Abstract
Antibody-dependent enhancement (ADE) is an important safety concern for vaccine development against dengue virus (DENV) and its antigenically related Zika virus (ZIKV) because vaccine may prime deleterious antibodies to enhance natural infections. Cross-reactive antibodies targeting the conserved fusion loop epitope (FLE) are known as the main sources of ADE. We design ZIKV immunogens engineered to change the FLE conformation but preserve neutralizing epitopes. Single vaccination conferred sterilizing immunity against ZIKV without ADE of DENV-serotype 1–4 infections and abrogated maternal–neonatal transmission in mice. Unlike the wild-type-based vaccine inducing predominately cross-reactive ADE-prone antibodies, B cell profiling revealed that the engineered vaccines switched immunodominance to dispersed patterns without DENV enhancement. The crystal structure of the engineered immunogen showed the dimeric conformation of the envelope protein with FLE disruption. We provide vaccine candidates that will prevent both ZIKV infection and infection-/vaccination-induced DENV ADE.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The crystal structures of ZIKV-sE-Z6 and ZIKV-sE-MutC-Z3L1 have been deposited in the Protein Data Bank (PDB) under accession codes 7BQ5 and 7BPK, respectively. The raw data for scBCR-seq have been deposited at https://www.scidb.cn/en/datalist?tag=1. Source data are provided with this paper. All other data supporting the findings of this study are available within the paper or from the corresponding author upon reasonable request.
References
Knipe, D. M., Howley, P. M. (eds) Fields Virology 6th edn (Lippincott Williams & Wilkins Health, 2013).
Baud, D., Gubler, D. J., Schaub, B., Lanteri, M. C. & Musso, D. An update on Zika virus infection. Lancet 390, 2099–2109 (2017).
Wang, Q. et al. Genetic and biological characterization of Zika virus from human cases imported through Shenzhen Port. Chin. Sci. Bull. 61, 2463–2474 (2016).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 375, 2321–2334 (2016).
Rasmussen, S. A., Jamieson, D. J., Honein, M. A. & Petersen, L. R. Zika virus and birth defects—reviewing the evidence for causality. N. Engl. J. Med. 374, 1981–1987 (2016).
van der Eijk, A. A. et al. Miscarriage associated with Zika virus infection. N. Engl. J. Med. 375, 1002–1004 (2016).
Cao-Lormeau, V. M. et al. Guillain–Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: a case-control study. Lancet 387, 1531–1539 (2016).
Miner, J. J. & Diamond, M. S. Zika virus pathogenesis and tissue tropism. Cell Host Microbe 21, 134–142 (2017).
Ma, W. et al. Zika virus causes testis damage and leads to male infertility in mice. Cell 167, 1511–1524.e10 (2016).
Govero, J. et al. Zika virus infection damages the testes in mice. Nature 540, 438–442 (2016).
Slon-Campos, J. L. et al. A protective Zika virus E-dimer-based subunit vaccine engineered to abrogate antibody-dependent enhancement of dengue infection. Nat. Immunol. 20, 1291–1298 (2019).
Fowler, A. M. et al. Maternally acquired Zika antibodies enhance dengue disease severity in mice. Cell Host Microbe 24, 743–750.e5 (2018).
Richner, J. M. et al. Modified mRNA vaccines protect against Zika virus infection. Cell 168, 1114–1125.e10 (2017).
Rey, F. A., Stiasny, K., Vaney, M. C., Dellarole, M. & Heinz, F. X. The bright and the dark side of human antibody responses to flaviviruses: lessons for vaccine design. EMBO Rep. 19, 206–224 (2018).
Halstead, S. B. & O’Rourke, E. J. Dengue viruses and mononuclear phagocytes. I. Infection enhancement by non-neutralizing antibody. J. Exp. Med. 146, 201–217 (1977).
Katzelnick, L. C. et al. Antibody-dependent enhancement of severe dengue disease in humans. Science 358, 929–932 (2017).
Halstead, S. B. Dengvaxia sensitizes seronegatives to vaccine enhanced disease regardless of age. Vaccine 35, 6355–6358 (2017).
Sridhar, S. et al. Effect of dengue serostatus on dengue vaccine safety and efficacy. N. Engl. J. Med. 379, 327–340 (2018).
Stettler, K. et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 353, 823–826 (2016).
George, J. et al. Prior exposure to Zika virus significantly enhances peak Dengue-2 viremia in rhesus macaques. Sci. Rep. 7, 10498 (2017).
Katzelnick, L. C. et al. Zika virus infection enhances future risk of severe dengue disease. Science 369, 1123–1128 (2020).
York, A. Zika virus enhances dengue risk. Nat. Rev. Microbiol. 18, 605 (2020).
Rogers, T. F. et al. Zika virus activates de novo and cross-reactive memory B cell responses in dengue-experienced donors. Sci. Immunol. 2, eaan6809 (2017).
Dejnirattisai, W. et al. Cross-reacting antibodies enhance dengue virus infection in humans. Science 328, 745–748 (2010).
Beltramello, M. et al. The human immune response to Dengue virus is dominated by highly cross-reactive antibodies endowed with neutralizing and enhancing activity. Cell Host Microbe 8, 271–283 (2010).
Dejnirattisai, W. et al. A new class of highly potent, broadly neutralizing antibodies isolated from viremic patients infected with dengue virus. Nat. Immunol. 16, 170–177 (2015).
Dai, L., Wang, Q., Song, H. & Gao, G. F. Zika virus envelope protein and antibody complexes. Subcell. Biochem. 88, 147–168 (2018).
Barba-Spaeth, G. et al. Structural basis of potent Zika–dengue virus antibody cross-neutralization. Nature 536, 48–53 (2016).
Rouvinski, A. et al. Recognition determinants of broadly neutralizing human antibodies against dengue viruses. Nature 520, 109–113 (2015).
Fibriansah, G. & Lok, S. M. The development of therapeutic antibodies against dengue virus. Antivir. Res. 128, 7–19 (2016).
Wang, Q. et al. Molecular determinants of human neutralizing antibodies isolated from a patient infected with Zika virus. Sci. Transl. Med. 8, 369ra179 (2016).
Zhao, H. et al. Structural basis of Zika virus-specific antibody protection. Cell 166, 1016–1027 (2016).
Abbink, P., Stephenson, K. E. & Barouch, D. H. Zika virus vaccines. Nat. Rev. Microbiol. 16, 594–600 (2018).
Xu, K. et al. Recombinant chimpanzee adenovirus vaccine AdC7-M/E protects against Zika virus infection and testis damage. J. Virol. 92, e01722-17 (2018).
Dai, L. et al. Structures of the Zika virus envelope protein and its complex with a flavivirus broadly protective antibody. Cell Host Microbe 19, 696–704 (2016).
Cherrier, M. V. et al. Structural basis for the preferential recognition of immature flaviviruses by a fusion-loop antibody. EMBO J. 28, 3269–3276 (2009).
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature 536, 474–478 (2016).
Abbink, P. et al. Durability and correlates of vaccine protection against Zika virus in rhesus monkeys. Sci. Transl. Med. 9, eaao4163 (2017).
Smith, S. A. et al. Dengue virus prM-specific human monoclonal antibodies with virus replication-enhancing properties recognize a single immunodominant antigenic site. J. Virol. 90, 780–789 (2016).
Tsai, W. Y. et al. High-avidity and potently neutralizing cross-reactive human monoclonal antibodies derived from secondary dengue virus infection. J. Virol. 87, 12562–12575 (2013).
Lazear, H. M. et al. A mouse model of Zika virus pathogenesis. Cell Host Microbe 19, 720–730 (2016).
Pierson, T. C. et al. The stoichiometry of antibody-mediated neutralization and enhancement of West Nile virus infection. Cell Host Microbe 1, 135–145 (2007).
Crill, W. D. & Chang, G. J. Localization and characterization of flavivirus envelope glycoprotein cross-reactive epitopes. J. Virol. 78, 13975–13986 (2004).
Bennett, K. M. et al. Hybrid flagellin as a T cell independent vaccine scaffold. BMC Biotechnol. 15, 71 (2015).
Deng, Y. Q. et al. A broadly flavivirus cross-neutralizing monoclonal antibody that recognizes a novel epitope within the fusion loop of E protein. PLoS ONE 6, e16059 (2011).
Oliphant, T. et al. Antibody recognition and neutralization determinants on domains I and II of West Nile Virus envelope protein. J. Virol. 80, 12149–12159 (2006).
van der Linden, V. et al. Description of 13 infants born during October 2015–January 2016 with congenital Zika virus infection without microcephaly at birth—Brazil. MMWR Morb. Mortal. Wkly Rep. 65, 1343–1348 (2016).
Fernandez, E. et al. Human antibodies to the dengue virus E-dimer epitope have therapeutic activity against Zika virus infection. Nat. Immunol. 18, 1261–1269 (2017).
Rouvinski, A. et al. Covalently linked dengue virus envelope glycoprotein dimers reduce exposure of the immunodominant fusion loop epitope. Nat. Commun. 8, 15411 (2017).
Crill, W. D. et al. Sculpting humoral immunity through dengue vaccination to enhance protective immunity. Front. Immunol. 3, 334 (2012).
Yang, C. et al. Characterization of two engineered dimeric Zika virus envelope proteins as immunogens for neutralizing antibody selection and vaccine design. J. Biol. Chem. 294, 10638–10648 (2019).
Ye, J., Ma, N., Madden, T. L. & Ostell, J. M. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–W40 (2013).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Read, R. J. Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallogr. D Biol. Crystallogr. 57, 1373–1382 (2001).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Aad, G. et al. Search for new particles in two-jet final states in 7 TeV proton-proton collisions with the ATLAS detector at the LHC. Phys. Rev. Lett. 105, 161801 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We thank the staff of BL17U and BL19U beamlines at National Center for Protein Sciences Shanghai and Shanghai Synchrotron Radiation Facility (Shanghai, China) for assistance during data collection. We are grateful to J. Jia (Institute of Biophysics, Chinese Academy of Sciences (CAS), China) for technical support during BD FACSAria II manipulation. We thank Y. Chen (Institute of Biophysics, CAS, China) for technical support with BIAcore experiments. We thank D. Zhou (Institut Pasteur of Shanghai, CAS, China) for providing us with the AdC7 vector. We thank G. Cheng (Tsinghua University, Beijing, China) and Q. Leng (Institut Pasteur of Shanghai, CAS, China) for providing AG6 mice for breeding. We thank R. Gong (Wuhan Institute of Virology, CAS) for providing ZIKV-sE dimer protein. We thank Y. Chen (China CDC) for breeding of AG6 and Ifnar1−/− mice. We thank C. Liu (Guangxi University, China) for technical support. This work is supported by the National Science and Technology Major Project (2016YFE0205800 and 2020YFA0907100) (L.D. and G.F.G.), the National Natural Science Foundation of China (NSFC) (81991494 and 81801648) (L.D.) and the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB29010202) (G.F.G). L.D. is supported by Youth Innovation Promotion Association CAS (2018113).
Author information
Authors and Affiliations
Contributions
L.D., J.Y. and G.F.G. designed the experiments. L.D., K.X., J.L., Q.H., J.S., Y.H., T.Z., P.G., X.L., H.Y., K.L., Q.W., Y.C. and J.Q. performed the investigations and assays. L.D., K.X., J.Y. and G.F.G. analyzed the data. L.D. and K.X. wrote the manuscript. L.D., K.X., Q.X., Q.W., J.Y. and G.F.G. discussed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.F.G, L.D., J.Y., K.X., Y.H., Q.W., Q.H. and J.L. are listed as the coinventors for the pending patent for the ZIKV vaccines described in this study. The other authors declare no competing interests.
Additional information
Peer review Information Nature Immunology thanks R. Rappuoli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. J. D. K. Wilson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Antibodies induced by ZIKV vaccines enhance DENV infections.
a, Neutralizing activities of ZIKV vaccines. 6-8-week old female BALB/c mice (n = 5 biologically independent samples) were immunized with a single dose of AdC7-M/E or AdC7-prM/E (1.6 × 1011 vp) via i.m. route. PBS was given as a sham vaccine. Sera were collected from blood at 4 weeks postvaccination. Magnitudes of NAbs were measured as MN50 titers. Data are means ±standard errors of means (SEM). P values were analyzed with One-way ANOVA test (ns, not significant). The dashed line indicates the LOD. b-e, ADE activities of the immune sera (n = 3 biologically independent samples). Enhancement curves of K562 cells infected with DENV1 (b), DENV2 (c), DENV3 (d), and DENV4 (e) in the presence of serially diluted mouse immune sera as indicated. Infected cells were quantified by flow cytometry and were normalized to the peak infectivity of FLE mAb Z6. Data are means ± SEM. f, ADE activities of the human mAb Z6. Enhancement curves of K562 cells infected with DENV1, 2, 3 and 4 in the presence of serially diluted Z6 as indicated.
Extended Data Fig. 2 Comparison of FLE mAbs bound to flavivirus-E.
a, Z6 bound to ZIKV-sE. Z6 Fab is colored in green and ZIKV-sE is colored in grey. FL is highlighted in magenta. b, 2A10G6 bound to ZIKV-sE (PDB: 5JHL). 2A10G6-Fab is colored in cyan and ZIKV-sE is colored in grey. FL is highlighted in magenta. c, E53 bound to WNV-sE (PDB: 3I50). E53 Fab is colored in orange and ZIKV-sE is colored in grey. FL is highlighted in magenta. d, The Z6, 2A10G6 and E53 bound to E are overlapped, showing different approaching angles. e, The E residues participating in interactions of antibodies are listed. The numbers of contacts are highlighted for the main contributors in each sE.
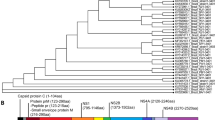
Extended Data Fig. 4 Phylogenetic trees flaviviruses, neutralizing titers in Ifnar−/− mice and ADE activities of the human mAb Z5.
a, Phylogenetic trees of 73 flaviviruses based on the amino acid sequences of the E. b-c, Groups of 5-week-old female Ifnar−/− mice (n = 7) were immunized with a single dose (1.6 × 1011 vp) of AdC7-M/E-WT, -M/E-MutB or -M/E-MutC via i.m. route. PBS was given as the sham vaccine. Sera were collected at week 4, 12 and 24. Serologic binding and neutralizing antibodies were detected. Data are means ± SEM. d, Groups of 5-week-old Female Ifnar−/− mice (n = 8-9) were immunized with a single dose (1.6 × 1011 vp) of AdC7-M/E-WT, -M/E-MutB or -M/E-MutC via i.m. route. PBS was given as the sham vaccine. Sera were collected at 4 weeks postimmunization. MN50 titers of NAbs were measured. Results were pooled from two independent experiments (related to Fig. 2b–e). Data are means ± SEM. P values were analyzed with two-tailed Student’s t test (ns, not significant; *, P < 0.05; P values are available in source data). The dashed line indicates the LOD. e, ADE activities of the ZIKV FLE mAb Z5. Enhancement curves of K562 cells infected with DENV1, 2, 3 and 4 in the presence of serially diluted Z5 as indicated. Infected cells were quantified by flow cytometry.
Extended Data Fig. 5 Antigen characterization of AdC7 vectored vaccines.
HEK293 cells were infected with AdC7 expressing ZIKV-M/E WT, MutB or MutC. Non-infected cells were used as the negative control. a, Flow cytometry detection of antigen expression and epitope display probed by FLE mAbs or neutralizing mAbs targeting epitopes on DI, DII or DIII. b, Capture ELISA quantifying the secretion of ZIKV antigen from AdC7 virus infected cells. Data are means of triplicates.
Extended Data Fig. 6 Protective efficacy of MutB/C-based vaccine against ZIKV infection at 3 DPI.
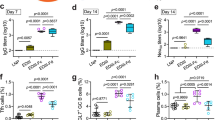
a-c, Groups of Ifnar1−/− mice (n = 4) (Female:male of 2:2 in both WT and MutB groups; 3:1 in MutC group; 4:0 in sham group) received a single immunization of 1.6 × 1011 vp of AdC7-M/E-WT, AdC7-M/E-MutB, AdC7-M/E-MutC via the i.m. route. PBS was given as the sham vaccine. Serum samples were collected for detection of ZIKV-E-specific IgG (a) and NAb titers (b). Thirty-days after vaccination, mice were challenged with 5 × 106 FFU of ZIKV-SMGC-1 via the i.p. route. At 3 DPI, tissues from the brain, spinal cord, testis, spleen, liver and eye were harvested for measurement of ZIKV loads (c). Data are means ± SEM. The dashed line indicates the LOD.
Extended Data Fig. 7 Evaluation of the sera from donor mice, related to Figs. 3f,g.
a,b, The pooled serum from donors were tested for ZIKV-E-specific IgG (a) and NAb titers (b). The dashed line indicates the LOD. c, ADE activities of the pooled donor sera in cell culture. Shown are the enhancement curves of K562 cells infected with DENV1, DENV2, DENV3, and DENV4, respectively, in the presence of serially diluted immunized mice sera as indicated. Infected cells were quantified by flow cytometry. Relative infectivity for each sample were normalized to the peak infectivity of mAb Z6.
Extended Data Fig. 8 FACS plots showing the gating strategy to isolate single ZIKV-E+ BGC.
The ZIKV-E-binding BGC (GL-7+ B220hi CD38lo IgD−CD93−CD138−) from the lymph nodes of a group of BALB/c mice primed with ZIKV vaccines was sorted by flow cytometry.
Extended Data Fig. 9 Amino acid sequences and alignment of FLE mAbs.
a, Amino acid sequence alignment of VH-region for mAb ZWT.1-3 and 6B6C-1. b, Amino acid sequence alignment of VL-region for mAb ZWT.1-3 and 6B6C-1. c, Amino acid sequence alignment of VH-region for mAb ZWT.4-6, 4G2 and 2A10G6. d, Amino acid sequence alignment of VL-region for mAb ZWT.4-5 and 4G2. e, Amino acid sequence alignment of VL-region for mAb ZWT.6 and 2A10G6.
Extended Data Fig. 10 Overview of E-MutC bound to Z3L1_scFv and model on virus particle.
a, Overall structure of complex of sE-MutC/Z3L1_scFv. The sE-MutC is shown as ribbon, Z3L1_scFvs are showed as surface. b, Left: the cryo-electron microscopy structure of the mature ZIKV particle (PDB: 5IZ7). Middle: ZIKV-E-MutC/Z3L1-scFv docked onto the the mature ZIKV particle (PDB: 5IZ7), Right: ZIKV-E-MutC docked onto the the mature ZIKV particle (PDB: 5IZ7). The Z3L1 footprint are highlighted in green.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1, 2, 4 and 5, Fig. 1, and PDB reports for 7BPK and 7BQ5.
Supplementary Table 3 (download XLSX )
Paired V gene usage.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
About this article
Cite this article
Dai, L., Xu, K., Li, J. et al. Protective Zika vaccines engineered to eliminate enhancement of dengue infection via immunodominance switch. Nat Immunol 22, 958–968 (2021). https://doi.org/10.1038/s41590-021-00966-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-021-00966-6
This article is cited by
-
Approach from the laboratory to the field: New strategies in the control of mosquito-borne infectious diseases
Science China Life Sciences (2025)
-
Bioprofiling and vaccine development for Zika virus
Science China Life Sciences (2025)
-
Convergence and divergence of B cell responses in two HIV-1 Env immunizations in Rhesus macaques
Communications Medicine (2025)
-
Rational design of flavivirus E protein vaccine optimizes immunogenicity and mitigates antibody dependent enhancement risk
Nature Communications (2025)
-
A single-dose circular RNA vaccine prevents Zika virus infection without enhancing dengue severity in mice
Nature Communications (2024)