Abstract
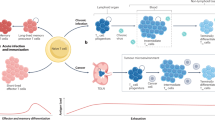
CD8+ T cells responding to chronic infections or tumors acquire an ‘exhausted’ state associated with elevated expression of inhibitory receptors, including PD-1, and impaired cytokine production. Exhausted T cells are continuously replenished by T cells with precursor characteristics that self-renew and depend on the transcription factor TCF1; however, their developmental requirements are poorly understood. In the present study, we demonstrate that high antigen load promoted the differentiation of precursor T cells, which acquired hallmarks of exhaustion within days of infection, whereas early effector cells retained polyfunctional features. Early precursor T cells showed epigenetic imprinting characteristic of T cell receptor–dependent transcription factor binding and were restricted to the generation of cells displaying exhaustion characteristics. Transcription factors BACH2 and BATF were key regulators with opposing functions in the generation of early precursor T cells. Overall, we demonstrate that exhaustion manifests first in TCF1+ precursor T cells and is propagated subsequently to the pool of antigen-specific T cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data are available in the main text or the Extended Data materials. All materials used in the present study are available upon request from the lead authors. Sequencing data generated for this study have been deposited in the Gene Expression Omnibus database with accession code GSE142687.
References
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Hashimoto, M. et al. CD8 T cell exhaustion in chronic infection and cancer: opportunities for interventions. Annu. Rev. Med. 69, 301–318 (2018).
Kuchroo, V. K., Anderson, A. C. & Petrovas, C. Coinhibitory receptors and CD8 T cell exhaustion in chronic infections. Curr. Opin. HIV AIDS 9, 439–445 (2014).
Klebanoff, C. A., Gattinoni, L. & Restifo, N. P. CD8+ T-cell memory in tumor immunology and immunotherapy. Immunol. Rev. 211, 214–224 (2006).
Baitsch, L. et al. Exhaustion of tumor-specific CD8+ T cells in metastases from melanoma patients. J. Clin. Invest. 121, 2350–2360 (2011).
Ahmadzadeh, M. et al. Tumor antigen–specific CD8 T cells infiltrating the tumor express high levels of PD-1 and are functionally impaired. Blood 114, 1537–1544 (2009).
Lugli, E., Galletti, G., Boi, S. K. & Youngblood, B. A. Stem, effector, and hybrid states of memory CD8+ T cells. Trends Immunol. 41, 17–28 (2020).
Kallies, A., Zehn, D. & Utzschneider, D. T. Precursor exhausted T cells: key to successful immunotherapy? Nat. Rev. Immunol. 20, 128–136 (2020).
Utzschneider, D. T. et al. High antigen levels induce an exhausted phenotype in a chronic infection without impairing T cell expansion and survival. J. Exp. Med. 213, 1819–1834 (2016).
Angelosanto, J. M., Blackburn, S. D., Crawford, A. & Wherry, E. J. Progressive loss of memory T cell potential and commitment to exhaustion during chronic viral infection. J. Virol. 86, 8161–8170 (2012).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8+ T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Man, K. et al. Transcription factor IRF4 promotes CD8+ T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity 47, 1129–1141.e5 (2017).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Wu, T. et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
He, R. et al. Follicular CXCR5-expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Leong, Y. A. et al. CXCR5+ follicular cytotoxic T cells control viral infection in B cell follicles. Nat. Immunol. 17, 1187–1196 (2016).
Hudson, W. H. et al. Proliferating transitory T cells with an effector-like transcriptional signature emerge from PD-1+ stem-like CD8+ T cells during chronic infection. Immunity 51, 1043–1058.e4 (2019).
Zander, R. et al. CD4+ T cell help is required for the formation of a cytolytic CD8+ T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042.e4 (2019).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e20 (2018).
Brummelman, J. et al. High-dimensional single cell analysis identifies stem-like cytotoxic CD8+ T cells infiltrating human tumors. J. Exp. Med. 215, 2520–2535 (2018).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Menner, A. J. et al. Id3 controls cell death of 2B4+ virus-specific CD8+ T cells in chronic viral infection. J. Immunol. 195, 2103–2114 (2015).
Kaech, S. M. & Wherry, E. J. Heterogeneity and cell-fate decisions in effector and memory CD8+ T cell differentiation during viral infection. Immunity 27, 393–405 (2007).
Pauken, K. E. et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science 354, 1160–1165 (2016).
Sen, D. R. et al. The epigenetic landscape of T cell exhaustion. Science 354, 1165–1169 (2016).
Oestreich, K. J., Yoon, H., Ahmed, R. & Boss, J. M. NFATc1 regulates PD-1 expression upon T cell activation. J. Immunol. 181, 4832–4839 (2008).
Lynn, R. C. et al. c-Jun overexpression in CAR T cells induces exhaustion resistance. Nature 576, 293–300 (2019).
Roychoudhuri, R. et al. BACH2 regulates CD8+ T cell differentiation by controlling access of AP-1 factors to enhancers. Nat. Immunol. 17, 851–860 (2016).
Sidwell, T. et al. Attenuation of TCR-induced transcription by Bach2 controls regulatory T cell differentiation and homeostasis. Nat. Commun. 11, 252 (2020).
Wei, J. et al. Targeting REGNASE-1 programs long-lived effector T cells for cancer therapy. Nature 576, 471–476 (2019).
Xin, G. et al. A critical role of IL-21-induced BATF in sustaining CD8-T-cell-mediated chronic viral control. Cell Rep. 13, 1118–1124 (2015).
Grusdat, M. et al. IRF4 and BATF are critical for CD8+ T-cell function following infection with LCMV. Cell Death Differ. 21, 1050–1060 (2014).
Utzschneider, D. T. et al. T cells maintain an exhausted phenotype after antigen withdrawal and population reexpansion. Nat. Immunol. 14, 603–610 (2013).
Ghoneim, H. E. et al. De novo epigenetic programs inhibit PD-1 blockade-mediated T cell rejuvenation. Cell 170, 142–157.e19 (2017).
Blank, C. U. et al. Defining ‘T cell exhaustion’. Nat. Rev. Immunol. 19, 665–674 (2019).
Chen, Z. et al. TCF-1-centered transcriptional network drives an effector versus exhausted CD8 T cell-fate decision. Immunity 51, 840–855.e5 (2019).
Diao, B. et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front. Immunol. 11, 827 (2020).
Miyazaki, M. et al. The opposing roles of the transcription factor E2A and its antagonist Id3 that orchestrate and enforce the naive fate of T cells. Nat. Immunol. 12, 992–1001 (2011).
Kometani, K. et al. Repression of the transcription factor Bach2 contributes to predisposition of IgG1 memory B cells toward plasma cell differentiation. Immunity 39, 136–147 (2013).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Itoh-Nakadai, A. et al. The transcription repressors Bach2 and Bach1 promote B cell development by repressing the myeloid program. Nat. Immunol. 15, 1171–1180 (2014).
Puglielli, M. T. et al. In vivo selection of a lymphocytic choriomeningitis virus variant that affects recognition of the GP33-43 epitope by H-2Db but not H-2Kb. J. Virol. 75, 5099–5107 (2001).
Battegay, M. et al. Quantification of lymphocytic choriomeningitis virus with an immunological focus assay in 24- or 96-well plates. J. Virol. Methods 33, 191–198 (1991).
Blattman, J. N. et al. Estimating the precursor frequency of naive antigen-specific CD8 T cells. J. Exp. Med. 195, 657–664 (2002).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
McCarthy, D. J. & Smyth, G. K. Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 25, 765–771 (2009).
Wu, D. et al. ROAST: rotation gene set tests for complex microarray experiments. Bioinformatics 26, 2176–2182 (2010).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank T. Mason for technical support and T. Gebhardt, S. Nutt and members of the Kallies lab for discussions. This work was funded by the National Health and Medical Research Council (project grant no. 1085151 and Senior Research Fellowship no. 1139607 to A.K.), the Swiss National Science Foundation (fellowship nos. P300PA_177907 to D.T.U., P400PM_180807 to S.S.G. and P300PB_177934 to P.M.G.) and the Novartis Foundation for Medical–Biological Research (fellowship to S.S.G.). D.T.U. is a Special Fellow of the Leukemia and Lymphoma Society (fellowship no. 3387-19). W.S .is supported by a WEHI Centenary Fellowship funded by a donation from CSL Ltd. We acknowledge the Melbourne Cytometry Platform for provision of flow cytometry services.
Author information
Authors and Affiliations
Contributions
D.T.U., S.S.G. and A.K. conceived the study, designed experiments, interpreted the results and wrote the manuscript. D.T.U. and S.S.G. performed the experiments with support from R.G. and P.M.G. A.V. generated the BACH2-overexpressing vector. D.C. and W.S. analyzed the sequencing data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports available. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 High levels of antigen promote precursor T cell generation and PD-1 and TOX expression.
a, Experimental scheme referring to b–d and Fig. 1a, b. Congenically marked P14 T cells were labelled with CellTrace Violet (CTV) and transferred into naïve mice, which were infected with either acute LCMV Armstrong (Arm) or chronic LCMV Docile (Doc). Spleens were analysed 48, 62 and 69 hours post infection (p.i.). For each experiment, all 12 biological replicates (four mice at 3 timepoints) were concatenated into one file in FlowJo. b, Representative FACS plots showing CTV-dilutions of P14 T cells from concatenated samples. c, d, Representative FACS plots showing PD-1 (c) and TOX expression (d) versus CTV-dilution (top). Samples from Docile infected mice are in black and from Armstrong in green. Mean fluorescent intensity (MFI) combined from two experiments (bottom). Data are representative of three individual experiments. Error bars indicate SEM. e–g, P14 T cells were transferred into congenically marked naïve recipient mice, which were infected with either LCMV Armstrong or Docile. e, Gating strategy for identification of P14 T cells. f, g, Representative histograms depicting PD-1 (f) and TOX (g) among TCF1+ precursor (TP, solid line, left) and TIM-3+ effector (TE, dashed line, right) P14 T cells on day 5 p.i. with high dose Docile (black) or Armstrong (green); grey represents host CD8+ T cells. Data are representative of at least three independent experiments with five mice per group.
Extended Data Fig. 2 Memory T cells in acutely-resolved infection express low levels of TOX and high amounts of IFN-γ.
P14 T cells were transferred into congenically marked naïve recipient mice, which were infected with either LCMV Armstrong or Docile and analysed on day 21 p.i. a, TOX expression of TCF1+ precursor (TP, black solid dots) and TIM-3+ effector (TE, black open dots) P14 T cells from Docile compared to memory P14 T cells from Armstrong (green solid dots) infection. b, Frequencies of IFN-γ producing TP and TE P14 T cells from Docile compared to memory P14 T cells from Armstrong after ex vivo re-stimulation with LCMV-derived gp33 peptide. TP and TE cell segregation based on Ly108 and TIM-3 expression. Symbols represent individual mice; lines connect P14 T cells within the same host. Data are representative of at least three independent experiments with at least four mice per group. Statistical analysis was performed with unpaired Student’s t test (two-tailed). ****p < 0.0001; ***p < 0.001.
Extended Data Fig. 3 Phenotypic characterization of ID3+ and ID3- T cells in chronic and acute infection.
Congenically marked Id3GFP P14 T cells were transferred into naïve mice, which were then infected with either LCMV Armstrong or Docile. a, TCF1 versus TIM-3 expression of P14 T cells from spleens on day 5 post LCMV Docile (top, black) or LCMV Armstrong (bottom, green) infection. b, ID3 versus TIM-3 expression of P14 T cells from spleens on day 5 post Docile or Armstrong infection. c, TCF1 versus TIM-3 expression of ID3+ TP and TIM-3+ TE cells from b. d, Expression of Granzyme B (GzmB), 2B4 and CD39 among ID3+ TP and TIM-3+ TE cells on day 5 post Docile or Armstrong infection. Histograms of ID3+TIM-3- TP cells in solid lines, ID3-TIM-3+ TE cells in dashed lines and host CD8+ T cells in grey are shown. Data are representative of at least two individual experiments with at least four mice each.
Extended Data Fig. 4 Transcriptional differences between ID3+ and ID3- P14 T cells in acute and chronic infection.
Congenically marked Id3GFP P14 T cells were transferred into naïve mice, which were then infected with either LCMV Armstrong or Docile. ID3+ and ID3- P14 T cells from Armstrong infected mice were FACS purified for RNAseq analysis as described in Fig. 4. a, b, Volcano plots show differentially expressed (DE) genes between ID3+ and ID3- P14 T cells obtained from Armstrong infected mice on day 5 p.i. (a) and day 21 p.i. (b). c, Number of DE genes in all comparisons including comparisons shown in Fig. 4a, b. Number of genes upregulated in ID3+ compared to ID3- P14 T cells are indicated in red and genes downregulated in ID3+ P14 T cells are indicated in blue.
Extended Data Fig. 5 Generation of a core exhaustion T cell gene signature.
Congenically marked Id3GFP P14 T cells were transferred into naïve mice, which were infected with either LCMV Armstrong or Docile. ID3+ and ID3- P14 T cells were FACS purified for RNAseq analysis as described in Fig. 4. a, b, Volcano plots show genes differentially expressed (DE) between ID3+ (a) and ID3- (b) P14 T cells from day 21 Docile and Armstrong infections. c, Venn diagram shows numbers of DE genes common (core exhaustion signature) or unique to each comparison. Numbers in red highlight genes upregulated in Docile samples and numbers in blue genes downregulated in Docile. d, Heatmap established by unsupervised clustering of all generated samples based on the ‘dysfunctional vs polyfunctional’ signature defined by Alfei et al. 2019 (ref. 11). Black (Docile) and green (Armstrong) boxes on top highlight origin of cells. Dotted squares highlight main two clusters; green square = polyfunctional phenotype, black square = exhausted phenotype. e, Hierarchical tree of unsupervised clustered heatmap. f, Gene set enrichment analysis of Docile derived day 5 TP versus TE cells using the ‘dysfunctional vs polyfunctional’ signature defined by Alfei et al. 2019 (ref. 11). Barcode plots based on ROAST tests including p values from the tests are shown; signature genes upregulated are shown in red and genes downregulated in blue.
Extended Data Fig. 6 The Pdcd1 gene locus displays early enhanced accessibility in T cells responding to LCMV Docile infection.
Congenically marked Id3GFP P14 T cells were transferred into naïve mice, which were then infected with either LCMV Armstrong or Docile. ID3+ and ID3- P14 T cells were FACS purified at days 5 and 21 p.i. and ATAC sequencing was performed. Chromatin accessibility of the Pdcd1 locus is shown. Red arrows highlight regions that are differentially accessible between T cells obtained from Armstrong (green tracks) compared to Docile infected mice (black tracks). ATAC seq analysis was performed with two experimental replicates at each time point. Open chromatin tracks are based on merged replicates.
Extended Data Fig. 7 BACH2 and BATF regulate early precursor T cell differentiation.
a, Congenically marked Id3GFP P14 T cells were transferred into naïve recipient mice, which were then infected with LCMV Docile. 5 days p.i., ID3+ TP and ID3- TE P14 T cells were purified and chromatin accessibility determined by ATAC sequencing. Chromatin accessibility of Bach2 locus of TP and TE cells. Grey box highlights regions that are differentially accessible comparing the two samples. b, c Bach2fl/flCd4Cre (Bach2-/-) and Cd4Cre (Control) mice were infected with Docile and analysed 5 days later. b, Absolute numbers of gp33-tetramer+CD8+ T cells. c, Plots show TCF1 and TIM-3 expression of gp33-tetramer+CD8+ T cells in Bach2-/- and control mice (left), graph shows absolute numbers of TCF1+ TP and TIM-3+ TE cells among gp33-tetramer+CD8+ T cells in spleens (right). d–g, Congenically marked BACH2-deficient (Bach2fl/flCd4Cre, labelled as Bach2-/-) and control P14 T cells (Cd4Cre, labelled as control) (b, c) or Batf-/- and Batf+/+ control P14 T cells (d, e) were co-transferred into naive mice, which were challenged with Armstrong and analysed 5 days later. b, d, TCF1 versus TIM-3 (left) and TCF1+ TP cell frequency (right) in spleens. c, e, Absolute numbers of TCF1+ TP and TIM-3+ TE cells. Numbers indicate fold increase in control compared to Bach2-/- (c) or Batf-/- cells (d). ATAC seq analysis was performed with two experimental replicates at each time point. Open chromatin tracks are based on merged replicates. Symbols in b–e represent individual mice; lines connect P14 T cells within the same host. Data are combined or representative of two independent experiments with at least three mice per group. Statistical analysis was performed with a paired Student’s t test (two-tailed). ****p < 0.0001; ***p < 0.001; **p < 0.01; ns, not significant. MFI, mean fluorescent intensity. Error bars indicate SEM.
Extended Data Fig. 8 Late precursor T cells retain higher levels of TOX and PD-1 after expansion in acute infection.
a, Experimental scheme. Congenically marked Id3GFP P14 T cells were transferred into naïve mice, which were infected with LCMV Docile. 5 or 26 days p.i., ID3+TIM-3- TP and ID3-TIM-3+ TE P14 T cells were FACS sorted and transferred into naïve mice, which were infected with Armstrong. Donor P14 T cells were analysed 7 days post infection. b, Fold expansion of donor T cell populations. c, d, MFI of TOX (c) and PD-1 (d) of donor P14 T cells. Symbols represent individual mice. Data are representative or combined of two experiments with at least 3 mice per group. Statistical analysis was unpaired Student’s t test (two-tailed). ****p < 0.0001; ***p < 0.001; **p < 0.01; ns, not significant.
Supplementary information
Rights and permissions
About this article
Cite this article
Utzschneider, D.T., Gabriel, S.S., Chisanga, D. et al. Early precursor T cells establish and propagate T cell exhaustion in chronic infection. Nat Immunol 21, 1256–1266 (2020). https://doi.org/10.1038/s41590-020-0760-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-020-0760-z
This article is cited by
-
High efficiency CRISPR knock-in demonstrates that TCF1 is insufficient to reverse T cell exhaustion
Nature Communications (2026)
-
BACH2 dosage establishes the hierarchy of stemness and fine-tunes antitumor immunity in CAR T cells
Nature Immunology (2026)
-
Dual Bcl-2/Bcl-xl inhibition via AZD0466 combines with immune checkpoint blockade to enhance anti-tumour activity
Cell Death & Disease (2026)
-
Regulators of CD8+ T cell exhaustion
Nature Reviews Immunology (2026)
-
Propagation of monocyte exhaustion memory and underlying mechanisms
Cell Communication and Signaling (2025)