Abstract
Diffuse large B cell lymphomas (DLBCLs) are a heterogeneous group of malignancies that can arise in lymph nodes or extranodal locations, including immune-privileged sites. Here, we applied highly multiplexed spatial transcriptomics and proteomics together with genomic profiling to characterize the immune microenvironment architecture of 78 DLBCL tumors. We define seven distinct cellular niches, each characterized by unique cellular compositions, spatial organizations and patterns of intercellular communication associated with niche-specific phenotypes of both T cells and tumor B cells. Among these, DLBCLs from immune-privileged sites showed abundant T cell infiltration into diffuse niches, where immune cells were intermixed with tumor B cells and bore transcriptional hallmarks of activation and effector function, suggesting that they may be primed for anti-tumor immunity. Spatial characterization of the DLBCL immune microenvironment, therefore, reveals cellular niches that foster divergent patterns of cell–cell interactions contributing to the phenotypic heterogeneity of both niche-resident tumor and immune cells.
Similar content being viewed by others
Main
DLBCLs are a common and heterogeneous group of mature B cell malignancies that typically arise in lymph nodes, sites in which stromal cells, antigen-presenting cells and other myeloid cells facilitate adaptive immune responses by T cells and B cells against pathogens. Lymph nodes are architecturally arranged to facilitate complex cell–cell interactions that control cell migration, expansion, clonal selection and negative feedback required for robust immune responses and their post-infection resolution. This architecture is lost in the setting of DLBCL, in which malignant B cells efface large portions of the lymph node, although many of the underlying cell types remain at variable frequencies and contribute to disease etiology.
Although most DLBCLs respond to first-line chemoimmunotherapy, approximately 40% of patients are either refractory or will relapse. The outcomes for these patients were historically poor1, but have significantly improved with the introduction of CD19 chimeric antigen receptor (CAR) T cells2,3 and, more recently, bispecific antibodies4,5. The breakthrough success of these immune therapies in some patients, but limited response in others, highlights the importance of understanding lymphoma immunobiology to identify strategies that can improve patient outcomes. Gene expression deconvolution-based approaches have shown that DLBCLs have significant interpatient heterogeneity in lymphoma microenvironment composition, which impacts patient outcomes but shows only a modest relationship to genetic or transcriptional subtypes6,7. Single-cell RNA sequencing (scRNA-seq) offers a substantial advantage over deconvolution-based approaches by directly measuring individual cells and has highlighted the diversity of the non-malignant T cell compartment of DLBCLs8,9,10,11. However, the tissue disaggregation approaches used in most studies result in the loss of non-hematopoietic and myeloid cells. Furthermore, both deconvolution and scRNA-seq approaches lose the 2D spatial organization of cellular niches (CNs) that foster distinct patterns of cell–cell interaction that contribute to the functional state of cells residing within each niche. Recent studies of DLBCL using imaging mass cytometry have uncovered characteristic patterns of cell type assembly into CNs, often defined by the predominance of a major cell type that is either in close proximity to or largely excluding tumor B cells12,13,14. These studies also suggested that tumor microenvironment characteristics may be associated with molecular features such as cell-of-origin subtypes, somatic mutations and patient outcomes12,13,14. However, spatial proteomics is typically limited by the number of antibodies available per platform and thus limits the discovery potential for immunological mechanisms underlying CN structure and resident cell phenotypes. Spatial transcriptomics is an emerging technology that leverages the advantages of single-cell transcriptomic analysis while retaining 2D tissue architecture15.
Here, we used spatial transcriptomics together with proteomic and genomic analysis to investigate the spatial immunology and pathobiology of DLBCL. We define distinct CNs that differ in frequency across tumors, related to both Epstein–Barr virus (EBV) infection status and the anatomical site of the tumor, and we show that residence within different CNs is associated with divergent functional states of T cells and tumor B cells owing to different patterns of cell–cell communication. This provides a framework for understanding stereotyped patterns of spatial organization in DLBCL and immunological interactions associated with heterogeneity in immune cell infiltration and function across patients.
Results
High-resolution cell typing by single-cell spatial transcriptomics
We assembled six tissue microarrays (TMAs) from excisional biopsies of 78 large B cell lymphomas and five controls (four tonsil, one lymph node) (Supplementary Tables 1 and 2 and Supplementary Fig. 1a). Large B cell lymphomas included 66 DLBCLs not otherwise specified; five EBV-positive DLBCLs not otherwise specified; four T cell/histiocyte-rich large B cell lymphomas; two primary mediastinal large B cell lymphomas; and one post-transplant lymphoproliferative disorder. Of these, 47 were previously untreated and 31 were relapsed–refractory cases (median prior line of therapy, 1; range, 1–6). High-quality whole-exome sequencing data were available for 75 samples (Supplementary Table 3), and each TMA was subjected to single-cell spatial transcriptomics using the NanoString CosMx platform with the 1K gene panel (949 probe sets; Supplementary Table 4) and spatial proteomics using a 31-antibody Co-Detection-by-InDEXing (CODEX) panel (Supplementary Table 5 and Fig. 1a).
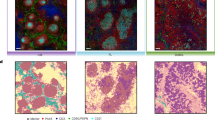
a, Schematic representation of the workflow, created with Biorender.com. Tissue cores from 78 B cell lymphoma samples and five normal tissue samples were assembled into six TMAs and underwent paired single-cell spatial transcriptomic and proteomic profiling using CosMx SMI and CODEX, respectively. Unsupervised cell clustering, spatial neighborhoods and niches, cell–cell communications, as well as association with clinical parameters were deeply dissected. b, Uniform manifold approximation and projection (UMAP) plot of major cell types identified in the CosMx SMI dataset. RBC, red blood cell. c, Canonical marker genes for each cell type shown in (b). d, Representative CosMx SMI (top) and CODEX (middle) images of samples from five different clinical conditions: IPS, EN-O, nodal (EBV-negative (EBV−)), nodal (EBV-positive (EBV+)) and normal tonsil. The regions in CODEX images that correspond to CosMx FOVs are highlighted. The major lineage compositions in each sample, as identified by CosMx and CODEX, are shown at the bottom.
After rigorous quality control, 1,322,740 high-quality, characterizable cells were retained from the CosMx profiling for downstream analysis. Following batch correction (Supplementary Fig. 1b–d), unsupervised clustering identified seven major cell types (Fig. 1b), as determined by canonical marker genes (Fig. 1c). Cells from each TMA were distributed across all cell types, indicating minimal batch effect (Supplementary Fig. 1b–d). To validate the accuracy of cell typing, we used our in-house large B cell lymphoma single-nucleus multiome (snMultiome) dataset, which contains all major lineages16. The major lineages (B cell, T cell, myeloid and non-hematopoietic) within our CosMx data showed high specificity in the expression of signatures from orthogonal snMultiome data and co-alignment within the joint embedding of the two datasets (Supplementary Fig. 1e,f). We further validated cell typing performance using a proteomic approach with CODEX, performed on the same TMAs but not on sequential sections, by comparing cellular frequencies between all cells from the core for CODEX and all cells from the field of view (FOV) for CosMx (Fig. 1d). Despite CosMx and CODEX being separated along the Z dimension, which prevents cell-level mapping, the four major lineages identified using a marker-based approach by CODEX (Extended Data Fig. 1a) showed a significant correlation with the cell types identified by CosMx (Spearman correlation, P < 0.05; Extended Data Fig. 1b). The strongest correlations were observed for B cell and T cell lineages, which were defined by three antibody markers in CODEX (Extended Data Fig. 1a,b). Both modalities confirmed a high degree of heterogeneity in cellular composition and tissue architecture across samples (Fig. 1d and Extended Data Fig. 1c). Spatial transcriptomic profiling with the CosMx platform is therefore a robust tool for exploring spatial composition and gene expression at single-cell resolution.
DLBCL harbors spatially distinct CNs
High-resolution clustering of CosMx data defined 19 cell types and states (Fig. 2a, Extended Data Fig. 2 and Supplementary Table 6), including a predominant (35.27% of cells) B cell cluster characterized by high expression of proliferation markers (TOP2A, PCNA) and immunoglobulin heavy-chain and light-chain restriction (Supplementary Fig. 2), therefore probably representing tumor B cells (C0_Tumor-B). We also identified three clusters of plasma cells distinguished by different immunoglobulin heavy-chain expression (C1_PC_IgG, C2_PC_IgA, C4_PC_IgM) and a CD44-expressing resting B cell population (C3_Resting-B). A single cluster of T cells was defined (C5_T), together with tumor-associated macrophages (TAMs) (C6_TAM_APOE_C1Q, C7_TAM_SPP1) and multiple other macrophage subsets (C8_Mac_DUSP1, C9_Mac_CXCL8, C10_Mac_MT2A). The non-hematopoietic compartment included diverse subsets such as follicular reticular cells (FRCs) (C11_FRC), high endothelial venule cells (C12_HEV), endothelial cells expressing von-Willebrand’s Factor (C13_Endothelial_VWF), vascular smooth muscle cells (C14_VSMC), follicular dendritic cells (C15_FDC) and a small number of epithelial cells (C17_Epithelial). Finally, we detected a small cluster of red blood cells (C18_RBC) and cells expressing heat-shock proteins that are hallmarks of cellular stress17 (C16_Stressed). Although the CosMx 1K platform lacks the transcriptome-wide coverage of scRNA-seq, it was able to markedly exceed the resolution of cell typing achievable with other spatial approaches, such as our 31-antibody CODEX panel.
a, High-resolution cell state identification in the CosMx SMI dataset. b, Schematic depiction of cellular neighborhood analysis, created with Biorender.com. A cellular neighborhood of 200 pixels in radius was identified for each cell. On top of that, the neighborhood compositions, spatial niches and neighborhood-based cell–cell communication analysis were performed. c, The neighborhood compositions of 19 different cell states in the CosMx SMI dataset, as represented by the sub-segmented barplots colored according to panel a. d, Identification of seven unique spatial niches from unsupervised clustering of cells based on cellular neighborhood compositions. The distribution of cells from spatial niches in the PCA space is shown. e, The cell state compositions of each spatial niche, represented by the sub-segmented barplots colored according to panel a. f, The representative DEGs in each spatial niche. g, Representative FOV images from CosMx SMI, each with a dominant spatial niche. CN cluster (above) is colored according to panel e, and cell type (below) is colored according to panel a.
A key advantage of single-cell spatial transcriptomics over scRNA-seq is the ability to map cells back onto the 2D space to identify cell neighbors and CNs. Indeed, with knowledge gained from spatial studies over the past decade18, it is increasingly better appreciated that groups of tissue-specialized cells work together as functional units rather than individually. In line with this notion, we performed cell neighborhood analysis for each cell and subsequently quantified the ability of immune cells to make functional contacts with one another and with tumor cells, using spatial statistics. A cellular neighborhood was defined as the collection of cells whose centroids lay within 200 pixels (24 µM) from the center of the neighborhood (Fig. 2b). The neighborhood structures were highly similar across different radius thresholds (Supplementary Fig. 3). As such, we selected the 200-pixel threshold, which included approximately 10–20 cells per neighborhood to maintain consistency with prior studies13,19. Accordingly, determination of the neighborhood compositions revealed that a cell state was always highly overrepresented in the neighborhoods centered by the same cell state (Fig. 2c). The propensity of like cells to spatially aggregate suggests the formation of distinct CNs. We therefore characterized reproducible patterns of spatial organization into niches by performing clustering of cell neighborhoods (Fig. 2d). This identified seven stereotypical CNs characterized by an enrichment of T cells (CN1_T; 8.76% of cells), plasma cells (CN2_PC; 2.34% of total tumor area), myeloid cells (CN3_Myeloid; 9.18% of total tumor area), stromal cells (CN4_Stromal; 6.93% of total tumor area), tumor B cells (CN5_Tumor-B; 14.21% of total tumor area), diffuse tumor cells amongst a heterogeneous immune infiltrate (CN6_Diffuse; 35.47% of total tumor area) or a mixed pattern of non-malignant cells with sporadic tumor cells (CN7_Mixed; 23.12% of total tumor area) (Fig. 2e and Extended Data Fig. 3a,b). The dominance of cell states within CN1–CN5 was reflected in their cell state-specific differentially expressed genes (DEGs), while CN6 and CN7 showed a mixed pattern in their DEGs reflective of their diverse cellular compositions (Fig. 2f). Representative FOVs for each cell niche and their corresponding cell states are shown in Fig. 2g and Extended Data Fig. 3c, and the spatial aggregations of major cell types (T cells or natural killer cells, B cells, myeloid and non-hematopoietic cells) within CN1–CN5 was validated using orthogonal CODEX images from the same samples (Extended Data Fig. 3d). In each specific cell niche (CN1–CN5), the dominant cell state occupied the center of the cell co-localization network, but no obvious center was observed in the mixed cell niche (CN7) (Extended Data Fig. 4). Interestingly, tumor B cells formed the center of the co-localization network within the diffuse cell niche (CN6), indicating contact between tumor B cells and diverse immune cell types within this microenvironment (Extended Data Fig. 4). The two niches with the highest fraction of tumor B cells were CN5 and CN6, representing densely packed tumor cells or diffuse tumor cells among immune infiltrate, respectively (Fig. 2e and Extended Data Fig. 3). The CN6 diffuse cell niche was present at variable degrees in 75 out of 78 tumors, representing a median of 33% of cells, whereas the CN5 dense tumor-B cell niche was more restricted, being observed in 58 out of 78 tumors and representing a median of 5% of cells (Supplementary Table 1 and Supplementary Fig. 4a).
No obvious associations were found between the relative representation of discrete cell niches within each tumor and other molecular characteristics, such as cell of origin subtype20, dark-zone signature21 or somatic mutation (Supplementary Fig. 4a). However, tumors harboring mutations in major histocompatibility complex class I (MHC-I) genes (B2M, HLA-A, HLA-B, HLA-C) tended to have a higher fraction of the diffuse cell niche (CN6) (Wilcoxon rank sum, P = 0.0118; Supplementary Fig. 4b), suggesting that immune evasion mechanisms may facilitate tumor B cell residence within an inflammatory niche. Comparison of the patterns of cell niche representation with previously described tumor microenvironment subtypes determined by deconvolution of bulk RNA sequencing7 showed only modest overlap. The previously described ‘depleted’ microenvironment subtype was enriched in seven out of eight tumors, with a high tumor area occupied by the CN5 dense tumor-B cell niche, while the ‘inflamed’ microenvironment subtype was enriched in six out of six and three out of four tumors, with a predominant CN3 myeloid-enriched cell niche or CN1 T cell-enriched cell niche, respectively (Supplementary Fig. 4a). However, both the ‘depleted’ and ‘inflamed’ microenvironment subtypes were also identified in tumors in which the CN6 diffuse and CN7 mixed cell niches were the dominant compartments (Supplementary Fig. 4a). Other subtypes, such as the ‘mesenchymal’ and ‘GC’ microenvironments, showed no clear pattern, demonstrating an overall lack of alignment between microenvironment characteristics directly measured by spatial transcriptomics and those inferred from deconvolution of bulk transcriptome data. Spatial transcriptomics, therefore, provides a more accurate construction of CNs in DLBCL that inform stereotypical patterns of cellular co-localization and interaction.
Niche-specific cell–cell communication
Although the initial round of unsupervised clustering defined only a single T cell cluster (C5_T), we reasoned that T cell phenotypes may differ within discrete cell niche structures owing to differing patterns of cell–cell communication from neighboring cells (Extended Data Fig. 5a). To explore this possibility, we conducted a supervised analysis of gene expression differences between T cells in CN1 (T cell-enriched), CN3 (myeloid-enriched) and CN5 (tumor B cell-enriched). The T cells within CN1 expressed the highest levels of naive/memory markers, high levels of cytotoxic markers and low levels of dysfunction markers (adjusted P < 0.05; Fig. 3a,b, Extended Data Fig. 5b and Supplementary Table 7), suggesting more functional and healthier T cell states. By contrast, T cells within CN3 showed lower expression of naive/memory markers but elevated levels of cytotoxic markers (adjusted P < 0.05; Fig. 3a and Extended Data Fig. 5c), along with high expression of LAG3, HAVCR2 (TIM3) and TNFRSF9 (4-1BB), indicative of an activated, effector phenotype combined with signs of dysfunction or exhaustion (adjusted P < 0.05; Fig. 3a and Supplementary Table 7). T cells within CN5 did not strongly express cytotoxic markers and instead expressed significantly higher levels of alternative markers of dysfunction, such as ENTPD1, TIGIT and PDCD1 (adjusted P < 0.05), suggesting that the tumor-enriched niche is suppressive to T cell function (Fig. 3a). To investigate the mechanisms of T cell recruitment and exclusion, we applied two approaches. First, we evaluated the expression of T cell chemotactic chemokines among the DEGs from each niche, revealing high expression of multiple chemotactic cytokines from diverse cell types, including APOE+C1Q+ TAMs (C6), tumor B cells (C0) and follicular reticular cells (FRCs) (C11) within both the T cell-enriched CN1 and myeloid-enriched CN3, but not within the tumor-enriched CN5 (Fig. 3b,c and Extended Data Fig. 5d,e). APOE+C1Q+ TAMs (C6) within the T cell-enriched niche (CN1) also expressed the highest level of antigen presentation genes (Fig. 3d), potentially contributing to the improved T cell health observed in this niche. Our second approach involved cell–cell communication analysis, which supported the prior result by demonstrating a significant enrichment of multiple chemokine–chemokine-receptor interactions within the T cell-enriched niche (CN1). These chemokines emanated from multiple cell types, with the corresponding chemokine receptors expressed on T cells (Extended Data Fig. 6a). Such interactions were absent in other cell niches, including the tumor-enriched niches, as summarized schematically in Fig. 3e. Inspection of single-molecule-resolution transcript expression within neighboring APOE+C1Q+ TAMs and T cells (Fig. 3f), or high endothelial venule cells and T cells (Extended Data Fig. 6b), clearly shows chemokine and chemokine receptor expression on neighboring cells, supporting the notion of local interactions within each niche. Cell–cell communication analysis also confirmed increased antigen presentation via MHC class II (MHC-II) to CD4 T cells within the T cell-rich niche (Extended Data Fig. 6c) and, by contrast, highlighted an enrichment of CD274–PDCD1 (PD-L1–PD-1) interactions within the myeloid-enriched and tumor-enriched niches, with the source of PD-L1 emanating from myeloid cells or tumor cells, respectively (Fig. 3g,h and Extended Data Fig. 6d,e). Together, these findings highlight substantial differences in T cell states and phenotypes across diverse CNs, which are probably influenced by distinct patterns of cell–cell communication within each niche.
a, The expression of naive/memory, cytotoxic and exhaustion markers in T cells from different spatial niches. Statistically significant DEGs are highlighted in dashed boxes. b, A representative FOV from CosMx SMI dominated by CN1, with abundant expression of cytotoxic markers in T cells, as well as T cell chemotaxis molecules from APOE+C1Q+ TAMs, tumor B cells and FRCs. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d. c, T cell chemotaxis molecules in APOE+C1Q+ TAMs, tumor B cells and FRCs from different spatial niches. d, The expression of MHC-II genes in APOE+C1Q+ TAMs residing in different spatial niches. e, Summary of T cell chemotaxis pathways in CN1 and CN5, created with Biorender.com. f, The expression of CXCL9 and CXCR3 in APOE+C1Q+ TAMs and T cells from the same FOV as panel b. g, Barplot depicting the normalized PD-L1–PD-1 interaction count in each spatial niche. P values were calculated with a Kruskal–Wallis test. h, A representative CN3-dominated FOV showing the PD-L1–PD-1 interaction between APOE+C1Q+ TAMs and T cells. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d.
Tumor B cell niches mirror germinal center interactions
Tumor B cells were found in all CNs, but were most enriched in CN5, where they largely occupied the spatial regions, and in CN6, where they were intermixed with a diverse immune infiltrate. To investigate the mechanisms contributing to these divergent spatial niches, we examined the characteristics of tumor B cells within CN5 and CN6. We observed higher expression of cell-cycle-related markers and the anti-apoptotic BCL2 oncogene in tumor B cells residing in CN5 compared to those in CN6 (adjusted P < 0.05; Fig. 4a,b, Extended Data Fig. 7a–c and Supplementary Tables 7 and 8), providing further evidence that rapid expansion of tumor B cells may contribute to the effacement observed in these regions. A supervised analysis of DEGs uncovered significantly higher expression of multiple genes implicated in lymphoma biology within CN5-resident tumor B cells, including genes involved in BCR signaling and epigenetic regulation, compared to CN6-resident tumor B cells (Extended Data Fig. 7d,e). Cell–cell communication analysis revealed additional differences between the CN5 and CN6 niches. The CXCL12–CXCR4 interactions were the most significant interaction in the tumor-enriched CN5 niche, with FRCs serving as a source of CXCL12 for CXCR4-expressing tumor B cells (Fig. 4c,d and Extended Data Fig. 7f) and both FRCs and APOE+C1Q+ TAMs serving as a CXCL12 source for T cells (Fig. 4e,f). By contrast, tumor B cells within CN1 showed significant CD40L–CD40 interactions with CD4 T cells (Extended Data Fig. 7g), with a subset also having higher expression of multiple immune checkpoint ligands (Extended Data Fig. 7h). The predominant tumor-B cell niche thus bears patterns that are reminiscent of the germinal center dark zone, which is regulated by CXCR4, harbors highly proliferative B cells and has low T cell abundance compared to the germinal center light zone, in which B cells are less proliferative and receive CD40-mediated co-stimulatory signals from CD4 T cells.
a, The expression of cell cycle markers and G2/M signature score in tumor B cells from different spatial niches. Statistically significant DEGs are highlighted in dashed boxes. The P value was calculated with a one-way ANOVA. b, A representative FOV showing extensive expressions of the cell cycle marker TUBB in tumor B cells from CN5. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d. c, The CXCL12–CXCR4 interaction between FRC and tumor B cells in the same FOV as panel b. d, The activity of CXCL12–CXCR4 interaction between all TME cells and tumor B cells in CN5 and CN6, with senders from the same spatial niche as tumor B cells. e, The activity of CXCL12–CXCR4 interaction between APOE+C1Q+ TAM or FRC and T cells in different spatial niches. f, A representative CN5-dominant FOV showing the interactions depicted in panel e. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d.
T cell dysfunction in EBV-positive DLBCL
EBV is a B cell transforming virus that is a driver of ~5% of DLBCL cases22. In healthy individuals, EBV-infected B cells are surveilled by T cells, which prevent malignant outgrowth. We therefore explored the T cell characteristics of nodal EBV-positive DLBCLs from immunocompetent individuals (that is, excluding post-transplant lymphoproliferative disorder; n = 5) by comparing them to nodal EBV-negative DLBCLs (n = 29) using available EBV status from clinical EBER in situ hybridization testing (Supplementary Table 1). EBV-positive DLBCLs harbored a significantly lower proportion of tumor B cells (C0) and a significantly higher proportion of T cells (C5) (Fig. 5a and Supplementary Table 9). As such, the tumor-enriched CN5 niche represented only 0.04% of cells in EBV-positive DLBCLs, while the T cell-enriched CN1 and mixed CN7 niches were significantly overrepresented (Fig. 5b and Extended Data Fig. 8a; hypergeometric P < 0.0001 for both CN1 and CN7), and T cells predominantly resided within these niches (Fig. 5c,d). In line with this observation, cell–cell communication analysis showed higher T cell chemotaxis signals and receipt of antigenic signals through MHC-I and MHC-II in EBV-positive DLBCLs (Extended Data Fig. 8b–d). This included closer interaction with tumor B cells, which more frequently neighbored T cells and APOE+C1Q+ TAMs in EBV-positive cases compared to neighboring other tumor B cells in EBV-negative cases (P < 0.001 for all three cell types; Fig. 5e, Extended Data Fig. 8e and Supplementary Table 10), as corroborated by orthogonal CODEX data (Extended Data Fig. 8f). Given that the proportions of CNs differed between EBV-negative and EBV-positive DLBCLs, and because niche residence is associated with functional differences, we controlled for this confounding factor by restricting our comparison of T cell function between EBV-negative and EBV-positive tumors to the T cell niche (CN1). Although the T cell niche (CN1) supported naive or memory T cells with low expression of exhaustion markers in EBV-negative DLBCLs, the expression levels of markers and signatures of cytotoxicity and exhaustion were significantly increased in EBV-positive DLBCLs (Fig. 5f–i and Supplementary Table 11). At single RNA molecule resolution with CosMx (Fig. 5j) and through protein staining with CODEX (Fig. 5k), it can be clearly observed that T cells within the CN1 T cell niche in EBV-positive DLBCLs exhibited markedly higher expression of effector molecules. Furthermore, T cells within EBV-positive DLBCLs co-expressed exhaustion markers such as HAVCR2 (Fig. 5j). Together, these results indicate that despite EBV-positive DLBCLs having an overrepresentation of T cell and mixed niches that are typically favorable for T cell function in EBV-negative DLBCLs, the T cells within EBV-positive tumors exhibit cytotoxic and exhaustion markers indicative of chronic stimulation and dysfunction.
a, Comparisons of the cellular proportions of tumor B cells and T cells in EBV-positive and EBV-negative nodal B cell lymphoma. P values were calculated with a Wilcoxon rank sum test. b, Pie chart representation of the CN compositions of EBV-positive and EBV-negative nodal lesions. c, Sub-segmented barplot depicting the CN allocations of T cells in nodal lesions with distinct EBV infection status. d, Representative FOVs from CosMx SMI showing the distinct cellular compositions in EBV-positive and EBV-negative nodal lesions. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d. e, The neighborhood structures of tumor B cells in EBV-positive and EBV-negative nodal lesions. f–i, The expression of representative cytotoxicity (f,g) or exhaustion (h,i) markers and signatures in T cells from EBV-positive and EBV-negative nodal lesions. P values for genes (GZMB, HAVCR2) were calculated with the Seurat FindAllMarkers function that uses a Wilcoxon rank sum test with Bonferroni correction. P values for signatures (cytotoxicity, exhaustion) were calculated with a Student’s t-test. j, Representative FOV images from CosMx SMI showing the expression of GZMB and HAVCR2 in T cells from EBV-positive and EBV-negative nodal lesions. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d. k, Representative CODEX images from the same sample as panel j, with a larger amount of Granzyme B+CD8+ T cells in EBV-positive nodal lesions.
T cell infiltration in DLBCL of immune-privileged sites
Despite arising in sites such as the central nervous system (CNS) and testis, which have restricted trafficking of immune cells, DLBCLs of immune-privileged sites (IPS) have been noted to have immune infiltration23,24. We explored the immunological characteristics of IPS DLBCL within our dataset by comparing 11 IPS DLBCLs (CNS, n = 5; testicular, n = 5; ocular, n = 1) with 28 nodal DLBCLs and 23 DLBCLs from other extranodal sites (EN-O; Supplementary Table 1). Splenic and EBV-positive cases were excluded from this analysis. Although IPS DLBCLs had a higher proportion of tumor B cells (C0) compared to nodal and EN-O DLBCLs, there was no significant difference in the frequency of T cells (C5) (Fig. 6a and Supplementary Table 12). APOE+C1Q+ TAMs (C6) and FRCs (C11) were more abundant in EN-O DLBCLs, but not significantly different between nodal and IPS DLBCLs (Fig. 6a and Supplementary Table 12). IPS and EN-O DLBCLs each showed a lower proportion of cells within tumor-enriched CN5 niches and a higher proportion of cells within diffuse CN6 and mixed CN7 niches (Fig. 6b and Extended Data Fig. 9a), while nodal DLBCLs had a higher fraction of tumor B cells that neighbored each other within tumor-enriched CN5 niches (Fig. 6c and Extended Data Fig. 6b). Despite being in sites of immune privilege, tumor B cells in IPS DLBCLs neighbored T cells to an approximately equal degree as nodal DLBCLs and significantly more frequently than in EN-O DLBCLs (Fig. 6d, Extended Data Fig. 9b and Supplementary Table 13). Furthermore, an approximately equal fraction of T cells in nodal and IPS DLBCLs reside within diffuse CN6 niches where T cells and other cells are intermixed with tumor B cells, while in EN-O DLBCLs, a greater fraction of T cells resided within mixed CN7 niches that contain fewer tumor B cells (Fig. 6e,f and Extended Data Fig. 9c). In addition, we observed a significantly higher expression of the cytotoxicity signature (Fig. 7a), a greater fraction of cells expressing effector molecules (GZMB and PRF1, adjusted P < 0.0001; Fig. 7b) and higher rates of proliferation (adjusted P < 0.0001; Fig. 7c–g and Extended Data Fig. 9d,e) in T cells from IPS DLBCLs compared to those from nodal or EN-O DLBCLs, which is suggestive of ongoing antigenic responses and expansion of T cells within IPS DLBCL. However, this was accompanied by a higher expression of markers related to T cell dysfunction, such as LAG3, HAVCR2, PDCD1 and ENTPD1 in T cells from IPS DLBCLs (Fig. 7h,i), with ligands such as CD274 (PD-L1) being highly expressed by other cell types within IPS DLBCL (Fig. 7j and Extended Data Fig. 9f), as well as significant Galectin-9–TIM3 (HAVCR2) signaling within IPS DLBCLs (Fig. 7k and Extended Data Fig. 9g), together indicative of an immune-suppressive microenvironment. Together, these data show that IPS DLBCLs have robust T cell infiltration within diffuse and mixed CNs, with strong proliferation and cytotoxicity but also accompanied by coinhibitory signals, highlighting IPS DLBCLs as being potentially amenable to immunotherapy.
a, Cellular compositions of B cell lymphoma developed in different anatomical sites, including nodal, IPS and EN-O lesions. The center line shows the median, boxes represent quartiles 1–3 and whiskers represent 1.5 times the interquartile range. P values were calculated using a Kruskal–Wallis test, with Dunn’s post hoc test for pairwise comparisons. b, Pie chart representation of the CN compositions of tumors in different anatomical sites. c,d, Boxplot representation of tumor B cell and T cell populations in the neighborhood of tumor B cells from different anatomical sites. The center line shows the median, boxes represent quartiles 1–3 and the whiskers represent 1.5 times the interquartile range. P values were calculated using a Kruskal–Wallis test, with Dunn’s post hoc test for pairwise comparisons. e, Sub-segmented barplot showing CN allocations of T cells from different tumor sites. f, Representative FOVs from CosMx SMI showing the distinct cellular compositions in tumors from different anatomical sites. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d.
a, The expression of the cytotoxicity signature in lymphomas of different tumor sites. P values were calculated using a pairwise t-test. b, The expression of cytotoxicity markers GZMB and PRF1 in T cells from nodal, IPS and EN-O lesions. P values were calculated comparing IPS to all other tumors using the FindAllMarkers function of Seurat, based upon a Wilcoxon rank sum test with Bonferroni correction. c, The expression of G2/M cell cycle signature according to tumor site. P values were calculated using a pairwise t-test. d, The expression of a representative cell cycle gene, HMGB2, according to tumor site. P value was calculated comparing IPS to all other tumors using the FindAllMarkers function of Seurat, based upon a Wilcoxon rank sum test with Bonferroni correction. e, Representative FOVs from CosMx SMI showing the expression of cytotoxicity marker GZMB and proliferation marker HMGB2, and in T cells from different tumor sites. Cell cluster is colored according to Fig. 2a, and CN cluster is colored according to Fig. 2d. f,g, Representative CODEX images from the same sample as panel e, showing Granzyme B+CD8+ T cells in IPS lesions (f) and the presence of proliferating Ki67+CD8+ T cells in multiple regions of the IPS sample (g). h,i, The expression of exhaustion signature (h) and representative exhaustion markers (i) in T cells from different tumor sites. In h, P values were calculated using a pairwise t-test. j,k, The PD-L1–PD-1 interaction (j) and Galectin-9–TIM3 interaction (k) between all other cells and T cells in different tumor sites.
Discussion
In this study, we have shown that DLBCLs consist of CNs with stereotypical patterns of co-association in communities that harbor distinct patterns of cell–cell communication, contributing to diversity in the proportions and functional states of each cell type. For example, T cells residing in T cell-enriched niches exhibited increased expression of naive/memory markers and reduced exhaustion markers compared to those in myeloid-enriched and tumor-B cell-enriched niches. These differences in T cell proportions and functional states were linked to variations in chemoattractant and coinhibitory signals within each niche, respectively. However, we noted that etiological factors, such as EBV and the anatomical location of the tumor, were also associated with different immune cell states even within the same CN that may make tumors more or less immunologically primed. These factors are therefore important to consider as we move towards more patient-tailored immunotherapeutic strategies with our current armamentarium of checkpoint-blocking antibodies, bispecific T cell engagers and CAR T cell therapies.
EBV-positive DLBCL arising in immunocompetent individuals has been recently recognized as a distinct clinical entity25 and is associated with inferior outcomes26,27. Although dysfunctional T cells have been observed to aggregate with EBV-positive B cells in a range of settings28,29, the CNs and patterns of cell–cell communication contributing to T cell dysfunction in EBV-positive DLBCLs have not been explored. We found an enrichment of T cell-enriched niche structures within EBV-positive DLBCLs that is associated with high levels of T cell expression of cytotoxic molecules, but accompanied by high expression of coinhibitory molecules and exhaustion signatures. This suggests that T cells in EBV-positive DLBCLs are primed for anti-lymphoma immunity and may be responsive to immunotherapeutic strategies, such as PD-1 checkpoint blockade30 or CAR T cell therapy31,32. In line with this possibility, PD-1 blockade in combination with chemoimmunotherapy has been found to have high activity in the frontline treatment of EBV-positive DLBCLs33.
Approximately 30% of DLBCLs arise in extranodal sites, including IPS such as the CNS, testis and eye34, and therefore do not share the same underlying lymph node architecture of nodal DLBCLs. Notably, the MYD88/CD79B (MCD) genetic subtype of DLBCL frequently involves extranodal sites, including IPS, and possesses mutations in human leukocyte antigen genes in 76% of cases35. Furthermore, CNS and testicular DLBCLs frequently bear amplification of the PD-L1 (CD274) locus36. These genetic observations suggest that IPS DLBCLs are subject to complex immunoediting despite residing in sites of immune privilege. Consistent with this suggestion, we found that IPS DLBCLs had strong immune cell infiltration, with the majority of cells residing within diffuse or mixed CNs that contain complex mixtures of diverse cell types. The immune infiltration observed in IPS DLBCLs is consistent with the dysfunction of the blood–brain barrier in CNS lymphomas and in line with the activity of both CD19 CAR T cells and bispecific antibodies in these lymphomas37,38, which both require the infiltration and engagement of T cells with tumor cells. T cells within IPS DLBCLs showed high expression of proliferation and cytotoxic molecules; however, like EBV-positive DLBCLs, they also displayed high expression of multiple coinhibitory molecules. In line with prior observations30, the genetic etiology of IPS DLBCLs36 and responsiveness of CNS lymphomas to PD-1 blockade39,40,41, we found the PD-1–PD-L1 axis to be most prominently active in IPS DLBCLs. However, our data also highlight strong expression of the LAG3 and TIM3 (HAVCR2) coinhibitory molecules that may also represent tractable therapeutic targets in this context.
The limitations of this study include our use of TMAs and restricted FOVs on the CosMx platform, which may introduce sampling bias and limit the perspective of intratumoral diversity. Furthermore, CosMx and CODEX analyses were not performed on serial sections, resulting in spatial data being separated along the Z dimension. This prevents direct cell-to-cell mapping and limits direct comparisons between the two platforms. Our analysis leveraged a heterogeneous cohort of patients that enabled investigation of unique factors, such as EBV status and extranodal sites, but prevented us from identifying associations between CN features and patient outcomes. Despite these limitations, the discovery of spatially confined patterns of DLBCL immunobiology highlights core concepts around the functional diversity of cell types within different CNs, identifies opportunities for immunotherapeutic interventions in EBV-positive and IPS DLBCLs and positions the field for prospective evaluation of the spatial architecture of lymphoma tumor microenvironments in important therapeutic contexts, such as bispecific antibody and CAR T cell therapy. Future studies using platforms with higher information content and larger sample cohorts will be required to distill key principles of tumor microenvironment architecture, its underlying biology and associated therapeutic opportunities, along with associations with molecular and clinical features of DLBCL. These large datasets will also serve as a foundation for training deep learning models or other artificial intelligence-based approaches capable of predicting gene expression and microenvironmental features from routine digitized hematoxylin and eosin-stained slides42,43,44 to yield clinically deployable, microenvironment-informed or integrative strategies for DLBCL classification.
Methods
Human research ethics approval
This study was approved by the Institutional Review Board of The University of Texas MD Anderson Cancer Center (protocol PA19-0420) using archival samples and a waiver of informed consent. The sex and age of study participants are provided in Supplementary Table 1. No compensation was provided to participants.
CosMx SMI assay
To obtain a quasi-3D transcriptomic dataset, formalin-fixed, paraffin-embedded (FFPE) tissue blocks were sectioned to 5 μm slices using a microtome. The sample processing, staining, imaging and cell segmentation were performed as previously described. In brief, tissue sections were placed on VWR Superfrost Plus Micro Slides (cat. no. 48311-703) for optimal adherence. Slides were then dried at 37 °C overnight, followed by deparaffinization, heat-induced antigen retrieval and proteinase-mediated permeabilization (https://nanostring.com/products/cosmx-spatial-molecular-imager/single-cell-imaging-overview). Then, 1 nM RNA-in situ hybridization probes were applied for hybridization at 37 °C overnight. After a stringent wash, a flow cell was assembled on top of the slide, and cyclic RNA readout on CosMx was performed (Human Universal Cell Characterization RNA 1K-plex panel). Before starting RNA detection cycling, morphology markers for visualization and cell segmentation were added, including pan-cytokeratin (PanCK), CD68, CD298/B2M, CD45 and DAPI. Over 40 FOVs (0.509 mm × 0.509 mm) were selected for data collection on each slice. The commercial CosMx optical system has an epifluorescent configuration based on a customized water objective (×22.7/1.1 NA) and uses widefield illumination, with a mix of lasers and light-emitting diodes (385 nm, 750 mW; 488 nm, 1 W; 530 nm, 1 W; 590 nm, 150 mW; 647 nm, 1 W) that allow imaging of DAPI, Alexa Fluor 488, Atto-532, Dyomics Dy-605 and Alexa Fluor 647, as well as removal of photocleavable dye components. The camera used was a Lucid Atlas10 ATX CMOS sensor (pixel size, 120 nm). A 3D multichannel image stack (eight frames) was obtained at each FOV location per imaging round, with a step size of 0.8 μm. Image registration, feature extraction, localization, decoding of individual transcripts and machine learning-based cell segmentation (developed upon Cellpose) were performed as previously described. The final segmentation mapped each transcript in the registered images to its corresponding cell, as well as to subcellular compartments (nucleus, cytoplasm, membrane), where the transcript was located.
CosMx data analysis
Data quality control and processing
The CosMx SMI data for TMA slides, including gene count matrices, metadata, FOV positions, cellular boundary coordinates and transcript coordinates, were downloaded from the Nanostring AtoMx Spatial Informatics Platform. The gene count matrix was used for transcriptomic analysis with the R software Seurat (v.5.0.1)45. Cells with a unique molecular identifier count of less than 20 or a gene count of less than ten were filtered out. Furthermore, cells lacking the expression of defined canonical markers were excluded from subsequent analyses. Finally, a total of 1,322,740 high-quality, characterizable cells were retained for downstream analysis, with a median gene count of 74 per cell and a median unique molecular identifier count of 121 per cell. Data normalization was performed using the SCTransform function in Seurat45. Linear dimension reduction based on highly variable genes was performed by principal component analysis (PCA), using the RunPCA function in Seurat46. Elbow plots were generated using the ElbowPlot function, and the number of dimensions used for downstream analysis was determined accordingly. Following this step, TMA-specific batch effects were evaluated, and data integration in the PCA space was performed using the R package Harmony (v.0.1.1)47. A nearest-neighbor graph was then constructed with the FindNeighbors function in Seurat45, and unsupervised cell clustering was conducted using the Louvain algorithm with the FindClusters function. At a resolution of 0.1, a total of 19 cell clusters were identified. Further non-linear dimension reduction was performed using the uniform manifold approximation and projection method48, implemented with the RunUMAP function in Seurat45.
Cell type and state identification
DEGs among cell clusters were identified using the FindAllMarkers function in Seurat45. The top DEGs of each cell cluster were manually reviewed. Additionally, the expression levels of canonical lineage markers, as reported previously17,49,50,51, were manually checked, and dot plots were generated using the DotPlot function in Seurat45. Based on these methods, cell clusters were annotated as T cells and natural killer cells (high expression of CD3D, CD3E, CD2 and so on), B cell and plasma cells (high expression MS4A1, CD79A, MZB1 and so on), myeloid cells (high expression of CD68, APOE, C1QB and so on), stromal cells (high expression of COL1A1, COL1A2 and so on), stressed cells (high expressing of HSP genes), epithelial cells (high expression of KRT genes) and erythrocytes (that is, red blood cells) (high expression of HBB, HBA1/2).
To further validate the accuracy of the identified cell types, the signatures of major cell types from an in-house-constructed B cell lymphoma atlas were used (see Supplementary Table 4 for the detailed signature). This atlas was based on single-nucleus RNA sequencing (snRNA-seq) data from 105 B cell lymphoma samples and contained 970,239 cells, ensuring its comprehensiveness and accuracy as a reference. Signature scores for each major cell type were calculated using the AddModuleScore function and visualized with the FeaturePlot function in Seurat45. Furthermore, we integrated the CosMx SMI and single-nucleus RNA sequencing datasets using Harmony47 and overlaid the two datasets in the joint embedding space for all cells as well as for each major cell type. Finally, we performed Spearman correlation analysis to evaluate the concordance of major cell types identified from CosMx SMI and CODEX datasets using the same set of samples.
Identification of cellular neighborhoods and spatial niches
We defined a cellular neighborhood as the collection of cells whose centroid lies within 200 pixels (24 µm) of the centroid of the center cell. Such a strategy was applied to the entire CosMx SMI dataset, with each retained high-quality cell being iteratively selected as the center. Based on this definition, most cellular neighborhoods contain ~10–20 cells. A cell-by-cell state composition matrix was then calculated, in which each column represents a center cell and each row represents the count of a cell state within that center cell’s neighborhood. We tested multiple neighborhood radii (from 100 pixels to 2,000 pixels) and obtained the corresponding neighborhood composition matrix. Then, pairwise Spearman correlation analysis was performed to assess the similarity of the neighborhood structures obtained using different neighborhood searching radii.
Subsequently, k-means clustering was performed based on the cell-by-cell state composition matrix, obtained using a neighborhood radius of 200 pixels, to cluster center cells based on the similarity of their neighborhood structures. Multiple resolutions were tested to optimize the clustering results. Then, clusters with similar structures were combined to reduce redundancy. This analysis identified seven cell clusters, each characterized by unique neighborhood compositions, which we defined as spatial CNs. Dimensionality reduction of the dataset was performed on the scaled neighborhood compositions using PCA, and the CN information was projected onto the PCA space for visualization.
Quantification of cellular enrichment
To quantify the enrichment of different cell states across different groups (that is, the neighborhood compositions of different cell states or the cellular compositions of different spatial niches) and determine whether a certain cell state is enriched or depleted, we calculated the ratio of observed to expected cell numbers for each cell state across different groups, as reported previously52. First, a contingency table of cell states by groups was calculated. Then, a chi-squared test was performed to calculate the expected distributions of cell states across the different groups and to evaluate the deviation of the actual distribution from the expected distribution. Following this, the Ro/e values for each cell type-group combination were calculated using the following formula:
Here, Ro/e represents the ratio of observed to expected cell numbers for a particular cell state within a given group. For a specific cell state, Ro/e > 1 indicates that cells of this state are observed more frequently than expected by random chance in that group (that is, enriched), whereas Ro/e < 1 indicates that the number of cells of this state is lower than expected (that is, depleted). The results were shown as heatmaps using the R package pheatmap (v.1.0.12).
Niche-specific cell co-localization network
The cell co-localization network in each spatial niche was calculated with the R package CellTrek (v.0.0.94)53. In brief, the function scoloc was used to construct a minimum spanning tree based on the Delaunay triangulation network for cells within each spatial niche. The co-localization graph of different cell states within each spatial niche was then visualized using the function scoloc_vis. In the graph, each node represents a cell state, and the width of the edge connecting two cell states represents their proximity within the specific niche.
Niche-specific cell functional state analysis
The FindAllMarkers function in the R package Seurat45 was applied to identify DEGs for different spatial niches as well as DEGs for specific cell types across different spatial niches. Specifically, the cell cycle signature was derived by intersecting the cell cycle metaprograms reported in a recent pan-cancer study46 with the CosMx SMI gene panel (see Supplementary Table 6 for the detailed signature). The T cell cytotoxicity and exhaustion signatures were derived by intersecting the DEGs of corresponding cell clusters from a pan-cancer T cell atlas17 with the CosMx SMI gene panel. The top 20–50 DEGs from the pan-cancer T cell atlas were used for constructing each signature, ensuring that at least ten genes overlapped with the CosMx SMI gene panel (see Supplementary Table 9 for the detailed signature). The signature scores were then calculated using the AddModuleScore function in Seurat45.
Neighborhood-based cell–cell communication analysis
For spatially resolved single-cell transcriptomic data, we inferred cell–cell communications within each cell’s neighborhood, leveraging spatial information and restricting cellular communication to adjacent regions. We used the human ligand–receptor databases from CellChat54, CellPhoneDB (v.2.0)55 and iTALK56, which together comprised a total of 4,028 ligand–receptor pairs. The crosstalk between each cell and its neighboring cells was quantified based on ligand–receptor interaction activity, calculated as follows:
where L stands for a ligand, R stands for the corresponding receptor, s denotes a sender cell expressing ligand L, r represents a receiver cell expressing the corresponding receptor R, ExpL,s stands for the expression value of ligand L in the sender cell s and ExpR,r stands for the expression value of the corresponding receptor in the receiver cell r. The calculations described above were implemented using the commu.cn.call function in Spyrrow (v.0.1; https://github.com/liuyunho/Spyrrow) in the Python environment (v.3.10.5). The results were visualized using the ggplot2 package (v.3.4.4) in R (detailed in the section ‘CosMx SMI data visualization’).
Calculation of normalized PD-L1–PD-1 interaction count and intensity
The normalized PD-L1–PD-1 interaction count was calculated as the total number of cell pairs of interest (myeloid–T cell pair or tumor–T cell pair) with a positive PD-L1–PD-1 interaction activity, divided by the total number of receiver cells (C5_T plus natural killer cells) in each spatial niche. The normalized PD-L1–PD-1 interaction intensity was calculated as the sum of PD-L1–PD-1 interaction activity between cell pairs of interest (myeloid–T cell pair or tumor–T cell pair) divided by the total number of receiver cells (C5_T plus natural killer cells) in each spatial niche. These two matrices assess the extent to which T cells were influenced by the PD-L1–PD-1 interaction in each spatial niche.
CosMx SMI data visualization
Cell patch plots were generated using the cell boundary coordinates, implemented with the function plot_cell_patch in the software Spyrrow. The single-molecule resolution gene expression plots were generated using both the cell boundary and transcript coordinates by plotting the transcripts on top of the corresponding cell patches. This was implemented with the plot_RNA_on_cell function in Spyrrow, with the transcripts plotted only for the cell states of interest. For the visualization of cell–cell communication results, the R package ggplot2 (v.3.4.4) was applied. For the communication between cell clusters in one or different spatial niches, dot plots were applied to show the proportion of cell pairs involved in the specified ligand–receptor interaction in a certain niche and the average interaction intensity (total interaction intensity normalized by the number of cell pairs).
CODEX assay
CODEX technology (Akoya Biosciences) was used for multiplex immunofluorescence marker detection on FFPE tissue sections.
Antibody conjugation
In addition to the commercially available conjugates from Akoya Biosciences, additional antibody conjugates were made in-house using barcodes and conjugation kits provided by Akoya, following the manufacturer’s protocol. In brief, each antibody in purified, carrier-free form was treated with the reduction solution (conjugation kit component) to open up the thiol groups and subsequently incubated with a unique DNA barcode for 2 h. Successful conjugations were first confirmed with a gel electrophoresis run, showing an upward shift in molecular weight. These conjugates were then tested at different dilutions for staining on a sample tissue expressing the target antigen. The staining was reviewed and approved by pathologists. A comprehensive list of the antibody conjugates used in this study, including the conjugates from Akoya, is provided in Supplementary Table 3.
Tissue staining
Tissue staining and imaging were performed according to the instructions by Akoya Biosciences. In brief, FFPE sections (4 μm thickness) mounted on coverslips were baked at 55–60 °C overnight. Once cooled down to room temperature (20–25 °C), the slides were rinsed in deionized water, followed by rinses in CODEX hydration buffer, and then placed in staining solution to incubate at room temperature for 30 min. Then, 200 μl of antibody conjugate mix consisting of CODEX staining solution, blocking solutions and antibody conjugates at different dilutions (see Supplementary Table 3) was added onto the tissue section and incubated for 3 h at room temperature. After incubation, the coverslips were washed in staining solution twice, post-fixed with 1.6% paraformaldehyde in storage buffer, cold 100% methanol and an Akoya final fixative with abundant washing steps in 1× PBS in between and after each step. The coverslips were then subject to imaging or short-term storage in storage solution at 4 °C for up to 5 days.
Imaging on the CODEX instrument
A 96-well reporter plate containing the corresponding reporters was prepared for imaging. Each well to be used in the plate contained three reporters that correspond to the antibodies used in each cycle, plus a DAPI nuclear stain. Two wells containing only DAPI were used as the first and last cycle for the correction of background, known as blanks. The plate was sealed with a foil plate seal to block it from light. During imaging, one coverslip was mounted on a perfusion stage resting on a Keyence fluorescence microscope (model BZ-X800) equipped with a ×20/0.75 NA objective. The solution exchanges were performed using a microfluidics instrument (Akoya Biosciences) and controlled through a software interface. The order of markers per cycle is listed in Supplementary Table 3. After completion of imaging cycles, the raw image tiles in.tiff format were fed through the CODEX image processor for stitching. Quality control of tissue, signals and staining was performed by an experienced pathologist.
CODEX data analysis
Cellular segmentation
Cellular segmentation was performed using a convolutional neural network-based approach with UNet++ architecture. As inputs, the nuclear marker DAPI and the most common membrane marker Na/K-ATPase were used. A cell instance mask, which includes each cell detected by the aforementioned network, was produced as an output of the cellular segmentation process. Cell instances that lack clearly visible membranes but contain nuclei were segmented with a slight extension around the nuclei. The segmentation quality for all samples was visually inspected by pathologists experienced in MxIF analysis. It was confirmed that the segmentation quality for all cores met the 90% threshold. To ensure accurate analysis, large segments that did not represent real cells and appeared as a result of artificial staining, blur or other artifacts were filtered out based on their size.
Mask generation
Signal masks for macrophages were generated by setting a threshold for CD68, CD11c, CD206 and HLA-DR markers. Vessel masks were generated with UNet++ with an EfficientNet-v2 encoder using SMA, CD31 and nucleus (DAPI/DRAQ5) markers. For cases in which results were unsatisfactory (that is, more than 10% of the signal area was lost or more than 10% of the detected area did not belong to the correct compartment or cell population), masks were corrected manually.
Cell typing
Cell classification was performed based on the expression of each marker and the distribution of markers in the cell’s contour, such as the cells’ mean marker intensity. Cell typing was performed across the entire set of cells in the analysis. A batch-balanced k-nearest neighbor tree57 was constructed, and further clustering was performed using the Leiden algorithm58 along with the Wilcoxon signed-rank test to assign specific cell types to the clusters. To ensure cell typing accuracy, a ‘gating’ strategy was used to verify that detected clusters of cells with a given phenotype did not contain cells lacking expression of the specified marker or any other inconsistencies with the assigned phenotype. This was achieved by comparing marker expression levels to a ‘threshold’ value, which was determined either through manual review of the image data or by inferring the value from the distribution of the given cluster.
A two-step strategy was then applied for cell typing. In the first step, major cell types were identified, including B cells (cells expressing CD45 along with one of the markers CD19, CD20 or CD79a), T cells (CD45+CD3+CD4+, CD45+CD3+CD8+ and CD45+CD3+FOXP3+ cells), macrophages (CD68+ cells), endothelial cells (CD31+) and stromal cells (FN1+ or SMA+). These major cell types were cross-validated by two experienced pathologists to ensure alignment with the observed fluorescent signals. In the second step, cell subtypes were identified: B cells were subdivided based on co-expression of CD10, CD21, CD23 or Bcl2; T cells were classified based on co-expression of PD-1, ICOS (for CD8+ and CD4+ cells) or Granzyme B (for CD8+ cells); and macrophages were subdivided based on co-expression of CD206. After identifying the detailed cell subtypes, all cell classifications were re-evaluated by pathologists to confirm accuracy.
Statistics and reproducibility
Comparisons of cellular and CN compositions between EBV-positive and EBV-negative nodal lesions were performed using the Wilcoxon rank sum test. Comparisons of cellular and CN compositions among different tumor anatomical sites were done with the Kruskal–Wallis test, with intergroup comparisons performed using Dunn’s test for multiple comparisons, applying the Holm method to adjust P values. For comparisons of signature score expressions between EBV-positive and EBV-negative nodal lesions, the Student’s t-test was conducted. Pairwise t-tests were used for comparisons of signature score expressions among different tumor sites, with the Benjamini–Hochberg method used to control the false discovery rate. For comparisons of individual gene expression levels among different clinical groups, the P values were obtained using the FindAllMarkers function in Seurat46, using the default Wilcoxon rank sum test. For all comparisons described above, two-sided P values were reported, and P < 0.05 was considered statistically significant. Unless otherwise specified, all bioinformatic and statistical analyses in this study were implemented in the R statistical environment (v4.2.0; https://www.r-project.org).
Sample size of the study was determined based on the sample availability. No statistical methods were used to determine sample size. All statistical comparisons were made using data from all applicable cases, with sample sizes described in the Results section and assignment of patients for each comparison group outlined in Supplementary Table 1. No data were excluded from the analyses, and the experiments were not randomized. Investigators were blinded to sample characteristics during data acquisition and processing. Individual cases were selected for display in figures based on their alignment with the overall trend from the full series.
Whole-exome sequencing and bulk RNA sequencing
A full description of the methods for whole-exome sequencing and RNA sequencing can be found in the Supplementary Information.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The CosMx SMI data can be accessed through the Gene Expression Omnibus (GEO) under accession number GSE289194. Assignment of patients to groups for comparisons based on EBV status or anatomical site is provided in Supplementary Table 1. Raw whole-exome and RNA sequencing data are available through the European Genome–Phenome Archive (EGA), under accession number EGAS50000001146.
Code availability
All data analysis was performed using publicly available software, packages and tools, as described in the Methods section. The computational code used in the analysis is available at https://github.com/Coolgenome/Lymphoma-spatial (https://doi.org/10.5281/zenodo.15675991)59, where a detailed description of the analysis flow, including specific code and functions used in the study, is provided to ensure transparency and reproducibility of the results.
References
Crump, M. et al. Outcomes in refractory diffuse large B-cell lymphoma: results from the international SCHOLAR-1 study. Blood 130, 1800–1808 (2017).
Abramson, J. S. et al. Lisocabtagene maraleucel for patients with relapsed or refractory large B-cell lymphomas (TRANSCEND NHL 001): a multicentre seamless design study. Lancet 396, 839–852 (2020).
Westin, J. R. et al. Survival with axicabtagene ciloleucel in large B-cell lymphoma. N. Engl. J. Med. 389, 148–157 (2023).
Thieblemont, C. et al. Epcoritamab, a novel, subcutaneous CD3xCD20 bispecific T-cell-engaging antibody, in relapsed or refractory large B-cell lymphoma: dose expansion in a phase I/II trial. J. Clin. Oncol. 41, 2238–2247 (2023).
Dickinson, M. J. et al. Glofitamab for relapsed or refractory diffuse large B-cell lymphoma. N. Engl. J. Med. 387, 2220–2231 (2022).
Steen, C. B. et al. The landscape of tumor cell states and ecosystems in diffuse large B cell lymphoma. Cancer cell 39, 1422–1437.e10 (2021).
Kotlov, N. et al. Clinical and biological subtypes of B-cell lymphoma revealed by microenvironmental signatures. Cancer Discov. 11, 1468–1489 (2021).
Ye, X. et al. A single-cell atlas of diffuse large B cell lymphoma. Cell Rep. 39, 110713 (2022).
Roider, T. et al. Dissecting intratumour heterogeneity of nodal B-cell lymphomas at the transcriptional, genetic and drug-response levels. Nat. Cell Biol. 22, 896–906 (2020).
Roider, T. et al. Multimodal and spatially resolved profiling identifies distinct patterns of T cell infiltration in nodal B cell lymphoma entities. Nat. Cell Biol. 26, 478–489 (2024).
Liu, N. et al. Single-cell landscape of primary central nervous system diffuse large B-cell lymphoma. Cell Discov. 9, 55 (2023).
Colombo, A. R. et al. Single-cell spatial analysis of tumor immune architecture in diffuse large B-cell lymphoma. Blood Adv. 6, 4675–4690 (2022).
Reiss, D. J. et al. High-plex imaging and cellular neighborhood spatial analysis reveals multiple immune escape and suppression patterns in diffuse large B-cell lymphoma. Leukemia 38, 1164–1168 (2024).
Wright, K. T. et al. Diffuse large B-cell lymphomas have spatially defined, tumor immune microenvironments revealed by high-parameter imaging. Blood Adv. 7, 4633–4646 (2023).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Li, X. et al. Large B cell lymphoma microenvironment archetype profiles. Cancer Cell 43, 1347–1364 (2025).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Gulati, G. S., D’Silva, J. P., Liu, Y., Wang, L. & Newman, A. M. Profiling cell identity and tissue architecture with single-cell and spatial transcriptomics. Nat. Rev. Mol. Cell Biol. 26, 11–31 (2025).
Nirmal, A. J. et al. The spatial landscape of progression and immunoediting in primary melanoma at single-cell resolution. Cancer Discov. 12, 1518–1541 (2022).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Alduaij, W. et al. Molecular determinants of clinical outcomes in a real-world diffuse large B-cell lymphoma population. Blood 141, 2493–2507 (2023).
Malpica, L. et al. EBV-positive diffuse large B-cell lymphoma, not otherwise specified: 2022 update on diagnosis, risk-stratification, and management. Am. J. Hematol. 97, 951–965 (2022).
Bashir, R., Chamberlain, M., Ruby, E. & Hochberg, F. H. T-cell infiltration of primary CNS lymphoma. Neurology 46, 440–444 (1996).
Leivonen, S. K. et al. T-cell inflamed tumor microenvironment predicts favorable prognosis in primary testicular lymphoma. Haematologica 104, 338–346 (2019).
Alaggio, R. et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 36, 1720–1748 (2022).
Keane, C. et al. The tumour microenvironment is immuno-tolerogenic and a principal determinant of patient outcome in EBV-positive diffuse large B-cell lymphoma. Eur. J. Haematol. 103, 200–207 (2019).
Bourbon, E. et al. Clinicopathological features and survival in EBV-positive diffuse large B-cell lymphoma not otherwise specified. Blood Adv. 5, 3227–3239 (2021).
Barros, M. H. M., Vera-Lozada, G., Segges, P., Hassan, R. & Niedobitek, G. Revisiting the tissue microenvironment of infectious mononucleosis: identification of EBV infection in T cells and deep characterization of immune profiles. Front. Immunol. 10, 146 (2019).
Qiu, M. Z. et al. Dynamic single-cell mapping unveils Epstein‒Barr virus-imprinted T-cell exhaustion and on-treatment response. Signal Transduct. Target Ther. 8, 370 (2023).
Goodman, A., Patel, S. P. & Kurzrock, R. PD-1–PD-L1 immune-checkpoint blockade in B-cell lymphomas. Nat. Rev. Clin. Oncol. 14, 203–220 (2017).
Nikolaenko, L., Herrera, A. F., Budde, E., Song, J. Y. & Shouse, G. EBV positive DLBCL is effectively treated with commercial CAR T cell therapy but is associated with higher rates of severe immune effector cell associated neurologic syndrome (ICANS): a single institution experience. Blood 138, 3862–3863 (2021).
Nair, R. et al. CD19 CAR-T outcomes in patients with EBV-positive DLBCL. Blood 140, 3800–3802 (2022).
Chen, X. et al. PD-1 blockade combined with R-CHOP in patients with newly diagnosed EBV-positive diffuse large B-cell lymphoma: a retrospective analysis. Blood 142, 4506–4507 (2023).
Roschewski, M., Phelan, J. D. & Jaffe, E. S. Primary large B-cell lymphomas of immune-privileged sites. Blood 144, 2593–2603 (2024).
Schmitz, R. et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N. Engl. J. Med. 378, 1396–1407 (2018).
Chapuy, B. et al. Targetable genetic features of primary testicular and primary central nervous system lymphomas. Blood 127, 869–881 (2016).
Cook, M. R. et al. Toxicity and efficacy of CAR T-cell therapy in primary and secondary CNS lymphoma: a meta-analysis of 128 patients. Blood Adv. 7, 32–39 (2023).
Godfrey, J. K. et al. Glofitamab stimulates immune cell infiltration of CNS tumors and induces clinical responses in secondary CNS lymphoma. Blood 144, 457–461 (2024).
Graber, J. J., Plato, B., Mawad, R. & Moore, D. J. Pembrolizumab immunotherapy for relapsed CNS Lymphoma. Leuk. Lymphoma 61, 1766–1768 (2020).
Yi, J. H., Kim, S. J., Kim, S. A., Jung, J. & Yoon, D. H. Nivolumab in relapsed or refractory primary CNS lymphoma: multicenter, retrospective study. Blood 142, 1772–1773 (2023).
Gavrilenko, A. N. et al. Nivolumab in primary CNS lymphoma and primary testicular lymphoma with CNS involvement: single center experience. Blood 136, 4 (2020).
Shmatko, A., Ghaffari Laleh, N., Gerstung, M. & Kather, J. N. Artificial intelligence in histopathology: enhancing cancer research and clinical oncology. Nat. Cancer 3, 1026–1038 (2022).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Schmauch, B. et al. A deep learning model to predict RNA-seq expression of tumours from whole slide images. Nat. Commun. 11, 3877 (2020).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Gavish, A. et al. Hallmarks of transcriptional intratumour heterogeneity across a thousand tumours. Nature 618, 598–606 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2020).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Cheng, S. et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell 184, 792–809.e23 (2021).
Wang, R. et al. Evolution of immune and stromal cell states and ecotypes during gastric adenocarcinoma progression. Cancer Cell 41, 1407–1426.e9 (2023).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Wei, R. et al. Spatial charting of single-cell transcriptomes in tissues. Nat. Biotechnol. 40, 1190–1199 (2022).
Jin, S. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Wang, Y. et al. iTALK: an R package to characterize and illustrate intercellular communication. Preprint at BioRxiv https://doi.org/10.1101/507871 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Dai, Y., Wang, L. Code for Multi-modal spatial characterization of tumor immune microenvironments identifies targetable inflammatory niches in diffuse large B-cell lymphoma. Zenodo https://doi.org/10.5281/zenodo.15675991 (2025).
Acknowledgements
This work was chiefly supported by the Futcher family. M.R.G. is a Scholar of the Leukemia and Lymphoma Society. A.K. is supported by a Lymphoma Research Foundation fellowship. L.W. was in part supported by the James P. Allison Institute and the Institute for Data Science in Oncology at the University of Texas MD Anderson Cancer Center.
Author information
Authors and Affiliations
Contributions
Y.D. analyzed data and prepared the manuscript. A.K. performed experiments and prepared the manuscript. D.C., A.W., J.H., D.W., A.P., C.J., A.S., K.N., A.B., A.C., J.W., C.F., K.K., T.Z. and F.V. collected data and samples. X.L. and Y.L. analyzed data. L.W. supervised the study and prepared the manuscript. M.G. conceived of the study, supervised the study and prepared the manuscript. All authors read and agreed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.C. reports research funding from Genmab, BMS, MorPhosys, Ipsen and ONO Pharmaceutical; honoraria from ONO Pharmaceutical and BeiGene. D.W., A.P., C.J., A.S., K.N. and A.B. are employees of BostonGene Corporation. J.W. reports research funding or advisory board membership with AbbVie, ADC Therapeutics, Allogene, AstraZeneca, BMS, Genentech, Janssen, Kite/Gilead, Morphosys/Incyte, Novartis, Nurix, Pfizer and Regeneron. C.R.F. reports consulting for AbbVie, Bayer, BeiGene, Celgene, Denovo Biopharma, Foresight Diagnostics, Genentech/Roche, Genmab, Gilead, Karyopharm, N-Power Medicine, Pharmacyclics/Janssen, SeaGen and Spectrum; research funding from 4D, AbbVie, Acerta, Adaptimmune, Allogene, Amgen, Bayer, Celgene, Cellectis EMD, Gilead, Genentech/Roche, Guardant, Iovance, Janssen Pharmaceutical, Kite, Morphosys, Nektar, Novartis, Pfizer, Pharmacyclics, Sanofi, Takeda, TG Therapeutics, Xencor, Ziopharm, Burroughs Wellcome Fund, Eastern Cooperative Oncology Group, National Cancer Institute, V Foundation and the Cancer Prevention and Research Institute of Texas; and stock options in Foresight Diagnostics and N-Power Medicine. M.R.G. reports research funding from Sanofi, Kite/Gilead, AbbVie and Allogene; Advisory board or consulting for AbbVie, Allogene, Arvinas, Desai, Bristol Myers Squibb and Johnson & Johnson; honoraria from Daiichi Sankyo and DAVA Oncology; and stock ownership of Melbridge Therapeutics and Shenandoah Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Ethel Cesarman and Dimitris Sidiropoulos for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Correlation between CosMX SMI and CODEX for major cell types.
a, Schematic representation of marker-based cell typing strategy for CODEX. b, Correlation scatter plots showing the enumeration of major cell types on the CosMx and CODEX platforms. Cell type enumeration for CosMx was performed by cluster-based classification within the field-of-view. Cell type enumeration for CODEX was performed by marker-based classification including the entire core. CosMx and CODEX were not performed on sequential sections, therefore represent different cells separated along the Z dimension. P-values were calculated with a Spearman rank-order correlation test. Shaded area represents the 95% confidence interval for the correlation line. c, Sub-segmented barplots representing the proportion of four major cell types identified by CosMx SMI and CODEX in tissue sections from the same set of samples. The two platforms showed overall high concordance in major cell type identification.
Extended Data Fig. 2 Expression of lineage markers in each cell type.
Bubble plot of key lineage markers expressed in each major cell type; related to Fig. 2a.
Extended Data Fig. 3 Unique spatial niches identified in CosMx SMI and CODEX.
a, Ro/e plot showing the enrichment or depletion of different cell states in each spatial niche. C5_T, C1_PC_IgG, C6_TAM_APOE_C1Q and C11_FRC were highly enriched in CN1 ~ 4, respectively. C0_Tumor-B were highly enriched in CN5 and CN6. No obvious enrichment of any single cell state was observed in CN7. b, Pie chart representation of the proportion of cells in the CosMx SMI dataset that resided in each spatial niche. c, Additional examples of cellular niches, corresponding to Fig. 2g. d, Representative CODEX images of tissue sections from the first five samples shown in Fig. 2g, with spatial enrichment of T cells, PCs, myeloid cells, stromal cells, and B cells, respectively.
Extended Data Fig. 4 Cell co-localization network in seven different spatial niches.
In each graph, every node represents a cell state, and the width of the edge connecting two cell states represents the proximity of the two cell states within the specific niche.
Extended Data Fig. 5 T cell neighborhood compositions, cytotoxicity, and chemotaxis signals in different spatial niches.
a, Sub-segmented barplots showing the neighborhood compositions of T cells in each spatial niche. b, The expression of cytotoxicity markers GZMB, GZMH and NKG7 in T cells from the same CN1 dominated FOV as Fig. 3b. c, The expression of cytotoxicity markers in T cells from a representative FOV dominated by CN3. d, The expression of T cell chemotaxis molecules CXCL10 and CCL2 in APOE+ C1Q+ TAM in the same CN1 dominated FOV as Fig. 3b. e, The expression of T cell chemotaxis molecules in APOE+ C1Q+ TAM, tumor-B cells and FRC in the same CN3 dominated FOV as panel c.
Extended Data Fig. 6 Niche-specific cell-cell communications effecting T cell chemotaxis and functionality.
a, The activity of T cell chemotaxis signaling axis in different spatial niches, with ligands coming from APOE+ C1Q+ TAM, tumor-B cells, FRC or HEV. b, A representative FOV image from CosMx SMI showing abundant CCL21/19 : CCR7 chemotaxis signaling between HEV and T cells in CN1. c, The activity of MHC II : CD4 signaling axis between APOE+ C1Q+ TAM and T cells in different spatial niches, with overall strongest signals in CN1. d, Normalized interaction intensity in each sample for PD-L1 : PD-1 interaction between APOE+ C1Q+ TAM (C6) and T cells (C5) [left] and between tumor-B cells (C0) and T cells (C5) [right] in each spatial niche, the former strongest in CN3 and the latter strongest in CN5. P-values were calculated with a Kruskal-Wallis test. e, A representative CN5 dominated FOV showing the PD-L1 : PD-1 interaction between tumor-B cells and T cells.
Extended Data Fig. 7 The expression profile of tumor-B cells in different spatial niches.
a, The expression of G1/S cell cycle signature score in tumor-B cells from different spatial niches. P-values were calculated with a one-way ANOVA. b, The expression of proliferation marker STMN1 and anti-apoptotic molecule BCL2 in tumor-B cells from the same FOV as Fig. 4b. c, The expression of proliferation and anti-apoptotic markers in tumor-B cells from a CN6 dominated FOV. d, The expression of therapeutic targets and genes of interest in tumor-B cells from different spatial niches. Dashed boxes represent adjusted P < 0.05 for the given cell type compared to all other cell types. e, Example of therapeutic targets co-expressed in tumor-B cells from the same FOV as Fig. 4b. f, The activity of CXCL12 : CXCR4 interaction between all TME cells and tumor-B cells in CN5 and CN6, with senders from the whole neighborhood of tumor-B cells, including cells both within and outside of the tumor-B cell-resident niche. g, The activity of CD40LG : CD40 interaction between all T cells and tumor-B cells in different spatial niches. h, The expression of immune checkpoint markers in tumor-B cells from different spatial niches, with strongest expressions observed in CN1-resident tumor-B cells.
Extended Data Fig. 8 Spatially resolved cellular landscape of nodal EBV+ DLBCL.
a, Boxplot representation of the proportion of cells within each FOV that reside within each niche, comparing EBV positive and negative nodal DLBCLs. Center line shows the median, boxes represent quartiles 1 – 3 and the whiskers represent 1.5 times the interquartile range. P-values were calculated using a Wilcoxon rank sum test. b, The activities of T cell chemotaxis signaling axes in EBV positive and negative nodal DLBCLs. Two representative axes were shown: CCL5 : CCR5, CXCL9 : CXCR3, with ligands from tumor-B cells, APOE+ C1Q+ TAM and FRC. c, The activities of B2M : CD3 interaction between tumor-B cells, APOE+ C1Q+ TAM, or FRC and T cells in EBV positive and negative nodal lesions. d, The activities of MHC II : CD4 interaction between APOE+ C1Q+ TAM and T cells in EBV positive and negative nodal lesions. e, Boxplot representation of tumor-B and T cell populations in the neighborhood of tumor-B cells from EBV positive and negative nodal lesions. Center line shows the median, boxes represent quartiles 1 – 3 and the whiskers represent 1.5 times the interquartile range. P-values were calculated using a Wilcoxon rank sum test. f, CODEX validation corresponding to Fig. 5j, showing the distinct cellular compositions of EBV positive and negative nodal lesions.
Extended Data Fig. 9 Spatially resolved cellular landscape of B cell lymphoma developed in different anatomical sites.
a, Boxplot representation of the proportion of spatial niches in tumors from different anatomical sites. Center line shows the median, boxes represent quartiles 1 – 3 and the whiskers represent 1.5 times the interquartile range. P-values were calculated using a Kruskal-Wallis test with Dunn’s post-hoc test for pairwise comparisons. b, The neighborhood structures of tumor-B cells from different anatomical sites. c, The corresponding CODEX images of samples shown in Fig. 6f, indicating the distinct cellular compositions in nodal, IPS, and EN-O lesions. d, The expression of G1/S cell cycle signature score from nodal, IPS, and EN-O lesions. P-values were calculated using a pairwise Student’s T test. e, Representative proliferation markers in T cells from nodal, IPS, and EN-O lesions. P-values were calculated comparing IPS to all other tumors using the FindAllMarkers function of Seurat, based upon a Wilcoxon rank sum test with Bonferroni correction. f, Decomposition of Fig. 7j, showing the PD-L1 : PD-1 interaction between each individual cell state and T cells in nodal, IPS, and EN-O lesions. g, Decomposition of Fig. 7k, showing the Galactin-9 : TIM-3 interaction between each individual cell state and T cells in nodal, IPS, and EN-O lesions.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Supplementary Figs. 1–4.
Supplementary Table (download XLSX )
Supplementary Tables 1–13.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dai, Y., Kizhakeyil, A., Chihara, D. et al. Multi-modal spatial characterization of tumor immune microenvironments identifies targetable inflammatory niches in diffuse large B cell lymphoma. Nat Genet 57, 2715–2727 (2025). https://doi.org/10.1038/s41588-025-02353-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02353-5