Abstract
RNA interference (RNAi) depends on DICER, an essential enzyme that processes RNA precursors into small regulatory RNAs. DICER cleaves RNA precursors according to the 5′-end counting rule, in which RNA length is measured from the 5′-end1,2,3. Previous work proposed a single 5′-end binding pocket that disfavours guanosine (5′-G), leading to cleavage inaccuracies4. Here we show that 5′-G promotes precise cleavage for many substrates. Using massively parallel dicing assays and cryo-electron microscopy, we identify a conserved guanosine-favoured (G-favoured) binding pocket in DICER, distinct from the previously described uridine-favoured (U-favoured) pocket. Together, these pockets influence the alignment between 21-nucleotide and 22-nucleotide cleavage registers, expanding the mechanism of small-RNA biogenesis in metazoan DICERs. We also find that conflicts between 5′-end binding and RNA-motif recognition can trigger RNA conformational adjustments that preserve accurate cleavage-site selection. In addition, conformational adjustments of the double-stranded RNA-binding domain (dsRBD) and PAZ domain help to align substrates with the catalytic centres for precise double-strand cleavage. These results show that the DICER cleavage mechanism integrates dual 5′-end binding pockets, RNA-motif influence and domain motions, advancing our understanding of microRNA biogenesis.
Similar content being viewed by others
Main
DICER is an evolutionarily conserved RNase III enzyme that has a central role in RNA silencing, a crucial gene regulatory mechanism that influences diverse biological processes. DICER processes precursor microRNAs (pre-miRNAs) and long double-stranded RNAs (dsRNAs) into small regulatory RNAs, microRNAs (miRNAs) and small interfering RNAs (siRNAs), which target mRNAs for degradation or translational repression through the RNA-induced silencing complex (RISC)1,2,3. This cleavage activity also drives short-hairpin RNA (shRNA) technology, in which synthetic shRNAs are processed by DICER into siRNA duplexes that mimic endogenous RNAi pathways5,6,7,8,9,10.
Over the past two decades, studies have elucidated key aspects of DICER’s cleavage mechanism, including 5′- and 3′-end counting rules that determine cleavage sites around 21–22 nucleotides (nt) from the RNA ends4,11,12,13,14,15,16,17. Structural studies have shown that the 3′-end binds within a conserved pocket in DICER’s PAZ domain, whereas the 5′-end interacts with a pocket in DICER’s platform domain4,14,17,18,19,20,21,22,23,24,25. In human, mouse and fly DICER enzymes, the 3′-end terminal phosphate interacts with a cluster of conserved tyrosines and basic residues4,17,18,19,21,22. Similarly, plant DICER-like enzymes (for example, Arabidopsis thaliana DCL1 and DCL3) have a conserved 3′-end docking pocket, highlighted by aligned conserved tyrosines, basic residues, and phenylalanine24,25. These findings suggest that the 3′-end binding pocket is conserved across DICER and DICER-like enzymes.
The previous effort to identify the 5′-end binding pocket in DICER used crystal structures of its PAZ–platform–connector helix cassette bound to siRNAs, revealing a phosphate-binding pocket17. However, these structures might not represent the dicing state. A cryo-electron microscopy (cryo-EM) study resolved DICER in the dicing state, detailing a 5′-end binding pocket for 5′-U, involving R790, R821 and R10034. This pocket disfavours guanosine (5′-G), owing to steric clashes with the arginine residue (R821), suggesting that 5′-G reduces cleavage accuracy4. Similarly, a dicing-state structure of fly Dcr-1 identified a 5′-end binding pocket for 5′-U, with R1027 (conserved with R821 in human DICER) located in this pocket21. In A. thaliana DCL3, the 5′-end binding pocket consists of histidine and arginine residues, which are not conserved with human DICER25. By contrast, the 5′-end binding pockets of mouse DICER, fly Dcr-2 and Arabidopsis thaliana DCL1 remain undefined19,22,23,24. These findings suggest that, unlike the conserved 3′-end binding pocket, the 5′-end binding pocket varies among enzymes and is challenging to determine. Moreover, structural studies of human DICER and fly Dcr-1 have relied on RNA substrates with fixed 5′-U, and the effects of diverse RNA ends have not been examined.
DICER uses fine-tuning mechanisms, such as sequence motifs (for example, mWCU, YCR and GYM), bulges, loops and mismatches, to achieve precise cleavage at either position 21 (DC21) or position 22 (DC22) from the 5′-end6,9,26,27,28,29,30,31,32,33,34,35,36,37. However, the interaction between these features and end-binding rules, as well as how DICER coordinates RNA motifs with end-binding preferences for precise cleavage, remain unclear.
In this study, using massively parallel dicing assays and single-particle cryo-EM, we show that 5′-G, contrary to previous reports, enhances cleavage precision at DC21 for many substrates. Cryo-EM reconstructions uncover a previously unrecognized G-favoured binding pocket, distinct from the U-favoured binding pocket, directing cleavage to DC22. This dual-pocket mechanism reshapes our understanding of small-RNA biogenesis, showing how DICER integrates end-binding rules with RNA features for precise cleavage. RNA motifs such as mWCU and YCR cooperate with the 5′-end binding rule to refine specificity, and conflicts induce RNA conformational changes that override end-binding preferences. Cryo-EM further reveals dynamic rearrangements of the dsRBD and PAZ domains, repositioning RNA for precise catalysis and providing a framework for RNA processing in silencing pathways.
DICER cleavage accuracy is enhanced by 5′-G
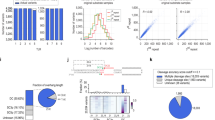
When testing pre-mir-517a, a pre-miRNA with a 5′-C (pre-mir-517a_CU), we observed cleavage at both the DC21 and the DC22 site (Fig. 1a–c). To assess the contribution of the 5′-nucleotide (5′-nt), we replaced the 5′-C with other nucleotides and examined dicing outcomes. Substituting 5′-C with 5′-A (pre-mir-517a_AU) or 5′-G (pre-mir-517a_GU) shifted cleavage predominantly to DC21, with 5′-G showing the highest accuracy, whereas 5′-U (pre-mir-517a_UU), like 5′-C, supported cleavage at both sites (Fig. 1b,c and Extended Data Fig. 1a). Because 5′-G and 5′-A pair with U on the 3′-strand, whereas 5′-C and 5′-U do not, these substitutions altered the overhang geometry. To isolate the effect of the 5′-nt from the overhang context, we tested pre-mir-517a variants with comparable overhangs. Notably, G–U (pre-mir-517a_GU) showed higher DC21 accuracy than U–G, whereas A–U favoured DC21 and U–A favoured DC22 (Fig. 1b,c). These findings refute the hypothesis that 5′-G reduces cleavage accuracy4, and reveal that the 5′-nt strongly influences DICER’s cleavage-site preference: 5′-G and 5′-A promote DC21, whereas 5′-C and 5′-U favour DC22.
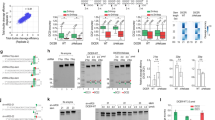
a,Schematic of pre-mir-517a substrates, illustrating terminal pairs (for example, G–U and C–U) at the 5′- and 3′-ends. Green and red arrowheads indicate DC21 and DC22 cleavage sites, located 21 and 22 nt from the 5′-end, producing fragments F1, F2 and F3. b, In vitro dicing assays for pre-mir-517a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21), whereas F2 is undetectable on gels. c, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-517a variants, based on four independent experiments. Statistical significance was determined using a two-tailed, two-sample t-test (**P < 0.01, ****P < 0.0001). d, Schematic of massively parallel dicing assays using pre-mir-324 groups with fixed 5′-nt (A, G, U or C) and randomized 3′-overhang. Cleaved products (F3) were cloned, sequenced and analysed. e, Cleavage patterns from pre-mir-324 groups with distinct 5′-end nucleotides. f, Cleavage sites (DC20–DC23) identified from sequencing results. n = 256 variants. g, Cleavage accuracy at DC21 and DC22 for pre-mir-324 groups, calculated from sequencing data. n = 64 for each comparing group. h, Diagrams of pre-mir-629, pre-mir-208a and variants, showing terminal pairs. Green and red arrowheads indicate DC21 and DC22 cleavage sites. i, In vitro dicing assays for pre-mir-629, pre-mir-208a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21); F2 is undetectable. Green arrowheads, DC21; red arrowheads, DC22. j, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-629 and pre-mir-208a variants, based on three independent experiments. Statistics as in c. k, Mechanistic model illustrating how 5′-nt affects DICER cleavage. Guanine (G) and adenine (A) favour DC21 cleavage, whereas uracil (U) and cytosine (C)—especially U—favour DC22 cleavage.
Parallel dicing assays confirm the effects of the 5′-nt
To generalize these findings, we performed massively parallel dicing assays using pre-mir-324, cleaved by DICER at DC20, DC21 and DC2238,39,40. We synthesized four groups of pre-mir-324 variants, each with a specific 5′-nt and randomized 3′-ends, producing 64 unique sequences per group (Fig. 1d). After DICER cleavage, F3 fragments were sequenced to identify cleavage sites (Fig. 1e and Extended Data Fig. 1b). The assays were highly reproducible, with strong correlation between replicates (Extended Data Fig. 1c,d). Sequencing confirmed that DICER cleaves pre-mir-324 mainly at DC21 and DC22 (Fig. 1f). Whereas the 3′-nt had no detectable effect in our randomized pre-mir-324 context (Extended Data Fig. 1e), the 5′-nt strongly influenced site preference: 5′-G yielded the highest DC21 accuracy, followed by 5′-A, whereas 5′-C and 5′-U favoured DC22 (Fig. 1g). These results confirm that 5′-G enhances, rather than impairs, cleavage accuracy by promoting DC21 cleavage, underscoring the prominent role of the 5′-nt in determining DICER’s cleavage-site preference.
Effects of the 5′-nt validated across pre-miRNAs
To validate the role of the 5′-nt in cleavage specificity, we synthesized pre-mir-324 variants with a fixed 3′-end and varying 5′-nts (Extended Data Fig. 1f). Consistent with the massively parallel assays, 5′-G and 5′-U promote cleavage at DC21 and DC22, respectively, whereas 5′-A and 5′-C support cleavage at both sites, with slight preferences for DC21 and DC22, respectively (Extended Data Fig. 1g,h). To isolate the influence of the 5′-nt, we generated variants with similar 2-nt overhang structures but differing base pairs (A–U versus U–A, or G–U versus U–G), confirming that 5′-G strongly favours DC21, 5′-U favours DC22 and 5′-A promotes DC21 more than 5′-U but less than 5′-G (Extended Data Fig. 1i–k). Finally, variants with similar 3′-nt overhangs but differing 5′-nts showed that 5′-A favoured DC21, whereas 5′-U and 5′-C supported DC22, with 5′-U showing the strongest bias (Extended Data Fig. 1l–n).
To test whether the effects of the 5′-nt apply to other pre-miRNAs, we examined pre-mir-629 and pre-mir-208a, which naturally have 5′-U and 5′-G, respectively (Fig. 1h). In pre-mir-629, DICER cleaves at both DC21 and DC22, with DC21 cleavage driven by a YCR motif. Changing 5′-U to 5′-A reduces DC22 cleavage, confirming that 5′-U enhances DC22 cleavage (Fig. 1i,j). Similarly, pre-mir-208a shows DC21 cleavage with 5′-G and DC22 cleavage with 5′-U (Fig. 1i,j). These findings show that 5′-G promotes DC21, whereas 5′-U promotes DC22 (Fig. 1k).
Cryo-EM shows DICER bound to 5′-G and 5′-U shRNAs
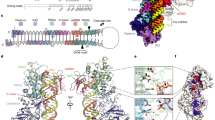
To examine how 5′-G and 5′-U influence DICER cleavage specificity at DC21 and DC22, we solved cryo-EM structures of DICER bound to 5′-G (26S-GU) or 5′-U (26S-UG) shRNAs (Fig. 2a,b). Adding Ca2+ during reconstitution trapped the dicing state by inhibiting catalysis while preserving the cleavage-ready conformation. Cryo-EM maps of the DICER–26S-GU and DICER–26S-UG complexes were obtained at 3.34-Å and 3.37-Å resolution, respectively, as determined by gold-standard Fourier shell correlation (GS-FSC) (Extended Data Fig. 2a–f and Supplementary Table 1). Multiple rounds of two-dimensional (2D) classification selected particles that resembled the dicing-state complex, consistent with previous studies4 (Extended Data Fig. 2a,b). A bias in particle orientation was observed, which is likely to be due to DICER’s asymmetric shape and mass distribution, which might reduce directional resolution and lead to anisotropic reconstructions (Extended Data Fig. 2f). Local-resolution analysis showed slightly higher resolution in protein densities than in RNA densities (Extended Data Fig. 2d). Using the published structural model4 (Protein Data Bank (PDB): 7XW2), we refined atomic structures of DICER in complex with both shRNAs, revealing detailed architecture and domain organization (Extended Data Fig. 2g). The 26S-UG and 26S-GU RNAs included mWCU and YCR motifs, promoting cleavage at DC2235,36, which allowed us to investigate the effects of 5′-nt and RNA motifs on DICER–RNA interactions (Fig. 2c and Extended Data Fig. 3a).
a, Schematic of human DICER domains: DUF283, PAZ, RIIIDa and RIIIDb (RNase III) and dsRBD, with amino acid boundaries labelled. b, Sequences of the 26S-UG and 26S-GU shRNAs that were used in cryo-EM studies. The mWCU and YCR motifs guide DC22 cleavage. c, Cleavage accuracy of DICER for 26S-UG and 26S-GU calculated from three independent assays, showing precise cleavage at DC22. d, Cryo-EM reconstructions of DICER–26S-UG and DICER–26S-GU complexes, with domains colour-coded. e, Structural comparison of DICER complexes reveals conformational shifts in dsRBD and PAZ after RNA binding. RMSD values show significant movement compared with apo-DICER (PDB: 7XW3) and DICER–pre-let-7a-1GYM (PDB: 7XW2). f, Alignment of cryo-EM densities shows that shRNA-bound complexes are more compact (57.7–58.5 Å) than DICER/pre-let-7a-1GYM (68.0 Å). g, PAZ and dsRBD translocate in the DICER–26S-UG and DICER–26S-GU complexes relative to DICER–pre-let-7a-1GYM, with RMSD values of 7.4 Å (PAZ) and around 5.0 Å (dsRBD), aligning the RNA substrate with the catalytic centres. h, PAZ adopts an ‘inner’ conformation in DICER–shRNA complexes, bending the 3′-overhang, unlike the ‘outer’ conformation in DICER–pre-let-7a-1GYM. i, Buried surface area analysis shows increased RNA–DICER interaction in shRNA-bound states, compared with DICER–pre-let-7a-1GYM. j, Proposed model: the PAZ-dsRBD ‘in mode’ aligns the RNA backbone for precise cleavage, whereas the ‘out mode’ can cause RNA misalignment.
The cryo-EM structures reveal the organization of DICER’s key domains, including the platform, PAZ, RIIIDa, RIIIDb, dsRBD, connector helix and bound RNA (Fig. 2d). Partial densities for helicase domains, probably Hel1, are also observed (Extended Data Fig. 3b). However, as in previous studies4,22, incomplete densities are seen for helicase residues (1–564), DUF283 (590–715) and certain loops in RIIIDa and RIIIDb (1389–1545 and 1588–1658). In both the DICER–26S-GU and the DICER–26S-UG complex, the shRNAs are fully docked at the catalytic centre, formed by intramolecular dimerization of RIIIDa and RIIIDb. The alignment of conserved acidic residues in RIIIDa (E1316, D1320, D1561 and E1564) and RIIIDb (E1705, D1709, D1810 and E1813) with the RNA cleavage site, along with calcium ions positioned between these residues and the RNA, confirms the dicing-ready conformation (Extended Data Fig. 3c).
dsRBD and PAZ domain conformational changes
To examine how DICER adapts during RNA binding and cleavage, we compared our RNA-bound dicing-state structures (DICER–26S-GU and DICER–26S-UG) with the previously reported dicing-state structure (PDB: 7XW2) and apo-DICER (PDB: 7XW3)4. Root-mean-square deviation (RMSD) analysis, measuring atomic displacement between corresponding residues, revealed significant structural variability in the dsRBD and PAZ domains, highlighting transitions between functional states (Fig. 2e). Although the overall conformational changes in the dsRBD are consistent with previous observations4 (7XW2), the inward motion of the PAZ domain is distinctly observed in our RNA-bound dicing-state structures.
Alignment of the RNA-bound structures (DICER–26S-GU and DICER–26S-UG) with the 7XW2 dicing-state structure reveals a more compact conformation in the RNA-bound states, with a narrower overall width (57.7–58.8 Å for DICER–26S-GU and DICER–26S-UG vs. 68.0 Å for 7XW2) (Fig. 2f). This compaction is driven by structural rearrangements in the PAZ domain, which shifts inwards by around 7.4 Å after RNA binding (Fig. 2g). Specifically, the α-helix (residues 968–976) that anchors the RNA 3′-end moves inward by 7.6–8.1 Å, and adjacent β-sheets shift by 5.0–5.3 Å (Extended Data Fig. 3d). This differential motion compresses the PAZ domain, and probably contributes to bending nucleotides near the 3′-end (Fig. 2h).
The dsRBD and PAZ domain move in coordination
Structural comparisons of our RNA-bound dicing-state structures (DICER–26S-UG and DICER–26S-GU) with the previously reported dicing-state structure (7XW2) reveal coordinated movements of the dsRBD and PAZ domains. Although the dsRBD undergoes similar changes in all RNA-bound states, compared with apo-DICER (Fig. 2e), it shifts around 5 Å closer to DICER’s longitudinal axis in the DICER–26S-UG and DICER–26S-GU structures compared with 7XW2 (Fig. 2g). Combined with the inward motion of the PAZ domain, this shift increases the buried surface area of the DICER–RNA complex (Fig. 2i). These rearrangements align the RNA duplex along DICER’s axis, optimizing the positioning of cleavage sites for efficient RNA processing (Fig. 2i).
We propose that the dsRBD and PAZ domains act like chopsticks, gripping and aligning RNA substrates for precise double cleavages on the 5′- and 3′-strands (Fig. 2j). In our shRNA-bound structures, the RNA duplex remains tightly aligned without expansion (Extended Data Fig. 3e). By contrast, in pre-let-7a-1GYM from 7XW2, the dsRBD and PAZ domains are farther from the RNA duplex, which would require RNA helix widening to align both cleavage sites with the catalytic centre for accurate cleavage (Extended Data Fig. 3f).
Conserved structure of 3′-end binding pockets
In our dicing-state structures, the terminal wobble pairs (G–U and U–G) are unpaired, allowing the 3′-nt and 5′-nt to interact with the PAZ and platform domains, respectively (Fig. 3a,b, surface-view panels). The 3′-nt consistently occupies the conserved 3′-end binding pocket, in which the terminal phosphodiester linkage interacts with conserved residues, including tyrosines (Y936, Y971, Y972 and Y976) and basic residues (R937 and K975) (Fig. 3a,b, 3′-end binding pocket panels). This configuration aligns with the results of previous structural studies4,14,19,21,22,23,24, confirming the conserved nature of the 3′-end recognition mechanism of DICER. Notably, no residues interact with the terminal base of the 3′-nt, indicating that its identity is unlikely to affect cleavage-site selection (Extended Data Fig. 1e).
a, Binding pockets in the DICER–26S-UG complex. Left, cryo-EM model of DICER bound to 26S-UG with colour-coded domains. Left middle, electrostatic surface view of the PAZ–platform region, showing the paths of the 5′- and 3′-ends. Right middle, close-up view of the 3′-end pocket, with A64 and labelled residues. Right, 5′-end pocket, showing U1 contacts with R821 and R1003. b, Binding pockets in the DICER–26S-GU complex. Panels as in a. The 3′-end pocket accommodates A64, whereas the 5′-end pocket binds to G1, with residues D991 and H992 indicated. c, The alignment of the RNA backbone is influenced by 5′-end binding. In DICER–26S-GU, the RNA backbone shifts one base pair upwards towards the catalytic centre, compared with DICER–26S-UG. d,e, In vitro dicing of pre-mir-517a_GU (d) and pre-mir-517a_CU (e) by wild-type (WT) and mutant DICER. Left, denaturing gels showing DC21 and DC22 products. Middle, quantification of cleavage accuracy; significance by two-tailed, two-sample t-test (***P < 0.001, ****P < 0.0001; NS, not significant). Right, pre-miRNA schematics with cleavage sites. f, Cryo-EM map of DICER(D991G/H992G)–26S-GU, with colour-coded domains and RNA duplex (26S-GU) indicated. g, Electrostatic surface view of the active-site region of the mutant DICER, highlighting the bound duplex and the 5′- and 3′-strand positions. h, Close-up view of the PAZ domain and the terminal pair G1–U62. i, Proposed mechanism: distinct 5′ binding pockets (5′-G-favoured for DC21 and 5′-U-favoured for DC22) determine cleavage-site selection.
Identification of a 5′-G-favoured binding pocket
Unlike the conserved 3′-end binding pocket, the 5′-nt shows distinct binding differences between the DICER–26S-UG and DICER–26S-GU structures (Fig. 3a,b, surface-view panels). In the DICER–26S-UG structure, the 5′-U occupies the previously identified pocket4, in which R1003 interacts with the 5′-phosphate and R821 recognizes the uridine base (Fig. 3a, 5′-end binding pocket panels, and Extended Data Fig. 4a). However, R790, which was previously thought to interact with uridine, shows no interaction, consistent with mutation studies suggesting that R790 is not essential for uridine recognition4.
Our DICER–26S-GU structure reveals a previously unidentified 5′-G binding pocket. Here, the 5′-phosphate of 5′-G is anchored by H992, while its guanine base forms hydrogen bonds with D991 (Fig. 3b, 5′-end binding pocket panels, and Extended Data Fig. 4a). D991 and H992 create a G-selective microenvironment that is distinct from the 5′-U site. Binding of 5′-G shifts the RNA by approximately one nucleotide upwards along the hairpin’s lower stem compared with 5′-U (Fig. 3c). With identical upper RNA sequences, this shift positions DC21 at DICER’s catalytic centre for 5′-G, whereas 5′-U positions DC22. These structural differences provide a mechanistic basis for how 5′-nt identity biases cleavage-site selection.
Validation of the 5′-G binding pocket
To confirm the functional importance of the 5′-G binding pocket, we mutated the key residues D991 and H992 (Extended Data Fig. 4b). The mutations D991G, H992G and D991G/H992G reduced DC21 cleavage in pre-miRNAs with 5′-G, showing that these residues are essential for the function of the 5′-G binding pocket (Fig. 3d,e and Extended Data Fig. 4c–e). These mutations did not affect DC22 cleavage in pre-miRNAs with 5′-U, confirming the pocket’s specificity for 5′-G substrates (Fig. 3d,e and Extended Data Fig. 4c–e). Although the overall cleavage efficiency remained unchanged, the mutations influenced site selection, suggesting that these residues have a role in adjusting DICER positioning between DC21 and DC22. We further expressed pre-mir-517a_GU or pre-mir-517a_UG in HCT116 DICER-knockout cells with wild-type DICER or DICER(D991G/H992G), and performed miRNA sequencing (Extended Data Fig. 4f). Wild-type DICER produced more cleavage at DC21 from pre-mir-517a_GU than from pre-mir-517a_UG, consistent with the in vitro findings. However, the D991G/H992G mutant yielded less DC21 cleavage from pre-mir-517a_GU than wild-type DICER, but similar levels of DC21 and DC22 cleavage from both variants, confirming that DC21 enhancement by 5′-G depends on D991 and H992 (Extended Data Fig. 4g). In cells, cleavage at DC22 predominates, in part owing to the influence of the cofactor TRBP (transactivation response element RNA-binding protein) (Extended Data Fig. 4h).
We determined the structure of the DICER(D991G/H992G) mutant bound to 26S-GU (Fig. 3f and Extended Data Fig. 5a–g). The map shows continuous density for the loop containing residues 991–992 but lacks side-chain density for D991 or H992, consistent with their substitution by glycine (Extended Data Fig. 5h). Unlike in the wild-type DICER–26S-GU complex, the 5′-G shifts from the 5′-G pocket towards the 5′-U pocket, but does not interact with R1003 or R821 as 5′-U does (Fig. 3g and Extended Data Fig. 5i). Instead, the 5′-G is retained by base pairing with the opposing U (Fig. 3h and Extended Data Fig. 5j). The 3′-nt remains in the conserved 3′-end pocket (Extended Data Fig. 5k). These findings provide further evidence for the role of D991 and H992 in forming the 5′-G binding pocket.
The 5′-G binding pocket is conserved across species
Sequence analysis shows that D991 and H992, forming the core of the 5′-G binding pocket, are conserved across DICER enzymes in several species (Extended Data Fig. 6a), suggesting that this is an evolutionarily conserved functional feature. Massively parallel dicing assays on fly Dcr-1, a homologue of human DICER, revealed similar cleavage specificity, with 5′-G promoting DC21 cleavage and 5′-U favouring DC22 (Extended Data Fig. 6b–f). Consistently, an analysis of miRNAs derived from pre-miRNAs with different 5′-nts (A, G, U or C) across species showed that 5′-G pre-miRNAs yield the most DC21 miRNAs, whereas 5′-U pre-miRNAs produce the most DC22 miRNAs (Extended Data Fig. 6g,h).
On the basis of these findings, we propose a dual-pocket model for 5′-end recognition in DICER: a G-favoured binding pocket (formed by D991 and H992) for DC21 cleavage, and a U-favoured binding pocket (stabilized by R1003 and R821) for DC22 cleavage. These pockets align the RNA substrate 21 or 22 nucleotides from the 5′-end with DICER’s catalytic centres, enabling flexible yet precise RNA processing (Fig. 3i).
Resolving 5′-end counting and motif rules
In the DICER–26S-UG structure, the 5′-U directs cleavage at DC22, consistent with mWCU–YCR motifs that also guide DC22 cleavage. By contrast, the DICER–26S-GU structure shows that the 5′-G directs cleavage at DC21, conflicting with the DC22-guiding mWCU–YCR motifs. However, despite this conflict, 26S-GU is ultimately cleaved at DC22 (Fig. 1c). To understand how DICER resolves this conflict, we compared the protein structure of DICER in the DICER–26S-GU and DICER–26S-UG structures. The overall protein architectures are nearly identical, with only minor adjustments depending on whether the 5′-nt is G or U (Fig. 4a). Specific changes occur in a helix near the 3′ cleavage site in RIIIDa, the PAZ helix and a loop in the 5′-end binding pocket (Extended Data Fig. 7a). These subtle shifts suggest that DICER resolves cleavage conflicts mainly through local rearrangements, rather than through large-scale conformational changes.
a, Structural comparison of the DICER–26S-UG and DICER–26S-GU complexes. RMSD heat maps show minor structural differences in the 5′ binding pocket, PAZ domain and RIIIDa near the 3′ cleavage site. b, Alignment of the 26S-UG and 26S-GU RNA structures with their corresponding structures from AlphaFold3 (AF3)-predicted models. c, RNA flexibility and DICER’s dsRBD interactions: the 26S-GU complex exhibits distortion of the RNA backbone, aligning DC22 with the catalytic centre. d, Pre-dicing state of DICER bound to pre-mir-517a_GU. Cryo-EM density shows the PAZ domain (gold) engaging the duplex end and the dsRBD (gold) positioned along the RNA duplex (green), with other domains in grey. e, Dicing state. The duplex (green) docks into RIIIDa and RIIIDb (RIIIDa/b) as the PAZ domain and dsRBD (red) reposition to clamp the RNA for cleavage. f, Model of domain motion. Arrows indicate the movements of the dsRBD and the PAZ domain during activation. g, Density view of the end of the RNA duplex, showing G1 and opposing U59. h, The RNA duplex 5′-end occupies the DC21 pocket; the boundary loop (red) lies between DC21 and DC22. Open circles indicate pocket positions. i, Close-up view of the 5′-end region, showing G1 and the 5′-phosphate (PO4) coordinated near residues D991 and H992. j, RNA helical geometry comparison, highlighting the duplex distortions in 26S-GU compared with AlphaFold3 and experimental models of pre-mir-517a_GU. k, Proposed mechanism. 5′-U anchor: the 5′-U pocket aligns the mWCU–YCR motif at DC22 for cleavage. 5′-G anchor: the 5′-G pocket favours DC21 cleavage, misaligned with the mWCU–YCR motif at DC22. Thus, motif-driven RNA conformational changes override the 5′-G rule, distort the RNA backbone and ensure DC22 cleavage.
RNA rearrangements dictate cleavage sites
Our analysis reveals that RNA undergoes substantial conformational changes upon interaction with DICER, and these are particularly evident in the DICER–26S-GU structure compared with DICER–26S-UG. Although the predicted secondary structures of 26S-GU and 26S-UG are similar (AlphaFold341; Fig. 4b), DICER induces distinct structural changes at the RNA ends and cleavage sites (Fig. 4b). At the ends, interactions with the 5′- and 3′-end binding pockets destabilize the base pairs of terminal nucleotides, anchoring them in specific positions4,14 (Fig. 4b). Near the cleavage site, interactions between RNA motifs (for example, mWCU and YCR) and DICER shape structural distortions. Although YCR does not directly interact with DICER residues, R1855 in dsRBD engages with the 19-CC mismatch of mWCU, aiding RNA backbone adjustments for cleavage positioning (Fig. 4c).
In the DICER–26S-UG structure, the 5′-U pocket positions DC22 at the catalytic centre, with a moderate separation between the 19-CC mismatch and R1855. In the DICER–26S-GU structure, the 5′-G pocket shifts the backbone upwards, at first aligning DC21 and bringing the 19-CC mismatch closer to R1855. Engagement of this motif imposes a register-specific strain: the 26S-UG backbone undergoes a subtle adjustment, whereas the 26S-GU backbone distorts more substantially to accommodate the interaction and realign DC22 at the catalytic centre (Fig. 4c and Extended Data Fig. 7b,c). This mechanism might help to explain how RNA-motif interactions can override the DC21 bias of the 5′-G pocket, favouring DC22.
In humans, pre-miRNAs with a 5′-G produced the highest levels of DC21 miRNAs, followed by 5′-A, 5′-C and 5′-U (Extended Data Fig. 7d). However, around 42% of 5′-G pre-miRNAs still produced DC22 miRNAs, and around 20% of 5′-U pre-miRNAs generated DC21 miRNAs. Notably, around 27.3% of 5′-G pre-miRNAs yielding DC22 miRNAs contained a YCR motif at DC22, whereas around 19% of 5′-U pre-miRNAs generating DC21 miRNAs had a YCR motif at DC2135,40 (Extended Data Fig. 7d). These findings show that strong motifs, such as YCR, can override the 5′-end counting rule to determine cleavage sites.
DICER dicing with 5′-G pre-miRNA
To structurally validate the role of the 5′-G pocket in promoting DC21 cleavage in a pre-miRNA context, we resolved the structure of DICER bound to pre-mir-517a_GU, cleaved at DC21 (Extended Data Fig. 8a–d). Both pre-dicing and dicing states were observed (Fig. 4d,e and Extended Data Fig. 8e). The pre-dicing state resembles previous structures14, whereas the dicing state matches those described here, confirming the movements of the dsRBD and the PAZ domain during cleavage (Fig. 4f). In the dicing-state structure, the terminal G–U pair is unpaired, with the 5′-G occupying the same pocket as in 26S-GU and engaging D991 and H992 (Fig. 4g–i). The 5′-G and mWCU motif at position 17 both promote DC21, reinforcing cleavage at the same site. Consequently, the RNA conformational shifts seen in 26S-GU are absent in pre-mir-517a_GU (Fig. 4j).
These observations generalize to how sequence motifs interface with the 5′-end counting rule. When mWCU–YCR motifs align with the 5′-end counting rule (for example, 5′-U with mWCU–YCR at DC22), DICER cleaves without notable structural changes in the RNA. However, when motifs conflict with the counting rule (for example, 5′-G with mWCU–YCR at DC22), the motif can override the rule by inducing RNA structural rearrangements. These distortions realign the cleavage site with DICER’s catalytic centre, allowing the motif to dictate cleavage (Fig. 4k).
Discussion
Our study refines the DICER 5′-end binding model. Unlike the previous proposal, which suggested that there is a single pocket that disfavours 5′-G4, we identify a 5′-G-favoured pocket and a distinct 5′-U-favoured pocket. The G-favoured pocket biases cleavage toward DC21, whereas the U-favoured pocket biases DC22. Structural and biochemical data support this dual-pocket mechanism, and highlight the prominent role of the 5′-nt in cleavage specificity. Whereas A and C show moderate selectivity, G and U strongly bias DC21 and DC22, respectively. This framework explains variation in register choice and enables prediction of how 5′-end identity influences site selection across substrates.
The 5′-G pocket is formed by D991 and H992, whereas the 5′-U pocket involves R1003 and R821. H992, which has previously been linked to 5′-phosphate recognition17 (PDB: 4NGG), anchors the 5′-phosphate of guanine. Our analysis of hydrogen-bonding interactions provides a mechanistic basis for the distinct preferences for 5′-G and 5′-U (Extended Data Fig. 9a). Both G and U can accept hydrogen bonds via specific oxygen atoms to interact with R821. G can form one hydrogen bond with R821 through O6, but steric hindrance by R821 prevents G from entering the 5′-U pocket4. By contrast, U can interact with R821 through two hydrogen bonds involving O2 and O4. Unlike U, G also donates hydrogen bonds through its exocyclic amino group (-NH2), enabling interaction with D991. Together, these factors help to explain the strong preference of the 5′-G pocket for G over U. A and C can form weaker hydrogen bonds with R821 or D991, mainly through nitrogen atoms. Because nitrogen is less electronegative than oxygen, these bonds are less stable, which accounts for the moderate selectivity of A and C for either pocket. Further structural and biochemical work will be needed to define the binding modes of A and C in detail.
Our cryo-EM structures reveal marked conformational changes in the PAZ and dsRBD domains, which are crucial for RNA positioning in DICER’s catalytic centres. For the RNAs analysed (26S-UG, 26S-GU and pre-mir-517a), the dsRBD adjusts its angle to align the RNA duplex along DICER’s longitudinal axis, whereas the PAZ domain compresses to achieve precise alignment, facilitating efficient double-stranded cleavage. By contrast, pre-let-7a-1 from a previous study4 requires RNA backbone expansion near the cleavage site to compensate for suboptimal alignment, thereby reducing the likelihood of mis- or single-site cleavage events. In pre-let-7a-1, the PAZ contacts the terminal nucleotide, relaxing the last three nucleotides; by contrast, in pre-mir-517a and 26S RNAs, the 3′-terminal nucleotides are curved, with overhang nucleotides contacting specific residues (Extended Data Fig. 9b). These patterns suggest that DICER adapts to different RNA substrates. Among animal DICER structures, the PAZ domain in fly Dcr-1 and mouse DICER moves closer to the longitudinal axis, whereas this movement is absent21,22,23 in fly Dcr-2, indicating that PAZ domain movement varies depending on enzyme–RNA interactions (Extended Data Fig. 9c).
This study shows that the 5′-nt is a key determinant of DICER cleavage at DC21 versus DC22, whereas the terminal 3′-nt generally has minimal effect. However, in 5′-A and 5′-G pre-miRNAs, the cleavage sites exhibit greater variability (Fig. 1f,g), suggesting that the 3′-end overhang has an influence. Future work should define how 3′-end sequence and structure modulate register choice. RNA motifs, such as mWCU and YCR, also contribute to cleavage specificity, either reinforcing or opposing 5′-end cues to guide DICER to specific sites. For example, in pre-mir-629, 5′-U favours DC22, whereas the YCR motif favours DC21, resulting in cleavage at both sites. Conversely, in pre-mir-517a, 5′-G or 5′-A cooperates with the mWCU motif to restrict cleavage to DC21, whereas 5′-U works with the YCR motif to direct cleavage at DC22 (Extended Data Fig. 10a). These findings highlight how DICER integrates competing signals. For instance, in 26S-UG, 5′-U, the mWCU–YCR motif and the long stem cooperatively enforce DC22 cleavage. In 26S-GU, 5′-G favours DC21, but the mWCU–YCR motif and long stem override this preference, directing cleavage to DC22; removing the motif or shortening the stem shifts cleavage toward DC21 (Extended Data Fig. 10b). Structurally, dsRBD engagement with the mWCU motif correlates with backbone distortions that realign the scissile site with the catalytic centres, providing a flexible mechanism for resolving competing cues. As in previous work4, we did not resolve unambiguous base-specific side-chain contacts for YCR, but YCR still biases cleaving towards DC22, consistent with motif-driven RNA conformational effects that may precede docking. Our study also highlights how conflicts—such as between 5′-G and mWCU–YCR at DC22—are resolved. Investigating conflicts that involve different 5′-ends, motifs (for example, mWCU or YCR), stem lengths or bulges in diverse RNA contexts would be valuable. Future high-throughput structural and artificial-intelligence-assisted approaches could sharpen mechanistic models of these interactions.
Our findings highlight the evolutionary conservation of the dual-pocket architecture across metazoans. Sequence analysis shows that the 5′-G and 5′-U pockets are preserved among DICER enzymes in these species, emphasizing their functional role in RNA processing. This dual-pocket mechanism is likely to have evolved to ensure precise and flexible RNA cleavage, enabling DICER to process a wide range of RNA substrates. Biochemical evidence, using fly Dcr-1 as a model, supports the presence of both pockets, mirroring human DICER. Examining these features in non-metazoan systems, such as plants or protozoa, could provide further insights into RNAi evolution.
The discovery of the 5′-G binding pocket, which promotes DC21 cleavage, offers valuable insights for shRNA design. Because shRNAs are typically produced by RNA polymerase III, which often incorporates a G at the first nucleotide, this feature can predispose substrates toward DC21 cleavage. Coordinating RNA elements (such as stem length) and RNA sequence motifs (such as mWCU or YCR) to enhance DC21 production could optimize DICER processing at DC21 in cells, thereby improving shRNA knockdown efficiency.
Methods
Plasmid construction
The pXG-10×His-DICER plasmid was generated by inserting a DNA sequence encoding human DICER (amino acids 25–1922) and a sequence encoding a 10-histidine tag at the N terminus of DICER into the pXG plasmid using the In-Fusion cloning kit (Takara). The pXG-10×His-DICER mutant variants were obtained through site-directed mutagenesis using the pXG-10×His-DICER plasmid as the template. Mutated sites were confirmed by Sanger sequencing. A list of the plasmids and oligonucleotides used for their construction is provided in Supplementary Table 2.
The Dcr-1-bacmids were prepared as follows. The DNA coding sequence of Drosophila melanogaster Dicer-1 (Dcr-1) was obtained from cDNA synthesized using random hexamers and total RNA extracted from D. melanogaster cells. The coding sequence of Dcr-1 was cloned into the pBIG plasmid42 using a restriction cloning scheme, resulting in the pBIG-Dcr-1 construct. The pBIG-Dcr-1 plasmids were introduced into DH10EMBacY E. coli cells to produce Dcr-1-bacmids. After blue–white selection on agar plates containing Bluo-gal, IPTG and antibiotics, Dcr-1-bacmids were isolated from white colonies testing positive. Purification of bacmids was performed using alkaline lysis and alcohol precipitation methods. The presence of the gene encoding Dcr-1 within the bacmids was confirmed by PCR using the pUC-M13 primer pair. The primers used for generating pBIG-Dcr-1 and amplifying the Dcr-1 coding sequence are listed in Supplementary Table 2.
Protein expression
Wild-type human DICER and mutant variants were expressed using the human cell system HEK293E, as previously described34,35,36. The DICER plasmids were prepared using the MaxiPrep kit (Thermo Fisher Scientific). HEK293E cells were cultured in 100-mm dishes in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 5% fetal bovine serum (FBS) at 37 °C. Each 100-mm dish was transfected with 10 µg of plasmid DNA and 30 µg of linear polyethyleneimine (L-PEI) as the transfection reagent. Cells were collected 72 h after transfection.
For the baculovirus system, Dcr-1-bacmids were transfected into Sf9 cells (provided by S. Dang), which were cultured in in a six-well plate in 2 ml ESF-921 (Expression Systems) per well using the CellFectin II reagent (Thermo Fisher Scientific) to generate the initial baculovirus stock (P0). The virus was subsequently amplified through two additional passages to achieve a viral titre that was sufficient for efficient protein expression. Sf9 cells infected with the appropriate amount of virus were collected 72 h after infection in a shaking incubator at 27 °C.
Protein purification
For DICER purification, we collected approximately 100 dishes (100-mm) of HEK293E cells expressing DICER. For Dcr-1, we collected around 200 ml cell culture of approximately 400 million insect cells expressing Dcr-1. The purification procedures for the two proteins were similar and are described below.
The cell pellets were resuspended in a lysis buffer at a ratio of 1:10 (cell pellet:buffer volume). The lysis buffer consisted of 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 4 mM β-mercaptoethanol and 10% glycerol, and was supplemented with RNase A and a protease inhibitor cocktail. After resuspension, the cells were subjected to a brief sonication step to disrupt the cell membranes. The lysates were then clarified by high-speed centrifugation at 18,000 rpm for 30 min.
The clarified supernatant was immediately applied to a pre-equilibrated Ni-NTA column. Unbound and nonspecifically bound proteins were removed with wash buffers containing either 150 mM NaCl or 1,000 mM NaCl, supplemented with 25 mM imidazole. His-tagged proteins were eluted from the Ni-NTA beads using an elution buffer (T150) containing 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 4 mM β-mercaptoethanol and 200 mM imidazole. The eluate was then applied to Q-Sepharose beads at 100 mM NaCl, and the bound proteins were eluted at 500 mM NaCl to achieve higher purity.
The partially purified protein was further processed using gel-filtration chromatography (Bio-Rad NGC). The final elution buffer consisted of 50 mM Tris (pH 7.5), 500 mM NaCl, 0.5 mM TCEP and 10% glycerol. Peak fractions were collected, pooled and concentrated using Centricon devices with a cut-off of 100 kDa. The concentrated protein was rapidly frozen in liquid nitrogen and stored at −80 °C for future use.
In vitro pre-miRNA dicing assay
The pre-miRNAs were synthesized using the method described in our previous studies34,35,36. The oligonucleotides used for each RNA were synthesized by a commercial company (BGI). We performed two sequential PCR reactions to generate IVT-DNA sequences containing the T7 promoter, hammerhead ribozyme sequence, pre-miRNA sequence and HDV ribozyme sequence. A total of 200 ng of IVT-DNA was added to a 20-µl in vitro transcription reaction using the MEGAscript T7 kit, and the reaction was incubated overnight at 37 °C. The RNA products were treated with 40 mM MgCl2 to activate the ribozyme reaction, resulting in pre-miRNAs with a 3′-phosphate and a 5′-OH group. The pre-miRNAs were then treated with T4 Polynucleotide Kinase (T4 PNK) to convert the 3′-phosphate and 5′-OH groups into 3′-OH and 5′-phosphate, respectively. The oligos that were used to produce pre-miRNAs are listed in Supplementary Table 3.
The in vitro pre-miRNA dicing assay was performed in a 10 µl reaction mixture containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 10% glycerol, 1 mM DTT and 2 mM MgCl2. Approximately 2 pmol of pre-miRNA was incubated with 1–2 pmol of purified recombinant DICER for 30 min at 37 °C. The reaction was terminated by adding 2× TBE loading buffer containing 10 µg ml−1 of proteinase K, followed by heat treatment at 50 °C for 15 min. The reaction mixture was then denatured at 95 °C for 5 min before being loaded onto a pre-run 15% urea–denaturing PAGE gel. Electrophoresis was performed for approximately 50 min at 300 V. The gel was stained with 0.1% SYBR Green II in TBE buffer for 8 min and imaged using the Bio-Rad Gel Doc XR+ system. Quantification of product band intensities was performed using Image Lab v.6.0.1 software.
Massively parallel dicing assays for randomized pre-mir-324 and library construction
Synthesis of randomized pre-mir-324
The in vitro synthesis of four randomized pre-mir-324 groups, each containing three randomized nucleotides at the 3′-end and one of four specific nucleotides at the 5′-end, was performed as follows. Note that we removed the 5′-bulged U near the 5′ cleavage site so that the F1 and F3 fragments generated by DICER have the same length, simplifying gel-based interpretation of cleavage. For each group, forward and reverse primers with overlapping regions were annealed and extended by a single-cycle PCR using the Klenow exo fragment to produce double-stranded DNA (dsDNA-1) containing the hammerhead ribozyme and pre-miRNA regions. Each group used a unique forward primer, and all groups shared the same reverse primer. In the second PCR step, a new set of primers was used. The reverse primer introduced three randomized nucleotides at the 3′-end of pre-mir-324, and the forward primer included the T7 promoter and the hammerhead ribozyme sequence. The resulting DNA from the second PCR, referred to as IVT-dsDNA for each group, contained the T7 promoter, hammerhead ribozyme sequence and pre-miRNA sequence. Approximately 400 ng of purified dsDNA from each group was used as the template for in vitro transcription. The oligonucleotides used for this process are listed in Supplementary Table 4.
The in vitro transcription reaction was performed at 37 °C for 12 h using the MEGAscript T7 kit (Thermo Fisher Scientific). Afterwards, 40 mM MgCl2 was added to activate the hammerhead ribozyme’s self-cleavage, separating the pre-miRNA sequence (now with a 5′-OH) from the ribozyme. The reaction mixture was subjected to three thermal cycles (72 °C for 1 min, 65 °C for 5 min and 37 °C for 10 min) to facilitate ribozyme activity. The RNA products were resolved on an 8% urea–denaturing PAGE gel (300 V, 40 min), and the pre-miRNA band was excised on the basis of its expected size. The RNA was extracted using an elution buffer (500 mM NaCl and 5 mM EDTA, pH 8.0) to prevent cation-dependent degradation and purified by isopropanol precipitation. The purified RNA was treated with T4 PNK (Thermo Fisher Scientific) in Thermo Buffer A to convert its 5′-OH to a 5′-phosphate. A final isopropanol purification yielded the randomized pre-miRNAs, which were stored at −80 °C for downstream assays.
Massively parallel dicing assays for randomized pre-mir-324
For the massively parallel dicing assays, 2 pmol of each of the four randomized groups of pre-mir-324 (groups A, U, G and C, based on the 5′-nt) were independently processed with approximately 1 pmol of recombinant human DICER or D. melanogaster Dcr-1 at 37 °C for 30 min. The reactions were terminated by adding 2× TBE sample loading buffer supplemented with 10 µg ml−1 proteinase K. The mixtures were incubated at 50 °C for 15 min, denatured and resolved on a 12% urea–denaturing PAGE gel, separating cleaved products from substrates. Cleaved product bands were excised from the gel and purified using an ethanol–isopropanol precipitation with GlycoBlue (Thermo Fisher Scientific) as a co-precipitant.
Construction of sequencing libraries
To prepare libraries for the original pre-mir-324 substrates, the circular ligation scheme was used. A total of 2 pmol of pooled RNA from all four groups (A, U, G and C) was ligated with a 4N-RA3 oligo using T4 RNA Ligase 2-truncated KQ (Thermo Fisher Scientific). The ligated RNA was reverse-transcribed with a 6N-R-RA3-cirRTP primer at 50 °C for 15 min using SuperScript IV Reverse Transcriptase (Invitrogen). After reverse transcription, the original RNA was degraded by treating the reaction with 0.1 M NaOH at 90 °C for 10 min. The cDNA was purified by 12% urea–denaturing PAGE gel fallowed by ethanol precipitation. The purified cDNA was circularized with CircLigase ssDNA ligase (Epicentre) and separated from linear cDNA on an 18% urea–denaturing PAGE gel. The circularized cDNA served as the template for a final PCR, performed using RP1 and RPx primers from the TruSeq Illumina system, to generate the DNA library for original substrates.
For the cleaved products, separate libraries were constructed for each of the four groups (A, U, G and C). Each RNA sample was ligated with the 4N-RA3 primer using T4 RNA Ligase 2-truncated KQ. After ligation, the samples were resolved on a 12% urea–denaturing PAGE gel to separate ligated products from unligated oligos. The ligated products were then ligated with the 4N-RA5 RNA oligo using T4 RNA Ligase 1. The double-ligated RNA was reverse-transcribed with the R-RA3 primer using Superscript IV Reverse Transcriptase. The resulting cDNA pools were used as templates for the final PCR with RP1 and RPIx primers from the TruSeq Illumina system, generating DNA libraries for cleaved products. Separate libraries were prepared for each of the four subgroups (A, U, G and C).
These libraries for both original substrates and cleaved products were sequenced using an Illumina NovaSeq 6000 in 150-bp paired-end mode (HaploX). The oligonucleotides used for library preparation are listed in Supplementary Table 4.
Analysis of massively parallel dicing assays for randomized pre-mir-324
The raw sequencing reads were processed using the following pipeline. First, the 3′ and 5′ adapter sequences were removed using the cutadapt tool43 with the command cutadapt -a TGGAATTCTCGGGTGCCAAGG -A GATCGTCGGACTGTAGAACTCTGAAC. Next, paired-end reads were joined using the fastq-join tool with default parameters. After obtaining the joined reads, low-quality reads were filtered out using the fastq_quality_filter tool with the parameters -q 20 -p 9044.
The reads were then collapsed using the fastx_collapser tool to remove duplicates that shared the same ligation barcode (http://hannonlab.cshl.edu/fastx_toolkit/index.html, v.0.0.13). After this, a second round of trimming with cutadapt was performed to remove the 4N/4N and 6N/4N ligation barcodes at the 5′- and 3′-ends of the product reads and original substrate reads, respectively.
The processed reads were mapped to the pre-mir-324 reference sequence using the BWA mapping toolkit45. Only reads that were perfectly mapped to a single variant (out of 64 variants for each subgroup) were selected for further analysis. These reads contained the randomized sequence at 3′-ends and the DICER cleavage sites at 5′-ends.
For each pre-mir-324 variant (for example, var1), mapped read counts in the substrate sample were normalized to the total substrate reads and converted to reads per million (RPM), denoted as Control(var1). In the product sample, mapped read counts for each cleaved product were similarly normalized to the total product reads and converted to RPM. A given variant can yield multiple cleaved products with distinct 5′-ends that correspond to different DICER cleavage sites. Let NPx denote the RPM of the cleaved product whose DICER cleavage site is at position x. Cleavage accuracy for position x within a variant is defined as the fraction of reads at x among all cleavage positions observed for that variant, Accuracyx(var1) = NPx/ΣiNPi. Cleavage efficiency for position x within a variant is defined as the fraction of product reads at x relative to the total substrate abundance of that variant, Efficiencyx(var1) = NPx/Control(var1).
Pre-miRNA structure analysis
To investigate the effect of the 5′-nt on selection of DICER cleavage sites in human pre-miRNA dicing, we used data from our previous study about the enrichment of the YCR motif for analysis35. Sequences and major cleavage sites of 566 human pre-miRNAs were collected from MirGeneDB40. The secondary structures of the pre-miRNA sequences were predicted using RNAfold (ViennaRNA Package)46. To pinpoint DICER cutting sites for each pre-miRNA, we analysed either the 5′-terminus of the mature 3′-strand miRNAs or the 3′-terminus of the mature 5′-strand miRNAs. The presence and position of YCR motifs in pre-miRNAs were identified previously35. The pre-miRNA sequences, their corresponding miRNA sequences and the YCR motifs identified are presented in Supplementary Table 5.
In vitro reconstitution of DICER with shRNAs
The chemically synthesized shRNAs 26S-GU (5′-pGGGAUAUUUCUCGCAGAUCUCAUGUGAAAAAAAAAACACAUGACAUCUGUGAGAAAUAUUCUUA) and 26S-UG (5′-pUGGAUAUUUCUCGCAGAUCUCAUGUGAAAAAAAAAACACAUGACAUCUGUGAGAAAUAUUCGUA) and pre-mir-517a_GU (5′-pGCUCUAGAUGGAAGCACUGUCUGUUGUAUAAAAGAAAAGAUCGUGCAUCCCUUUAGAGUGU) were obtained from GenCefe and dissolved in RNase-free water to a final concentration of around 100 µM.
To assemble the RNA–protein complex, 20 pmol of DICER protein was mixed with 60 pmol of shRNA or pre-mir-517a_GU in assembly buffer containing 50 mM Tris-HCl (pH 8), 150 mM NaCl, 0.5 mM TCEP, 2 mM Ca2+ and 5% glycerol. The reaction, performed in a 10-µl PCR tube, was incubated on ice for three hours before being loaded onto EM grids.
Because shRNA binds to DICER at an estimated 1:1 ratio, a 3:1 RNA:protein ratio was used to ensure nearly complete occupancy of DICER by shRNA. Sample homogeneity was assessed by negative staining immediately before EM grid freezing.
Preparation of cryo-EM samples
The assembled samples for both DICER–26S-GU and DICER–26S-UG complexes were prepared using the same protocol. An aliquot of approximately 4 µl sample was applied to glow-discharged Quantifoil R2/2 300-mesh Cu grids. After application, the sample was blotted at 100% humidity and 4 °C, followed by vitrification in liquid ethane using a Vitrobot Mark IV (Thermo Fisher Scientific; blot force 0, wait time 30 s, blotting time 4 s with blotting paper no.2) at the Biological Cryo-EM Center at Hong Kong University of Science and Technology (HKUST). Cryo-EM sample preparation for DICER–pre-mir-517a_GU and DICER(D991G/H992G)–26S-GU followed the same protocol as that for DICER–26S-GU, except that we used Quantifoil R1.2/1.3 400-mesh Au grids. Grids were screened on a Glacios (Thermo Fisher Scientific) at 200 keV, and those with evenly distributed particles and a suitable ice thickness were selected for data collection.
Data collection was done using a 300-kV Titan Krios G3i cryo-TEM microscope (Thermo Fisher Scientific) located at the Biological Cryo-EM Center at HKUST. Microscope settings: Gatan K3 direct electron detector in counting mode; nominal magnification 81,000× (physical pixel size 1.051 Å). Exposure: total dose 50 e− Å−2, fractionated into 40 frames (3.1 s total); dose rate of around 17.7 e− per pixel per second and around 1.25 e− Å−2 per frame. Defocus: −1.0 to −2.4 µm.
For the DICER–26S-GU complex, 23,300 movies were collected from 5 datasets. The DICER–26S-UG complex included 11,241 movies from 3 datasets, DICER(D991G/H992G)–26S-GU had 16,972 movies from 2 datasets and DICER–pre-mir-517a_GU had 14,244 movies from 3 datasets. Detailed data collection parameters for these complexes are provided in Supplementary Table 1.
Cryo-EM data processing and 3D refinement
All image processing was performed in cryoSPARC v.4.6.2 (Structura Biotechnology)47. Movies were corrected using Patch motion correction with default settings and binned to the physical pixel size, and CTF parameters were estimated per micrograph using Patch CTF estimation (with default setting, fit range of around 4–25 Å). Micrographs with poor CTF fits, thick ice or contamination were excluded after manual inspection.
Particles were first identified by blob picking (radii 100–250 Å) and extracted in 256-pixel boxes, followed by multiple rounds of 2D classification to remove junk and retain views characteristic of the DICER–RNA dicing state. High-quality 2D classes were used as templates for template-based auto-picking (particle diameter 200 Å), after which additional 2D cleaning was performed. Cleaned particle sets of around 181,000 particles were seeded for ab initio reconstruction (C1 symmetry) to obtain initial volumes. The appropriate initial volume resembling DICER–RNA complexes was selected and refined with non-uniform refinement to obtain the final maps.
The final DICER–26S-UG map (641,317 particles) reached a resolution of 3.34 Å by GS-FSC, and the DICER–26S-GU map (1,755,133 particles) reached 3.37 Å. The map of DICER(D991G/H992G)–26S-GU (787,381 particles) reached a resolution of 3.29 Å by GS-FSC. The maps of DICER–pre-mir-517a_GU in the pre-dicing (1,272,937 particles) and dicing (475,650) state reached resolutions of 3.00 Å and 3.21 Å by GS-FSC, respectively. Maps were sharpened with CryoTEN (default settings) to obtain final maps for model building48.
Model building
The published model of the DICER protein in the dicing state with pre-let-7a-1GYM (PDB: 7XW2) was used as the initial protein model for the DICER–26S-GU, DICER–26S-UG and DICER–pre-mir-517a_GU in dicing-state maps. The initial RNA models for 26S-UG, 26S-GU and pre-mir-517a_GU were generated using AlphaFold3 for three-dimensional (3D) RNA structure prediction41. The refined DICER–26S-GU model served as the starting model for DICER(D991G/H992G)–26S-GU; D991G and H992G mutations were introduced in ChimeraX (v.1.7)49. For the DICER–pre-miR-517a_GU pre-dicing state, the apo structure (PDB: 7XW3) was used as the initial model. These initial models were aligned with the cryo-EM density maps using the Fit-in-Map tool in ChimeraX v.1.7, followed by manual refinement in Coot (WinCoot v.0.9.8.96)49,50. The manually fitted models were further refined using phenix.real_space_refine in PHENIX (v.1.20.1)51. Model validation was done with phenix.validation_cryoem51.
All figures presented in this study were generated using ChimeraX v.1.749 and PyMOL (Schrödinger)52.
Small-RNA analysis
The DNA sequences coding for pri-miRNA (pre-miRNA sequences with a 20-nt extension on both ends) were cloned into the pcDNA3 vector using a ligation strategy. Detailed primer information is provided in Supplementary Table 2. HCT116 DICER-knockout cells (provided by N. Kim) were cultured in six-well plates using McCoy’s 5A medium supplemented with 10% FBS (Gibco). Transfections were performed with 1.5 µg of either pXG-DICER-WT or pXG-DICER-D991G-H992G, along with 0.25 µg of pcDNA3-pri-mir-517a_GU or pcDNA3-pri-mir-517a_UG, using lipofectamine. Total RNA was extracted 48 h after transfection using TRIzol reagent (Invitrogen).
We constructed RNA libraries from isolated small RNA fragments obtained from 4 µg of total RNA per sample using a 12% urea–PAGE gel. Library preparation was performed using the NEBNext Small RNA Library Prep Set for Illumina (NEB, E7330S). In brief, the purified small RNA fragments were first ligated to an adenylated 3′ adapter (AppAGATCGGAAGAGCACACGTCT-NH2). To prevent excess adapter from interfering with subsequent steps, a reverse complementary oligonucleotide was used. The 3′-ligated RNAs were then ligated to a 5′ adapter (GUUCAGAGUUCUACAGUCCGACGAUC). After adapter ligation, the RNAs were reverse-transcribed into cDNA, which was subsequently amplified by PCR using indexed primers to generate DNA libraries. Each sample was prepared in three biological replicates.
The small-RNA libraries were sequenced using the Illumina NovaSeq 6000 platform in 150-bp paired-end mode (HaploX). For sequencing data analysis, the adapters were first removed from read1 and read2 using the commands cutadapt -a AGATCGGAAGAGCACACGTCT and cutadapt -a GATCGTCGGACTGTAGAACTCTGAAC, respectively45. The reads were then concatenated using fastq-join, and low-quality reads were excluded using fastq_quality_filter with the parameters -q 20 -p 9044. The resulting reads were mapped to a customized reference containing pri-miRNA sequences using Bowtie253. Reads mapping to pri-miRNA sequences were selected for further analysis. The starting positions of reads mapped to the 3′-miRNA regions were used to identify DICER cleavage sites. IsomiR frequency was calculated as the ratio of the positional RPM to the sum of all positional RPMs. We categorized the isomiRs into three groups—DC21, DC22 and DC-other—which correspond to DICER cleavage at DC21, DC22 and other positions, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data and processed data for massively parallel dicing assays have been deposited at the Gene Expression Omnibus (GEO) under accession number GSE296721. Structural models of DICER–26S-GU, DICER–26S-UG, DICER(D991G/H992G)–26S-GU, DICER–pre-mir-517a_GU (pre-dicing state) and DICER–pre-mir-517a_GU (dicing state) have been deposited in the PDB under the accession codes 9V42, 9V43, 21CQ, 21CB and 21CN, respectively. The density maps for the above models have been deposited in the Electron Microscopy Data Bank (EMDB) under the following accession codes: EMD-64764 (DICER–26S-GU), EMD-64765 (DICER–26S-UG), EMD-67575 (DICER(D991G/H992G)–26S-GU), EMD-67570 (DICER–pre-mir-517a_GU, pre-dicing) and EMD-67573 (DICER–pre-mir-517a_GU, dicing). A complete list of accession codes is provided in Supplementary Table 1. Additional structural models analysed in this study are available in the PDB under the accession codes: 7XW2, 7XW3, 8DG5, 8DGI, 7W0E, 7W0B, 7YYN, 7YZ4 and 7ZPI.
Code availability
Analysis codes are available at https://github.com/mkngo2797/pocket_project.
References
Bartel, D. P. Metazoan microRNAs. Cell 173, 20–51 (2018).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Shang, R., Lee, S., Senavirathne, G. & Lai, E. C. microRNAs in action: biogenesis, function and regulation. Nat. Rev. Genet. 24, 816–833 (2023).
Lee, Y. Y., Lee, H., Kim, H., Kim, V. N. & Roh, S. H. Structure of the human DICER–pre-miRNA complex in a dicing state. Nature 615, 331–338 (2023).
Bofill-De Ros, X. & Gu, S. Guidelines for the optimal design of miRNA-based shRNAs. Methods 103, 157–166 (2016).
Paddison, P. J., Caudy, A. A., Bernstein, E., Hannon, G. J. & Conklin, D. S. Short hairpin RNAs (shRNAs) induce sequence-specific silencing in mammalian cells. Genes Dev. 16, 948–958 (2002).
Rao, D. D., Vorhies, J. S., Senzer, N. & Nemunaitis, J. siRNA vs. shRNA: similarities and differences. Adv. Drug Deliv. Rev. 61, 746–759 (2009).
Setten, R. L., Rossi, J. J. & Han, S. P. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 18, 421–446 (2019).
Siolas, D. et al. Synthetic shRNAs as potent RNAi triggers. Nat. Biotechnol. 23, 227–231 (2005).
Taxman, D. J., Moore, C. B., Guthrie, E. H., & Huang M. T.-H. In RNA Therapeutics: Function, Design, and Delivery. Methods in Molecular Biology Vol. 269 (ed. Sioud, M) 139–156 (Humana Press, 2010).
Zhang, H., Kolb, F. A., Brondani, V., Billy, E. & Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 21, 5875–5885 (2002).
Zhang, H., Kolb, F. A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Vermeulen, A. et al. The contributions of dsRNA structure to Dicer specificity and efficiency. RNA 11, 674–682 (2005).
Liu, Z. et al. Cryo-EM structure of human Dicer and its complexes with a pre-miRNA substrate. Cell 173, 1191–1203 (2018).
MacRae, I. J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006).
Park, J. E. et al. Dicer recognizes the 5′ end of RNA for efficient and accurate processing. Nature 475, 201–205 (2011).
Tian, Y. et al. A phosphate-binding pocket within the platform–PAZ–connector helix cassette of human Dicer. Mol. Cell 53, 606–616 (2014).
Sinha, N. K., Iwasa, J., Shen, P. S. & Bass, B. L. Dicer uses distinct modules for recognizing dsRNA termini. Science 359, 329–334 (2018).
Yamaguchi, S. et al. Structure of the Dicer-2–R2D2 heterodimer bound to a small RNA duplex. Nature 607, 393–398 (2022).
MacRae, I. J., Zhou, K. & Doudna, J. A. Structural determinants of RNA recognition and cleavage by Dicer. Nat. Struct. Mol. Biol. 14, 934–940 (2007).
Jouravleva, K. et al. Structural basis of microRNA biogenesis by Dicer-1 and its partner protein Loqs-PB. Mol. Cell 82, 4049–4063 (2022).
Zapletal, D. et al. Structural and functional basis of mammalian microRNA biogenesis by Dicer. Mol. Cell 82, 4064–4079 (2022).
Su, S. et al. Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD. Nature 607, 399–406 (2022).
Wei, X. et al. Structural basis of microRNA processing by Dicer-like 1. Nat. Plants 7, 1389–1396 (2021).
Wang, Q. et al. Mechanism of siRNA production by a plant Dicer–RNA complex in dicing-competent conformation. Science 374, 1152–1157 (2021).
Gu, S. et al. The loop position of shRNAs and pre-miRNAs is critical for the accuracy of dicer processing in vivo. Cell 151, 900–911 (2012).
Schopman, N. C. T., Liu, Y. P., Konstantinova, P., ter Brake, O. & Berkhout, B. Optimization of shRNA inhibitors by variation of the terminal loop sequence. Antiviral Res. 86, 204–211 (2010).
Starega-Roslan, J., Galka-Marciniak, P. & Krzyzosiak, W. J. Nucleotide sequence of miRNA precursor contributes to cleavage site selection by Dicer. Nucleic Acids Res. 43, 10939–10951 (2015).
Mcintyre, G. J., Yu, Y. H., Lomas, M. & Fanning, G. C. The effects of stem length and core placement on shRNA activity. BMC Mol. Biol. 12, 34 (2011).
Kim, D. H. et al. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat. Biotechnol. 23, 222–226 (2005).
Rose, S. D. et al. Functional polarity is introduced by Dicer processing of short substrate RNAs. Nucleic Acids Res. 33, 4140–4156 (2005).
Soifer, H. S. et al. A role for the Dicer helicase domain in the processing of thermodynamically unstable hairpin RNAs. Nucleic Acids Res. 36, 6511–6522 (2008).
Starega-Roslan, J. et al. Structural basis of microRNA length variety. Nucleic Acids Res. 39, 257–268 (2011).
Nguyen, T. D., Trinh, T. A., Bao, S. & Nguyen, T. A. Secondary structure RNA elements control the cleavage activity of DICER. Nat. Commun. 13, 2138 (2022).
Le, C. T., Nguyen, T. D. & Nguyen, T. A. Two-motif model illuminates DICER cleavage preferences. Nucleic Acids Res. 52, 1860–1877 (2024).
Le, T. N. Y., Le, C. T. & Nguyen, T. A. Determinants of selectivity in the dicing mechanism. Nat. Commun. 15, 8989 (2024).
Lee, Y.-Y., Kim, H. & Kim, V. N. Sequence determinant of small RNA production by DICER. Nature 615, 323–330 (2023).
Kim, H. et al. A mechanism for microRNA arm switching regulated by uridylation. Mol. Cell 78, 1224–1236 (2020).
Griffiths-Jones, S., Saini, H. K., Van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, 154–158 (2008).
Fromm, B. et al. MirGeneDB 2.0: the metazoan microRNA complement. Nucleic Acids Res. 48, D132–D141 (2020).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Weissmann, F. et al. biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. Proc. Natl Acad. Sci. USA 113, E2564–E2569 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads EMBnet. J. 17, 10–12 (2011).
Aronesty, E. Comparison of sequencing utility programs. Open Bioinform. J. 7, 1–8 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Selvaraj, J., Wang, L. & Cheng, J. CryoTEN: efficiently enhancing cryo-EM density maps using transformers. Bioinformatics 41, btaf092 (2025).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
DeLano, W. L. Pymol: an open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 40, 82–92 (2002).
Langmead, B. & Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Acknowledgements
We thank Y. Zhang and H. Ho for advice on sample preparation and data collection; S. Dang and his laboratory members at HKUST for providing the insect cell expression system and guidance on structural determination; K. H. Bui, L. Wang and M. D. Nguyen for advice on EM data processing and structural determination; T. Xie for providing RNA and DNA materials for the fly study; N. Kim for providing HCT116 cells; and all our laboratory members for discussions and contributions, particularly T. D. Nguyen and M. N. Le. M.K.N. is supported by a Hong Kong PhD Fellowship from the Research Grants Council. This research was funded by the Research Grants Council of Hong Kong (grant number 16103525).
Author information
Authors and Affiliations
Contributions
M.K.N., C.T.L. and T.A.N. conceived and designed the study. M.K.N. performed the biochemical and structural experiments and prepared the figures. All authors contributed to data analysis, interpretation of results and manuscript drafting. T.A.N. supervised the project and secured the necessary funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks En-Zhi Shen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Massively parallel assay and validation experiments on pre-mir-324.
a, SDS–PAGE of purified human DICER and Drosophila melanogaster Dcr-1. b, Library construction scheme for massively parallel dicing assays. Substrate libraries were generated by circular ligation product libraries used RA3/RA5 ligation. Reverse transcription and PCR amplification steps are shown for both substrate and product libraries. Detailed methodologies are described in the Methods section. c, Summary of variants recovered from massively parallel dicing of randomized pre-mir-324. Each group (A, U, G, C) has 64 theoretical variants; recovery was complete. d, Reproducibility of the massively parallel dicing assay. The global cleavage efficiency of DICER was calculated across three replicates for each group (A, U, G, and C). Green dots represent individual variants. High Pearson correlation (r) demonstrates assay reproducibility. e, Effect of the 3′-terminal nucleotide on DICER cleavage-site accuracy. Box plots represent the DC21 and DC22 cleavage accuracy across variants with different 3′-terminal nucleotides. f,i,l, Sequences and diagrams of pre-mir-324 variants. Green and red arrowheads mark cleavage at DC21 and DC22 positions, respectively. g,j,m, In vitro dicing assays for pre-mir-324 variants. h,k,n, Quantification of DC21 accuracy from three independent repeats of the assays in g,j,m. DC21 accuracy was calculated as DC21 products divided by total cleaved products.
Extended Data Fig. 2 Three-dimensional reconstruction and model building of DICER–shRNA complexes.
a, Summary of data collection and electron microscopy data processing workflow. b, Representative micrograph and 2D class averages of DICER–shRNA complexes in the dicing state. c, 3D electron density map of DICER–shRNA complexes. d, Local-resolution analysis of the density maps. The resolution ranges from ~2.5 Å in well-resolved regions to ~10 Å in flexible regions. e, GS-FSC curves. Map resolutions at the 0.143 criterion are 3.37 Å and 3.34 Å for two complexes. f, Particle orientation heat map showing the distribution of particle orientations used in the 3D reconstruction. g, Map-model fitting of the DICER–shRNA complexes. The density map (mesh form) is overlaid with the atomic model. Protein domains (platform, PAZ, connector helix, RIIIDa, RIIIDb, and dsRBD) and ssRNAs (26S-GU, 26S-UG) are coloured as in Fig. 2a,c.
Extended Data Fig. 3 Dicing of 26S shRNAs and characteristics of DICER–26S complexes.
a, In vitro dicing assay of 26S-UG and 26S-GU shRNAs by DICER. F1 and F3 are 22 nt, while F2 is 20 nt. b, Partial density of the helicase domain observed in the cryo-EM maps of DICER–shRNA complexes. The helicase (Hel1) is outlined in dashed circles, with adjacent domains (PAZ, platform, RIIIDa, RIIIDb, and dsRBD) labelled in distinct colours. c, Detailed view of the cleavage site (DC22) in the DICER–shRNA complexes. Calcium ions (green spheres) and the four catalytic acidic residues (sticks). Black arrowheads indicate hydrolysed phosphates at the cleavage sites. The cleavage sites align with the hydrolysis products observed from the in vitro dicing assay. d, Movement of the helix and β-sheet in the PAZ domain of DICER–shRNA complexes relative to DICER (PDB:7XW3), illustrated by RMSD values. e, Comparison of RNA expansion in pre-let-7a-1GYM (PDB: 7XW2) versus shRNAs (26S-UG and 26S-GU). The RNA duplex in pre-let-7a-1GYM expands near the catalytic centre, whereas the shRNAs remain compact. Alignment of AlphaFold3-predicted RNA (AF3-shRNAs) with experimental structures highlights structural differences. f, RNA conformation change analysis in PDB: 7XW2. The resolved RNA (blue) is compared with the AlphaFold3-predicted RNA (salmon). Calcium ions (red spheres) are positioned at the 3′ and 5′ catalytic centres. Measured distances from the calcium ions to the hydrolysed phosphates are displayed, showing structural differences in RNA alignment.
Extended Data Fig. 4 Identification of a 5′-G-favoured binding pocket.
a, 5′-end docking region and interacting models for terminal nucleotides in DICER–shRNA complexes. Top, DICER–26S-UG: surface cutaway highlighting the boundary loop (red), DC22 pocket, and modelled interactions of the 5′-terminal nucleotide U1 with residues R821 and R1003. Bottom, DICER–26S-GU: analogous views showing the boundary loop and DC21 pocket with the 5′-terminal nucleotide G1 positioned near residues D991 and H992. b, SDS–PAGE of purified DICER variants. c, Sequences and predicted secondary structures of pre-mir-208a and pre-mir-324 substrates. d, In vitro dicing of pre-mir-208a variants by WT and mutant DICER. Representative denaturing gels (DC21/DC22 labelled) and cleavage-site accuracy quantification from three replicates (right). Two-tailed, two-sample t-tests; significance: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; n.s., not significant. e, As in d, for pre-mir-324 variants. Gels (left/middle) and accuracy quantification (right). Same statistics as above. f, Cellular assay workflow. HCT116 DICER-knockout cells were transfected with pri-mir-517a (GU or UG) plus either WT DICER or the G-pocket mutant D991G-H992G. Small-RNA libraries from total RNA profiled DICER cleavage products. g, IsomiR distributions from the cellular assay. Bars show frequencies of DC21 (green), DC22 (red), and other isomiRs (grey) for each pre-mir-517a terminal pair and DICER variant. n = 3 biological replicates. Statistics: two-tailed, two-sample t-tests; **** p < 0.0001; n.s., not significant. h, In vitro dicing of pre-mir-517a with varied terminal pairs by DICER–TRBP. Left: representative denaturing gel (DC21/DC22 labelled). Right: cleavage-site accuracy for DC21 (green) and DC22 (red), n = 3. Statistics: two-tailed, two-sample t-tests; **** p < 0.0001; n.s., not significant.
Extended Data Fig. 5 Three-dimensional reconstruction and model building of the DICER(D991G/H992G)–26S-GU complex.
a, Overview of cryo-EM data collection and processing. Movies were motion-corrected, CTF-estimated, and subjected to particle picking, 2D/3D classification, non-uniform refinement, and CryoTEN map sharpening. The final reconstruction used 787,381 particles and reached 3.29 Å. b, Representative micrograph and selected 2D class averages of the dicing-state complex. c, Front and side views of the 3D density map showing the overall protein-RNA architecture. d, GS-FSC curve. The global resolution at the 0.143 criterion is 3.29 Å. e, Local-resolution estimates, ranging from ~2.5 Å in rigid regions to ~10 Å in flexible portions; front and side views illustrate spatial variation. f, Particle orientation heat map showing the distribution of particle orientations used in the 3D reconstruction. g, Map-model fit. The sharpened density (mesh) overlaid with the atomic model, with domains labelled: Platform, PAZ, Connector helix, RIIIDa, RIIIDb, and dsRBD; RNA (26S-GU, light blue). h, Close-up of the 5′-terminal G1 adjacent to the boundary loop and the G-pocket mutations (G991, G992; positions of D991/H992 mutated to Gly). i, Docking of the duplex relative to the boundary loop (red) and the DC21/DC22 pockets (open circles). The 26S-GU RNA is shown in blue. j, View of the 5′-end highlighting G1 and U62 positioned near the boundary loop. k, 3′-end pocket in the PAZ domain showing A64 and its contacts with surrounding residues.
Extended Data Fig. 6 Massively parallel dicing assay of Dcr-1 on pre-mir-324.
a, Sequence alignment of DICERs highlighting conserved residues in the 5′- and 3′-end binding pockets. 5′-pocket: R821, D991, H992, R1003; 3′-pocket: Y936, R937, Y956, Y971, Y972, K975, Y976 (shaded red). b, Number of recovered variants. The assay recovered all 256/256 expected variants, with equal contributions from each subgroup (A, U, G, C). c, Reproducibility of three independent dicing assays. Orange dots represent individual variants, and the Pearson correlation coefficient (r) reflects consistency between replicates. d, Cleavage sites (DC20–DC23) identified from sequencing results. e, Cleavage accuracy at DC21 and DC22 for pre-mir-324 groups, based on sequencing data. Left: RNAs grouped by 5′-nt; Right: RNAs grouped by 3′-nt. f, Similar effect of the 5′-nt on cleavage-site selection between DICER and Dcr-1. Variants were grouped by 5′-nt, and DC21 was calculated for each variant. g, Dicer cleavage across species for pre-miRNAs with different 5′-nt. Left: Workflow for cross-species analysis (34 species; MirGeneDB), assigning 5′-nt identity (A, U, G, C) to pre-miRNAs, predicting secondary structures, annotating major cleavage sites (DC21 or DC22), and comparing cleavage patterns. Right: Box-and-whisker plots show the proportion of pre-miRNAs cleaved at DC21 (top) or DC22 (bottom) by 5′-nt. Statistics: two-tailed, two-sample t-tests; **** p < 0.0001; n.s., not significant. h, Per-species distributions of cleavage outcomes. Top, proportion of pre-miRNAs cleaved at DC21 across species; 5′-G (green) versus others (grey). Bottom, proportion cleaved at DC22; 5′-U (red) versus others (grey).
Extended Data Fig. 7 RNA conformational changes during the dicing process.
a, Local structural comparisons between DICER–26S-GU (green) and DICER–26S-UG (salmon). Left, A-helix in RIIIDa (residues 1339–1363). Middle, A-helix in the PAZ domain (1017–1028). Right, boundary loop in the 5′-end pocket (986–998). Root-mean-square deviations (RMSDs) are indicated for each region. b, RNA helical trajectory differs between complexes. Superposition of DICER–26S-GU (green), DICER–26S-UG (red), and AF3-predicted shRNAs (grey) with selected inter-helical distances labelled, illustrating how RNA adjusts to facilitate precise cleavage. c, Close-ups show interactions involving R1855 and C19-C44 in DICER–26S-GU (left, green) and DICER–26S-UG (right, red). d, Coordination of 5′-nt and YCR motif in pre-miRNAs. Left, proportions of pre-miRNAs cleaved at DC21 (green), DC22 (red), or other sites (grey) grouped by 5′-nt (A, U, G, C; sample sizes below). Right, subsets enriched for YCR motifs show biases toward DC22 (top pie; n = 33) or DC21 (bottom pie; n = 42); remaining pre-miRNAs are categorized as no YCR or others.
Extended Data Fig. 8 Cryo-EM analysis of DICER–pre-mir-517a_GU.
a, Data processing workflow. From 4,024 + 5,460 + 4,760 movies, particles were subjected to motion correction, CTF estimation, curated exposure selection, blob/template picking, iterative 2D/3D classification, ab initio reconstruction, non-uniform refinement, and CryoTEN map sharpening. Two conformational groups were resolved: pre-dicing (1,272,937 particles; 3.00 Å) and dicing (475,650 particles; 3.21 Å). Representative 2D classes and GS-FSC curves are shown. b, Pre-dicing state. Composite map with segmented densities for domains: Helicase, DUF283, dsRBD, D-linker, Connector helix, Platform, PAZ, RIIIDa, RIIIDb, and the pre-mir-517a_GU. c, Dicing state. Overall map and individual domain segments as in b, highlighting rearrangements accompanying catalytic engagement. d, Secondary-structure model of pre-mir-517a_GU indicating terminal ends and DC21. e, Active-site views. Close-ups around the cleavage centre in RIIIDa/RIIIDb show coordination of catalytic residues and metal ions with nearby nucleotides. Residues are labelled; putative scissile-bond positions are indicated.
Extended Data Fig. 9 DICER end recognition, 3′-flanking contacts and conserved PAZ domain movement.
a, Model for 5′-end recognition by DICER. The 5′-nt engages two alternative binding pockets. Left, a 5′-G-favoured pocket where H992 provides a positive charge and D991 acts as an H-bond acceptor, forming a hydrogen-bonding web with purine edges. Right, a 5′-U-favoured pocket where R1003 contributes positive charge and R821 serves as an H-bond donor compatible with uridine. b, Y-cluster contacts with the 3′-overhang region differ between RNAs. Top, DICER bound to pre-let-7a-1GYM (orange). Overall view (left) and close-ups (middle, right) show defined interactions between the 3′-overhang region and the Y-cluster pocket. Bottom, DICER bound to pre-mir-517a_GU (green). Overall view (left) and close-ups (middle, right) highlight altered positioning of the 3′-overhang region near the Y-cluster. Question marks indicate weaker or ambiguous density/contacts relative to pre-let-7a-1GYM. c, Superposition of DICER homologues in dicing and apo states reveals PAZ-domain movement; apo conformations are shown in grey.
Extended Data Fig. 10 Contribution of R1855 to pre-miRNA dicing outcomes.
a, The presence of mWCU and YCR motifs in pre-mir-629 and pre-mir-517a. b, The in vitro dicing assays. Left, denaturing gels of in vitro dicing with WT DICER and R1855 mutants (R1855A/E/K) using shRNA substrates that variably include the YCR motif, mWCU motif, and/or extended stem. Right, schematics of substrates and quantification of cleavage accuracy (DC21, DC22, others) from the three independent repeats.
Supplementary information
Supplementary Figure (download PDF )
Raw gel images used in the main and Extended Data figures.
Supplementary Table 1 (download XLSX )
Cryo-EM data collection and model validation statistics.
Supplementary Table 2 (download XLSX )
Oligos for DNA cloning and mutagenesis.
Supplementary Table 3 (download XLSX )
Oligos for randomized pre-mir-324.
Supplementary Table 4 (download XLSX )
Oligos for synthesis of pre-miRNAs.
Supplementary Table 5 (download XLSX )
Pre-miRNA sequences and their associated YCR motifs.
Supplementary Table 6 (download XLSX )
Raw count of small-RNA sequencing data for pre-mir-517a.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ngo, M.K., Le, C.T. & Nguyen, T.A. DICER cleavage fidelity is governed by 5′-end binding pockets. Nature 653, 611–620 (2026). https://doi.org/10.1038/s41586-026-10211-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-026-10211-5