, J., Men, Z. & Ma, X. Microbiota governs host chenodeoxycholic acid glucuronidation to ameliorate bile acid disorder induced diarrhea. Microbiome 13, 36 (2025).
Mostafa, I. et al. Proof-of-concept study of the efficacy of a microbiota-directed complementary food formulation (MDCF) for treating moderate acute malnutrition. BMC Public Health 20, 242 (2020).
Ni, J., Wu, G. D., Albenberg, L. & Tomov, V. T. Gut microbiota and IBD: causation or correlation?. Nat. Rev. Gastroenterol. Hepatol. 14, 573–584 (2017).
Schlechte, J. et al. Dysbiosis of a microbiota–immune metasystem in critical illness is associated with nosocomial infections. Nat. Med. 29, 1017–1027 (2023).
DeGruttola, A. K., Low, D., Mizoguchi, A. & Mizoguchi, E. Current understanding of dysbiosis in disease in human and animal models. Inflamm. Bowel Dis. 22, 1137–1150 (2016).
Bleich, R. M. et al. A consortia of clinical E. coli strains with distinct in vitro adherent/invasive properties establish their own co-colonization niche and shape the intestinal microbiota in inflammation-susceptible mice. Microbiome 11, 277 (2023).
von Mentzer, A. & Svennerholm, A. M. Colonization factors of human and animal-specific enterotoxigenic Escherichia coli (ETEC). Trends Microbiol. 32, 448–464 (2024).
Maier, L. et al. Unravelling the collateral damage of antibiotics on gut bacteria. Nature 599, 120–124 (2021).
Isaac, S. et al. Short- and long-term effects of oral vancomycin on the human intestinal microbiota. J. Antimicrob. Chemother. 72, 128–136 (2016).
van Zyl, W. F., Deane, S. M. & Dicks, L. M. T. Molecular insights into probiotic mechanisms of action employed against intestinal pathogenic bacteria. Gut Microbes 12, 1831339 (2020).
Shepherd, E. S., DeLoache, W. C., Pruss, K. M., Whitaker, W. R. & Sonnenburg, J. L. An exclusive metabolic niche enables strain engraftment in the gut microbiota. Nature 557, 434–438 (2018).
Xue, C. et al. Tryptophan metabolism in health and disease. Cell Metab. 35, 1304–1326 (2023).
Agus, A., Planchais, J. & Sokol, H. Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724 (2018).
Kim, C.-S., Jung, S., Hwang, G.-S. & Shin, D.-M. Gut microbiota indole-3-propionic acid mediates neuroprotective effect of probiotic consumption in healthy elderly: a randomized, double-blind, placebo-controlled, multicenter trial and in vitro study. Clin. Nutr. 42, 1025–1033 (2023).
Wu, H. et al. Strategies for high cell density cultivation of Akkermansia muciniphila and its potential metabolism. Microbiol. Spectr. 12, e02386–02323 (2023).
Lei, C. et al. Lemon exosome-like nanoparticles-manipulated probiotics protect mice from C. diff infection. iScience 23, 101571 (2020).
Dimopoulou, C. et al. An engineered Escherichia coli Nissle 1917 increase the production of indole lactic acid in the gut. FEMS Microbiol. Lett. https://doi.org/10.1093/femsle/fnad027 (2023).
Lin, L. P. et al. Post-ingestion conversion of dietary indoles into anticancer agents. Natl Sci. Rev. 9, nwab144 (2022).
da Silva, D. A. V. et al. Biocide susceptibility and antimicrobial resistance of escherichia coli isolated from swine feces, pork meat and humans in Germany. Antibiotics https://doi.org/10.3390/antibiotics12050823 (2023).
Zhou, Q. et al. The effect of indole-3-lactic acid from Lactiplantibacillus plantarum ZJ316 on human intestinal microbiotain vitro. Foods 11, 3302 (2022).
Chitra, G., Franklin, D. S., Sudarsan, S., Sakthivel, M. & Guhanathan, S. Indole-3-acetic acid/diol based pH-sensitive biological macromolecule for antibacterial, antifungal and antioxidant applications. Int. J. Biol. Macromol. 95, 363–375 (2017).
Yongyan, C., Xiaoxiao, W. & Huan, Y. Indole-3-carboxaldehyde protects mouse from Shigella infection. Acta Microbiol. Sin. 62, 4019–4029 (2022).
de Macedo, V., Meneghete, B. P., Koaski, J. C., Albuquerque, A. S. & Fachi, M. M. Doxycycline for multidrug-resistant gram-negative bacterial infection treatment: a scoping review. J. Glob. Infect. Dis. 15, 95–100 (2023).
Slingerland, C. J., Kotsogianni, I., Wesseling, C. M. J. & Martin, N. I. Polymyxin stereochemistry and its role in antibacterial activity and outer membrane disruption. ACS Infect. Dis. 8, 2396–2404 (2022).
Jones, R. N., Stilwell, M. G., Wilson, M. L. & Mendes, R. E. Contemporary tetracycline susceptibility testing: doxycycline MIC methods and interpretive criteria (CLSI and EUCAST) performance when testing Gram-positive pathogens. Diagn. Microbiol. Infect. Dis. 76, 69–72 (2013).
van Hal, S. J., Lodise, T. P. & Paterson, D. L. The clinical significance of vancomycin minimum inhibitory concentration in Staphylococcus aureus infections: a systematic review and meta-analysis. Clin. Infect. Dis. 54, 755–771 (2012).
Yilmaz, S., Nyerges, A., van der Oost, J., Church, G. M. & Claassens, N. J. Towards next-generation cell factories by rational genome-scale engineering. Nat. Catal. 5, 751–765 (2022).
Sinha, A. K. et al. Dietary fibre directs microbial tryptophan metabolism via metabolic interactions in the gut microbiota. Nat. Microbiol. https://doi.org/10.1038/s41564-024-01737-3 (2024).
Xu, J. et al. Probiotic-inspired nanomedicine restores intestinal homeostasis in colitis by regulating redox balance, immune responses, and the gut microbiome. Adv. Mater. 35, e2207890 (2023).
Xu, Z. et al. Converting organosulfur compounds to inorganic polysulfides against resistant bacterial infections. Nat. Commun. 9, 3713 (2018).
Shen, X. et al. Nano-decocted ferrous polysulfide coordinates ferroptosis-like death in bacteria for anti-infection therapy. Nano Today 35, 100981 (2020).
Meng, X. et al. High-performance self-cascade pyrite nanozymes for apoptosis–ferroptosis synergistic tumor therapy. ACS Nano 15, 5735–5751 (2021).
Cao, H. et al. Deciphering the catalytic mechanism of peroxidase-like activity of iron sulfide nanozymes. ACS Appl. Mater. Interfaces 16, 30958–30966 (2024).
Cao, H. et al. Metastable iron sulfides: a versatile antibacterial candidate with multiple mechanisms against bacterial resistance. Acc. Mater. Res. 4, 115–132 (2023).
Fang, L. et al. Metastable iron sulfides gram-dependently counteract resistant Gardnerella vaginalis for bacterial vaginosis treatment. Adv. Sci. 9, e2104341 (2022).
Bai, M. et al. Diversity of lactic acid bacteria and Bifidobacterium in the intestine of healthy Mongolians. Microbiol. China 46, 2697–2709 (2019).
Ye, Q. et al. Maternal intestinal L. vaginalis facilitates embryo implantation and survival through enhancing uterine receptivity in sows. Microbiome 13, 145 (2025).
Feng, L. et al. The development of intestinal lactic acid bacteria in piglets as determined by high-throughput sequencing. Anim. Biotechnol. 34, 911–920 (2023).
Ruiz-Moyano, S., Martín, A., Benito, M. J., Nevado, F. P. & de Guía Córdoba, M. Screening of lactic acid bacteria and bifidobacteria for potential probiotic use in Iberian dry fermented sausages. Meat Sci. 80, 715–721 (2008).
Zeng, Y. et al. Liberation of daidzein by gut microbial β-galactosidase suppresses acetaminophen-induced hepatotoxicity in mice. Cell Host Microbe 31, 766–780 (2023).
Wang, Z. et al. Lactobacillus vaginalis alleviates DSS induced colitis by regulating the gut microbiota and increasing the production of 3-indoleacrylic acid. Pharmacol. Res. 213, 107663 (2025).
Giordani, B. et al. Functional and safety profile of Limosilactobacillus vaginalis and development of oral fast-disintegrating tablets for gut microbiota modulation. Pharmaceutics https://doi.org/10.3390/pharmaceutics17081011 (2025).
Giordani, B. et al. Limosilactobacillus vaginalis exerts bifidogenic effects: a novel postbiotic strategy for infant prebiotic supplementation. Nutrients https://doi.org/10.3390/nu15204433 (2023).
Sun, M., Wu, W., Liu, Z. & Cong, Y. Microbiota metabolite short chain fatty acids, GPCR, and inflammatory bowel diseases. J. Gastroenterol. 52, 1–8 (2017).
Feng, S. et al. Impaired function of the intestinal barrier in a novel sub-health rat model. Mol. Med. Rep. 13, 3459–3465 (2016).
Scott, S. A., Fu, J. & Chang, P. V. Microbial tryptophan metabolites regulate gut barrier function via the aryl hydrocarbon receptor. Proc. Natl Acad. Sci. USA 117, 19376–19387 (2020).
Miyamoto, K., Sujino, T. & Kanai, T. The tryptophan metabolic pathway of the microbiome and host cells in health and disease. Int. Immunol. 36, 601–616 (2024).
Jiang, T., Liu, N., Jiang, Y.-W., Ye, P.-P. & Xu, W.-M. A practical synthesis of indole-2-carboxylic acid. Org. Prep. Proced. Int. 49, 476–478 (2017).
Fan, L., Wang, Y., Zheng, P. & Sun, J. Methanol dehydrogenase, a key enzyme of one-carbon metabolism: a review. Chin. J. Biotechnol. 37, 530–540 (2021).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Horch, M. et al. Resonance Raman spectroscopy on [NiFe] hydrogenase provides structural insights into catalytic intermediates and reactions. J. Am. Chem. Soc. 136, 9870–9873 (2014).
Galinato, M. G., Whaley, C. M. & Lehnert, N. Vibrational analysis of the model complex (μ-edt)[Fe(CO)3]2 and comparison to iron-only hydrogenase: the activation scale of hydrogenase model systems. Inorg. Chem. 49, 3201–3215 (2010).
Tosi, L. & Danon, J. Infrared spectral evidence of π-bonding in the Fe(CN)5NO−2 ion. Inorg. Chem. 3, 150–151 (1964).
Abdullah Al Awadh, A. Biomedical applications of selective metal complexes of indole, benzimidazole, benzothiazole and benzoxazole: a review (from 2015 to 2022). Saudi Pharm. J. 31, 101698 (2023).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Alwazeer, D. & Cachon, R. Ion-selective electrode integrated in small-scale bioreactor for continuous intracellular pH determination in Lactobacillus plantarum. Folia Microbiol. 65, 467–473 (2020).
Molina-Gutierrez, A., Stippl, V., Delgado, A., Gänzle, M. G. & Vogel, R. F. In situ determination of the intracellular pH of Lactococcus lactis and Lactobacillus plantarum during pressure treatment. Appl. Environ. Microbiol. 68, 4399–4406 (2002).
Cao, F. et al. Artificial-enzymes-armed Bifidobacterium longum probiotics for alleviating intestinal inflammation and microbiota dysbiosis. Nat. Nanotechnol. 18, 617–627 (2023).
Bielecka, M., Biedrzycka, E., Biedrzycka, E., Smoragiewicz, W. & Smieszek, M. Interaction of Bifidobacterium and Salmonella during associated growth. Int. J. Food Microbiol. 45, 151–155 (1998).
Singh, P. et al. Intestinal microbial communities associated with acute enteric infections and disease recovery. Microbiome 3, 45 (2015).
Laursen, M. F. et al. Bifidobacterium species associated with breastfeeding produce aromatic lactic acids in the infant gut. Nat. Microbiol. 6, 1367–1382 (2021).
Sparfel, L., Ratodiarivony, S., Boutet-Robinet, E., Ellero-Simatos, S. & Jolivet-Gougeon, A. Akkermansia muciniphila and alcohol-related liver diseases. a systematic review. Mol. Nutr. Food Res. 68, e2300510 (2024).
Niu, Y. et al. Blautia coccoides is a newly identified bacterium increased by leucine deprivation and has a novel function in improving metabolic disorders. Adv. Sci. 11, e2309255 (2024).
Sun, D. et al. Early-life ruminal microbiome-derived indole-3-carboxaldehyde and prostaglandin D2 are effective promoters of rumen development. Genome Biol. 25, 64 (2024).
Fang, Z. et al. Bifidobacterium longum mediated tryptophan metabolism to improve atopic dermatitis via the gut-skin axis. Gut Microbes 14, 2044723 (2022).
Cheng, J. et al. Molecular insights into the one-carbon loss oxidation of indole-3-acetic acid. ACS Catal. 14, 8528–8540 (2024).
Konopelski, P. & Mogilnicka, I. Biological effects of indole-3-propionic acid, a gut microbiota-derived metabolite, and its precursor tryptophan in mammals’ health and disease. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23031222 (2022).
Wlodarska, M. et al. Indoleacrylic acid produced by commensal peptostreptococcus species suppresses inflammation. Cell Host Microbe 22, 25–37.e26 (2017).
Wang, G. et al. Microbiota-derived indoles alleviate intestinal inflammation and modulate microbiome by microbial cross-feeding. Microbiome 12, 59 (2024).
Gupta, S. K. et al. Microbiota-derived tryptophan metabolism: impacts on health, aging, and disease. Exp. Gerontol. 183, 112319 (2023).
Bai, D. et al. Eubacterium coprostanoligenes alleviates chemotherapy-induced intestinal mucositis by enhancing intestinal mucus barrier. Acta Pharm. Sin. B 14, 1677–1692 (2024).
Zarkan, A. et al. Inhibition of indole production increases the activity of quinolone antibiotics against E. coli persisters. Sci. Rep. 10, 11742 (2020).
Rezwan, F., Lan, R. & Reeves, P. R. Molecular basis of the indole-negative reaction in Shigella strains: extensive damages to the tna operon by insertion sequences. J. Bacteriol. 186, 7460–7465 (2004).
Lee, J. H., Wood, T. K. & Lee, J. Roles of indole as an interspecies and interkingdom signaling molecule. Trends Microbiol. 23, 707–718 (2015).
Nikaido, E. et al. Effects of indole on drug resistance and virulence of Salmonella enterica serovar Typhimurium revealed by genome-wide analyses. Gut Pathog. 4, 5 (2012).
Yan, Y. et al. Research progress on antibacterial activities and mechanisms of natural alkaloids: a review. Antibiotics https://doi.org/10.3390/antibiotics10030318 (2021).
Lin, Z., Wu, J., Wang, J., Levesque, C. L. & Ma, X. Dietary Lactobacillus reuteri prevent from inflammation mediated apoptosis of liver via improving intestinal microbiota and bile acid metabolism. Food Chem. 404, 134643 (2023).
Wu, Y. et al. Galactooligosaccharides and Limosilactobacillus reuteri synergistically alleviate gut inflammation and barrier dysfunction by enriching Bacteroides acidifaciens for pentadecanoic acid biosynthesis. Nat. Commun. 15, 9291 (2024).
Xu, J. & Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5?. Bioinformatics 26, 889–895 (2010).
Chae, J. P., Pajarillo, E. A., Hwang, I. C. & Kang, D. K. Construction of a bile-responsive expression system in Lactobacillus plantarum. Food Sci. Anim. Resour. 39, 13–22 (2019).
Wu, J. et al. Clostridium butyricum alleviates weaned stress of piglets by improving intestinal immune function and gut microbiota. Food Chem. 405, 135014 (2023).
Kresse, G. & Hafner, J. Ab initio molecular dynamics for liquid metals. Phys. Rev. B 47, 558–561 (1993).
Kresse, G. & Hafner, J. Ab initio molecular-dynamics simulation of the liquid-metal–amorphous-semiconductor transition in germanium. Phys. Rev. B 49, 14251–14269 (1994).
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comput. Mater. Sci. 6, 15–50 (1996).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169–11186 (1996).
Kresse, G. & Joubert, D. From ultrasoft pseudopotentials to the projector augmented-wave method. Phys. Rev. B 59, 1758–1775 (1999).
Blöchl, P. E. Projector augmented-wave method. Phys. Rev. B 50, 17953–17979 (1994).
Perdew, J. P. et al. Erratum: Atoms, molecules, solids, and surfaces: applications of the generalized gradient approximation for exchange and correlation. Phys. Rev. B 48, 4978 (1993).
Perdew, J. P. et al. Atoms, molecules, solids, and surfaces: applications of the generalized gradient approximation for exchange and correlation. Phys. Rev. B 46, 6671–6687 (1992).
Vosko, S. H., Wilk, L. & Nusair, M. Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis. Can. J. Phys. 59, 1200 (1980).
Grimme, S. Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J. Comput. Chem. 27, 1787–1799 (2006).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Terranova, U. & de Leeuw, N. H. Aqueous Fe2S2 cluster: structure, magnetic coupling, and hydration behaviour from Hubbard U density functional theory. Phys. Chem. Chem. Phys. 16, 13426–13433 (2014).
Dzade, N. Y., Roldan, A. & de Leeuw, N. H. The surface chemistry of NOx on mackinawite (FeS) surfaces: a DFT-D2 study. Phys. Chem. Chem. Phys. 16, 15444–15456 (2014).
Santos-Carballal, D., Roldan, A., Grau-Crespo, R. & de Leeuw, N. H. First-principles study of the inversion thermodynamics and electronic structure of FeM2X4 (thio)spinels (M=Cr, Mn, Co, Ni; X=O, S). Phys. Rev. B 91, 195106 (2015).
Haider, S., Roldan, A. & de Leeuw, N. H. Catalytic dissociation of water on the (001), (011), and (111) surfaces of violarite, FeNi2S4: a DFT-D2 study. J. Phys. Chem. C 118, 1958–1967 (2014).
Dzade, N. Y., Roldan, A. & de Leeuw, N. H. Adsorption of methylamine on mackinawite (FES) surfaces: a density functional theory study. J. Chem. Phys. https://doi.org/10.1063/1.4822040 (2013).
Dzade, N. Y., Roldan, A. & de Leeuw, N. H. Activation and dissociation of CO2 on the (001), (011), and (111) surfaces of mackinawite (FeS): a dispersion-corrected DFT study. J. Chem. Phys. 143, 094703 (2015).
Hendrik, J. M. & James, D. P. Special points for Brillouin-zone integrations. Phys. Rev. B 13, 5188 (1976).
Blöchl, P. E., Jepsen, O. & Andersen, O. K. Improved tetrahedron method for Brillouin-zone integrations. Phys. Rev. B 49, 16223–16233 (1994).
Anisimov, V. I., Korotin, M. A., Zaanen, J. & Andersen, O. K. Spin bags, polarons, and impurity potentials in La2−xSrxCuO4 from first principles. Phys. Rev. Lett. 68, 345–348 (1992).
Dudarev, S. L., Botton, G. A., Savrasov, S. Y., Humphreys, C. J. & Sutton, A. P. Electron-energy-loss spectra and the structural stability of nickel oxide: an LSDA+U study. Phys. Rev. B 57, 1505–1509 (1998).
Haider, S., Grau-Crespo, R., Devey, A. J. & de Leeuw, N. H. Cation distribution and mixing thermodynamics in Fe/Ni thiospinels. Geochim. Cosmochim. Acta 88, 275–282 (2012).
Roldan, A., Santos-Carballal, D. & de Leeuw, N. H. A comparative DFT study of the mechanical and electronic properties of greigite Fe3S4 and magnetite Fe3O4. J. Chem. Phys. https://doi.org/10.1063/1.4807614 (2013).
Devey, A. J., Grau-Crespo, R. & de Leeuw, N. H. Electronic and magnetic structure of Fe3S4: GGA+U investigation. Phys. Rev. B 79, 195126 (2009).
Roldan, A. et al. Bio-inspired CO2 conversion by iron sulfide catalysts under sustainable conditions. Chem. Commun. 51, 7501–7504 (2015).
Tasker, P. W. The stability of ionic crystal surfaces. J. Phys. C 12, 4977–4984 (1979).
Santos-Carballal, D., Roldan, A., Grau-Crespo, R. & de Leeuw, N. H. A DFT study of the structures, stabilities and redox behaviour of the major surfaces of magnetite Fe3O4. Phys. Chem. Chem. Phys. 16, 21082–21097 (2014).
Devey, A. J., Grau-Crespo, R. & de Leeuw, N. H. Combined density functional theory and interatomic potential study of the bulk and surface structures and properties of the iron sulfide mackinawite (FeS). J. Phys. Chem. C 112, 10960–10967 (2008).
Lin, Z. S. et al. Nanozymes modulate probiotic tryptophan metabolism to prevent Salmonella infection in mammalian models. figshare https://doi.org/10.6084/m9.figshare.29652707 (2025).
Acknowledgements
We thank Z. Wang (Institute of Atomic and Molecular Physics, Jilin University) for their technical support in the DFT simulations. We thank P. He (State Key Laboratory of Animal Nutrition and Feeding, Department of Animal Nutrition and Feed Science, China Agricultural University) and A. Liu (State Key Laboratory of Animal Nutrition and Feeding, Department of Animal Nutrition and Feed Science, China Agricultural University) for liquid chromatography–mass spectrometry. We thank P. Liu (College of Animal Sciences, Fujian Agriculture and Forestry University) for the piglet sample collection. This work is supported by the National Key Research and Development Program of China (grant no. 2023YFD1301103 for B.D.), the Special Fund for Strategic Pilot Technology of Chinese Academy of Sciences (grant no. XDC0120200 for L.G.) and the National Natural Science Foundation of China (grant nos. 22121003 for L.G. and 32260859 for B.D.).
Author information
Authors and Affiliations
Contributions
B.D., L.G. and Z.L. conceived of this study. Z.L. performed the bioinformatics and statistical analyses on the sequencing data, and performed and analysed the results of redox potential assays, Raman spectrometry and FTIR. Z.L. and Y.F. designed the biological experiments and carried out the experiments with J.W., Q.W., L.W. and Y.Z. L.C., H.C., J.J. and Y.G. conducted the DFT calculations. Z.L., B.D. and L.Z.G. wrote the paper with input from Y.F., L.C., H.C. and Y.G. All authors approved the final draft of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Nico Claassens, Till Strowig and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Microbiota sequence enriches microbial ferroptosis pathway induced by nFeS.
a Relative protein expression of claudin-1, glutathione peroxidase (GPX4), and NADPH oxidase 1 (NOX1) in the gut epithelium (n = 3 samples). b Sobs index of alpha diversity between CON and nFeS group in genus and family levels (n = 5 piglets). c Venn plot analysis. d Relative abundance of Lactobacteriaceae in family level (n = 5 piglets). e Relative abundance of Fusicatenibacter in genus level (n = 5 piglets). f Antibacterial effect of nFeS to L. vaginalis. g, h MDA level (n = 5 L. vaginalis) and ROS species generation (n = 3 L. vaginalis) change in nFeS-treated L. vaginalis. i Growth curve of L. vaginalis. j Comparison of absorbance (OD600) of L. vaginalis treated with nFeS or Na2S3 (n = 3 L. vaginalis). k Correlation between bacteria killed by nFeS (related to Fig. 1l, m) with levels of serum DAO and IL-1β. l Faecal SCFA levels (n = 5 piglets). m Correlation between bacteria killed by nFeS (related to Fig. 1l, m) and faecal SCFA levels. n Spearman analysis between differential bacteria abundance and relative gene levels, serum DAO and IL-1β levels, and the functions of the bacterial communities (ferroptosis, pentose phosphate pathway, cytochrome c oxidase, and glutamate synthase). These functions of the bacterial communities were predicted with PICRUSt2 of 16S rRNA sequencing using the Kyoto Encyclopaedia of Genes and Genomes (KEGG) database and had significant difference between the CON and nFeS groups. P values were calculated using a two-sided unpaired t-test. Spearman analysis of k, m, n are at the two-sided 99% confidence interval, and the significance marked in the figures. Data were shown as the standard error of the mean. Exact P values of a-d, d-e, g, j, and l are displayed in the figure. Each experiment was repeated independently for three times with similar results.
Extended Data Fig. 2 Antibacterial action of nFeS towards S. aureus, S. Typhimurium, and E. coli correlates to ferrous iron.
a Antibacterial ability of nFeS was shown as antibacterial diameter in the agar plate included S. aureus, S. Typhimurium, and E. coli. DOX was doxycycline. The diameter of the inhibition zone is measured in millimetres. b Antibacterial ability of nFeS to suppress the growth of S. aureus, S. Typhimurium, and E. coli in liquid culture media (n = 3 bacteria per treatment). c Concentration of ferrous iron ion accumulated in the supernatant of S. aureus, S. Typhimurium, E. coli in the different doses of nFeS (n = 3 bacteria per treatment). d, e Antibacterial ability of FeCl2 and Na2S3 to suppress the growth of S. aureus, S. Typhimurium, and E. coli in liquid culture media, respectively (n = 3 bacteria per treatment). f Generation of ROS in S. Typhimurium with nFeS treatment (n = 3 bacteria per treatment). g Changes of MDA level in S. aureus, S. Typhimurium, and E. coli with nFeS treatment (n = 5 bacteria per treatment). P values were calculated using a two-sided unpaired t-test. Data were shown as the standard error of the mean. Exact P values are displayed in the figure. Each experiment was repeated independently for three times with similar results.
Extended Data Fig. 3 nFeS along with L. vaginalis improves gut epithelial heath, without inducing host ferroptosis.
a Faecal iron element was detected every day from the first gavage day to the 7th day of the end of gavage. Mice that received 5 mg/kg body weight (BW) nFeS for 7 days adapted a concentration balance of nFeS in the intestine, and excreted out by a-five-day after stopping gavage (n = 3 mice). b Changes of MDA level in gut epithelium of mice (n = 6 mice). c Relative mRNA levels of GPX4, FTH-1, NCOA4, ACSL4, NOX1, and COX2 in gut epithelium of mice (n = 6 mice). d Changes of colonic length (n = 6 mice). e Serum IL-1β and IL-6 levels (n = 6 mice). f Faecal SCFA levels (n = 6 mice). g Relative protein levels of FTH-1 and GPX4 in the gut epithelium of mice (n = 3 samples). h Relative mRNA levels of COX2, ACSL4, and NCOA4 in gut epithelium of mice (n = 6 mice). i Changes of MDA level in gut epithelium of mice (n = 6 mice). j ROS generation in the gut epithelial cells of mice (n = 3 mice). P values were calculated using ordinary one-way ANOVA followed by Tukey’s test (b-d, g) or Benjamini, Krieger & Yekutieli (e, f) for multiple comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001). Each experiment was repeated independently for three times with similar results.
Extended Data Fig. 4 Antibacterial tryptophan-indole metabolites of L. vaginalis are beyond 3-HAA, ILA, I3CA, and metabolites of serotonin pathway.
a, b Antibacterial abilities of L. vaginalis (included supernatant and sediment) towards S. aureus, S. Typhimurium, and E. coli tested in agar plate (n = 3 L. vaginalis). c PCoA of Bray-Curtis dissimilarity matrices and Venn plot were generated from the non-targeted metabolomics data of L. vaginalis, weaned piglets, and mice. d Kynurenine and serotonin pathways are showed as microbial pathways in tryptophan pathway27. 3-HAA is the end product of the kynurenine/IDO signalling pathway, and 5HIAA and melatonin are the end products of the serotonin pathway. e Relative abundance of 3-HAA, 5-HTP, 5-HT, 5-HIAA, and melatonin in L. vaginalis, weaned piglets, and mice. 3-HAA and 5-HT were not captured in L. vaginalis metabolites. 5-HTP was not captured in metabolites of weaned piglets. 5-HIAA and melatonin were not detected in metabolites of weaned piglets and L. vaginalis (n = 4 (CON/SM + LV) or 5 (SM/SM + LV+nFeS) mice per group, 5 piglets per group, and 6 L. vaginalis per group). f Common pathway, I3C/I3CA/I2CA, in L. vaginalis, weaned piglets, and mice. g Antibacterial ability of ILA to inhibit growth of S. aureus, S. Typhimurium, and E. coli. h Antibacterial abilities of I3CA alone and I3CA co-treated with nFeS to inhibit growth of S. aureus, S. Typhimurium, and E. coli (n = 3 samples). P values were calculated using a two-sided unpaired t-test for two-group comparisons and ordinary one-way ANOVA followed Benjamini, Krieger & Yekutieli for multiple comparisons. Data were shown as the standard error of the mean. The diameter of the inhibition zone is measured in millimetres (mm). Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001). Each experiment was repeated independently for three times with similar results. Panel f was created with BioRender.com.
Extended Data Fig. 5 nFeS enhances the transcription level of sugar transporters and the metabolic level of TCA cycle to promote L. vaginalis viability.
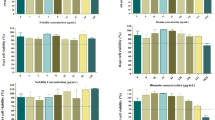
a PCoA analysis of transcriptome in the L. vaginalis on Bray-Curtis dissimilarity matrices. b Venn plot of transcriptome in the L. vaginalis between the CON and nFeS group. c Changes of differential up-regulated genes in L. vaginalis with nFeS treatment among the first 20 altered genes. d KEGG enrichment of transcriptome in L. vaginalis with nFeS treatment. e Changes of monosaccharide compounds in the metabolic pathway of L. vaginalis (n = 6 L. vaginalis). f Relative abundance of significantly changed intermediate metabolites of TCA cycle in L. vaginalis (n = 6 L. vaginalis). g Changes of saccharide compounds in the metabolic pathway of weaned piglets (n = 5 piglets). h Relative abundance of significant change intermediate metabolites of TCA cycle in weaned piglets (n = 5 piglets). i Changes of monosaccharide compounds in the metabolic pathway of mice (n = 4 (CON/SM + LV) or 5 (SM/SM + LV+nFeS) mice per group). j Relative abundance of significant change intermediate metabolites of TCA cycle in mice (n = 4 (CON/SM + LV) or 5 (SM/SM + LV+nFeS) mice per group). P values were calculated using a two-sided unpaired t-test for two-group comparisons and ordinary one-way ANOVA followed Benjamini, Krieger & Yekutieli for multiple comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001). Each experiment was repeated independently for three times with similar results. Panels e, g and i were created with BioRender.com.
Extended Data Fig. 6 Alcohol dehydrogenase is the upstream enzyme to produce I3C as the substrate for nFeS catalysis in L. vaginalis.
a Amino acid sequences of methanol dehydrogenase in Lactobacillus spp. were compared with 7 kinds of alcohol dehydrogenases (adH1-7) in L. vaginalis by TM-align comparisons. The adH3 gene was identified to potentially be the gene encoding methanol dehydrogenase in L. vaginalis. b Establishment of pUC19-pLV-adH-OE and pUC19-adH-Mut recombinant plasmids to overexpress or knock out adH3 (L. vaginalisOE or L. vaginalisMut). c Relative mRNA level of adH (n = 3 L. vaginalisWT/OE/Mut). d Detection of I3C, I3CA, and I2CA concentration in L. vaginalisOE and L. vaginalisMut (supernatant and sediment) (n = 3 L. vaginalisWT/OE/Mut). e Detection of I3C, I3CA, and I2CA concentration in L. vaginalisWT, L. vaginalisOE, and L. vaginalisMut (bacteria sediment) receiving nFeS treatment (n = 3 L. vaginalisWT/OE/Mut). f Characterization of nFeS in L. vaginalis by transmission electron microscopy (Scale bar at 100 nm). P values were calculated using a two-sided unpaired t-test for two-group comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure. Each experiment was repeated independently for three times with similar results. Panel b was created with BioRender.com.
Extended Data Fig. 7 Recyclability of nFeS in the conversion from I3C to I2CA confirms its catalytic behavior.
a nFeS and Fe3O4 were reacted with I3C and I3CA, respectively, then recycled nFeS and Fe3O4 to next reaction with new weigh I3C and I3CA. b-e I3C, I3CA, and I2CA concentration at the end of five times of recycled reaction. nFeS is represented as Fe3S4 here. f Different concentrations of I3C as substrates reacted with nFeS (After 5 times recycled), the generation rate of I2CA per unit time was used to fit the Michaelis–Menten curve of enzyme kinetic reaction. g Concentration changes of I2CA in the reaction between I3C and nFeS during 240 min. h Redox potential of nFeSOx/nFeSRe and I3CA/I3C couples. i Standard redox potentials of NAD+/NADH, nFeSOx2/nFeSRe2, O2/HO2−, I3CA/I3C, O2/OH−, nFeSOx1/nFeSRe1, O2/H2O2, and O2/H2O. Ox represents the oxidation state. Re indicates the reduced state. Each experiment was repeated independently for three times with similar results. Panel a was created with BioRender.com.
Extended Data Fig. 8 I2CA enables ferrous iron outside of L. vaginalis and synergizes with nFeS against pathogenic bacteria.
a Concentration of ferrous ions accumulated in the supernatant of L. vaginalis (removed the nFeS) in the different doses of nFeS. b Concentration of ferrous ions accumulated in the supernatant of L. vaginalisWT, L. vaginalisOE, L. vaginalisMut. c Changes of intracellular pH of bacteria in different extracellular pH conditions. d Intracellular pH values of different bacteria under nFeS treatment. e Changes of intracellular pH of bacteria, which were allowed to grow to a relative density (OD600 = 0.8) followed by nFeS treatment (1 mg/ml) and/or I2CA (1 mg/ml) for 12 hours. f Characterization of bactericidal abilities of I2CA, I3CA, combined utilization of I2CA and nFeS to S. aureus, S. Typhimurium, E. coli (after treating 8 hours), scale bar at 1 µm. g Level of AKP in the supernatant of S. aureus, S. Typhimurium, E. coli (after treating 24 hours) (n = 6 (CON/12CA/12CA+nFeS) or n = 4 (I3CA) samples per group). h Turbidity and growth of S. aureus, S. Typhimurium, E. coli in treatment of I2CA, I3CA, I2CA+nFeS for 24 hours, respectively. P values were calculated using ordinary one-way ANOVA followed Benjamini, Krieger & Yekutieli for multiple comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001). Each experiment was repeated independently for three times with similar results.
Extended Data Fig. 9 nFeS is biocompatible with multiple probiotics.
Relative abundance of significant changed probiotics in colonic chyme of weaned piglets (n = 5 piglets). P values were calculated using ordinary one-way ANOVA or Kruskal-Wallis test followed Benjamini, Krieger & Yekutieli for multiple comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001).
Extended Data Fig. 10 Pathogenic and opportunistic pathogenic bacteria are sensitive to single or combined nFeS with L. vaginalis.
Relative abundance of significant changed pathogenic and opportunistic bacteria in colonic chyme of weaned piglets (n = 5 piglets). P values were calculated using ordinary one-way ANOVA followed Benjamini, Krieger & Yekutieli for multiple comparisons. Data were shown as the standard error of the mean. Exact P values are displayed in the figure (P < 0.0001; the exact P value is < 0.0001).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–10, Results 1–6 and Tables 1–8.
Source data
Source Data Figs. 1–6 (download XLSX )
Statistical source data for Figs. 1–6.
Source Data Figs. 1, 2, 5 and 6 (download PDF )
Uncropped microscopy images for Figs. 1, 2, 5, and 6.
Source Data Extended Data Fig. 1-8 (download XLSX )
Statistical source data for Extended Data Figs. 1–8.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Figs. 6 and 8 (download PDF )
Uncropped microscopy images for Extended Data Figs. 6f and 8f.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, Z., Feng, Y., Chen, L. et al. Nanozymes modulate probiotic tryptophan metabolism to prevent Salmonella infection in mammalian models. Nat Microbiol 10, 3272–3289 (2025). https://doi.org/10.1038/s41564-025-02176-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02176-4