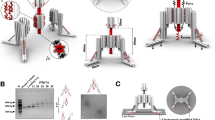
Abstract
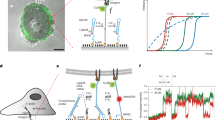
Force-induced changes in protein structure and function mediate cellular responses to mechanical stresses that are important in human development, physiology and diseases. However, existing methods to study proteins under mechanical force are generally single-molecule techniques unsuitable for biochemical and structural analysis. Taking advantage of DNA nanotechnology, including the well-defined geometry of DNA origami and the programmable mechanics of DNA hairpins, we built a nanodevice to apply controlled forces to proteins. This device was used to study the R1-R2 segment of the talin1 rod domain as a model protein. R1-R2 consists of two α-helical bundles that reversibly unfold under tension to expose binding sites for the cytoskeletal protein vinculin. Electron microscopy confirmed tension-dependent protein extension, and biochemical analysis demonstrated enhanced vinculin binding under tension. Using the device in pull-down assays with cell lysates, we identified filamins as novel tension-dependent talin binders. The DNA nanodevice thus provides a valuable molecular tool for studying mechanosensitive proteins on a biochemical scale.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the data that support the findings of this study are available within the article and its Supplementary Information. Source data (statistical data and uncropped gels and blots for Figs. 1–5 and Extended Data Figs. 1–4, 6–8 and 10) are provided with this paper. Source data are provided with this paper.
References
Mierke, C. T. Extracellular matrix cues regulate mechanosensing and mechanotransduction of cancer cells. Cells 13, 96 (2024).
Orr, A. W., Helmke, B. P., Blackman, B. R. & Schwartz, M. A. Mechanisms of mechanotransduction. Dev. Cell 10, 11–20 (2006).
Hoffman, B. D., Grashoff, C. & Schwartz, M. A. Dynamic molecular processes mediate cellular mechanotransduction. Nature 475, 316–323 (2011).
Jin, P., Jan, L. Y. & Jan, Y. N. Mechanosensitive ion channels: structural features relevant to mechanotransduction mechanisms. Annu Rev. Neurosci. 43, 207–229 (2020).
Romani, P., Valcarcel-Jimenez, L., Frezza, C. & Dupont, S. Crosstalk between mechanotransduction and metabolism. Nat. Rev. Mol. Cell Biol. 22, 22–38 (2021).
Grashoff, C. et al. Measuring mechanical tension across vinculin reveals regulation of focal adhesion dynamics. Nature 466, 263–266 (2010).
Hu, Y. et al. DNA-based ForceChrono probes for deciphering single-molecule force dynamics in living cells. Cell 187, 3445–3459.e15 (2024).
Ren, Y. et al. Force redistribution in clathrin-mediated endocytosis revealed by coiled-coil force sensors. Sci. Adv. 9, eadi1535 (2023).
Tao, A. et al. Identifying constitutive and context-specific molecular-tension-sensitive protein recruitment within focal adhesions. Dev. Cell 58, 522–534.e7 (2023).
Zhang, Y., Ge, C., Zhu, C. & Salaita, K. DNA-based digital tension probes reveal integrin forces during early cell adhesion. Nat. Commun. 5, 5167 (2014).
Fisher, T. E., Oberhauser, A. F., Carrion-Vazquez, M., Marszalek, P. E. & Fernandez, J. M. The study of protein mechanics with the atomic force microscope. Trends Biochem. Sci. 24, 379–384 (1999).
Bustamante, C. J., Chemla, Y. R., Liu, S. & Wang, M. D. Optical tweezers in single-molecule biophysics. Nat. Rev. Methods Primers 1, 25 (2021).
Choi, H. K., Kim, H. G., Shon, M. J. & Yoon, T. Y. High-resolution single-molecule magnetic tweezers. Annu. Rev. Biochem. 91, 33–59 (2022).
Ishijima, A. et al. Simultaneous observation of individual ATPase and mechanical events by a single myosin molecule during interaction with actin. Cell 92, 161–171 (1998).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Dey, S. et al. DNA origami. Nat. Rev. Methods Primers 1, 13 (2021).
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2018).
Fisher, P. D. E. et al. A programmable DNA origami platform for organizing intrinsically disordered nucleoporins within nanopore confinement. ACS Nano 12, 1508–1518 (2018).
Fu, J. et al. Multi-enzyme complexes on DNA scaffolds capable of substrate channelling with an artificial swinging arm. Nat. Nanotechnol. 9, 531–536 (2014).
Zeng, Y. C. et al. Fine tuning of CpG spatial distribution with DNA origami for improved cancer vaccination. Nat. Nanotechnol. 19, 1055–1065 (2024).
Mills, A. et al. A modular spring-loaded actuator for mechanical activation of membrane proteins. Nat. Commun. 13, 3182 (2022).
Nickels, P. C. et al. Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354, 305–307 (2016).
Wang, Y. et al. A nanoscale DNA force spectrometer capable of applying tension and compression on biomolecules. Nucleic Acids Res. 49, 8987–8999 (2021).
Darcy, M. et al. High-force application by a nanoscale DNA force spectrometer. ACS Nano 16, 5682–5695 (2022).
Praetorius, F. et al. Biotechnological mass production of DNA origami. Nature 552, 84–87 (2017).
Jia, Y. L., Chen, L. M., Liu, J., Li, W. & Gu, H. Z. DNA-catalyzed efficient production of single-stranded DNA nanostructures. Chem 7, 959–981 (2021).
Kramm, K. et al. DNA origami-based single-molecule force spectroscopy elucidates RNA polymerase III pre-initiation complex stability. Nat. Commun. 11, 2828 (2020).
Sun, Z., Guo, S. S. & Fassler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 215, 445–456 (2016).
Goult, B. T., Yan, J. & Schwartz, M. A. Talin as a mechanosensitive signaling hub. J. Cell Biol. 217, 3776–3784 (2018).
Hytonen, V. P. & Vogel, V. How force might activate talin’s vinculin binding sites: SMD reveals a structural mechanism. PLoS Comput. Biol. 4, e24 (2008).
Yao, M. et al. Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation. Sci. Rep. 4, 4610 (2014).
Yao, M. et al. The mechanical response of talin. Nat. Commun. 7, 11966 (2016).
Papagrigoriou, E. et al. Activation of a vinculin-binding site in the talin rod involves rearrangement of a five-helix bundle. EMBO J. 23, 2942–2951 (2004).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Funke, J. J. & Dietz, H. Placing molecules with Bohr radius resolution using DNA origami. Nat. Nanotechnol. 11, 47–52 (2016).
Xiong, Q. et al. DNA origami post-processing by CRISPR-Cas12a. Angew. Chem. Int. Ed. 59, 3956–3960 (2020).
Aksel, T., Yu, Z., Cheng, Y. & Douglas, S. M. Molecular goniometers for single-particle cryo-electron microscopy of DNA-binding proteins. Nat. Biotechnol. 39, 378–386 (2021).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Wagenbauer, K. F. et al. How we make DNA origami. ChemBioChem 18, 1873–1885 (2017).
Poppleton, E. et al. Design, optimization and analysis of large DNA and RNA nanostructures through interactive visualization, editing and molecular simulation. Nucleic Acids Res. 48, e72 (2020).
Woodside, M. T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl Acad. Sci. USA 103, 6190–6195 (2006).
Liu, J. & Yan, J. Unraveling the dual-stretch-mode impact on tension gauge tethers’ mechanical stability. J. Am. Chem. Soc. 146, 7266–7273 (2024).
Bercy, M. & Bockelmann, U. Hairpins under tension: RNA versus DNA. Nucleic Acids Res. 43, 9928–9936 (2015).
Marko, J. F. & Siggia, E. D. Stretching DNA. Macromolecules 28, 8759–8770 (1995).
Kumar, A. et al. Talin tension sensor reveals novel features of focal adhesion force transmission and mechanosensitivity. J. Cell Biol. 213, 371–383 (2016).
Austen, K. et al. Extracellular rigidity sensing by talin isoform-specific mechanical linkages. Nat. Cell Biol. 17, 1597–1606 (2015).
Chung, M., Zhou, K., Powell, J. T., Lin, C. & Schwartz, M. A. DNA-based molecular clamp for probing protein interactions and structure under force. ACS Nano 18, 27590–27596 (2024).
Lin, C., Perrault, S. D., Kwak, M., Graf, F. & Shih, W. M. Purification of DNA-origami nanostructures by rate-zonal centrifugation. Nucleic Acids Res. 41, e40 (2013).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Evans, E. & Ritchie, K. Strength of a weak bond connecting flexible polymer chains. Biophys. J. 76, 2439–2447 (1999).
Zhou, J., Kang, X., An, H., Lv, Y. & Liu, X. The function and pathogenic mechanism of filamin A. Gene 784, 145575 (2021).
Kumar, A. et al. Filamin A mediates isotropic distribution of applied force across the actin network. J. Cell Biol. 218, 2481–2491 (2019).
Aissaoui, N. et al. Modular imaging scaffold for single-particle electron microscopy. ACS Nano 15, 4186–4196 (2021).
Pfaff, M., Liu, S., Erle, D. J. & Ginsberg, M. H. Integrin beta cytoplasmic domains differentially bind to cytoskeletal proteins. J. Biol. Chem. 273, 6104–6109 (1998).
Rief, M., Clausen-Schaumann, H. & Gaub, H. E. Sequence-dependent mechanics of single DNA molecules. Nat. Struct. Biol. 6, 346–349 (1999).
Amiram, M. et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 33, 1272–1279 (2015).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2010).
Driscoll, T. P., Ahn, S. J., Huang, B., Kumar, A. & Schwartz, M. A. Actin flow-dependent and -independent force transmission through integrins. Proc. Natl Acad. Sci. USA 117, 32413–32422 (2020).
Chanduri, M. et al. Cellular stiffness sensing through talin 1 in tissue mechanical homeostasis. Sci. Adv. 10, eadi6286 (2024).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Yan, J., Yao, M., Goult, B. T. & Sheetz, M. P. Talin dependent mechanosensitivity of cell focal adhesions. Cell. Mol. Bioeng. 8, 151–159 (2015).
Acknowledgements
We thank Yl Zhang for helping with the statistical thermodynamic model. This work was supported by a National Institutes of Health grant (R01HL155543) to M.A.S. and C.L., a Department of Defense/Army Research Office MURI grant (W911NF1410403) to M.A.S., a Smith Family Foundation Odyssey Award to C.L., and a fellowship from the American Heart Association (20POST35080107) to M.C.
Author information
Authors and Affiliations
Contributions
M.A.S. conceived the project. K.Z., M.C., J.T.P., M.A.S. and C.L. designed the approach. K.Z., M.C., S.P., J.T.P. and Q.Y. performed experiments. K.Z., M.C. and J.C. analysed data. J.L., Y.X., C.L. and M.A.S. provided resources and supervised the study. K.Z. and C.L. wrote the original draft. K.Z., M.C., M.A.S. and C.L. edited the paper. All authors participated in the discussions and approved the paper. K.Z. and M.C. contributed equally.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Zheng Liu, Alf Mansson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Design and validation of the left DNA Spring.
a) Schematics showing sequences of a reconfigurable handle (left, that is, DNA spring) and a fixed handle (right) in a DNA origami device. The reconfigurable handle is converted from extended to folded state by TMSD. b) Schematics showing the formation of a tension-loaded (that is, extended) spring and its activation. HP: hairpin strand with a 17-bp stem (76.5% GC content) and a 14-nt loop; TR: target strand that binds to HP to form tension-loaded spring; HD: displacement strand that triggers HP folding. Minimum free energy (MFE) structures are predicted by NUPACK (25 °C, 100 mM Na+ and 12 mM Mg2+). Base pair mismatches are designed to lower energy barriers for HP/TR binding and TMSD. c) Validation of reactions shown in b by non-denaturing PAGE. TR is Cy5-labeled. HP*: A 105-nt staple strand carrying the HP sequence and BG-tether-binding sequence.
Extended Data Fig. 2 Fluorophore-quencher based DNAts.
a) DNAts (F1/2 ≈ 8.3 pN) consisting of a 12-bp stem (17% GC content) and a dT8 loop. Its maximum end-to-end distance in the folded state is ~16 nm, accounting for the lengths of BHQ1-DNA and AF488-DNA (21 nt each) and the stem diameter (2 nm). b) A snapshot of DNAts simulation by oxDNA. The relaxed 3-arm junction leads to a smaller end-to-end distance. c) Validation of DNAts assembly (lanes 1–4) by non-denaturing PAGE. As a control, an oligonucleotide with complementary stem-loop sequence was added in 10-fold excess during assembly (lanes 5–8). d) DNAts incorporation and unfolding in DNA devices with a 12.8-pN spring and various gap sizes examined by agarose gel electrophoresis (upper: AF488 channel, middle: Cy5 channel, lower: ethidium bromide channel). DNA devices with DNAts missing BHQ1 (Ctrl) and with complete DNAts before (-F) and after (+F) TMSD were electrophoresed. Note that DNAts in the 9 and 12 nm gap, but not the 16 or 6 nm gap, unzipped efficiently and that DNA devices with a 16 nm gap contained more quencher-free DNAts than those with a narrower gap. Part of the gel (9 nm gap) is shown in Fig. 1c.
Extended Data Fig. 3 Rate of DNAts unfolding and TMSD.
a) Schematics of the experiment setup for simultaneously monitoring DNAts unfolding and target strand displacement by AF488 and Cy5 dequenching, respectively. AF488: Green star; Cy5: light purple star; BHQ1: black circle; Iowa Black RQ: black square. b) Traces of AF488 (green) and Cy5 (light purple) fluorescence increase following the addition of displacement strand (HD, marked by arrows) at various concentrations (1.5–75: 1 molar ratio to target strand). DNAts with theoretical unfolding forces of 8.3 and 6.1 pN were mounted in DNA devices with a 9-nm gap and a 12.8-pN DNA spring. Reaction half time (t1/2) is defined as the time for fluorescence to reach half of the maximum. Curves are guide for the eye.
Extended Data Fig. 4 Mounting and unzipping DNAts in DNA origami device.
a) Incorporation efficiencies of DNAts (F1/2 ≈ 8.3 pN) into DNA origami device (gap=9 nm) assessed by agarose gel electrophoresis at increasing DNAts:origami molar ratios. b) Incorporation efficiencies of the 8.3 pN DNAts into DNA origami devices with various handle configurations (see schematics below the gel images) assessed by agarose gel electrophoresis. DNAts is labeled with AF488 but without BHQ1; DNA origami device is labeled with Atto647N. Comparing the AF488/Atto647N ratios of the DNA device bands shows ~16% higher AF488 signal on DNA devices with two extended handles (gap: 9 nm) than those with only one handle, which we attribute to ~16% of the devices erroneously capturing two DNAts in its cavity. c) DNAts designed with different GC contents and/or mismatch to tune their unfolding forces. d) Left: images of agarose gel (upper: AF488 channel, middle: Cy5 channel, lower: ethidium bromide channel) in which the DNA devices with DNAts missing BHQ1 (Ctrl) and with complete DNAts before (-F) and after (+F) TMSD were electrophoresed. Right: DNAts fluorescence quantified from the gel images (normalized to DNAts without BHQ1). DNA spring F1/2 ≈ 12.8 pN; gap=9 nm. The data for unzipping 8.3 pN DNAts (also shown in Fig. 1c) are shown here again for comparison.
Extended Data Fig. 5 TEM characterization of S-R1-R2-H and Origami-R1-R2.
a) an AlphaFold model (left; SNAP-tag: green, R1-R2: cyan, HaloTag: red) and an electron micrograph of S-R1-R2-H. b) protein end-to-end distances (Figs. 3c, 3d) fitted with Gaussian functions. c) and d) show Origami-R1-R2 before (5,748 auto-picked particles) and after (7,185 auto-picked particles) force application, respectively. Circles in the zoomed-out images contain protein molecules selected for averaging. Averaged images are centered on the U-frame cavity where S-R1-R2-H is expected. Particle numbers are shown for each class. Selected averaged TEM images are shown in Fig. 3. Scale bars: a. 50 nm, c. and d. 100 nm. Box sizes: c. 30 nm, d. 50 nm.
Extended Data Fig. 6 DNAts stretched by various spring configurations.
a) DNAts (F1/2 ≈ 8.3 pN) pulled by single-spring devices examined by agarose gel electrophoresis. DNAts was mounted in a 9-nm gap and pulled by an unstructured DNA strand (max tension ≈ 6.1 pN), a weak hairpin (HP, F1/2 ≈ 8.8 pN) or a strong HP (F1/2 ≈ 12.8 pN). DNA devices with DNAts missing BHQ1 (Ctrl) and with complete DNAts before (-F) and after (+F) spring activation were electrophoresed. b) DNAts (F1/2 ≈ 8.3 pN) pulled by dual-spring devices (6-nm gap) examined by agarose gel electrophoresis. DNA devices with DNAts missing BHQ1 (Ctrl) and with complete DNAts pulled by the neither (L-/R-), left (L+/R-), right (L-/R+) or both springs (L+/R+) were electrophoresed. Spring activation via TMSD was performed at room temperature (21 °C) for samples on the left gel and at 37 °C for the right. Bar graphs in a and b show DNAts fluorescence quantified from the corresponding gel images (normalized to DNAts without BHQ1).
Extended Data Fig. 7 Controls and quantification for pulldown experiments.
a) Vinculin knockdown efficiency. Normalizing the vinculin band intensities by the GAPDH band intensities on a western blot of wild-type (WT) and vinculin-depleted (Vin-/-) cell lysates shows a knockdown efficiency of ~87%. b) FlnA-GFP pulldown by an elastic peptide (GPGGA)8, dubbed F40, shows no apparent differences on western blot before (-F) and after (+F) force application to F40. The gel on the right contains 8 times as much sample as the left. c) Western blot showing force-activated R1-R2 binding to FlnA-GFP, but not GFP, in lysates of NIH3T3 cells overexpressing the corresponding protein. Ctrl: DNA device without proteins, -F/+F: Origami-R1-R2 before/after force application. d) Purified FlnA-GFP examined by SDS–PAGE. e) SDS–PAGE of purified S-R1-R2-H (unconjugated, 87.3 kDa) and VinD1-FLAG (expressed in E. coli, 33.5 kDa) loaded with a range of known concentrations, as well as the mixture of a pulldown experiment loaded in two lanes with 2-fold concentration difference. f) Determining concentrations of S-R1-R2-H (top) and VinD1-FLAG (bottom) in the pulldown sample. Band intensities of the purified proteins (black dots) measured from e were used to construct calibration lines by linear regression. The intensities of the corresponding protein bands in the pulldown sample (crosses, measured from the 1× pulldown lane) were then used to determine the concentrations of S-R1-R2-H released from DNA devices and the pulled-down VinD1-FLAG (expressed in 293 T cells, 30.9 kDa). The resulting VinD1:R1R2 molar ratio is 163 nM/412 nM≈0.40. For experiments in b, c and e, DNA spring F1/2 ≈ 12.8 pN, gap=6 nm.
Extended Data Fig. 8 Design and validation of the right DNA spring.
a) Schematics showing the formation of a tension-loaded (that is, extended) right spring and its activation. HP: hairpin strand with a 12-bp stem (75% GC content) and a 7-nt loop; TR: target strand that binds to HP to form tension-loaded spring; HD: displacement strand that triggers HP folding. Minimum free energy (MFE) structures are predicted by NUPACK (25 °C, 100 mM Na+ and 12 mM Mg2+). Base pair mismatches are designed to lower energy barriers for HP/TR binding and TMSD. b) Validation of reactions shown in a by non-denaturing PAGE.
Extended Data Fig. 9 TEM images of S-R1-R2-H in the dual-spring DNA device.
The four groups of images show proteins in the relaxed (L-/R-) and various extended conditions (L+/R-, L-/R+ , L+/R+ ). L/R denotes the left/right springs and -/+ denotes the loaded/activated states. Scale bars for zoomed-in and zoomed out images are 50 and 100 nm, respectively.
Extended Data Fig. 10 Effect of Y27632 on intracellular filamin A-talin colocalization.
Cells were seeded on fibronectin (10 µg/mL) for 8 h and treated with DMSO (control) and 10 µM of Y27632 (ROCK inhibitor) for 15 min, washed and fixed. Top: Representative images of cells treated with DMSO (left) and Y27632 (right). Immuno-fluorescence of filamin A and talin are pseudo-colored cyan and magenta, respectively. Images are 92 by 92 µm2. Below: Quantification of the fraction of FA-localized filamin A (left), FA size (middle), and filamin A to talin ratio at FA (right) in cells treated with DMSO and Y27632. Group means were compared using one-way ANOVA (n = 78 DMSO-treated cells, n = 54 inhibitor-treated cells). Data are presented as mean ± s.d.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Table 1 (download XLSX )
Oligonucleotides for building and operating the DNA nanodevice.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, K., Chung, M., Pandey, S. et al. DNA nanodevice for analysis of force-activated protein extension and interactions. Nat. Nanotechnol. 21, 259–267 (2026). https://doi.org/10.1038/s41565-025-02086-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41565-025-02086-w