Abstract
Antibody–drug conjugates enable highly specific delivery of potent cytotoxics to biomarker-expressing cells. In parallel, advances in DNA circuitry and DNA–protein conjugates have allowed programmable integration of molecular inputs and signal amplification via hybridization chain reactions (HCRs). Here we present a system using affibody–DNA and aptamer–DNA conjugates to execute a Boolean logic operation on cell-surface biomarkers, resulting in amplified payload delivery using an HCR of DNA–drug conjugates. Proximity-induced assembly of the biomarker binders generates the initiator that triggers an HCR. The resulting assembly undergoes endocytosis, enabling controlled payload release of drugs conjugated to the DNA with cathepsin-cleavable linkers. We show that DNA–drug conjugates achieve targeted delivery with >100-fold amplification relative to the input biomarkers using fluorescence quantifications. We also identify payloads that strongly influence delivery efficiency and demonstrate delivery of different drug combinations. Finally, we show that biomarker-triggered HCRs can recruit generic antibodies. This modular technology enables tailored combinations of biomarker inputs and drug outputs toward more precise and personalized treatment.
Similar content being viewed by others
Main
Antibody–drug conjugates (ADCs) represent a cutting-edge approach in targeted cancer therapy, uniquely combining the specificity of monoclonal antibodies (mAbs) with the potent cytotoxicity of chemotherapeutic agents1,2,3,4. ADCs consist of three components: a monoclonal antibody specific to a tumor-associated antigen, a cytotoxic drug (payload) and a linker that binds the drug to the antibody. Notwithstanding their clinical success, ADC efficacy in solid tumors is often limited by poor tissue penetration. The large molecular weight of IgGs (approximately 150 kDa) presents a challenge for penetration through the tumor matrix5. The slow diffusion leads to a ‘binding site barrier’ where rapid target binding prevents deeper tumor penetration, leading to ADC accumulation at the periphery of the tumor6,7. These factors have prompted efforts to identify smaller proteins with antibody-like binding properties, so-called affinity proteins8. Among affinity proteins, affibodies stand out for their small size (<10 kDa) and high affinity and specificity9,10. Their utility has been demonstrated in multiple clinical programs, including applications in radioimaging11,12,13,14,15,16,17 and fluorescence-guided surgery18. An affibody dimer targeting HER2 conjugated to a cytotoxic agent has also been reported for selective delivery of the drug, in analogy to ADCs19. Another limitation of ADCs is the drug loading capacity, with most second-generation ADCs carrying only 2–4 drug molecules. The clinical success of trastuzumab deruxtecan (Enhertu)20,21,22 with a drug antibody ratio of 7–8, suggests that a higher payload can be beneficial. However, achieving higher payloads with other cytotoxic agents remains challenging without altering the properties of the antibody.
In parallel, engineered DNA circuits23,24,25 and hybridization chain reactions (HCRs)26 have emerged as powerful tools for biomarker detection and quantification, offering unprecedented sensitivity, specificity and versatility in molecular diagnostics. These nucleic acid-based systems exploit the inherent programmability of DNA, allowing for the design of complex, autonomous enzyme-free reactions that can amplify signals in the presence of target biomolecules23,24,25. In particular, HCRs enable an isothermal self-assembly of DNA hairpins into long, polymeric chains upon the introduction of a trigger sequence: typically connected to the recognition of a biomarker of interest, making it suitable for an amplified response in biological application27,28,29,30,31,32. In addition, engineered DNA circuits can incorporate logic gates, feedback loops, and cascades, allowing for sophisticated processing of molecular inputs to produce an output. However, for therapeutic application, the range of outputs has been limited33,34,35,36,37,38,39,40,41.
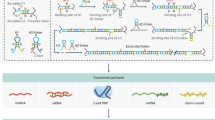
Herein we combine affibodies and aptamers with DNA to engineer an HCR in response to different biomarkers. While the DNA hairpins are cell impermeant, we demonstrate that the cell-surface-associated HCR product can undergo endocytosis. We further show that a cytotoxic drug conjugated to the hairpin via a cathepsin-cleavable linker induces cell-specific toxicity (Fig. 1). We also demonstrate that HCRs can be used to recruit antibodies to a target cell.
Drugs of interest are conjugated to DNA hairpins capable of HCRs. Specific binders to biomarkers of interest are conjugated to a split initiator such that the HCR takes place only if the two biomarkers are present in proximity.
Results
HCR and affibody–DNA conjugate
We initiated our investigation with affibodies (Z) targeting epidermal growth factor receptor (EGFR) (ZEGFR)42 and programmed death-ligand 1 (PD-L1) (ZPD-L1)17, two well-established biomarkers in cancer therapies. These markers have been extensively validated in therapeutic applications. The affibodies (58 amino acids (aa)) were synthesized via standard solid-phase peptide synthesis with an ε-azido-lysine at the C terminus of the affibody for subsequent conjugation to DNA through azide-DBCO coupling (see Supplementary Fig. 1 for explicit structures of all conjugates used in this study, methionine residues in affibodies were replaced by Nle to avoid oxidation products). This modification was positioned on a helix opposite to the target-binding interface to preserve binding specificity. A linker of 32 Å was used to provide flexibility and space for the affibody to accommodate the formation of the DNA structures necessary for HCR initiation and propagation. For the HCR, we used a previously reported system43 optimized for minimal leakage, rapid propagation and compatibility with split initiators (see Supplementary Fig. 2 for a diagram of all hybridization reactions and sequences). We validated the propagation of the HCR reaction using a full-length initiator loaded on streptavidin beads, comparing the fluorescence intensity between the addition of only the first hairpin labeled with fluorescein (H1Fl) to the same experiment with both hairpins (H1Fl + H2Fl, Extended Data Fig. 1a). The difference in integrated fluorescence corresponds to HCR propagation of 103 hairpins per initiator. Agarose gel analysis of the HCR initiated with the affibodies conjugated to the full-length initiator (ZEGFR-I or ZPD-L1-I, Extended Data Fig. 1b), showed that the conjugate did not interfere with the HCR and that minimal leakage was observed without initiation (lane 4) but complete consumption of hairpins was observed with the Z-I (lanes 5–8). Quantification of the HCR reaction on cells (A-431, EGFR+, Extended Data Fig. 1c) afforded a signal amplification of 132-fold, which was slightly higher than on beads (Extended Data Fig. 1a).
Logic-gated HCR
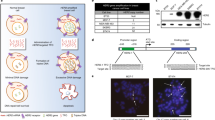
We examined a logic-gated response by splitting the initiator into two DNA strands (split initiator 1: S1 hybridized stem region of H1 and split initiator 2: S2 at toehold part), such that the HCR reaction can only proceed if both biomarkers (EGFR and PD-L1) are present in proximity leading to a three-way junction (3WJ) between S1 and S2 (Fig. 2a). Given that EGFR and PD-L1 do not dimerize spontaneously, the receptor density and membrane mobility need to account for proximity. An open question is the optimal number of nucleotides in the stem of the 3WJ. If the stem is too short, the transient interaction of the split initiators may be too short lived to initiate the HCR. Conversely, an excessively long stem could cause nonspecific initiation and result in AND-gate leakage. To address this, we tested three different stem lengths (4 nt, 6 nt and 8 nt) and evaluated HCR efficiency. To differentiate the outcome of HCR triggered by receptor proximity (AND-gated HCR) and untriggered HCR (AND-gate leakage), we compared two pairs of affibodies; ZPD-L1:ZEGFR (expected to initiate propagation) and ZTaq:ZEGFR (negative control). We measured the level of propagation by comparing the signal intensity of H1Fl alone and H1Fl + H2Fl as previously done with the full-length initiator. Each affibody is also labeled with a spectrally resolved fluorophore to validate its engagement with the biomarker (Cy3 for ZEGFR, ATTO-647N for ZPD-L1 or ZTaq, Fig. 2b). Quantification of fluorescence signal (Fig. 2c) indicated that a 4-nucleotide (nt) 3WJ was too short for efficient initiation, while the 8-nt was too long and led to more background signal than the 6-nt 3WJ. The use of a 6-nt stem was further validated with the alternative negative control of the AND-gate: ZTaq:ZPD-L1 (Fig. 2b). Collectively, the data support the ability to trigger the HCR conditionally on the presence of two biomarkers (AND gate) with an amplification of greater than tenfold. The lower level of amplification observed within the 30 min of HCR in this system versus the experiment performed with the full-length initiator (Fig. 2c versus Extended Data Fig. 1c) is a consequence of the lower concentration of effective initiator in the former (lower density of PD-L1 versus EGFR) and the possibility that not all available receptor pairs are close enough to initiate an HCR. Finally, the robustness of the two-biomarker AND-gated HCR was tested across different cell lines selected for the expression of both biomarkers (A-431, see Supplementary Fig. 3 for analysis of biomarkers), predominantly a single biomarker (U-251 and MDA-MB-468) or none (MCF-7). A strong amplification from HCR was observed only for A-431 (Fig. 2d,e for quantification); while a very weak HCR was observed on U-251 cells, consistent with the low level of EGFR expression observed in the Cy3 channel.
a, Schematic diagram of AND-gated response to two biomarkers using a split initiator HCR. b, Comparison of the HCR performance and specificity as a function of the 3WJ length (4 nt, 6 nt, 8 nt) in the split initiator on A-431 cells by microscopy imaging. c, Integrated fluorescein intensity across each cell membrane in the nine conditions represented in b plus an extra condition (ind: images not displayed in b comparing ZEGFR + ZPD-L1 and ZEGFR + ZTaq; AND +: logic gate fulfilled) (cells n = 255, 485, 295, 322, 749, 283, 63, 220, 534 and 306, respectively. The data were collected from 3, 4, 4, 3, 4, 4, 2, 3, 4 and 4 biological replicates, respectively). d, Specificity of AND-gated response across different cancer cell lines displaying different biomarker combinations. e, Integrated fluorescein intensity across each cell membrane in conditions in the six conditions represented in d (cells n = 322, 749, 37, 35, 48, and 35, respectively, 7 and 8 not quantified due to lack of signal in all channels. The data were collected from 3 (biological), 4 (biological), 3, 2, 3, 2, 3 and 3 technical replicates, respectively). Scale bar: 10 µm. Each hairpin was conjugated with one fluorescein molecule at the 5′ terminus. Fl: fluorescein channel; Merged channel: overlay of Fl, Cy3 and Atto647N. For graphs, the annotated folds were calculated based on the median. Graph legend: red line, median; black dashed line, first and third quartiles. Annotated P values calculated using a Kruskal–Wallis’ test plus a post hoc two-tailed Dunn’s test.
To test the generality of the approach, we used the same design to target a second pair of biomarkers: EGFR:PTK7. Based on the availability of a well-established DNA-aptamer that binds selectively to PTK7, ApPTK744,45, we investigated this biomarker combination (Fig. 3a) using the same workflow as with EGFR:PD-L1. As previously, a strong HCR was observed exclusively when the two biomarkers are targeted (ZEGFR:ApPTK7) but not with the control using either a nonbinding affibody (ZTaq:ApPTK7) or a scrambled aptamer (ZEGFR:Apscr) (Fig. 3b,e). The system was tested across a panel of cell lines selected for their expression of both EGFR and PTK7 (A-431 and MDA-MB-468) or a single biomarker (HCT-116 with weak expression of PTK7) or neither biomarker (MCF-7). Only the cell lines displaying both biomarkers yielded a statistically significant response with median amplification reaching 59-fold. A-431 cells yielded a stronger fluorescence signal than MDA-MB-468 (59-fold versus 13-fold, respectively), consistent with the lower expression level of PTK7 relative to A-431.
a, Schematic diagram of reaction performed in b–d and quantified in e–g, respectively. b, Microscopy imaging of response to EGFR and PTK7 on A-431, MDA-MB-468, HCT-116, MCF-7; quantified in e. c, Microscopy imaging of response to EGFR using split initiator requiring proximal binding to two EGFR on A-431, MDA-MB-468; quantified in f. d, Microscopy imaging of response to PTK7 using split initiator requiring proximal binding to two PTK7 on A-431, MDA-MB-468; quantified in g. e, Integrated fluorescein intensity across each cell membrane in the conditions in b (n = 105, 184, 105, 32, 129, 159, 74, 23 and 12 cells, respectively; Data collected from 3, 6, 3, 3, 3, 3, 5, 3, 3 and 4 technical replicates, respectively). f, Integrated fluorescein intensity across each cell membrane in the conditions in c (n = 409, 746, 381, 348, 190, 245, 160 and 130 cells (1 replica each) from left to right). g, Integrated fluorescein intensity across each cell membrane in the conditions in d (n = 173, 277, 226, 169, 92, 137, 239 and 114 cells (1 replicate) from left to right). Scale bar: 10 µm. Merged channel: overlay of Fl, Cy3 and Atto647N. For graphs, the annotated folds were calculated based on the median. Graph legend: red line, median; black dashed line, first and third quartiles. Annotated P values calculated using a Kruskal–Wallis test plus a post hoc two-tailed Dunn’s test. For the combination of tests, compare each column corresponding to the vertical black lines with the column indicated by the vertical red arrows.
Finally, we explored whether the split initiator system could also be used to report on a single biomarker using the same affibody, or aptamer, conjugated independently to either split initiator. This experiment differentiates itself to the one shown in Fig. 2c using a full-length initiator conjugated to the affibody in three ways. First, it requires sufficient receptor density and mobility for the split initiator system to work; second, the HCR product will be anchored to the cell surface by the interaction of two binders to the biomarker rather than one; third, the HCR can only be initiated at the cell surface on the 3WJ formation. As shown in Fig. 3c,d and quantified in Fig. 3f,g, A-431 and MDA-MB-468 both yielded a strong HCR response while the controls with a single nonbinding affibody (ZTaq) or nonbinding aptamer (Apscr) did not. The experiment targeting EGFR on A-431 afforded the highest level of amplification measured in this study (median of 217 propagation steps in the HCR). It is interesting to note that the amplification ratio between A-431 and MDA-MB-468 using PTK7 as single biomarker (67 versus 16, ratio of 4) or using the PTK7-EGFR dual biomarker (59 versus 13, ratio of 4.5) is different from using EGFR as a single biomarker (217 versus 111, ratio of 2) suggesting that PTK7 is the limiting biomarker in the dual biomarker amplification. To further probe the response, we pretreated A-431 cells with cetuximab to trigger internalization of EGFR in a dose-dependent manner with a linear correlation between EGFR staining and cetuximab (half-maximum inhibitory concentration (IC50) 1.23 nM, quantification supplied in Supplementary Fig. 4). Cells were then treated as shown in Fig. 3 and the level of HCR quantified (Extended Data Fig. 2). A clear correlation between the HCR signal and quantification of EGFR is observed.
Collectively, the data indicate that the design is general and enables HCRs triggered by two biomarkers using two affibodies, an affibody and an aptamer, or two aptamers.
DDC and targeted delivery
To assess the payload delivery following an HCR response, we used monomethyl auristatin E (MMAE) linked to either H1 or H2 (HM) with a cathepsin-cleavable linker, as used in ADCs46. In analogy to ADCs, where the drug loading is often adjusted to 3–4 drugs per mAb, the hairpins were also conjugated with 1 or 3 equivalents of MMAE (HM1 or HM3, Fig. 4a). The use of different hairpin combinations thus allows a gradient of drug loading, with H1Fl + H2M1 delivering the lowest loading and H1M1 + H2M3 delivering the highest loading. HCR performed with the maximal drug loading using ZEGFR-I for 30 minutes led to dramatic cell death (<8% cell viability after 48 h). The same experiment using ZTaq-I, which does not bind to any biomarker (Fig. 4b), showed no cytotoxicity. As a further control, cells treated with ZEGFR-I and H1Fl + H2Fl did not show growth inhibition, ruling out toxicity arising from EGFR binding or HCR product. To better gauge the differences between various drug loading combinations, we performed the cell viability assay at different concentrations of hairpins (Fig. 4b). Since the level of HCR amplification in the 30-min timeframe will be proportional to the concentration of the hairpin, a dose response should be obtained. To our surprise, the response between drug loading and cytotoxicity was not linear, rather, the presence of a hairpin with three MMAE (M3) was a determinant factor. Notably, H2M3 in combination with H1Fl (a total of 3 drugs per HCR cycle) is vastly more active (~13-fold) than H2M1 in combination with H1M1 (total of 2 drugs per HCR cycle) and 50-fold more active than H2M1 in combination with H1Fl (1 drug per HCR cycle). Based on these results, we turned to microscopy to verify whether a difference in HCR propagation could account for the higher toxicity of HCRs performed with H1Fl + H2M3, H1Fl + H2M1 and H1Fl + H1Fl (Fig. 4c). Quantification of the fluorescence showed that H2M3 yielded a statistically significant difference in fluorescence (roughly twofold) but not sufficient to account for the difference in cytotoxicity (roughly 50-fold). We also observed a striking difference in the fluorescence pattern with a less uniform fluorescence distribution across the membrane obtained with H2M3 versus H2M1, which we had not observed in any of the previous HCRs performed. Quantification of the coefficient of variation of the fluorescence signal across the cell perimeter showed a statistically significant difference in the fluorescence distribution between these two HCRs (that is, performed with H2M3 versus H2M1). A marginal difference in the coefficient of variation was observed between HCRs performed with H2M1 versus H2Fl (Fig. 4c). The same change in distribution was also evident in the Cy3 channel reporting on EGFR, suggesting that the HCR induces an aggregation at the cell surface that accounts for most of the cell-surface EGFR (see Supplementary Fig. 5 for a full set of images including controls included in the quantification and calculated coefficient of variation of Cy3 fluorescence–EGFR labeling). Similar results were also observed using the dual biomarker with split initiator system (Fig. 4d and Supplementary Fig. 6 for a full set of images including controls). It is interesting to note that in this case, the irregular fluorescence distribution across the cell perimeter is more pronounced with ApPTK7 than with ZEGFR (for quantitative analysis, see Supplementary Fig. 7) consistent with PTK7 being the limiting factor in the AND-gated HCR.
a, Schematic representation of experiments performed in b–d and structure of cytotoxic used in this study. b, Cell viability (A-431) as a function of hairpin concentration using different drug loading (data presented as mean ± s.d., from n = 4 biological replicates; data fitted to a four-parameter logistic, IC50 values presented as mean ± 95% CI; NA, not applicable). c, Microscopy imaging of EGFR-triggered HCR performed with different combination of hairpins on A-431 and quantification of fluorescein signal intensity and coefficient of variation across the cell membrane (n = 256, 421, 500 and 231 cells (1 replicate) from left to right; the coefficient of variation analysis is based on the same cells in each group). d, Microscopy imaging of EGFR-PTK7-triggered HCR performed with different combination of hairpins on A-431 and quantification of fluorescein signal intensity and coefficient of variation across the cell membrane (n = 363, 304 and 404 cells from left to right; the coefficient of variation analysis is based on the same cells in each group; data collected from 4, 5, 7 and 3 technical replicates, respectively). e, Cellular uptake (internalization) of HCR product under different treatment and quantification fluorescein intensity arising from HCR uptake (n = 274, 180 and 291 cells from top to bottom of the graph; data collected from 6, 4 and 6 technical replicates, respectively). Scale bars 10 µm. Merged channel: overlay of Fl, Cy3 and Atto647N/Cell Mask Deep Red and Hoechst. For graphs, the annotated folds were calculated based on the median. Graph legend: red line, median; black dashed line, first and third quartiles. NS, not significant, annotated P values calculated using a Kruskal–Wallis test plus a post hoc two-tailed Dunn’s test.
We next analyzed the internalized HCR product by treating cells with a nuclease (Benzonase) after 30 min of HCR reaction under nonpermeabilizing condition to remove all extracellular HCR, measuring the internalized fluorescein (Fig. 4e). Given that all the HCRs were performed with H1Fl, quantification of fluorescein is a direct measure of the internalization of the HCR product, that is, drug loading from H2. We observed a striking difference between the experiments using H2M3 versus H2M1 with a greater than tenfold higher uptake when H2M3 was used instead H2M1. Considering that the use of H2M3 delivers three times more drug than H2M1, this observation (more than tenfold higher delivery and threefold higher payload) accounts for the difference in cytotoxicity between HCRs performed H2M1 and H2M3 (IC50 in Fig. 4b). An HCR using H1Fl and H2M3 triggered by ZTaq-I (no cell-surface binding) or ZEGFR-I on MCF-7 cells (no EGFR expression) showed no internalization, indicating that the HCR product in solution does not provide sufficient cellular anchoring and, hence, cannot deliver a drug (Supplementary Fig. 8). We next investigated the exatecan derivative Dxd47 (Fig. 4a), another cytotoxic drug used in ADCs, notably trastuzumab. We performed HCR with H1Fl and H2D3, a hairpin with three copies of Dxd per hairpin using the same conjugation chemistry as used for H2M3. The combination H1Fl + H2D3 did not show the same change in fluorescence distribution as observed with H2M3 (Supplementary Fig. 9). Collectively, the data support a unique property to a hairpin functionalized with three MMAE conjugates. Comparing the physicochemical properties of the MMAE-cleavable-linker conjugate to Dxd-cleavable-linker conjugate, logP stands as the most notable difference: 2.14 for the MMAE-linker conjugate and 0.88 for the Dxd-linker conjugate (see Supplementary Information 5.6 for measurements). Given the logP value of the MMAE, it can be anticipated to partition favorably in the cellular membrane following the HCR initiated at the membrane. We speculate that this partitioning leads to an aggregation when three copies per hairpin are used, but not if a single copy per hairpin is used, and this aggregation facilitates, or even promotes, endocytosis. To characterize the delivery pathway, we next performed the experiment using H1 loaded with a fluorogenic cathepsin substrate (H1FQ) and H2M3 (Extended Data Fig. 3 and Supplementary Fig. 10 for controls showing that EGFR dimerization via the 3WJ is insufficient to promote internalization). We observed a gradual rise of florescence reaching a plateau at 5 h, consistent with the material of the HCR being trafficked through the endosome–lysosome (Extended Data Fig. 3b). It is interesting to note the time-release effect that accounts for drug exposure for >5 h following a 30 min HCR treatment. Pretreatment with a cathepsin inhibitor (K-777, Extended Data Fig. 3c) or an inhibitor of endocytosis (dynasore, Extended Data Fig. 3d) further supports an endocytosis and lysosomal trafficking. Finally, analysis of colocalization with lysosomes shows a high colocalization (Supplementary Fig. 11) corroborating this pathway as the major axis of delivery of HCR product. Finally, we next asked whether a simple peptide unrelated to MMAE but with analogous physicochemical properties might also trigger delivery. We used the sequence VVIGPF with a reduced C terminal alcohol and N-methylate amides or an inactive MMAF analog modified at the C terminal to prevent tubulin binding (Extended Data Fig. 4). HCR performed with both alternatives (H2PNM3 or H2IA3) showed the same cell-surface aggregation as with H2M3. Notably, treatment with H1M1 and H2PNM3 yielded a sevenfold better IC50 (28 nM) compared with H1M1 and H2M1, despite the latter containing twice the MMAE payload, demonstrating that nontoxic bystander peptides can be used for payload delivery.
We next investigated the fidelity of the AND-gated delivery of cytotoxic across different biomarker combinations and drug combinations (Fig. 5). Targeting EGFR dimer with the split initiator system (ZEGFR + ZEGFR), produced a dramatic cytotoxicity on A-431 (high expression level of EGFR), while no significant reduction of viability was observed otherwise (Fig. 5a). This result closely parallels the results obtained by microscopy quantification of fluorescence (Fig. 3c) but demonstrates that adding three copies of MMAE does not compromise the specificity of the HCR. The combined targeting of PTK7 and EGFR demonstrated an excellent selectivity for targeted cells (A-431: high biomarker expression; MDA-MB-468: moderate biomarker expression) over cell lines that are otherwise sensitive to the cytotoxic used (MMAE) but do not have significant expression levels of both biomarkers (HCT-116, MCF-7, HeLa-GFP) (Fig. 5b). Similarly, the combination of PD-L1 and EGFR showed a marked selectivity for the cell line that expresses both biomarkers (A-431) over others (U-251, MDA-MB-468, MCF-7) (Fig. 5c). In all the experiments, none of the controls using a nonbinding affibody or a scrambled sequence aptamer showed significant cytotoxicity compared to the target group. Finally, we asked whether the HCR could be used to deliver a combination of drugs (MMAE and Dxd, Fig. 5d) using H1D1 and H2M3. A dramatic difference in cytotoxicity was observed between the group using the appropriate biomarker binder versus control binder. Comparing this result to the same experiment using H1M1 and H2M3 (Fig. 5d versus Fig. 5b) supports an efficient delivery of the combination of cytotoxic agent. Thus, the benefits of H2M3 or H2PNM3 in terms of drug delivery can be harnessed to deliver a diversity of drugs through H1, irrespective of their lipophilicity.
a, A-431 viability in response to logic-gated HCR responding to two proximal EGFR using MMAE in DDC (n = 3 biological replicates; data presented as mean ± s.d.). b, Viability of different cell lines presenting different combinations of biomarkers in response to logic-gated HCR (PTK7 and EGFR) using MMAE in DDC (n = 3, 6, 5, 3, 3 and 6 biological replicates from left to right; data presented as mean ± s.d.). c, Viability of different cell lines presenting different combination of biomarkers to logic-gated HCR (PD-L1 and EGFR) using MMAE in DDC (n = 3, 3, 3, 2, 2, 7, 3 and 3 biological replicates from left to right; data presented as mean ± s.d.). d, A-431 viably in response to logic-gated HCR delivering two different cytotoxic (n = 2, and 5 biological replicates from left to right; Data presented as mean ± s.d.). e, Evaluation selective drug delivery in a mixed cell population (A-431: PTK7+ and EGFR+; HeLa-GFP: PTK7− and EGFR−) after 30 min of treatment followed by a change of media (t = 0) and 48 h posttreatment. f, Cell count plot as a function of time, color indicates the cell lines, numbers indicate the conditions as shown in e (n = 2 biological replicates). Scale bars: 100 µm, the overlayed images of blue (Hoechst 33342), green (GFP/FITC), yellow (Cy3) and red (Atto647N) are displayed. For DIC images, arrows show representative examples of a cell. Annotated P values calculated using ordinary one-way analysis of variance test with a post hoc two-tailed Tukey test. For the combination of tests, compare of each column corresponding to the vertical black lines with the column indicated by the vertical red arrows.
In addition, we evaluated the selectivity of AND-gated HCR-mediated drug delivery by microscopy using a mixed cell population (Fig. 5e). A-431 cells were cocultured with a HeLa cell line constitutively expressing tubulin-GFP (HeLa-GFP) and treated to an HCR conditional on PTK7 and EGFR, that is targeting A-431, using hairpins functionalized with either a cytotoxic (MMAE) or fluorophores (Fl). After a 30-min treatment, the media was exchanged, which removes the material that has not taken part in biomarker-mediated HCR, and imaged (t = 0). Since the aptamer targeting PTK7 (ApPTK7-S1) bears a red fluorophore (Atto647N), cells can be differentiated by the expression of GFP (HeLa) or staining of PTK7 (A-431) in both treatment groups. In the case where the HCR was performed with H1Fl + H2Fl, merged images showed a yellow signal due to the overlap of green (fluorescein isothiocyanate, FITC) and red (PTK7 staining) fluorescence. Cells can also be identified by differential interference contrast (DIC), A-431 showing a stronger response than HeLa (marked with colored arrow in Fig. 5e, see Supplementary Fig. 12 for individual channels and merged fluorescence and DIC images). After 48 hours, cells were treated with a nuclear stain (Hoechst) and the Atto647N-labeled PTK7 aptamer to facilitate cell counting and discrimination. Visually, the three control groups (no treatment, hairpin–drug conjugates without HCR initiation and AND-triggered HCR without drug conjugates) displayed a comparable proliferation of HeLa cells and A-431 cells, respectively. Notably, the group subjected to AND-gated HCR with MMAE-functionalized hairpins exhibited a dramatically reduced number of A-431 cells. Quantification indicates that HeLa proliferated at comparable rate in the drug treated group than in the control groups (Fig. 5f). These findings suggest minimal bystander effects from neighboring cells that internalized MMAE, highlighting the selectivity of the AND-triggered HCR approach.
Collectively, the results demonstrate that it is possible to use an HCR to deliver payloads into cells from DNA–drug conjugates (DDCs) in analogy to ADCs. The result also points to special features present in H2M3 that strongly influence the extent of intracellular delivery.
Protein recruitment as output of AND-gated HCR
In addition to delivering cytotoxic drugs through direct functionalization of the hairpins, we explored whether HCR could be used to recruit proteins48,49. We investigated the selective capture of antibodies on HCR products, given the bivalent interaction of monoclonal antibodies (mAbs) and the importance of appropriate ligand spacing to maximize bivalent engagement, and therefore full affinity48,49 (Fig. 6). Using a commercial humanized anti-fluorescein mAb, we first evaluated its affinity for a monovalent DNA conjugate (H1Fl) and compared it to DNA duplexes with FITC at both ends (bivalent) with 21-nt spacing (~70 Å) and 42-nt spacing (~140 Å). These distances correspond to the two possible geometric arrangements in the HCR, assuming a long persistence length and ignoring mesh-like arrangements. Notably, while the anti-FITC mAb showed very weak binding to H1Fl (surface plasmon resonance (SPR) could not be fitted to an accurate dissociation constant, KD), transitioning from the monovalent to the bivalent interaction with the DNA duplexes displaying two FITC afforded good affinity, with a slight preference for the 42-nt spacing (24.4 nM versus 33.8 nM for 21-nt spacing, Fig. 6a). Furthermore, the HCR product demonstrated an even greater gain in affinity (3.2 nM, a tenfold improvement relative to 21-nt bivalent duplex, KD are reported as the concentration of FITC). This enhanced affinity likely arises from an increased probability of rebinding following dissociation events at a given binding site (see Supplementary Fig. 13 for a full set of SPR measurements).
a, SPR sensorgrams and binding constants of different fluorescein-labeled (FITC) DNA assemblies (measured by SPR immobilizing the anti-FITC at low loading on SPR surface. Concentrations tested: 400 nM to 12.5 nM, twofold dilutions). ND, not determined. b, Schematic representation of experiments performed to obtain images shown in c. c, Microscopy images of A-431 treated with following treatment with ZEGFR-S2 and ApPTK7-S1 and different combinations of hairpin, mAb and protein G. d, Integrated intensity of DyLight 405 (protein G) in shown in c (n = 195, 367, 320, 363, 351 and 290 cells from left to right; data were collected from 3, 5, 5, 5, 5 and 4 technical replicates, respectively). Scale bars 10 µm; each hairpin was conjugated with one fluorescein molecule at the 5′ terminus. Merged channel: overlay of DyLight 405, Fl, Cy3 and Atto647N/Cell Mask Deep Red. For graphs, the annotated folds were calculated based on the median. Graph legend: red line, median; black dashed line, first and third quartiles. Annotated P values calculated using a Kruskal–Wallis test plus a post hoc Dunn’s test. RU, response units in SPR sensorgram.
To test whether this dramatic gain of affinity could be used to capture the anti-FITC mAb at the cell surface, we measured the recruitment of antibodies following an AND-gated HCR (EGFR and PTK7) with H1Fl + H2Fl (Fig. 6b). The experiment compared an anti-FITC mAb to an anti-GAL4 mAb using protein G-DyLight 405 conjugate for the detection of recruited mAbs (Fig. 6c). A bright DyLight 405 signal was observed exclusively on cells treated with all the components for the HCR and anti-FITC mAb. It is interesting to note that the fluorescence pattern across the cell surface (Fl-DNA channel and Atto647N-PTK7 channel) is different in the group treated with anti-FITC mAb + protein G (second column) from the group treated with Anti-GAL4 (third column) or protein G alone (fifth column). This resembles the pattern obtained with HCR using H2M3 (Fig. 4c,d) and suggests that mAb and protein G treatment lead to a reorganization of the HCR product. Given that mAb can bind across branches of HCR polymers and protein G can bind multiple Fc fragments50, we speculate that such treatment led to a reorganization of the HCR network. This reorganization is also visible in the channels of the aptamer targeting PTK7.
Discussion
The results collectively demonstrate that DNA assembly at the cell surface enables the selective delivery of multiple payloads using cleavable linkers with a proven track record in ADCs. The use of split initiators to trigger the HCR restricts the assembly exclusively to an AND logic-gated response, requiring the presence of a specific combination of biomarkers51. The 3WJ forming the split initiator required a duplex stability achieved with six base pairs for high fidelity AND-gated response and this held true for three biomarker combinations, suggesting it is general for tight binders as used in this study. Both miniproteins, such as affibodies, and aptamers can be used to achieve a combinatorial output for biomarker selectivity. We show that three copies of MMAE on a hairpin significantly enhance the internalization of the payload, and this strategy can be extended to codeliver multiple cytotoxic agents. There is considerable interest in ADCs with dual or multiple payloads to combat drug resistance or to achieve synergistic therapeutic effects52,53,54,55,56,57,58. The approach described here paves the way to a personalized strategy, where biomarker selection and payload combinations can be customized on demand. Several bispecific ADCs have been reported to enhance therapeutic performance relative to single-biomarker targeting and the first-in-class bispecific ADC (EGFR-HER3) is in clinical development59,60,61. The biomarkers used in the present study—EGFR, PD-L1 and PTK7—are either established therapeutic targets or are undergoing clinical evaluation62,63,64. Beyond ADCs, antibody-dependent cellular cytotoxicity has emerged as a promising therapeutic avenue. Our demonstration that monoclonal antibodies can be selectively recruited following an HCR suggests that HCR could also be leveraged to engage effector cells for HCR-dependent, cell-mediated cytotoxicity65 or targeted mucin degradation66. This work complement alternative approaches using exclusively proteins for logic-gated response to multiple antigens67,68,69.
One limitation of the present study is the use of unmodified DNA to perform the HCR, as DNA exhibits suboptimal metabolic stability in plasma (t1/2 2 h, see Supplementary Fig. 14 for gel analysis)70,71. A straightforward solution is to use mirror image DNA (l-DNA)72,73,74,75. Assembly of antibody fragments using l-DNA76 or PNA77 have also been reported. Another limitation is the use of an unmodified aptamer targeting PTK744,78. This cannot be resolved with l-DNA; however, various chemical modifications that enhance the metabolic stability of aptamers have been demonstrated79,80 and are of interest for hybridization-driven assemblies41,81,82.
Currently, ADCs in clinical use exhibit heterogeneity in both the position and number of payloads, with their engineering constrained by protein expression. While advances in conjugation chemistry have mitigated this heterogeneity4,83,84,85,86, the present system relies exclusively on materials produced by synthetic chemistry. This approach enables precise control over purification, modification and conjugation. In addition, the high solubility of DNA ensures consistent physicochemical properties across different hairpin–payload combinations, facilitating a modular strategy to access different combinations on demand.
In parallel, advances in radioimmunodiagnosis and radioimmunotherapy are converging toward pretargeting strategies that combine the tumor-specific recognition of tagged affinity proteins with medicinal radioisotopes for high-contrast imaging and high therapeutic indices. Unlike conventional radioimmunoconjugates, pretargeted approaches separate the tumor-recognition step from the payload delivery step, thereby minimizing normal tissue exposure87. Recent studies have highlighted the potential of hybridization-driven delivery of radioisotopes87,88,89, but the use of logic-gated response or amplification have not been harnessed thus far. Our findings suggest that additional specificity and imaging contrast could be achieved using the logic-gated HCR described here.
In this study, we demonstrated targeted drug delivery using DDCs actuated by HCR. By integrating affinity miniproteins (affibodies) and aptamers with DNA circuits, we engineered a biomarker-responsive system capable of amplifying therapeutic responses in a controlled and specific manner. The ability to achieve AND-gated HCR initiation, driven by the presence of two biomarkers or the density of a single biomarker, may advance selective drug delivery by benefiting from amplification and modularity of biomarker combination and combinatorial payload. We anticipate that the approach will contribute to the future of ADCs advances with smaller affinity proteins and high drug loading. Beyond drug delivery, we demonstrated the ability of HCR to recruit antibodies selectively, opening new avenues for immune-based therapies by integrating DNA nanotechnology with precision medicine.
Method
Synthesis of affibody–ssDNA or aptamer–ssDNA conjugates
To single-stranded DNA (ssDNA)-DBCO (1 equiv., in general 200 µM, 50 μl) in 0.8 M triethylammonium acetate buffer (TEAA, original from 1 M TEAA buffer, pH 7.0) were added affibody-N3 (1.5 equiv., in general 1 mM in 15 μl of H2O) or aptamer-N3 (0.5 equiv.) and 20% CH3CN (v:v, compared to the final volume). The solution was then incubated overnight protected from light at 1,000 rpm shaking at 37 °C. Reaction progress was monitored by matrix-assisted laser desorption/ionization. On completion, the reaction mixture was directly purified by reverse-phase high-performance liquid chromatography (HPLC) (mobile phase A: 0.1 M TEAA in H2O; mobile phase B: 0.1 M TEAA in CH3CN), followed by lyophilization. The lyophilized product was then desalted using a Nap-5 column (GE-17-0853-01), lyophilized again to provide the desired ligand–ssDNA conjugate as a pure lyophilized powder. For more detailed information and characterization, see Supplementary Information 9.2.
Synthesis of hairpin–drug conjugates
To hairpin-NH2 (500 µM in 0.5 M TEAA buffer, 1 equiv., roughly 120 μl) were added N,N-diisopropylethylamine (DIPEA) (2% v:v to TEAA buffer) and a linker bearing a NHS-activated acid and one or three copies of BCN (bicyclo[6.1.0]nonyne) (100 equiv. or 20 equiv., in dry dimethylformamide, roughly 100 μl). The solution was then incubated for 4 h with 1,000 rpm shaking at 37 °C. Subsequently, an additional equivalent amount of DIPEA and NHS-activated linker was added, and the reaction was allowed to continue shaking overnight under the same conditions. For hairpin-BCN, the excess organic reagents were removed by ethanol precipitation and the white precipitated was redissolved in 1 M TEAA, then purified by reverse-phase HPLC (mobile phase A: 0.1 M TEAA in H2O; mobile phase B: 0.1 M TEAA in CH3CN) followed by lyophilization. For hairpin-BCN3, reaction mixtures were purified by HPLC directly. The obtained hairpin-BCN or hairpin-BCN3 (1 equiv.) was then conjugated with prodrug-N3 (2 equiv. or 5 equiv.) by a strain promoted azide–alkyne cycloaddition reaction according to the similar procedure as ligand–ssDNA conjugates to provide the hairpin–drug conjugate. For more detailed information and characterization, see Supplementary Information 9.3.
HCR on beads
To 10 μl of initiator-loaded streptavidin beads (for preparation details, see Supplementary Information 3.1) in HCR+ buffer (5 mM MgCl2, 10 mM Na2HPO4, 140 mM NaCl in Milli-Q water, pH 7.4, +1 mg ml−1 BSA) were added 30 μl of HCR buffer+, 5 μl of 5 μM H1Fl and 5 μl of 5 μM H2Fl (hairpins were individually annealed at 95 °C for 5 min, then placed on ice 30 min before using). For the controls, HCR buffer+ was added to make the final reaction volume as 50 µl. The mixtures were incubated at 37 °C, in the dark with shaking (600 rpm) for 30 min. The beads were then washed twice with HCR buffer by centrifugation, then resuspend in HCR buffer and analyzed by microscopy (confocal SP8 with a ×63 oil immersion objective).
Procedure for HCR on cells with full-length initiator
A-431 cells were preseeded to μ-slide 18-well ibidiTreat (ref. no. 81816) 24 h before the experiments (10,000 per well). Hairpins (10 µM) in HCR buffer (5 mM MgCl2, 10 mM Na2HPO4, 140 mM NaCl in Milli-Q water, pH 7.4) were annealed (95 °C for 5 min, then placed on ice) 30 min before using. For HCR, cells were washed twice with HCR buffer++ (HCR buffer + 1 mg ml−1 BSA + 4.5 g l−1 glucose), then 25 nM ZEGFR-I in 100 μl of HCR buffer+ was added and incubated for 15 min (37 °C in a cell incubator). Subsequently, cells were washed twice with HCR buffer+. Then, freshly prepared solutions of 500 nM H1Fl or 500 nM H1Fl and H2Fl in 100 μl of HCR buffer+ were added to the corresponding wells, and the cells were placed into the incubator for an additional 30 min under 37 °C. After HCR, cells were washed three times by HCR buffer before imaging (confocal SP8 with ×63 oil immersion objective). For more detailed information, see Supplementary Information 3.3.
Procedure for HCR with split systems
Cells were seeded to μ-slide 18-well ibidiTreat 24 h before the experiments (for A-431 and MDA-MB-468 cells, 10,000 per well; for U-251 and MCF-7 cells, 5,000 per well; for HCT-116 cells, 2,500 per well). Hairpins (10 µM in HCR buffer) were annealed (95 °C for 5 min, then placed on ice) 30 min before using. Cells were washed twice with HCR buffer++, then a solution of freshly prepared 50 nM Ligand1-S1 and 50 nM Ligand2-S2 in 50 μl of HCR buffer++ were added and incubated for 15 min (37 °C in a cell incubator). Subsequently, a freshly prepared solution of 1,000 nM H1Fl and 1,000 nM H2Fl in 50 μl of HCR buffer++ was added (working concentration of H1 or H2: 500 nM), and the cells were placed into the incubator (37 °C) for an additional 30 min. After HCR, cells were washed three times by HCR buffer before imaging (confocal SP8 with a ×63 oil immersion objective). For more detailed information, see Supplementary Information 4.
Cytotoxicity assay of HCR with full-length initiator
A-431 cells were preseeded to a 96-well plate 24 h before the experiments (5,000 per well). Hairpins (10 µM in HCR buffer) were annealed (95 °C for 5 min, then placed on ice) 30 min before using. 25 nM ZEGFR-I or ZTaq-I in 100 μl of cell culture medium (FluoroBrite DMEM + 2 mM L-glutamine + 10% FCS + 1% penicillin–streptomycin) was added to the corresponding well and incubated for 15 min at 37 °C in a cell incubator. Then cells were washed twice with cell culture medium, before adding 100 μl of freshly prepared hairpin solution (H1 + H2) at the designed concentration gradients in cell culture medium (starting from 1,000 nM of each hairpin, in general, serial twofold dilution) and incubated for an additional 30 min at 37 °C in a cell incubator. After HCR, cells were washed 3 times with cell culture medium, then cultured for additional 48 h and cell viability was quantified using CCK-8 (Abbkine, ref. KTA1020-1000TEST). For more detailed information, see Supplementary Information 5.2.
Cellular uptake (internalization) study of HCR product
The HCR (concentration of each hairpin: 1,000 nM; A-431 cells were preseeded to μ-slide 18-well ibidiTreat) was conducted using the same procedure as above. After the 30 min of HCR and removal of excess hairpins, the cells were treated with a cocktail containing Hoechst, Benzonase (Merck, cat. no. 70746-4) and Cell Mask Deep Red (Invitrogen, cat. no. C10046) in 100 μl of HCR buffer at 37 °C for 15 min, then fixed with 4% PFA before imaging (SP8, ×63 oil immersion objective). For more detailed information and data with longer cellular uptake time after HCR, see Supplementary Information 5.5. For detailed information regarding the protocols for the trafficking of the HCR product via fluorophore-quencher probe, see Supplementary Information 5.8 and 5.9.
Cytotoxicity study of HCR with split systems
Cells were seeded to 96-well plate 24 h before the experiments (for A-431 cells, 5,000 per well; for MDA-MB-468 cells 7,500 per well; for U-251, MCF-7 and HeLa-GFP cells, 2,500 per well; for HCT-116 cells, 1,250 per well). Hairpins (10 µM in HCR buffer) were annealed (95 °C for 5 min, then placed on ice) 30 min before using. A solution of freshly prepared Ligand1-S1 and Ligand2-S2 in (50 nM) in 50 μl of cell culture medium was added to the corresponding wells and incubated for 15 min at 37 °C in a cell incubator. Then, 50 μl of hairpin solution (2,000 nM H1 + 2,000 nM H2, or others, depending on the specific experimental group design) in cell culture medium were added to the corresponding wells and incubated for 30 min at 37 °C in a cell incubator. After HCR, cells were washed 3 times with cell culture medium, then cultured for an additional 48 h. Cell viability was quantified using CCK-8. For more detailed information, see Supplementary Information 6.1.
Procedure for mAb treatment
The cells were pretreated with split-HCR or H1Fl with ZEGFR-S1 and ApPTK7-S2 according to ‘Procedure for HCR with split systems’. After HCR or H1Fl treatment for 30 min, cells were washed three times with cell culture medium. Subsequently, 100 μl of different protein solutions (25 nM Antibody and 100 nM Protein G-DyLight 405, 25 nM Anti-FITC or 100 nM Protein G-DyLight 405, depending on the design) in cell culture medium were added to the corresponding wells and incubated for an extra 30 min at 37 °C in a cell incubator. Finally, cells were washed three times with HCR buffer before imaging (nonfocal SP8 with a ×63 oil immersion objective). For more detailed information, see Supplementary Information 7.2.
Image analysis
All image analysis and quantification were performed through custom scripts in ImageJ stardist and ilastik and deposited on Zenodo.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available within this paper and its Supplementary Information. The raw data are available via Zenodo at https://doi.org/10.5281/zenodo.17293902 (ref. 90).
References
Beck, A., Goetsch, L., Dumontet, C. & Corvaïa, N. Strategies and challenges for the next generation of antibody drug conjugates. Nat. Rev. Drug Discov. 16, 315–337 (2017).
Fu, Z. W., Li, S. J., Han, S. F., Shi, C. & Zhang, Y. Antibody drug conjugate: the ‘biological missile’ for targeted cancer therapy. Signal Transduct. Tar. 7, 93 (2022).
Dumontet, C., Reichert, J. M., Senter, P. D., Lambert, J. M. & Beck, A. Antibody-drug conjugates come of age in oncology. Nat. Rev. Drug Discov. 22, 641–661 (2023).
Tsuchikama, K., Anami, Y., Ha, S. Y. Y. & Yamazaki, C. M. Exploring the next generation of antibody-drug conjugates. Nat. Rev. Clin. Oncol. 21, 203–223 (2024).
Xu, S. Internalization, trafficking, intracellular processing and actions of antibody-drug conjugates. Pharm. Res. 32, 3577–3583 (2015).
Bordeau, B. M., Yang, Y. J. & Balthasar, J. P. Transient competitive inhibition bypasses the binding site barrier to improve tumor penetration of trastuzumab and enhance T-DM1 efficacy. Cancer Res. 81, 4145–4154 (2021).
Rubahamya, B., Dong, S. & Thurber, G. M. Clinical translation of antibody drug conjugate dosing in solid tumors from preclinical mouse data. Sci. Adv. 10, 1894 (2024).
Gebauer, M. & Skerra, A. Engineered protein scaffolds as next-generation antibody therapeutics. Curr. Opin. Chem. Biol. 13, 245–255 (2009).
Löfblom, J. et al. Affibody molecules: engineered proteins for therapeutic, diagnostic and biotechnological applications. FEBS Lett. 584, 2670–2680 (2010).
Ståhl, S. et al. Affibody molecules in biotechnological and medical applications. Trends Biotechnol. 35, 691–712 (2017).
Orlova, A. et al. Tumor Imaging using a picomolar affinity HER2 binding affibody molecule. Cancer Res. 66, 4339–4348 (2006).
Feldwisch, J. et al. Design of an optimized scaffold for affibody molecules. J. Mol. Biol. 398, 232–247 (2010).
Andersson, K. G. et al. Feasibility of imaging of epidermal growth factor receptor expression with ZEGFR:2377 affibody molecule labeled with Tc using a peptide-based cysteine-containing chelator. Int. J. Oncol. 49, 2285–2293 (2016).
Da Pieve, C. et al. Efficient radiolabeling of Z affibody molecule for imaging of HER3 positive tumors. Bioconj. Chem. 27, 1839–1849 (2016).
Han, J. Y. et al. Therapeutic efficacy and imaging assessment of the HER2-targeting chemotherapy drug Z-pemetrexed in lung adenocarcinoma xenografts. Invest. New Drug 38, 1031–1043 (2020).
Alhuseinalkhudhur, A. et al. Kinetic analysis of HER2-binding ABY-025 Affibody molecule using dynamic PET in patients with metastatic breast cancer. EJNMMI Res. 10, 21 (2020).
Liang, Z. G., Hu, X. W., Hu, H. Y., Wang, P. & Cai, J. Novel small 99-Tc-labeled affibody molecular probe for PD-L1 receptor imaging. Front. Oncol. 12, 1017737 (2022).
de Souza, A. L. R. et al. Fluorescent affibody molecule administered at a microdose level labels EGFR expressing glioma tumor regions. Mol. Imaging Biol. 19, 41–48 (2017).
Serwotka-Suszczak, A. M. et al. A conjugate based on Anti-HER2 diaffibody and auristatin E targets HER2-positive cancer cells. Int. J. Mol. Sci. 19, 3676 (2018).
Tsurutani, J. et al. Targeting HER2 with trastuzumab deruxtecan: a dose-expansion, Phase I study in multiple advanced solid tumors. Cancer Discov. 10, 688–701 (2020).
Modi, S. et al. Trastuzumab deruxtecan in previously treated HER2-positive breast cancer. New Engl. J. Med. 382, 610–621 (2020).
Shitara, K. et al. Discovery and development of trastuzumab deruxtecan and safety management for patients with HER2-positive gastric cancer. Gastric Cancer 24, 780–789 (2021).
Yin, P., Choi, H. M. T., Calvert, C. R. & Pierce, N. A. Programming biomolecular self-assembly pathways. Nature 451, 318–322 (2008).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Dirks, R. M. & Pierce, N. A. Triggered amplification by hybridization chain reaction. Proc. Natl Acad. Sci. USA 101, 15275–15278 (2004).
Jung, C. & Ellington, A. D. Diagnostic applications of nucleic acid circuits. Acc. Chem. Res. 47, 1825–1835 (2014).
Koos, B. et al. Proximity-dependent initiation of hybridization chain reaction. Nat. Commun. 6, 7294 (2015).
Zhao, Y. X., Chen, F., Li, Q., Wang, L. H. & Fan, C. H. Isothermal amplification of nucleic acids. Chem. Rev. 115, 12491–12545 (2015).
Li, J., Green, A. A., Yan, H. & Fan, C. H. Engineering nucleic acid structures for programmable molecular circuitry and intracellular biocomputation. Nat. Chem. 9, 1056–1067 (2017).
Chang, X. et al. Construction of a multiple-aptamer-based DNA logic device on live cell membranes via associative toehold activation for accurate cancer cell identification. J. Am. Chem. Soc. 141, 12738–12743 (2019).
Zhang, C. et al. Cancer diagnosis with DNA molecular computation. Nat. Nanotechnol. 15, 709–715 (2020).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Kim, K. T., Angerani, S., Chang, D. L. & Winssinger, N. Coupling of DNA circuit and templated reactions for quadratic amplification and release of functional molecules. J. Am. Chem. Soc. 141, 16288–16295 (2019).
Emanuelson, C., Bardhan, A. & Deiters, A. DNA logic gates for small molecule activation circuits in cells. ACS Synth. Biol. 13, 538–545 (2024).
Prinzen, A. L. et al. Amplified self-immolative release of small molecules by spatial isolation of reactive groups on DNA-minimal architectures. Angew. Chem. Int. Ed. 59, 12900–12908 (2020).
Wang, D., Li, S. H., Zhao, Z. L., Zhang, X. B. & Tan, W. H. Engineering a second-order DNA logic-gated nanorobot to sense and release on live cell membranes for multiplexed diagnosis and synergistic therapy. Angew. Chem. Int. Ed. 60, 15816–15820 (2021).
Li, S. P. et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger. Nat. Biotechnol. 36, 258–264 (2018).
Dockerill, M. et al. Development of supramolecular anticoagulants with on-demand reversibility. Nat. Biotechnol. 43, 186–193 (2025).
Yu, H. X. et al. Aptameric hirudins as selective and reversible EXosite-ACTive site (EXACT) inhibitors. Nat. Commun. 15, 3977 (2024).
Tolmachev, V. et al. Imaging of EGFR expression in murine xenografts using site-specifically labelled anti-EGFR In-DOTA-Z Affibody molecule: aspect of the injected tracer amount. Eur. J. Nucl. Med. Mol. I. 37, 613–622 (2010).
Ang, Y. S., Qiu, X. Z., Yam, H. M., Wu, N. J. & Yung, L. Y. L. Enzyme-free and isothermal discrimination of microRNA point mutations using a DNA split proximity circuit with turn-on fluorescence readout. Biosens. Bioelectron. 217, 114727 (2022).
He, A. X. et al. Structure- based investigation of a DNA aptamer targeting PTK7 reveals an intricate 3D fold guiding functional optimization. Proc. Natl Acad. Sci. USA 121, E2404060121 (2024).
Shangguan, D. et al. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl Acad. Sci. USA 103, 11838–11843 (2006).
Doronina, S. O. et al. Development of potent monoclonal antibody auristatin conjugates for cancer therapy. Nat. Biotechnol. 21, 778–784 (2003).
Li, W. et al. Synthesis and evaluation of camptothecin antibody-drug conjugates. ACS Med. Chem. Lett. 10, 1386–1392 (2019).
Melkko, S., Dumelin, C. E., Scheuermann, J. & Neri, D. On the magnitude of the chelate effect for the recognition of proteins by pharmacophores scaffolded by self-assembling oligonucleotides. Chem. Biol. 13, 225–231 (2006).
Ranallo, S., Prévost-Tremblay, C., Idili, A., Vallée-Bélisle, A. & Ricci, F. Antibody-powered nucleic acid release using a DNA-based nanomachine. Nat. Commun. 8, 15150 (2017).
Kato, K. et al. Model for the complex between protein-G and an antibody Fc fragment in solution. Structure 3, 79–85 (1995).
Dannenfelser, R. et al. Discriminatory power of combinatorial antigen recognition in cancer T cell therapies. Cell Syst. 11, 215–228 (2020).
Maruani, A. et al. A plug-and-play approach to antibody-based therapeutics a chemoselective dual click strategy. Nat. Commun. 6, 6645 (2015).
Strop, P. et al. Site-specific conjugation improves therapeutic index of antibody drug conjugates with high drug loading. Nat. Biotechnol. 33, 694–696 (2015).
Levengood, M. R. et al. Orthogonal cysteine protection enables homogeneous multi-drug antibody-drug conjugates. Angew. Chem. Int. Ed. 56, 733–737 (2017).
Kumar, A. et al. Synthesis of a heterotrifunctional linker for the site-specific preparation of antibody-drug conjugates with two distinct warheads. Bioorg. Med. Chem. Lett. 28, 3617–3621 (2018).
Yamazaki, C. M. et al. Antibody-drug conjugates with dual payloads for combating breast tumor heterogeneity and drug resistance. Nat. Commun. 12, 3528 (2021).
Journeaux, T. & Bernardes, G. J. L. Homogeneous multi-payload antibody-drug conjugates. Nat. Chem. 16, 854–870 (2024).
Liu, B. et al. Antibody–bottlebrush conjugates unlock diverse payloads for targeted cancer therapy. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02772-z (2025).
Andreev, J. et al. Bispecific antibodies and antibody-drug conjugates (ADCs) bridging HER2 and prolactin receptor improve efficacy of HER2 ADCs. Mol. Cancer Ther. 16, 681–693 (2017).
Dong, Y. J. et al. Novel bispecific antibody-drug conjugate targeting PD-L1 and B7-H3 enhances antitumor efficacy and promotes immune-mediated antitumor responses. J. Immunother. Cancer 12, e009710 (2024).
Ma, Y. X. et al. BL-B01D1, a first-in-class EGFR-HER3 bispecific antibody- drug conjugate, in patients with locally advanced or metastatic solid tumours: a first-in-human, open-label, multicentre, phase 1 study. Lancet Oncol. 25, 901–911 (2024).
Liu, Y. S. et al. Glycoproteomic analysis of prostate cancer tissues by SWATH mass spectrometry discovers N-acylethanolamine acid amidase and Protein Tyrosine Kinase 7 as signatures for tumor aggressiveness. Mol. Cell. Proteomics 13, 1753–1768 (2014).
Maitland, M. L. et al. First-in-human study of PF-06647020 (cofetuzumab pelidotin), an antibody-drug conjugate targeting Protein Tyrosine Kinase 7, in advanced solid tumors. Clin. Cancer Res. 27, 4511–4520 (2021).
Dessaux, C., Ganier, L., Guiraud, L. & Borg, J. P. Recent insights into the therapeutic strategies targeting the pseudokinase PTK7 in cancer. Oncogene 43, 1973–1984 (2024).
Oostindie, S. C. et al. Logic-gated antibody pairs that selectively act on cells co-expressing two antigens. Nat. Biotechnol. 40, 1509–1519 (2022).
Pedram, K. et al. Design of a mucin-selective protease for targeted degradation of cancer-associated mucins. Nat. Biotechnol. 42, 597–607 (2024).
Gunnoo, S. B. et al. Creation of a gated antibody as a conditionally functional synthetic protein. Nat. Commun. 5, 4388 (2014).
Lajoie, M. J. et al. Designed protein logic to target cells with precise combinations of surface antigens. Science 369, 1637–1643 (2020).
Kofoed, C. et al. Programmable protein ligation on cell surfaces. Nature 645, 793–800 (2025).
Dovgan, I. et al. On the use of DNA as a linker in antibody-drug conjugates: synthesis, stability and potency. Sci. Rep. 10, 7691 (2020).
Mrcher, A., Nijenhuis, M. A. D. & Gothelf, K. V. A wireframe DNA cube: antibody conjugate for targeted delivery of multiple copies of monomethyl auristatin E. Angew. Chem. Int. Ed. 60, 21691–21696 (2021).
Urata, H., Shinohara, K., Ogura, E., Ueda, Y. & Akagi, M. Mirror-image DNA. J. Am. Chem. Soc. 113, 8174–8175 (1991).
Damha, M. J., Giannaris, P. A. & Marfey, P. Antisense L/D-oligodeoxynucleotide chimeras—nuclease stability, base-pairing properties, and activity at directing ribonuclease-H. Biochemistry 33, 7877–7885 (1994).
Schaudy, E., Somoza, M. M. & Lietard, J. l-DNA duplex formation as a bioorthogonal information channel in nucleic acid-based surface patterning. Chem. Eur. J. 26, 14310–14314 (2020).
Shearer, V., Yu, C. H., Han, X. & Sczepanski, J. T. The clinical potential of L-oligonucleotides: challenges and opportunities. Chem. Sci. 15, 18239–18258 (2024).
Zhou, L. J. et al. Self-assembled L-DNA linkers for rapid construction of multi-specific antibody-drug conjugates library. Angew. Chem. Int. Ed. 62, e202302805 (2023).
Kazane, S. A. et al. Self-assembled antibody multimers through peptide nucleic acid conjugation. J. Am. Chem. Soc. 135, 340–346 (2013).
Shangguan, D. H., Cao, Z. H. C., Li, Y. & Tan, W. H. Aptamers evolved from cultured cancer cells reveal molecular differences of cancer cells in patient samples. Clin. Chem. 53, 1153–1155 (2007).
Kratschmer, C. & Levy, M. Effect of chemical modifications on aptamer stability in serum. Nucleic Acid Ther. 27, 335–344 (2017).
Han, Y. Q. et al. Molecular programming design of glyconucleic acid aptamer with high stability. Adv. Sci. 12, 2408168 (2024).
Zhang, Q. et al. Aptamer-based nongenetic reprogramming of CARs enables flexible modulation of T cell-mediated tumor immunotherapy. ACS Cent. Sci. 10, 813–822 (2024).
Skaanning, M. K. et al. Self-assembly of ultrasmall 3D architectures of (L)-acyclic threoninol nucleic acids with high thermal and serum stability. J. Am. Chem. Soc. 146, 20141–20146 (2024).
Chalker, J. M., Bernardes, G. J. L., Lin, Y. A. & Davis, B. G. Chemical modification of proteins at cysteine: opportunities in chemistry and biology. Chem. Asian J. 4, 630–640 (2009).
Bernardim, B. et al. Stoichiometric and irreversible cysteine-selective protein modification using carbonylacrylic reagents. Nat. Commun. 7, 13128 (2016).
Walsh, S. J. et al. Site-selective modification strategies in antibody-drug conjugates. Chem. Soc. Rev. 50, 1305–1353 (2021).
Ochtrop, P. et al. Compact hydrophilic electrophiles enable highly efficacious high DAR ADCs with excellent PK profile. Chem. Sci. 14, 2259–2266 (2023).
Honarvar, H. et al. Feasibility of affibody molecule-based PNA-mediated radionuclide pretargeting of malignant tumors. Theranostics 6, 93–103 (2016).
Oroujeni, M. et al. Affibody-mediated PNA-based pretargeted cotreatment improves survival of trastuzumab-treated mice bearing HER2-expressing xenografts. J. Nucl. Med. 63, 1046–1051 (2022).
Yan, J. X. et al. Radionuclide therapy of bevacizumab-based PNA-mediated pretargeting. Nucl. Med. Commun. 45, 901–909 (2024).
Chen, S.-K. et al. DNA-drug conjugates (DDCs) for computed delivery. Zenodo https://doi.org/10.5281/zenodo.17293902 (2026).
Acknowledgements
This work was supported, in part, by the Swiss National Science Foundation (188406,219316, N.W.), NCCR Chemical Biology (grant no. 185898, N.W.) and EMBO (postdoctoral fellowship grant no. ALTF 1015-2020 to E.E.W.). We thank D. Danylchuk for his assistance with the synthesis of the fluorophore-quencher probe. We thank D. Moreau and V. Mercier for their assistance in microscopy images acquisition and analysis.
Author information
Authors and Affiliations
Contributions
All the synthetic materials used in this study were designed and prepared by S.-K.C., M.L.-T., F.R. and E.E.W. Cell-based experiments were designed and performed by S.-K.C., M.L.-T., F.R. Statistical analysis was performed by S.-K.C., J.C.G. and F.R. SPR measurements were performed by S.-K.C. and M.D. Physical characterization of synthetic products were performed by S.-K.C., M.L.-T., F.R., E.E.W., M.D. and S.B. Data analysis was performed by all authors. N.W. conceived and supervised the work. The paper was written by S.-K.C., M.L.-T., F.R. and N.W. S.-K.C., M.L.-T. and F.R. had equal contributions to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Gonçalo Bernardes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Validation and quantification of HCR.
a, Schematic diagram and microscopy images of HCR on beads (fluorescein channel and bright field merged channel, scale bar 20 µm); Graph: integrated fluorescein intensity across each bead surface after signal propagation (HCR) or under no propagation (H1-only) conditions; Median-fold between conditions: 103x (nHCR = 55 beads; nH1=50 beads, from two independent experimental replicates). The P value was calculated using a Mann-Whitney U-test. b, Agarose gel electrophoresis analysis of HCR initiated by affibody-DNA conjugates (ladder in lane 1, 50bp ladder ref: 13525 from Norgen Biotek). c, Schematic representation and microscopy imaging on A-431 cells after signal propagation (HCR) or under no propagation (H1-only) conditions. Green channel images the fluorescein in DNA; yellow channel images the Cy3 conjugated to ZEGFR-I. Scale bar 10 µm; Graph: integrated fluorescein intensity across each cell-membrane. Median fold between conditions: 132x (nHCR = 477 cells; nH1= 362 cells, from 5 and 4 technical replicates respectively). Graph legend: red line (median); black dashed line (1st and 3rd quartile). The P value was calculated using a Mann-Whitney U-test. See Supplementary Fig. 1 for explicit structures of synthetic molecules.
Extended Data Fig. 2 HCR response in A-431 cells pre-treated with varying concentration of cetuximab to reduce available EGFR density.
a, Fluorescence signal on A-431 cells of either ZEGFR-S1 and ZEGFR-S2 or HCR at increasing concentration of cetuximab. b, Quantification of split initiator and HCR signal over increasing concentrations of Cetuximab fitted to a four-parameters logistic. Vertical bars represent the standard deviation of the mean response (n = 3 biological replicates). c, Single cell fluorescence correlation plot between the split initiator density and HCR output at three different concentrations of cetuximab. The higher slope observed in the absence of cetuximab (higher EGFR density) than at intermediate of high cetuximab concentration reflects a higher probability of HCR initiation with higher receptor density (all HCR performed at the same hairpin concentration). For all panels, in yellow Cy3 signal corresponding to ZEGFR-S1 and ZEGFR-S2. In green, FITC signal corresponding to H1Fl and H2Fl (HCR). In cyan, nuclei stain with Hoechst. Scale bar = 10 μm. See Supporting Information section 4.1. for a detailed protocol.
Extended Data Fig. 3 Kinetics of linker cleavage and trafficking of HCR product.
a, Schematic representation of the cellular uptake investigation. In brief, by functionalizing H2 with a fluorophore-quencher (ATTO 647N-Iowa black RQ) system connected by a cathepsin-cleavable linker, we designed a fluorogenic probe (H1FQ) responsive to the activity of cathepsin B. H1FQ was implemented in the circuitry, and the delivery route was investigated either by observing an increase in the fluorescence signal overtime, or its decrease upon treatment with dynamin or cathepsin inhibitors. b, On the left, microscopy images of A-431 cells treated with ZEGFR-S1, ZEGFR-S2, H1FQ and H2M3 at 0.5, 2, 4, and 6 h. On the right, a graph showing the relative fluorescence increase normalized by the intensity at the first timepoint (30 min) throughout 10 h of imaging. Vertical bars represent the standard deviation of the mean fluorescent signal (n = 3 biological replicates). c, On the left, microscopy images of A-431 cells treated with ZEGFR-S1, ZEGFR-S2, H1FQ and H2M3 after one hour of incubation with K-777. The images show a difference in the fluorogenic probe activation between treated and untreated cells at 0.5, 2, 4, and 6 h. On the right a graph showing the decrease in fluorescence signal upon cathepsin inhibition achieved by increasing the doses of K-777 (4 h timepoint). Vertical bars represent the standard deviation of the mean fluorescent signal (n = 3 biological replicates). P values were calculated using ordinary one-way ANOVA d. On the left, microscopy images of A-431 cells treated with ZEGFR-S1, ZEGFR-S2, H1FQ, H2M3, and Dynasore at 0.5, 1.5, 2, and 2.5 h. On the right a graph showing the decrease in fluorescence signal upon dynamin inhibition at different timepoints after the HCR. Vertical bars represent the standard deviation of the mean fluorescent signal (n = 3 biological replicates). For all images, red is used to denote ATTO 647N signal (fluorogenic cathepsin probe), cyan for nuclei staining with Hoechst; For all images: Scale bar 10 µm.
Extended Data Fig. 4 Alternative hairpin functionalization with non-cytotoxic peptide for efficient cellular delivery.
a, Schematic representation of experiments performed in panels b-c and structure of caged compounds used in this study. b, Cell viability (A-431) as a function of hairpin concentration using different drug loading. Data presented as mean ± s.d., from n = 3 biological replicates. Data fitted to a four-parameters logistic, IC50 values presented as mean ± 95% CI. (n.a.: not applicable). c, Microscopy imaging of EGFR-triggered HCR performed with different combination of hairpins on A-431 and quantification of fluorescein signal intensity and coefficient of variation across the cell membrane (n = 233, 243, 89, 229 and 267 cells from left to right; the coefficient of variation analysis is based on the same cells in each group). The data was collected from 5, 5, 3, 6 and 6 technical replicates from left to right. For all images: Scale bar 10 µm; Fl: fluorescein channel; Merged channel: Overlay of Fl, Cy3 and Atto647N/Cell Mask Deep Red and Hoechst. For Graphs: The annotated folds were calculated based on the median; Graph legend-red line (median), black dashed line (1st and 3rd quartile); annotated P values calculated using a Kruskal-Wallis test plus a post-hoc two-tailed Dunn’s test.
Supplementary information
Supplementary Information (download PDF )
Supplementary figures, tables, physical characterization of compounds and synthesis method.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, SK., López-Tena, M., Russo, F. et al. DNA–drug conjugates enable logic-gated drug delivery amplified by hybridization chain reactions. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-026-03044-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-026-03044-0