Abstract
Loss-of-function variants in the lipid transporter ABCA7 substantially increase the risk of Alzheimer’s disease1,2, yet how they impact cellular states to drive disease remains unclear. Here, using single-nucleus RNA-sequencing analysis of human brain samples, we identified widespread gene expression changes across multiple neural cell types associated with rare ABCA7 loss-of-function variants. Excitatory neurons, which expressed the highest levels of ABCA7, showed disrupted lipid metabolism, mitochondrial function, DNA repair and synaptic signalling pathways. Similar transcriptional disruptions occurred in neurons carrying the common Alzheimer’s-associated variant ABCA7 p.Ala1527Gly3, predicted by molecular dynamics simulations to alter the ABCA7 structure. Induced pluripotent stem (iPS)-cell-derived neurons with ABCA7 loss-of-function variants recapitulated these transcriptional changes, displaying impaired mitochondrial function, increased oxidative stress and disrupted phosphatidylcholine metabolism. Supplementation with CDP-choline increased phosphatidylcholine synthesis, reversed these abnormalities and normalized amyloid-β secretion and neuronal hyperexcitability—key Alzheimer’s features that are exacerbated by ABCA7 dysfunction. Our results implicate disrupted phosphatidylcholine metabolism in ABCA7-related Alzheimer’s risk and highlight a possible therapeutic approach.
Similar content being viewed by others
Main
After APOE4, rare loss-of-function (LoF) mutations in ABCA7 caused by premature-termination codons (PTCs) are among the strongest genetic risk factors for Alzheimer’s disease (AD), with an odds ratio of approximately 2 (refs. 1,2). Common single-nucleotide polymorphisms in ABCA7 also moderately increase AD risk2,3, suggesting that ABCA7 dysfunction contributes substantially to disease risk in the broader population. Despite their significance, the precise cellular mechanisms through which ABCA7 LoF variants affect AD risk remain poorly defined.
ABCA7 functions by transporting phospholipids across cell membranes, thereby maintaining membrane asymmetry and facilitating lipid transport within the brain4,5,6. Mouse model studies suggest that ABCA7 dysfunction promotes amyloid deposition, impairs amyloid clearance by astrocytes and microglia and increases glial inflammatory responses7,8,9,10. Moreover, recent research in human cell lines and tissues has identified disturbances in lipid metabolism as potential mechanisms linking ABCA7 dysfunction to AD risk11,12,13. However, systematic analyses of ABCA7 LoF effects across various human brain cell types have not yet been performed. Investigations specifically addressing known ABCA7 PTC variants found in patients have been limited, with most existing studies focusing broadly on complete ABCA7 knockouts8,11,14.
Single-nucleus RNA sequencing (snRNA-seq) has effectively identified cell-type-specific transcriptional changes linked to other AD-associated genes, such as APOE and TREM215,16,17, and provided insights into disease mechanisms and therapeutic targets. Here we generated a cell-type-specific transcriptomic snRNA-seq atlas of ABCA7 LoF variants in the post-mortem human prefrontal cortex (PFC). Using this resource, we identified the cell-type-specific correlates of ABCA7 LoF variants in the human brain—particularly in neurons—and experimentally investigated these predictions in human neurons harbouring ABCA7 PTC variants.
Single-cell atlas of ABCA7 LoF variants
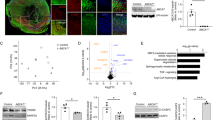
To investigate the cell-type-specific impacts of ABCA7 LoF variants in AD, we selected 12 carriers of rare ABCA7 PTC variants, including splice (c.4416+2T>G and c.5570+5G>C), frameshift (p.Leu1403fs, p.Glu709fs) and nonsense (p.Trp1245*, p.Trp1085*) variants, from the ROSMAP cohort (Fig. 1a,b and Supplementary Tables 1 and 2). ABCA7 PTC variants are presumed to cause ABCA7 haploinsufficiency, and were associated with lower ABCA7 protein levels in the PFC compared with in matched non-carriers, in a previously available dataset in which a subset overlapped with our sequencing patients (Extended Data Fig. 1a and Supplementary Table 3). Twenty-four matched controls without ABCA7 PTC variants were selected on the basis of AD pathology, age, sex, APOE genotype and cognitive status (Fig. 1c, Extended Data Fig. 1b,c, Supplementary Table 4 and Supplementary Note 1). The absence of rare damaging variants in other AD-associated genes2 was confirmed and a subset of genotypes was verified using Sanger sequencing (Methods and Extended Data Fig. 1d).
a, ABCA7 gene structure indicating studied variant locations. Exons are indicated by rectangles; introns are indicated by lines. The pie chart shows the frequency of ABCA7 PTC-variant carriers in the ROSMAP cohort. b, Human snRNA-seq cohort overview. The diagram was created using BioRender. c, Metadata summary of the snRNA-seq cohort. n = 36 individuals. This snRNA-seq experiment was performed once. d, 2D UMAP of ABCA7 LoF gene perturbation scores (S = −log10[P] × sign(log2[fold change (FC)]); unadjusted Limma-Voom P values; n = 12 (LoF) and n = 24 (control) individuals), restricted to genes with |S| > 1.3. Red, S > 1.3; blue, S < −1.3; point size ∝ |S|. The top 10 genes are labelled. e, 2D UMAP coloured by gene cluster assignment (Gaussian mixture model). f, For each cluster, the top two enriched pathways (GO BP; hypergeometric enrichment, one-sided, unadjusted P < 0.01; relative to all genes in UMAP) and top five genes (highest absolute mean S across cell types among genes in enriched pathways, P < 0.01) are shown. g, Cell-type-specific scores per gene cluster (SC), calculated as the mean perturbation score (S) of all genes in each cluster. *FDR-adjusted P < 0.01, |SC| > 0.25. Ast, astrocytes; Ex, excitatory neurons; In, inhibitory neurons; Mic, microglia; Oli, oligodendrocytes.
Raw snRNA-seq data from the BA10 region of the PFC were available for ten non-carrier samples from a previous study18. Fresh-frozen PFC BA10 tissues from the remaining individuals underwent snRNA-seq analysis using the 10x Genomics Chromium platform. After extensive quality control—including genotype–transcriptome matching to confirm sample identities and rule out potential sample swaps and correcting for batch effects (Extended Data Fig. 1e, Supplementary Fig. 1 and Supplementary Table 5)—our final dataset consisted of 102,710 high-quality cells from an initial total of 150,456 cells, representing inhibitory neurons, excitatory neurons, astrocytes, microglia, oligodendrocytes and oligodendrocyte precursor cells (OPCs; Supplementary Fig. 2). A small putative vascular cell cluster did not meet our quality thresholds and was excluded from further analysis.
We next analysed cell-type-specific transcriptional changes associated with ABCA7 LoF variants. We identified 2,389 genes with nominal evidence of perturbation (P < 0.05), suggesting possible transcriptional changes across six major neural cell types after controlling for covariates and focusing on genes detected in more than 10% of cells per type (Supplementary Table 6). We visualized these perturbations in two dimensions, revealing clear transcriptional patterns across cell types (Fig. 1d and Extended Data Fig. 2a). To nominate biological pathways that may be affected by ABCA7 LoF, we clustered genes on the basis of their proximity in this two-dimensional (2D) visualization, as closer genes exhibited similar perturbation patterns (Fig. 1e). Each gene cluster was then analysed for enrichment of biological pathways using the Gene Ontology Biological Process database, highlighting candidate functional themes disrupted by ABCA7 LoF, including cellular stress and apoptosis, synaptic function, DNA repair and metabolism (Fig. 1f and Supplementary Table 7).
Specifically, microglia exhibited marked downregulation of stress-response genes (such as HSPH1, cluster 11), a trend also observed, although less prominently, in neurons and OPCs (Fig. 1g). Microglia and astrocytes showed increased expression of transcriptional regulatory genes (clusters 9 and 10, respectively) (Fig. 1g). OPCs and oligodendrocytes displayed changes in inflammatory signalling pathways (for example, IL10RB, cluster 0; STAT2, cluster 8) (Fig. 1g). Neurons demonstrated increased expression of DNA-repair genes (such as FANCC, cluster 12) and reduced expression of synaptic transmission genes (for example, NLGN1 and SHISA6, cluster 1) (Fig. 1g). Excitatory neurons uniquely exhibited enhanced expression of cellular respiration genes (such as NDUFV2, cluster 7) and decreased expression of triglyceride biosynthesis genes (such as PPARD, cluster 5) (Fig. 1g). Overlaps in gene perturbations across cell types are summarized in Extended Data Fig. 3.
Together, these results highlight extensive cell-type-specific transcriptional disruptions associated with ABCA7 LoF in the human PFC. This single-cell atlas serves as a valuable resource to nominate pathways and genes for future investigation, and is accessible through the Single Cell Portal and Synapse (accession IDs: SCP3182, syn53461705).
ABCA7 LoF profiles in excitatory neurons
Our snRNA-seq data revealed that excitatory neurons express the highest levels of ABCA7 among major neural cell types in the brain (Extended Data Fig. 4a,b). We validated these expression patterns using an independent dataset19 (Supplementary Table 3), confirming significantly higher ABCA7 expression in neuronal versus glial populations from the human temporal cortex. Expression profiles of control genes that are known to be neuron or glia specific matched expectations (Extended Data Fig. 4c).
Given this expression profile, we hypothesized that excitatory neurons may be particularly impacted by ABCA7 LoF variants. To identify transcriptional correlates in excitatory neurons, we performed gene set enrichment analysis (GSEA) using WikiPathways (472 pathways), and we identified 34 candidate ABCA7 LoF-perturbed pathways (P < 0.05) involving 268 unique genes (Supplementary Table 8). To minimize redundancy and clearly identify biological themes, we grouped these genes into non-overlapping clusters through graph partitioning (Fig. 2a,b, Supplementary Fig. 3 and Supplementary Note 2). This analysis revealed eight biologically meaningful clusters highlighting three major themes: (1) energy metabolism and lipid homeostasis (clusters PM.0 and PM.1); (2) DNA damage and cellular stress responses (clusters PM.2, PM.3, PM.4 and PM.5); and (3) synaptic signalling (cluster PM.7) (Fig. 2a). Layer-specific analysis indicated consistent transcriptional perturbation patterns across cortical layers (Extended Data Fig. 5).
a, Kernel density plots of gene perturbation scores (S = −log10[P] × sign(log2[FC]); unadjusted Limma-Voom P values; n = 12 (LoF), n = 24 (control) individuals) per Kernighan–Lin cluster. Positive S indicates increased expression in ABCA7 LoF. The solid lines show the cluster means; the top pathways are indicated. Kernighan–Lin clustering performed on leading-edge genes from perturbed pathways (fGSEA, WikiPathways; unadjusted P < 0.05; Methods). b, Kernighan–Lin gene–pathway graph related to a: genes (circles) and pathways (squares) are indicated. c, Schematic of the ABCA7 gene, highlighting the p.Ala1527Gly variant (purple arrow). Cohort overview for snRNA-seq data from post-mortem PFC, comparing carriers of ABCA7 Gly1527 (≥1 allele) with Gly1527 non-carriers, that is, Ala1527 carriers (data from ref. 21). The diagram was created using BioRender. d, Perturbation of ABCA7 LoF-associated gene clusters (from a) in excitatory neurons from Ala1527 (n = 227 individuals) versus Gly1527 (n = 133 individuals) carriers (fGSEA analysis of ABCA7 LoF clusters 0–7). Normalized enrichment scores (NES) are shown. The top unadjusted P values are indicated. Positive scores indicate upregulation in Gly1527 variant carriers. e, The closed-conformation ABCA7 structure, highlighting the simulated domain (residues 1517–1756, yellow) and the lipid bilayer (orange). The inset shows Ala1527 (grey) and Gly1527 (purple). f, The root mean squared deviation (r.m.s.d.) of the ABCA7 domain (from e) with the Ala1527 (grey) or Gly1527 (purple) variant, relative to the closed-conformation reference during simulations. Inset: the average positional fluctuations of Cα atoms. Statistical analysis was performed using a two-sided Mann–Whitney U-test. g, Projection of Cα atom positional fluctuations onto first two principal components for the Ala1527 (top, grey) and Gly1527 (bottom, purple) variants during simulations.
Clusters PM.0 and PM.1 were enriched for genes involved in lipid metabolism, mitochondrial function and oxidative phosphorylation (OXPHOS). Specifically, cluster PM.0, including genes associated with lipid homeostasis (such as NR1H3, ACLY and PPARD), was downregulated, whereas cluster PM.1, comprising mitochondrial complex genes (such as COX7A2 and NDUFV2), was upregulated. Clusters PM.2, PM.3 and PM.6 contained upregulated DNA damage response and replication genes (such as RECQL, TLK2 and BARD1). Clusters PM.4 and PM.5 encompassed genes associated with proteasomal degradation, ciliogenesis, apoptosis and inflammation, exhibiting mixed directional regulation. Similarly, cluster PM.7, linked to synaptic and developmental pathways, contained both upregulated and downregulated genes (Fig. 2a).
ABCA7 LoF and p.Ala1527Gly overlap
ABCA7 LoF variants significantly increase AD risk but are rare, and therefore contribute to a small fraction of AD cases1,13. To assess whether transcriptional patterns associated with ABCA7 LoF extend to more common variants, we analysed carriers of the prevalent ABCA7 missense variant p.Ala1527Gly (rs3752246; minor allele frequency ≈ 0.18) within the ROSMAP cohort (Fig. 2c and Supplementary Table 3). Although annotated as the reference allele, Gly1527 represents the less-frequent allele associated with moderately increased AD risk (odds ratio = 1.15 (95% confidence interval 1.11–1.18))2,3,20. We analysed existing snRNA-seq data21 from human PFC samples of 133 Gly1527 carriers and 227 non-carriers, ensuring no overlap with the previous ABCA7 LoF cohort.
We observed directional transcriptional perturbations in excitatory neurons from Gly1527 carriers that were consistent with those previously associated with ABCA7 LoF variants across all identified clusters (PM.0–7) (Fig. 2a,d). Notably, clusters related to DNA damage (PM.3) and proteasomal function (PM.4) demonstrated evidence of upregulation, suggesting similar cellular stress and genomic instability signatures associated with ABCA7 LoF (Fig. 2a,d). Moreover, we found evidence for downregulation in lipid metabolism (PM.0) and modest upregulation in mitochondrial function (PM.1), aligning with changes seen in ABCA7 LoF neurons (Fig. 2a,d).
To explore structural explanations for these shared transcriptional patterns, we conducted molecular dynamics simulations, comparing Ala1527 and Gly1527 variants in the two extreme ABCA7 conformations—the fully closed, ATP-bound state (Fig. 2e–g) and the fully open, ATP-unbound state (Extended Data Fig. 6). Both conformations were embedded within a lipid bilayer and simulated over a 300 ns timescale to evaluate the local structural impact of the Gly1527 variant. The Gly1527 variant exhibited increased structural flexibility specifically in the ATP-bound closed state, characterized by pronounced conformational fluctuations compared with the Ala1527 variant (Fig. 2f,g). Given that the ATP-bound closed conformation is proposed to facilitate lipid presentation to apolipoproteins22,23, the increased flexibility of the Gly1527 variant may reduce lipid extrusion efficiency, consistent with recent experimental findings23. Both variants remained structurally stable in the ATP-unbound open state (Extended Data Fig. 6c–e). These results are further supported by analyses of ϕ/ψ dihedral angle distributions and secondary structure persistence, as described in Supplementary Note 3 and Extended Data Fig. 7. These structural insights, together with our transcriptomics data, suggest that both rare, high-effect ABCA7 LoF variants and common, moderate-risk variants may influence AD risk through similar ABCA7-dependent mechanisms, indicating broader relevance of ABCA7 dysfunction in AD.
Conserved signatures in ABCA7 LoF iNs
To experimentally validate the effects of ABCA7 LoF predicted by our single-cell dataset, we generated two isogenic iPS cell lines homozygous for distinct ABCA7 LoF variants using CRISPR–Cas9 editing (Fig. 3a and Supplementary Fig. 4). One variant, p.Glu50fs*3, introduces an early frameshift mutation, while the other, p.Tyr622*, represents a clinically relevant AD-associated variant1. Both variants likely produce severely truncated ABCA7 proteins or trigger nonsense-mediated decay; however, transcript rescue mechanisms such as exon skipping cannot be excluded24. We differentiated these iPS cells into induced neurons (iNs) through doxycycline-inducible NGN2 expression25 (Supplementary Fig. 5a). Wild-type (WT) and ABCA7 LoF neurons both expressed neuronal markers, formed robust neuronal processes within 2–4 weeks (Supplementary Fig. 5b,c), and exhibited electrophysiological activity (Extended Data Fig. 8a–e). ABCA7 LoF neurons demonstrated increased excitability, firing action potentials at lower thresholds than WT neurons (Extended Data Fig. 8f,g), consistent with neuronal hyperexcitability observed in AD.
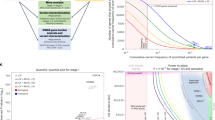
a, iPS cell (iPSC)-derived isogenic iNs with ABCA7 PTC variants (exon 3: p.Glu50fs*3; exon 15: p.Tyr622*). The gene schematic shows exons (rectangles) and introns (lines). The diagram was created using BioRender. Confocal MAP2 staining is shown. b, Correlation of gene perturbation scores (S = −log10[P] × sign(log2[FC]); unadjusted P values were computed using Limma-Voom) from bulk mRNA-seq data. n = 2 (WT) and n = 5 (for each LoF line) wells. c, Kernighan–Lin clustering of leading-edge genes from perturbed pathways in WT versus p.Tyr622* iNs (fGSEA, WikiPathways; FDR-adjusted P < 0.05). For the gene–pathway graph, genes (circles) and pathways (squares) are indicated. d, Heat map (Jaccard index) comparing Kernighan–Lin (K–L) clusters from p.Tyr622* iNs and post-mortem neurons (from Fig. 2a,b). Upregulated (red) and downregulated (blue) clusters in LoF neurons are indicated. FDR-adjusted P was calculated by permutation (1,000 iterations, one-sided). e, Kernel density plots of gene perturbation scores per cluster. Positive S indicates increased expression in p.Tyr622* iNs. The solid lines show the cluster means. The top pathways are indicated. f, Volcano plot of genes encoding mitochondrial proteins (MitoCarta); genes with FDR-adjusted P < 0.05 (Limma-Voom) in WT versus p.Tyr622* iNs are coloured. The top ten upregulated and downregulated genes are labelled. g, Seahorse mitochondrial uncoupled OCR (%). n = 18 (WT), n = 17 (p.Tyr622*) and n = 13 (p.Glu50fs*3) wells; two experiments. h, MitoHealth intensity. n = 8 (WT), n = 11 (p.Tyr622*) and n = 9 (p.Glu50fs*3) wells; around 3 × 103 cells per condition; three experiments. Statistical analysis was performed using a linear mixed-effects model. Maximum-intensity projections are shown with NeuN/GFP clipped at the 90th percentile, γ-corrected (γ = 0.5). i, The average TMRM intensity per 75th percentile mask (n = 4 (WT) and n = 5 (p.Tyr622*) wells; mean projection over time). j, The average CellROX intensity per 75th percentile mask. n = 10 wells per genotype. k, Differentially abundant lipid species in WT versus p.Tyr622* iNs (coloured by class); species are labelled if FDR-adjusted P < 0.05, |log[FC]| > 1, two-sided t-test, unequal variances assumed. n = 10 (WT) and n = 8 (p.Tyr622*) wells. For a–k, experiments were carried out after 4 weeks of differentiation; wells represent technical replicates. For g–j, analysis was performed using two-sided t-tests following Shapiro–Levene tests; the box plots show the median (centre line), interquartile range (IQR) (box limits) and 1.5 × IQR (whiskers). For g–j, datapoints represent the per-well mean. Experiments were performed once (a–f, j and k) or at least twice (g–i). Scale bars, 62 μm (a) and 125 μm (h–j).
We next examined whether transcriptional signatures identified in ABCA7 LoF post-mortem neurons were recapitulated in iNs using bulk mRNA-seq after 4 weeks of differentiation (Supplementary Table 9). Transcriptional perturbations between the p.Glu50fs*3 and p.Tyr622* variants were strongly correlated (Pearson correlation = 0.84; Fig. 3b). GSEA revealed 15 significantly perturbed pathways for each variant (false-discovery rate (FDR)-adjusted P < 0.05; WikiPathways; Supplementary Table 10). Kernighan–Lin partitioning of these pathways identified nine transcriptional clusters perturbed in WT versus p.Tyr622* and ten clusters in WT versus p.Glu50fs*3 (Fig. 3c, Extended Data Fig. 9a and Supplementary Table 10). These clusters showed substantial overlap between the two variants, with eight out of nine clusters from p.Tyr622* significantly overlapping with eight out of ten clusters from p.Glu50fs*3 (FDR-adjusted P < 0.05; Extended Data Fig. 9b). Moreover, we observed significant concordance with transcriptional signatures from post-mortem excitatory neurons, with five out of nine p.Tyr622*-associated clusters and seven out of ten p.Glu50fs*3-associated clusters overlapping significantly with post-mortem ones, predominantly with concordant directional changes (Fig. 3d and Extended Data Fig. 9c). For example, consistent with post-mortem findings, p.Tyr622* iNs exhibited downregulated clusters associated with lipid metabolism (T.9 and T.13) and upregulated clusters related to cell cycle regulation and proteasomal activity (T.8 and T.14) compared with WT neurons (Fig. 3e). A mitochondrial cluster (T.10) showed the strongest overlap with post-mortem neurons (cluster PM.1), being consistently upregulated in both variant lines (Fig. 3d and Extended Data Fig. 9c). Together, these data support a causal relationship between ABCA7 LoF variants and multiple transcriptional disruptions in excitatory neurons, particularly affecting proteostasis, cell cycle, lipid metabolism and mitochondrial function.
ABCA7 LoF variants impact mitochondria
To investigate mitochondrial alterations in ABCA7 LoF iNs in more depth, we analysed the expression of 1,136 genes encoding mitochondrial proteins from the MitoCarta database (Supplementary Table 3). Upregulated genes in p.Tyr622* neurons included mitochondrial apoptosis pathway genes (such as CASP3 and BID) and OXPHOS subunits, previously identified in clusters PM.1 and T.10 (Fig. 3f and Supplementary Table 11). Conversely, downregulated genes were enriched for β-oxidation (ACAD and CPT), mitochondrial metabolite transport (SLC25) and oxidative stress detoxification (CAT) (Fig. 3f and Supplementary Table 11). Similar gene expression profiles were observed in the p.Glu50fs*3 variant (Extended Data Fig. 9e).
We directly assessed mitochondrial function by measuring the oxygen-consumption rates (OCRs) using the Seahorse assay (Extended Data Fig. 10a,b). During OXPHOS, OCR-driven proton movement across the inner mitochondrial membrane maintains mitochondrial membrane potential (ΔΨm) and supports ATP production (Extended Data Fig. 10c). To account for differences in cell viability and mitochondrial abundance, we analysed internally normalized OCR ratios. Spare respiratory capacity—representing the mitochondria’s ability to increase respiration in response to demand—was comparable between WT and ABCA7 LoF neurons (Extended Data Fig. 10d,e). However, ABCA7 LoF neurons showed significantly reduced uncoupled mitochondrial respiration, defined as the proportion of basal oxygen consumption dedicated to maintaining membrane potential lost due to proton leakage rather than ATP synthesis26, compared with WT neurons (Fig. 3g and Extended Data Fig. 10f). The uncoupled mitochondrial OCR in WT neurons (approximately 20%; Fig. 3g) aligns with previous reports for neurons and other cell types27,28, indicating that ABCA7 LoF neurons exhibited abnormally low mitochondrial uncoupling. Consistent with this finding, expression of UCP2—a mitochondrial uncoupling protein expressed in the brain29—was reduced in ABCA7 LoF neurons (Extended Data Fig. 10g).
As reduced mitochondrial uncoupling can result in an elevated ΔΨm, we assessed ΔΨm using MitoHealth and TMRM dyes, both of which accumulate in mitochondria proportionally to the membrane potential. Both dyes showed higher fluorescence—indicative of elevated ΔΨm—in ABCA7 LoF neurons compared with in WT neurons (Fig. 3h,i). Signal specificity was confirmed by decreased fluorescence after FCCP-induced depolarization (Extended Data Fig. 10h). As mitochondrial uncoupling regulates reactive oxygen species (ROS) generation27, we next measured oxidative stress using CellROX dye. ABCA7 LoF neurons showed significantly increased fluorescent CellROX signal compared with WT neurons (Fig. 3j). Together, these data indicate that ABCA7 LoF variants decrease mitochondrial uncoupling, resulting in elevated membrane potential and increased oxidative stress.
ABCA7 LoF alters PC balance
As ABCA7 functions as a lipid transporter, we used liquid chromatography coupled with mass spectrometry (LC–MS) to examine lipid profiles in WT and ABCA7 LoF iNs (Supplementary Table 12). Comparing WT and p.Glu50fs*3 iNs revealed significant changes across several lipid classes, including neutral lipids, phospholipids, sphingolipids and steroids (Extended Data Fig. 11a,b). Notably, triglycerides (TGs), particularly long-chain polyunsaturated species, were frequently elevated in p.Glu50fs*3 iNs (Extended Data Fig. 11b,c).
Consistent with ABCA7’s known transport of phospholipids5,23, phosphatidylcholines (PCs)—key structural membrane components and potential ABCA7 substrates6—were prominently affected, with approximately 22% of detected PC species perturbed in p.Glu50fs*3 iNs (Extended Data Fig. 11b). Analysis by fatty acid saturation showed significant enrichment of saturated PCs among upregulated species (hypergeometric P = 0.026; Extended Data Fig. 11d). By contrast, several highly unsaturated polyunsaturated fatty acid (PUFA)-containing PCs showed decreased abundance (for example, PC(44:7) and PC(38:7); Extended Data Fig. 11e,f).
Lipidomics analysis in p.Tyr622* iNs (conducted in positive-ionization mode) revealed similarly increased saturated PCs (hypergeometric P = 0.044; Fig. 3k and Extended Data Fig. 11g,h). However, PUFA-containing PCs and long-chain TGs were not reliably detected in this analysis (Extended Data Fig. 11i,j).
De novo PC synthesis occurs through the Kennedy pathway, followed by fatty acyl chain remodelling through the Lands cycle, mediated by LPCAT enzymes, with LPCAT3 specifically introducing PUFA chains30. LPCAT3 expression was reduced in both p.Tyr622* and p.Glu50fs*3 iNs compared with in WT iNs (Extended Data Fig. 11k,l), consistent with elevated saturated PC levels. Overall, these results indicate that ABCA7 LoF neurons accumulate neutral lipids, including long-chain polyunsaturated TGs and sterol lipids (zymosteryl), and show enriched saturated PC content.
CDP-choline reverses ABCA7 LoF effects
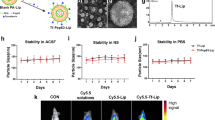
Previous studies showed that inhibition of de novo PC synthesis through the Kennedy pathway increased PC saturation in yeast, whereas boosting this pathway using exogenous choline reversed APOE4-induced lipid defects, including saturation changes, in human astrocytes31,32. We therefore tested whether choline supplementation could similarly mitigate ABCA7 LoF-induced phenotypes in iNs. To test this, we first treated p.Tyr622* iNs with CDP-choline for 2 weeks and performed targeted LC–MS analysis. CDP-choline treatment elevated extracellular CDP-choline from undetectable to detectable levels (Extended Data Fig. 12a and Supplementary Table 13). Furthermore, CDP and choline specifically accumulated in medium conditioned by treated p.Tyr622* cells (Extended Data Fig. 12a), indicating extracellular hydrolysis. Intracellular choline was significantly increased after treatment (Extended Data Fig. 12b), along with elevated expression of choline transporters (Extended Data Fig. 12c), confirming successful choline uptake by p.Tyr622* iNs.
We hypothesized that increased intracellular choline would enhance PC synthesis. Indeed, lipidomic analysis showed elevated levels of choline-containing phospholipids—particularly PCs, lysophosphatidylcholines (LPCs) and sphingomyelins—alongside a reduction in a single TG species, with other neutral lipid species showing a similar downward trend (Fig. 4a and Supplementary Table 12). Correspondingly, PCYT1B, the rate-limiting enzyme of Kennedy pathway-mediated PC synthesis, showed increased expression (Extended Data Fig. 12c). Moreover, LPCAT enzymes, including LPCAT3, exhibited higher expression after treatment (Extended Data Fig. 12d), consistent with increases to several unsaturated PC species (Fig. 4a and Extended Data Fig. 12e). These findings suggest that CDP-choline enhances the synthesis and remodelling of choline-containing lipids in ABCA7 LoF iNs.
a, Differentially abundant lipid species in p.Tyr622* iNs with or without CDP-choline. Species are labelled if unadjusted P < 0.05, |log[FC]| > 1 (two-sided t-test, equal variances assumed). n = 5 wells per condition. b, Correlation of gene scores comparing WT versus p.Tyr622* iNs and p.Tyr622* iNs with or without CDP-choline (n = 2 (WT), n = 5 (p.Tyr622* + H2O) and n = 5 (p.Tyr622* + CDP-choline); experiment from Fig. 3). c, Kernighan–Lin clustering of leading-edge genes in p.Tyr622* iNs with or without CDP-choline (fGSEA, WikiPathways; FDR-adjusted P < 0.05). Gene–pathway graph: genes (circles) and pathways (squares) are indicated. d, Heat map (Jaccard index) comparing Kernighan–Lin clusters in p.Tyr622* iNs with or without CDP-choline and WT versus p.Tyr622* iNs (Fig. 3c,e). Upregulated (red) and downregulated (blue) clusters in p.Tyr622* iNs + CDP-choline relative to p.Tyr622* iNs or in p.Tyr622* iNs relative to WT are indicated; FDR-adjusted permutation P values (1,000 iterations, one-sided). e, Kernel density plots of gene scores per cluster. Positive S represents an increase in the p.Tyr622*+CDP-choline condition. The solid lines show the cluster means. The top pathways are indicated. f, Volcano plot of genes encoding mitochondrial proteins. Genes with FDR-corrected P < 0.05 (Limma-voom) in p.Tyr622* iNs with or without CDP-choline are indicated in colour. Bold font indicates shared top genes with Fig. 3f. g, Seahorse mitochondrial uncoupled OCR (%). n = 6 (p.Tyr622* + H2O) and 8 (p.Tyr622* + CDP-choline) wells. h, The average TMRM intensity per masked region (75th percentile threshold; n = 8 wells per condition). i, The average CellROX intensity per masked region (75th percentile threshold; n = 10 wells per condition; same experiment as in Fig. 3j). j, Secreted Aβ, cortical organoids (182-day culture, with or without 1 mM CDP-choline for 4 weeks). n = 20 (WT), n = 19 (p.Tyr622*) and n = 14 (p.Tyr622* + CDP-choline) organoids. k, Spontaneous action potentials in dissociated cortical organoids (150 day culture, with or without 100 µM CDP-choline for 2 weeks). n = 7 (WT), n = 13 (p.Tyr622*) and n = 9 (p.Tyr622* + CDP-choline) cells. Statistical analysis was performed using two-sided Mann–Whitney U-tests following a Shapiro test. For a–i, 4-week differentiation was performed. +CDP-choline indicates treatment with 100 µM CDP-choline during the last 2 weeks; wells represent technical replicates. For g–k, the box plots show the median (centre line), IQR (box limits) and 1.5 × IQR (whiskers). For g–j, statistical analysis was performed using two-sided t-tests following Shapiro–Levene tests. Experiments were performed once (g, i and k) and at least twice (a–f, h and j). Scale bars, 125 μm (h and i).
Next, we characterized changes induced by CDP-choline treatment using LC–MS-based metabolomics and bulk RNA-seq. Although many metabolites altered by treatment could not be annotated, principal component analysis indicated that CDP-choline treatment reversed the separation between WT and p.Tyr622* iNs along the first principal component (PC1; Extended Data Fig. 12f). Transcriptomic analysis further demonstrated clear separation between treated and untreated samples (Extended Data Fig. 12g). Notably, the transcriptional signature of CDP-choline treatment negatively correlated with the p.Tyr622* signature (Fig. 4b), indicating partial restoration toward the WT state. Kernighan–Lin cluster analysis comparing untreated versus treated p.Tyr622* samples revealed significant overlap in seven out of nine clusters identified between p.Tyr622* and WT, with five clusters showing reversed directional changes after treatment (Fig. 4c–e and Supplementary Table 10).
Specifically, clusters associated with proteasomal and ribosomal functions (T+C.25, T+C.31), previously upregulated in p.Tyr622* iNs, were downregulated by CDP-choline treatment (Fig. 4e). Importantly, mitochondrial cluster T+C.26, strongly overlapping with mitochondrial cluster T.10 observed in post-mortem data (PM.1), also reversed after treatment (Fig. 4e). Analysis of MitoCarta genes confirmed a significant reversal in the expression of genes encoding mitochondrial proteins (Extended Data Fig. 12h), including reduced apoptosis-related genes (BID and CASP3; Fig. 3f), restored mitochondrial metabolic signatures (Supplementary Table 14) and elevated mitochondrial fusion regulators (MFN2 and OPA1), which support mitochondrial biogenesis and function33. Overall, CDP-choline treatment significantly reversed gene expression changes associated with ABCA7 LoF.
To assess whether CDP-choline treatment could restore mitochondrial uncoupling to WT levels, we repeated the Seahorse assay on p.Tyr622* iNs with and without treatment (Extended Data Fig. 12i,j). CDP-choline treatment significantly increased uncoupled respiration in p.Tyr622* iNs to WT levels (Fig. 4g), without altering the spare respiratory capacity (Extended Data Fig. 12k). Supporting this result, both TMRM and MitoHealth dyes showed lower fluorescence—indicative of decreased ΔΨm—in treated cells compared with in untreated cells (Fig. 4h and Extended Data Fig. 12l). Moreover, CDP-choline significantly reduced oxidative stress, as indicated by decreased CellROX fluorescence (Fig. 4i).
CDP-choline reduces AD phenotypes
Finally, we evaluated whether CDP-choline treatment could ameliorate key AD-associated phenotypes, as ABCA7 dysfunction has been linked to altered amyloid-β (Aβ) processing13. p.Tyr622* iNs secreted significantly higher Aβ40 and showed a trend toward increased Aβ42 secretion, although the absolute levels remained relatively low (Extended Data Fig. 13a). To examine the effects in a model with stronger pathology, we differentiated p.Tyr622* and WT lines into cortical organoids matured for approximately 6 months, a stage at which robust Aβ secretion was observed (approximately twofold to fourfold higher than iNs; Extended Data Fig. 13b,c). Treatment for 4 weeks with 1 mM CDP-choline normalized Aβ40 and Aβ42 secretion in p.Tyr622* organoids to WT levels (Fig. 4j); this effect was concentration and duration dependent (Extended Data Fig. 13c). Furthermore, CDP-choline treatment at 100 µM significantly reduced neuronal hyperexcitability in dissociated cortical organoids, as assessed by electrophysiology (Fig. 4k).
Discussion
Here we generated a transcriptional atlas to identify potential cell-type-specific effects of ABCA7 LoF variants in the human PFC. Excitatory neurons expressed the highest levels of ABCA7 and showed transcriptional alterations in pathways related to lipid biosynthesis, mitochondrial respiration and cellular stress, including upregulation of DNA-damage-response genes, as well as changes in inflammatory and synaptic genes. Experimental validation of predictions from this atlas revealed that ABCA7 LoF variants impaired mitochondrial uncoupling, elevated mitochondrial membrane potential and increased ROS levels in human iNs. Consistent with ABCA7’s function as a phospholipid transporter, ABCA7 LoF altered PC composition in these neurons, characterized by increased saturated PCs and reduced highly polyunsaturated PCs. CDP-choline treatment increased PC synthesis, elevated expression of PC-remodelling enzymes, and corrected mitochondrial uncoupling deficits, mitochondrial membrane potential and oxidative stress. Furthermore, CDP-choline supplementation reduced neuronal hyperexcitability and amyloid-β secretion.
Our findings indicate that ABCA7 LoF neurons accumulate saturated PC species, consistent with recent reports of phospholipid saturation imbalance in human amyotrophic lateral sclerosis and frontotemporal dementia neurons34. Treatment with CDP-choline, which boosts de novo PC synthesis, effectively reduced the downstream effects of ABCA7 LoF but did not fully normalize the lipid profile. Enhancing de novo PC synthesis with CDP-choline may broaden the diversity of PC species by providing additional substrates for the Lands cycle. This cycle rapidly remodels the PC pool, generating diverse saturated and unsaturated PCs35,36, potentially counteracting an excess of saturated species. Although further studies are needed to fully characterize these lipid alterations and their functional implications, and investigate the specificity of CDP-choline treatment, our data suggest that disrupted PC metabolism may contribute to neuronal dysfunction associated with ABCA7 LoF variants.
PCs are abundant components of biological membranes, including mitochondrial membranes. Changes in their fatty acyl chain composition impact mitochondrial bioenergetics, dynamics and membrane potential37. Mitochondrial dysfunction, including impaired mitochondrial uncoupling, is increasingly linked to ageing and neurodegeneration, although its specific role in AD remains unclear38. Neurons heavily depend on mitochondrial OXPHOS to meet their energy demands39, and regulated mitochondrial uncoupling supports neuronal health by controlling mitochondrial membrane potential, reducing ROS and promoting mitochondrial biogenesis40,41. Elevated oxidative stress resulting from impaired uncoupling could contribute to neuronal DNA damage and inflammation observed in AD brains42, both of which are transcriptionally evident in ABCA7 LoF carriers.
Further studies are needed to clarify how ABCA7 regulates PC composition. Previous research suggests that ABCA7 transports PC and its Lands cycle derivatives, LPCs6. We speculate that impaired ABCA7 floppase activity, which moves phospholipids between membrane leaflets, might initially cause PC accumulation at the inner leaflet. This buildup could trigger compensatory downregulation of PC synthesis. Supporting this idea, the Kennedy pathway’s rate-limiting enzyme, CTP:phosphocholine cytidylyltransferase, is activated after binding to PC-deficient membranes43. Such conditions could shift the existing PC pool toward more saturated species, as previously shown in yeast31. Impaired floppase activity involving PCs, LPCs or other phospholipids transported by ABCA7, such as phosphatidylserines23, might also indirectly alter PC composition by changing membrane fluidity and curvature44. These membrane changes could then affect lipid metabolism by modulating enzymes such as LPCAT3, which helps to maintain levels of unsaturated phospholipids45,46.
Consistent with our findings linking PC imbalance to mitochondrial dysfunction in ABCA7 LoF neurons, a recent independent study reported mitochondrial impairment associated with phosphatidylglycerol deficiency in ABCA7-deficient neurospheroids11, further emphasizing lipid metabolism as a therapeutic target. Here we demonstrate that CDP-choline—a safe and widely available dietary supplement47—reverses key aspects of ABCA7 LoF-induced neuronal dysfunction, including AD pathology and neuronal hyperexcitability. Recent studies from our laboratory similarly linked PC and fatty acid saturation imbalances to APOE4-associated dysfunction32 and cognitive resilience to AD pathology48, highlighting broad relevance for PC disruptions in AD risk.
Supporting the therapeutic potential of targeting phospholipid metabolism in AD, dietary choline supplementation in APP/PS1 mouse models significantly reduced amyloid pathology49, aligning with our findings in cortical organoids. Moreover, recent studies in Drosophila amyotrophic lateral sclerosis/frontotemporal dementia models demonstrated that in vivo overexpression of fatty acid desaturases improved survival, probably through correction of phospholipid saturation imbalances34. Notably, a recent epidemiological study also linked higher dietary choline intake to reduced AD risk in humans50, further highlighting the promise of targeting PC pathways therapeutically in AD.
Our data also suggest that the common missense variant p.Ala1527Gly may produce effects convergent with ABCA7 LoF. Genetic interactions between mild ABCA7 dysfunction and other AD risk factors, such as APOE4, could substantially amplify AD risk51. Collectively, these findings align with a growing body of literature, including recent work on APOE416,52,53, highlighting that lipid metabolic disruptions are central to AD pathogenesis and identifying additional genotypes that may benefit from modifying phospholipid metabolism.
Methods
Isolation of nuclei from post-mortem brain tissue
Batch 1 nuclei (BA10 region, frozen tissue) were isolated according to a protocol adapted from a previous study18, performed entirely at 4 °C or on ice. In brief, tissue was homogenized (700 µl homogenization buffer: 320 mM sucrose, 5 mM CaCl2, 3 mM Mg(CH3COO)2, 10 mM Tris-HCl pH 7.8, 0.1 mM EDTA pH 8.0, 0.1% IGEPAL CA-630, 1 mM β-mercaptoethanol, 0.4 U µl−1 recombinant RNase inhibitor (Clontech)) using a Wheaton Dounce tissue grinder (15 strokes, loose pestle), filtered (40 µm cell strainer), then mixed 1:1 with working solution (diluent (30 mM CaCl2, 18 mM Mg(CH3COO)2, 60 mM Tris pH 7.8, 0.6 mM EDTA, 6 mM β-mercaptoethanol) and OptiPrep density-gradient solution (Sigma-Aldrich, D1556-250ML), 1:5). The sample was layered onto an OptiPrep density gradient consisting of 750 µl of 30% OptiPrep (1.5:1 ratio of working solution:homogenization buffer) above 300 µl of 40% OptiPrep (4:1 ratio of working solution:homogenization buffer), centrifuged (10,000g, 5 min, 4 °C) and nuclei were collected from the 30/40% interface (100 µl). Nuclei were washed twice (1 ml PBS, 0.04% BSA, 300 g, 3 min), resuspended (100 µl PBS, 0.04% BSA), counted (C-Chip hemocytometer) and diluted to 1,000 nuclei per µl (PBS 0.04% BSA).
Batch 2 nuclei (fresh post-mortem PFC BA10 tissue) were prepared as part of a previous study18.
Informed consent and Anatomical Gift Act consent were obtained, including repository consent to allow sharing of data and biospecimens. Rush University Medical Center IRB approved protocols (Religious Orders Study, Rush Memory and Aging Project).
Droplet-based snRNA-seq
Batch 1 libraries were prepared using Chromium Single Cell 3′ Reagent Kits v3 (10x Genomics) and sequenced on the NovaSeq 6000 S2 (paired-end, 28 + 91 bp, 8-nt index) system. Each sample was sequenced twice across two lanes to increase the depth. Batch 2 libraries were prepared using the Chromium Single Cell 3′ Reagent Kits v2 and sequenced using the NextSeq 500/550 High Output v2 kits (150 cycles), as previously described18. All raw reads were processed together for alignment and gene counting.
Variant calling and ROSMAP participant selection
We selected 36 individuals from the ROSMAP cohort, a longitudinal study of ageing and dementia54. Whole-genome sequencing (WGS) variant calls (n = 1,249 available ROSMAP samples) were downloaded from Synapse (syn11724057) for genes with rare damaging variants linked to AD: SORL1, TREM2, ABCA7, ATP8B4, ABCA1 and ADAM102. For participants with multiple WGS samples, the highest-quality sample was chosen (Genomic Quality Score). Samples with sex mismatches or genotype inconsistencies were excluded (see Synapse accession syn12178037). Only variants passing quality control (FILTER_PASS) were used.
PTC variants flagged as splice, frameshift, nonsense, missense or premature stop variants annotated as loss-of-function (LOF) were identified. For ABCA7, known LoF variants from the literature were captured, except for c.5570+5G>C, which was manually added. Additional WGS details (library preparation, quality control, annotations, impact predictions) can be viewed at Synapse (syn10901595).
We selected 12 individuals (LoF samples) who carried ABCA7 PTC variants, had no PTC variants in the other candidate genes listed above and had fresh-frozen tissue available from Rush University. Moreover, we chose 24 matched controls without any PTC variants in ABCA7 or the other listed genes. Controls were matched by age, sex and pathology.
Read counting and alignment
Libraries were demultiplexed using the MIT BioMicroCenter BMC/BCC 1.8 pipeline (updated 9 December 2020; https://openwetware.org/wiki/BioMicroCenter:Software#BMC-BCC_Pipeline). Fastq reads were aligned to the human reference genome (GRCh38) and counted using Cell Ranger (v.6.1.2; 10x Genomics) with intron counting enabled and an expected cell count of 5,000 per sample. The default parameters were otherwise used. Counts from all samples were aggregated using a custom script, yielding a total of 150,456 cells.
Sample-swap analysis
Sample-swap analysis was performed using an established pipeline (MBV; QTLtools v1.1)55, comparing allelic concordance between genomic (VCF) and transcriptomic (BAM, generated by Cell Ranger) data. We specifically analysed chromosome 19 variants (location of ABCA7). Each single-cell sample matched the expected WGS sample clearly, showing higher concordance (fewer mismatches) compared with all other ROSMAP WGS samples (examples are shown in Extended Data Fig. 1e).
Cell filtering metrics
Aggregated counts underwent quality control before cell annotation. Cells with fewer than 500 or more than 10,000 detected genes (count > 0) were removed. Next, we filtered cells by mitochondrial fraction (total mitochondrial counts divided by total gene counts), a measure of nuclear integrity. We log-transformed mitochondrial fractions and fitted a Gaussian mixture model (GMM, sklearn GaussianMixture) to identify and remove cells assigned to the GMM component with the highest mean mitochondrial fraction. This step removed approximately 20,000 low-quality cells.
We next considered cells in marker-gene expression space defined by known major cell-type markers for human PFC: astrocytes (159 markers), excitatory neurons (113), inhibitory neurons (83), microglia (97), oligodendrocytes (179), OPCs (143) and vascular cells (124)1 (Supplementary Table 3). Marker counts were normalized to total library size, mean-centred and scaled to unit variance. Incremental principal component analysis (sklearn IncrementalPCA) reduced dimensionality (top 50 principal components). Visually, cells projected onto the first two principal components formed distinct Gaussian-like clusters. Assuming each Gaussian cluster corresponded to a distinct brain cell type, we fitted another GMM to the projected data. The resulting ten clusters aligned clearly with known brain cell types.
Cells poorly modelled by this GMM (log-probability < −100) and two clearly outlying clusters were removed. These excluded cells had lower total counts and higher mitochondrial fractions, suggesting low quality. This step removed approximately 12,000 cells, leaving a final dataset of 118,668 cells.
Gene filtering metrics
Downstream analyses included only nuclear-encoded, protein-coding genes (total 19,384) based on Ensembl GRCh38p12 annotations.
Cell type annotations
We first corrected variance due to sequencing batch and individual-of-origin by applying Harmony56 to the top 50 principal components from the quality-controlled data. Using Harmony-corrected principal components, we computed a neighbourhood graph (default Scanpy parameters)57 and clustered cells with the Leiden algorithm (Scanpy implementation)58.
Major cell types (excitatory neurons, inhibitory neurons, astrocytes, microglia, oligodendrocytes, OPCs, vascular cells) were assigned to Leiden clusters by computing cell-type-specific marker gene enrichment. Specifically, we calculated enrichment scores as the average log-ratio of expression for marker genes inside versus outside each cluster and assigned labels based on the highest enrichment.
We then subclustered each major cell type using the Leiden algorithm and removed subclusters with excessively high mitochondrial fraction or extreme total counts. Thresholds were set at two s.d. above the mean for these metrics within each major cell type. Removed clusters were small, poorly represented across individuals and weakly connected on manual inspection.
Individual-level filtering
After all of the previous quality-control steps, six individuals with fewer than 500 cells were excluded from further analyses, leaving 24 control individuals and 12 individuals with ABCA7 LoF. None of these excluded individuals carried ABCA7 PTC variants, and their removal did not substantially affect clinical variable distributions across genotypes.
Differential gene expression
Pseudo-bulk gene expression values were generated by summing cell-level counts per gene per individual (matrix multiplication). For each major cell type, we considered genes detected in >10% of cells. Counts were normalized by TMM (edgeR), and residual mean-variance trends were removed using Limma-Voom. Unknown variance was modelled through surrogate variable analysis (SVA). Differential expression analysis (Limma: lmFit, eBayes, topTable) was performed separately for each major cell type using the following linear model for each gene (Gi):
where n is the number of surrogate variables determined by num.sv() per cell type and ABCA7 LoF indicates individuals carrying ABCA7 LoF variants. Additional covariates (defined in Supplementary Note 1) included sex, NFT, amyloid burden, age at death, PMI, sequencing batch and APOE4 status.
Gene perturbation projections across cell types
We computed cell-type-specific gene perturbation scores summarizing differential expression significance and direction associated with ABCA7 LoF as S = sign(log2[FC]) × −log10(P), where positive log2[FC] indicates upregulation in ABCA7 LoF. Scores for genes not detected in >10% of cells per cell type were set to zero. Genes with \(| S| > 1.3\) in at least one of six major cell types (excitatory neurons, inhibitory neurons, astrocytes, microglia, oligodendrocytes and OPCs) were projected from 6D perturbation-score space into 2D using UMAP (Python umap).
Genes were clustered in the resulting 2D embedding using Gaussian mixture modelling (Python sklearn). Clusters were annotated by hypergeometric enrichment (Python gseapy) for Gene Ontology Biological Process pathways (Supplementary Table 3), using all genes in the embedding as background. Pathways with enrichment P < 0.01 were selected for naming each cluster. Per-cell-type perturbation scores for each cluster were calculated as the mean gene score within clusters. Statistical significance was assessed by permuting cluster assignments (100,000 permutations).
Gene-set enrichment and Kernighan–Lin pathway clustering
Genes were ranked by perturbation scores S (see the ‘Gene perturbation projections across cell types’ section). Fast GSEA (fGSEA; R implementation59) with 10,000 permutations tested enrichment of WikiPathways gene sets (Supplementary Table 3) among differentially expressed genes. Only gene sets with 5–1,000 genes were considered.
To simplify gene–pathway associations, we constructed a bipartite graph using genes from the fGSEA leading-edge (LE) subset (268 genes, enriched at P < 0.05 in ABCA7 LoF excitatory neurons) and WikiPathways associated with ≥4 LE genes. We treated gene–pathway grouping as a graph partitioning problem (Supplementary Note 2). Among three graph-partitioning algorithms tested (Supplementary Note 2), the METIS and the Kernighan–Lin algorithms showed the lowest loss and highly comparable performance (within 1.8% loss; Rand index = 0.98 after 5.0 × 104). We selected the Kernighan–Lin algorithm because it consistently outperformed the METIS algorithm across a wider range of graph sizes. The Kernighan–Lin algorithm was implemented in Python as described previously60 with the parameters C=0, KL_modified=True, random_labels=True, unweighted=True, and K=50 to partition the graph into eight groups. We performed 5.0 × 104 random initiations and selected the lowest-loss solution.
Graph layouts were computed using the spring layout algorithm (networkx, 10,000 iterations) and visualized using matplotlib. Representative pathways for each cluster were identified by averaging ABCA7 LoF perturbation scores (S) of genes in the cluster connected directly to each pathway. Pathways with ≥5 intracluster gene connections are highlighted in the figures.
Excitatory neuronal layer annotation
Excitatory neurons were annotated by cortical layer using published marker gene sets61 (Supplementary Table 3) according to the procedures described in the ‘Cell type annotations’ section. In brief, the normalized expression matrix was filtered to include only layer-specific marker genes and cells expressing ≥15% of these genes. Dimensionality was reduced using iterative principal component analysis, followed by batch-effect correction using Harmony. A neighbourhood graph was constructed, and cells were clustered using the Leiden algorithm. Clusters enriched for layer-specific markers (average log-transformed FC > 0.1) were labelled accordingly, while ambiguous clusters were excluded. Layers 5 and 6 were combined into a single ‘L5/6’ category. Annotations were validated using independent marker genes62 (Supplementary Table 3). Layer-specific differential expression analysis was performed as described in the ‘Differential gene expression’ section, followed by gene-set enrichment analysis (fGSEA, as described in the ‘Gene-set enrichment and Kernighan–Lin pathway clustering’ section) testing enrichment of ABCA7 LoF-associated gene clusters identified by Kernighan–Lin clustering (as described in the ‘Gene-set enrichment and Kernighan–Lin pathway clustering’ section).
ABCA7 p.Ala1527Gly variant calling and gene–pathway clustering comparisons
Participants carrying the ABCA7 p.Ala1527Gly variant with available PFC snRNA-seq data from a previous study (Supplementary Table 3) were identified using methods described in the ‘Variant calling and ROSMAP participant selection’ section. Differential expression was computed as described in the ‘Differential gene expression’ section, followed by fGSEA to test enrichment of ABCA7 LoF-associated gene clusters identified by Kernighan–Lin clustering (see the ‘Gene-set enrichment and Kernighan–Lin pathway clustering’ section).
Culture and generation of human isogenic iPS cells
A control parental iPS cell line (AG09173; 75-year-old female individual, APOE3/3 genotype) was generated previously by the Picower Institute iPSC Facility63. Two ABCA7 LoF isogenic lines were derived from AG09173: ABCA7 p.Glu50fs*3, containing a novel premature stop codon in exon 3 (generated by Synthego), and ABCA7 p.Tyr622*, containing a patient-derived mutation (Y622*)64 generated in-house by CRISPR–Cas9 editing.
For the ABCA7 p.Tyr622* line, an sgRNA targeting ABCA7 (oligos: forward, 5′-CACCGCCCCTACAGCCACCCGGGCG-3′; reverse, 5′-AAACCGCCCGGGTGGCTGTAGGGGC-3′; designed at http://crispr.mit.edu) was cloned into pSpCas9-2A-GFP (PX458, Addgene, 48138) as previously described65. The plasmid was confirmed by Sanger sequencing, then nucleofected (Amaxa, Lonza Human Stem Cell Nucleofector Kit I, program A-23) along with 15 μg of a single-stranded oligodeoxynucleotide template into dissociated AG09173 iPS cells (Accutase, Thermo Fisher Scientific; 10 μM ROCK inhibitor, Tocris). Cells (around 5 × 106) were sorted (BD FACS Aria IIU, Whitehead Institute), plated at single-cell density in medium supplemented with penicillin–streptomycin (Gemini Bio-products) and ROCK inhibitor. Colonies were expanded, screened by genomic DNA extraction (DNeasy Blood & Tissue Kit, Qiagen, 69504) and Sanger sequencing to confirm the Y622* mutation (Supplementary Table 16).
All iPS cell lines were regularly tested for karyotypic normality (Cell Line Genetics) and cultured at 37 °C, 5% CO2, in feeder-free conditions using mTeSR-1 medium (StemCell Technologies, 85850) on Matrigel-coated plates (Corning; hES-cell-qualified, 354277). Cells were passaged at 60–80% confluence using ReLeSR (StemCell Technologies, 05872) onto Matrigel-coated plates at a 1:6 to 1:24 split ratio.
rTTA and NGN2 virus production
HEK293T cells were seeded at 5 × 106 cells per 10 cm plate and transfected using a third-generation lentiviral system. Per plate, transfection mixtures contained 10 µg plasmid DNA (EF1a-rtTA-Hygro, Addgene 66810, or pLV-TetO-hNGN2-eGFP-Puro, Addgene, 79823), 5 µg pMDLg/pRRE, 2.5 µg pRSV-Rev, 2.5 µg MD2.G and 48 µl polyethyleneimine (1 mg ml−1) diluted in 600 µl OptiMEM (Thermo Fisher Scientific, 51-985-034). Mixtures were incubated 20 min at room temperature, added dropwise to cells and replaced with fresh medium after 16 h. Virus-containing supernatant collected 72 h after transfection was clarified (3,000g, 5 min, 4 °C) and the supernatant was ultracentrifuged (Beckman Optima L-90K Ultracentrifuge, SW32Ti rotor, 25,000 rpm, 2 h), resuspended in 1 ml PBS per 10 cm plate and stored at −80 °C.
Lentivirus-mediated NGN2 induction in iPS cells and drug treatments
iPS cells were dissociated into single-cell suspensions (Cell Dissociation Buffer, Life Technologies, 13151-014), resuspended in mTeSR1 medium with ROCK inhibitor (Rockout; Abcam, ab285418), and plated onto Matrigel-coated six-well plates at 50–60% confluence after 24 h. After 1 day, cells were co-transduced overnight with 80 µl each of pLV-TetO-hNGN2-eGFP-Puro and EF1a-rtTA-Hygro lentivirus per well. NGN2 expression was induced 24 h later with doxycycline (1 µg ml−1) and ROCK inhibitor. Puromycin selection was performed 24 h after viral transduction. Immature neurons were replated on PDL/laminin-coated plates (1 × 106 cells per well in six-well plates, or 5 × 104 cells per well in 96-well plates), and maintained in BrainPhys neuronal medium (StemCell Technologies, 05793) with Neurocult SM1 neuronal supplement (StemCell Technologies, 05711), (N2-supplement-A StemCell Technologies, 07152), laminin (1 µg ml−1) and doxycycline (1 µg ml−1). Half-medium changes were performed every 3–4 days, and cultures were matured for 28 days before experiments.
Neurons were treated with cytidine 5′-diphosphocholine (CDP-choline, Millipore Sigma-Aldrich, 30290) at a final concentration of 100 µM starting at day 14, continuing with each medium change until day 28. Choice of treatment concentration and duration was based on a previous study by our laboratory32.
Cortical organoid generation
Dorsal cortical organoids were generated as previously described66. In brief, iPS cells at 80–90% confluence were dissociated into single-cell suspensions (1 × 105 cells per ml) in mTeSR with 10 µM ROCK inhibitor, seeded at 100 µl per well in PrimeSurface 96 Slit-well plates (S-Bio, MS9096SZ) and induced to differentiate using neural induction medium consisting of DMEM/F12 (Life Technologies, 11330-032), 100 mM GlutaMAX (Life Technologies, 35050-061), 0.1 mM 2-mercaptoethanol (Sigma-Aldrich, M3148), 1% penicillin–streptomycin (Life Technologies, 15070-063) and 10 µM SB-431542 (R&D Systems, 1614), and 2.5 µM dorsomorphin (Sigma-Aldrich, P5499-CONF) with daily medium changes (days 0–5). The medium was then switched (days 6–16) to neural differentiation medium (Neurobasal A, B27 supplement, GlutaMAX, penicillin–streptomycin, human recombinant EGF and FGF2, 20 ng ml−1 each), with daily changes until day 16, then every other day until day 25. From day 25 onwards, EGF and FGF2 were replaced with 20 ng ml−1 each of BDNF and NT3, with medium changes twice weekly after day 45.
Confocal imaging experiments
All confocal images were acquired on a Zeiss LSM900 microscope using ZEN software.
For mitochondrial health staining, live cells were incubated with MitoHealth dye (Thermo Fisher Scientific, H10295) according to the manufacturer’s protocols for 30 min at 37 °C, fixed (4% paraformaldehyde/4% sucrose, 15 min, room temperature), permeabilized (0.1% Triton-X, 5 min), blocked (2% BSA, Fisher Bioreagents, BP9703) and incubated overnight at 4 °C with NeuN antibody (1:500), followed by incubation with secondary antibodies (1:1,000) for 2 h and Hoechst (1:2,000, Invitrogen, H3570) for 10 min (Supplementary Table 17). Images were captured as z stacks (1 µm intervals).
Live imaging of mitochondrial membrane potential used TMRM (0.1 µM, 30 min at 37 °C; Thermo Fisher Scientific, I34361), followed by imaging before and immediately after adding the mitochondrial uncoupler FCCP (1 µM; Cayman Chemical, 15218). ROS were assessed by live staining with CellROX Orange (5 µM, 30 min at 37 °C; Thermo Fisher Scientific, C10443). TMRM and CellROX images were acquired as single optical sections.
For immunostaining, iNs cultured on coverslips and cortical organoid cryosections (20 µm) were fixed (4% formaldehyde, 10 min), permeabilized (0.2% Triton X-100) and blocked (10% BSA, 1 h), and incubated overnight at 4 °C with primary antibodies (MAP2 and NeuN, both 1:1,000). Alexa-Fluor-conjugated secondary antibodies (1:500) and Hoechst (1:1,000) were used for visualization. Coverslips were mounted with Fluoromount-G, and images were captured as single optical sections.
For visualization, confocal images were pseudocoloured to enhance the signal contrast; representative unprocessed images are provided in Supplementary Fig. 6.
Confocal image quantification
Confocal images (.czi format; 8 or 16 bits; voxel size: 1 × 0.62 × 0.62 µm) were loaded into Python (aicsimageio) and normalized to floating-point format [0,1]. Acquisition settings were consistent within each imaging batch.
For fixed z-stack images, NeuN-positive cell bodies were segmented in 3D using the pre-trained cyto2 model (Cellpose67). The segmentation quality was manually verified (blinded), and low-quality images were excluded. Cell-level fluorescence intensities were computed as probability-weighted sums of voxel intensities, using segmentation-derived voxel probabilities. Measurements from multiple differentiation batches (independent staining and imaging experiments) were combined by uniformly sampling cells per condition per batch, batch-wise z-scaling fluorescence values, and including batch and well-of-origin indicator variables in downstream analyses. Clipping was minimal (<0.1%), and the confocal microscope response was assumed linear. A linear mixed-effects model (mixedlm() from statsmodels) tested cell-level fluorescence intensities, modelling genotype or treatment as a fixed effect and well of origin as a random effect.
For single-plane live imaging (TMRM, CellROX), images were binarized at the 75th percentile intensity threshold per channel to identify regions occupied by neuronal soma or processes, according to established methodology68. Mean fluorescence intensities were quantified within these masked areas. For time-course imaging, images were spatially aligned by Fourier-based registration (phase cross-correlation), with alignment accuracy confirmed manually. A mask from the baseline (pre-FCCP) TMRM image (75th percentile threshold) was consistently applied across timepoints. For all live-imaging experiments, masked regions (wells) were treated as individual observations in statistical tests. Batch-wise z-scaling was not required here, as data were not combined across batches for these experiments.
One outlier (p.Tyr622*+H20; value 0.34) was identified and removed in the TMRM p.Tyr622* (with or without CDP-choline) experiment using the interquartile range (IQR) method (values outside Q1−2 × IQR or Q3 + 2 × IQR) and removed for plotting convenience. This did not affect the statistical significance of the results.
Aβ ELISA assays
Culture media were collected and analysed for Aβ40 and Aβ42 levels using enzyme-linked immunosorbent assay (ELISA) kits (Thermo Fisher Scientific, KHB3481 and KHB3441, respectively) according to the manufacturer’s protocols. For 4-week-old iNs, media were flash-frozen before analysis. For cortical organoids (aged 5–6 months; days 176–182), media were analysed immediately after collection following 3–4 weeks of treatment with 500 µM or 1 mM CDP-choline.
Electrophysiology recordings
Electrophysiology recordings were performed using the Axon Multiclamp 700B amplifier and Clampex 11.2 software (Molecular Devices). Cells were visualized using infrared differential interference contrast imaging (Olympus BX-50WI microscope), placed in a recording chamber and perfused continuously at 2 ml min−1 (32 °C) with oxygenated artificial cerebrospinal fluid (containing 125 mM NaCl, 2.5 mM KCl, 1.2 mM NaH2PO4·H2O, 2.4 mM CaCl2·2H2O, 1.2 mM MgCl2·6H2O, 26 mM NaHCO3, and 11 mM d-glucose).
Action potentials were elicited by injecting current steps in current-clamp mode. Whole-cell currents were recorded from a holding potential of −80 mV by stepping to various voltages in voltage-clamp mode. Spontaneous firing was recorded in cell-attached configuration. Recordings were filtered at 1 kHz (four-pole Bessel filter), digitized at 10 kHz with a Digidata 1550B interface (Molecular Devices). Pipette solution contained 120 mM K-gluconate, 5 mM KCl, 2 mM MgCl2·6H2O, 10 mM HEPES, 4 mM ATP and 0.2 mM GTP. Data were analysed using pClamp 11.2 and GraphPad Prism 10.
For electrophysiology recordings from cortical organoids, day 150 organoids were dissociated using Accutase (StemCell Technologies, 07920, 40 min, 37 °C), plated onto #1 glass coverslips (Thermo Fisher Scientific, 50-194-4702) coated with PDL, laminin and Matrigel, and maintained in 2D culture with or without 100 µM CDP-choline for 2 weeks before recordings.
Spontaneous action potential outliers were identified using the IQR method (values outside Q1–Q2 × IQR or Q3 +Q2 × IQR) and removed, resulting in the exclusion of two datapoints (9.38 in p.Tyr622*; 6.15 in p.Tyr622* + CDP-choline). Cells recording zero spontaneous potentials (likely glial) were also excluded.
Seahorse metabolic assays and OCR analysis
iPS-cell-derived neurons were differentiated directly in Seahorse XFe96/XF Pro microplates for 28 days before metabolic assays on a Seahorse XFe96 Analyzer. Seahorse XF cell mito stress and oxidation stress tests were conducted according to manufacturer’s protocols using final drug concentrations of 2.5 µM oligomycin, 1 µM FCCP and 0.5 µM rotenone/antimycin. Data were analysed using XFe Assay v.2.6.3.5 software.
OCRs were monitored over time, with curves visually inspected (blinded) to exclude wells not responsive to drug injections. The following OCR metrics were computed from integrals of OCR curves between specific experimental intervals: (1) basal respiration (before oligomycin injection); (2) proton leak (after oligomycin, before FCCP); (3) maximal respiration (after FCCP, before rotenone/antimycin); (4) relative uncoupling (proton leak divided by basal respiration); and (5) spare respiratory capacity (maximal respiration divided by basal respiration).
mRNA-seq and analysis of iNs
Total RNA was extracted from iNs using the RNeasy Mini Kit (Qiagen). RNA quality was assessed (Fragment Analyzer, Agilent), and only samples with RNA quality number > 9.5 were selected. Full-length cDNA libraries were generated (SMART-seq v4 kit, Takara Bio), and sequencing libraries prepared (Nextera XT DNA Library Preparation Kit, Illumina) for sequencing on the Element AVITI platform (Element Biosciences; 75 bp paired-end reads with dual 8-nucleotide indexes) at the MIT BioMicro Center.
Sequencing data were processed through the MIT BioMicro Center BMC/BCC pipeline v1.8 (updated 6 June 2023; https://openwetware.org/wiki/BioMicroCenter:Software#BMC-BCC_Pipeline). Reads were adapter-trimmed (Trim Galore, Nextera-specific settings, minimum overlap 3 bases), aligned to the human reference genome (GRCh38.p14, GENCODE release 47; STAR aligner), and counted (featureCounts, paired-end settings). Read counts were summarized at the exon level and aggregated by gene identifier.
Differential expression analysis (edgeR, limma-voom) retained protein-coding genes expressed at ≥1 CPM in ≥1 sample, normalized counts and used linear modelling with empirical Bayes moderation with contrasts based on experimental conditions (treatment/genotype). fGSEA (10,000 permutations) of WikiPathways gene sets (Supplementary Table 3) was performed using ranked differentially expressed genes (score: sign(log[FC]) × −log10[P]), as described above (see the ‘Gene-set enrichment and Kernighan–Lin pathway clustering’ section). Significant pathways (adjusted P < 0.05) were identified, and leading-edge genes underwent gene–pathway clustering (Kernighan–Lin heuristic, described above). Gene–pathway cluster similarity was assessed by computing Jaccard indices based on pathways and genes assigned to each Kernighan–Lin cluster. The significance of observed overlaps was determined empirically through comparison to 1,000 random permutations, with P values adjusted using the Benjamini–Hochberg method to control the FDR.
LC–MS lipidomics on iNs
iPS-cell-derived neurons were washed in cold PBS, scraped, centrifuged (2,000g, 5 min), counted, pelleted to equal number and resuspended in cold methanol (2 ml). Biphasic extraction was performed by sequentially adding cold chloroform (4 ml) and cold water (2 ml), vortexing after each addition, then centrifuging (3,000 rcf, 10 min) for phase separation. Samples prepared at the Harvard Center for Mass Spectrometry were similarly processed from provided pellets (in 500 µl methanol), supplemented with additional methanol (1.5 ml) and chloroform (4 ml), sonicated (10 min), mixed with water (2 ml) and centrifuged (800 rcf, 10 min, 4 °C). Upper aqueous phases were collected for metabolomics, while chloroform phases were reserved for lipidomics. At least one blank control (no cells) was included in each extraction run. All LC–MS analyses were performed by the Harvard Center for Mass Spectrometry.
Extracted samples were dried under nitrogen, fully evaporated, resuspended in chloroform (scaled by biomass (cell count); ≥60 µl), and split equally for positive and negative ionization analyses (or unsplit if only positive mode). After centrifugation (18,000 rcf, 20 min, 4 °C), the supernatants were transferred into microinserts for LC–MS.
LC–MS analyses were performed on an Orbitrap Exactive plus MS (Thermo Fisher Scientific) consistent with an Ultimate 3000 LC (Thermo Fisher Scientific) in positive- and negative-ionization modes (in WT versus p.Tyr622* only in positive mode), in top five automatic data-dependent MS/MS mode. Chromatography separation was performed on the Biobond C4 column (4.6 × 50 mm, 5 µm particle size; Dikma Technologies). The flow rate began at 100 µl min−1 with 0% mobile phase B (MB) for the initial 5 min, followed by an increase to 400 µl min−1 over the next 50 min with a linear gradient of MB from 20% to 100%. The column was subsequently washed at 500 µl min−1 for 8 min with 100% MB, then re-equilibrated for 7 min at 500 µl min−1 using 0% MB. For positive-ion mode, mobile phases consisted of buffer A (MA: 5 mM ammonium formate, 0.1% formic acid and 5% methanol in water) and buffer B (MB: 5 mM ammonium formate, 0.1% formic acid, 5% water and 35% methanol in isopropanol). For negative-ion mode, buffer A (MA) contained 0.03% ammonium hydroxide and 5% methanol in water, and buffer B (MB) contained 0.03% ammonium hydroxide, 5% water and 35% methanol in isopropanol.
Lipids were identified, and their signals integrated using the Lipidsearch software (v.4.2.27, Mitsui Knowledge Industry, University of Tokyo). Integrations and peak quality were curated manually. Peak areas were background-corrected (subtracting 3× median blank peak areas; negative values set to zero). Statistical analyses were performed using Welch’s t-tests (unequal variance) to compare different cell lines, and Student’s t-tests (equal variance) for treatment comparisons within identical genetic backgrounds.
LC–MS metabolomics on iNs
Samples were dried under nitrogen, evaporated completely and resuspended in biomass-scaled volumes (≥20 µl) of 50% acetonitrile in water. After centrifugation (maximum speed, 10 min), consistent volumes (12 or 15 µl, depending on batch) of supernatants were transferred to microinserts. The remainder of the sample volumes was combined to create a pool sample used for MS2/MS3 data acquisition.
LC–MS metabolomics analyses were performed at the Harvard Center for Mass Spectrometry using a Vanquish LC system coupled with an ID-X mass spectrometer (Thermo Fisher Scientific). Samples (5 µl injection) were analysed on a ZIC-pHILIC peek-coated column (150 mm × 2.1 mm, 5 µm particle size; Sigma-Aldrich) held at 40 °C. Mobile phases comprised buffer A (20 mM ammonium carbonate and 0.1% ammonium hydroxide in water) and buffer B (97% acetonitrile in water). The gradient initiated at 93% B, decreasing linearly to 40% B over 19 min, further decreasing to 0% B over the subsequent 9 min, held at 0% B for 5 min, returned to 93% B within 3 min and finally was re-equilibrated at 93% B for 9 min. The flow rate was held constant at 0.15 ml min−1, except for an initial 30 s ramp from 0.05 to 0.15 ml min−1. MS data were acquired in polarity-switching mode at 120,000 resolution, with an AGC target of 1 × 105, covering an m/z range from 65 to 1,000. MS1 acquisition used polarity switching for all samples. MS2 and MS3 analyses were performed on pooled samples using the AcquireX DeepScan method, with five reinjections each in positive- and negative-ion modes separately. A mixture containing standards of targeted metabolites was prepared and analysed immediately after the sample runs for targeted metabolite analysis.
Data were analysed using Compound Discoverer 3.2 (Thermo Fisher Scientific). Metabolite identification was based either on MS2/MS3 spectral matching against a local mzVault library and corresponding retention times from pure standards (level 1), or spectral matching using mzCloud (level 2). Each metabolite identification was manually inspected. Blank samples were used to exclude background compounds (compounds for which the area in at least one sample was not higher than three times the area in the blanks). Median-centred peak areas were scaled to zero-mean and unit variance before principal component analysis. The Harvard Center for Mass Spectrometry identified three samples with notably low overall metabolite intensities, which were subsequently excluded from downstream analyses.
LC–MS metabolomics on medium
Medium samples (100 µl each) were transferred into microcentrifuge tubes containing 1 ml of methanol and incubated at −20 °C for 2 h. After incubation, the samples were centrifuged at 18,000 rcf for 20 min at −9 °C, and the supernatants were transferred into new tubes and evaporated to dryness under nitrogen flow. The dried samples were resuspended in 50 µl of 30% acetonitrile in water containing 2 mM medronic acid, centrifuged again at 18,000 rcf for 20 min at 4 °C and the resulting supernatants were transferred into glass microinserts for LC–MS analysis.
Peak areas from targeted metabolite analysis of media samples were compared for CDP, CDP-choline and choline. To ensure accurate detection, solvent blanks were analysed: CDP and CDP-choline were not detected in these blanks, while choline was detected at levels several orders of magnitude lower than in medium samples.
Molecular dynamics simulations
ABCA7 structures (unbound-open and bound-closed conformations; Protein Data Bank (PDB): 8EE6, 8EOP) containing the G1527 variant were retrieved from the PDB. The A1527 variant was generated by mutation (Gly to Ala) using PyMOL v.2.0. ABCA7 residues 1517–1756 were embedded in a DPPC membrane (CHARMM-GUI) and oriented according to the OPM database. Four simulations were performed (GROMACS 2022.3; CHARMM36M force field; Supplementary Table 15).
The protein–membrane system was solvated in a cubic box with a minimum distance of 1.0 nm between the protein and the box edge, using the TIP3P water model. Energy minimization was performed using the steepest descent algorithm with a maximum force threshold of 1,000 kJ mol−1 nm−1 to relieve any steric clashes or bad contacts. The system was equilibrated in six phases, each 125 ps long, to equilibrate volume (NVT) and pressure (NPT). The production run, 300 ns long, was performed in the NPT ensemble at 323 K using a V-rescale thermostat and 1 bar using the Parrinello–Rahman barostat. A 2 fs time step with H-bond constraints was used with periodic boundary conditions applied in all directions. Long-range electrostatics were handled using the particle mesh Ewald method with a cut-off of 1.0 nm for non-bonded interactions.
The r.m.s.d. was calculated to monitor the conformational stability of a given structure over the course of the simulation by comparing the position of Cα at time t under simulation to its reference position (in 8EOP or 8EE6). The ϕ and ψ dihedral angles were calculated using the gmx rama tool, followed by post-processing. Secondary structure analysis was performed using gmx dssp -hmode dssp, with subsequent post-processing using custom Python scripts. Visualization of the trajectories was carried out using VMD v.1.94 software. Principal component analysis was conducted on Cα atom positional fluctuations to identify the major conformational changes during the simulation.
Eukaryotic cell lines
Human iPS cell lines used in this study were generated by the Picower Institute for Learning and Memory iPSC core. The initial parental cell line (AG09173) was obtained from the Coriell Institute. HEK293T cells (ATCC, CRL-3216) were sourced from ATCC. iPS cell lines were confirmed by cell marker staining, RNA-seq and karyotyping. No further authentication of HEK293T cells was performed. All cell lines used here tested negative for mycoplasma contamination.
Use of large-language models
ChatGPT (OpenAI) was used to edit portions of the manuscript text for brevity and clarity, and to assist in generating selected plotting code.
Ethics statement
The study protocol involving the use of human stem cells was approved by the Coriell Institutional Review Board (Coriell IRB) in compliance with DHHS regulations (45 CFR Part 46). The initial cell lines were obtained from the Coriell Institute, which ensured that informed consent was received from all donors. Donors were informed that their tissue donations would be used for the creation of cell lines intended for educational and research purposes, and that all biological materials would be anonymized. For post-mortem human brain samples, informed consent was obtained from each participant, and the Religious Orders Study and Rush Memory and Aging Project were approved by an Institutional Review Board (IRB) of Rush University Medical Center.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The primary post-mortem human omics dataset generated in this study was deposited in the AD Knowledge Portal under Synapse ID syn53461705. As the data contain potentially re-identifiable human information, access is controlled. Researchers with a Synapse account can request the dataset by submitting a data use certificate (DUC) at https://help.adknowledgeportal.org/apd/Data-Use-Certificates.2623373330.html; requests are reviewed within approximately 1 week, and approved users may download the data for 12 months under the terms of the DUC. Redacted data can also be explored through the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP3182). Bulk mRNA-seq fastq files and count matrices are publicly available in the Gene Expression Omnibus: GSE299277. All other data are deposited through the Open Science Framework (https://osf.io/pqr9m/). External datasets analysed (but not generated) in this work are listed in Supplementary Table 3 with URLs and accession IDs.
Code availability
All code used in this study is available on GitHub (https://github.com/djunamay/ABCA7lof2) and deposited at Zenodo69 (https://doi.org/10.5281/zenodo.15722817).
References
Steinberg, S. et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat. Genet. 47, 445–447 (2015).
Holstege, H. et al. Exome sequencing identifies rare damaging variants in ATP8B4 and ABCA1 as risk factors for Alzheimer’s disease. Nat. Genet. 54, 1786–1794 (2022).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Abe-Dohmae, S. et al. Human ABCA7 supports apolipoprotein-mediated release of cellular cholesterol and phospholipid to generate high density lipoprotein. J. Biol. Chem. 279, 604–611 (2004).
Wang, N. et al. ATP-binding cassette transporter A7 (ABCA7) binds apolipoprotein A-I and mediates cellular phospholipid but not cholesterol efflux. J. Biol. Chem. 278, 42906–42912 (2003).
Tomioka, M. et al. Lysophosphatidylcholine export by human ABCA7. Biochim. Biophys. Acta 1862, 658–665 (2017).
Satoh, K., Abe-Dohmae, S., Yokoyama, S., St George-Hyslop, P. & Fraser, P. E. ATP-binding cassette transporter A7 (ABCA7) loss of function alters Alzheimer amyloid processing. J. Biol. Chem. 290, 24152–24165 (2015).
Kim, W. S. et al. Deletion of Abca7 increases cerebral amyloid-β accumulation in the J20 mouse model of Alzheimer’s disease. J. Neurosci. 33, 4387–4394 (2013).
Fu, Y., Hsiao, J.-H. T., Paxinos, G., Halliday, G. M. & Kim, W. S. ABCA7 mediates phagocytic clearance of amyloid-β in the brain. J. Alzheimers Dis. 54, 569–584 (2016).
Aikawa, T. et al. ABCA7 haplodeficiency disturbs microglial immune responses in the mouse brain. Proc. Natl Acad. Sci. USA 116, 23790–23796 (2019).
Kawatani, K. et al. ABCA7 deficiency causes neuronal dysregulation by altering mitochondrial lipid metabolism. Mol. Psychiatry 29, 809–819 (2023).
Liu, Y. et al. Plasma lipidome is dysregulated in Alzheimer’s disease and is associated with disease risk genes. Transl. Psychiatry 11, 344 (2021).
Duchateau, L., Wawrzyniak, N. & Sleegers, K. The ABC’s of Alzheimer risk gene ABCA7. Alzheimers Dement. 20, 3629–3648 (2024).
Tayran, H. et al. ABCA7-dependent induction of neuropeptide Y is required for synaptic resilience in Alzheimer’s disease through BDNF/NGFR signaling. Cell Genom. 4, 100642 (2024).
Brase, L. et al. Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers. Nat. Commun. 14, 2314 (2023).
Blanchard, J. W. et al. APOE4 impairs myelination via cholesterol dysregulation in oligodendrocytes. Nature 611, 769–779 (2022).
Sayed, F. A. et al. AD-linked R47H-TREM2 mutation induces disease-enhancing microglial states via AKT hyperactivation. Sci. Transl. Med. 13, eabe3947 (2021).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Welch, G. M. et al. Neurons burdened by DNA double-strand breaks incite microglia activation through antiviral-like signaling in neurodegeneration. Sci. Adv. 8, eabo4662 (2022).
Naj, A. C. et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat. Genet. 43, 436–441 (2011).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385 (2023).
Le T. M. L. et al. Cryo‐EM structures of human ABCA7 provide insights into its phospholipid translocation mechanisms. EMBO J. 42, e111065 (2022).
Fang, S.-C. et al. Structural insights into human ABCA7-mediated lipid transport. Structure 33, 583–593 (2025).
De Roeck, A. et al. Deleterious ABCA7 mutations and transcript rescue mechanisms in early onset Alzheimer’s disease. Acta Neuropathol. 134, 475–487 (2017).
Ho, S.-M. et al. Rapid Ngn2-induction of excitatory neurons from hiPSC-derived neural progenitor cells. Methods 101, 113–124 (2016).
Divakaruni, A. S. & Jastroch, M. A practical guide for the analysis, standardization and interpretation of oxygen consumption measurements. Nat. Metab. 4, 978–994 (2022).
Divakaruni, A. S. & Brand, M. D. The regulation and physiology of mitochondrial proton leak. Physiology 26, 192–205 (2011).
Jain, A. et al. Mitochondrial uncoupling proteins protect human airway epithelial ciliated cells from oxidative damage. Proc. Natl Acad. Sci. USA 121, e2318771121 (2024).
Kumar, R., T, A., Singothu, S., Singh, S. B. & Bhandari, V. Uncoupling proteins as a therapeutic target for the development of new era drugs against neurodegenerative disorder. Biomed. Pharmacother. 147, 112656 (2022).
Zhao, Y. et al. Identification and characterization of a major liver lysophosphatidylcholine acyltransferase. J. Biol. Chem. 283, 8258–8265 (2008).
Boumann, H. A. et al. Depletion of phosphatidylcholine in yeast induces shortening and increased saturation of the lipid acyl chains: evidence for regulation of intrinsic membrane curvature in a eukaryote. Mol. Biol. Cell 17, 1006–1017 (2006).
Sienski, G. et al. APOE4 disrupts intracellular lipid homeostasis in human iPSC-derived glia. Sci. Transl. Med. 13, eaaz4564 (2021).
Westermann, B. Mitochondrial fusion and fission in cell life and death. Nat. Rev. Mol. Cell Biol. 11, 872–884 (2010).
Giblin, A. et al. Neuronal polyunsaturated fatty acids are protective in ALS/FTD. Nat. Neurosci. 28, 737–747 (2025).
Yaghmour, M. H., Sajeevan, T., Thiele, C. & Kuerschner, L. Phosphatidylcholine synthesis and remodeling in brain endothelial cells. J. Lipid Res. 66, 100773 (2025).
O’Donnell, V. B. New appreciation for an old pathway: the lands cycle moves into new arenas in health and disease. Biochem. Soc. Trans. 50, 1–11 (2022).
Decker, S. T. & Funai, K. Mitochondrial membrane lipids in the regulation of bioenergetic flux. Cell Metab. 36, 1963–1978 (2024).
Crivelli, S. M., Gaifullina, A. & Chatton, J.-Y. Exploring the role of mitochondrial uncoupling protein 4 in brain metabolism: implications for Alzheimer’s disease. Front. Neurosci. 18, 1483708 (2024).
Morant-Ferrando, B. et al. Fatty acid oxidation organizes mitochondrial supercomplexes to sustain astrocytic ROS and cognition. Nat. Metab. 5, 1290–1302 (2023).
Korshunov, S. S., Skulachev, V. P. & Starkov, A. A. High protonic potential actuates a mechanism of production of reactive oxygen species in mitochondria. FEBS Lett. 416, 15–18 (1997).
Andrews, Z. B., Diano, S. & Horvath, T. L. Mitochondrial uncoupling proteins in the CNS: in support of function and survival. Nat. Rev. Neurosci. 6, 829–840 (2005).
Welch, G. & Tsai, L.-H. Mechanisms of DNA damage-mediated neurotoxicity in neurodegenerative disease. EMBO Rep. 23, e54217 (2022).
Cornell, R. B. Membrane lipid compositional sensing by the inducible amphipathic helix of CCT. Biochim. Biophys. Acta 1861, 847–861 (2016).
Takada, N. et al. Phospholipid-flipping activity of P4-ATPase drives membrane curvature. EMBO J. 37, e97705 (2018).
Ballweg, S. et al. Regulation of lipid saturation without sensing membrane fluidity. Nat. Commun. 11, 756 (2020).
Ariyama, H., Kono, N., Matsuda, S., Inoue, T. & Arai, H. Decrease in membrane phospholipid unsaturation induces unfolded protein response. J. Biol. Chem. 285, 22027–22035 (2010).
Gavrilova, S. I., Kolykhalov, I. V., Ponomareva, E. V., Fedorova, Y. B. & Selezneva, N. D. Clinical efficacy and safety of choline alfoscerate in the treatment of late-onset cognitive impairment. Zh. Nevrol. Psikhiatr. Im. S S Korsakova 118, 45–53 (2018).
Mathys, H. et al. Single-cell multiregion dissection of Alzheimer’s disease. Nature 632, 858–868 (2024).
Velazquez, R. et al. Lifelong choline supplementation ameliorates Alzheimer’s disease pathology and associated cognitive deficits by attenuating microglia activation. Aging Cell 18, e13037 (2019).
Karosas, T. et al. Dietary choline intake and risk of Alzheimer’s dementia in older adults. J. Nutr. 155, 2322–2332 (2025).
Wang, H., Bennett, D. A., De Jager, P. L., Zhang, Q.-Y. & Zhang, H.-Y. Genome-wide epistasis analysis for Alzheimer’s disease and implications for genetic risk prediction. Alzheimers Res. Ther. 13, 55 (2021).
Haney, M. S. et al. APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia. Nature 628, 154–161 (2024).
Victor, M. B. et al. Lipid accumulation induced by APOE4 impairs microglial surveillance of neuronal-network activity. Cell Stem Cell 29, 1197–1212 (2022).
Bennett, D. A. et al. Religious orders study and rush memory and aging project. J. Alzheimers Dis. 64, S161–S189 (2018).
Fort, A. et al. MBV: a method to solve sample mislabeling and detect technical bias in large combined genotype and sequencing assay datasets. Bioinformatics 33, 1895–1897 (2017).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kernighan, B. W. & Lin, S. An efficient heuristic procedure for partitioning graphs. Bell Syst. Tech. J. 49, 291–307 (1970).
He, Z. et al. Comprehensive transcriptome analysis of neocortical layers in humans, chimpanzees and macaques. Nat. Neurosci. 20, 886–895 (2017).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Lin, Y.-T. et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1141–1154 (2018).
De Roeck, A., Van Broeckhoven, C. & Sleegers, K. The role of ABCA7 in Alzheimer’s disease: evidence from genomics, transcriptomics and methylomics. Acta Neuropathol. 138, 201–220 (2019).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Sloan, S. A., Andersen, J., Pașca, A. M., Birey, F. & Pașca, S. P. Generation and assembly of human brain region-specific three-dimensional cultures. Nat. Protoc. 13, 2062–2085 (2018).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Esteras, N., Adjobo-Hermans, M. J. W., Abramov, A. Y. & Koopman, W. J. H. Visualization of mitochondrial membrane potential in mammalian cells. Methods Cell Biol. 155, 221–245 (2020).
von Maydell, D. et al. Code for ‘ABCA7 variants impact phosphatidylcholine and mitochondria’. Zenodo https://doi.org/10.5281/zenodo.15722817 (2025).
Acknowledgements
The results published here are in whole or in part based on data obtained from the AD Knowledge Portal. We thank the individuals who donated post-mortem brain samples, and their families, for enabling this research; Y. Zhou, E. McNamara and T. V. Garvey for administrative support; U. Geigenmüller for reviewing and editing the manuscript and for discussions; R. Pinals, M. Victor, J. Davila-Velderrain and R. Firenze for discussions and input on the manuscript; D. A. Bennett and G. Klein for providing the post-mortem samples; the staff at the MIT SuperCloud and Lincoln Laboratory Supercomputing Center for providing HPC and consultation resources that have contributed to the research results reported within this paper; the staff at the Harvard Center for Mass Spectrometry (HCMS) for lipidomic and metabolomic sample runs and quantifications and C. Vidoudez for consultation on the lipidomic data analysis and experimental procedures; the members of the MIT BioMicro Center for bulk and single-nucleus RNA-seq runs; and S. Levine and his team for consultations on data processing. This work was supported in part by the Cure Alzheimer’s Fund, The Freedom Together Foundation, the Carol and Gene Ludwig Family Foundation, James D. Cook and NIH grants R56-AG081376,RF1-AG062377, RF1-AG054321, RO1-AG054012 (L.-H.T.) and RO1-AG075901 (principal investigator, E. Fraenkel). Graphic illustrations were generated using BioRender under agreements PA28FD1SCF, IW28ETKEWW, CA28ETIAJQ, DU28ETMKEJ and RS28ETN9LG.
Author information
Authors and Affiliations
Contributions
D.v.M. and L.-H.T. designed the study, with L.-H.T. supervising the overall project and acquiring funding alongside M.K. Experimental work included snRNA-seq experiments performed by D.v.M., J.M.B., H.M. and L.L. Seahorse assays conducted by S.E.W. Differentiation and maintenance of iPS cell lines, iNs and cortical organoids were conducted by S.E.W., C.S., P.-C.P. and O.K.; S.E.W. and P.-C.P. prepared samples for LC–MS analysis and performed amyloid ELISA experiments and neuronal marker staining. P.-C.P. conducted TMRM and CellRox assays and S.E.W. performed MitoHealth assays. P.-C.P., C.S. and O.K. prepared bulk RNA samples. L.L. conducted electrophysiological recordings and analysis, and C.J.Y. generated the p.Tyr622* cell line. A.S. performed molecular dynamics simulations, analyses and visualizations and D.v.M. performed formal data analysis and visualization, with contributions from C.A.B., G.L. and A.S. Experimental and technical support was provided by A.N.S., M.L., G.S.M., G.W., A.G., N.L. and G.S.; C.-C.C. and D.L. helped with revision experiments. D.v.M., S.E.W. and L.-H.T. wrote the first draft of the manuscript. D.v.M. and L.-H.T. wrote the revised draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Johan Auwerx, Konstantin Khodosevich and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Overview of human snRNA-seq cohort.
a, ABCA7 and RBFOX3 (NeuN) protein levels from postmortem human PFC (Supplementary Table 3) comparing control (N = 180 individuals) vs. ABCA7 LoF carriers (N = 5, “All”) and subset overlapping with snRNA-seq cohort (N = 6 control, N = 4 ABCA7 LoF, “Subset”). b, Distribution of continuous clinical variables (Supplementary Note 1) comparing control (N = 24) vs. ABCA7 LoF carriers (N = 12). c, Distribution of discrete metadata variables comparing control (N = 24) vs. ABCA7 LoF carriers (N = 12); Fisher’s exact tests (R stats::fisher.test, extended to r × c tables). d, Sanger sequencing validating three ABCA7 LoF variants in genomic DNA from ABCA7 LoF carriers and controls. Variant location indicated by black box; WGS sample IDs shown. e, Validation plots demonstrating concordance between SNP calls from WGS and snRNA-seq libraries per individual. Extreme outlier points (dark blue) indicate correct matches. f, 2D UMAP projection of single-cell gene expression coloured by transcriptionally defined cell type. a,b: boxplots show median, IQR (box), whiskers=1.5×IQR; two-sided Mann–Whitney U tests. a–f: experiments performed once.
Extended Data Fig. 2 Expanded view of ABCA7 LoF gene perturbation landscape.
a, 2D UMAP of ABCA7 LoF gene perturbation scores (S = −log10(p) × sign(log2(FC)); unadjusted Limma-Voom p-values, N subjects=12 LoF, 24 control), restricted to genes with |S | > 1.3. Red: S > 1.3, blue: S < −1.3; point size ∝ |S|. Top 50 genes labelled.
Extended Data Fig. 3 Shared differentially expressed genes across cell types.
a, Heatmap indicating overlap of differentially expressed genes (unadjusted Limma-Voom p < 0.05 in ≥3 cell types). b, Functional annotations of genes shown in the heatmap (same gene order as panel a).
Extended Data Fig. 4 Neuronal expression of ABCA7 in postmortem human brain.
a, ABCA7 detection rate (counts > 0) per major cell type in postmortem PFC (snRNA-seq; N cells=Ex 42,014, In 14,806, Ast 7,158, Mic 5,441, Oli 28,078, Opc 5,213). b, Normalized ABCA7 expression comparing glial cells (mean per individual across Oli, Opc, Ast, Mic) vs. neuronal cells (mean per individual across Ex, In) from snRNA-seq (N = 24 control, 12 LoF). Two-sided paired Wilcoxon test following Shapiro test. c, Normalized expression of indicated genes comparing NeuN− vs. NeuN+ cells (N = 6 individuals; 3 control, 3 AD, γH2AX− cells; data from ref. 19, Supplementary Table 3). Two-sided paired t-test following Shapiro test. a–b: boxplots show median, IQR (box), whiskers=1.5×IQR.
Extended Data Fig. 5 Annotation of excitatory neurons from postmortem snRNA-seq dataset by cortical layer.
a, UMAP visualization of excitatory neurons annotated by cortical layers (Leiden clustering; N cells=42,014 from ABCA7 LoF snRNA-seq cohort). b, Heatmap showing enrichment of cortical layer-specific marker genes (from ref. 61) across annotated layers. Colours indicate average marker gene expression (log2(fold-change)) of each layer’s marker genes relative to all other clusters. c, Heatmap validating layer annotations using an independent set of cortical layer marker genes (from ref. 62). Colours represent average marker gene expression (log2(fold-change)) relative to all other clusters. d, Perturbation of ABCA7 LoF-associated gene clusters identified in all excitatory neurons (Fig. 2a), stratified by cortical layer (N subjects=24 control, 12 LoF; fGSEA analysis of clusters 0–7). Normalized enrichment scores (NES) and unadjusted p-values shown.
Extended Data Fig. 6 Molecular dynamics simulations of ABCA7 open conformations with p.Ala1527Gly substitution.
a, Open-conformation ABCA7 protein structure highlighting simulation domain (residues 1517–1756, yellow). Inset shows expanded view of structures with Ala1527 (grey) and Gly1527 (purple) variants. b, Further expanded inset from panel a. c, Root mean squared deviations (RMSD) of the open-conformation ABCA7 domain (panel b) carrying Ala1527 (grey) or Gly1527 (purple), measured relative to open-reference structure during simulations. d, Projection of Cα atom positional fluctuations onto first two principal components during simulations for Ala1527 (top, grey) and Gly1527 (bottom, purple) variants. e, Violin plot showing average positional fluctuations of Cα atoms; Mann-Whitney test, two-sided; **** = p < =1e-4.
Extended Data Fig. 7 Local conformational fluctuations and secondary structure changes induced by the p.Ala1527Gly substitution in ABCA7 open and closed conformations.
a, Phi vs. Psi dihedral angle distribution of residue 1527 over simulation time in open and closed ABCA7 conformations. b, Overall Phi vs. Psi angle distributions of residue 1527 across the entire simulation, comparing open and closed conformations. c, Time-resolved secondary structure assignments for residues 1517–1537. Alpha-helical regions highlighted in red; other colours indicate distinct secondary structures. d, Fraction of alpha-helical content for residues 1517–1537 during simulations. A value of 1 indicates continuous alpha-helical structure throughout duration of the simulation. e, Structural alignment of closed-conformation ABCA7 (purple; PDB ID: 8EOP) with ABCA1 (cyan; PDB ID: 7TBW). Gly1527 (ABCA7) and corresponding residue Val1646 (ABCA1) indicated as spheres. f, Structural alignment of closed-conformation ABCA7 (purple; PDB ID: 8EOP) with ABCA4 (green; PDB ID: 7LKZ). Gly1527 (ABCA7) and corresponding residue Ile1671 (ABCA4) indicated as spheres. a,d: G1527 refers to the ABCA7 structure with the risk variant (as present in the reference structures; Supplementary Table 15); G1527A refers to the ABCA7 structure with the mutated Gly→Ala change made to the reference structure in PyMOL.
Extended Data Fig. 8 Electrophysiological characterization of iPSC-derived neurons harbouring ABCA7 PTC variants.
a, Representative sweeps showing action potentials elicited by 800 ms current injections in patched iNs. b, Action potential frequency (mean ± s.e.m.) elicited by varying injected currents in 4-week-old iNs; N cells = 10 WT, 13 p.Tyr622*, 23 p.Glu50fs*3. c, Representative sweeps of inward (top) and outward (bottom) currents recorded in 4-week-old WT neurons (N cells = 23). d, Quantification of currents from panel c. e, Resting membrane potential (mV) in 4-week-old WT, ABCA7 p.Tyr622*, and ABCA7 p.Glu50fs*3 iNs. f, Rheobase (pA) in 4-week-old WT, ABCA7 p.Tyr622*, and ABCA7 p.Glu50fs*3 iNs; WT vs p.Tyr622* p = 0.0424; WT vs p.Glu50fs*3 p = 0.0200. g, Action potential frequency elicited by indicated current injections in 4-week-old WT, ABCA7 p.Tyr622*, and ABCA7 p.Glu50fs*3 iNs; 10 pA: WT vs p.Glu50fs*3 p = 0.0491; 15 pA: WT vs p.Tyr622* p = 0.0003 and WT vs p.Glu50fs*3 p = 0.0109; 20 pA: WT vs p.Tyr622* p = 0.0007 and WT vs p.Glu50fs*3 p = 0.0160. e–g: n = 24 WT, 13 p.Tyr622*, 23 p.Glu50fs*3 4-week-old iNs. Bar plots indicate mean ± s.e.m. P values by two-way ANOVA indicated as: P < 0.05 (*), P < 0.001 (***).
Extended Data Fig. 9 mRNA-seq analysis of p.Glu50fs*3 iNs.
a, K/L clustering of leading-edge genes from perturbed pathways in WT vs. p.Glu50fs*3 iNs (fGSEA, Wikipathways; FDR-adjusted p < 0.05; N wells=2 WT, 5 p.Glu50fs*3). Gene-pathway graph: genes (circles), pathways (squares). b, Heatmap (Jaccard index) comparing K/L clusters from p.Glu50fs*3 iNs vs. p.Tyr622* iNs (from Fig. 3c). Average per-cluster gene score S (S = −log10(p)*sign(log2(FC))) indicated; \(\bar{S} > 0\) (red); \(\bar{S} < 0\) (blue); FDR-adjusted permutation p-values (1000 iterations, one-sided). c, Heatmap (Jaccard index) comparing K/L clusters from p.Glu50fs*3 iNs vs. postmortem neurons (Fig. 2a,b). Red/blue indicates direction as described in c; FDR-adjusted permutation p-values (1000 iterations, one-sided). d, Kernel density plots of gene perturbation scores S; unadjusted Limma-Voom p-values; N wells=2 WT, 5 per LoF) per cluster. Positive S indicates increased expression in p.Glu50fs*3. Solid lines show cluster means; top enriched pathways indicated. e, Correlation of gene perturbation scores S for genes encoding mitochondrial proteins (MitoCarta database, Supplementary Table 3; N wells=2 WT, N = 5 per LoF line). a–e: 4-week differentiation; wells=technical replicates; same mRNAseq experiment as Fig. 3b; experiment performed once.
Extended Data Fig. 10 Analysis of oxygen consumption rates (OCR) in WT vs ABCA7 LoF iNs.
a, Seahorse OCR curves (one representative experiment from Fig. 3g). Lines indicate per-condition mean; error bars represent 95% confidence intervals (N wells=10 WT, 7 per LoF line). b, Representative per-well OCR traces from panel a. c, Simplified schematic of mitochondrial bioenergetics in electrical terms: oxygen consumption by the respiratory chain generates a proton current (I) that builds the proton-motive force (Δp, voltage V); proton return through ATP synthase (cyan) for ATP production or through uncoupling protein (red) provides parallel resistances (R) that dissipate Δp; BioRender agreement # RS28ETN9LG; based on information in27. d, Schematic depicting measurements of maximal and basal OCR used to calculate Spare Respiratory Capacity (SRC). e, Schematic showing measurement of uncoupled OCR (%). f, SRC computed for WT, ABCA7 p.Glu50fs*3, and ABCA7 p.Tyr622* iNs (N wells=18 WT, 17 p.Tyr622*, 13 p.Glu50fs*3; two experiments, same as Fig. 3g). Two-sided Mann–Whitney U test (WT vs. p.Tyr622*) and two-sided t-test (WT vs. p.Glu50fs*3, unequal variances), following Shapiro/Levine tests. g, UCP2 mRNA expression by genotype (N wells=2 WT, 5 per LoF; unadjusted Limma-Voom p-values). h, Average TMRM intensity per masked region (75th percentile threshold; N wells=4 WT, 5 p.Tyr622*; mean projection over time), under baseline conditions and after FCCP addition. Same experiment and images as Fig. 3i. Two-sided t-test assuming equal variance, following Shapiro/Levine tests for both comparisons. f, h: boxplots show median, IQR (box), whiskers=1.5×IQR. a–h: 4-week iN differentiation; g: experiment performed once, a, b, f, h ≥twice.
Extended Data Fig. 11 LC-MS lipidomics of ABCA7 LoF iNs.
a, Volcano plot of WT vs p.Glu50fs*3 showing differentially abundant lipid species, coloured by class. b, Table summarizing the significant species by lipid subclass. c, Fold-change distribution for triglycerides (TG) grouped by fatty-acid chain length and saturation (WT vs p.Glu50fs*3). d, Volcano plot highlighting perturbed phosphatidylcholines (PCs) that contain saturated or monounsaturated fatty acids (SFA/MUFA). e, As d but for PCs with polyunsaturated fatty acids (PUFA). f, Fold-change distribution for PCs grouped by chain length and saturation (WT vs p.Glu50fs*3). g, Table of significantly altered lipid species in WT vs p.Tyr622*, grouped by subclass (same experiment as Fig. 3k). h, Volcano plot comparing WT vs p.Tyr622* with saturated PCs highlighted in blue. i, TG fold-change distribution for WT vs p.Tyr622*. j, PC fold-change distribution for WT vs p.Tyr622*. k,l, mRNA expression changes of LPCAT genes in p.Tyr622* and p.Glu50fs*3 iNs relative to WT (unadjusted p by Limma-Voom; N wells=2 WT, 5 per LoF line). a-l: iNs differentiated for 4 weeks. a-j: Two-sided t-test assuming unequal variance; FDR-adjusted p < 0.05 and |log2(FC)| > 1 used to define significance. N wells= 10 WT, 8 p.Tyr622*, 6 p.Glu50fs*. Wells are technical replicates. g-l: experiment performed once, a-f ≥twice.
Extended Data Fig. 12 Treatment of p.Tyr622* iNs with CDP-choline.
a, Choline metabolites in media (targeted LC-MS; N wells=2 media-only, 4 cell-conditioned); “N/F”=not detected. b, Intracellular choline metabolites (targeted LC-MS; N = 8 wells/condition, 4 blanks). c, Differentially expressed choline synthesis and transport genes in p.Tyr622* ± CDP-choline iNs. d, Differentially expressed LPCAT genes in p.Tyr622* ± CDP-choline iNs. e, Phosphatidylcholine species log2 fold-changes by fatty acid chain length and saturation in p.Tyr622* ± CDP-choline (lipidomics; N = 5 wells/condition). f, PCA of untargeted LC-MS metabolite profiles from WT and p.Tyr622* ± CDP-choline iNs (N wells=9 WT, 7 per p.Tyr622* condition). g, PCA of mRNA-seq data from p.Tyr622* ± CDP-choline iNs. h, Correlation of gene perturbation scores (S = −log10(p) × sign(log2(FC)); unadjusted Limma-Voom p-values) for genes encoding mitochondrial proteins in WT vs p.Tyr622* and p.Tyr622* ± CDP-choline iNs (MitoCarta; Supplementary Table 3; N = 2 WT, 5 per p.Tyr622* condition). i, Seahorse OCR curves; lines=mean, error bars=95% CI. j, Representative OCR traces from panel i. k, SRC in p.Tyr622* ± CDP-choline (two-sided t-test, equal variances after Shapiro/Levene tests). l, MitoHealth intensity per NeuN+ volume (N = 11 wells p.Tyr622*+H2O, 12 p.Tyr622*+CDP-choline; ~3 × 10³ cells/condition; three experiments; linear mixed-effects model). Visualization: maximum projections, NeuN/GFP clipped at maximum, γ-corrected (γ = 0.5). Individual points=well averages; same experiment as Fig. 3h. a,b: two-sided t-tests (equal variances). c,d,g: mRNA-seq; unadjusted Limma-Voom p-values; N = 5 wells/condition. i–k: N wells=8 p.Tyr622*+CDP-choline, 6 p.Tyr622*+H2O (same experiment as Fig. 4g). a,b,k,l: boxplots=median, IQR (box), whiskers=1.5×IQR. a–l: 4-week iN differentiation; last 2 weeks, 100 µM CDP-choline. a-l: wells=technical replicates. a,b,e,f,i–k: experiments repeated once, c,d,g,h,l ≥twice.
Extended Data Fig. 13 CDP-choline treatment in cortical organoids.
a, Amyloid-β (Aβ40, Aβ42) levels quantified by ELISA from media of 4-week-old iNs (N wells=12 WT, 8 p.Tyr622*). Aβ40: two-sided t-test (equal variances); Aβ42: two-sided Mann–Whitney U test, following Shapiro–Levene tests. b, Representative images of cortical organoid slices by genotype; 5.5 months. c, Amyloid-β (Aβ40, Aβ42) levels quantified by ELISA from cortical organoid media (176-day-old), grouped by genotype and treatment (500 µM or 1 mM CDP-choline for 3 weeks). Samples correspond to organoids in Fig. 4k, analysed one week prior to assays presented there (N organoids=20 WT, 20 p.Tyr622*+H2O, 15 p.Tyr622*+500 µM CDP-choline, 14 p.Tyr622*+1 mM CDP-choline). Statistical comparisons (two-sided t-tests): Aβ40: WT vs p.Tyr622* (unequal variances), p.Tyr622*±500 µM (unequal variances), p.Tyr622*±1 mM (equal variances). Aβ42: all comparisons (equal variances). All tests followed Shapiro–Levene tests. a,c: boxplots=median, IQR (box), whiskers=1.5×IQR. a-c: experiments performed once.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–3, Supplementary Tables 1–3, 11 and 14–17, and Supplementary Figs. 1–6.
Supplementary Table 4 (download CSV )
Metadata for ROSMAP cohort individuals analysed. Sample_Id: snRNA-seq sample identifier. ABCA7LoF: indicates the presence (1) or absence (0) of ABCA7 premature termination variant (Methods). Passed_QC: snRNA-seq quality control status (Methods). Additional variables are described in Supplementary Note 1.
Supplementary Table 5 (download CSV )
Technical parameters from Cell Ranger analysis for snRNA-seq samples.
Supplementary Table 6 (download CSV )
Differential expression analysis results on snRNA-seq data for genes comparing ABCA7 LoF variant carriers to controls within each cell type. The columns report the log[FC], average expression (AveExpr) and statistical significance (t-statistic, P value, adjusted P value).
Supplementary Table 7 (download CSV )
Summary of gene clusters identified from the UMAP projection of ABCA7 LoF-associated differential expression scores across cell types. Each cluster lists enriched Gene Ontology pathways, the number of overlapping genes, hypergeometric enrichment P values and associated genes contributing to each pathway enrichment.
Supplementary Table 8 (download XLS )
Enriched pathways identified by fGSEA (WikiPathways database; cutoff P < 0.05) comparing ABCA7 LoF to control excitatory neurons. The columns include pathway, raw and adjusted P values, NES, leading-edge genes, cluster assignments (Kernighan–Lin partitioning) and cluster annotations.
Supplementary Table 9 (download CSV )
Differential expression analysis results from mRNA-seq comparing WT, p.Tyr622* with or without CDP-choline treatment and p.Glu50fs*3 iNs. The columns report the log[FC], average expression, t-statistic, P value and adjusted P value.
Supplementary Table 10 (download XLS )
Enriched pathways identified by fGSEA (WikiPathways database; cut-off Padj < 0.05) comparing WT, p.Tyr622* with or without CDP-choline treatment, and p.Glu50fs*3 iNs. The columns include pathway, raw and adjusted P values, NES, leading-edge genes, cluster assignments (Kernighan–Lin partitioning) and cluster annotations.
Supplementary Table 12 (download CSV )
Differential lipid abundance results comparing WT, p.Tyr622* with or without CDP-choline treatment, and p.Glu50fs*3 iNs. The columns include lipid species (name), lipid group (grp), lipid class, log2[FC], raw and FDR-corrected P values, and group comparisons. Undefined indicates a signal below the blank threshold.
Supplementary Table 13 (download CSV )
Targeted metabolomics of p.Tyr622* iNs and culture medium comparing the CDP-choline-treated and untreated conditions. The columns show peak areas for CDP, CDP-choline and choline in samples and blanks.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
von Maydell, D., Wright, S.E., Pao, PC. et al. ABCA7 variants impact phosphatidylcholine and mitochondria in neurons. Nature 647, 462–471 (2025). https://doi.org/10.1038/s41586-025-09520-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09520-y