Abstract
Ferroptosis has emerged as an actionable target to eliminate therapy-resistant and metastatic cancers1. However, which ferroptosis surveillance systems may offer a therapeutic window to leverage redox maladaptation in cancer remains unclear. In melanoma, glutathione peroxidase 4 (GPX4) impedes ferroptosis during haematogenous metastasis, but is dispensable during lymphatic metastasis2. Here, using a metastatic mouse melanoma model selected for lymph node metastasis, we show that lymph-node-derived metastatic cells exhibit markedly diminished expression of glutamate–cysteine ligase (GCLC) and reduced glutathione (GSH) levels relative to their parental counterparts. This metabolic shift occurs within the hypoxic lymphatic niche. Under comparable low-oxygen conditions, GPX4 undergoes ubiquitination and proteasomal degradation. In response, lymph node metastatic cells acquire increased reliance on ferroptosis suppressor protein 1 (FSP1), which is localized with perinuclear lysosomes. These findings reveal that the reduced reliance on the GPX4 axis enables melanoma cells to shift toward FSP1 dependency. Notably, intratumoural monotherapy with selective FSP1 inhibitors (viFSP1 and FSEN1) effectively suppresses melanoma growth in lymph nodes, but not in subcutaneous tumours, emphasizing a microenvironment-specific dependency on FSP1. Thus, targeting FSP1 in the lymph nodes holds strong potential for blocking melanoma progression.
Similar content being viewed by others
Main
Unrestrained iron-dependent lipid oxidation triggers ferroptosis—a non-apoptotic cell death modality with far-reaching implications for human disease, including neurodegeneration, ischaemia–reperfusion injury and cancer3,4. GPX4 is the guardian of ferroptosis and uses GSH to detoxify and reduce lipid oxidation5,6. Thus, depletion of cellular cysteine or GPX4 inhibition, among other processes, can result in the lethal accumulation of oxidized lipids predominantly on specific polyunsaturated-fatty-acid-containing phospholipids, eventually leading to rupture of the plasma membrane7.
Inducing ferroptosis has emerged as a possible strategy to target therapy-resistant and metastatic cancers8. Nonetheless, cancer cells adeptly leverage effective oxidation–reduction adaptations to mitigate uncontrolled lipid oxidation, including the FSP1–ubiquinone pathway, among others9,10,11,12. In the context of cancer metastasis to lymph nodes (LNs), melanoma cells are transiently protected from ferroptosis by incorporating high levels of oleic acid into their phospholipid membranes2. Notably, whereas metastasizing melanoma cells in the blood are GPX4 dependent and undergo death by oxidative stress2,13,14, metastasizing melanoma cells in the lymph are GPX4 independent but remain protected against ferroptosis2.
Comparing distinct microenvironmental contexts in which cancer cells display differential sensitivity or dependency on GPX4 provides a route to elucidate the complex and still poorly understood mechanisms governing ferroptosis vulnerability and resistance. To explore this, we used an in vivo mouse model of melanoma metastasis selected for LN metastasis compared with subcutaneous (s.c.; primary tumour) lines15. This model revealed a mechanism of context-dependent regulation of ferroptosis resistance mediated by the LN microenvironment and highlighted potential therapeutic opportunities to inhibit tumour growth in LNs.
LN colonization shifts ferroptosis defences
To investigate how LN colonization influences ferroptosis surveillance dependencies in melanoma, we used an in vivo mouse model of melanoma metastasis, generated by selecting for LN metastases across nine generations using a strategy adapted from other organ-specific metastasis models15. C57BL/6J mice were s.c. implanted with the syngeneic melanoma cell line B16-F0, hereafter, the parental line. Spontaneous LN metastases were isolated, expanded ex vivo and reimplanted into naive mice over nine generations, generating nearly 300 unique LN metastatic lines15 (Fig. 1a). Analysis of metastatic incidence in 30 cell lines across different generations revealed that the late-generation LN metastatic lines (LN7, LN8 and LN9, from generations 7 to 9) exhibit an increased incidence of spontaneous LN metastases compared with the parental lines15.
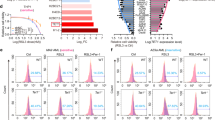
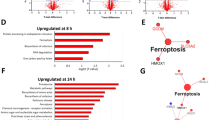
a, Schematic of the generation of LN metastatic melanoma lines through serial in vivo selection across nine generations. The diagram was created using BioRender. b,c, The transcript levels of Gclc (b) and Fsp1 (c) across LN generations. TPM, transcripts per million. d, Immunoblot of ACSL3, ACSL4, GCLC, FSP1, xCT and GPX4 in B16-F0 (parental), early (LN1) and late (LN7–9) generations of LN metastatic lines. e–l, Individual (e,g,i,k) and grouped (f,h,j,l) quantifications of GCLC (e,f), FSP1 (g,h), GPX4 (i,j) and ACSL4 (k,l) protein expression levels across LN generations. Each replicate represents an independent experiment. For e–l, data are mean ± s.d. Statistical analysis was performed using one-way analysis of variance (ANOVA) followed by Dunnett’s test (e, g, i and k) and two-sided unpaired Student’s t-tests with Welch’s correction (f, h, j and l).
To understand how LN colonization altered the propensity of melanoma cells to undergo cell death, we conducted RNA-sequencing (RNA-seq) analyses of parental and late-generation LN metastatic lines (Extended Data Fig. 1a). Analysis comparing transcript expression of key cell death genes between the late (LN7–9) versus early (parental, LN1 and LN2) generations of LN metastatic lines revealed differences in ferroptosis, necroptosis, autophagy and apoptosis-associated genes. Notably, the most downregulated gene among all of these cell death pathways was Gclc (Extended Data Fig. 1b), which is required for de novo GSH synthesis and can contribute to protection from ferroptosis by replenishing GSH, a co-factor of GPX4, among other actions16,17. By contrast, Fsp1 (also known as Aifm2) was upregulated among ferroptosis-related genes in late-generation LN metastatic lines (Extended Data Fig. 1b).
Gene expression analysis showed a progressive decrease in Gclc (Fig. 1b), and a progressive increase in Fsp1 across LN line generations (Fig. 1c), corroborated by analysis using quantitative PCR (qPCR; Extended Data Fig. 1c). Protein levels demonstrated lower levels of GCLC and higher levels of FSP1 in late generations compared with the parental line (B16-F0) and early generation 1 (LN1-18IL) (Fig. 1d–h). These correlative changes suggest that serial passaging through the LNs confers decreases in Gclc and increases in Fsp1 expression. No significant differences were detected in the mRNA levels of Gpx4, Acsl4, Acsl3 or Slc7a11 (the gene that encodes the functional subunit of system \({{\rm{x}}}_{{\rm{c}}}^{-}\) or xCT) (Extended Data Fig. 1d–h). However, protein levels of GPX4 and ACSL4 were significantly reduced, along with modest reductions in ACSL3 and system \({{\rm{x}}}_{{\rm{c}}}^{-}\), in late-generation lines compared with in parental lines (Fig. 1d,i–l and Extended Data Fig. 1i,j).
Next, to assess the role of the LN environment in the regulation of FSP1, GCLC and GPX4, B16-F10 WT cells were injected either s.c. or i.n. (intranodal) into C57BL/6J mice. The LN environment is characterized by low oxygen availability, with oxygen concentrations ranging from approximately 1 to 3%, and transient reductions to as low as 0.5% (ref. 18). Accordingly, we used HIF-1α as a marker to assess tumour hypoxia and found that i.n. tumours exhibited higher HIF-1α levels compared with s.c. tumours, consistent with lower oxygen availability in the LN environment (Extended Data Fig. 1k–m). Consistent with LN metastatic lines, i.n. tumours exhibited reduced GCLC and GPX4 protein levels compared with s.c. tumours (Extended Data Fig. 1k,m–o), along with a non-significant trend toward increased FSP1 expression in WT cells (Extended Data Fig. 1k,p). A similar reduction in GCLC and GPX4 was observed in B16-F10 Fsp1-knockout (KO) lines injected i.n. (Extended Data Fig. 1l). Notably, both B16-F10 WT and Fsp1-KO cells formed tumours with 100% incidence. These findings indicate that the LN microenvironment induces decreases in GCLC and GPX4 independently of FSP1.
FSP1 contributes to ferroptosis resistance in a GPX4-independent manner by reducing coenzyme Q10 (ubiquinone) to ubiquinol, thereby neutralizing lipid radicals9,10. While its role in metastasis is unclear, FSP1 is overexpressed in several cancers, including melanoma9,10. TCGA data in metastatic melanoma demonstrated a negative correlation between FSP1 and GCLC expression (Extended Data Fig. 1q).
To further investigate this relationship, we analysed FSP1, GCLC and GPX4 protein levels using immunohistochemistry (IHC) in a human melanoma tissue microarray (TMA) comprising primary tumours and LN metastases. While the correlations observed were modest and these IHC analyses do not distinguish between expression tumour cell expression and immune cell expression, FSP1 trended toward a positive correlation with GCLC in primary tumours but trended toward a negative correlation with both GCLC and GPX4 in LN metastases (Extended Data Fig. 1r,s,u). By contrast, GCLC and GPX4 remained positively correlated in both the s.c. and LN contexts (Extended Data Fig. 1t,u). However, these modest correlations should be cautiously interpreted, as total expression levels may not necessarily reflect FSP1 functional activity.
Epigenetic and NRF2 influences on GCLC and FSP1
We next examined potential epigenetic and transcriptional influences on GCLC and FSP1 expression in the LN metastatic lines. Epigenetic dysregulation is a common feature of human cancers, which contributes to tumorigenesis and maintenance of malignant phenotypes19. Previously, it has been described that LN colonization induces significant epigenetic changes in melanoma LN metastatic cells15 (Extended Data Fig. 2a). Assay for transposase-accessible chromatin using sequencing (ATAC–seq) analysis revealed reduced chromatin accessibility at the Gclc transcription start site and the promoter in LN metastatic lines compared with the parental line (Extended Data Fig. 2b). By contrast, no differences in chromatin accessibility at the Fsp1 promoter were observed in LN metastatic lines compared with the parental line, but there was an increase in chromatin accessibility at distant regions (putative enhancers) (Extended Data Fig. 2c). Given the modest changes in chromatin accessibility, we further evaluated potential transcriptional regulation, initially focusing on NRF2 due to its established role in oxidative stress.
NRF2 is a key regulator of the oxidative stress response and controls the expression of several genes involved in ferroptosis regulation, including Gclc, Slc7a11 and Lrp820. NRF2 has also been shown to influence Fsp1 expression in specific contexts21. We therefore next measured the expression of NRF2 and its primary downstream targets in parental versus LN metastatic lines. Although Nrf1, Nrf2 and Keap1 mRNA levels were not significantly different across LN generations (Extended Data Fig. 2d–g), key NRF2-target genes associated with ferroptosis were differentially changed in late versus early LN metastatic lines (Extended Data Fig. 2h). Protein levels of NRF2 were also decreased in the late LN metastatic lines compared with in the parental line (Extended Data Fig. 2i–k). Moreover, NRF2 overexpression in parental cells significantly increased GCLC, GPX4 and FSP1 levels, albeit to a lesser extent (Extended Data Fig. 2l,m).
Together, these results indicate that reduced Gclc expression in LN metastatic cells may result from epigenetic modification at the Gclc locus and reduced NRF2 expression and activity. By contrast, elevated Fsp1 mRNA and reduced GPX4 protein levels in LN metastatic lines are probably driven by NRF2-independent mechanisms, involving epigenetic and post-translational regulation, respectively.
GPX4 dependency in vitro versus in vivo
We next sought to understand how sensitivities to ferroptosis inducers changed across the LN generations. To test this, we evaluated the ferroptosis sensitivity of these lines in vitro and, in this context, LN metastatic lines exhibited greater sensitivity than their parental counterparts to GPX4 inhibitors RSL3 and ML210, as well as to the system xc− inhibitor erastin-2 (Extended Data Fig. 3a–h). Consistently, LN metastatic lines showed elevated lipid oxidation (as detected by BODIPY-C11) after RSL3 treatment in vitro (Extended Data Fig. 3i). By contrast, LN metastatic cells (LN7-1134BL) isolated from spontaneous LN metastases compared with those isolated from the s.c. tumour were less sensitive to erastin-2 or GPX4 inhibition ex vivo (Extended Data Fig. 3j), confirming that the LN-mediated protection from ferroptosis persists in these cell lines, consistent with our previous work2.
Also consistent with our previous findings2, pretreatment of parental cells with albumin-bound oleic acid in vitro fully rescued viability after RSL3 treatment (Extended Data Fig. 3k). However, oleic acid pretreatment only partially rescued cell viability in LN metastatic lines (Extended Data Fig. 3k), which correlated with a trend toward reduced ACSL3 protein levels in some LN metastatic lines and reduced sensitivity under pharmacological inhibition of ACSLs (Extended Data Fig. 3l,m). This model therefore provides a valuable system to investigate the durable cellular adaptations arising during LN colonization, which are distinct yet complementary to the transient metabolic responses mediated by oleic acid protection2.
Reduced GSH in LN metastatic cells
Ferroptosis is a metabolic form of cell death in which GSH has a critical protective role as a key co-substrate required for GPX4 activity22. To investigate whether metabolic changes contribute to the increased sensitivity of LN metastatic lines to GPX4 or system \({{\rm{x}}}_{{\rm{c}}}^{-}\) inhibition in vitro, we performed unbiased metabolomics, revealing distinct clustering between the parental and LN metastatic lines (Fig. 2a). Over 50 metabolites, including several mitochondrial-associated metabolites, were significantly altered in LN metastatic cells (P < 0.001; Fig. 2b). However, we did not observe significant differences between parental and LN metastatic cell lines using the Seahorse assay (Extended Data Fig. 4a). Metabolomic analysis identified significant differences in metabolites involved in GSH synthesis between parental and LN metastatic lines (Fig. 2b,c), including reduced glutamate (Fig. 2d,e), and reduced and oxidized GSH (Fig. 2f–i). These alterations were confirmed by both liquid chromatography coupled with tandem mass spectrometry (LC–MS/MS) and an independent luminescence-based assay (Fig. 2j). Although GCLC expression was reduced in LN metastatic lines (Fig. 1b,d–f), the expression of other GSH synthesis enzymes, such as GCLM and GSS, was not significantly altered (Extended Data Fig. 4b,c).
a, Principal component analysis (PCA) of metabolomic profiles from the B16-F0 (parental) and LN1, 7–9 lines. b, The top 25 differentially altered metabolites in LN7–9 compared with in the LN1 and parental lines. c, Diagram of the GSH-synthesis pathway. The diagram was created using BioRender. d,f,h, LC–MS/MS quantification of glutamate (d), GSH (f) and GSSG (h). e,g,i, Grouped quantification of d, f and h, respectively. j, Luminescence-based GSH quantification with or without cysteine depletion. For d–j, n = 3 independent experiments. For d–j, data are mean ± s.d. Statistical analysis was performed using one-way ANOVA followed by Dunnett’s test (d, f, h and j) and two-sided unpaired Student’s t-tests with Welch’s correction (e, g and i). GSSG, glutathione disulfide.
Given the cysteine requirement for de novo GSH synthesis through GCLC, we tested GSH levels under cysteine-depleted conditions. Depletion of L-cysteine reduced GSH levels in both the parental and LN metastatic lines, with a greater reduction observed in LN metastatic cells (Fig. 2j), indicating impaired cysteine-dependent GSH synthesis. Integration of transcriptomic and metabolomic profiles further confirmed that GCLC expression and GSH metabolism were among the most differentially regulated genes and pathways in LN metastatic cells compared with in the parental cells (Extended Data Fig. 4d,e). These findings suggest that reduced GCLC expression in LN metastatic lines may contribute to impaired GSH synthesis.
Oxygen modulates GPX4 protein levels
The lymph and LN microenvironment contains several ferroptosis-modulating factors, including low free iron, elevated oleic acid and reduced oxygen levels (1–3%)2,18. We next examined whether oleic acid, oxygen or GSH levels modulate GCLC, GPX4 and FSP1 expression in parental melanoma cells to assess their contributions to ferroptosis resistance. Oleic acid supplementation did not alter GPX4, GCLC or FSP1 protein levels under standard culture conditions (21% O2) (Extended Data Fig. 5a). Notably, exposure to 1% O2 levels reduced GPX4 protein levels, independent of oleic acid (Extended Data Fig. 5a), suggesting that oxygen may contribute to the decreased GPX4 expression observed in the LN metastatic lines.
Given that GSH levels are elevated in lymph relative to plasma2, we next tested whether exogenous GSH (GSH-ethyl ester, GSHee) could recapitulate the protein expression patterns observed in LN metastatic lines. GSHee increased GPX4 levels under 21% O2 levels but did not alter GCLC or FSP1 expression (Extended Data Fig. 5b,c). Under 1% O2 levels, GPX4 was also reduced but partially rescued by GSHee (Extended Data Fig. 5b,c). However, GCLC overexpression did not restore GPX4 levels in parental and LN metastatic lines under 21% or 1% O2 levels (Extended Data Fig. 5d–g), and GCLC inhibition through L-BSO further decreased GPX4 only under hypoxia, with no effect under 21% O2 (Extended Data Fig. 5h,i). Similarly to the GCLC overexpression, Gclc-KO lines did not exhibit reduced GPX4 under 1% O2 compared to Gclc-WT lines (Extended Data Fig. 5j,k).
A time-course experiment at 1% O2 confirmed that GPX4 protein levels decreased progressively over time under 1% O2 (Fig. 3a,b), which was rapidly reversed after reoxygenation (Fig. 3c,d), Similarly, CoCl2 treatment—a chemical inducer of hypoxia—led to similar effects, including HIF-1α stabilization and reduction of GPX4 protein levels (Extended Data Fig. 5l). Decreases in GPX4 protein levels under lower oxygen availability was observed in both parental and LN metastatic cells (Extended Data Fig. 5m,n) and in multiple mouse and human melanoma lines (Extended Data Fig. 5o). Moreover, 5% O2 levels also reduced GPX4 protein levels in parental and LN metastatic lines compared with 21% O2 (Extended Data Fig. 5p–s). Together, these findings reinforce our previous observations, highlighting oxygen availability as a critical regulator of the GPX4 surveillance axis in melanoma.
a, Immunoblot of HIF-1α and GPX4 in B16-F0 under 1% O2 for 16, 24 and 48 h. b, Quantification of GPX4 from a. c, Immunoblot of HIF-1α and GPX4 after reoxygenation. B16-F0 cells were cultured under 1% O2 for 24 h, then re-exposed to 21% O2 for 2, 4 or 8 h. d, Quantification from c. e, Confocal microscopy analysis of GPX4 (green) and mitochondria (MitoView; magenta) under 21% and 1% O2 for 24 h. Scale bars, 50 μm. f, Subcellular fractionation of GPX4 under 21% and 1% O2 for 24 h. g,i, GPX4 protein levels in B16-F0 and LN7-1134BL cells treated with proteasome inhibitors (BTZ (10 nM) (g) or MG-132 (0.5 μM) (i)) under 21% or 1% O2 for 24 h. h,j, Quantification of the experiments in g and i, respectively. k, Gpx4 mRNA levels with or without BTZ (10 nM) under 21% or 1% O2 for 24 h. l, GPX4 immunoprecipitation and ubiquitination in the LN7-1134BL line under 21% or 1% O2 for 16 h. LE, long exposure; SE, short exposure. m, The cell viability of B16-F0 and LN metastatic lines treated with ML-210 under 21% or 1% O2 for 48 h. n, The total GSH levels in parental and LN metastatic lines under 21% or 1% O2 for 24 h. n = 3 (a–e, g–l and n) and n = 2 (f) independent experiments. For m, n = 3 technical replicates, representative of 1 of 3 independent experiments. For b, d, h, j, k and n, data are mean ± s.d. Statistical analysis was performed using Kruskal–Wallis tests followed by Dunn’s multiple-comparison test (b, d, h and j) and one-way ANOVA followed by Tukey’s (k) or Šidák’s (m and n) post hoc test.
Low oxygen promotes GPX4 degradation
Although GPX4 is mainly cytoplasmic, it has been reported in the nucleus, mitochondria and at additional subcellular sites23,24. Confocal immunofluorescence analyses and subcellular fractionation revealed that, under 1% O2, GPX4 protein levels were reduced primarily in the cytoplasm while nuclear and mitochondrial GPX4 levels were maintained (Fig. 3e,f). To understand the underlying molecular mechanism by which oxygen availability may regulate GPX4 cytoplasmic levels in melanoma, we first examined the involvement of known GPX4 regulators, including antioxidants, selenium25, autophagic GPX4 degradation26,27 and ubiquitin-proteosome system (UPS)-mediated degradation of GPX428,29,30. Treatment with antioxidants (N-acetyl cysteine) and the ferroptosis inhibitor liproxstatin-1 did not rescue GPX4 protein levels under 1% O2 conditions (Extended Data Fig. 6a,b), whereas selenium supplementation and inhibition of autophagy by chloroquine (CQ) partially rescued GPX4 levels in these conditions (Extended Data Fig. 6c–e). By contrast, the proteosome inhibitors bortezomib (BTZ) and MG-132 significantly rescued GPX4 protein levels in the parental (B16-F0) and LN metastatic lines (LN7-1134BL) under low O2 levels (Fig. 3g–j). Similarly, inhibition of the ubiquitin-activating enzyme (E1) using NSC 624206 rescued GPX4 protein levels under 1% O2, therefore implicating UPS-mediated degradation of GPX4 (Extended Data Fig. 6f,g). To exclude transcriptional effects, we measured Gpx4 and Nrf2 mRNA levels and observed no significant changes after 1% O2 or BTZ treatment (Fig. 3k and Extended Data Fig. 6h). To confirm that GPX4 is ubiquitinated and regulated by UPS under lower O2 levels, we evaluated the ubiquitination of GPX4 under 21% and 1% O2. Exposure to 1% O2 induced HIF-1α and reduced GPX4 protein levels; however, immunoprecipitation of endogenous GPX4 revealed increased ubiquitination specifically under 1% but not under 21% O2 (Fig. 3l and Extended Data Fig. 6i). Together, these results demonstrate that oxygen availability regulates GPX4 protein stability through UPS-mediated degradation, contributing to the reduced GPX4 levels observed in LN metastatic melanoma cells.
Oxygen modulates ferroptosis sensitivity
Given that LN metastatic lines are more sensitive to RSL3, ML-210 and erastin-2 in standard culture conditions (21% O2) compared with the parental line in vitro (Extended Data Fig. 3a–h) and that GPX4 protein levels are reduced by low oxygen availability, we next examined whether oxygen availability modulates the sensitivity to GPX4 and system \({{\rm{x}}}_{{\rm{c}}}^{-}\) inhibitors. Under 1% O2, both parental and LN metastatic lines exhibited reduced sensitivity to GPX4 inhibition (ML-210 or RSL3) and system \({{\rm{x}}}_{{\rm{c}}}^{-}\) inhibition (erastin-2) (Fig. 3m and Extended Data Fig. 6j,k). Moreover, GSH levels were reduced under 1% O2 in both the parental lines and the LN metastatic lines (Fig. 3n). These findings highlight the importance of considering LN microenvironmental factors, such as oxygen availability, that significantly modulate the GCLC–GSH–GPX4 axis and melanoma cell susceptibility to ferroptosis. Given the observed upregulation of FSP1 in LN metastatic lines, we next investigated its regulation and therapeutic potential in melanoma cells within the LNs.
Lysosomal FSP1 in LN metastatic lines
FSP1 has an important protective role neutralizing lipid oxidation9,10. However, the regulation and the pathophysiological contexts in which FSP1 may be targeted in cancer are still unclear. FSP1 undergoes N-myristylation and membrane localization for its anti-ferroptotic action9,10; we therefore first characterized the localization of FSP1 in the LN metastatic lines. FSP1 was significantly located perinuclearly in LN metastatic cells compared to the parental line (Fig. 4a,b). Protein N-myristoylation is a fatty acylation catalysed by N-myristoyltransferases (NMTs). The myristoyl group added to the protein is crucial for cellular localization and signal transduction31. FSP1 perinuclear association was reduced by IMP-1088, a potent N-myristoyltransferase inhibitor, supporting the relevance of FSP1 myristoylation in the association with perinuclear endomembranes in LN metastatic lines (Fig. 4a,b).
a, Confocal images of FSP1–OFP (green) in B16-F0 (parental), LN7-1134BL, LN8-1194BR and LN9-1315BL cells with or without IMP-1088 (0.1 μM) for 24 h. Nuclei (N; DAPI; blue) are outlined with a dotted line. b, Quantification of perinuclear FSP1–OFP in the B16-F0, LN7, LN8 and LN9 lines. B16-F0: n = 194 (−IMP-1088), n = 193 (+IMP-1088); LN7-1134BL: n = 187 (−IMP-1088), n = 221 (+IMP-1088); LN8-1194BR: n = 211 (−IMP-1088), n = 193 (+IMP-1088); LN9-1315BL: n = 148 (−IMP-1088), n = 162 (+IMP-1088). c, Co-localization of FSP1–OFP (green) with lysosomal (LAMP1; magenta), Golgi (RCAS1; red) or ER (ERp72; red) markers. Fluorescence intensity profiles were measured along the arrows shown in the images. Nuclei (DAPI; blue) are outlined with a dotted line. d, Orthogonal immunofluorescence view of FSP1–OFP (green) and LAMP1 (magenta) stack images. Nuclei (DAPI; blue) are outlined with a dotted line. e, Immunoblot of lysosome-enriched and whole-cell extracts. LAMP1, LAMP2 and LIMPII were used as lysosomal markers and γ-tubulin served as a marker of whole-cell extract. f, Quantification of FSP1 protein levels in whole-cell (n = 9), lysosomal-enriched (n = 5), Golgi-enriched (n = 4) and ER-enriched (n = 4) extracts. g, Quantification of FSP1–OFP (green) and LAMP1 (magenta) co-localization in B16-F0, LN7-1134BL, LN8-1194BR and LN9-1315BL cells with or without IMP-1088 (0.1 μM) for 24 h. B16-F0: n = 105 (−IMP-1088), n = 68 (+IMP-1088); LN7-1134BL: n = 104 (−IMP-1088), n = 101 (+IMP-1088); LN8-1194BR: n = 94 (−IMP-1088), n = 76 (+IMP-1088); LN9-1315BL: n = 63 (−IMP-1088), n = 64 (+IMP-1088). h, Confocal microscopy of FSP1–OFP (green) and Lysotracker (magenta) in LN8-1194BR with or without IMP-1088 (0.1 μM) for 24 h. Nuclei (Hoechst; blue) are outlined with a dotted line. i, Quantification of FSP1–OFP and Lysotracker co-localization from h. n = 92 (−IMP-1088) and n = 94 (+IMP-1088). j, Confocal images of LN7-1134BL cells expressing FSP1–OFP WT or G2A mutant, co-stained with LAMP1 (magenta) and DAPI (blue). n = 3 (a–d, g and j) and 4 (h and i) independent experiments. For g and i, data are shown as violin plots with all points; the median is indicated by a dashed line. For b and f, data are mean ± s.d. Scale bars, 50 μm (a), 10 μm (c) and 5 μm (d, h and j). Statistical analysis was performed using one-way ANOVA with Šidák’s multiple-comparison test (b), Tukey’s test (g), Kruskal–Wallis with Dunn’s post-test (f) or two-sided unpaired Student’s t-tests with Welch’s correction (i).
FSP1 has been shown to localize at the plasma membrane, lipid droplets, perinuclear structures and mitochondria9,10. In the LN metastatic cells, perinuclear FSP1 did not co-localize with lipid droplets or mitochondria (Extended Data Fig. 7a). Given that FSP1 was located with the perinuclear region of LN metastatic lines, we further examined its association with perinuclear organelles, including endoplasmic reticulum (ER), Golgi and lysosomes. Immunofluorescence analysis revealed that FSP1 localized primarily with perinuclear lysosomes in the LN metastatic lines (Fig. 4c). Immunofluorescence stack analysis of FSP1 and LAMP1 confirmed that FSP1 is distributed throughout the entire lysosome compartment (Fig. 4d). Biochemical fractionation and enrichment of lysosomes, Golgi and ER confirmed a significant increase in lysosomal FSP1 in the LN metastatic lines compared with in the parental cell lines, whereas FSP1 levels in the Golgi and ER were comparable between the parental and LN metastatic lines (Fig. 4e,f and Extended Data Fig. 7b,c).
FSP1 co-localized with perinuclear lysosomes under both 21% (Fig. 4g–i and Extended Data Fig. 7d,e) and 1% O2 (Extended Data Fig. 7f) conditions. NMT1 and NMT2 activity were required for the FSP1 association with lysosomes (Fig. 4g–i and Extended Data Fig. 7d–f). Indeed, the FSP1 G2A mutant, which cannot undergo N-myristoylation, confirmed that N-myristoylation is essential for lysosomal localization of FSP1 in LN metastatic lines (Fig. 4j and Extended Data Fig. 7g). FSP1 association with lysosomes was also observed in SK-MEL5 and MeWo cells—two human metastatic melanoma lines isolated from axillary LNs32—as well as in A-375 cells (isolated from primary tumour) (Extended Data Fig. 7h). Thus, FSP1 association with perinuclear lysosomes is conserved in mouse and human (Extended Data Fig. 7i) and is generalizable to contexts beyond the LN environment, as observed in the A-375 cell line.
To investigate whether lysosomal activity influences FSP1 protein levels, we used bafilomycin A (BafA1) and CQ to inhibit lysosomal activity. FSP1 protein levels remained unchanged after BafA1 treatment in the parental line and the LN metastatic lines (Extended Data Fig. 8a–d). Similar results were observed with CQ treatment (Extended Data Fig. 8e,f). Together, these data suggest that the lysosomal association of FSP1 does not contribute to its degradation.
Lysosomes have recently been recognized as an organelle involved in the regulation and execution of ferroptosis33,34. Recent evidence indicates that exposure to hypoxic conditions leads to increased lysosomal pH35. Notably, FSP1 enzymatic activity remains largely unaffected by pH fluctuations in a concentration-dependent manner, maintaining its functionality even under the acidic environment typical of lysosomes or other organelles (Extended Data Fig. 8g,h). A recent study developed fentomycins—a synthetic family of small molecules34. Fentomycin is composed of a lysosome-targeting moiety conjugated to an iron-activating ligand, enabling activation of lysosomal iron and induction of ferroptosis. Indeed, sublethal doses of fentomycin-1 increased FSP1 mRNA levels in HT-1080 cells34. Given the lysosomal localization of FSP1 in the LN metastatic cells, we hypothesized that the absence of FSP1 would sensitize lysosomes to lipid oxidation induced by fentomycins. Indeed, LN7-1134BL Fsp1-KO cells (Extended Data Fig. 8i,j) showed increased lipid oxidation after fentomycin-3 treatment compared with wild-type cells (Extended Data Fig. 8k). These data suggest that the LN environment not only induces FSP1 expression, but also functionally engages FSP1 at lysosomal membranes to suppress lipid oxidation under ferroptotic stress.
FSP1 and GCLC inhibition impairs viability in vitro
To determine whether FSP1 is a targetable vulnerability in LN lines, we first compared ferroptosis resistance in WT and Fsp1-KO parental and LN metastatic lines under RSL3 or ML-210 treatment at 21% and 1% O2. As expected, Fsp1-KO lines were more sensitive to RSL3 treatment under 21% O2 (Extended Data Fig. 9a–d). Under 1% O2, although the overall sensitivity to GPX4 inhibition was reduced, Fsp1-KO lines compared with WT lines exhibited significantly diminished protection (Fig. 5a,b and Extended Data Fig. 9b). Notably, RSL3 protection was decreased across LN generations in Fsp1-KO lines, with LN9 Fsp1-KO cells exhibiting the highest sensitivity under 1% O2 (Fig. 5b). A similar trend was observed with ML-210 treatment, in which only Fsp1-KO LN metastatic lines remained sensitive under 1% O2 (Extended Data Fig. 9e,f).
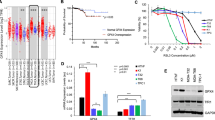
a,b, Cell viability of B16-F0 and LN7-1134BL WT and Fsp1-KO lines (a) or LN9-1315BL WT and Fsp1-KO lines (b) treated with RSL3 under 1% O2 for 48 h. c, Lipid oxidation levels (BODIPY-C11ox/red) of B16-F0 WT, B16-F0 Fsp1-KO, LN7-1134BL WT and LN7-1134BL Fsp1-KO lines under 21% (n = 4) or 1% O2 (n = 4) with or without liproxstatin-1 (1 μM) (n = 3) for 24 h. d, Cell viability of the B16-F0 and LN8-1194BR lines treated with viFSP1 (30 μM), BSO (1 mM), liproxstatin-1 (1μM) or combinations of which under 1% O2 for 48 h. e,f, Cell viability of MeWo (e) and SK-MEL5 (f) cells treated with FSP1 inhibitors (iFSP1 and FSEN1, 10 μM; icFSP1 and viFSP1, 15 μM) with or without BSO (100 μM) and with or without liproxstatin-1 (1 μM) under 21% (e and f) or 1% O2 (f) for 24 h. g, The end-point SK-MEL5 tumour volume after intratumoural treatment with vehicle (n = 19), icFSP1 (n = 8), viFSP1 (n = 10), BSO (n = 14), or combinations of BSO with icFSP1 (n = 8) or viFSP1 (n = 10). Data are normalized to the vehicle treatment. h, End-point SK-MEL5 tumour volumes in mice treated intratumourally with vehicle (n = 9) or FSEN1 (n = 8) normalized to the vehicle treatment. i, End-point tumour volumes comparing i.n. versus s.c. injections of LN7-1134BL WT and Fsp1-KO cells treated with vehicle or viFSP1. i.n.: WT: vehicle (n = 7), viFSP1 (n = 10); Fsp1 KO: vehicle (n = 9), viFSP1 (n = 8). s.c.: WT: vehicle (n = 7), viFSP1 (n = 8); Fsp1 KO: vehicle (n = 8), viFSP1 (n = 5). j, Regulation of GCLC, GPX4 and FSP1 in melanoma within LNs. The diagram was created using BioRender. For a, b and d–f, n = 3 technical replicates, representative of 1 of 3 independent experiments. For g–i, n = 2 independent experiments. For a–c, data are mean ± s.d. For g–i, the box and whisker plots show all points, with the whiskers showing the minimum to maximum values. The box limits represent the first and third quartiles (Q1–Q3), and the centre line indicates the median (Q2). Statistical significance was determined using Kruskal–Wallis tests followed by Dunn’s post hoc test (c), one-way ANOVA followed Tukey’s multiple-comparison test (d) with Šidák’s multiple-comparison test (e–g and i) and two-sided unpaired Student’s t-tests with Welch’s correction (h).
To confirm the relevance of FSP1 under lower O2 availability contexts, we measured lipid oxidation in both WT and Fsp1-KO cells from parental and LN7 lines under both 21% and 1% O2 conditions. Exposure to 1% O2 increased BODIPY-C11 staining in all lines (Fig. 5c), but to an even greater extent in the LN7-1134BL Fsp1-KO line (Fig. 5c), therefore emphasizing the functional importance of FSP1 under reduced O2 availability.
Several small-molecule inhibitors of FSP1 have recently been developed, including iFSP110, FSEN136 and icFSP137, that primarily target human FSP1 through different mechanisms. viFSP1 is the first cross-species inhibitor for FSP1 effective against mouse and human FSP138. viFSP1 in combination with lower doses of RSL3 significantly reduced the viability of LN metastatic lines, but not in the parental line under 1% O2 levels (Extended Data Fig. 9g). However, FSP1 inhibition alone was insufficient to trigger ferroptosis in vitro8, including in LN metastatic lines (Extended Data Fig. 9h).
Although GCLC expression and GSH levels are reduced in LN metastatic lines, they are not entirely absent, suggesting a targetable opportunity in combination with FSP1 inhibition. GCLC has gained attention as a ferroptosis target17,39. However, the contexts in which GCLC inhibition is effective remain unclear. To assess the impact of dual targeting, we tested genetic and pharmacologic inhibition of FSP1 combined with pharmacologic GCLC inhibition using L-BSO. This combination significantly decreased LN8-1194BR viability under 1% O2 compared to the parental cells (Fig. 5d). Fsp1 KO in parental B16-F0 cells had no effect on viability after treatment with L-BSO, whereas Fsp1-KO in LN7-1134BL led to reduced viability under 1% O2, highlighting a selective FSP1 dependency (Extended Data Fig. 9i).
Similar to mouse LN metastatic lines, human melanoma lines derived from LNs, including MeWo and SK-MEL532, exhibited increased sensitivity to L-BSO and FSP1 inhibitors (iFSP1, FSEN1, icFSP1, viFSP1) compared with the primary tumour-derived A-375 line (Fig. 5e,f and Extended Data Fig. 9j,k). Notably, SK-MEL5 was sensitive to combined FSP1 and GCLC inhibition under both 21% and 1% O2—an effect rescued by liproxstatin-1 (Fig. 5f). These in vitro findings support co-targeting FSP1 and GCLC in vivo to reduce tumour growth in LNs.
FSP1 inhibition reduces LN tumour growth
To examine the pharmacological relevance of targeting of FSP1 and/or GCLC in melanoma LN tumours, SK-MEL5 cells were injected i.n. into the popliteal LN of NSG mice. Once palpable, mice were treated daily with intratumoural administration of vehicle, L-BSO, icFSP1 or icFSP1 + L-BSO for 14 days (Extended Data Fig. 10a). Notably, treatment with BSO, icFSP1 or their combination did not result in a significant reduction in tumour size (Fig. 5g and Extended Data Fig. 10c–f). By contrast, viFSP1 monotherapy significantly reduced i.n. tumour burden (Fig. 5g and Extended Data Fig. 10c–f). However, co-treatment with BSO and viFSP1 did not produce an enhanced effect compared with viFSP1 alone in vivo, despite the combined treatment showing greater efficacy in vitro. These results further highlight the context-dependent differences in FSP1 targetability, therefore emphasizing the importance of evaluating the effects of FSP1 inhibitors both in vitro and in vivo.
To further validate the specificity and therapeutic potential of FSP1 inhibition in vivo, we tested FSEN1, a chemically distinct and human-specific FSP1 inhibitor36,40. FSEN1 was administered using the same formulation and dosing regimen as viFSP1 (Extended Data Fig. 10a). Notably, FSEN1 also significantly reduced i.n. tumour growth (Fig. 5h and Extended Data Fig. 10g,h) and extended overall survival in NSG mice (Extended Data Fig. 10i). These findings further support the in vivo efficacy of pharmacological FSP1 inhibition, particularly within the LN microenvironment, using structurally distinct compounds.
To examine potential off-target effects, we used the syngeneic models with LN7-1134BL Fsp1 WT and KO lines injected i.n. or s.c. into immunocompetent C57BL/6J mice (Extended Data Fig. 10b). Daily local administration of viFSP1 significantly reduced i.n. tumour growth in WT tumours at both the experimental end point (and at timepoints matched to the end point of the s.c. experiments), while no response was observed in Fsp1-KO tumours (Fig. 5i and Extended Data Fig. 10j–m), confirming that viFSP1 activity is on-target. Fsp1-KO tumours exhibited slower growth and, in some cases, regression, suggesting that genetic deletion of Fsp1 sensitizes LN-derived cells in the LNs (Fig. 5i and Extended Data Fig. 10j–m).
To assess whether this dependency is specific to the LN microenvironment, we compared s.c. tumour growth in the same model. In this context, viFSP1 treatment resulted in only a modest reduction in tumour growth in WT tumours and had no effect in Fsp1-KO tumours (Fig. 5i and Extended Data Fig. 10n–q). Indeed, no reduction in the tumour growth was observed in the Fsp1-KO tumours treated or not with viFSP1 (Fig. 5i and Extended Data Fig. 10n–q). These findings indicate that FSP1 dependency is markedly enhanced within the LN environment compared to s.c. tumours.
Given the relevance of FSP1 in LN tumours, we evaluated the contribution of FSP1 to metastatic colonization. Experimental metastasis through intravenous injection of LN7 Fsp1-WT or -KO cells resulted in a modest increase in lung colonization in mice injected with the Fsp1-KO lines (Extended Data Fig. 10r), indicating that the loss of FSP1 does not decrease overall survival of metastasizing melanoma cells in the bloodstream. However, in spontaneous models of metastasis, although there were no differences in primary tumour growth of mice implanted s.c. with LN7 Fsp1-WT or -KO cells (Extended Data Fig. 10s; consistent with Fig. 5i), mice bearing LN7 Fsp1-KO compared with WT tumours had a significantly decreased incidence of tumour-draining LN metastasis (Extended Data Fig. 10t,u). These findings suggest that melanoma cells in the LNs represent a context in which targeting FSP1 holds potential for limiting metastatic progression.
Discussion
Here we identify a vulnerability of LN metastatic melanoma cells to pharmacological inhibition of FSP1 as a monotherapy in vivo. In LN metastatic lines, we show that GCLC and GSH are reduced (Figs. 1 and 2) and GPX4 undergoes oxygen-dependent ubiquitin–proteasome-mediated degradation (Fig. 3). We show that the release of melanoma cell dependence on the GPX4 surveillance axis in LNs leads to increased functional reliance on FSP1, which remains intact and accumulates with perinuclear lysosomes through N-myristoylation (Fig. 4).
Notably, FSP1 inhibition is insufficient to reduce the viability of LN lines in vitro (Fig. 5). By contrast, both pharmacological inhibition and genetic deletion of FSP1 significantly impair melanoma growth within LNs, but not in s.c. sites (Fig. 5), emphasizing FSP1 dependencies of cancer cells that arise in vivo. Consistent with this finding, a complementary study demonstrates that FSP1 inhibition in vivo, but not in vitro, significantly reduced lung cancer survival, further indicating that the dependency of FSP1 inhibition differs between the in vitro and in vivo contexts41.
Our findings identify a distinct, durable adaptation that emerges during LN colonization: a shift from GPX4 to FSP1 dependence. The extent to which intratumoural heterogeneity influences this shift in FSP1 dependency remains to be determined. Additional limitations of this work include understanding why FSP1 small-molecule inhibitors have increased efficacy in vivo compared with in vitro. One possibility is that the low oxygen availability, high oleic acid levels in the lymph environment and/or lower GSH levels in the LN metastatic cells drive increased dependency on FSP1 beyond what can be achieved in vitro. Another possibility is that the in vivo environment allows favourable pharmacokinetics of the FSP1 small-molecule inhibitors. The differential efficacies between icFSP1 and viFSP1/FSEN1 may reflect compartmentalized mechanisms of action of these small molecules in vivo, which could account for differences in efficacy; this remains an area of active investigation. Furthermore, FSP1’s association with lysosomes raises numerous yet-to-be explored questions regarding the mechanisms underlying this location, including how FSP1 may protect lysosomes from lipid oxidation.
Our findings indicate that there are contexts that offer promise for FSP1 targetability in circumstances in which GPX4 is endogenously downregulated. Furthermore, our findings hold considerable opportunity for understanding and therapeutically targeting the nuanced physiological context-dependency of ferroptosis42, which has implications for pathophysiological disease states that extend beyond the scope of cancer, such as neurodegeneration and ischaemia–reperfusion injury characterized by heightened ferroptosis vulnerabilities1. Regulation of FSP1 activity and changes in FSP1 subcellular distribution emerge as promising targets to sensitize melanoma cells in LNs to ferroptosis, thereby reducing cancer progression.
Methods
Cell lines
B16-F0 (ATCC; CRL-6322) and its LN metastatic derivatives: NBF0-LN1-18IL, NBF0-LN7-1112AR, NBF0-LN7-1120BL, NBF0-LN7-1134BL, NBF0-LN8-1194BR, NBF0-LN8-1198AR, NBF0-LN8-1205BL, NBF0-LN9-1315BL and NBF0-LN9-1358IR—were provided by the Reticker-Flynn Laboratory. For simplicity, these cell lines are referred to throughout the manuscript as: B16-F0, LN1-18IL, LN7-1112AR, LN7-1120BL, LN7-1134BL, LN8-1194BR, LN8-1198AR, LN8-1205BL, LN9-1315BL and LN9-1358IR, respectively. B16F10 wild-type (WT), B16F10 Fsp1-KO and B16F10 Gpx4-KO cells were obtained from the Conrad Laboratory. B16-F0 Fsp1-KO, LN7-1134BL Fsp1-KO, LN9-1315BL Fsp1-KO, B16-F0 Gclc-overexpression, LN7-1134BL Gclc-overexpression, B16-F0 Gclc-KO and B16-F0 Nrf2-overexpression lines were generated in this study. Human melanoma cell lines MeWo, SK-MEL-5, A375, murine melanoma lines Yale University Melanoma Model (YUMM) 3.3 and YUMM 5.2, and HEK293T cells were purchased from ATCC. All cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, 11885076) supplemented with 10% FBS (Thermo Fisher Scientific, 26400044) and 1% penicillin–streptomycin (Thermo Fisher Scientific, 15140122). All of the other lines were authenticated by ATCC using STR profiling. Cells were routinely tested for mycoplasma contamination using MycoStrip (InvivoGen, rep-mys-50).
Chemicals
RSL3 (HY-100218A), erastin-2 (HY-139087), iFSP1 (HY-136057), BTZ (HY-10227) and PEG300 (HY-Y0873) were purchased from MedChemExpress. ML-210 (S0788), MG-132 (S2619) and icFSP1 (E1535) were acquired from Selleck Chemicals. Rotenone (R8875), oligomycin (75351), antimycin A (A8674), L-BSO (B2515), N-acetyl cysteine (A9165), Na2SeO3 (S5261), CQ (C6628) and PEG400 (202398) were obtained from Sigma-Aldrich. FCCP (15218), MTT (21795), GSHee (14953), liproxstatin-1 (17730), IMP-1088 (25366), NSC 624206 (20569), FSEN1 (38025), viFSP1 (39927) and triacsin C (10007448) were obtained from Cayman Chemical Company. MitoView Fix 640 (70082) and LipidSpot 488 (70065) were sourced from Biotium. Lipofectamine 3000 (L3000015), Bodipy 581/591 C11 (D3861), SYTOX Green (S7020), Lysotracker Deep Red (L12492) and NucBlue Live ReadyProbes Reagent (R37605) were from Thermo Fisher Scientific.
Plasmids
pCMV3-FSP1-OFP plasmid (MG52065-ACR) was obtained from Sino Biological. Lenti-luciferase-P2A-neo (Addgene, 105621), psPAX2 (Addgene, 12260), pMD2.G (Addgene, 12259) and PX458 (Addgene, 48138) were obtained from Addgene. Custom constructs including pTWIST-mFSP1-G2A-OFP, pLVX-EF1α-GCLC-IRES-Hygro, and pLVX-EF1α-NRF2-IRES-Hygro were synthesized by Twist Bioscience and cloned into expression vectors using Gibson Assembly.
Generation of stable cell lines
Stable cell lines expressing luciferase, GCLC or NRF2 were generated through lentiviral transduction followed by antibiotic selection. Lentivirus was produced by co-transfecting HEK293T cells with 5 µg of either Lenti-luciferase-P2A-neo, pLVX-EF1α-GCLC-IRES-Hygro or pLVX-EF1α-NRF2-IRES-Hygro, combined with 5 µg psPAX2 and 0.5 µg pMD2.G using Lipofectamine 3000. Virus-containing supernatants were collected every 24 h for 48 h, filtered and supplemented with 8 µg ml−1 Polybrene (Sigma-Aldrich, H9268). Target cells were infected and subsequently selected with either 1,500 µg ml−1 G418 or 1,000 µg ml−1 hygromycin B for 6 days to establish stable populations.
CRISPR–Cas9-mediated gene KO
To generate Fsp1- or Gclc-KO cell lines in B16-F0 and its LN metastatic derivatives, sgRNAs were designed with BbsI-compatible overhangs and cloned into the PX458 Cas9-GFP vector. The sgRNA sequences were as follows: Fsp1 (CACCGGCGGCTGCCAGCCAGCTGC) and Gclc (CACCGGGGAGTTACATGATCGA). sgRNA insertion was confirmed by whole-plasmid sequencing. Cells were transfected with PX458-sgRNA constructs using Lipofectamine 3000 and GFP-positive cells were sorted by flow cytometry and expanded. Transfection and cell sorting was repeated a second time to generate a pure population for expansion prior to validation. KOs were validated by western blotting and Sanger sequencing (Extended Data Fig. 8i,j for FSP1 and Extended Data Fig. 5j for GCLC).
LN9-1315BL Fsp1-KO cell lines were generated by lentiviral transduction using the LCv2_Blast vector containing mouse Fsp1 sgRNA 1 (sequence: CACCGCCGTGCACGTGGTGATCGT), previously validated43. Transduced cells were selected with 5 µg ml−1 blasticidin. KO validation is shown in Extended Data Fig. 9c.
Western blot analysis
Cell lysates (15–20 μg protein) were separated by SDS–PAGE, transferred onto PVDF membranes (Bio-Rad, 1620177), blocked with 5% non-fat milk in TBS-T or PBS-T, and incubated with primary antibodies overnight at 4 °C in 5% non-fat milk in PBS-T. After washes, the membranes were incubated with HRP-conjugated secondary antibodies and proteins detected by enhanced chemiluminescence (Thermo Fisher Scientific, 32106). The following antibodies were used: ACSL3 (Abcam, ab151959, 1056272-1, WB,1:5,000, Ms), ACSL4 (Santa Cruz Biotechnology, A-5, I1222, WB,1:200, Ms), actin (MP Biomedical, 691001, 0101008716, WB, 1:20,000, Ms and Hu), FSP1 (Proteintech, 20886-1-AP, 00111298, WB,1:2,000, KD validated in-house, Ms and Hu), anti-mouse IgG HRP (Cell Signaling, 7076S, 36, WB, 1:5,000), anti-rabbit IgG HRP (Cell Signaling, 7074S, 33, WB, 1:5,000), COX IV (Cell Signaling, 4850, 11, WB, 1:1000, Ms), GAPDH (Santa Cruz Biotechnology, 6C5, J2523, WB, 1:20,000, Ms), GCLC (Santa Cruz Biotechnology, H-5, J0621, WB, 1:2,000, KO validated in-house), GPX4 (Abcam, ab125066, lot 1000287-43, WB, 1:2,000, KO validated in-house), HIF-1α (Cell Signaling, 36169, 5, WB, 1:1,000), LAMP1 (Abcam, ab24170, GR3235630-1, WB, 1:1,000, Ms), LAMP2A (Abcam, ab18528, 1029399-1, WB, 1:1,000, Ms), LC3 (Cell Signaling, 3868, 14, WB, 1:1,000), LIMPII (Proteintech, 27102-1-AP, WB), NRF2 (Proteintech, 16396-1-AP, 00116728, WB, 1:5,000), NRF2 (Proteintech, 80593-1-RR, 23013625, WB, 1:1,000), PDIA3 (AMAB90988, WB, 1:200), RCAS1 (Cell Signaling, 12290S, D2B6N, 6, WB, 1:1,000), SCL7a11/xCT (Cell Signaling, 98051, 1, WB, 1:300), ubiquitin (Cell Signaling, 43124T, 4, WB, 1:1,000), γ-tubulin (Cell Signaling, T5326, WB, 1:1,000).
Immunoprecipitation and ubiquitination detection
B16-F0 and LN7 1134BL cells were incubated under normoxic (21% O2) or hypoxic conditions (1% O2) for 16 h. Proteins were extracted with RIPA buffer plus protease and phosphatase inhibitors. For denatured immunoprecipitation, lysates were heated to 95 °C for 5 min. Both native and denatured lysates were incubated with anti-GPX4 antibody (Proteintech, 67763-1-Ig, 10027815) or mouse IgG control (Proteintech, B900620) overnight at 4 °C, followed by incubation with anti-mouse IgG Sepharose beads (Cell Signaling, 5946) for 6 h at 4 °C. Beads were washed with RIPA buffer and analysed by immunoblotting using the anti-ubiquitin antibodies (Cell Signaling, 43124T, 4, WB, 1:1,000).
IHC analysis
A TMA containing primary cutaneous melanoma and LN metastases (ME551; TissueArray.com) was used to assess the expression of GCLC, GPX4 and FSP1. The sections were stained with antibodies against GPX4 (Abcam, ab125066, 1:500), GCLC (Santa Cruz, sc-390811, 1:500) and FSP1 (Proteintech, 68049-1-Ig, 1:500) using the Zytomed Permanent AP Red Kit (ZUC001-125) according to the manufacturer’s instructions, followed by counterstaining with haematoxylin. The slides were scanned with an Axio Scan.Z1 slide scanner (Zeiss). Quantification of AP Red signal intensity was performed using QuPath (v.0.5) with uniform thresholding parameters across all samples.
FSP1 enzyme activity
NADH consumption assays were performed in PBS (Gibco, 14190094) containing 15 or 25 nM recombinant non-myristoylated human FSP1, 100 μM menadione (Sigma-Aldrich, M5625) and 200 μM NADH43. The pH of the final reaction was adjusted from 4.0 to 9.0 by titrating PBS with HCl or NaOH. After the addition of FSP1, the absorbance at 340 nm was recorded every 20 s at 37 °C using the SpectraMax M5 microplate reader (Molecular Devices). Reactions lacking NADH or enzyme were included for background correction. Data were normalized and fitted using GraphPad Prism 10.
Confocal fluorescence microscopy
Cells plated on coverslips were transfected with FSP1-OFP using Lipofectamine 3000. After 16 h, cells were treated with IMP-1088 (0.1 μM) for 24 h. Cells were fixed (4% paraformaldehyde), permeabilized (0.1% Triton X-100), and incubated overnight with primary antibodies in 3% BSA/PBS and then with by Alexa-Fluor-conjugated secondary antibodies. For live-cell imaging, cells were plated on 30-mm glass-bottom dishes, transfected as described above, and incubated with Lysotracker (50 nM) and NucBlue Live ReadyProbes reagent during the final 30 min of IMP-1088 treatment. Images were captured with a Nikon Eclipse Ti confocal microscope using consistent settings for comparisons and analysed with Fiji software. Antibodies and stains used included Alexa Fluor 647 donkey anti-rat (Thermo Fisher Scientific, A48272, YK388772, IF, 1:500), Alexa Fluor 488 goat anti-rabbit (Thermo Fisher Scientific, A32731, YI374177, IF, 1:500), Alexa Fluor 546 goat anti-rabbit (Thermo Fisher Scientific, A11010, 2570547, IF, 1:500), ERp72 (Cell Signaling, 5033, 4, IF, 1:200) GPX4 (Abcam, ab125066, 1000287-7, IF, 1:100, KO validated in-house) from Abcam; LAMP1 (Thermo Fisher Scientific, 14-1071-82, 2698949, IF, 1:50), RCAS1 (Cell Signaling, 12290, 6, IF, 1:200), MitoView Fix640 (70082-50 μg, 23M0201-1215003) and LipidSpot 488 (70065, 22L0820) from Biotium.
Lipid oxidation assays
Cells (60,000 per well) were seeded in 12-well plates one day before treatment. Cells were treated with 0.5 µM RSL3 for 4 h or 1% O2 for 24 h, washed with PBS, trypsinized and resuspended in PBS containing 1.5 µM C11-BODIPY 581/591 (Invitrogen, D3861). After 30 min incubation at 37 °C, cells were washed, incubated with DAPI, filtered through a 70-µm strainer and analysed on the BD LSR Fortessa flow cytometer. Excitation was performed at 488 nm, detecting oxidized BODIPY (FITC, 525/40 nm) and reduced BODIPY (PE, 585/42 nm). At least 10,000 events were analysed per sample. Data were processed using FlowJo software, and the lipid oxidation ratio (FITC/PE ratio) was calculated as (median FITC-A − median FITC-A unstained)/(median PE-A − median PE-A unstained). The flow cytometry gating strategies for the lipid oxidation assays are presented in Supplementary Fig. 2.
Cell viability and cell death assays
Cells (2,500–3,000 per well) were seeded into 96-well plates. Viability was measured using MTT assay 24 h (erastin-2) or 48 h (RSL3, ML-210, viFSP1 + BSO and Triacsin C) after treatment. Cell death was monitored every 3 h using SYTOX Green (25 nM) in the Incucyte S3 (Sartorius) system.
Isolation of lysosome-enriched fractions
Lysosome-enriched fractions were isolated using the Lysosome Isolation Kit (Abcam, ab234047) according to the manufacturer’s protocol. In brief, 2 × 107 cells were washed and centrifuged at 600g for 10 min and the supernatant was removed. Cells were resuspended in Lysosome Isolation Buffer, vortexed and incubated on ice for 2 min. Complete cell disruption was obtained using a dounce homogenizer. After adding Lysosome Enrichment Buffer, the homogenate was centrifuged at 500g for 10 min at 4 °C. The supernatant was added to the top of a discontinuous gradient density and an ultracentrifugation at 145,000g for 2 h at 4 °C was performed. The lysosome-enriched fraction was present in the top 10% of the gradient volume. For western blot analyses, the protein content of the lysosomal-enriched gradient supernatant was quantified using the Qbit 1 fluorometer (Thermo Fisher Scientific) and a protein quantification kit (Thermo Fisher Scientific, Q33212). Equal total protein amounts of total cell extracts and lysosome-enriched extracts were loaded for comparison for western blot analyses.
Isolation of Golgi-enriched fractions
Golgi-enriched fractions were isolated using the Golgi enrichment extraction kit (Invent, GO-037) according to the manufacturer’s instructions. In brief, filter cartridges were placed and cooled on ice for several minutes. Then, 2 × 107 cells were trypsinized and collected by centrifugation at 500g, washed with 1× PBS and centrifuged again at 500g. The pellet was resuspended in buffer A with vigorous shaking. The filter cartridge was capped, the tube inverted several times and centrifuged at 16,000g for 30 s. The tube was then centrifuged at 4 °C at 5,000g for 5 min without removing the filter. The filter was then removed and the supernatant transferred to a fresh tube and centrifuged at 4 °C at 16,000g for 30 min. The supernatant was then transferred to a fresh tube. An equivalent in volume of buffer B was added to the supernatant, the resulting mixture incubated on ice for 15 min and then centrifuged at 8,000g for 5 min. The pellet was then resuspended in buffer A and mixed by pipetting up and down 50 times and subsequently centrifuged at 8,000g for 5 min. The supernatant was then transferred to a fresh tube and ice old buffer C was added, mixed by vortexing for 20 s and incubated on ice for 20 min. The tube was then centrifuged at 8,000g for 10 min and the supernatant removed. The pellet was resuspended Laemmli buffer for subsequent western blot analysis. For western blot analyses, the protein content of the Golgi-enriched extracts was quantified using the Qbit 1 fluorometer (Thermo Fisher Scientific) and a protein quantification kit (Thermo Fisher Scientific, Q33212). Equal total protein amounts of total cell extracts and lysosome-enriched extracts were loaded for comparison for western blot analyses.
Isolation of ER-enriched fraction
ER were isolated using the ER enrichment extraction kit (Novus Biologicals, NBP2-29482) according to the manufacturer’s instructions. In brief, 500 µl of 1 × isosmotic homogenization buffer followed by 5 µl of 100× PIC were added to a pellet of 2 × 107 cells. The resulting suspension was centrifuged at 1,000g for 10 min at 4 °C. The supernatant was transferred to a clean centrifuge tube and centrifuged at 12,000g for 15 min at 4 °C. The floating lipid layer was discarded. The supernatant was centrifuged in a clean centrifuge tube using an ultracentrifuge at 90,000g for 1 h. The resulting pellet contained the total ER fraction (rough and smooth). The pellet was resuspended Laemmli buffer for subsequent western blot analysis. For western blot analyses, the protein content of the ER-enriched extracts was quantified using the Qbit 1 fluorometer (Thermo Fisher Scientific) and a protein quantification kit (thermo Fisher Scientific, Q33212). Equal total protein amounts of total cell extracts and lysosome-enriched extracts were loaded for comparison for western blot analyses.
Mitochondrial/cytoplasmic fractionation
Mitochondrial and cytoplasmic fractions were obtained using a mitochondria isolation kit for mammalian cells (89874) from Thermo Fisher Scientific according to the manufacturer’s instructions.
RNA-seq analyses
RNA-seq data were generated and analysed as described previously15. Raw sequencing reads were trimmed and quality-filtered using Trimmomatic and FastQC, respectively. Transcript abundance was quantified with Salmon v.0.7.2 using quasi-mapping mode and corrected for sequence, GC and positional biases, using the mouse genome GRCm38 GENCODE release M11. TPM values were computed using tximport and renormalized after removing mitochondrial transcripts. Differential expression analysis was performed using DESeq2 with regularized log-transformed counts. Hierarchical clustering and PCA analyses used Spearman correlations from the top 1,000 highly variable genes. Heat maps (Extended Data Fig. 1a) were generated using heatmap3 from the top 200 differentially expressed genes. Data have been deposited in the Gene Expression Omnibus (GEO: GSE117529).
ATAC–seq analyses
ATAC–seq analyses were conducted as described previously15. In brief, cells were permeabilized and DNA was transposed using Tn5 transposase. Libraries were purified, amplified and sequenced (NovaSeq, 2 × 100 cycles, around 50 million paired reads per sample). Reads were mapped to mm10 (hisat2), duplicates removed (Picard) and peaks were called using MACS2. Normalized coverage was visualized in IGV. Transcription factor activity and motif enrichment were assessed with Chromvar and HOMER, respectively. Data were deposited at the GEO (GSE117529).
RNA isolation and qPCR analyses
RNA was extracted using the RNeasy Plus Mini Kit (Qiagen, 74134), and cDNA was synthesized using the iScript Reverse Transcription Supermix (Bio-Rad, 1708841). qPCR was performed using the iTaq Universal SYBR Green Supermix (Bio-Rad, 1725121) on the BioRad CFX96 system. The primers used were as follows: mNRF2_F: AACGACAGAAACCTCCATCTAC; mNRF2_R: AGTAAGGCTTTCCATCCTCATC; mFSP1_F: GCAATGAGTATCGGGAGTACAT; mFSP1_R: GTAGGCAGAGCTGTTGATCTT; mGPX4_F: ACTGACGTAAACTACACTCAGC; mGPX4_R: GGAAGGCCAGGATTCGTAAA; RNA pol II_F: ACTGTGCGGAACTCCATCAA; RNA pol II_R: AGCCAGGTTCTGGAACTCAA; mPPIB_F: CATCAAGGACTTCATGATCCA; mPPIB_R: ATAGATGCTCTTTCCTCCTGTG. RNA pol II and PPIB amplification were used as reference genes. PPIB was used as a housekeeping gene for qPCR analyses of parental and LN metastatic lines, while RNA Pol II was used for qPCR analyses of BTZ treatment under 21% and 1% O2 conditions.
Metabolite extraction and LC–MS analysis
For metabolite extraction, 5 × 105 cells were seeded into 6-well plates and cultured for 24 h. The medium was then aspirated, and cells were washed with cold normal saline (9 g l−1 sodium chloride). Immediately, 400 µl of extraction buffer (methanol:acetonitrile:water, 40:40:20, with 0.5% formic acid) was added per well, and the plates were incubated on ice for 5–10 min. The samples were neutralized with 35 µl of 15% ammonium bicarbonate (NH4HCO3), cells were scraped and lysates were transferred to 1.5 ml tubes and centrifuged at 16,000 rpm for 15 min. A total of 80 µl of supernatant was transferred to LC–MS vials, and 20 µl from each sample was pooled to generate a quality control sample. All of the extracts were stored at –80 °C until analysis.
Metabolites were analysed using a Q Exactive HF mass spectrometer (Thermo Fisher Scientific) coupled to hydrophilic interaction chromatography (HILIC). Separation was performed using an XBridge BEH Amide XP column (2.5 µm, 2.1 × 150 mm) with a guard column (2.5 µm, 2.1 × 5 mm; Waters). Mobile phase A consisted of water:acetonitrile (95:5) and mobile phase B comprised water:acetonitrile (20:80), both containing 10 mM ammonium acetate and 10 mM ammonium hydroxide. The gradient was as follows: 0–3 min, 100% B; 3.2–6.2 min, 90% B; 6.5–10.5 min, 80% B; 10.7–13.5 min, 70% B; 13.7–16 min, 45% B; 16.5–22 min, 100% B. The flow rate was 0.3 ml min−1. The autosampler was maintained at 4 °C and the column at 30 °C. The injection volume was 5 µl. Needle washes were performed between injections using acetonitrile:methanol:water (4:4:2, v/v/v).
MS1 scans were acquired from m/z 70 to 1,000 with polarity switching and a resolution of 120,000 (at m/z 200). Other MS parameters were as follows: sheath gas, 40; auxiliary gas, 10; sweep gas, 2; spray voltage, 3.5 kV; capillary temperature, 300 °C; S-lens RF level, 45; maximum injection time, 500 ms; AGC target, 3 × 106.
Raw data were converted to mzXML format using msConvert and analysed in El-Maven (Elucidata) for targeted metabolite identification based on accurate mass and retention time, using an in-house standard library. Data were normalized to protein content and analysed in MetaboAnalyst 6.0 (https://www.metaboanalyst.ca).
GSH measurements
Cells (5,000 per well) were seeded into 96-well plates, and GSH levels were assessed using the GSH/GSSG-Glo assay (Promega, V6611). Parallel cell viability assessments were used for data normalization.
Seahorse assay
Cells (5,000 per well) were seeded in 96-well plates and analysed using the Seahorse XF24 system. Oxygen consumption rates were measured sequentially after oligomycin (1 μM), FCCP (1 μM) and rotenone/antimycin A (0.5 μM each). Data were normalized to protein content.
s.c. and i.n. tumour models
Mice were housed under sterile conditions with sterilized standard chow and water provided ad libitum and maintained under a 12 h–12 h light–dark cycle and 22 ± 2 °C, 55 ± 5% humidity. Animals were allocated randomly to treatment groups, and the samples were processed in an arbitrary order. No formal randomization or blinding was applied. The maximum permitted tumour diameter of 2.0 cm was not exceeded in any of the experiments. All procedures complied with institutional ethical guidelines and were approved by the Institutional Animal Care and Use Committee of the Harvard T.H. Chan School of Public Health (protocol IS00003460) or the Stanford University Institutional Animal Care and Use Committee (protocol APLAC-34518).
For s.c. injections, 2 × 105 B16-F10 WT Luc, B16-F10 Fsp1-KO Luc, or LN7 1134BL WT or Fsp1-KO cells were suspended in 100 µl of DMEM without phenol red and injected into either the right or left flank of 6–8-week-old male or female C57BL/6J or C57BL/6N mice44.
For i.n. injections, 1 × 104 SK-MEL5 or LN7 1134BL WT or Fsp1-KO cells were injected into the popliteal LN of 6–8-week-old NSG or C57BL/6J mice. To visualize the lymphatics, 2% Evans Blue dye (Sigma-Aldrich, E2129) was injected into the footpad 5 min before the procedure. Mice were injected with buprenorphine and anesthetized with isoflurane, and a 5–10 mm incision was made in the region of the right popliteal LN. The node was identified by Evans Blue staining, immobilized with forceps and 1 × 104 cells in 10 µl of 1× PBS were injected into the LN using a 27 G Hamilton syringe. Successful injection was confirmed by visible swelling of the node. Incisions were closed with surgical glue (VetBond Tissue Adhesive, 3M, 1469SB) and the mice were monitored for signs of pain or distress for 5 days45.
Once tumours were palpable in ≥50% of mice (around 1 week after injection), 10 µl of vehicle or drug solution was administered daily through intratumoural (i.n. or s.c.) injection into tumour-bearing sites. Treatment groups included: L-BSO (1 mM; Thermo Fisher Scientific, 235520050), icFSP1 (0.025 mg (2.5 mg ml−1); Selleckchem, E1535), L-BSO + icFSP1 (1 mM + 0.025 mg (2.5 mg ml−1)), viFSP1 (0.025 mg (2.5 mg ml−1); MedChemExpress, HY-163002), L-BSO + viFSP1 (1 mM + 0.025 mg (2.5 mg ml−1)) and FSEN1 (0.025 mg (2.5 mg ml−1); MedChemExpress, HY-153629). L-BSO was dissolved in 0.9% sodium chloride (saline; Quality Biology, 114-055-101). icFSP1 was formulated in 55% PBS (Corning, VWR45000-430) and 45% PEG300 (MedChemExpress, HY-Y0873). viFSP1 and FSEN1 were formulated in 20% DMA, 40% PEG400 and 40% of 50% 2-hydroxypropyl-β-cyclodextrin (2HPβCD) in water.
Tumour diameters were measured daily using callipers until any tumour reached around 1.5 cm in its largest dimension, which defined the experimental end point. At the end point, all of the mice in the cohort were euthanized in accordance with approved protocols. Tumour diameters and weights were recorded, and tissues were collected and frozen for downstream analyses.
Experimental lung metastasis was evaluated through intravenous delivery of cancer cells in the lateral tail vein of tumour-naive mice. A total of 2 × 106 LN7-1134BL WT or Fsp1-KO cells was resuspended in 200 µl of DMEM without phenol red and injected into the lateral tail vein of 8-week-old female C57BL/6N mice using a 27-gauge needle44. Mice were euthanized 14 days after injection, and the lungs were inflated with PBS using a 25-gauge needle inserted into the trachea, and the lungs were removed for visible counting of metastatic nodules identified by melanin.
For LN spontaneous metastasis assays, 2 × 105 LN7 1134BL WT or Fsp1-KO cells were suspended in 100 µl DMEM (without phenol red) and injected s.c. into the right or left flank of 6–8-week-old male or female C57BL/6J or C57BL/6N mice. Mice were euthanized 24 days after injection and the draining LNs were collected and classified as metastatic (LN+) or non-metastatic (LN−) based on the presence of melanin-containing melanoma cells44.
Bioinformatics analysis
Correlation analyses used tools available online (https://hgserver1.amc.nl/). Metabolomic data were analysed using MetaboAnalyst 6.0 (https://www.metaboanalyst.ca/).
Joint pathway analysis transcriptomic and metabolomic datasets showing significant alterations (P < 0.05, |Fold Change| > 1) between parental (B16-F0) and LN (LN8) clones underwent joint pathway enrichment analysis using MetaboAnalyst. Parameters included integrated metabolic pathways, hypergeometric test, degree centrality topology and pathway-level P-value combination. Pathways were considered significant at P < 0.05 and impact > 0.2 (normalized degree centrality), with at least two significantly altered metabolites.
Correlation analysis gene–metabolite correlations were calculated using the cor.test function (R stats package v.3.6.2). Analysis focused on highly interconnected genes and metabolites within the KEGG glutathione metabolism pathway modules (glutathione biosynthesis and ferroptosis protection), obtained using the MetaboSignal package (v.1.32.1) and the cluster_walktrap algorithm from the igraph package (v.2.0.2). Only late LN tumour generations were included due to sample size limitations.
Bayesian inference of directed acyclic graphs (DAGs) was used to identify cause–effect networks among genes and metabolites across tumour generations (early: B16-F0, F018IL; late: LN7, LN8, LN9). DAG networks were inferred using the BiDAG package (v.2.1.4) with Bayesian Gaussian equivalent scoring and order Markov Chain Monte Carlo structure learning. Networks were averaged over 100 iterations to account for inference variability, assigning edge probabilities based on inference frequency.
Software for Illustrations
Illustrations were generated using FIJI (2.0.0-rc-69/1.52n), Prism (10.5.0) and BioRender (http://biorender.com). Figures created using BioRender include Figs. 1a, 2c and 5j and Extended Data Figs. 7i and 10a,b.
Statistical analysis
Data are presented as mean ± s.d. Statistical analyses were performed using GraphPad Prism v.10.5.0 (GraphPad Software) and included unpaired two-sided Student’s t-tests with Welch’s correction, one-way ANOVA with Dunnett’s, Tukey’s or Šidák’s multiple-comparisons tests, Kruskal–Wallis tests followed by Dunn’s post hoc test, log-rank (Mantel–Cox) tests for survival analyses and contingency analysis using χ2 with Fisher’s exact test. P < 0.05 was considered to be statistically significant. Sample sizes (n) refer to biological or technical replicates as defined in individual figure legends. Numbers independent biological replications are indicated in the figure legends, with the exception of Fig. 1, for which replicates are noted here: for Fig 1e,g, B16-F0 (n = 30), LN1-18IL (n = 30), LN7-1112AR (n = 9), LN7-1120BL (n = 9), LN7-1134BL (n = 9), LN8-1194BR (n = 12), LN8-1198AR (n = 12), LN8-1205BL (n = 12), LN9-1315BL (n = 6), LN9-1358IR (n = 6); Fig. 1f,h, parental (n = 30), LN (n = 75); Fig. 1i, B16-F0 (n = 7), LN1-18IL (n = 7), LN7-1112AR (n = 4), LN7-1120BL (n = 3), LN7-1134BL (n = 4), LN8-1194BR (n = 3), LN8-1198AR (n = 3), LN8-1205BL (n = 3), LN9-1315BL (n = 7), LN9-1358IR (n = 7); Fig. 1j, parental (n = 7), LN (n = 34); (k) B16-F0 (n = 15), LN1-18IL (n = 15), LN7-1112AR (n = 6), LN7-1120BL (n = 6), LN7-1134BL (n = 6), LN8-1194BR (n = 6), LN8-1198AR (n = 6), LN8-1205BL (n = 6), LN9-1315BL (n = 6), LN9-1358IR (n = 6); Fig. 1l, parental (n = 15), LN (n = 48).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Metabolomic data are presented in Supplementary Table 1. RNA-seq and ATAC–seq raw data have been deposited at the GEO (GSE117529). Source data and uncropped blot scans are provided in Supplementary Fig. 1. Source data are provided with this paper.
References
Stockwell, B. R. Ferroptosis turns 10: emerging mechanisms, physiological functions, and therapeutic applications. Cell 185, 2401–2421 (2022).
Ubellacker, J. M. et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature 585, 113–118 (2020).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Jiang, X., Stockwell, B. R. & Conrad, M. Ferroptosis: mechanisms, biology and role in disease. Nat. Rev. Mol. Cell Biol. 22, 266–282 (2021).
Seiler, A. et al. Glutathione peroxidase 4 senses and translates oxidative stress into 12/15-lipoxygenase-dependent and AIF-mediated cell death. Cell Metab. 8, 237–248 (2008).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Conrad, M. & Pratt, D. A. The chemical basis of ferroptosis. Nat. Chem. Biol. 15, 1137–1147 (2019).
Nakamura, T. & Conrad, M. Exploiting ferroptosis vulnerabilities in cancer. Nat. Cell Biol. 26, 1407–1419 (2024).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 575, 688–692 (2019).
Doll, S. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019).
Garcia-Bermudez, J. et al. Squalene accumulation in cholesterol auxotrophic lymphomas prevents oxidative cell death. Nature 567, 118–122 (2019).
Liang, D. et al. Ferroptosis surveillance independent of GPX4 and differentially regulated by sex hormones. Cell 186, 2748–2764 (2023).
Le Gal, K. et al. Antioxidants can increase melanoma metastasis in mice. Sci. Transl. Med. 7, 308re8 (2015).
Piskounova, E. et al. Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527, 186–191 (2015).
Reticker-Flynn, N. E. et al. Lymph node colonization induces tumor-immune tolerance to promote distant metastasis. Cell 185, 1924–1942 (2022).
Asantewaa, G. et al. Glutathione synthesis in the mouse liver supports lipid abundance through NRF2 repression. Nat. Commun. 15, 6152 (2024).
Kang, Y. P. et al. Non-canonical glutamate-cysteine ligase activity protects against ferroptosis. Cell Metab. 33, 174–189 (2021).
Schito, L. & Rey, S. Hypoxia orchestrates the lymphovascular-immune ensemble in cancer. Trends Cancer 8, 771–784 (2022).
Portela, A. & Esteller, M. Epigenetic modifications and human disease. Nat. Biotechnol. 28, 1057–1068 (2010).
Takahashi, N. et al. 3D culture models with CRISPR screens reveal hyperactive NRF2 as a prerequisite for spheroid formation via regulation of proliferation and ferroptosis. Mol. Cell 80, 828–844 (2020).
Koppula, P. et al. A targetable CoQ-FSP1 axis drives ferroptosis- and radiation-resistance in KEAP1 inactive lung cancers. Nat. Commun. 13, 2206 (2022).
Zheng, J. & Conrad, M. The metabolic underpinnings of ferroptosis. Cell Metab. 32, 920–937 (2020).
Pfeifer, H. et al. Identification of a specific sperm nuclei selenoenzyme necessary for protamine thiol cross-linking during sperm maturation. FASEB J. 15, 1236–1238 (2001).
Moreno, S. G. et al. Testis-specific expression of the nuclear form of phospholipid hydroperoxide glutathione peroxidase (PHGPx). Biol. Chem. 384, 635–643 (2003).
Ingold, I. et al. Selenium utilization by GPX4 is required to prevent hydroperoxide-induced ferroptosis. Cell 172, 409–422 (2018).
Sun, Y. et al. Fin56-induced ferroptosis is supported by autophagy-mediated GPX4 degradation and functions synergistically with mTOR inhibition to kill bladder cancer cells. Cell Death Dis. 12, 1028 (2021).
Xue, Q. et al. Copper-dependent autophagic degradation of GPX4 drives ferroptosis. Autophagy 19, 1982–1996 (2023).
Chu, L. K. et al. Autophagy of OTUD5 destabilizes GPX4 to confer ferroptosis-dependent kidney injury. Nat. Commun. 14, 8393 (2023).
Dong, K. et al. HOIP modulates the stability of GPx4 by linear ubiquitination. Proc. Natl Acad. Sci. USA 119, e2214227119 (2022).
Fan, Y. et al. PRMT5-mediated arginine methylation stabilizes GPX4 to suppress ferroptosis in cancer. Nat. Cell Biol. 27, 641–653 (2025).
Yuan, Y. et al. Protein lipidation in health and disease: molecular basis, physiological function and pathological implication. Signal Transduct. Target. Ther. 9, 60 (2024).
Carey, T. E. et al. Cell surface antigens of human malignant melanoma. Proc. Natl Acad. Sci. USA 73, 3278–3282 (1976).
Saimoto, Y. et al. Lysosomal lipid peroxidation contributes to ferroptosis via lysosomal membrane permeabilization. Nat. Commun. 16, 3554 (2025).
Cañeque, T. et al. Activation of lysosomal iron triggers ferroptosis in cancer. Nature 642, 492–500 (2025).
Sekine, H. et al. PNPO–PLP axis senses prolonged hypoxia in macrophages by regulating lysosomal activity. Nat. Metab. 6, 1108–1127 (2024).
Hendricks, J. M. et al. Identification of structurally diverse FSP1 inhibitors that sensitize cancer cells to ferroptosis. Cell Chem. Biol. 30, 1090–1103 (2023).
Nakamura, T. et al. Phase separation of FSP1 promotes ferroptosis. Nature 619, 371–377 (2023).
Nakamura, T. et al. Integrated chemical and genetic screens unveil FSP1 mechanisms of ferroptosis regulation. Nat. Struct. Mol. Biol. 30, 1806–1815 (2023).
Eaton, J. K. et al. The enzyme glutamate-cysteine ligase (GCL) is a target for ferroptosis induction in cancer. Preprint at bioRxiv https://doi.org/10.1101/2024.04.28.591552 (2024).
Zhang, S. et al. Cocrystal structure reveals the mechanism of FSP1 inhibition by FSEN1. Proc. Natl Acad. Sci. USA 122, e2505197122 (2025).
Wu, K. et al. Targeting FSP1 triggers ferroptosis in lung cancer. Nature https://doi.org/10.1038/s41586-025-09710-8 (2025).
Ubellacker, J.M. & Dixon, S.J. Prospects for ferroptosis therapies in cancer. Nat Cancer 8, 1326–1336 (2025).
Mishima, E. et al. A non-canonical vitamin K cycle is a potent ferroptosis suppressor. Nature 608, 778–783 (2022).
Breuer, C. B. et al. Spontaneous and experimental models of lymph node metastasis. Nat. Protoc. https://doi.org/10.1038/s41596-025-01200-5 (2025).
Sabatier, M. et al. Lymphatic collection and cell isolation from mouse models for multiomic profiling. Nat. Protoc. 20, 884–901 (2025).
Acknowledgements
This work was supported by the Melanoma Research Program Department of Defense Team Science Award (J.M.U. and N.E.R.-F.), the Ludwig Center at Harvard (J.M.U.), the Melanoma Research Foundation Young Career Investigators Award (J.M.U.), R01CA282202 (J.M.U.), NIH T32DK128781 Predoctoral Fellowship (C.S.F. and A.C.), Find the Cause Breast Cancer Foundation (J.M.U.), the METAvivor Early Career Investigator Award (N.E.R.-F.), NIH DP2 AI177915 (N.E.R.-F.), Arc Institute Fellowship and NCI F31CA301857 (C.B.B.). S.H. acknowledges support from R56DK140139. R.R. is supported by Ligue Contre le Cancer (Equipe Labellisée), Fondation Charles Defforey-Institut de France, Fondation Bettencourt Schueller, Institut National du Cancer, Agence Nationale de la Recherche and the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant agreement no. 647973). M. Conrad is supported by DFG Priority Program SPP 2306 (CO 291/9-1, 461385412; CO 291/10-1, 461507177; CO 291/9-2, CO 291/10-2, CO 291/14-1) and the CRC TRR 353 (CO 291/11-1; 471011418), and the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement no. GA 884754). A.T. acknowledges the support of an Emmy Noether Award from the German Research Foundation (DFG, 467788900), the Ministry of Culture and Science of the State of North Rhine-Westphalia (NRW-Nachwuchsgruppenprogramm) and an ERC starting grant (METATARGET, 101078355). A.T. holds the Peter Hans Hofschneider endowed Professorship of Molecular Medicine from the Stiftung Experimentelle Biomedizin. J.P.F.A. acknowledges the EU-H2020 (ERC-Consolidator, DeciFERR) and the support of the Deutsche Forschungsgemeinschaft (DFG) SPP2306.
Author information
Authors and Affiliations
Contributions
M.P., N.E.R.-F. and J.M.U. conceived the study and supervised the research. M.P., C.B.B., N.E.R.-F. and J.M.U. designed the experiments. M.P. performed experiments with support from M. Chaufan and K.J.S.; T.N. and B.P. generated the B16F10 Fsp1-KO cells. C.B.B. generated the B16-F0 and LN7 1134BL Fsp1-KO lines and the Gclc-KO and Gclc-overexpressing lines. M.P. generated the LN9-1315BL Fsp1-KO and Nrf2-overexpression lines. S.M. performed lysosomal, Golgi and ER fractionation and assessed lipid oxidation under fentomycin-3 treatment. T.C. and L.C. synthesized fentomycin-3. A.T., F.C. and L.M.N.M. designed and performed IHC analyses on the TMA. E.M. performed hFSP1 in vitro assays. In vivo studies were conducted by M.P., M. Chaufan, C.B.B., M.S., K.J.S., C.S.F., M.K., M.Y., A.C., M.L., L.Z. and M.F. Data analysis was conducted by M.P., C.S.F., M. Chaufan, S.M., M.K., J.M.-L., Y.L., M.S., M.F., N.C., S.H., S.M., A.T., L.M.N.M., R.R. and J.M.U.; M.P. and J.M.U. wrote the manuscript with input from S.M., J.P.F.A., A.W., M. Conrad, R.R. and N.E.R.-F.
Corresponding author
Ethics declarations
Competing interests
M. Conrad, B.P., T.N., R.R., S.M., T.C. and L.C. are listed as inventors for some of the compounds described here, and M. Conrad and B.P. are co-founders and shareholders of ROSCUE Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Hozumi Motohashi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Expression of ferroptosis-related genes in LN metastatic melanoma lines.
a, Heat map of differentially expressed genes (log2(Fold change) in parental (B16-F0), early (LN1), and late (LN7-9) generations of LN metastatic lines (RNA-seq from ref. 15). b, Volcano plot of cell death–related genes in late (LN7–9) vs. early (Parental, LN1–2) generations. c,d, qPCR of Fsp1 (c) and Gpx4 (d) mRNA levels in B16-F0 (Parental), LN7, LN8, and LN9 lines. e–h, Transcript levels of Gpx4 (e), Acsl4 (f), Acsl3 (g), and Slc7a11/xCT (h) across LN generations (R2 < 0.2 indicate negligible correlations). i,j, Quantification of ACSL3 (i) and xCT (j) immunoblots from Fig. 1d. k,l, Immunoblots of HIF-1α, GCLC, FSP1, and GPX4 in SubQ vs. IN tumours from B16-F10 WT (k) and FSP1 KO (l), (validation of FSP1 KO lines shown in Extended Data Fig. 8i). m–o, Quantification of HIF-1α (m), GCLC (n), and GPX4 (o) from (k–l). n = 9 mice for SubQ tumours; n = 10 mice for IN tumours. p, Quantification of FSP1 from (k). n = 4 mice for SubQ tumours; n = 5 mice for IN tumours. q, Correlation of FSP1 and GCLC mRNA expression in metastatic melanoma (Matta dataset, n = 198, R2 platform). r–t, Correlation of FSP1 and GCLC (r), FSP1 and GPX4 (s), and GCLC and GPX4 (t) protein levels assessed by immunohistochemistry (IHC) of a human melanoma tissue microarray (ME551, TissueArray), including primary tumours (n = 25) and lymph node (LN) metastases (n = 22). u, Representative IHC images from samples analysed in panels r-t. PT: primary tumours. Data in c, d, i, j, m-p show mean ± s.d. n = 3 independent experiments for c, d, i, j. Statistical significance was determined by Kruskal-Wallis followed by Dunn’s post hoc test (i, j); one-way ANOVA followed Dunn’s post hoc test (c, d), or two-sided unpaired Student’s t-test with Welch’s correction (m-p).
Extended Data Fig. 2 Epigenetic and transcriptional regulation of Gclc and Fsp1 in LN metastatic cells.
a, Heat map of chromatin accessibility in B16-F0 (parental) (n = 4) and LN7 (n = 4) lines. b,c, Chromatin accessibility at Gclc (b) and Fsp1 (c) loci; green shading marks peaks at promoter (b) or putative enhancer (c). d–f, Transcript levels of Nrf1 (d), Nrf2 (e), and Keap1 (f) across LN generations. g, qPCR of Nrf2 mRNA levels in B16-F0, LN7, LN8, and LN9 lines. h, Heat map of canonical NRF2 target genes in late (LN7–9) vs. early (Parental, LN1–2) lines. *indicates significantly changed genes. i, Immunoblot of NRF2 expression across parental, LN1, and LN7–9 lines. j,k, Quantification of immunoblots in (i). l, Immunoblot of NRF2, GCLC, FSP1, and GPX4 in B16-F0 WT and NRF2-overexpressing (NRF2 O.E) lines. m, Quantification of immunoblot from (l). n = 3 independent experiments for g, i-m. Data in g, j, k and m shown mean ± s.d. Statistical analysis by one-way ANOVA followed by Dunnett’s test (g, j); two-sided unpaired Student’s t-test for grouped comparisons (k, m).
Extended Data Fig. 3 GPX4 dependency differs in vitro versus in vivo.
a,b, Cell viability of B16-F0 and LN metastatic lines treated with increasing concentrations of RSL3 (a) or 0.5 μM (b) for 48 h. c, Cell death assays using Sytox Green in B16-F0, LN7, and LN9 cells treated with RSL3 (0.5 or 1 μM) for 16 h. d,e, Cell viability of B16-F0 and LN metastatic lines treated with increasing concentrations of ML-210 (d) or 2.5 μM (e) for 48 h. f,g, Cell viability of B16-F0 and LN metastatic lines treated with increasing concentrations of Erastin-2 (f) or 0.5 μM (g) for 24 h. h, Cell death assays in B16-F0, LN8, and LN9 lines treated with Erastin-2 (0.5 or 1 μM) for 16 h. i, Lipid oxidation levels (BODIPY-C11 ox/red) in B16-F0, LN7, LN8, and LN9 lines treated with RSL3 (0.5 μM) for 4 h. j, Ex vivo treatment of LN7 1134BL cells isolated from primary tumours (PT) or spontaneous lymph node (LN) metastases with DMSO, Erastin-2 (1 μM), RSL3 (1 μM), or ML-210 (2.5 μM) for 24 h. k, Cell viability heat map of B16-F0, LN7, LN8, and LN9 lines ± RSL3 (1 μM), ± oleic acid (OA, 125 μM), or combinations for 24 h. l,m, Cell viability of B16-F0 and LN metastatic lines treated with increasing concentrations of Triacsin C (l), or 0.5 μM (m) for 24 h. n = 3 technical replicates, representative of 1 of 3 independent experiments for a-i, k-m; n = 9 (3 technical replicates) from 3 independent experiments for j. Data in a-j and l,m shown mean ± s.d. Statistical analysis by one-way ANOVA followed by Dunnett’s test (b, e, g, m) or Sidak’s multiple comparisons test (c, h, j, k); two-way ANOVA followed by Tukey’s test (i).
Extended Data Fig. 4 Mitochondrial stress profiling with integrated transcriptomic and metabolomic analysis in parental and LN metastatic cells.
a, Mitochondrial stress test (Seahorse) of B16-F0, LN1, and late LN metastatic lines. mean ± s.d; n = 4 technical replicates, representative of 1 of 2 independent experiments. b,c, Transcript levels of Gclm (b) and Gss (c) across LN generations. d, KEGG glutathione metabolism pathway showing fold changes (LN8 vs. B16-F0); red and blue circles indicate up- and downregulated elements. e, Average directed acyclic graphs (DAGs) showing glutathione-gene/metabolite relationships for early (B16-F0, LN1) and late (LN7–9) lines. Beige, metabolites; teal, genes; edge weight = inference strength.
Extended Data Fig. 5 Oxygen regulates GPX4 independently of oleic acid and glutathione.
a, Immunoblot of GCLC, FSP1, GPX4 in B16-F0 ± oleic acid (125 μM) under 21% or 1% O2 for 24 h. b, Immunoblot of HIF-1α, GCLC, FSP1, GPX4 ± GSHee (10 mM) under 21% or 1% O2 for 24 h. c, GPX4 quantification from (b). d,f, Immunoblots of HIF-1α, GCLC, GPX4 in B16-F0 (d) or LN7 1134BL (f) WT and GCLC-OE cells under 21% or 1% O2 for 24 h. e,g, Quantification of GPX4 from (d) and (f). h,i, Immunoblot and quantification of GPX4 in cells ± L-BSO under 21% or 1% O2 for 24 h. j,k, Immunoblots and quantification of GPX4 in B16-F0 WT and GCLC KO cells under 21% or 1% O2 for 24 h. l, Immunoblot of HIF-1α and GPX4 in B16-F0 and LN metastatic lines ± CoCl2 (200 μM) for 24 h. m, Immunoblot of HIF-1α, NRF2, ACSL3, GCLC, and GPX4 in B16-F0 and LN metastatic lines under 21% or 1% O2 for 24 h. n, Immunoblot of GCLC, and GPX4 in B16-F0 and LN metastatic lines under 21% or 1% O2 for 48 h. o, Immunoblots of HIF-1α, and GPX4 from mouse and human melanoma lines under 21% or 1% O2 levels. p-r, Immunoblot of HIF-1α, GCLC, FSP1, and GPX4 from B16-F0 (p), LN7 1134BL (q), and LN9 1315BL (r) under 21% O2 and 5% O2 for 24 h. s, Quantification of GPX4 protein levels from p,q, and r. n = 3 independent experiments for a-s. Data in c, e, g, i, k and s shown mean ± s.d. Statistical analysis by one-way ANOVA with Tukey’s test (c, e, g, i, k) or Sidak’s test (s).
Extended Data Fig. 6 GPX4 is downregulated by UPS under low oxygen levels, independent of antioxidants, selenium, or autophagy.
a-c, Immunoblots of GPX4 from B16-F0 treated with NAC (1 or 5 mM) (a), Liproxstatin-1 (1 μM) (b) or selenium (15 or 50 nM) (c) under 21% or 1 % O2 for 24 h. d, Immunoblot of GPX4 from B16-F0 treated with chloroquine (CQ) (20 μM) under 21% or 1 % O2 for 24 h. e, Quantification from (d). f, Immunoblot of GPX4 from B16-F0 treated with NSC 624206 (10 μM) under 21% or 1 % O2 for 24 h. g, Quantification from (f). h, Nrf2 mRNA levels under 21% and 1% O2 ± BTZ (10 nM). i, GPX4 immunoprecipitation and ubiquitination in B16-F0 under 21% or 1% O2 for 16 h. j,k, Cell viability heatmaps following treatment with RSL3 (j) for 48 h or Erastin-2 (k) for 24 h under 21% or 1% O2. n = 3 independent experiments for a-i; n = 3 technical replicates, representative of 1 of 3 independent experiments for j, k. Data in e, g, and h shown mean ± s.d. Statistical analysis was performed using the Kruskal–Wallis test followed by Dunn’s post hoc test (e, g), or one-way ANOVA followed by Tukey’s test (h) or Sidak’s test (j, k).
Extended Data Fig. 7 FSP1 preferentially localizes to perinuclear lysosomes in LN metastatic lines.
a, Left, representative confocal microscopy of FSP1-OFP (Green) with lipid droplets (LipidSpot: Red), mitochondria (Mitoview: Magenta), and nuclei (Hoechst; Blue) in LN7 and LN8 lines. Right, histogram of the fluorescence intensity profile across the arrow. b,c, Immunoblots of Golgi- (b) and ER-enriched (c) fractions. RCAS1 and PDI3A were used as Golgi and ER markers, respectively and γ-tubulin served as a whole-cell extract control. ER extract (b), n = 4; Golgi extract (c), n = 4 independent experiments. d, Representative confocal microscopy of FSP1-OFP (Green) and LAMP1 (Magenta) ± IMP-1088 (0.1 μM) in B16-F0, LN7, LN8, and LN9 lines for 24 h. e, Representative confocal microscopy of FSP1-OFP (Green) and Lysotracker (Magenta) in B16-F0, LN7, LN8, and LN9 lines. f, Representative confocal microscopy of FSP1-OFP (Green) and Lysotracker (Magenta) localization in LN8 1194BR ± IMP-1088 (0.1 μM) under 1% O2 for 24 h. g, Representative confocal microscopy of FSP1-OFP WT and FSP1 OFP G2A (Green) localization with LAMP1 (Magenta) in LN9 1315BL. h, Representative confocal microscopy of FSP1-OFP (Green) and Lysotracker (Magenta) in SK-MEL5, MeWo and A-375 cells. i, FSP1 subcellular localization in LN metastatic lines. The diagram was created using BioRender. 3 independent experiments for a, d-h. Scale bar 10 μm (a, d, e, f, h) and 5 μm (g).
Extended Data Fig. 8 FSP1 protects lysosomes from lipid oxidation without undergoing lysosomal degradation.
a, c, Immunoblots of FSP1 and LC3I/II ± Bafilomycin A1 (10 nM) in B16-F0 (a) and LN7 (c) under 21% or 1 % O2 for 24 h. b,d, Quantification of FSP1 from (a, c). e,f, Immunoblot and quantification of FSP1 and LC3I/II ± chloroquine (20 μM) under 21% or 1 % O2 for 24 h. g,h, In vitro enzymatic activity of recombinant human FSP1 at 15 nM (g) or 25 nM (h) across a range of pH values. i, Immunoblot analysis of FSP1 protein levels in B16-F0 and LN7 1134BL WT and FSP1 KO lines (top) and B16-F10 WT and FSP1 KO (bottom). j, Analysis of the histogram from the sequencing of the LN7 1134BL WT and LN7 1134BL FSP1 KO lines. In green, the deletion observed in the KO models. The sequence and position of the sgRNA is at the top. k, Lipid oxidation levels (BODIPY-C11 ox/red) from LN7 1134BL WT, and LN7 1134BL FSP1 KO lines ± Fentomycin-3 (7 μM) for 6 h. n = 3 independent experiments for a-f; 1 experiment for g, h; n = 5 independent experiments for k. Data in b, d, f, k shown mean ± s.d. Statistical analysis by Kruskal–Wallis test with Dunn’s multiple comparisons (b, d, f), and one-way ANOVA with Sidak’s test (k).
Extended Data Fig. 9 FSP1 and GCLC inhibition reduces LN cell viability under 21% and 1% O2.
a, Cell viability of B16-F0 and LN7 1134BL WT and FSP1 KO lines treated with RSL3 under 21% O2 for 48 h. b, Cell viability of B16-F0 WT, B16-F0 FSP1 KO, LN7 1134BL WT, and LN7 1134BL FSP1 KO lines ± RSL3 (0.1 μM and 0.25 μM) under 21% or 1% O2 for 48 h. c, Immunoblot of FSP1 protein levels in LN9 1315BL WT and LN9 1315BL FSP1 KO clones. d, Cell viability of LN9 1315BL WT and FSP1 KO lines treated with RSL3 under 21% O2 for 48 h. e, Cell viability of B16-F0 and LN7 1134BL WT and FSP1 KO lines treated with ML-210 under 1% O2 for 48 h. f, Cell viability assay of B16-F0 WT, B16-F0 FSP1 KO, LN7 1134BL WT, and LN7 1134BL FSP1 KO lines ± ML-210 (2.5 μM) under 1% O2 for 48 h. g, Cell viability of B16-F0 and LN7 1134BL cells ± RSL3 (0.25 μM), viFSP1 (15 μM or 30 μM), or combinations under 21% and 1% O2 for 48 h. h, Cell viability of B16-F0 and LN metastatic lines treated with viFSP1 under 21% O2 for 48 h. i, Cell viability of B16-F0, LN7 1134BL WT and FSP1 KO lines ± BSO (1 mM and 3 mM) under 1% O2 for 48 h. j,k, Cell viability of A-375 cells ± iFSP1 (10 μM) (j) or FSEN1 (10 μM) (k) ± BSO (1 mM) or combinations. n = 3 technical replicates for a, b, d-f, h-k or n = 6 technical replicates for g, representative of 1 of 3 independent experiments. Data in a,b, d-k shown mean ± s.d. Statistical significance was determined using one-way ANOVA followed by Sidak’s multiple comparisons test (b, f, g, i–k).
Extended Data Fig. 10 Targeting FSP1 or its genetic deletion reduces intranodal tumour growth and LN metastasis.
a, Schematic of the experimental design for intranodal implantation of SK-MEL5 cells into NSG mice, followed by treatment with FSP1 inhibitors, BSO, or their combinations. The diagram was created using BioRender. b, Schematic of the experimental design for intranodal or subcutaneous implantation of LN7 1134BL WT or FSP1 KO cells into C57BL/6J mice treated with viFSP1. The diagram was created using BioRender. c–h, Representative tumour growth curves from one of the two independent experiments corresponding to Fig. 5g (c–f) and Fig. 5h (g,h). i, Kaplan–Meier survival curves of NSG mice treated intranodally with vehicle (n = 9) or FSEN1 (n = 8). j–q, Representative tumour growth curves from one of the two independent experiments from Fig. 5i showing intranodal (j–m) and subcutaneous (n–q) tumour growth of LN7 1134BL WT and FSP1 KO lines ± viFSP1. r, Quantification of lung metastases (number of nodules) from intravenous injection of LN7 1134BL WT (n = 19) and FSP1 KO cells (n = 21). Data combined 2 independent experiments (Exp1: white dots; Exp2: grey dots). s, Subcutaneous tumour growth curves for LN7 1134BL WT and FSP1 KO cells. Representative curves from 1 of 2 independent experiments. t, Quantification of spontaneous lymph node metastases in draining lymph nodes from 2 independent experiments including subcutaneous LN7 1134BL WT (n = 11) or FSP1 KO tumours (n = 13). u, Representative images of LNs from 1 of 2 independent experiments. Statistical significance was determined using the log-rank (Mantel–Cox) test for survival curve comparisons (i), Mann-Whitney test (r) and contingency analysis using Chi-square with Fisher’s exact test (t).
Supplementary information
Supplementary Figs. 1 and 2 (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Table 1 (download XLSX )
Raw metabolomic data pertaining to Fig. 2.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Palma, M., Chaufan, M., Breuer, C.B. et al. Lymph node environment drives FSP1 targetability in metastasizing melanoma. Nature 649, 477–486 (2026). https://doi.org/10.1038/s41586-025-09709-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09709-1
This article is cited by
-
Single-cell omics in tumor lymph node metastasis: mechanisms and therapeutic implications
Molecular Cancer (2026)
-
Cell death: targeting ferroptosis in cancer
Signal Transduction and Targeted Therapy (2026)
-
Targeting FSP1 triggers ferroptosis in lung cancer
Nature (2026)
-
Triggering ferroptosis to tackle cancer
Nature Reviews Drug Discovery (2026)
-
In vivo models bring FSP1 inhibitors to life
Nature Cell Biology (2026)