Abstract
Chronic hepatitis B virus (HBV) infection is marked by dysfunctional HBV-specific CD8+ T cells, and restoring their effector activity is a major therapeutic goal. Here, we generated HBV-specific CD4+ T cell receptor transgenic mice to show that CD4+ effector T cells can prevent and reverse the CD8⁺ T cell dysfunction induced by hepatocellular priming. This rescue enhances antiviral CD8+ T cell function and suppresses viral replication. CD4+ T cell help occurs directly within the liver, independent of secondary lymphoid organs, and requires local antigen recognition. Kupffer cells, rather than dendritic cells, are the critical antigen-presenting platform. CD4+ T cells license Kupffer cells via CD40–CD40L interactions, triggering interleukin (IL)-12 and IL-27 production. IL-12 expands the CD4+ T cell pool, while IL-27 is essential for CD8+ T cell rescue. Exogenous IL-27 similarly restores HBV-specific CD8+ T cell function in mice and in T cells isolated from chronically infected patients. These findings identify IL-27 as a tractable immunotherapeutic target in chronic HBV infection.
Similar content being viewed by others
Main
HBV is a noncytopathic, hepatotropic DNA virus that can establish persistent infections, often progressing to cirrhosis and hepatocellular carcinoma1. Viral clearance relies primarily on the adaptive immune system, particularly CD8+ effector T (Teff) cell functions, as HBV elicits limited innate responses during early phases of infection1,2,3. However, in most neonatal or perinatal infections, and in some adult-onset cases, virus-specific CD8+ T cell responses are weak and dysfunctional, leading to viral persistence1,2,4. Restoring CD8+ Teff cell activity is, therefore, a central goal of curative therapy for chronic hepatitis B5.
Although naive CD8+ T cells are generally thought to be primed in secondary lymphoid organs (SLOs), studies have shown that HBV-specific CD8+ T cells can also be primed directly in the liver6,7,8. However, liver-primed CD8+ T cells often enter a dysfunctional state, proliferating but failing to acquire effector functions6. Multiple lines of evidence—from chimpanzee depletion studies9 to human cohort data10,11,12,13—suggest that CD4+ T cell help is essential for an effective antiviral CD8+ T cell response. Yet, the molecular and cellular mechanisms involved during chronic antigen exposure remain poorly defined.
Classical models portray CD4–CD8 cooperation as a synchronous, tri-cellular interaction in SLOs, where antigen-specific CD4+ and CD8⁺ T cells engage the same dendritic cell (DC) to achieve optimal responses14,15,16,17,18. More recent work suggests a staged model in which CD4+ T cell help is delivered after CD8+ T cell priming and in a separate SLO compartment19. Whether pre-activated (effector) CD4+ T cells can deliver restorative signals directly within nonlymphoid tissues, such as the chronically infected liver, remains unknown and therapeutically relevant.
Here, by combining existing mouse models of hepatitis B with newly generated HBV-specific CD4+ T cell antigen receptor (TCR) transgenic mice, we dissect how antigen-experienced CD4+ effector T cells reprogram Kupffer cells (KCs) to revive dysfunctional CD8+ T cells in a tolerogenic hepatic environment. Defining this liver-centric help circuit identifies key cytokine mediators and provides mechanistic rationale for immunotherapeutic strategies aimed at achieving a functional cure for chronic HBV.
Results
Generation and validation of HBV-specific CD4+ TCR transgenic mice
We began our investigation by generating an HBV-specific CD4+ TCR transgenic (Tg) mouse. Wild-type (WT) mice were immunized with the I-Ab-restricted immunodominant Env126–138 peptide from the preS2 region of the HBV genome20. Splenocytes were subsequently isolated from immunized mice and restimulated with the Env126–138 peptide to enrich for antigen-specific T cells (Extended Data Fig. 1a). Single-cell sorting of these T cells (Extended Data Fig. 1b) allowed us to identify and clone the TCR alpha and beta chains. After confirming the TCR’s specificity and reactivity to the Env126–138 peptide (Extended Data Fig. 1c), the cloned TCR alpha and beta chains were inserted into TCR expression vectors21 (Extended Data Fig. 1d) and injected into fertilized C57BL/6 eggs, generating two mouse lines expressing the Env126–138 Tg TCR chains. Crossbreeding of these lines produced HBV-specific CD4+ TCR Tg mice (Env126 mice).
Flow cytometry confirmed the presence of HBV-specific CD4+ T cells in primary and secondary lymphoid organs of Env126 mice (Fig. 1a). Tetramer staining verified Tg TCR expression on CD4+ T cells, enabling ex vivo detection of HBsAg-specific Env126 CD4+ T cells (Fig. 1a). These cells made up between 8% and 30% of total CD4+ T cells and exhibited a naive phenotype (CD44loCD62LhiCD69−CD25−) comparable to WT CD4+ T cells (Fig. 1b and Extended Data Fig. 1e).
a, Representative flow cytometry plots showing Env126–138 tetramer+ CD4+ T cells in WT mice and Env126 TCR Tg mice. Cells were stained with Env126–138 peptide-loaded tetramer (I-Ab Env126–138) and human peptide-loaded tetramer as a control (I-Ab CLIP). b, Frequency and mean fluorescence intensity (MFI) of indicated markers on splenic CD4+ T cells in Env126 Tg and WT mice. c, Design: splenocytes of Env126 or WT mice were in vitro stimulated with cognate peptide, PMA–ionomycin or anti-CD3/anti-CD28 Dynabeads and analyzed by flow cytometry for cytokine production and proliferation with CellTrace Violet (CTV). Non-stimulated splenocytes were included as a control (Ctrl). d, Representative flow cytometry plots showing the frequency of TNF-producing CD4+ T cells in WT or Env126 mice after 4 h of in vitro stimulation. e, Representative flow cytometry plots showing the frequency of CD44+ and Ki-67+ CD4+ T cells in WT or Env126 mice after 4 h of in vitro stimulation with Env126–138 peptide. f, Representative plots of CTV-labeled WT or Env126 CD4+ T cells after 48 h of in vitro stimulation. Each peak represents a cell division and is indicated with a different shade of color. Numbers indicate the frequency of cells within the peak of events that did not undergo cell division. g, Proliferation index (total number of divisions divided by the number of cells that went into division) of WT and Env126 CD4+ T cells after 48 h of in vitro stimulation. h, Representative histograms of surface expression of indicated markers in WT and Env126 CD4+ T cells after 48 h of in vitro stimulation. i, Representative flow cytometry plot showing the frequency of Env126–138 tetramer+ CD4+ T cells in the spleen of Env126 or Env126 Rag1−/− mice. CD4+ T cells were identified as live, CD45+, CD8−CD4+; Env126 CD4+ T cells were identified as live, CD45+, CD8−CD4+Env-Tet+. a–h, n = 3 (WT), 5 (Env126). Data are representative of at least three independent experiments. i, n = 5 (Env126), 5 (Env126 Rag1−/−). Results in g are expressed as the mean ± s.e.m. P values were calculated with two-way analysis of variance (ANOVA) with Bonferroni’s post hoc test. ****P < 0.0001.
Next, to assess HBsAg-specific reactivity of Env126 TCR in CD4+ T cells, splenocytes from Env126 and WT mice were stimulated in vitro with either the Env126–138 peptide or polyclonal stimuli (phorbol 12-myristate 13-acetate (PMA)–ionomycin or anti-CD3/anti-CD28 beads) for 4 h and 48 h (Fig. 1c). Only Env126 CD4+ T cells were activated by peptide stimulation, as evidenced by tumor necrosis factor (TNF) production and CD44 and Ki-67 upregulation (Fig. 1d,e and Extended Data Fig. 1f). After 48 h, Env126 CD4+ T cells proliferated and upregulated CD44, CD69 and CD25 in response to Env126–138 peptide, while WT cells only responded to polyclonal stimuli (Fig. 1f–h and Extended Data Fig. 1g).
To prevent the development of polyclonal endogenous T cells, Env126 mice were bred onto a Rag1−/− background. Resulting CD4+ T cells stained uniformly positive for the Env126 tetramer (Fig. 1i) and displayed similar phenotype and function to their WT background counterparts (Supplementary Fig. 1). These mice were used for all subsequent experiments, providing an ideal system for in vivo functional studies.
CD4+ Teff cells reverse CD8+ T cell dysfunction
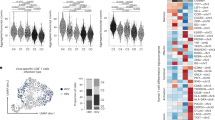
To assess the ability of CD4+ Teff cells to support intrahepatic CD8+ T cell responses, we developed an in vivo activation strategy to generate Env126 CD4+ Teff cells for adoptive transfer experiments (Extended Data Fig. 2a). Naive Env126 CD4+ T cells were transferred into Rag1−/− mice that were subsequently infected with a recombinant vesicular stomatitis virus expressing the HBV envelope glycoprotein (rVSVEnv)22 (Extended Data Fig. 2a). After 7 days, Env126 CD4+ T cells differentiated into CD62LloCD44hiT-bet+CXCR3+ effector cells, producing interferon gamma (IFNγ), TNF and IL-2 upon peptide restimulation (Extended Data Fig. 2b–f). Single-cell RNA sequencing (scRNA-seq) confirmed the Env126 Teff phenotype and identified two clusters (Extended Data Fig. 3a–c) expressing transcripts encoded by type 1 helper T (TH1) cell-associated genes, including Cxcr3, Tbx21, Ifng, Tnf, Il2, Il12rb1 and Il12rb2 (Extended Data Fig. 3a–f and Supplementary Table 1) with differences limited to proliferative state (Extended Data Fig. 3b,g,h).
To evaluate whether Env126 CD4+ Teff cells could prevent CD8+ T cell dysfunction induced by hepatocellular priming6,7,23,24, we co-transferred Env126 CD4+ Teff cells and naive Env28 CD8+ naive T (Env28 TN) cells23 into HBV replication-competent Tg mice (hereafter HBV Tg mice)25 (Fig. 2a). While Env28 TN cells alone, as expected6,7,23, failed to induce serum alanine aminotransferase (sALT) elevation (Fig. 2b), co-transfer with Env126 CD4+ Teff cells caused liver damage (Fig. 2b), increased intrahepatic leukocyte (IHL) and Env28 CD8+ T cell numbers (Fig. 2c,d), enhanced Env28 CD8+ Teff cell functions, including IFNγ, TNF and granzyme B (Grzm-B) production (Fig. 2e,f), increased proliferation (Ki-67 expression), reduced inhibitory marker expression (PD-1, Lag-3 and CTLA-4) and upregulation of activation markers (CD44, CD25, CD107a, Sca-1; Fig. 2g–i and Extended Data Fig. 4a,b). Delayed transfer (day 7) of Env126 CD4+ Teff cells still increased CD8+ T cell number and function, demonstrating that CD4+ T cells can also reverse established CD8+ T cell dysfunction (Extended Data Fig. 4c–g). Despite these effects in the liver, Env126 CD4+ Teff cells did not affect the number of Env28 CD8+ T cells in SLOs (Extended Data Fig. 4h), suggesting that CD4+ T cell help to CD8+ T cells was localized in the liver. Histological and confocal analyses revealed that when Env28 CD8+ T cells were co-transferred with Env126 CD4+ Teff cells into HBV Tg mice, the T cells formed clusters scattered throughout the liver lobules (Fig. 2j). This distribution pattern closely resembled that seen during acute, self-limited HBV infection26. In contrast, Env28 CD8+ T cells transferred alone tended to accumulate around periportal areas (Fig. 2j), a pattern characteristic of chronic HBV infection27. Co-transfer reduced HBV replication, as demonstrated by Southern blot analysis (Fig. 2k), decreased cytoplasmic HBcAg staining (Fig. 2l) and reduced viremia (Fig. 2m).
a, Design: Env126 Teff cells were transferred at a 1:1 ratio (106:106) with naive Env28 T cells in HBV Tg mice (Co-T). Single transfer of Env28 and untreated mice (phosphate-buffered saline (PBS)) were used as controls. b, sALT in indicated groups of mice and time points. c, IHL numbers at day 5 after cell transfer. d, Frequency and numbers of Env28 T cells. e,f, Frequency (e) and numbers (f) of IFNγ-, TNF- and Grzm-B-producing Env28 T cells. g, Representative flow cytometry plots showing the frequency of control and Env28 T cells expressing Ki-67 at medium (med) and high (high) levels. h, Frequency of Ki-67hi Env28 T cells. i, Radar plot showing normalized MFI values of indicated markers on Env28 T cells. Asterisks refer to the comparison between Env28 and Co-T conditions. j, Immunohistochemical representative micrographs of liver sections: hematoxylin and eosin (H&E), cytokeratin-7 (bile ducts) and Ki-67 stainings. Scale bars, 100 μm. Bottom, confocal immunofluorescence micrographs of liver sections showing: DsRed+/+ Env28 T cells (red), CD38+ sinusoids (white), F4/80+ KCs. Scale bars, 200 μm. k, HBV DNA quantification by Southern blot. Bands indicate integrated transgene (TG), relaxed circular (RC), double-stranded linear (DS) and single-stranded (SS) HBV DNA. l, Representative HBcAg immunohistochemistry (IHC) micrographs of liver sections. Scale bars, 100 μm. m, Serum HBV DNA quantification. n, Env126 Teff cell numbers. o, MFIs of CD44, CD62L, T-bet and Ki-67 on hepatic Env126 Teff cells. p, Representative flow cytometry plots showing the frequency of Env126 Teff cells producing IFNγ and TNF, expressing CXCR3 and T-bet. Control CD4+ T cells are shown in black. n = 3 (PBS), 4 (Env28) and 5 (Co-T). Data are representative of five independents and are expressed as the mean ± s.e.m. Data in b and m were analyzed using two-way ANOVA with Bonferroni’s post hoc test. Data in all other graphs were analyzed using a two-tailed t-test or one-way ANOVA with Bonferroni’s post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. dLNs, draining lymph nodes.
Further analysis revealed that, as expected, Env126 CD4+ Teff cells preferentially accumulated in the liver, rather than in SLOs, in HBV Tg mice (Fig. 2n) and maintained their activated phenotype (Fig. 2o), their TH1 profile, and their ability to produce antiviral cytokines (for example, IFNγ and TNF; Fig. 2p). Similar results were obtained when lower, more physiological numbers of Env126 CD4+ Teff cells and HBV-specific naive CD8+ T cells were co-transferred in HBV Tg mice (Extended Data Fig. 4i–n).
Importantly, when Env126 CD4+ Teff cells were transferred alone into HBV Tg mice, no T cell-mediated liver immunopathology was observed (Extended Data Fig. 5a–e) and HBV replication was not affected (Extended Data Fig. 5f,g), indicating that the Env28 CD8+ T cells were the primary effectors responsible for the observed antiviral effects in the co-transferred HBV Tg mice.
In conclusion, our findings demonstrate that CD4+ Teff cells can prevent or reverse hepatocellular priming-induced CD8+ T cell dysfunction, prompting investigation into the antigen-presenting cells (APCs) and cytokine signals involved.
CD4+ T cell help requires antigen recognition but not epitope linkage
To determine whether CD4+ Teff cell help extends to CD8+ T cells with different antigen specificities from those of the CD4+ T cells, we co-transferred Env126 CD4+ Teff cells with Cor93 CD8+ naive T (Cor93 TN) cells, whose TCR recognizes an H-2Kb-restricted epitope within the HBV core protein (HBcAg23,24) into HBV Tg mice (Fig. 3a). Co-transfer induced acute liver injury (Fig. 3b) and increased IHL numbers (Fig. 3c). Cor93 CD8+ T cells, like Env28 CD8+ T cells, expanded and were phenotypically reinvigorated in the liver when co-transferred with Env126 CD4+ Teff cells (Fig. 3d and Extended Data Fig. 6a–d). HBV replication was also suppressed in these mice (Fig. 3e and Extended Data Fig. 6e), and the T cells formed clusters scattered throughout the liver lobules (Fig. 3f and Extended Data Fig. 6f), demonstrating that Env126 CD4+ Teff cells can provide heterologous help, enhancing the antiviral responses of CD8+ T cells regardless of their viral TCR specificity.
a, Experimental design. b, sALT in indicated groups of mice and time points. c, IHL numbers at day 5 after cell transfer. d, Numbers of hepatic Cor93 and IFNγ+ Cor93 T cells in indicated groups of mice. e, HBV DNA quantification by Southern blot. Bands indicate integrated TG, RC, DS and SS HBV DNA. f, Immunohistochemical representative micrographs of liver sections: H&E, cytokeratin-7 (bile ducts) and Ki-67 stainings. Scale bars, 100 μm. g, Experimental design. h, sALT in indicated groups of mice and time points. i, IHL numbers. j, Numbers of hepatic Cor93 and Env28 T cells at day 5 after transfer. k, Numbers of IFNγ-producing Cor93 and Env28 T cells. l, Histograms and representative plots of Ki-67 MFI values on control, Env28 and Cor93 CD8+ T cells. m, Experimental design. n, sALT in indicated groups of mice and time points. o, IHL numbers. p, Numbers of hepatic Cor93 and IFNγ+ Cor93 T cells. q, MFIs of indicated markers on control and Cor93 CD8+ T cells. r, Immunohistochemical representative micrographs of liver sections: H&E, cytokeratin-7 (bile ducts) and Ki-67. Scale bars, 100 μm. s, Representative HBcAg IHC micrographs of liver sections. Scale bars, 100 μm. t, Numbers of hepatic Env126 Teff cells at day 5 after transfer. a–f, n = 3 (PBS), 4 (Cor93) and 5 (Co-T). Data are representative of at least four independent experiments. g–l, n = 3 (PBS), 4 (CD8) and 4 (Co-T). Data are representative of at least three independent experiments. m–t, n = 3 (PBS), 4 (Ctrl Cor93), 3 (MUP Cor93), 4 (Ctrl Co-T), 4 (MUP Co-T) and 4 (MHC-II−/− Co-T). Data are representative of at least two independent experiments. Data are expressed as the mean ± s.e.m. Data in b, h and n were analyzed using two-way ANOVA with Bonferroni post hoc test. Data in all other graphs were analyzed using a two-tailed t-test or one-way ANOVA with Bonferroni’s post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001.
Previous studies have suggested that optimal CD4+ T cell help requires the recognition of linked epitopes17,28. To test whether epitope linkage was required in our model, we co-transferred Env126 CD4+ Teff cells with Env28 TN and Cor93 TN into 107-5 mice (ref. 23; expressing the HBV envelope proteins) crossed with MUP-core (expressing the HBV core protein)29 F1 mice (Fig. 3g). In contrast to HBV Tg mice, which produce virions containing both envelope and core proteins within the same particle, the 107-5 mice crossed with MUP-core F1 mice produce these proteins independently. Despite lack of physical linkage between viral epitopes, co-transferred mice exhibited elevated sALT levels (Fig. 3h) and increased IHL numbers (Fig. 3i). Both CD8+ T cell populations showed enhanced proliferation and effector functions in the presence of Env126 CD4+ Teff cells (Fig. 3j–l). These findings suggest that CD4+ T cell help might not be reliant on epitope linkage, aligning with evidence that this requirement is dispensable, especially at high antigen doses17.
Next, we investigated whether CD4+ T cell help required direct antigen recognition and TCR engagement by CD4+ T cells. To test this, we performed adoptive transfer experiments using either HBV Tg MHC-II−/− mice or MUP-core mice (expressing the HBV core protein; Fig. 3m). This setup allowed us to assess Env126 Teff cell help under conditions in which either their cognate antigen was present but not presented on MHC class II molecules (HBV Tg MHC-II−/− mice) or where MHC-II was broadly expressed on APCs but the cognate antigen was absent (MUP-core mice), preventing TCR engagement. Env126 CD4+ Teff cell transfer led to elevated sALT levels only in HBV Tg control mice, but not in HBV Tg MHC-II−/− or MUP-core mice, and there was no increase in IHL numbers in these latter groups (Fig. 3n,o). Additionally, HBV-specific CD8+ T cells accumulated and exhibited enhanced effector functions only in HBV Tg control mice co-transferred with Env126 CD4+ Teff cells (Fig. 3p,q). Histological analyses showed that, without antigen recognition by CD4+ T cells, T cells failed to form intra-parenchymal clusters, instead clustering around portal tracts (Fig. 3r). This was associated with a failure to suppress HBV replication, as shown by persistent cytoplasmic HBcAg staining (Fig. 3s). Furthermore, the number of Env126 CD4+ Teff cells was significantly reduced in MHC-II−/− mice and MUP-core mice, confirming that their proliferation and helper functions depend on direct antigen recognition (Fig. 3t).
These experiments elucidate that intrahepatic CD4+ Teff cell help to HBV-specific CD8+ T cells in HBV Tg mice relies on antigen recognition by CD4+ T cells but does not require cognate epitope linkage. This help extends to CD8+ T cells with different antigen specificities, underscoring the critical role of direct TCR–MHC class II interactions in mediating CD4+ T cell support for antiviral CD8+ T cells. These findings raise important questions about which APC presents antigens to CD4+ T cells and whether this occurs in the liver or in SLOs.
CD4+ T cells help CD8+ T cells through extra-lymphoid licensing of APCs
Although the data described thus far establish a critical role of CD4+ T cell help in shaping CD8+ T cell function, it remains unclear whether this support is delivered within SLOs, or can occur within the liver itself. To address this, we used a model in which HBV-specific CD8+ T cell priming is restricted to the liver6 (Fig. 4a and Extended Data Fig. 7a). We co-transferred Env126 CD4+ Teff cells with HBV-specific CD8+ TN cells into either control HBV Tg mice or HBV Tg mice that had undergone splenectomy and anti-CD62L treatment to block recirculation of CD8+ TN cells through SLOs6 (Fig. 4a and Extended Data Fig. 7a). In both conditions, co-transfer led to elevated sALT levels (Fig. 4b), increased IHL accumulation (Fig. 4c), expansion and activation of HBV-specific CD8+ T cells (Fig. 4d–h) and formation of scattered intrahepatic T cell clusters (Fig. 4i and Extended Data Fig. 7b) alongside a reduction in cytoplasmic HBcAg staining (Fig. 4j), suggesting that CD4+ Teff cells can provide local help toward CD8+ T cells within the liver. Consistent with their effector phenotype, the number and functionality of Env126 CD4+ Teff cells were unaffected by impaired SLO recirculation (Fig. 4k–m). Similar results were observed when FTY720 treatment was used to prevent lymphocyte egress from SLOs (Extended Data Fig. 7c–h).
a, Experimental design. b, sALT in indicated groups of mice and time points. c, IHL numbers at day 5 after cell transfer. d, Numbers of Cor93 T cells in the liver. e, Numbers of IFNγ+ and Grzm-B+ Cor93 T cells. f, Frequency of IFNγ+ Cor93 T cells. g,h, MFIs of indicated markers on Cor93 T cells. i, Liver immunohistochemical representative micrographs at day 5 after cell transfer: H&E, cytokeratin-7, Ki-67 stainings. Scale bars, 100 μm. j, Representative HBcAg IHC liver micrographs at day 5 after cell transfer. Scale bars, 100 μm. k,l, Numbers of hepatic Env126 and IFNγ+ Env126 CD4+ Teff cells. m, Frequency of IFNγ+ Env126 Teff cells. n, Heat map of normalized MFI (nMFI) of indicated markers on KCs and DCs at day 5 after cell transfer. MFI values were normalized separately (lowest value set to 0 and highest to 100). o, Experimental setup. Livers were analyzed 24 h after cell transfer. p, Rendered confocal immunofluorescence representative liver micrographs from co-transferred mice: DsRed+/+Env28 T cells (red), EGFP+/+Env126 Teff cells (green), CD38+ sinusoids (white), F4/80+ KCs (blue). Scale bars, 100 μm and 10 μm. q, Time-coded shapes of DsRed+/+Env28 T (red) and EGFP+/+Env126 Teff (green) cell tracks of Supplementary Video 1. r, Single-cell speeds and arrest coefficients of Env28 T cells non-interacting or engaged in three-cell clusters (interacting; cutoff, <2.5 µm min−1). s, Representative multiplex immunofluorescence liver image from a chimpanzee acutely infected with HBV at week 20 after infection: CD4+ T cells (green), CD8+ T cells (red), macrophages (KP1, white), nuclei (DAPI, blue). Scale bar, 10 µm. Individual channels are displayed in Supplementary Fig. 3. a–n, n = 3 (PBS), 4 (Cor93), 4 (Ctr Co-T), 4 (-SLOs Co-T). Data are representative of four independent experiments. o–r, n = 5. r, n = 82 tracks (not interacting), 28 tracks (interacting). Data are representative of at least two independent experiments. Data are expressed as the mean ± s.e.m. Data in b were analyzed using two-way ANOVA with Bonferroni’s post hoc test. Data in all other graphs were analyzed using two-tailed t-test or one-way ANOVA with Bonferroni’s post hoc test. *P < 0.05, **P < 0.01, ****P < 0.0001.
We next sought to identify which hepatic APC serves as the cellular platform for CD4+ Teff cell help delivery to CD8+ T cells. Because KCs and hepatic DCs are the main APCs in the liver, we examined their activation following adoptive transfer of Env126 CD4+ Teff cells and HBV-specific CD8+ T cells. Our results demonstrated that, under the co-transfer condition, KCs exhibited markedly higher upregulation of MHC machineries and key co-stimulatory molecules (including CD40, CD80, CD86 and CD70) compared to hepatic DCs. This enhanced activation, which is critical for antigen presentation and T cell priming16, was particularly evident when compared to the single CD8+ T cell transfer condition (Fig. 4n and Extended Data Fig. 7i–k).
To investigate whether KCs serve as the platform for intrahepatic CD4+–CD8+ T cell cooperation, we performed confocal and multiphoton intravital microscopy in HBV Tg mice 24 h after transfer (Fig. 4o). Both imaging approaches revealed that Env126 CD4+ Teff cells and HBV-specific CD8+ T cells frequently interact with KCs within the liver, even with the same KC (Fig. 4p), forming triadic structures. Notably, HBV-specific CD8+ T cells in proximity to KCs and Env126 CD4+ Teff cells predominantly exhibited reduced motility and higher arrest coefficients, indicative of antigen sensing and active engagement, while non-interacting ones remained highly motile (Fig. 4q–r, Supplementary Video 1 and Supplementary Fig. 2). These interactions persisted for over 20 min, aligning with the estimated 30-min duration required for stable immunological synapse formation between naive T cells and APCs30. While HBV-specific CD8+ T cells formed long, stable interactions with KCs, Env126 CD4+ Teff cells established similarly stable interactions but migrated between KCs in a stop-and-go manner, likely facilitating the licensing process (Supplementary Video 1).
To extend these findings, we analyzed liver biopsy samples from a previously published chimpanzee model of acute HBV infection9. In these samples, we qualitatively observed consistent CD4+–CD8+–KC triadic interactions across multiple time points during the acute phase in a chimpanzee that naturally resolved the infection (Fig. 4s and Supplementary Fig. 3). While these data are limited in number and scope and did not allow for comprehensive quantification of APC–T cell interactions, they offer a unique window into hepatic immune responses in higher primates. Within these constraints, our observations are consistent with the possibility that intrahepatic CD4+–CD8+ T cell cooperation involving KC may also occur during natural HBV infection.
Thus, our findings demonstrate that CD4+ Teff cell-mediated help to CD8+ T cells can occur within the liver, independently of SLO involvement. While DCs are traditionally considered primary mediators of CD4+–CD8+ T cell cooperation, they remained phenotypically poorly affected by the presence of Env126 CD4+ Teff cells in our setting. Instead, KCs emerged as the principal APC facilitating intrahepatic T cell cooperation, unveiling an extra-lymphoid mechanism of immune modulation that may be critical in chronically infected tissues.
CD4+ T cell help to CD8+ T cells requires KCs
To determine which APCs mediate CD4+ Teff cell help to HBV-specific CD8+ T cells, we performed selective depletion experiments. First, we depleted DCs using diphtheria toxin in HBV Tg mice reconstituted with CD11c-iDTR-GFP bone marrow (Fig. 5a,b and Extended Data Fig. 8a–e). Despite efficient DC depletion, Env126 CD4+ Teff cells continued to provide help to HBV-specific CD8+ T cells, as indicated by sustained liver damage (elevated sALT), increased IHL accumulation and improved effector function and phenotype of HBV-specific CD8+ T cells (Fig. 5c–e and Extended Data Fig. 8f,g). Inflammatory infiltrates persisted throughout the liver parenchyma, and HBV replication remained suppressed (Fig. 5f and Extended Data Fig. 8h). Moreover, DC depletion did not affect the number, phenotype or functionality of Env126 CD4+ Teff cells (Fig. 5g and Extended Data Fig. 8i–k).
a, Experimental design. b, Hepatic DC numbers at day 5 after cell transfer. c, sALT in indicated groups of mice and time points. d, IHL numbers. e, Hepatic Cor93 T cell and IFNγ+ Cor93 CD8+ T cell numbers. f, Immunohistochemical representative liver micrographs at day 5 after cell transfer: H&E, cytokeratin-7, cleaved caspase-3 stainings. Scale bars, 100 μm. g, Hepatic Env126 Teff cell numbers at day 5 after transfer. h, Experimental setup. i, KC numbers at day 5 after cell transfer. j, Representative flow cytometry plots showing frequency of F4/80+TIM4+YFP+ KCs at day 5 after cell transfer. k, DC numbers at day 5 after cell transfer. l, sALT in indicated groups of mice and time points. m, IHL numbers. n,o, Env28 T cells and IFNγ+ Env28 T cell numbers. p, Representative flow cytometry plots showing the frequency of IFNγ+ Env28 T cells. q, Immunohistochemical representative liver micrographs: H&E, cytokeratin-7, Ki-67 stainings. Scale bars, 100 μm. r, Representative HBcAg IHC liver micrographs at day 5 after cell transfer. Scale bars, 100 μm. s, Serum HBV DNA quantification of indicated groups of mice and time points. t, Hepatic Env126 Teff cell numbers at day 5 after cell transfer. a–g, n = 3 (PBS), 3 (Cor93), 4 (+DCs Co-T), 5 (-DCs Co-T). Data are representative of at least two independent experiments. h–t, n = 3 (PBS), 3 (Cor93), 5 (+KCs Co-T), 6 (-KCs Co-T). Data are representative of at least three independent experiments. Data are expressed as the mean ± s.e.m. Data in c and l were analyzed using two-way ANOVA with Bonferroni post hoc test. Data in all other graphs were analyzed using a two-tailed t-test or one-way ANOVA with Bonferroni’s post hoc test. **P < 0.01, ***P < 0.001, ****P < 0.0001. BMT, bone marrow transplant. DT, diphtheria toxin.
We next depleted KCs using diphtheria toxin in HBV Tg mice crossed with CLEC4f-iDTR-YFP mice31 (Fig. 5h). In these mice, diphtheria toxin specifically depletes KCs while sparing other resident macrophages and DCs (Fig. 5i–k and Extended Data Fig. 9a–e). In mice co-transferred with Env126 CD4+ Teff cells and HBV-specific CD8+ TN cells, KC depletion prevented liver damage (Fig. 5l), led to decreased IHL accumulation (Fig. 5m) and severely impaired HBV-specific CD8+ T cell expansion, differentiation and cytokine production (Fig. 5n–p). In the absence of KCs, CD8+ T cells accumulated around portal tracts and were unable to induce T cell-mediated liver immunopathology or suppress HBV replication (Fig. 5q–s and Extended Data Fig. 9f). KC depletion also significantly reduced the accumulation and expansion of Env126 CD4+ Teff cells in the liver (Fig. 5t). Similar results were obtained using clodronate liposomes for KC depletion, confirming the specificity of these findings (Extended Data Fig. 9g–l).
In conclusion, these data demonstrate that KCs, but not DCs, can serve as a cellular platform through which CD4+ Teff cells rescue dysfunctional CD8+ T cells in a tolerogenic hepatic environment.
CD4+ T cells license KCs via CD40L to trigger IL-27 production
Given that CD40 signaling on APCs is a well-established mediator of CD4+ T cell help to CD8+ T cells14,15, we investigated its relevance by blocking CD40L in HBV Tg mice co-transferred with Env126 CD4+ Teff cells and HBV-specific CD8+ TN cells (Fig. 6a). CD40L blockade prevented sALT elevation and IHL accumulation (Fig. 6b,c), as Env126 CD4+ Teff cells failed to promote CD8+ T cell proliferation and effector functions (Fig. 6d). It also impaired Env126 CD4+ Teff cell expansion (Fig. 6e) and abolished upregulation of MHC and co-stimulatory molecules in KCs (Supplementary Fig. 4a), underscoring the importance of the CD40L–CD40 axis in the CD4+ T cell-mediated licensing of KCs, which is essential for reversing CD8+ T cell dysfunction. Similar results were obtained with the administration of an Fc-mutated version of anti-CD40L (MR-1-CP032), which prevents potential Fc-mediated depletion of CD40L-expressing cells (Supplementary Fig. 4b–h).
a, Experimental design. b, sALT in indicated groups of mice and time points. c, IHL numbers. d, Env28 T cell and IFNγ+ Env28 T cell numbers. e, Hepatic Env126 CD4+ Teff cell frequency. f, Relative quantification of IL-12 in total liver tissue. g, Experimental design. h, sALT in indicated groups of mice and time points. i, IHL numbers. j, Hepatic Cor93 T cell numbers at day 5 after transfer. k, IFNγ+ and Grzm-B+ hepatic Cor93 T cells. l, Immunohistochemical representative liver micrographs. Scale bars, 100 μm. m, Representative HBcAg IHC liver micrographs. Scale bars, 100 μm. n, Hepatic Env126 Teff cell frequency. o, Sca-1 MFI on Cor93 T cells. p, Relative quantification of IL-27 in total liver at day 5 after cell transfer. q, Experimental design. r, sALT in indicated groups of mice and time points. s, IHL numbers. t, Hepatic Cor93 T cell and IFNγ+ Cor93 T cell numbers. u, Representative flow cytometry plots showing the frequency of IFNγ+ Cor93 T cells. v, Immunohistochemical representative liver micrographs. Scale bars, 100 μm. w, Representative HBcAg IHC liver micrographs. Scale bars, 100 μm. x, Hepatic Env126 Teff cell frequency. y, Relative quantification of IL-27 in total liver tissue at day 5 after cell transfer (Fig. 5h–t). a–f, n = 3 (PBS), 3 (Env28), 4 (Isotype Co-T) and 3 (anti-CD40L (CD40L) Co-T); representative of three independent experiments. g–p, n = 3 (PBS), 3 (Cor93), 4 (Isotype Co-T), 4 (anti-IL-12 (aIL-12) Co-T), 4 (aCD40L Co-T); representative of three independent experiments. q–x, n = 3 (PBS), 3 (Cor93), 4 (Isotype Co-T), 4 (aIL-27 Co-T); representative of two independent experiments. y, n = 3 (PBS), 3 (Cor93), 5 (+KCs Co-T), 6 (−KCs Co-T). Data are expressed as the mean ± s.e.m. Data in b, h and r were analyzed using two-way ANOVA with Bonferroni’s post hoc test. Data in all other graphs were analyzed using a two-tailed t-test or one-way ANOVA with Bonferroni post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. NS, not significant.
We next investigated downstream mediators. CD40–CD40L interactions are known to induce IL-12 production by myeloid cells, a cytokine crucial for sustaining TH1 and CD8+ T cell effector responses32. Indeed, transfer of Env126 CD4+ Teff cells to HBV Tg mice markedly increased hepatic IL-12 levels, which was fully prevented by CD40L blockade (Fig. 6f). To explore the potential involvement of IL-12, we co-transferred Env126 CD4+ Teff cells and HBV-specific CD8+ TN cells into HBV Tg mice treated with either anti-CD40L or anti-IL-12 blocking antibodies (Fig. 6g). In mice treated with anti-IL-12, liver damage (Fig. 6h), IHL (Fig. 6i) and numbers and effector functions of HBV-specific CD8+ T cells (Fig. 6j,k) were significantly reduced, although somewhat less pronounced compared to CD40L blockade. While CD40L blockade fully abolished T cell-mediated liver immunopathology, IL-12 blockade primarily reduced the size and extent of intra-parenchymal inflammatory foci without fully preventing their formation (Fig. 6l). Similarly, the reduction in HBV replication was less pronounced with IL-12 blockade than with CD40L blockade (Fig. 6m). IL-12 blockade also diminished Env126 CD4+ T cell expansion (Fig. 6n and Supplementary Fig. 4i,j), consistent with the hypothesis that IL-12 supports Env126 CD4+ T cell proliferation and expansion.
Further analysis revealed that Sca-1, a key surface marker associated with enhanced CD8+ Teff cell functions33, was subtantially downregulated following CD40L blockade (Fig. 6o and Supplementary Fig. 4k). Since IL-27 is known to induce Sca-1 expression and shape CD8+ T cell activation and differentiation34,35, we examined its regulation. Interestingly, CD40L blockade not only reduced Sca-1 expression but also prevented the upregulation of IL-27 in the liver following the adoptive transfer of Env126 CD4+ Teff cells (Fig. 6p), highlighting the role of the CD40L–CD40 axis in driving effective T cell cooperation in the liver, with IL-27 potentially serving as a key mediator.
To further investigate the role of IL-27, we co-transferred Env126 CD4+ Teff cells and HBV-specific CD8+ TN cells into HBV Tg mice treated with a blocking anti-IL-27 antibody (Fig. 6q). IL-27 blockade completely prevented sALT elevation (Fig. 6r), IHL accumulation (Fig. 6s), HBV-specific CD8+ T cell expansion and effector differentiation (Fig. 6t,u), effectively abolishing liver immunopathology and antiviral activity (Fig. 6v,w). Notably, unlike CD40L or IL-12 blockade, IL-27 blockade did not affect Env126 CD4+ Teff cell expansion or effector functions (Fig. 6x), suggesting that IL-27 acts directly on CD8+ T cells.
To determine whether KCs are the intrahepatic source of IL-27, we measured IL-27 levels in the livers of HBV Tg CLEC4f-iDTR-YFP mice, which were co-transferred with T cells and depleted of KCs using diphtheria toxin (Fig. 5h–t). In the absence of KCs, the increase in IL-27 levels was completely abolished, confirming that KCs are required for intrahepatic IL-27 production (Fig. 6y).
In summary, CD4+ T cells license KCs via CD40L–CD40 interactions, enabling KCs to produce IL-12 and IL-27. While IL-12 primarily supports CD4+ Teff cell expansion, IL-27 is crucial for reversing dysfunctional HBV-specific CD8+ T cells, revealing a mechanism by which CD4+ T cells can reprogram hepatic APCs to restore CD8+ T cell function within a tolerogenic tissue environment.
Therapeutic potential of IL-27 for CD8+ T cell restoration
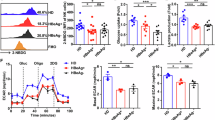
To assess the therapeutic relevance of our findings, we administered recombinant IL-27 (rIL-27) to HBV Tg mice that had been transferred with HBV-specific naive CD8+ T cells (Fig. 7a). IL-27 treatment induced sALT elevation, increased IHL accumulation (Fig. 7b,c) and promoted robust expansion and differentiation of Cor93 T cells into IFNγ-producing cytotoxic effectors (Fig. 7d–f and Extended Data Fig. 10a,b). These activated T cells formed intra-parenchymal clusters throughout the liver, effectively suppressing HBV replication (Fig. 7g,h). Furthermore, in vitro experiments confirmed that T cell reinvigoration by IL-27 required TCR signaling and that IL-27 alone was ineffective without antigenic stimulation (Extended Data Fig. 10c,d). Notably, systemic administration of IL-27 did not induce observable toxic effects, further supporting its favorable safety profile (Extended Data Fig. 10e–m).
a, Experimental design. b, sALT in indicated groups of mice and time points. c, IHL numbers. d, Frequency and numbers of hepatic Cor93 T cells. e, IFNγ+ and Grzm-B+ Cor93 T cell numbers. f, Representative flow cytometry plots showing the frequency of Sca-1+, CD107a+ (LAMP1), IFNγ+ Cor93 T cells. Control CD8+ T cells are shown in black. g, Immunohistochemical representative liver micrographs of indicated groups of mice: H&E, cytokeratin-7, cleaved caspase-3, Ki-67 stainings. Scale bars, 100 μm. h, Representative HBcAg IHC liver micrographs. Scale bars, 100 μm. i, Experimental setup: PBMCs were isolated from 12 individuals with CHB and stimulated with HBV peptide pools or with hCMV-UL55 peptides as a control. Cells were expanded in vitro for 10 days with or without rIL-27. j,k, Frequency of HBV-specific (core, S and polymerase combined) and hCMV-specific (UL55) T cells (n = 12 individuals with CHB) expanded in the absence (full blue) or presence (empty blue) of rIL-27. Each dot represents one donor. j, Individual results from single individuals. l, Pie charts representing the number and proportion of individuals with CHB with higher or lower/equal response to HBV (core, S, POL peptide pools individually tested or combined) and hCMV-UL55 in T cells expanded in the presence of rIL-27 in comparison to expansion without rIL-27. m, ELISpot wells from donor no. 2045 of T cell lines expanded in the presence or absence of rIL-27 and stimulated with the indicated HBV peptide pools. n, Flow cytometry plots showing unstimulated (medium) and HBV-S-stimulated CD4+ and CD8+ T cells expanded in the presence of rIL-27. a–h, n = 3 (Ctrl), 3 (PBS), 3 (rIL-27). Data are representative of at least three independent experiments and are expressed as the mean ± s.e.m. i–l, n = 12. Each dot represents one donor. Paired data were analyzed by paired t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. n, n = 1. Donor no. 2045.
We next investigated whether IL-27 could restore dysfunctional HBV-specific T cell responses in individuals with chronic HBV infection (CHB). To this end, peripheral blood mononuclear cells (PBMCs) from 12 individuals with CHB (4 HBeAg+, 8 anti-HBe+; Supplementary Table 2 and Supplementary Fig. 5) were cultured with or without rIL-27 and stimulated with overlapping peptide pools spanning the entire HBV proteome. Following 10 days of in vitro expansion, the frequency of IFNγ-secreting cells (spot-forming cells (SFCs)) specific to individual HBV proteins (core, PreS1/S2, S, X, and polymerase) was measured using an IFNγ ELISpot assay (Fig. 7i). In parallel, PBMCs were stimulated with human cytomegalovirus (hCMV) UL55 peptides as a control to confirm that IL-27 selectively enhances HBV-specific responses. IL-27 increased the frequency of IFNγ-SFCs specific to HBV peptides in most individuals (92%), with no comparable enhancement in CMV UL55-specific responses (Fig. 7j,k). Enhancement was particularly notable in two HBeAg+ donors (Fig. 7k, donors no. 2123, 1981), although statistical comparisons were limited by sample size. Polymerase-specific responses showed the most consistent improvements (8 of 9 individuals), but improvements were also observed for S-specific and core-specific responses (Fig. 7l and Supplementary Fig. 5). Notably, while some individuals showed an across-the-board increase in HBV-specific responses with IL-27, others exhibited selective enhancement for specific antigens. In donor no. 2045, IL-27 markedly expanded HBV-S-specific responses, as evidenced by a substantial increase in IFNγ-SFCs and intracellular cytokine staining of HBV-specific T cells (Fig. 7k,m and Supplementary Fig. 5), demonstrating improved functional capacity of HBV-S-specific CD8+ T cells (Fig. 7n).
Collectively, these findings from human samples suggest that IL-27 may have therapeutic potential in augmenting the functional capacity of HBV-specific T cells in patients with chronic HBV infection, offering a promising avenue for restoring antiviral immunity in this setting.
Discussion
In this study, we show that antigen-experienced CD4+ Teff cells can prevent or reverse the dysfunction imposed on HBV-specific CD8+ T cells by hepatocellular priming, restoring effector function and suppressing viral replication. This rescue is mediated through a liver-intrinsic circuit in which CD4+ T cells license KCs via CD40–CD40L engagement, leading to IL-12 and IL-27 production. IL-12 amplifies the helper pool, while IL-27 is necessary and sufficient to reinvigorate dysfunctional CD8+ T cells. These findings provide a mechanistic explanation for the long-standing correlation between robust CD4+ T cell immunity and HBV control in chimpanzees9 and humans10,11,36,37,38,39,40,41,42, and identify a tractable therapeutic axis.
Our results extend the classical model of CD4-mediated help within SLOs16,43. Although canonical models emphasize synchronous tri-cellular encounters of CD4+ T cells, CD8+ T cells and DCs15,18,44, more recent work supports a staged paradigm in which CD4+ T cells help follow CD8+ T cell priming, often in separate anatomical compartments19,45,46. By isolating the effector phase and decoupling priming events, we demonstrate that pre-activated CD4+ T cells can deliver restorative signals through local licensing of hepatic APCs, revealing that the liver itself can serve as an autonomous site for CD4+ T cell-mediated immune modulation. This aligns with emerging evidence that CD4+ T cells can license APCs outside SLOs, particularly in persistent infections and cancer47,48,49,50.
Within this framework, KCs emerge as the dominant antigen-presenting platform, extending previous work from our laboratory7. KCs outnumber hepatic DCs by approximately 10:1, and are strategically positioned within liver sinusoids to efficiently sense blood-borne antigens51,52. Unlike periportal DCs53, KCs have direct access to antigens released from hepatocytes and initiate immune response without requiring trafficking to lymph nodes54,55. Hepatic DCs may also possess intrinsic functional limitations compared to their lymphoid counterparts. Together, these features may explain why KCs, rather than DCs, support intrahepatic T cell cooperation in chronic viral infections7,8,31,56.
Work has shown that hepatic cDC1 can also be licensed by CD4+ T cells to amplify CD8+ T cell responses within periportal niches57. Although we cannot exclude similar processes during HBV infection, English et al.57 observed CD4+ T cell help only after lymph node priming of both T cell subsets. In contrast, our data show that CD4+ Teff cells can act directly within the liver to revive dysfunctional CD8+ T cells primed locally, without prior coordination in SLOs. While neither model fully recapitulates the complexity of human HBV infection, our findings are particularly relevant for chronic infection, where intrahepatic CD8+ T cell dysfunction predominates and local reactivation through CD4+ T cell-mediated licensing may offer a viable therapeutic strategy16.
Beyond confirming that CD40 engagement restores intrahepatic T cell function23, our work identifies IL-27 as the critical cytokine mediating CD8+ T cell rescue. CD40–CD40L interactions trigger IL-27 release from licensed KCs, and IL-27’s ability to balance effector activation with regulation has been documented across viral infections and cancer58,59,60,61,62. In the chronically infected liver, we show that IL-27 is indispensable: blocking IL-27 collapses CD8+ T cell rescue, whereas administering IL-27 reinstates effector function. Thus, CD4+ T cells harness IL-27 to override the liver’s tolerogenic bias, offering a therapeutic strategy.
The translational potential of this axis is underscored by our ex vivo experiments with PBMCs from chronically infected patients, where IL-27 consistently enhanced HBV-specific CD8+ responses. Together with IL-27’s favorable safety profile63,64, these results position IL-27 as an attractive, low-toxicity lever for immune-based HBV therapy.
Several limitations should be acknowledged. Our model does not recapitulate the full course of HBV infection, including naive T cell priming, and no small-animal model currently enables study of early HBV-specific T cell priming under productive infection. Using pre-activated CD4+ T cells allowed us to isolate the effector phase but cannot capture all aspects of natural infection. Similarly, rVSV immunization, while effective for TH1 polarization, may imprint functional properties distinct from hepatotropic priming. Finally, while TCR transgenic systems enable mechanistic dissection, they simplify the TCR repertoire and may not fully reflect endogenous human responses.
In summary, we delineate a liver-centric mechanism whereby CD4+ Teff cells license KCs to produce IL-27 and restore antiviral CD8+ T cell function. These findings broaden the paradigm of T cell help beyond lymphoid organs and nominate the KC–IL-27 axis as a promising target for immune-based strategies aimed at functional cure of chronic HBV infection.
Methods
Mice
C57BL/6, CD45.1 (inbred C57BL/6), BALB/c, Thy1.1 (CBy.PL(B6)-Thya/ScrJ), Rag1−/− (B6.129S7-Rag1tm1Mom/J), CD11c-iDTR (B6.FVB-1700016L2RikTg(Itgax-DTR/EGFP)57Lan/J), CAG-eGFP (C57BL/6-Tg(CAG-EGFP)1Osb/J) and CAG-DsRed (B6.Cg-Tg(CAG-DsRed*MST)1Nagy/j) mice were purchased from Charles River or The Jackson Laboratory. Clec4f-iDTR-YFP mice were generated as previously described31. MHC-II−/− (B6.129S2-H2dlAb1Ea/J) mice were obtained through the Swiss Immunological Mutant Mouse Repository (Zurich, Switzerland). MUP-core transgenic mice (lineage MUP-core 50, inbred C57BL/6, H-2Kb) have been previously described6,29. HBV replication-competent transgenic mice (lineage 1.3.32, inbred C57BL/6, H-2Kb) have been previously described25. HBV transgenic mice were used as C57BL/6 or C57BL/6 mice crossed with BALB/c H-2bxd F1 hybrids. Cor93 TCR transgenic mice (lineage BC10.3, inbred CD45.1) have been previously described23. Env28 TCR transgenic mice (lineage 6C2.36, inbred Thy1.1 BALB/c) have been previously described23. Env126 TCR transgenic mice (Rag1−/− CD45.1/2 C57BL/6 inbred) were generated in this laboratory, in collaboration with BioNTech. For intravital imaging experiments, Cor93 and Env28 TCR transgenic mice were crossbred with the CAG-DsRed lineage and Env126 TCR transgenic mice were crossbred with the CAG-eGFP lineage for homozygous fluorescent protein expression. Mice were housed under specific pathogen-free conditions and used at 8–10 weeks of age. No randomization was used in group allocation; experimental groups were predefined based on genetic background or treatment. In all experiments, mice were matched for age, sex and (for the 1.3.32 animals) serum HBeAg levels before experimental manipulations. In selected experiments, 1.3.32 mice were matched for serum HBV DNA levels before experimental manipulations. All experimental animal procedures were approved by the Institutional Animal Committee of the San Raffaele Scientific Institute and are compliant with all relevant ethical regulations. Mice were maintained on an ad libitum diet (VRF1 (P), Special Diets Services, 801900) in specific pathogen-free-controlled housing, with a 12-h light–dark cycle and stable environmental conditions.
Bone marrow chimeras
Bone marrow chimeras were generated by treatment of HBV transgenic mice with four consecutive doses of 20 mg per kg body weight of busulfan (1,4-butanediol dimethanesulfonate; Sigma-Aldrich, B2635) before receiving the new bone marrow (6 h after last dose); mice were allowed to reconstitute for at least 8 weeks before use.
Single-cell sorting
For Env126 transgenic mouse generation, WT C57BL/6 mice were immunized weekly with 100 µg Env126–138 peptide and 50 µg polyI:C for 3 weeks by subcutaneous injection. Seven days after the third injection, 5 × 106 splenocytes were cultured for 15 h in the presence of 4 µg ml−1 peptide or with 2 µg ml−1 concanavalin A (Sigma-Aldrich). Activated cells were labeled using IFNγ-secretion assay-APC kit and were further enriched via anti-APC-microbeads according to the manufacturer’s protocol (Miltenyi, 130-090-433, 130-090-855) using the magnetic-activated cell sorting technique. CD4+CD8−IFNγhi single T cells (anti-CD4, clone GK1.5, eBioscience, 48-0041-82 and anti-CD8α clone 53-6.7, BD Biosciences, 560777) were sorted (BD FACSAria) into wells of 96-well V-bottom plates (Greiner Bio-One) containing 6 μl of a mild hypotonic cell lysis buffer per well consisting of 0.2% Triton X-100, 0.2 μl RiboLock RNase inhibitor (Thermo Scientific, EO0381), 5 ng poly(A) carrier RNA (Qiagen, 1068337) and 1 μl dNTP mix (10 mM, Biozym, 331520) in RNase-free water. Plates were sealed, centrifuged and stored at −65 °C to −85 °C directly after sorting.
For Env126 CD4+ TN cell and Teff cell sorting, single-cell suspensions were prepared from mouse liver and spleen according to previously described protocols. Cells were subsequently stained with the following panel: LIVE/DEAD Fixable Far-Red dye (Invitrogen, L34973), CD11b-eFluor450 (M1/70, eBioscience, 48-5698-82), CD19-eFluor450 (1D3, eBioscience, 48-0193-82), NK1.1-Pacific Blue (PK136, BioLegend, 108722), CD8a-eFluor450 (53-6.7, eBioscience, 48-0081-82), CD45.1-Alexa Fluor 488 (A20, BioLegend, 110718), CD45.2-PE (104, BioLegend, 109808), Ter119-PE-Cy7 (TER-119, BioLegend, 116222), B220-PE-Cy7 (RA3-6B2, BD Bioscience, 552772) and CD4-Alexa Fluor 647 (GK15, BioLegend, 100426). Cells were resuspended at a concentration of approximately 15–20 × 106 cells per ml in sorting buffer (PBS with 0.04% BSA Invitrogen UltraPure, AM2616). Live CD4+ CD45.1+, CD45.2+, CD11b−, CD19−, NK1.1−, CD8a−, Ter119− and B220− cells originally from Env126 CD45.1.2 mice were sorted using a BD FACSAria cell sorter equipped with a 100-µm nozzle at 20 PSI and maintained at 4 °C. Sorted target cells were collected in 1.5-ml microcentrifuge tubes pre-coated overnight with 1% UltraPure BSA at 4 °C. The final cell purity exceeded 98%.
Cloning and selection of antigen-specific TCRs
Plates with sorted CD4+CD8−IFNγhi T cells were thawed, and template-switch cDNA synthesis was performed as described65 with adapted primers specific to murine TCRα and TCRβ constant genes. TRA and TRB fragments were further amplified66, sequenced and the respective V(D)J junctions analyzed using the IMGT/V-Quest tool. DNAs of novel and productively rearranged corresponding TCR chains were digested using NotI and cloned into pST1 vectors67 containing the appropriate constant region for in vitro transcription of complete TCRα/β chains. The reactivity of the isolated TCRα/β chains was validated in active C57BL/6 T cells transfected with respective TCR mRNA, using Env126–138 pulsed bone marrow-derived DCs in an IFNγ ELISpot assay, as previously described68. TCR7 (alpha chain: V7-4*2orV7D-4J15C and beta chain: V4D2J1-6C1) was used for the generation of the Env126 Tg mouse.
Viruses and viral vectors
Mice were injected intravenously (i.v.) with 106 plaque-forming units of replication-competent recombinant VSV expressing HBsAg (rVSVEnv) vector 24 h after Env126 CD4+ T cell adoptive transfer. All infectious work was performed in designated BSL-3 workspaces, in accordance with institutional, national and international guidelines.
T cell isolation, adoptive transfer and in vivo treatments
In selected experiments, WT C57BL/6 mice were immunized subcutaneously with 100 μg of Env126–138 peptide and 50 μg of polyI:C immune adjuvant at three time points every 7 days.
Isolation of CD8+ T cells from spleens of Cor93 or Env28 mice and CD4+ T cells from spleens of Env126 transgenic mice were performed as previously described6. Isolation of CD4+ T cells from spleens of Env126 transgenic mice was performed by negative immunomagnetic selection using the EasySep Mouse CD4+ T Cell Isolation Kit (StemCell Technologies). Mice were injected i.v. with 106 HBV-specific (Cor93 or Env28) naive CD8+ T cells and/or HBV-specific effector CD4+ T cells (Env126 Teff cells). In the indicated experiments, mice were splenectomized and treated i.v. with 200 μg of anti-CD62L monoclonal antibody (clone MEL-14, BioXcell) 48 h and 4 h before cell injection and 24 h after cell transfer. Splenectomy was performed according to standard procedures69 and as previously described6. For FTY720 treatment, mice were injected with 1 mg per kg body weight of fingolimod (Sigma) intraperitoneally (i.p.) every 48 h throughout the experiment. In selected experiments, mice were treated i.p. with busulfan and reconstituted i.v. with bone marrow from CD11c-iDTR-GFP mice (107). In selected experiments, DCs and KCs were depleted by injecting i.p. 25 ng per gram body weight of diphtheria toxin (Sigma-Aldrich) 3 days and 1 day before T cell transfer and 1 day and 3 days after cell transfer. In the indicated experiments, KCs were depleted by i.v. injection of clodronate-containing liposomes (10 μl per gram body weight; Liposoma B’V) 2 days before T cell injection70,71. In the indicated experiments, mice were treated i.p. with 250 μg of anti-CD40L monoclonal antibody (clone MR-1) or 250 μg anti-CD40L monoclonal antibody (clone D265A), 250 μg of anti-IL-12 p40 monoclonal antibody (clone C17.8), 250 μg of anti-IL-27 p28 monoclonal antibody (clone MM27.7B1) and 2.5 μg of rIL-27 (Fc: LALA-PG-KIH heterodimer, Adipogen). All monoclonal antibodies were purchased from BioXcell, unless otherwise indicated.
In vitro cell activation
In the indicated experiments, Env126 CD4+ T cells were cultured in vitro under 4 h of stimulation in lymphocyte complete medium: RPMI 1640, penicillin–streptomycin (100 IU ml−1 and 100 µg ml−1, Corning), 2 mM l-glutamine (Corning), 50 µM 2-mercaptoehanol (Sigma-Aldrich), HEPES 10 mM (Corning), NEAA (non-essential amino acids, Corning) at 100 µM. HBsAg126–138 was used at a concentration of 1 µg ml−1. Ionomycin calcium salt (Sigma-Aldrich), was used at a concentration of 1 µg ml−1 in combination with PMA (Sigma-Aldrich) at a concentration of 50 ng ml−1. In the indicated experiments, Env126 CD4+ T cells were cultured in vitro for 48 h in complete medium with HBsAg126–138 peptide (1 µg ml−1) or with Dynabeads Mouse T-Activator CD3/CD28 (Thermo Fisher Scientific, 11456D) following the manufacturer’s instructions. For ex vivo intracellular staining to assess cytokine production before FACS analyses, cell suspensions were incubated for 4 h at 37 °C in complete medium, with HBsAg126–138, HBsAg28–39 or HBcAg93–100 at 2 µg ml−1 with 1 µg ml−1 brefeldin A (Sigma-Aldrich) and Monensin (BioLegend) according to the manufacturer’s instructions. In selected experiments, in vitro cultured CD8+ T cells were stimulated for 48 h with rIL-27 at 25 ng ml−1 (R&D Systems, 7430-ML).
In vitro cell labeling
In the indicated experiments, WT or Env126 CD4+ T cells were labeled with CellTrace Violet Cell Proliferation Kit (Invitrogen, C34571) following the manufacturer’s protocol. Briefly, WT or Env126 CD4+ T cells were isolated, washed and resuspended in PBS at a concentration of 106 cells per ml in the working dye solution for 20 min at 37 °C. After the incubation, complete medium was added to the cell suspension that was rested at room temperature for 5 min to dilute and remove free remaining dye. Labeled cells were then resuspended in complete medium ready to be cultured.
Cell isolation and flow cytometry
Briefly, cells were plated at 2 × 106 cells per well in a 96-well U-bottom tissue culture plate with cognate peptide (2 µg ml−1) for 4 h in an incubator in complete RPMI medium containing 10 µg ml−1 of brefeldin A and 10 U ml−1 of IL-2 (EL-4 supernatant). At the end of the stimulation, cells were washed and stained as described above. Cell viability was assessed by staining with Viability 405/520 fixable dye (Miltenyi, 130-130-404) and LIVE/DEAD Fixable Far-Red dye (Invitrogen, L34973). In each experiment, Env28 CD8+ T cells were identified as live, CD45+, CD8+CD4− Thy1.1+; Env126 CD4+ T cells were identified as live, CD45+, CD8−CD4+ CD45.2+CD45.1+; and Cor93 CD8+ T cells were identified as live, CD45+, CD8+CD4− CD45.1+.
Staining of cell surface markers was performed with Brilliant Stain buffer (BD Biosciences, 566349) according to the manufacturer’s instructions. Antibodies are described in this order: target, clone name, catalog number and dilution to ensure precise identification of the reagent used. Antibodies from BD Biosciences: anti-CD45.2 (104, 564616; 1:200 dilution), anti-Thy1.1 (OX-7, 740917; 1:200 dilution), anti-CD8 (53-6.7, 612898; 1:300 dilution), anti-B220 (RA3-6B2, 564662; 1:300 dilution), anti-CD19 (1D3, 562291; 1:200 dilution), anti-CD3 (145-2C11, 562286; 1:100 dilution), anti-Ly6G (1A5, 562700; 1:300 dilution), anti-CD49b (DX5, 562453; 1:100 dilution), anti-CD44 (IM7, 569705, 560781; 1:200 dilution), anti-CD69 (H1.2F3, 612793, 552879; 1:100 dilution), anti-CD86 (GL1, 564199; 1:100 dilution), anti-IA-IE (M5/114.15.2, 748846, 2G9, 569244; 1:300 dilution), anti-H2-kB (AF6-88.5, 742861; 1:200 dilution), anti-CD40 (44986, 745218; 1:100 dilution), anti-TIM4 (21H12, 742774; 1:100 dilution), anti-CD28 (37.51, 740466; 1:100 dilution), anti-CD70 (FR70, 740931; 1:100 dilution), anti-Ly6A/E (D7, 756372; 1:100 dilution), anti-Ly108 (13G3, 755697; 1:100 dilution), anti-TIGIT (1G9, 744212; 1:100 dilution), anti-NKG2A/C/E (20d5, 740549; 1:100 dilution), anti-2B4 (2B4, 740671; 1:100 dilution), anti-Bcl-6 (K112-91, 562401; 1:75 dilution), anti-IFNγ (XMG1.2, 562333; 1:200 dilution), anti-T-bet (O4-46, 569089; 1:75 dilution) and anti-TNF (MP6-XT22, 563943; 1:100 dilution). From BioLegend: anti-CD45 (30-F11, 103108; 1:400 dilution), anti-CD45.1 (clone A20, 110743; 1:300 dilution), anti-CD4 (GK1.5, 100480; 1:200 dilution), anti-CD62L (MEL-14, 104438, 104453; 1:200 dilution), anti-CD80 (16-10A1, 104738; 1:100 dilution), anti-CD25 (PC61, 102075, 102020; 1:100 dilution), anti-CD11c (QA18A72, 161107; 1:100 dilution), anti-CD11b (M1/70, 101285; 1:100 dilution), anti-CD44 (IM7, 103028; 1:200 dilution), anti-F4/80 (BM8, 123110, 123130; 1:100 dilution), anti-CD64 (X54-5/7.1, 139311; 1:100 dilution), anti-CD69 (H1.2F3, 104527; 1:100 dilution), anti-ICAM-1 (YN1/1.7.4, 116116; 1:100 dilution), anti-ESAM (1G8/ESAM, 136203; 1:100 dilution), anti-CD206 (C068C2, 141719; 1:100 dilution), anti-Lag-3 (C9B7W, 125248; 1:100 dilution), anti-CD39 (Duha59, 143819; 1:100 dilution), anti-CD31 (390, 102424; 1:100 dilution), anti-IL-2 (JES6-5H4, 503837; 1:100 dilution), anti-CD107a (1D4B, 121608; 1:80 dilution), anti-ICOS (15F9, 107716; 1:100 dilution), anti-PD-1 (29F.1A12, 125253; 1:100 dilution), anti-NK1.1 (S17016D, 156529; 1:100 dilution), anti-KLRG1 (2F1/KLRG1, 138410; 1:100 dilution), anti-CXCR3 (S18001A, 155923; 1:100 dilution), anti-CTLA-4 (UC10-4B9, 106314; 1:75 dilution), anti-4-1BB (17B5, 107105; 1:100 dilution), anti-OX40 (OX-86, 119414; 1:100 dilution), anti-TNF (MP6-XT22, 506313; 1:200 dilution), anti-TIM3 (RMT3-23, 119738; 1:100 dilution), anti-CXCR5 (L138D7, 145532; 1:75 dilution), anti-CXCR6 (SA051D1, 151111; 1:100 dilution), anti-IFNγ (XMG1.2, 505830; 1:200 dilution). From eBioscience: anti-CD4 (RM4-5, 48-0042-82; 1:200 dilution), anti-CD27 (LG.7F9, 25-0271-80; 1:100 dilution), anti-Ki-67 (SolA15, 48-5698-82; 1:400 dilution). Antibodies from Invitrogen: anti-Foxp3 (FJK-16s, 53-5773-82; 1:100 dilution), anti-CD11c (N418, 25-0114-82; 1:100 dilution), anti-T-bet (4b10, 25-5825-80; 1:75 dilution), anti-TOX (TXRX10, 50-6502-82; 1:100 dilution), anti-Grzm-B (GB11, GRB04, GRB05; 1:80 dilution). Human antibodies included: anti-CD3 (clone HIT3a, BioLegend, 300306; 1:40 dilution), anti-CD4 (clone SK3, BD Bioscience, 563875; 1:20 dilution), anti-CD8 (clone RPA-T8, BD Bioscience, 557746; 1:100 dilution) and anti-IFNγ (clone B27, BioLegend, 506510; 1:20 dilution).
Recombinant dimeric H-2Ld–Ig or H-2Kb–Ig fusion proteins (BD Biosciences) complexed with peptides derived from HBsAg (Env28–39, IPQSLDSWWTLS, Primm) or from HBcAg (Cor93–100, MGLKFRQL, Primm), respectively, were prepared according to the manufacturer’s instructions. Dimer staining was performed as described and used to quantify HBV-specific CD8+ T cells (Env28 and Cor93)6. Peptide HBs126–138-loaded tetramer (I-Ab Env126–138) and human CLIP peptide-loaded negative control (I-Ab CLIP) were provided by the National Institutes of Health (NIH) Tetramer Core Facility. Tetramer staining was performed by incubating cell suspensions at 37 °C for 1 h according to the manufacturer’s instructions. Flow cytometry staining for all transcription factors and intranuclear proteins was performed using Foxp3/Transcription Factor Staining Buffer Set (eBioscience, 00-5523-00), following the manufacturer’s instructions. All flow cytometry analyses have been performed in FACS buffer comprising PBS with 2 mM EDTA and 2% FBS (Corning). Flow cytometry analyses were performed on the BD FACSCanto Clinical Flow Cytometry System, BD FACSymphony A5 Cell Analyzer or Cytek Aurora. Flow cytometry data were analyzed with FlowJo software (TreeStar, v10).
Southern blot analysis
Genomic DNA was isolated from frozen livers using the phenol–chloroform method and analyzed for intrahepatic HBV DNA contents by Southern blotting, as previously described7. No molecular weight markers were included, as the migration pattern of HBV DNA forms, including RC, DS, SS DNA and integrated TG, has been well characterized25. Band identity was assigned based on electrophoretic mobility and comparison to these widely recognized standards. All blots are shown for qualitative purposes only.
Biochemical analysis
The extent of hepatocellular injury was monitored by the measuring of sALT activity at different and multiple time points after experimental manipulation and cell transfer, as already described6. Normal intervals of sALT activity are between 20 and 80 U l−1.
RT–PCR
Total RNA was extracted from frozen livers using the ReliaPrep RNA Tissue Miniprep System (Promega), according to the manufacturer’s instructions, as described6. Genomic DNA contamination was removed using TURBO DNA-free DNase (Ambion). Around 1 μg of total RNA was reverse transcribed with Superscript IV Vilo (Life Technologies) before qPCR analysis for mouse Il27 (TaqMan Mm00461163, Life Technologies) and Il12b (TaqMan Mm 01288989, Life Technologies). All experiments were performed in triplicate and normalized to the reference gene GAPDH (TaqMan Mm99999915, Life Technologies). All nucleic acids were purified from serum using the MiniElute Virus Spin Kit (Qiagen). Viremia was determined by plotting a standard curve using HBV plasmid DNA and the Core TaqMan probe (forward TACCGCCTCAGCTCTGTATC, reverse CTTCCAAATTAACACCCACCC, probe TCACCTCACCATACTGCACTCAGGCAA). Reactions were run and analyzed on the QuantStudio 5 instrument (Life Technologies).
Confocal immunofluorescence and histochemistry
At the time of autopsy, mouse livers were perfused via the inferior vena cava with 10 ml of PBS as liver pieces were surgically removed and instantly fixed in paraformaldehyde at 4% concentration for 16–20 h. The solutions were then dehydrated in 30% sucrose before their embedding into OCT freezing media (Bio-Optica). Liver sections were obtained with a Leica CM1520 Cryostat (20 µm per section). Then, liver sections were stained as already described6. Antibodies used included anti-F4/80 Alexa Fluor 647 (Invitrogen, MF48021) and anti-CD38 Brilliant Violet 421 (BioLegend, 102732). All images were acquired using a Mavig RS-G4 confocal microscope.
For all the immunohistochemical staining, liver pieces were harvested in zinc formalin solution and transferred into 70% ethanol solutions 24 h later. All the tissues were then embedded in paraffin and stained as previously described6. Primary antibodies included monoclonal rat anti-Ki-67 (clone TEC-3, Dako, M7249), polyclonal rabbit anti-HBcAg (Dako, B0586), monoclonal rabbit anti-cytokeratin 7 (clone ERP17078, Abcam, ab181598), monoclonal rabbit anti-CD3 (clone SP7, Abcam, ab16669) and polyclonal rabbit anti-cleaved caspase-3 (Asp175, CST, 9661). Bright-field images were acquired using an Aperio Scanscope CS2 microscope and ImageScope software (Leica Biosystems), following the manufacturer’s guidelines. In each experiment, histochemical analysis was conducted for every mouse using the median-left liver lobe, following a standardized protocol established in collaboration with our institutional histopathology core facility. The entire lobe section was scanned to avoid selection bias and ensure inclusion of all tissue regions. For qualitative assessment, regions of interest were selected from these whole-lobe scans based on tissue integrity and staining quality, each covering an area of approximately 600 µm². These regions of interest were chosen to capture diverse hepatic microenvironments (for example, periportal, pericentral and mid-zonal regions), rather than isolated or extreme features. Selection was performed independently by two investigators (blinded to condition) and confirmed by a pathologist to ensure consistency and reproducibility. All HBcAg IHC stainings are presented as a qualitative assessment of antigen distribution, illustrating its presence or absence across conditions. The number of animals and sections analyzed per group is indicated in each figure legend. Immunohistochemical imaging analyses were performed with QuPath 0.5.0-x64 software.
Intravital multiphoton microscopy
Liver intravital multiphoton microscopy was performed as described6. Liver sinusoids were visualized by injecting nontargeted Quantum Dots 655 (Invitrogen) i.v. during image acquisition. Images were acquired with a LaVision BioTec TriMScope II coupled to a Nikon Ti-U inverted microscope enclosed in a custom-built environmental chamber (Life Imaging Services) that was maintained at 37–38 °C with heated air. Fluorescence excitation was provided by two tunable femtosecond-pulsed titanium–sapphire lasers (680–1,080 nm, 120-fs pulse width, 80-MHz repetition rate, Ultra II, Coherent), an Optical Parametric Oscillator (1,000–1,600 nm, 200-fs pulse width, 80-MHz repetition rate, Chameleon Compact OPO, Coherent). The setup includes four non-descanned photomultiplier tubes (Hamamatsu H7422-40 GaAsP High Sensitivity PMTs and Hamamatsu H7422-50 GaAsP High Sensitivity red-extended PMT from Hamamatsu Photonics K.K.) and a ×25, 1.05-NA, 2-mm working distance, water-immersion multiphoton objective (Olympus). For four-dimensional analysis of cell migration, stacks of 5–13 square xy sections (1,024 × 1,024 pixel) sampled with 4-μm z-spacing were acquired every 20 s for 20 min. Sequences of image stacks were transformed into volume-rendered, four-dimensional time-lapse movies with Imaris (Bitplane, v9.1.2). The three-dimensional (3D) positions of the cell centroids were segmented by the semiautomated cell tracking algorithm of Imaris.
The single-cell speed was calculated as the mean 3D velocity per cell track, while the instantaneous velocity represents the 3D velocities of each single step in a track. The arrest coefficient was defined as the percentage of time a track instantaneous velocity remains below a given motility threshold (here 5 μm min−1). The track displacement (distance between the initial and the final position of a cell) was divided by the total track length to obtain the corrected straightness, a measure of the directionality of a track. Finally, instantaneous speed, turning angle changes, mean displacement versus time and the motility coefficient were calculated from the x, y and z coordinates of the cell centroids using custom-designed scripts in RStudio.
scRNA-seq library preparation
Env126 CD4+ Teff cells and Env126 CD4+ TN cells were fixed according to the Evercode Fixation (Parse Biosciences) protocol, and single-cell RNA libraries were prepared using the Evercode split-pool combinatorial barcoding technology approach using the ParseBio Evercode WT v2 kit. Sequencing was performed on the Illumina NovaSeq 6000 sequencer with the S4 flow cell 200 cycle kit according to the sequencing protocol provided by the Evercode kit (v1.5) at a desired depth of 30,000 reads per cell.
scRNA-seq data analysis
Parse Biosciences analysis pipeline ‘split-pipe’ (v.1.1.0p) was used with default parameters to perform demultiplexing, alignment of sequencing reads against the GRCm38.93 genome and generation of the cell-by-gene count matrix. A raw counts matrix of the aggregated libraries containing the counts for all the samples was imported into R (version 4.2.3) and processed with custom scripts. The ‘isOutlier’ function from the ‘scuttle’ package72 was used to identify outlier and low-quality cells in an unbiased way. For each of the following features, cells with values greater or lower than four median absolute deviations were discarded: library size, number of features, percentage of mitochondrial gene contamination, percentage of ribosomal gene contamination, percentage of hemoglobin gene contamination, percentage of platelet gene contamination. After quality filtering, only cells with a number of features greater than 200 and genes expressed in at least three cells were kept. Doublet removal was performed with the scDblFinder R package73; only the cells automatically classified as a ‘singlet’ by the algorithm in the package were kept. Cells that were not CD4+ T cells were discarded from each sample individually by calling the ‘SingleR’ function using the ‘celldex MouseRNAseqData’ dataset74,75 aided by manual curation. A Seurat (version 4.3.0)76 object was created for each filtered sample and merged into a single dataset. The complete dataset was normalized and scaled following the standard Seurat workflow (scaling to 10,000 and log transformation log1p). The top 3,000 genes identified by Seurat’s FindVariableFeatures using the vst method were used for principal component analysis. Uniform manifold approximation and projection reduction was then applied on the first 30 principal components. Graph-based clustering was performed to cluster cells according to their gene expression profile with the FindClusters function in Seurat with default parameters, and a resolution value of 0.2 was chosen. Differential gene expression was calculated with the FindAllMarkers function in Seurat. Gene-set or signature scores were obtained with the AddModuleScore Seurat function. CD4+ T cells signatures were obtained from Kiner et al.77. Heat maps were prepared with the ComplexHeatmap package; feature plots were generated with the function FeaturePlot_scCustom from the scCustomize R package78,79. Gene-set enrichment analysis was performed using the fgsea R package (v.1.8.0)80 on all genes ranked by log2 fold change (from FindMarkers Seurat output) from the selected comparison. P values were calculated using 1,000,000 permutations and were corrected using the Benjamini–Hochberg method. Gene sets with a false discovery rate <0.05 were considered significant. R packages used for data analysis were scuttle (v.1.8.4), scDblFinder (v1.12.0), SingleR (v2.0.0), Seurat (v4.3.0), scCustomize (v1.1.3), celldex (v1.8.0).
Study participants and approval
A total of 12 individuals with chronic HBV infection (HBsAg+) were included (Supplementary Table 2), of whom 4 had HBeAg-positive and 8 had HBeAg-negative chronic HBV infection. All 12 individuals were treatment-naive, had no known history of hepatitis (normal ALT) and were non-cirrhotic with a normal FibroScan result. Supplementary Table 2 summarizes the available clinical and virological parameters. Blood donors were recruited from the viral hepatitis clinic at The Royal London Hospital. Written informed consent was obtained from all participants. The study was conducted in accordance with the Declaration of Helsinki and approved by the Barts and the London NHS Trust local ethics review board and the NRES Committee London–Research Ethics Committee (reference 10/H0715/39) and by the Singapore National Healthcare Group ethical review board (DSRB 2008/00293).
Clinical and virological parameters
On recruitment to the study, viral serology and HBV DNA levels were tested. HBsAg, HBeAg and anti-HBe levels were measured with a chemiluminescent microparticle immunoassay (CMIA; Architect Assay, Abbott Diagnostics). HBV DNA levels in serum were quantified by real-time PCR (COBAS AmpliPrep/COBAS TaqMan HBV test v2.0; Roche Molecular Diagnostics).
HBV peptide library
A set of 15-mer peptides overlapping by ten amino acids was used to identify HBV-specific T cells. The peptides covered the sequences of the HBV protein core, envelope (S) and polymerase (Pol) of genotypes A, B, C and D and were purchased from T Cell Diagnostics. Similarly, 15-mer peptides overlapping by ten amino acids that covered the sequence of hCMV protein UL55 were used to identify CMV-specific T cells and were purchased from GenScript.
PBMC isolation and T cell culture
PBMCs were isolated from peripheral blood by Ficoll gradient and cryopreserved. Cells were thawed and T cell lines were generated as follows: 20% of PBMCs were pulsed with 10 μg ml−1 of the overlapping HBV or CMV peptides for 1 h at 37 °C, subsequently washed, and co-cultured with the remaining cells in AIM-V medium (Gibco; Thermo Fisher Scientific) supplemented with 2% AB human serum (Gibco; Thermo Fisher Scientific). T cell lines were cultured for 10 days in the presence of 20 U ml−1 of rIL-2 (R&D Systems) with or without 50 ng ml−1 of rIL-27. Half of the medium was refreshed on day 4 and day 8.
ELISpot assays
ELISpot assays for the detection of IFNγ-producing cells were performed on in vitro-expanded T cell lines using either HBV peptide pool core, envelope (S), polymerase 1 (Pol 1), polymerase 2 (Pol 2) or CMV peptide pool UL55. T cell lines were incubated overnight at 37 °C with pools of HBV/CMV peptides (2 μg ml−1), where final dimethylsulfoxide concentrations did not exceed 0.2%. IFNγ ELISpot assays (Millipore) were performed as previously described81. Spot-forming units were quantified with ImmunoSpot 7.0.26.0.
Liver histology from HBV-infected chimpanzees
Liver tissue from an HBV-infected chimpanzee (A3A005) was analyzed. All relevant details regarding sex, age, and body weight before infection are available in the published dataset9. The animal was handled in accordance with humane care and use guidelines established by the Animal Research Committees at the NIH, The Scripps Research Institute and Bioqual Laboratories. It was housed individually at Bioqual Laboratories, an AAALAC International-accredited institution under contract with the National Institute of Allergy and Infectious Diseases. The chimpanzee was inoculated with 104 genome equivalents of HBV, a dose that typically results in a self-limited infection9. Liver tissue samples, measuring 5 to 10 mm in length, were obtained via needle biopsy and fixed in 10% zinc formalin for histological analysis. Formalin-fixed paraffin-embedded liver sections were subjected to multiplex immunofluorescence staining using the Opal detection system (Akoya Biosciences) and visualized with fluorescence microscopy. Sections were first deparaffinized and subjected to heat-induced epitope retrieval using Bond Epitope Retrieval Solution (Leica Biosystems). Immunostaining was performed on an automated Leica Bond RX platform using ready-to-use primary antibodies for CD4 (clone 4B12, Leica PA0427), CD8 (clone C8/144B, Dako) and CD68 (clone KP1, Ventana 790-2931). Images were acquired using an HT 2.0 Akoya scanner at a magnification of ×20. Image analysis was performed using Phenochart software (Akoya Biosciences) to assess spatial interactions within the hepatic microenvironment.
Statistical analyses
All results are expressed as means ± s.e.m. All statistical analyses were performed in Prism (GraphPad Software, v10). Statistical tests, sample size and P values are indicated in each figure. No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications6,7,24. Data distribution was assumed to be normal, but this was not formally tested. Data collection and analysis were not performed blind to the experimental conditions. No data were excluded from the analyses unless classified as outliers using the ROUT method (Q = 5%), which combines robust regression and false discovery rate control. This method was used to objectively remove technical artifacts while minimizing subjective bias.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
scRNA-seq data for mouse CD4+ T cells have been deposited in the Gene Expression Omnibus under the accession code GSE283038. The processed scRNA-seq data that support the findings of this study are available in Figshare via https://doi.org/10.6084/m9.figshare.27894906. Source data are provided with this paper. All other data supporting the findings of this study are available within the article and its Supplementary Information.
Code availability
The scripts used to analyze the mouse scRNA-seq data can be accessed at https://doi.org/10.6084/m9.figshare.27894906.
References
Iannacone, M. & Guidotti, L. G. Immunobiology and pathogenesis of hepatitis B virus infection. Nat. Rev. Immunol. 22, 19–32 (2022).
Kawashima, K., Andreata, F., Beccaria, C. G. & Iannacone, M. Priming and maintenance of adaptive immunity in the liver. Annu. Rev. Immunol. 42, 375–399 (2024).
Venzin, V., Beccaria, C. G., Andreata, F., Fumagalli, V. & Iannacone, M. Intrahepatic immunity to hepatitis B virus. J. Hepatol. 81, 911–913 (2024).
Bertoletti, A. & Ferrari, C. Adaptive immunity in HBV infection. J. Hepatol. 64, S71–S83 (2016).
Maini, M. K. & Burton, A. R. Restoring, releasing or replacing adaptive immunity in chronic hepatitis B. Nat. Rev. Gastroenterol. Hepatol. 16, 662–675 (2019).
Bénéchet, A. P. et al. Dynamics and genomic landscape of CD8+ T cells undergoing hepatic priming. Nature 574, 200–205 (2019).
De Simone, G. et al. Identification of a Kupffer cell subset capable of reverting the T cell dysfunction induced by hepatocellular priming. Immunity 54, 2089–2100 (2021).
Warren, A. et al. T lymphocytes interact with hepatocytes through fenestrations in murine liver sinusoidal endothelial cells. Hepatology 44, 1182–1190 (2006).
Asabe, S. et al. The size of the viral inoculum contributes to the outcome of hepatitis B virus infection. J. Virol. 83, 9652–9662 (2009).
Penna, A. et al. Long-lasting memory T cell responses following self-limited acute hepatitis B. J. Clin. Invest. 98, 1185–1194 (1996).
Ferrari, C. et al. Cellular immune response to hepatitis B virus-encoded antigens in acute and chronic hepatitis B virus infection. J. Immunol. 145, 3442–3449 (1990).
Penna, A. et al. Predominant T-helper 1 cytokine profile of hepatitis B virus nucleocapsid-specific T cells in acute self-limited hepatitis B. Hepatology 25, 1022–1027 (1997).
Jung, M. C. et al. Activation of a heterogeneous hepatitis B (HB) core and e antigen-specific CD4+ T-cell population during seroconversion to anti-HBe and anti-HBs in hepatitis B virus infection. J. Virol. 69, 3358–3368 (1995).
Schoenberger, S. P., Toes, R. E. M., Van Dervoort, E. I. H., Offringa, R. & Melief, C. J. M. T-cell help for cytotoxic T lymphocytes is mediated by CD40–CD40L interactions. Nature 393, 480–483 (1998).
Bennett, S. R. M. et al. Help for cytotoxic-T-cell responses is mediated by CD40 signalling. Nature 393, 478–480 (1998).
Borst, J., Ahrends, T., Bąbała, N., Melief, C. J. M. & Kastenmüller, W. CD4+ T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Mitchison, N. A. & O’Malley, C. Three-cell-type clusters of T cells with antigen-presenting cells best explain the epitope linkage and noncognate requirements of the in vivo cytolytic response. Eur. J. Immunol. 17, 1579–1583 (1987).
Ridge, J. P., Di Rosa, F. & Matzinger, P. A conditioned dendritic cell can be a temporal bridge between a CD4+ T-helper and a T-killer cell. Nature 393, 474–478 (1998).
Jobin, K. et al. A distinct priming phase regulates CD8 T cell immunity by orchestrating paracrine IL-2 signals. Science 388, eadq1405 (2025).
Mancini, M., Hadchouel, M., Tiollais, P. & Michel, M. -L. T cells after DNA-based immunization-secreting γ transgenic mouse model by IFN-expression in a hepatitis B surface antigen regulation of hepatitis B virus mRNA. J. Immunol. 161, 5564–5570 (1998).
Kouskoff, V., Signorelli, K., Benoist, C. & Mathis, D. Cassette vectors directing expression of T transgenic mice cell receptor genes in transgenic mice. J. Immunol. Methods 180, 273–280 (1995).
Cobleigh, M. A., Wei, X. & Robek, M. D. A vesicular stomatitis virus-based therapeutic vaccine generates a functional CD8 T cell response to hepatitis B virus in transgenic mice. J. Virol. 87, 2969–2973 (2013).
Isogawa, M., Chung, J., Murata, Y., Kakimi, K. & Chisari, F. V. CD40 Activation rescues antiviral CD8+ T cells from PD-1-mediated exhaustion. PLoS Pathog. 9, e1003490 (2013).
Andreata, F. et al. Therapeutic potential of co-signaling receptor modulation in hepatitis B. Cell 187, 4078–4094 (2024).
Guidotti, L. G., Matzke, B., Schaller, H. & Chisari, F. V. High-level hepatitis B virus replication in transgenic mice. J. Virol. 69, 6158–69 (1995).
Guidotti, L. G. et al. Viral clearance without destruction of infected cells during acute HBV infection. Science 284, 825–829 (1999).
Ishak, K. et al. Histological grading and staging of chronic hepatitis. J. Hepatol. 22, 696–699 (1995).
Smith, C. M. et al. Cognate CD4+ T cell licensing of dendritic cells in CD8+ T cell immunity. Nat. Immunol. 5, 1143–1148 (2004).
Guidotti, L. G., Martinez, V., Lor, Y. -T., Rogler, C. E. & Chisari, F. V. Hepatitis B virus nucleocapsid particles do not cross the hepatocyte nuclear membrane in transgenic mice. J. Virol. 68, 5469–5475 (1994).
Mempel, T. R., Henrickson, S. E. & Von Andrian, U. H. T-cell priming by dendritic cells in lymph nodes occurs in three distinct phases. Nature 427, 154–159 (2004).
Scott, C. L. et al. Bone marrow-derived monocytes give rise to self-renewing and fully differentiated Kupffer cells. Nat. Commun. 7, 10321 (2016).
Celia, M. et al. Ligation of CD40 on dendritic cells triggers production of high levels of interleukin-12 and enhances T cell stimulatory capacity: T–T help via APC activation. J. Exp. Med. 184, 747–752 (1996).
DeLong, J. H. et al. Cytokine- and TCR-mediated regulation of T cell expression of Ly6C and Sca-1. J. Immunol. 200, 1761–1770 (2018).
Yoshida, H. & Hunter, C. A. The immunobiology of interleukin-27. Annu. Rev. Immunol. 33, 417–443 (2015).
Huang, Z. et al. IL-27 promotes the expansion of self-renewing CD8+ T cells in persistent viral infection. J. Exp. Med. 216, 1791–1808 (2019).
Smyk-Pearson, S. et al. Spontaneous recovery in acute human hepatitis C virus infection: functional T-cell thresholds and relative importance of CD4 help. J. Virol. 82, 1827–1837 (2008).
Urbani, S. et al. Outcome of acute hepatitis C is related to virus-specific CD4 function and maturation of antiviral memory CD8 responses. Hepatology 44, 126–139 (2006).
Webster, G. J. M. et al. Incubation phase of acute hepatitis B in man: dynamic of cellular immune mechanisms. Hepatology 32, 1117–1124 (2000).
Jung, M. -C. et al. Activation of a heterogeneous hepatitis B (HB) core and e antigen-specific CD4+ T-cell population during seroconversion to Anti-HBe and Anti-HBs in hepatitis B virus infection. J. Virol. 69, 3358–3368 (1995).
Dong, Y. et al. CD4+ T cell exhaustion revealed by high PD-1 and LAG-3 expression and the loss of helper T cell function in chronic hepatitis B. BMC Immunol. 20, 27 (2019).
Fisicaro, P. et al. Antiviral intrahepatic T-cell responses can be restored by blocking programmed death-1 pathway in chronic hepatitis B. Gastroenterology 138, 682–693 (2010).
Hoogeveen, R. C. et al. Hepatitis B virus-specific CD4 T cell responses differentiate functional cure from chronic surface antigen+ infection. J. Hepatol. 77, 1276–1286 (2022).
Castellino, F. & Germain, R. N. Cooperation between CD4+ and CD8+ T cells: when, where, and how. Ann. Rev. Immunol. 24, 519–540 (2006).
Ferris, S. T. et al. cDC1 prime and are licensed by CD4+ T cells to induce anti-tumour immunity. Nature 584, 624–629 (2020).
Hor, J. L. et al. Spatiotemporally distinct interactions with dendritic cell subsets facilitates CD4+ and CD8+ T cell activation to localized viral infection. Immunity 43, 554–565 (2015).
Eickhoff, S. et al. Robust anti-viral immunity requires multiple distinct T cell-dendritic cell interactions. Cell 162, 1322–1337 (2015).
Zander, R. et al. CD4+ T cell help is required for the formation of a cytolytic CD8+ T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042 (2019).
Ahrends, T. et al. CD4+ T cell help confers a cytotoxic T cell effector program including coinhibitory receptor downregulation and increased tissue invasiveness. Immunity 47, 848–861 (2017).
Magen, A. et al. Intratumoral dendritic cell–CD4+ T helper cell niches enable CD8+ T cell differentiation following PD-1 blockade in hepatocellular carcinoma. Nat. Med. 29, 1389–1399 (2023).
Espinosa-Carrasco, G. et al. Intratumoral immune triads are required for immunotherapy-mediated elimination of solid tumors. Cancer Cell 42, 1202–1216 (2024).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396 (2022).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the Kupffer cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654 (2019).
English, K. et al. The liver contains distinct interconnected networks of CX3CR1+ macrophages, XCR1+ type 1 and CD301a+ type 2 conventional dendritic cells embedded within portal tracts. Immunol. Cell Biol. 100, 394–408 (2022).
Ficht, X. & Iannacone, M. Immune surveillance of the liver by T cells. Sci. Immunol 5, eaba2351 (2020).
Crispe, I. N. Liver antigen-presenting cells. J. Hepatol. 54, 357–365 (2011).
Krenkel, O. & Tacke, F. Liver macrophages in tissue homeostasis and disease. Nat. Rev. Immunol. 17, 306–321 (2017).
English, K. et al. A hepatic network of dendritic cells mediates CD4+ T cell help outside lymphoid organs. Nat. Commun. 15, 1261 (2024).
Kastelein, R. A., Hunter, C. A. & Cua, D. J. Discovery and biology of IL-23 and IL-27: related but functionally distinct regulators of inflammation. Annu. Rev. Immunol. 25, 221–242 (2007).
Murugaiyan, G. & Saha, B. IL-27 in tumor immunity and immunotherapy. Trends Mol. Med. 19, 108–116 (2013).
Chudnovskiy, A. et al. Proximity-dependent labeling identifies dendritic cells that drive the tumor-specific CD4+ T cell response. Sci. Immunol. 9, eadq8843 (2024).
Martin, E. et al. Role of IL-27 in Epstein–Barr virus infection revealed by IL-27RA deficiency. Nature 628, 620–629 (2024).
Bréart, B. et al. IL-27 elicits a cytotoxic CD8+ T cell program to enforce tumour control. Nature https://doi.org/10.1038/s41586-024-08510-w (2025).
Zhu, J. et al. IL-27 gene therapy induces depletion of Tregs and enhances the efficacy of cancer immunotherapy. JCI Insight 3, e98745 (2018).
Yoshimoto, T. et al. Potential clinical application of interleukin-27 as an antitumor agent. Cancer Sci. 106, 1103–1110 (2015).
Sahin, U. et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature 585, 107–112 (2020).
Paret, C. et al. CXorf61 is a target for T cell based immunotherapy of triple-negative breast cancer. Oncotarget 6, 25356–25367 (2015).
Holtkamp, S. et al. Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells. Blood 108, 4009–4017 (2006).
Vormehr, M. et al. A non-functional neoepitope specific CD8+ T-cell response induced by tumor derived antigen exposure in vivo. Oncoimmunology 8, 1553478 (2019).
Reeves, J. P., Reeves, P. A. & Chin, L. T. Survival surgery: removal of the spleen or thymus. Curr. Protoc. Immunol. 2, 10 (1992).
Sitia, G. et al. Kupffer cells hasten resolution of liver immunopathology in mouse models of viral hepatitis. PLoS Pathog. 7, e1002061 (2011).
Andreata, F. et al. Isolation of mouse Kupffer cells for phenotypic and functional studies. STAR Protoc. 2, 100831 (2021).
McCarthy, D. J., Campbell, K. R., Lun, A. T. L. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Germain, P. L., Lun, A., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res 10, 979 (2021).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Benayoun, B. A. et al. Remodeling of epigenome and transcriptome landscapes with aging in mice reveals widespread induction of inflammatory responses. Genome Res. 29, 697–709 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Kiner, E. et al. Gut CD4+ T cell phenotypes are a continuum molded by microbes, not by TH archetypes. Nat. Immunol. 22, 216–228 (2021).
Marsh, S. scCustomize: custom visualizations & functions for streamlined analyses of single cell sequencing. R package version 2.1.2. https://samuel-marsh.github.io/scCustomize/ (2024).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Tan, A. T. et al. Host ethnicity and virus genotype shape the hepatitis B virus-specific T-cell repertoire. J. Virol. 82, 10986–10997 (2008).
Acknowledgements
We thank A. Fiocchi, S. Grassi, M. Mainetti and S. D. Palma for technical support; M. Tinelli, S. Cristiano and M. Silva for secretarial assistance; and the members of the Iannacone laboratory for helpful discussions. We thank M. D. Robek (Albany Medical College) for providing rVSVEnv, and B. Steege (TCR cloning) and T. Holdt (TCR screening ELISPOT) for technical support. We thank F. Chisari (The Scripps Research Institute) for sharing several transgenic mouse lineages including 1.3.32, 107-5, MUP-core, BC10.3 and 6C2.36. We acknowledge S. F. Wieland (University of Basel) and F. Chisari for providing the slides from the livers of HBV-infected chimpanzees that were generated thanks to the scientific collaboration between F. Chisari and R. H. Purcell (NIH) and described in Asabe et al.9.
Confocal immunofluorescence histology was carried out at Alembic, an advanced microscopy laboratory established by the San Raffaele Scientific Institute and the Vita-Salute San Raffaele University. Part of flow cytometry analyses and single-cell sorting was carried out at FRACTAL, a flow cytometry resource and advanced cytometry technical applications laboratory established by the San Raffaele Scientific Institute. We acknowledge the PhD program in Basic and Applied Immunology and Oncology at Vita-Salute San Raffaele University, as V.V. conducted this study as partial fulfilment of a PhD in Molecular Medicine within that program.
This work was supported by: an EMBO fellowship (ALTF180-2020) to C.G.B.; Italian Ministry of Health Grant GR-2019-12368956 to F.M.; European Research Council (ERC) Synergy Grant 101118936, Italian Ministry for University and Research Grants (Project no. PE00000007 INF-ACT) and Fondazione Prossimo Mio to V.F.; Fondazione Prossimo Mio to C.L.; Giovanni Armenise Harvard Foundation Career Development Award, the Italian Association for Cancer Research (AIRC) Start-Up Grant 27564, the ERC Starting Grant 101116224 to M.D.G.; Italian Ministry of University and Research grants PRIN-P2022YKH9R, PRIN-20209Y5YFZ and PE00000007 (INF-ACT), and by the Italian Ministry of Health (MOH) Grant GR-2021-12372615 to M.K.; MOH Singapore Translational Research (STAR) Investigator Award MOH-001633 (MOH-STaR24jan-0004) to A.B., N.L.B. and S.O.; Italian Ministry for University and Research (grant PE00000007 INF-ACT, grant PRIN 20224NMLXK and grant PRIN P2022Z8HNC), AIRC grant 22737 and donations from Fondazione SAME and Fondazione Prossimo Mio to L.G.G.; ERC Advanced Grant 101141363, ERC Proof of Concept Grant 101138728, Italian Association for Cancer Research (AIRC) Grants 19891 and 22737, Italian Ministry for University and Research Grants PE00000007 (INF-ACT) and PRIN 2022FMESXL to M.I.
Author information
Authors and Affiliations
Contributions
Conceptualization: V.V., C.G.B. and M.I.; Investigation: V.V., C.G.B., C.P., P.D., E.B., L.G., F.M., V.F., M.G., C.L., P.D.L., K.R., J.P., S.O., A.C., K.K. and M.R.; Formal analysis: V.V., C.G.B., P.D., F.M., C.L., N.L.B., A.B. and F.V.; Visualization: V.V.; Writing—first draft: V.V. and C.G.B.; Writing—final version: M.I., with input from all authors; Funding acquisition: M.I., L.G.G., M.K., D.I. and M.D.G.; Methodology: T.A.O., M.G., G.C., P.T.F.K., F.P., M.P. and U.S.; Supervision: M.I.; Project administration: M.I.
Corresponding author
Ethics declarations
Competing interests
K.R. and T.A.O. are stock owners, and employees at BioNTech. U.S. is stock owner and management board member at BioNTech. P.T.F.K. has acted as a consultant/advisor for Aligos, Assembly Biosciences, Gilead Sciences, GlaxoSmithKline and Bluejay and received educational grants from Aligos, Gilead Sciences and Vir Biotechnology. N.L.B. and A.B. are the founders and hold stocks of T Cell Diagnostics, a biotech company developing T cell tests for viral infection. A.B. is the cofounder of LION TCR, a company developing T cell therapy against viral-related cancers. A.B. consulted and served on the advisory boards of Brii Biosciences, Assembly Biosciences and Gilead Sciences. L.G.G. participated in boards, advisory boards and consultantships for Genenta Science, Epsilen Bio, Gilead Sciences, Antios Therapeutics, Aligos Therapeutics, Medicxi, Chroma Medicine and Ananda Immunotherapies. M.I. participates in advisory boards/consultantships or receives funding from Gilead Sciences, GentiBio, BlueJay Therapeutics, BioNTech, Excision BioTherapeutics, Moderna, GSK and Curie.Bio. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Robert Thimme and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation of TCR transgenic Env126 mice.
(a) Experimental design: WT mice were immunized with Env126-138 peptide and polyI:C at three time points. At day 21 post immunization, splenocytes were isolated and restimulated ex vivo o/n with the cognate peptide before cell sorting. (b) Sorting strategy and frequency of IFN-ɣ producing CD4+ T cells after 4 h and o/n in vitro peptide restimulation and subsequent IFN-ɣ+ cell enrichment prior sorting. As a control, isolated splenocytes were cultured with an irrelevant Ag (CtrlAg, OVAII) and Concavalin A (ConA). (c) RNA encoding TCR α and β chains were electroporated into activated WT CD4+ T cells (2×105 cells) and tested with Env126-138 peptide via IFNγ ELISpot in the presence of peptide pulsed bone marrow-derived cells (BMDCs, 1×104 cells). OVAII peptide was used as control (Ctrl-Ag). (d) Transgenic constructs maps generated after cloning of the Env126-138-specific TCR alpha and beta chain sequences into cassette plasmids21. (Upper) Plasmid cassette (pTalphaCass) for TCRα gene expression comprises the Vα (variable alpha region) sequence of the specific Env126-138 TCR (green), the the α constant region (blue) downstream enhancers components (black). (Lower) Plasmid cassette (pTbetaCass) for TCR β gene expression comprises the Vβ (variable beta region) sequence of the specific Env126-138 TCR (orange), the β constant region (blue) downstream enhancers components (black). (e) MFIs of indicated markers on splenic WT or Env126 CD4+ T cells. (f) MFIs of CD44 and Ki-67 on ex vivo 4 h restimulated Env126 CD4+ T cells or WT CD4+ T cells in indicated conditions. WT unstimulated CD4+ T cells were included as a control (Ctrl). (g) MFIs of indicated markers on splenic polyclonal WT or Env126 CD4+ T cells after 48 h in vitro stimulation with indicated stimuli. WT unstimulated CD4+ T cells were included as a control (Ctrl). In (e-g) CD4+ T cells were identified as Live, CD45+, B220−CD19−, CD3+, CD8−CD4+. Env126 CD4+ T cells were identified as Live, CD45+, B220−CD19−, CD3+, CD8−CD4+Env-Tet+. (e-g) n = 3 (WT), 5 (Env126). Results are expressed as mean ± SEM and are representative of at least 3 independent experiments. P values were calculated with two-way ANOVA with Bonferroni post-test. ** P<0.01, *** P<0.001.
Extended Data Fig. 2 In vivo generation of effector Env126 CD4+ T cells.
(a) Experimental design. Rag1−/− recipient mice were injected intravenously with 105 naïve Env126 CD4+ T cells. After 24 h, a group of Rag1−/− recipient mice was infected with rVSVEnv. Spleens were surgically collected and analyzed 7 days post infection (p.i.) by flow cytometry. (b) Absolute numbers of Env126 T cells in the spleen of indicated groups of mice at day 7 p.i. (c) Radar plot of the normalized MFIs of indicated markers on Env126 T cells. Polyclonal CD4+ T cells from untreated WT mice were included as a control (PBS). Asterisks refer to the results obtained from the statistical analysis of the comparison between Env126 and Env126 + rVSVEnv experimental groups. (d) Histograms of the MFIs represented in (b) calculated on Env126 T cells or WT CD4+ T cells as a control (PBS). (e) Frequency of Env126 T cells or control WT CD4+ T cells expressing the indicated marker o producing the indicated cytokine after 4 h in vitro peptide (Env126-138) restimulation. (f) Representative flow cytometry plots showing the frequency of Env126 CD4+ T cells (yellow) producing IFN-ɣ and TNF-α (left plot), expressing T-bet and producing IL-2 (right plot) after 4 h in vitro peptide (Env126-138) restimulation. Each plot is compared with the same frequencies in WT CD4+ T cells as a control (black). Env126 CD4+ T cells were defined as Live, CD45+, B220−CD19−, CD8−CD4+, CD45.1+CD45.2+; control CD4+ T cells from untreated WT control mice were identified as Live, CD45+, B220−CD19−, CD8−CD4+. (a-f) n = 3 (PBS), 3 (Env126), 5 (Env126 + rVSVEnv) (b, d, e) Data are representative of at least 6 independent experiments. Data are expressed as mean ± SEM. P values were calculated with two-way ANOVA with Bonferroni post-test. * P<0.05, ** P< 0.01, *** P< 0.001, **** P<0.0001.
Extended Data Fig. 3 Single-cell RNA sequencing of effector Env126 CD4+ T cells.
(a) Top panel: UMAP projection of 8894 CD4+ T Env126 cells coloured by the condition: Env126 CD4+ TEFF (yellow), Env126 CD4+ TN (blue); bottom panel: UMAP visualization of 8894 Env126 CD4+ T cells colored by unsupervised Louvain clustering. (b) Heatmap of scaled and clusters-averaged expression values from the three unsupervised-identified clusters displayed in (a) across a curated TH1 set of genes. (c) Feature plot representation of the normalized expression level of selected genes. (d) Violin plot representation of CD4+ T helper signatures extracted from Kiner et al. 2021 showing the enrichment of the naïve, T helper 1, T helper 2, T helper 17, T follicular helper signatures. (e) Barplot displaying the normalized enrichment score (NES) and adjusted pvalue of the GO biological processes related to the signatures displayed in (d) enriched in cluster 1 + cluster 2. (f) GSEA plot displaying the results of the enrichment of the most significant gene set from MSigDB M5 (MSigD B 2024.1 release) of the biological processes related to the T helper 1 signature displayed in (d), in the list of genes pre-ranked based on the comparison between cluster 1 + cluster 2 (positive log2FC) versus cluster 0 (negative log2FC). (g) Violin plot representation of Mki67 gene expression stratified by Env126 CD4+ T cells clusters defined in (a). (h) Feature plot representation of the normalized expression level of Mki67 gene. The central line within the box represents the median of the data, while the lower and upper edges of the box correspond to the first (Q1) and third (Q3) quartiles, respectively, thus defining the interquartile range (IQR). The whiskers extend to the most extreme values that fall within 1.5 times the IQR from the quartiles, and any observations beyond this range are plotted individually as outliers (d, g). p.adjust represents FDR adjusted values calculated using two-tailed Benjamini–Hochberg procedure (e, f). In (d, g) no statistical analysis was performed. n = 5 (Env126 TN), 5 (Env126 TEFF). Data shown are representative of one experiment.
Extended Data Fig. 4 CD4+ T cells prevent and revert CD8+ T cell dysfunction.
(a) MFIs of indicated markers (Fig. 2I) on Env28 T cells or control CD8+ T cells. (b) MFIs of indicated markers (of Fig. 2I) on Env28 T cells in indicated conditions or control CD8+ T cells. (c) Experimental design: HBV-sp CD8+ TN cells were transferred in HBV Tg mice transferred 7 days later with Env126 CD4+ TEFF. Single transfer of CD8+ T cells (CD8) and untreated HBV Tg mice (PBS) were used as controls. (d) sALT at indicated time points post CD4+ TEFF transfer. (e) IHL in indicated groups at day 5 post CD4+ TEFF transfer. (f) Frequency and numbers of hepatic HBV-specific CD8+ T cells at day 5 post Env126 CD4+ TEFF transfer. (g) Numbers of IFN-γ producing hepatic HBV-specific CD8+ T cells t day 5 post Env126 CD4+ TEFF transfer. (h) Env28 T cells frequency in spleen and liver draining lymph nodes at day 5 post cell transfer (refers to Fig. 2A). (i) Experimental design: Env126 CD4+ TEFF were transferred in 1:1 ratio (2,5×105:2,5×105 or 5×104: 5×104) with naïve HBV-specific CD8+ T cells in HBV Tg mice (Co-T). Single transfer of CD8+ T cells (CD8) and untreated HBV Tg mice (PBS) were used as controls. (j) Hepatic HBV-specific CD8+ T cells numbers at day 5 post transfer. (k) Numbers of IFNγ producing hepatic HBV-specific CD8+ T cells at day 5 post transfer. (l) Grzm-B producing hepatic HBV-specific CD8+ T cells numbers at day 5 post transfer. (m) Representative flow cytometry plots showing the frequency of Sca-1+ HBV-specific CD8+ T cells at day 5 post transfer. (n) Sca-1 MFI on HBV-specific CD8+ T cells or control CD8+ T cells. (a-b, h) n = 3 (PBS), 4 (Env28), 5 (Co-T). Data are representative of at least 10 independent experiments. (c-g) n = 3 (PBS), 4 (CD8), 6 (Co-T). (j-n) n = 2 (PBS), 2 (CD8 5×104), 4 (Co-T 5×104), 3 (CD8 2,5×105), 3 (Co-T 2,5×105). Data are representative of at least 3 independent experiments. Data are expressed as mean ± SEM. P values were calculated with two-way ANOVA with Bonferroni post-test. * P<0.05, ** P<0.01, *** P<0.001.
Extended Data Fig. 5 Effector CD4+ T cells do not mediate liver immunopathology.
(a) Experimental design: Env126 CD4+ TEFF were generated as described in Extended Data Fig. 2 and transferred in HBV Tg mice (Env126, yellow dots). As a control, untreated HBV Tg mice (PBS, grey) were included in the experiment. Livers were collected 5 days post cell transfer. (b) sALT in indicated groups of mice at indicated time points. (c) Numbers of IHL isolated from indicated groups of mice at day 5 post cell transfer. (d) Frequency and absolute numbers of endogenous CD8+ T cells in HBV Tg mice in indicated conditions at day 5 post Env126 CD4+ TEFF transfer. (e) Immunohistochemical representative micrographs of liver sections from the indicated groups of mice at day 5 post cell transfer in HBV Tg mice. Micrographs are showing the H&E, cytokeratin-7 (bile ducts) and Ki-67 and cleaved-Caspase3 staining of liver sections. Scales bars represent 100μm. (f) Representative HBcAg IHC micrographs of liver sections from indicated groups of mice at day 5 post Env126 CD4+ TEFF transfer. Scales bars represent 100μm. The staining is presented as a qualitative assessment of antigen distribution, illustrating its presence or absence across conditions. (g) HBV DNA quantification by Southern blot in livers of indicated mice. Bands corresponding to integrated transgene (TG), relaxed circular (RC), double-stranded linear (DS), and single-stranded (SS) HBV DNA are indicated by arrows. n = 3 (PBS), 3 (Env126). Data are representative of at least 3 independent experiments.
Extended Data Fig. 6 Effector CD4+ T cells help is heterologous.
(a) Frequency of IFNγ, TNF and granzyme-B producing hepatic Cor93 CD8+ T cells in indicated groups of mice at day 5 post cell transfer after in vitro peptide restimulation. (b) Representative flow cytometry plots showing the frequency of IFNγ producing hepatic Cor93 CD8+ T cells in indicated groups of mice at day 5 post cell transfer after in vitro peptide restimulation. Endogenous CD8+ T cells from untreated HBV Tg mice were included as a control (PBS, black plot). (c) Absolute numbers of TNF and Grzm-B producing hepatic Cor93 CD8+ T cells in indicated conditions at day 5 post cell transfer after in vitro peptide restimulation. (d) Radar plot of the mean of normalized MFIs of indicated markers on hepatic Cor93 CD8+ T cells. Polyclonal CD8+ T cells from untreated HBV Tg mice were included as a control (PBS). Asterisks refer to the results obtained from the statistical analysis of the comparison between Cor93 and Co-T experimental conditions. (e) Representative HBcAg IHC micrographs of liver sections from indicated groups of mice at day 5 post cell transfer. Scales bars represent 100μm. The staining is presented as a qualitative assessment of antigen distribution, illustrating its presence or absence across conditions. (f) Representative CD3 IHC micrographs of liver sections from indicated groups of mice at day 5 post cell transfer. Scales bars represent 100μm. n = 3 (PBS), 4 (Cor93), 5 (Co-T). Data are representative of at least 4 independent experiments. Data are expressed as mean ± SEM. Data were analysed using two-tailed t-test or one-way ANOVA with Bonferroni post-test. * P<0.05, ** P<0.01, *** P<0.001.
Extended Data Fig. 7 Effector CD4+ T cells help CD8+ T cells outside SLOs.
(a) Representative flow cytometry plots showing frequency of Cor93 T cells in the liver draining lymph nodes 24 h post transfer. (b) Immunohistochemical representative micrographs of liver sections at day 5 post cell transfer. Micrographs show cleaved-Caspase3 staining. Scales bars represent 100μm. (c) Experimental design: Env126 CD4+ TEFF were transferred 1:1 (106:106) with naïve HBV-specific CD8+ T cells in HBV Tg mice splenectomized and treated with anti-CD62L antibody (Co-T) or treated with FTY720 (Co-T FTY). Single transfer of HBV-specific CD8+ T cells in mice splenectomized and treated with anti-CD62L (CD8) or treated with FTY (Cor93 FTY) were included in the experiment. Untreated HBV Tg mice (PBS) were used as a control. (d) Total lymphocytes count in peripheral blood at day 5 post cell transfer. (e) Numbers of hepatic Cor93 T cells at day 5 post transfer. (f) Numbers of hepatic Env126 CD4+ TEFF at day 5 post cell transfer. (g) Numbers of IFNγ producing hepatic HBV-specific CD8+ T cells at day 5 post cell transfer. (h) MFIs of indicated markers on Cor93 T cells or control CD8+ T cells (PBS). (i) Representative flow cytometry plots showing the gating strategy used to identify KCs and DCs among liver non parenchymal cells. Lin comprises CD3+NK1.1+CD19+Ly6G+CD49b+ cells. (j) Representative histogram Fold Change values (FC) of MFIs values on KCs in the liver of co-transferred (Co-T) mice versus Env28 transferred mice (Env28); refers to Fig. 4n. (k) MFIs of indicated markers on KCs and DCs at day 5 post cell transfer (refers heatmap in Fig. 4n). (a, b) n = 4 (Cor93), 4 (Ctr Co-T), 4 (-SLOs Co-T). Data are representative of at least 4 independent experiments. (c-h) n = 3 (PBS), 3 (Cor93), 4 (Co-T), 3 (Cor93 FTY), 4 (Co-T FTY) (k) n = 3 (PBS), 4 (Cor93), 4 (Ctr Co-T). Data are representative of at least 2 independent experiments. Data are expressed as mean ± SEM. Data were analyzed using one-way ANOVA with Bonferroni post-test or two-tailed t-test. * P<0.05, ** P<0.01.
Extended Data Fig. 8 DCs are dispensable for intrahepatic cooperation of CD4+ CD8+ T cells.
(a) Total chimerism on CD45+ cells in the blood of HBV Tg mice transplanted with CD11c-iDTR-GFP bone marrow at indicated time points (b) Frequency of GFP+ DCs cells out of total DCs in the blood of HBV Tg mice transplanted with CD11c-iDTR-GFP bone marrow at indicated time points. (c) Frequency of DCs in the liver of indicated groups of mice at day 5 post cell transfer. (d) Representative flow cytometry plots showing the frequency of DCs in the liver of indicated groups of mice at day 5 post cell transfer. Color scale indicate GFP MFI. (e) Frequency and absolute numbers of KCs in the liver of indicated groups of mice at day 5 post cell transfer. (f) Absolute numbers of granzyme-B+ Cor93 CD8+ T cells in the liver of indicated groups of mice at day 5 post cell transfer. (g) Representative histograms showing the MFIs of indicated markers on Cor93 CD8+ T cells in the liver of indicated groups of mice at day 5 post cell transfer. (h) Representative HBcAg IHC micrographs of liver sections from indicated groups of mice at day 5 post cell transfer. The staining is presented as a qualitative assessment of antigen distribution, illustrating its presence or absence across conditions. (i) Frequency of Env126 CD4+ TEFF in the livers of indicated groups of mice at day 5 post cell transfer. Scales bars represent 100μm. (j) Representative flow cytometry plots showing the frequency of IFN-g producing Env126 CD4+ TEFF in in the livers of indicated groups of mice at day 5 post cell transfer and after in vitro peptide restimulation. (k) Representative histograms showing the MFIs of indicated markers in Env126 CD4+ TEFF in the livers of indicated groups of mice at day 5 post cell transfer. n = 3 (PBS), 3 (Cor93), 4 (+DCs Co-T), 5 (-DCs Co-T). Data are representative of at least 2 independent experiments. (c, f) Data are expressed as mean ± SEM. Data were analyzed using one-way ANOVA with Bonferroni post-test. * P<0.05, ** P<0.01, **** P<0.0001.
Extended Data Fig. 9 KCs depletion impairs intrahepatic T cell cooperation.
(a) Frequency of KCs in the liver at day 5 post T cell transfer. (b) Representative flow cytometry plots showing the frequency of hepatic DCs at day 5 post T cell transfer. Colour scale indicate YFP MFI. (c) Frequency of hepatic DCs at day 5 post T cell transfer. (d) Frequency of hepatic resident macrophages (rMΦ, identified as F4/80+TIM4−) in indicated groups of mice at day 5 post cell transfer. (e) Frequency of macrophages in the spleen of indicated groups of mice at day 5 post cell transfer. (f) HBV DNA quantification by Southern blot in livers of indicated groups of mice at day 5 post cell transfer. Bands corresponding to integrated transgene (TG), relaxed circular (RC), double-stranded linear (DS), and single-stranded (SS) HBV DNA are indicated by arrows. (g) Experimental setup: Env126 CD4+ TEFF were transferred 1:1 (106:106) with naïve Cor93 T cells in HBV Tg mice (Ctr Co-T) and in HBV Tg mice treated with a single dose of clodronate liposomes (CLL) 48 h before cell transfer (CLL Co-T). Single transfer of CD8+ T cells (Cor93) and untreated HBV Tg mice (PBS) were used as controls. (h) Representative flow cytometry plots showing the frequency of KCs at day 5 post T cell transfer. (i) Frequency of KCs at day 5 post cell transfer. (j) Frequency of hepatic resident macrophages and DCs in indicated groups of mice at day 5 post T cell transfer. (k) IHL numbers in indicated groups of mice at day 5 post cell transfer. (l) Absolute numbers of hepatic Cor93 T cells and IFN-g+ Cor93 T cells at day 5 post cell transfer. (a-f) n = 3 (PBS), 3 (Cor93), 5 (+KCs Co-T), 6 (-KCs Co-T). Data are representative of at least 3 independent experiments. (g-l) n = 3 (PBS), 3 (Cor93), 4 (Ctr Co-T), 4 (CLL Co-T). Data are representative of at least 2 independent experiments. Data are expressed as mean ± SEM and analysed using two-tailed t-test or one-way ANOVA with Bonferroni post-test. * P<0.05, ** P<0.01, *** P<0.001, **** P<0.0001.
Extended Data Fig. 10 IL-27 ameliorate HBV-specific CD8+ T cell function with minimal toxicity and off-target effects.
(a-b) MFIs of indicated markers on hepatic Cor93 T cells at day 5 post cell transfer. (c) Experimental design: Cor93 naïve CD8+ T cells were transferred in HBV Tg mice. After 5 days, Cor93 CD8+ T cells were isolated from the liver and put in culture in the presence or absence of the cognate peptide and/or rIL-27. Analyses were performed 48 h after in vitro culture. (d) Representative flow cytometry plots showing the frequency of Sca-1+ and Granzyme-B producing Cor93 T cells after 48 h of in vitro culture. (e) Experimental design: HBV Tg mice were treated with rIL-27 (rIL-27) and compared with untreated HBV Tg mice (PBS). (f) sALT at indicated time points. (g) IHL numbers at day 5 post cell transfer. (h) Frequency of hepatic CD8+ T, CD4+ T and NK cells 5 days after treatment. (i) Numbers of hepatic CD8+ T, CD4+ T and NK cells in indicated groups of mice 5 days after treatment. (j) Frequency of Ki-67+ CD8+ T cells and Ki-67 MFI on hepatic CD8+ T cells 5 days post treatment. (k) Frequency of Sca-1+ CD8+ T cells and Sca-1 MFI on hepatic CD8+ T cells 5 days post treatment. (l) Frequency of Grzm-B producing CD8+ T and NK cells 5 days after treatment. (m) Representative MFI histograms of indicated markers on CD8+ T cells 5 days after treatment. (a-b) n = 3 (Ctr), 3 (PBS), 3 (rIL-27). Data are representative of at least 3 independent experiments. (c-e) n = 2 (WT), 5 (Cor93). Data are representative of at least 2 independent experiments. Each in vitro culture condition was performed in triplicate. (f-m) n = 3 (PBS), 3 (rIL-27). Data are representative of at least 2 independent experiments. CD8+ T cells were identified as Live, CD45+, CD3+, CD8+CD4−. CD4+ T cells were identified as Live, CD45+, CD3+, CD4+CD8−. NK cells were identified as Live, CD45+, CD3−, NK1.1+NKp46+. All data are expressed as mean ± SEM and were analysed using two-tailed t-test or one-way ANOVA with Bonferroni post-test. * P<0.05, ** P<0.01, *** P<0.001.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–5 and Supplementary Table 2
Supplementary Table 1 (download XLSX )
Differentially expressed genes from scRNA-seq of Env126 Teff cells. Table of differentially expressed genes obtained from the comparison cluster 1 + cluster 2 versus cluster 0. Results are from the FindMarkers function in Seurat (related to Extended Data Fig. 3).
Supplementary Video 1 (download MP4 )
Multiphoton intravital microscopy in the livers of HBV Tg mice 24 h after the transfer of Env126 CD4+ Teff cells and HBV-specific CD8+ T cells. Intravital multiphoton microscopy image sequence showing DsRed+/+ Env28 CD8+ T cells (red), EGFP+/+ Env126 CD4+ Teff cells (green) and KCs (white) at 24 h after T cell transfer. Arrowheads point to Env28 CD8+ T cells that stably interact with KCs (red) and to Env126 CD4+ Teff cells that establish similarly stable interactions while migrating between KCs (white). Time is shown in minutes and seconds. Scale bar, 5 µm.
Source data
Source Data Fig. 2 (download TIF )
Unprocessed Southern blot gel.
Source Data Fig. 3 and Extended Data Fig. 5 (download TIF )
Unprocessed Southern blot gel.
Source Data Extended Data Fig. 9 (download TIF )
Unprocessed Southern blot gel.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Venzin, V., Beccaria, C.G., Perucchini, C. et al. CD4+ T cells license Kupffer cells to reverse CD8+ T cell dysfunction induced by hepatocellular priming. Nat Immunol 26, 1352–1366 (2025). https://doi.org/10.1038/s41590-025-02199-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02199-3
This article is cited by
-
CD209a regulates metabolic dysfunction-associated steatotic liver disease development through macrophage STAT3 signaling pathway
Cellular & Molecular Biology Letters (2025)
-
Role of cccDNA in the Quest for HBV Cure- Implications for HIV-HBV Coinfection
Current HIV/AIDS Reports (2025)