Abstract
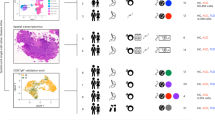
Fibrosis is a major complication of Crohn’s disease (CD) marked by excess deposition of extracellular matrix, leading to stricturing and functional impairment. As mechanistic characterization and therapeutic options are lacking, we paired single-cell and spatial transcriptomics in 61 samples from 21 patients with CD and 10 patients without inflammatory bowel disease (IBD). Intestinal strictures were characterized by increased immune cells, including IgG+ plasma cells, CCR7-hi CD4+ T cells and inflammatory fibroblasts. Spatial transcriptomics showed that key subsets colocalize within diseased tissues and identified additional populations such as interstitial cells of Cajal and enteric neurons. Furthermore, we mapped gene expression onto intestinal biogeography, finding that known genetic risk loci are enriched within discrete spatial modules, defined by the presence of inflammatory fibroblasts and lymphoid follicles. Altogether, our datasets chart the key transcriptomic and cellular networks in stricturing CD and highlight the spatial organization of multicellular genetic risk factors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Single-cell and spatial transcriptomics datasets generated in this study were deposited in the controlled access repository Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP2959/human-cd-fibrosis-study-using-single-cell-and-spatial-data), dbGAP with accession phs003943.v1.p1 (http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs003943.v1.p1) and Zenodo (https://zenodo.org/records/14509802?preview=1&token=eyJhbGciOiJIUzUxMiJ9.eyJpZCI6ImQ4NmNjYzliLTI4MGItNGYyZi04ODM3LTRjMGNiMzgyYzkzNyIsImRhdGEiOnt9LCJyYW5kb20iOiI2YzFhOTNkZjg0MzljZTFkZTZhYjc5MmU2MjNmNjk0YSJ9.t978Tj9Z5deFC_C7yDytHe_QxS0wXmgT_chPqwZkm0XDXp0Q3zYyPnhfRGa75fk_dIF6muaLJWUOwOiYhjBJiA). The reference genome used for Cell/SpaceRanger alignments can be accessed at https://cf.10xgenomics.com/supp/cell-exp/refdata-gex-GRCh38-2020-A.tar.gz.
Code availability
All original code is available on Zenodo at https://doi.org/10.5281/zenodo.15212391 (ref. 49).
References
Lewis, J. D. et al. Incidence, prevalence, and racial and ethnic distribution of inflammatory bowel disease in the United States. Gastroenterology 165, 1197–1205 (2023).
Graham, D. B. & Xavier, R. J. Pathway paradigms revealed from the genetics of inflammatory bowel disease. Nature 578, 527–539 (2020).
Kugathasan, S. et al. Prediction of complicated disease course for children newly diagnosed with Crohn’s disease: a multicentre inception cohort study. Lancet 389, 1710–1718 (2017).
D’Alessio, S. et al. Revisiting fibrosis in inflammatory bowel disease: the gut thickens. Nat. Rev. Gastroenterol. Hepatol. 19, 169–184 (2022).
Smillie, C. S. et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 178, 714–730 (2019).
Martin, J. C. et al. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell 178, 1493–1508 (2019).
Elmentaite, R. et al. Cells of the human intestinal tract mapped across space and time. Nature 597, 250–255 (2021).
Kong, L. et al. The landscape of immune dysregulation in Crohn’s disease revealed through single-cell transcriptomic profiling in the ileum and colon. Immunity 56, 444–458.e5 (2023).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Mukherjee, P. K. et al. Stricturing Crohn’s disease single-cell RNA sequencing reveals fibroblast heterogeneity and intercellular interactions. Gastroenterology 165, 1180–1196 (2023).
Levantovsky, R. M. et al. Multimodal single-cell analyses reveal mechanisms of perianal fistula in diverse patients with Crohn’s disease. Med 5, 886–908 (2024).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Tsukui, T., Wolters, P. J. & Sheppard, D. Alveolar fibroblast lineage orchestrates lung inflammation and fibrosis. Nature 631, 627–634 (2024).
Buechler, M. B. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021).
Li, S. et al. An integrated map of fibroblastic populations in human colon mucosa and cancer tissues. Commun. Biol. 5, 1326 (2022).
Chalkidi, N., Paraskeva, C. & Koliaraki, V. Fibroblasts in intestinal homeostasis, damage, and repair. Front. Immunol. 13, 924866 (2022).
Jasso, G. J. et al. Colon stroma mediates an inflammation-driven fibroblastic response controlling matrix remodeling and healing. PLoS Biol. 20, e3001532 (2022).
Korsunsky, I. et al. Cross-tissue, single-cell stromal atlas identifies shared pathological fibroblast phenotypes in four chronic inflammatory diseases. Med 3, 481–518 (2022).
Garrido-Trigo, A. et al. Macrophage and neutrophil heterogeneity at single-cell spatial resolution in human inflammatory bowel disease. Nat. Commun. 14, 4506 (2023).
Cadinu, P. et al. Charting the cellular biogeography in colitis reveals fibroblast trajectories and coordinated spatial remodeling. Cell 187, 2010–2028 (2024).
Schneider, S. et al. Single nucleus sequencing of human colon myenteric plexus–associated visceral smooth muscle cells, platelet derived growth factor receptor α cells, and interstitial cells of Cajal. Gastro Hep Adv. 2, 380–394 (2023).
Suchanski, J. et al. Podoplanin increases the migration of human fibroblasts and affects the endothelial cell network formation: a possible role for cancer-associated fibroblasts in breast cancer progression. PLoS ONE 12, e0184970 (2017).
Waizenegger, J. S. et al. Role of growth arrest-specific gene 6-Mer axis in multiple myeloma. Leukemia 29, 696–704 (2015).
Abdollahi, P., Köhn, M. & Børset, M. Protein tyrosine phosphatases in multiple myeloma. Cancer Lett. 501, 105–113 (2021).
Drokhlyansky, E. et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622 (2020).
De Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Wynn, T. A. & Ramalingam, T. R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 18, 1028–1040 (2012).
Burger, A. et al. Towards a clinically-based common coordinate framework for the human gut cell atlas: the gut models. BMC Med. Inform. Decis. Mak. 23, 36 (2023).
Baumgart, D. C. & Le Berre, C. Newer biologic and small-molecule therapies for inflammatory bowel disease. N. Engl. J. Med. 385, 1302–1315 (2021).
Friedrich, M. et al. IL-1-driven stromal–neutrophil interactions define a subset of patients with inflammatory bowel disease that does not respond to therapies. Nat. Med. 27, 1970–1981 (2021).
Tian, L., Chen, F. & Macosko, E. Z. The expanding vistas of spatial transcriptomics. Nat. Biotechnol. 41, 773–782 (2023).
Lako, A. & Rodig, S. HTAPP_OCT embedding of fresh tumor tissue. protocols.io dx.doi.org/10.17504/protocols.io.4nzgvf6 (2019).
Li, B. et al. Cumulus provides cloud-based data analysis for large-scale single-cell and single-nucleus RNA-seq. Nat. Methods 17, 793–798 (2020).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Cham, L. B. et al. Single cell analysis reveals a subset of cytotoxic-like plasmacytoid dendritic cells in people with HIV-1. iScience 26, 107628 (2023).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Chu, T., Wang, Z., Pe’er, D. & Danko, C. G. Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat. Cancer 3, 505–517 (2022).
Liu, J. et al. Jointly defining cell types from multiple single-cell datasets using LIGER. Nat. Protoc. 15, 3632–3662 (2020).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Ramilowski, J. A. et al. A draft network of ligand–receptor-mediated multicellular signalling in human. Nat. Commun. 6, 7866 (2015).
Baker, E. A. G., Schapiro, D., Dumitrascu, B., Vickovic, S. & Regev, A. In silico tissue generation and power analysis for spatial omics. Nat. Methods 20, 424–431 (2023).
Kong, L. & Smillie, C. S. Single-cell and spatial transcriptomics of stricturing Crohn’s disease highlights a fibrosis-associated network. Zenodo https://doi.org/10.5281/zenodo.15212391 (2025).
Acknowledgements
We thank T. Reimels for editorial assistance; A. Slamin, S. Maldonado and the Broad Genomics Platform for assistance with scRNA-seq experiments; L. Besse and C. McCabe for logistical support; and D. Chafamo, Y. Klindziuk, W. Hwang and S. Fleming for assistance with data processing. This work was funded by the National Institutes of Health (P30 DK043351 (to R.J.X. and C.S.S.) and RC2 DK135492 (to R.J.X.)), the Leona M. and Harry B. Helmsley Charitable Trust (to R.J.X.) and the Pew Biomedical Scholars Award (to C.S.S.). This publication is part of the Gut Cell Atlas Consortium funded by the Leona M. and Harry B. Helmsley Charitable Trust and is supported by a grant from Helmsley to the Broad Institute (to R.J.X.; www.helmsleytrust.org/gut-cell-atlas/).
Author information
Authors and Affiliations
Contributions
S.S., D.B.G. and R.J.X. conceptualized the project. L.K. performed single-cell analyses with assistance from J.D. S.S. and G.T.C. processed samples and performed scRNA-seq under supervision from T.M.D., D.B.G. and J.D. A.S. and J.L. performed spatial transcriptomics experiments under supervision from T.M.D. V.T. analyzed spatial transcriptomics datasets with assistance from S.G. and under the supervision of C.S.S. A.R.S. provided histological annotations and guidance. H.K., C.C., L.B., R.R., A.N.A. and R.H. provided clinical samples and information. S.W.T. coordinated consent, sample and metadata acquisition with assistance from M.E.K. under supervision from H.L. E.J.C. processed raw sequencing data under supervision from C.B.M.P. L.K., J.D., C.S.S. and R.J.X. wrote the paper with input from all authors. R.J.X. obtained funding and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
R.J.X. is cofounder of Jnana Therapeutics, Scientific Advisory Board member at Nestlé, Magnet BioMedicine and Arena BioWorks, and Board Director at MoonLake Immunotherapeutics; these organizations had no roles in this study. S.S. is currently an employee of Vertex Pharmaceuticals, which also had no role in this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Shalev Itzkovitz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Annotation and subclustering of scRNA-seq data.
a–d, UMAP and subcluster annotations of epithelial cells (a), myeloid cells (b), T cells (c, including a subcluster of plasmacytoid dendritic cells that initially clustered with T cells) and stromal cells (d). The epithelial UMAP uses the original embeddings, while the others use subcluster-specific embeddings. e–h, Dot plots showing relevant markers gene (columns) expression in the indicated subclusters (rows) of epithelial cells (e), myeloid cells (f), T cells (g) and stromal cells (h). Dot color indicates normalized expression, and dot size indicates the fraction of cells expressing the gene within the subcluster. i–l, UMAP visualization of all cells from scRNA-seq data, colored by tissue state (i, indicating stricture, non-stricture, inflamed or control), fraction (j, epithelial fraction or underlying tissue), location (k, small intestine, ascending colon or colon) and collection procedure (l, biopsy or resection).
Extended Data Fig. 2 Cell types and samples in scRNA-seq data.
a, Heatmaps show the cell-type correlation (Pearson correlation) between this paper and our previous CD paper8. Left, terminal ileum; right, colon. b, Separate PCoAs of Bray–Curtis dissimilarities per fraction. Left, samples from immune fraction; right, samples from epithelial fraction. c, Barplots show significant differences in composition for resection (blue) relative to biopsy (yellow) samples in the epithelial compartment. *padj. < 0.05, **padj. < 0.01 (Wald test from Dirichlet regression with FDR correction; Methods). Blue asterisks indicate overrepresentation in resection vs. biopsy samples, while red asterisks indicate underrepresentation. Error bars are s.e.m. The total number of cells contributing to each bar is also shown. d,e, Similar to c, but from stromal and immune compartments. f, Barplots contrast the number of cells from resection and biopsy samples.
Extended Data Fig. 3 Cell-type distribution by stricture status in scRNA-seq data.
a, Barplots show significant differences in cell-type frequency for non-IBD (green) and non-stricture (blue) samples relative to stricture (red) samples in immune, stromal epithelial and epithelial fraction immune compartments. *padj. < 0.05, **padj. < 0.01 (Wald test from Dirichlet regression with FDR correction; Methods). Blue asterisks indicate overrepresentation in non-stricture or non-IBD vs. stricture samples, while red asterisks indicate underrepresentation. Error bars are s.e.m. The total number of cells contributing to each bar is also shown. b, Similar to a, only using paired stricture and non-stricture samples (Methods).
Extended Data Fig. 4 Differential gene expression across non-stricture, stricture and high IAF samples.
a, Ranked ratio between the total number of DEGs (as shown in Fig. 2c) and number of analyzed cells per cell type. b, Volcano plots for inflammatory fibroblasts, with up to 15 DEGs (FDR < 0.05) per direction labeled. c, Number of DEGs between paired high IAF and non-stricture samples, displayed by cell type. Compare with number of DEGs between all stricture and non-stricture samples in Fig. 2c. d, Fractions of cell types in which KEGG pathways are significantly enriched (FDR < 0.05) within each compartment and by each comparison, split by enrichment direction.
Extended Data Fig. 5 Spatial distribution of cell types across all tissue sections.
For non-stricturing (top) and stricturing (bottom) tissue sections (columns), the spatial distributions of spot clusters, radial axis scores and the cell lineage proportions for B cells, epithelial cells, fibroblasts, myeloid cells, pericytes and T cells across all profiled spots. Cell-type proportions in each spot were estimated from the deconvolution with BayesPrism. Muscle cells were absent from the single-cell atlas and not included in the deconvolution, which substituted them with the most transcriptionally similar cell types, pericytes and myofibroblasts. Radial axis scores were congruent with the proportions of epithelial cells and pericytes (muscle). Scale bars, 2 × 2.5 mm (all images).
Extended Data Fig. 6 Spatial clustering and gene expression of inflammatory and collagen-hi fibroblasts.
a, For select cell types (x axis), the distribution of the radial axis scores of spots that contain high proportions of the cell type (>10% frequency estimated by the BayesPrism deconvolution). For clarity, only cell types with statistically significant differences from other cells are shown. Wilcoxon test, p values: epithelium (n = 21,408, p = 0), IgG+ plasma (n = 3,027, p = 1 × 10−95), CD63+CD81+ macrophage (n = 3,226, p = 2 × 10−64), tissue fibroblast (n = 15,640, p = 3 × 10−113), inflammatory fibroblast (n = 4,341, p = 1 × 10−82), venous endothelium (n = 4,252, p = 1 × 10−9), arterial endothelium (n = 1,384, p = 8 × 10−17), muscle (n = 46,040), p < 1 × 10−300), tissue macrophages (n = 1,671, p = 2 × 10−92) and collagen-hi fibroblasts (n = 1,798, p = 3 × 10−113). Boxplot quantiles: 25%, 50% and 75%; whiskers: 1.5 IQR. Adjusted p values: **<1 × 10−3, ***<1 × 10−10. b,c, DEGs for fibroblast subsets. For DEGs in inflammatory (n = 1,368; b) and collagen-hi (n = 111; c) fibroblasts relative to all other fibroblasts (n = 9,084 total), volcano plots showing the significance (y axis) and fold change in mean gene expression (x axis), computed from the single-cell reference atlas. The labels of select genes with statistically significant differential expression are provided (statistical significance assessed using a regularized logistic regression model; Methods). d, For select cell types (x axis), the distribution of the number of UMIs per single cell (y axis). For clarity, the 19 cell types with the highest number of UMIs per cell are shown, with all other cells combined into the ‘Other’ group. Boxplot quantiles: 25%, 50% and 75%; whiskers: 1.5 IQR. Total number of cells (left to right): 43; 8; 95; 14; 239; 24,147; 21,380; 551; 285; 6,465; 3; 111; 854; 2,993; 1,882; 1,140; 74; 662; 1,368; and 284,703. e, Interactions between inflammatory fibroblasts and T cells (top) and epithelial cells (bottom). Zoomed H&E images for samples V10A14-143_D (top row) and V11Y24-011_C (bottom row), overlaid with the gene expression levels of the induced target gene (IL11 or IL24, left column), the proportions of the ‘expressing’ cell across spots (inflammatory fibroblast, middle column) and the proportions of the ‘context’ cell across spots (T cells or epithelial cells, right column). Scale bar, 0.5 mm (all images).
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Table 1: Clinical metadata and sample information. Description of each individual and sample profiled in the study. Supplementary Table 2: DEGs for cell subsets during disease. DEGs in this table are genes with the adjusted P value for the discrete component <0.1 (that is, FDR < 0.1). Supplementary Table 3: KEGG enrichment results. Supplementary Table 4: Correlation of cell types between IAF score and cell composition. Supplementary Table 5: Genes that are significantly correlated with IAF scores in IgG+ plasma and arterial endothelial cells. Supplementary Table 6: Clinical metadata, sample information and spot annotations. Description of the individuals, samples and spots profiled in this study. Supplementary Table 7: Cell-type proportions and DEGs used to annotate spatial clusters. Cell-type proportions estimated with BayesPrism, and the top 50 DEGs per spatial cluster estimated using MAST. Supplementary Table 8: Spatially enriched ligand–receptor interactions between cell types. Expression statistics for all significant ligand–receptor interactions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kong, L., Subramanian, S., Segerstolpe, Å. et al. Single-cell and spatial transcriptomics of stricturing Crohn’s disease highlights a fibrosis-associated network. Nat Genet 57, 1742–1753 (2025). https://doi.org/10.1038/s41588-025-02225-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02225-y