Abstract
Mitochondrial stress pathways protect mitochondrial health from cellular insults1,2,3,4,5,6,7,8. However, their role under physiological conditions is largely unknown. Here, using 18 single, double and triple whole-body and tissue-specific knockout and mutant mice, along with systematic mitochondrial morphology analysis, untargeted metabolomics and RNA sequencing, we discovered that the synergy between two stress-responsive systems—the ubiquitin E3 ligase Parkin and the metalloprotease OMA1—safeguards mitochondrial structure and genome by mitochondrial fusion, mediated by the outer membrane GTPase MFN1 and the inner membrane GTPase OPA1. Whereas the individual loss of Parkin or OMA1 does not affect mitochondrial integrity, their combined loss results in small body size, low locomotor activity, premature death, mitochondrial abnormalities and innate immune responses. Thus, our data show that Parkin and OMA1 maintain a dual regulatory mechanism that controls mitochondrial fusion at the two membranes, even in the absence of extrinsic stress.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Supplementary Fig. 1 includes uncropped immunoblot images corresponding to Figs. 3g and 4l, as well as to Extended Data Figs. 6a,c,e, 8b and 10a, with molecular weight markers indicating how blots were cropped for the final figures. RNA-seq data generated in this study are available in Gene Expression Omnibus under accession code PRJNA1157386. Materials generated in this study are available from the corresponding authors on request. Source data are provided with this paper.
Code availability
We did not generate any custom code for the RNA-seq analyses in this study; we utilized software packages with prewritten code and default parameters. We performed pairwise differential gene expression analysis using the R package DESeq2 1.20.0 and applied the Benjamini–Hochberg method for false discovery rate correction through RNAseqChef 1.1.2.
References
Konig, T. et al. MIROs and DRP1 drive mitochondrial-derived vesicle biogenesis and promote quality control. Nat. Cell Biol. 23, 1271–1286 (2021).
Yang, Q. et al. LONP-1 and ATFS-1 sustain deleterious heteroplasmy by promoting mtDNA replication in dysfunctional mitochondria. Nat. Cell Biol. 24, 181–193 (2022).
Vargas, J. N. S., Hamasaki, M., Kawabata, T., Youle, R. J. & Yoshimori, T. The mechanisms and roles of selective autophagy in mammals. Nat. Rev. Mol. Cell Biol. 24, 167–185 (2023).
Murata, D., Arai, K., Iijima, M. & Sesaki, H. Mitochondrial division, fusion and degradation. J. Biochem. 167, 233–241 (2020).
Friedman, J. R. & Nunnari, J. Mitochondrial form and function. Nature 505, 335–343 (2014).
Youle, R. J. & van der Bliek, A. M. Mitochondrial fission, fusion, and stress. Science 337, 1062–1065 (2012).
Murata, D., Roy, S., Lutsenko, S., Iijima, M. & Sesaki, H. Slc25a3-dependent copper transport controls flickering-induced Opa1 processing for mitochondrial safeguard. Dev. Cell 59, 2578–2592 (2024).
Murata, D. et al. Mitochondrial Safeguard: a stress response that offsets extreme fusion and protects respiratory function via flickering-induced Oma1 activation. EMBO J. 39, e105074 (2020).
Prashar, A. et al. Lysosomes drive the piecemeal removal of mitochondrial inner membrane. Nature 632, 1110–1117 (2024).
Koyano, F. et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510, 162–166 (2014).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Sauve, V. et al. Mechanism of parkin activation by phosphorylation. Nat. Struct. Mol. Biol. 25, 623–630 (2018).
Trempe, J. F. et al. Structure of parkin reveals mechanisms for ubiquitin ligase activation. Science 340, 1451–1455 (2013).
Sarraf, S. A. et al. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496, 372–376 (2013).
Giacomello, M., Pyakurel, A., Glytsou, C. & Scorrano, L. The cell biology of mitochondrial membrane dynamics. Nat. Rev. Mol. Cell Biol. 21, 204–224 (2020).
Dorn, G. W. 2nd Mitofusins as mitochondrial anchors and tethers. J. Mol. Cell. Cardiol. 142, 146–153 (2020).
Panicker, N., Ge, P., Dawson, V. L. & Dawson, T. M. The cell biology of Parkinson’s disease. J. Cell Biol. 220, e202012095 (2021).
Pallanck, L. & Greenamyre, J. T. Neurodegenerative disease: pink, parkin and the brain. Nature 441, 1058 (2006).
Kitada, T. et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608 (1998).
Valente, E. M. et al. Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 304, 1158–1160 (2004).
Ehses, S. et al. Regulation of OPA1 processing and mitochondrial fusion by m-AAA protease isoenzymes and OMA1. J. Cell Biol. 187, 1023–1036 (2009).
Head, B., Griparic, L., Amiri, M., Gandre-Babbe, S. & van der Bliek, A. M. Inducible proteolytic inactivation of OPA1 mediated by the OMA1 protease in mammalian cells. J. Cell Biol. 187, 959–966 (2009).
Ishihara, N., Fujita, Y., Oka, T. & Mihara, K. Regulation of mitochondrial morphology through proteolytic cleavage of OPA1. EMBO J. 25, 2966–2977 (2006).
Wai, T. et al. Imbalanced OPA1 processing and mitochondrial fragmentation cause heart failure in mice. Science 350, aad0116 (2015).
Acin-Perez, R. et al. Ablation of the stress protease OMA1 protects against heart failure in mice. Sci. Transl. Med. 10, eaan4935 (2018).
Guo, X. et al. Mitochondrial stress is relayed to the cytosol by an OMA1-DELE1-HRI pathway. Nature 579, 427–432 (2020).
Fessler, E. et al. A pathway coordinated by DELE1 relays mitochondrial stress to the cytosol. Nature 579, 433–437 (2020).
Shammas, M. K. et al. OMA1 mediates local and global stress responses against protein misfolding in CHCHD10 mitochondrial myopathy. J. Clin. Invest. 132, e157504 (2022).
Ahola, S. et al. OMA1-mediated integrated stress response protects against ferroptosis in mitochondrial cardiomyopathy. Cell Metab. 34, 1875–1891 (2022).
Lee, Y., Dawson, V. L. & Dawson, T. M. Animal models of Parkinson’s disease: vertebrate genetics. Cold Spring Harb. Perspect. Med. 2, a009324 (2012).
Quiros, P. M. et al. Loss of mitochondrial protease OMA1 alters processing of the GTPase OPA1 and causes obesity and defective thermogenesis in mice. EMBO J. 31, 2117–2133 (2012).
Sliter, D. A. et al. Parkin and PINK1 mitigate STING-induced inflammation. Nature 561, 258–262 (2018).
Matheoud, D. et al. Intestinal infection triggers Parkinson’s disease-like symptoms in Pink1(-/-) mice. Nature 571, 565–569 (2019).
Zhang, Z. et al. The dynamin-related GTPase Opa1 is required for glucose-stimulated ATP production in pancreatic beta cells. Mol. Biol Cell 22, 2235–2245 (2011).
Tamura, Y., Itoh, K. & Sesaki, H. SnapShot: mitochondrial dynamics. Cell 145, 1158 (2011).
Schrepfer, E. & Scorrano, L. Mitofusins, from mitochondria to metabolism. Mol. Cell 61, 683–694 (2016).
Cipolat, S., Martins de Brito, O., Dal Zilio, B. & Scorrano, L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc. Natl Acad. Sci. USA 101, 15927–15932 (2004).
Mullen, R. J., Buck, C. R. & Smith, A. M. NeuN, a neuronal specific nuclear protein in vertebrates. Development 116, 201–211 (1992).
Wai, T. et al. The membrane scaffold SLP2 anchors a proteolytic hub in mitochondria containing PARL and the i-AAA protease YME1L. EMBO Rep. 17, 1844–1856 (2016).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Newman, L. E. & Shadel, G. S. Mitochondrial DNA release in innate immune signaling. Annu. Rev. Biochem. 92, 299–332 (2023).
Sauer, J. D. et al. The N-ethyl-N-nitrosourea-induced Goldenticket mouse mutant reveals an essential function of Sting in the in vivo interferon response to Listeria monocytogenes and cyclic dinucleotides. Infect. Immun. 79, 688–694 (2011).
Yamada, T. et al. Mitochondrial stasis reveals p62-mediated ubiquitination in Parkin-independent mitophagy and mitigates nonalcoholic fatty liver disease. Cell Metab. 28, 588–604 (2018).
Wakabayashi, J. et al. The dynamin-related GTPase Drp1 is required for embryonic and brain development in mice. J. Cell Biol. 186, 805–816 (2009).
Postic, C. et al. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic beta cell-specific gene knock-outs using Cre recombinase. J. Biol. Chem. 274, 305–315 (1999).
Genin, E. C. et al. CHCHD10 mutations promote loss of mitochondrial cristae junctions with impaired mitochondrial genome maintenance and inhibition of apoptosis. EMBO Mol. Med. 8, 58–72 (2016).
Yamada, T., Dawson, T. M., Yanagawa, T., Iijima, M. & Sesaki, H. SQSTM1/p62 promotes mitochondrial ubiquitination independently of PINK1 and PRKN/parkin in mitophagy. Autophagy 15, 2012–2018 (2019).
Yamada, T. et al. Prevention and regression of megamitochondria and steatosis by blocking mitochondrial fusion in the liver. iScience 25, 103996 (2022).
Kageyama, Y. et al. Parkin-independent mitophagy requires Drp1 and maintains the integrity of mammalian heart and brain. EMBO J. 33, 2798–2813 (2014).
Stichel, C. C. et al. Parkin expression in the adult mouse brain. Eur. J. Neurosci. 12, 4181–4194 (2000).
Kitada, T. et al. Impaired dopamine release and synaptic plasticity in the striatum of PINK1-deficient mice. Proc. Natl Acad. Sci. USA 104, 11441–11446 (2007).
Chen, H. et al. Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J. Cell Biol. 160, 189–200 (2003).
Kageyama, Y. et al. Mitochondrial division ensures the survival of postmitotic neurons by suppressing oxidative damage. J. Cell Biol. 197, 535–551 (2012).
Shin, J. H. et al. PARIS (ZNF746) repression of PGC-1alpha contributes to neurodegeneration in Parkinson’s disease. Cell 144, 689–702 (2011).
Itoh, K. et al. Brain-specific Drp1 regulates postsynaptic endocytosis and dendrite formation independently of mitochondrial division. eLife 8, e44739 (2019).
Yamada, T., Adachi, Y., Fukaya, M., Iijima, M. & Sesaki, H. Dynamin-related protein 1 deficiency leads to receptor-interacting protein kinase 3-mediated necroptotic neurodegeneration. Am. J. Pathol. 86, 2798–2802 (2016).
Nguyen, T. et al. Uncovering the role of N-acetyl-aspartyl-glutamate as a glutamate reservoir in cancer. Cell Rep. 27, 491–501 (2019).
Elgogary, A. et al. Combination therapy with BPTES nanoparticles and metformin targets the metabolic heterogeneity of pancreatic cancer. Proc. Natl Acad. Sci. USA 113, E5328–E5336 (2016).
Udupa, S. et al. Upregulation of the glutaminase II pathway contributes to glutamate production upon glutaminase 1 inhibition in pancreatic cancer. Proteomics 19, e1800451 (2019).
Acin-Perez, R. et al. A novel approach to measure mitochondrial respiration in frozen biological samples. EMBO J. 39, e104073 (2020).
Etoh, K. & Nakao, M. A web-based integrative transcriptome analysis, RNAseqChef, uncovers the cell/tissue type-dependent action of sulforaphane. J. Biol. Chem. 299, 104810 (2023).
Kam, T. I. et al. Poly(ADP-ribose) drives pathologic alpha-synuclein neurodegeneration in Parkinson’s disease. Science 362, eaat8407 (2018).
Acknowledgements
We thank N. Senoo and S. M. Claypool at Johns Hopkins University School of Medicine for assistance with measurement of OCR. We also thank past and present members of the Iijima and Sesaki laboratories for helpful discussions and technical assistance. We acknowledge the joint participation by the Adrienne Helis Malvin Medical Research Foundation Parkinsons Disease Program (no. M-2019), through its direct engagement in the continuous active conduct of medical research, in conjunction with Johns Hopkins Hospital and Johns Hopkins University School of Medicine. T.M.D. is the Leonard and Madlyn Abramson Professor in Neurodegenerative Diseases. H.S. is the Ethan and Karen Leder CIM/HAP Scholar. This work was supported by NIH grants to H.S. (no. R35GM144103), M.I. (no. R35GM131768) and T.Y. (no. P20GM104320), as well as by grants from the Human Aging Project and Adrienne Helis Malvin Medical Research Foundation to H.S.
Author information
Authors and Affiliations
Contributions
T.Y., A.I., M.I. and H.S. conceived the study. T.Y. and A.I. performed most experiments, with assistance from Y.A. D.M. conducted immunoblot analysis, qPCR and the mitochondrial fusion assay. H.W. analysed dopaminergic neurons, dopamine metabolites and motor neurons. C.Z. and P.K. performed metabolomics. F.I. purified RNA for RNA-seq. A.I., M.I. and H.S. analysed most RNA-seq data, with assistance from S.B. M.I., H.S., D.C.C., T.L., T.M.D., P.M.Q. and C.L.-O. provided reagents and animals. H.S., M.I., T.M.D., V.L.D. and A.L. supervised the study. H.S., M.I., T.Y., A.I., H.W. and A.L. wrote and edited the manuscript. All authors participated in discussing and finalizing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jason Locasale and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation of Dele1−/− mice.
(a) To delete the Dele1 gene in the mouse genome, exon 2 was targeted using the CRISPR-Cas9 editing system with the indicated gRNA. (b) The location of the targeted site in the Dele1 gene is shown. (c) DNA sequencing confirmed that the genome editing removed 10 nucleotides at the position from 45 to 54 and introduced a stop codon after adding four amino acids.
Extended Data Fig. 2 Representative images of mitochondria in the zona incerta, midbrain, dorsal pallidum, lateral cortex, hippocampus, olfactory bulb, striatum, and cerebellum.
Brain sections from the indicated mouse lines were analyzed by laser confocal immunofluorescence microscopy with anti-PDH antibodies. Mice were analyzed at 6 weeks of age.
Extended Data Fig. 3 Representative images of mitochondria in the kidney, brown adipose tissue (BAT), skeletal muscle, lung, intestine, and spleen.
Tissue sections from the indicated mouse lines were analyzed by laser confocal immunofluorescence microscopy with anti-PDH antibodies. Mice were analyzed at 6 weeks of age.
Extended Data Fig. 4 Quantification of cells that contain enlarged mitochondria.
Values are mean ± SD (n = 4 for all tissues in the mouse lines, except for the following: zona incerta, midbrain, dorsal pallidum, and lateral cortex in 5 Parkin−/−Oma1−/−Mfn1+/− mice; BAT in 3 Oma1−/− mice; and lung in 3 WT, 3 Parkin−/−Oma1−/−, and 3 Parkin−/−Oma1−/−Mfn1+/− mice). Significance was calculated using one-way ANOVA with post-hoc Tukey: *p < 0.05, **p < 0.01, ***p < 0.001. Mice were analyzed at 6 weeks of age.
Extended Data Fig. 5 Megamitochondria are formed in neurons of Parkin−/−Oma1−/− mice.
(a) Quantification of cells that contain enlarged mitochondria in the pons and medulla in Parkin−/−Oma1−/− mice (mean ± SD, n = 3 mice). (b) Frozen sections of the pons/medulla from WT and Parkin−/−Oma1−/− mice were analyzed using laser confocal immunofluorescence microscopy with anti-PDH antibodies alongside cell type markers: NeuN (neurons), Iba1 (microglia), GFAP (astrocytes), and PECAM1 (vascular cells). (c) Megamitochondria-containing cells positive for each marker were quantified (mean ± SD, n = 3 mice). (d and e) Frozen sections of WT pons/medulla were analyzed using laser confocal immunofluorescence microscopy with antibodies against PDH and NeuN, with or without antigen retrieval using 1 mM EDTA. Two NeuN antibodies were used: 26975-1-AP from Proteintech in (d) and 24307 from Cell Signaling in (e). DNA was co-stained with DAPI. Significance was calculated using two-tailed Student’s t-test in (a) and one-way ANOVA with post-hoc Tukey (c): ***p < 0.001. Mice were analyzed at 6 weeks of age.
Extended Data Fig. 6 Western blotting analysis.
(a) The pons/medulla, cerebellum, and liver from WT and Parkin−/− mice were analyzed by Western blotting with the indicated antibodies. The asterisk indicates non-specific bands detected by the anti-Mfn1 antibodies. (b) Quantification of band intensity (mean ± SD, n = 3). (c) Confirmation of the specificity of the anti-Mfn1 antibodies was performed. Mfn1 was knocked down in mouse embryonic fibroblasts using two distinct siRNAs and subjected to Western blotting with the specified antibodies. (d) Quantification of band intensity (mean ± SD, n = 3). (e) The tissues from WT and Oma1−/− mice were analyzed by Western blotting with the specified antibodies. For Pgam5, P indicates the precursor form, and M in the mature form. (f) Quantification of band intensity (mean ± SD, n = 3). Significance was determined using two-tailed Student’s t-test in (b, f) and one-way ANOVA with post-hoc Tukey (d): *p < 0.05, **p < 0.01, ***p < 0.001. Mice were analyzed at 6 weeks of age.
Extended Data Fig. 7 Comparison of the metabolomic landscapes.
(a) Volcano plot of metabolomic data from WT and Parkin−/−Oma1−/− pons/medulla. None of the 188 metabolites identified showed significant changes in the Parkin−/−Oma1−/− pons/medulla compared to WT, using an FDR threshold of 0.05 indicated by the dotted line (n = 4 WT and 5 Parkin−/−Oma1−/− mice). (b) Log2 fold change and FDR of each metabolite involved in the TCA cycle, energy metabolism, glycolysis/gluconeogenesis, purine/pyrimidine metabolism, and amino acids. (c) Activity of each of the nine TCA cycle enzymes: aconitase (ACO), isocitrate dehydrogenase (IDH), oxoglutarate dehydrogenase (OGDH), succinyl-CoA ligase (SUCLG), fumarase (FUMH), malate dehydrogenase (MDH), citrate synthase (CS), succinate dehydrogenase (SDH), and pyruvate dehydrogenase (PDH) in WT and Parkin−/−Oma1−/− pons/medulla (mean ± SD, n = 3 mice). (d) Oxygen consumption rates (OCRs) in WT and Parkin−/−Oma1−/− brain (mean ± SD, n = 3 mice). Mice were analyzed at 6 weeks of age.
Extended Data Fig. 8 Principal component analysis and Western blotting.
(a) Principal component analysis of bulk RNA-seq data. (b) Postnuclear supernatants and mitochondrial fractions from WT and Parkin−/−Oma1−/− pons/medulla were analyzed by Western blotting using antibodies against Tim23 and α-tubulin. (c) qPCR analysis of the mitochondrial fraction from WT and Parkin−/−Oma1−/− pons/medulla (mean ± SD, n = 3). Significance was determined using two-tailed Student’s t-test.
Extended Data Fig. 9 qRT-PCR analysis.
qRT-PCR analysis of Ifit3, Usp18, Oasl2, Ddx60, Bst2, and Sting1 in the pons/medulla of WT and Parkin−/−Oma1−/− mice (mean ± SD, n = 5). Significance was determined using two-tailed Student’s t-test: *p < 0.05, **p < 0.01, ***p < 0.001.
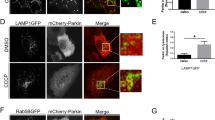
Extended Data Fig. 10 Measurement of mitochondrial fusion status.
(a) Western blotting of Drp1 KO MEFs transduced with lentiviruses carrying Opa1 and/or Mfn1. The asterisk indicates non-specific bands of anti-Mfn1 antibodies. Quantification of band intensity is shown (mean ± SD, n = 3). (b) Drp1 KO MEFs carrying matrix-targeted photoactivatable GFP (mito-PA-GFP), along with Opa1 and/or Mfn1, were stained with tetramethylrhodamine ethyl ester (TMRE). mito-PA-GFP in a single mitochondrion (indicated by the boxes) in the cell periphery was photoactivated, and images were obtained every 1 min at a single focal plane. Photoactivation was performed every 1 min on the same mitochondrion to maintain signal intensity. Representative images before and after (0 and 15 min) photoactivation are shown. (c) The mitochondrial fusion status was calculated based on the relative area containing photoactivated mito-PA-GFP signals over the total mitochondria stained with TMRE at 15 min (mean ± SD, n = 35 cells for Drp1 KO, 32 cells for Drp1 KO + Opa1, 34 cells for Drp1 KO + Mfn1, and 31 cells for Drp1 KO + Opa1 + Mfn1). Statistical analysis was performed using one-way ANOVA with post-hoc Tukey test: *p < 0.05, ***p < 0.001.
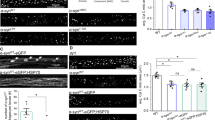
Extended Data Fig. 11 Analysis of dopaminergic and motor neurons.
(a) Tyrosine hydroxylase (TH) immunohistochemistry images of dopaminergic neurons in the substantia nigra pars compacta (SN) and striatum (STR) of WT and Parkin−/−Oma1−/− mice are shown. Nissl staining was included for the SN. (b, c) Stereological counting of TH-positive neurons (b) and Nissl-positive neurons (c) in the SN (mean ± SD, n = 3 mice). (d) Quantification of TH-positive fiber density in the STR (mean ± SD, n = 9 from 3 mice). (e–h) Measurements of dopamine (e), 3,4-dihydroxyphenylacetic acid (f), homovanillic acid (g), and 3-methoxytyramine (h) (mean ± SD, n = 4 mice). (i) Immunofluorescence microscopy of the spinal cord using anti-choline acetyltransferase antibodies. The cervical, thoracic, and lumbar regions were analyzed. Boxed areas are enlarged. (j) The number of motor neurons per 2,500 µm length of each spinal cord region is shown (mean ± SD, n = 4 mice). Significance was determined using two-tailed Student’s t-test in (b–h, j): **p < 0.01, ***p < 0.001.
Supplementary information
Supplementary Table 1 (download XLSX )
Comparison of amounts of metabolites in WT and Parkin−/−Oma1−/− pons/medulla. Values are normalized to the mean of WT for each metabolite (n = 4 WT and 5 Parkin−/−Oma1−/− mice). Significance was calculated using the two-tailed Student’s t-test with Benjamini–Hochberg multiple testing corrections.
Supplementary Table 2 (download XLSX )
RNA-seq gene count data and the results of GSEA and DEG analysis using RNAseqChef (n = 3 mice).
Supplementary Video 1 (download MOV )
A representative example of general locomotor activity of WT, Parkin−/−, Oma1−/− and Parkin−/−Oma1−/− mice at 6 weeks of age. The three larger, active mice are WT, Parkin−/− and Oma1−/−; the small, inactive mouse is Parkin−/−Oma1−/−.
Supplementary Video 2 (download MOV )
A representative example of general locomotor activity of Parkin−/−Oma1−/− and Parkin−/−Oma1−/−Opa1+/− mice at 6 weeks of age. The smaller, inactive mouse is Parkin−/−Oma1−/−; the larger, active mouse is Parkin−/−Oma1−/−Opa1+/−.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yamada, T., Ikeda, A., Murata, D. et al. Dual regulation of mitochondrial fusion by Parkin–PINK1 and OMA1. Nature 639, 776–783 (2025). https://doi.org/10.1038/s41586-025-08590-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08590-2
This article is cited by
-
Organelle-specific regulation of ferroptosis
Biology Direct (2026)
-
The hidden driver of osteoporosis: mitochondria-dominated inflammation
Cell Communication and Signaling (2025)
-
Upregulated astrocytic HDAC7 induces depression-like disorders via deacetylating PINK1 and inhibiting mitophagy
Journal of Neuroinflammation (2025)
-
An energy metabolism-engaged nanomedicine maintains mitochondrial homeostasis to alleviate cellular ageing
Nature Nanotechnology (2025)
-
Putting the brakes on mitochondrial fusion to prevent escape of mitochondrial DNA
Nature (2025)