Abstract
Plasmacytoid dendritic cells (pDCs) are major producers of type I/III interferons. As interferons are crucial for antiviral defense, pDCs are assumed to play an essential role in this process; however, robust evidence supporting this dogma is scarce. Genetic or pharmacological manipulations that eliminate pDCs or disrupt their interferon production often affect other cells, confounding interpretation. Here, to overcome this issue, we engineered pDC-less mice that are specifically and constitutively devoid of pDCs by expressing diphtheria toxin under coordinated control of the Siglech and Pacsin1 genes, uniquely coexpressed in pDCs. pDC-less mice mounted protective immunity against systemic infection with mouse cytomegalovirus and showed higher survival and less lung immunopathology to intranasal infection with influenza virus and SARS-CoV-2. Thus, contrary to the prevailing dogma, we revealed that pDCs and their interferons are dispensable or deleterious during several viral infections. pDC-less mice will enable rigorously reassessing the roles of pDCs in health and disease.
Similar content being viewed by others
Main
Vertebrate antiviral immunity depends on type I and III interferons (IFN-I/IIIs), which activate STAT1, STAT2 and IRF9 and the downstream induction of IFN-stimulated genes (ISGs). ISGs reinforce intrinsic immunity in virtually all nucleated cells or promote innate and adaptive antiviral immunity1.
During viral infections, IFN-I/III responses are generally beneficial but can turn deleterious when not properly controlled in time and space, promoting virus-induced immunopathology, immunosuppression or susceptibility to secondary bacterial infections1,2,3. Determining which cell types produce IFN-I/IIIs, how and whether they play beneficial or harmful roles, could allow us to mitigate the detrimental effect specifically and preserve the protective functions of IFN-I/IIIs.
Infected cells mainly produce IFNβ and low amounts of only a few IFNα subtypes, induced upon cytosolic or nuclear sensing of viral genome intermediates through helicases or the cGAS enzyme converging on the STING–IRF3 signaling pathway. This function can be inhibited by cell-intrinsic viral immune-evasion mechanisms1,4. In contrast, plasmacytoid dendritic cells (pDCs) are resistant to viral infection and uniquely produce high amounts of all subtypes of IFN-I/IIIs rapidly upon engulfment and endosomal TLR7/9-dependent sensing of nucleic acids from viral particles or infected cells1,5. Thus, pDCs are thought to be critical for vertebrate antiviral immunity4,6,7,8; however, robust evidence supporting this dogma is scarce5.
In humans, primary immune deficiencies abrogating pDCs ability to produce IFN-I/IIIs, including loss-of-function mutations in the TLR7, MYD88, IRAK4 or IRF7 genes, increase susceptibility to Mycobacterium tuberculosis and a few other pyogenic bacteria but not to viruses except influenza A (IAV) and SARS-CoV-2 (refs. 9,10,11,12,13,14,15,16,17). In contrast, patients with inborn errors in genes necessary for IFN-I/III responses, including IFNAR1/2, STAT1/2 and IRF9, are more broadly susceptible to viral infections18,19. Thus, pDCs but not IFN-I/III responses seem largely redundant for antiviral immunity in humans in the current hygiene and healthcare context.
In mice, the role of pDCs in antiviral immunity has often been extrapolated from the study of animals deficient for Tlr7/9 or Myd88, with contrasting results reported depending on the virus and mouse strains, the dose or route of inoculation and the type of readout. Side-by-side comparisons with mice deficient for IFN-I/III responses have rarely been performed1,20.
Whether in humans or mice, genetic inactivation of TLR7/9, MyD88, IRAK4 or IRF7 affects other cells and biological processes beyond pDC-derived IFN-I/III production. For example, TLR7 and IRF7 are expressed and functional for IFN-I induction in monocytes/macrophages, and MYD88 is required for cellular responses to the interleukin (IL)-1 cytokine family members5.
Most of the approaches previously used to deplete pDCs in vivo in mice harbor off-target effects, preventing drawing robust conclusions on the specific role of pDCs5,21. pDC depletion can be achieved with high specificity and efficacy in hBDCA2-DTR mice, upon diphtheria toxin (DT) administration22; however, DT triggers IFN-I production by hDTR+ pDCs before causing their cell death, which can confound data interpretation during an acute treatment and elicit an interferonopathy upon repeated DT injections23. Therefore, there is still an unmet scientific need to determine rigorously whether and how pDCs modulate host antiviral defense in vivo.
Here, we generated and characterized mutant mice constitutively and specifically devoid of pDCs. These pDC-less mice retained the ability to develop protective antiviral immunity during systemic infection with mouse cytomegalovirus (MCMV), a natural rodent pathogen. They revealed a deleterious role of pDCs during respiratory viral infections with the IAV Scotland H3N2 strain24 or SARS-CoV-2, both of which induce a severe lung immunopathology associated with a local cytokine storm. We demonstrated that IFN-I production by pDCs was deleterious in the IAV infection model. Thus, our study challenges the prevailing dogma whereby pDCs are generally protective in antiviral immunity. pDC-less mice will be a critical tool for future studies aiming at rigorously determining the physiological functions of pDCs.
Results
pDC-less mice are constitutively and specifically devoid of pDCs
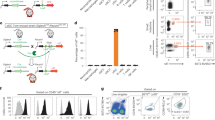
To specifically target pDCs in mice, we recently validated an intersectional genetics approach based on the unique coexpression in mouse pDCs of the Siglech and Pacsin1 genes. We generated reporter pDC-Tom mice, as SiglechiCre × Pacsin1 LoxP-STOP-LoxP-tdTomato animals, and verified that tdTomato fluorescence was exclusively expressed in pDCs25. Here, we adapted this strategy by replacing the tdTomato sequence by the gene encoding the A subunit of diphtheria toxin (DTA), to generate Pacsin1LoxP-STOP-LoxP-DTA mice (Pacsin1LSL-DTA) (Fig. 1a). In SiglechiCre × Pacsin1LSL-DTA mice, the floxed STOP sequence should be removed and DTA lethally expressed only in Siglech+Pacsin1+ cells, namely pDCs. Indeed, pDCs were ablated in both lymphoid and nonlymphoid organs of these mice, hereafter called pDC-less mice (Fig. 1b and Extended Data Fig. 1a). pDC-less mice developed and grew normally (Extended Data Fig. 1b), harbored unaltered cellularity of lymphoid and nonlymphoid organs (Fig. 1c), and normal development of other immune cell types (Fig. 1d,e). We used a staining panel optimized to discriminate transitional dendritic cells (tDCs; CD11b−Ly6D−/lowCX3CR1+ dendritic cells) from pDCs (CD11cloBst2hiLy6DhiCX3CR1neg cells) and conventional dendritic cells (cDCs)25,26,27 (Extended Data Fig. 1c). In pDC-less mice, pDCs were absent, cDC2s unaffected and cDC1s slightly but significantly increased (Fig. 1f). The tDCs expressing low levels of CD11c were slightly reduced, consistent with expression in pDC-Tom mice of tdTomato in late pDC-committed precursors including pro-pDC/tDC25,27. Thus, in pDC-less mice, pDCs are specifically and constitutively absent.
a, Scheme illustrating the strategy used to generate pDC-less mice. LoxP is the sequence recognized by Cre recombinase. ‘STOP’ corresponds to a transcriptional stop sequence. b, Dot plots showing the absence of lin−CD11b−CD11cintBst2hi pDCs in cells isolated from indicated organs of CTR versus pDC-less mice. pDC percentages (mean ± s.e.m.) were pooled from two independent experiments (n = 5 for each strain). c, Absolute numbers of total cells isolated from the indicated organs of CTR versus pDC-less mice (n = 9 for the spleen and lymph nodes (LNs) and n = 5 for the BM and liver). d–f, Percentages within CD45+ splenic cells of indicated lymphoid (d) and myeloid (e,f) lineages isolated from CTR versus pDC-less mice. RPM, red pulp macrophage; cMo, conventional monocyte. The data shown (mean + s.e.m.) are pooled from two independent experiments (n = 9 CTR and pDC-less (d,e) and n = 9 CTR and n = 8 pDC-less (f)). An unpaired and nonparametric two-sided multiple t-test (Mann–Whitney U-test) with Holm–Sidak method for correction was used for the statistical analysis.
During MCMV infection, pDCs are the main IFN-I source but dispensable for intrinsic and innate antiviral immunity
During systemic MCMV infection, IFN-I peak production occurs 1.5 days post infection (p.i.), mainly from pDCs22,28,29,30. Consistently, both IFNα and IFNβ were reduced to undetectable levels in the sera of MCMV-infected pDC-less mice compared to control (CTR) animals, whereas many other proinflammatory cytokines were unaffected (Fig. 2a and Extended Data Fig. 2a). pDC-less mice survived MCMV infection, even at high viral loads (Fig. 2b and Extended Data Fig. 2b) and efficiently controlled viral dissemination (Fig. 2c). The expression of ISGs, such as Mx2, was higher in the spleen than in the lungs and liver, peaking in the spleen and lungs at 1.5 days p.i., while being delayed by 12 h in the liver (Extended Data Fig. 2c). The ISGs Mx2, Ifit2, Irf7 and Isg15 were similarly upregulated in the organs of infected CTR and pDC-less mice (Fig. 2d and Extended Data Fig. 2d), although we observed a trend toward pDC-dependent expression of Irf7 in the spleen and of Mx2 and Isg15 in the liver. In infected Ifnar1KO mice, ISGs were poorly induced in most of the organs studied. In infected pDC-less mice, Il28b induction was reduced in the spleen but not in the liver (Extended Data Fig. 2d), consistent with Il28a/b expression in IFN-I-producing splenic pDCs30. During MCMV infection, the IFNs produced by pDCs can promote the activation of natural killer (NK) cells and cDCs, two innate immune lineages crucial for host defense20,22,31,32,33; however, infection-induced upregulation of Granzyme B, IFNγ and CD69 in NK cells or of CD86 in cDC1s and cDC2s was not affected in pDC-less mice (Fig. 2e and Extended Data Fig. 2e).
CTR and pDC-less mice were infected (intraperitoneally) with MCMV Smith (1 × 105 p.f.u.) and analyzed p.i. a, Cytokine concentrations were analyzed at 0 and 1.5 days p.i. Each dot represents an individual. Data (mean ± s.e.m.) are pooled from two independent experiments (n = 3 for 0, n = 7 for 1.5) and analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction method. b, Survival of CTR and pDC-less mice infected with 1.4 × 105 p.f.u. of MCMV Smith was analyzed in a Kaplan–Meier curve with a log-rank (Mantel–Cox) test (n = 10 for both strains). c, Viral titers were measured by RT–qPCR in the spleen, liver and lungs of CTR and pDC-less mice 3 days p.i. Data are pooled from two independent experiments (spleen n = 9 CTR and 7 pDC-less, liver n = 7 CTR and 6 pDC-less, lungs n = 8 for both strains). Means are indicated by a horizontal bar. An unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction method was used for the statistical analysis. d, Expression of indicated genes was analyzed by RT–qPCR in indicated organs of uninfected or MCMV-infected CTR, pDC-less and Ifnar1KO mice at indicated days p.i. Data were normalized to expression level of Actin gene. The data (mean + s.e.m.) shown are from two independent experiments (n = 4 for each condition and time point). A nonparametric one-way analysis of variance (ANOVA) (Kruskal–Wallis test with Dunn’s correction) was used for the statistical analysis. e, The percentages of Granzyme B (GzmB)+, IFNγ+ and CD69+ splenic NK cells (mean ± s.e.m.) were determined in CTR and pDC-less mice at indicated days p.i. (n = 4 for 0, 8 for 1.5, n = 7 for 2 and n = 11 for 3 days p.i. for both strains, except n = 8 for IFNγ staining 3 p.i.). Data were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction.
Thus, during systemic MCMV infection, despite being the main IFN-I source, pDCs are dispensable for mounting protective intrinsic and innate antiviral immunity.
pDCs and the response to IFN-I are detrimental in respiratory IAV infection
Enhanced susceptibility of IRF7-deficient patients to IAV has been attributed to defective IFN-I production by pDCs12,14. The situation seemed more complex in IAV-infected pDC-depleted mice, with conflicting studies reporting protective, dispensable or detrimental roles of pDCs34,35,36,37,38,39; however, in these studies, other cells were depleted, confounding result interpretation. In addition, both viral strain and mouse genetic background affected host resistance to IAV infection. In contrast with 129S7 mice, upon respiratory infection with most IAV strains C57BL/6 mice did not develop IFN-I-dependent immunopathology, likely due to their low IFN-I production37. Thus, to address the role of pDCs during IAV infection by harnessing our pDC-less mice generated on the C57BL/6 genetic background, we used the H3N2 A/Scotland/20/74 viral strain that induces a severe, Myd88-dependent, pneumonia in CTR C57BL/6 mice24. When infected with a dose of Scotland IAV lethal for 50% of the CTR animals (LD50), all pDC-less mice survived (Fig. 3a and Extended Data Fig. 3a). We extended our analyses to hBDCA2-DTR mice22. Upon DT administration, most of the infected hBDCA2-DTR mice survived, whereas 80% of the CTR mice died (Fig. 3b and Extended Data Fig. 3b). Thus, the use of two distinct mutant mouse strains allowing the specific depletion of pDCs consistently demonstrated that pDCs are deleterious during respiratory infection with Scotland IAV. The loss of pDCs affected neither the control of IAV replication, as measured by IAV M1 expression, nor the induction of ISGs (Irf7, Isg15 and Rsad2), in the lungs (Fig. 3c–e) or brain (Supplementary Fig. 1a,b). Brain inflammation was similarly mild in CTR and pDC-less mice (Supplementary Fig. 1c–g). These results excluded a key role of pDCs in intrinsic antiviral immunity and in the neuropathy that can be induced during respiratory IAV infection.
Mice were infected with 145 p.f.u. (intranasally; i.n.) of IAV strain H3N2A/Scotland/20/74. a, Survival of indicated mice. b, hBDCA2-DTR mice were inoculated with 15 ng g−1 of DT 2 days after IAV infection. n = 10 in a, n = 9 for both strains in b. Kaplan–Meier survival plots show the percentages of survival. c–e, Expression of IAV M1 gene (c) or of indicated ISGs (e) was analyzed by RT–qPCR in lungs of uninfected or IAV-infected CTR and pDC-less mice at indicated days p.i. Data were normalized to expression level of Actin gene. The data (mean + s.e.m.) shown are from two independent experiments. c, n = 6 CTR and 5 pDC-less at 0, 3 at 1, 5 at 2, 14 at 3, 8 CTR and 4 pDC-less at 6, 3 CTR and 4 pDC-less at 8 days p.i. d, Normalized expression (log2) of indicated genes based on bulk RNA-seq samples obtained by whole lung samples isolated from CTR and pDC-less mice at day 0 (n = 3 per strain), 3 and 6 (n = 4 per strain) after IAV infection. e, Expression of indicated genes was analyzed by RT–qPCR. n = 4 CTR and 5 pDC-less at 0, 3 per strain at 4 and 4 per strain at 8 days p.i. Data in c–e were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction method. f,g, IAV-infected mice were monitored daily to assess their morbidity and survival. n = 15 CTR, 8 Ifnar1KO, 7 Ifngr1KO, 9 Ifnl2/3KO and 9 Stat1KO mice (f) and n = 10 CTR, 9 Tlr7KO mice (g). Kaplan–Meier survival plot shows the percentages of survival of indicated mouse strains. log-rank (Mantel–Cox) test was used for statistical analysis in a,b,f,g.
C57BL/6 and most other laboratory murine strains harbor a loss-of-function mutation in the Mx1 gene encoding the main restriction factor against IAV. Congenic C57BL/6 mice expressing a functional Mx1 survive high loads of IAV infection36. This was proposed to depend on Mx1 upregulation in lung epithelial cells in response to pDC IFN-I, as viral titers increased in the lungs of IAV-infected Mx1+ mice depleted of Bst2+ cells, including pDCs36. Thus, we tested whether restoring a functional Mx1 in pDC-less mice would reveal a beneficial role of pDCs against respiratory infection with Scotland IAV. We generated Pacsin1LSL-DTA;Mx1+/+ mice and crossed them with SiglechiCre mice to obtain Mx1+ pDC-less mice. We also generated Mx1+;Ifnar1KO animals. CTR B6.A2G-Mx1+ animals, Mx1+ pDC-less and Mx1+;Ifnar1KO mice all survived IAV infection with high viral loads (106 plaque-forming units (p.f.u.)) and did not differ in weight loss (Extended Data Fig. 3c), despite a trend toward increased viral titers in Mx1+;Ifnar1KO mice at day 2 p.i. (Extended Data Fig. 3d). Thus, during Scotland IAV infection, a functional Mx1 allele confers enhanced resistance to C57BL/6 mice irrespective of the presence of pDCs and of IFN-I-responsiveness. We then investigated the role of IFN-I/IIIs and IFNγ in the resistance to Scotland IAV infection of mice on a classical (Mx1-deficient) C57BL/6 genetic background. When infected with the LD50, 90% of the Ifnar1KO and Ifngr1KO mice survived, whereas most of the Stat1KO and Ifnl2/3 KO mice succumbed (Fig. 3f and Extended Data Fig. 3e). Thus, the overall response to IFN-I is deleterious in C57BL/6 mice during respiratory infection with Scotland IAV, whereas the response to IFN-III is protective, consistent with previous observations in 129S7 mice infected with the X31 IAV strain37,38. The administration of synthetic Tlr7 antagonists inhibits IAV-dependent lung immunopathology in X31-infected 129S7 mice, suggesting that the deleterious IFN-Is were produced via Tlr7 triggering39. Consistent with this hypothesis, most of the Tlr7KO C57BL/6 mice infected with Scotland IAV survived (Fig. 3g and Extended Data Fig. 3f).
In conclusion, respiratory infection of C57BL/6 mice with the pathogenic Scotland IAV strain induces a life-threatening pneumonia promoted by pDCs, Tlr7, IFN-I and IFNγ responses, whereas host resistance relies on Stat1 and IFN-III responses.
During IAV infection, lung pDCs increase in numbers and are a major source of IFNα
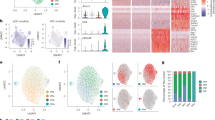
We next investigated whether and when lung pDCs produced IFN-I during IAV infection, by using Ifnb1EYFP reporter mice30,40. pDCs, tDCs and cDCs were rare within lung CD45+ cells in uninfected mice, whereas their percentages increased starting from day 4 p.i. (Fig. 4a and Extended Data Fig. 4a,b). Only pDCs expressed YFP (Extended Data Fig. 4a) with the fraction of positive cells peaking between day 4 and 6 p.i. (Fig. 4b). We then evaluated the impact of pDC loss on lung IFN-Is. First, we analyzed the expression of genes encoding IFN-I/IIIs, in bulk RNA-seq samples from the lungs of CTR versus pDC-less mice, at 0, 3 and 6 days p.i. (Extended Data Fig. 4c). IFN-I/III gene expression was detectable exclusively in infected mice. Ifnb1, Ifna4, Ifnl2 and Ifnl3 were induced to comparable levels in CTR and pDC-less mice, whereas the induction of Ifna1, Ifna2, Ifna6 and Ifna5 was decreased in pDC-less mice (Extended Data Fig. 4c). Starting day 3 p.i., both IFNα and IFNβ proteins were detectable in bronchioalveolar lavages (BAL), with IFNα levels significantly reduced in the absence of pDCs, whereas IFNβ levels were comparable (Fig. 4c). Thus, pDCs are a major source of IFNα in infected lungs, whereas other cells can produce the bulk of IFNβ and IFN-III. Crossing Ifnb1EYFP and pDC-Tom reporter mice generated SCRIPT animals that enable discriminating in tissues IFN+ pDCs (Tom+YFP+) from their IFN− counterparts (Tom+YFP−)25. Heterogeneous CD45 staining distinguished lowly inflamed lung areas (CD45low) from inflammatory foci (CD45high, highly inflamed areas) (Fig. 4d,e,j and Extended Data Fig. 4d,e). Highly inflamed areas contained the highest numbers of infected cells (expressing viral hemagglutinin; HA+) (Fig. 4d,e,k and Extended Data Fig. 4f). Both CD45+ and HA+ cell numbers increased during infection. pDC frequency increased in highly inflamed areas, starting from day 4 p.i. (Fig. 4f,g,l and Extended Data Fig. 4g). IFN+ pDCs were detected only in highly inflamed areas and peaked at day 4 p.i. (Fig. 4h,i,m and Extended Data Fig. 4h). Thus, during respiratory IAV infection, lung pDCs produce IFN-Is locally in highly inflamed and infected areas but at relatively late time points.
a,b, Ifnb1EYFP mice were infected (i.n.) with 160 p.f.u. of IAV and analyzed at different time points p.i. pDC (a) and YFP+ pDC (b) percentages within lung CD45+ cells (data presented as mean ± s.e.m.) isolated from infected Ifnb1EYFP mice were determined. n = 2 at 0, 3 at 2, 6 at 4 and 6, 4 at 8 days p.i. In a,b, a nonparametric one-way ANOVA (Kruskal–Wallis test followed by Dunn’s post hoc test) was used for statistical analysis. c, IFNα and IFNβ concentrations were analyzed in the BAL of CTR and pDC-less mice uninfected (0) or at indicated days p.i. Each dot represents an individual. The data shown (mean ± s.e.m.) are pooled from two independent experiments (n = 3 per strain at 0, 3 CTR and 2 pDC-less at 2, 6 CTR and 7 pDC-less for IFNα and 7 per strain for IFNβ at 3, 6 per strain at 6 and 7 per strain at 8 days p.i.). Data were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction. d–i, Representative images of lung sections of SCRIPT mice at 4 p.i. Sections were stained with anti-CD45 (white), anti-HA IAV (orange), anti-Tomato (Tom; purple) and anti-YFP (green). On each image acquired, CD45 mean fluorescence intensity (MFI) in highly inflamed areas was at least twice that of lowly inflamed areas. j–m, The numbers of CD45+ cells (j), HA (IAV)+ cells (k), pDCs (l) and YFP+ pDCs (m) per mm² were quantified in highly and lowly inflamed areas. The data shown (mean ± s.e.m.) are from five SCRIPT mice at 2, 5 at 3, 10 at 4 and 4 at 6 p.i., with two lung sections analyzed per mouse. An unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction was used for statistical analysis.
IFN production by pDCs is responsible for their deleterious role during IAV infection
We then investigated the specific role of the IFN-I produced by pDCs in IAV-induced pneumonia. None of the existing Cre-driver mice allowed to conditionally inactivate floxed genes specifically in pDCs25,41. To overcome this roadblock, we used pDC-Shield Bone Marrow Chimera (SBMC) mice (Fig. 5a). We lethally irradiated the hind legs of pDC-less mice, protecting the rest of their body with a lead shield. This promoted partial engraftment of donor bone marrow (BM) cells, preferentially reconstituting the immune cell lineages lacking in the recipient mice, in our case pDCs (Fig. 5b). This strategy allows studying the impact of the loss of expression of a candidate gene specifically in pDCs, by using knockout animals as BM donors. As cytokine production by pDCs is Myd88-dependent41, we generated CTR, pDC-less and Myd88KO pDC-SBMC and infected them with IAV. Half of the CTR pDC-SBMC mice succumbed, whereas all pDC-less and most of the Myd88KO pDC-SBMC mice survived (Fig. 5c and Extended Data Fig. 5a). Most of the Tlr7KO pDC-SBMC mice also survived IAV infection (Fig. 5d and Extended Data Fig. 5b). In pDCs, the inactivation of TLR7/Myd88 abrogates the production of most cytokines, whereas IRF7 deficiency abolishes exclusively IFN production41. Thus, we generated Irf7KO pDC-SBMC mice and infected them with IAV. Most of the Irf7KO and pDC-less pDC-SBMC mice survived, whereas 60% of the CTR pDC-SBMC mice died (Fig. 5e and Extended Data Fig. 5c). Compared to CTR pDC-SBMC mice, Irf7KO and pDC-less pDC-SBMC mice displayed reduced morbidity, and lower clinical scores based on their posture, mobility, fur appearance and respiration (Extended Data Fig. 5d), although lung histological analysis did not show any significant difference in lesion scoring (Extended Data Fig. 5e and Supplementary Table 1). CTR, Irf7KO and pDC-less pDC-SBMC displayed similar viral loads (Fig. 5f). Ifna1, Ifna2 and Ifna5 expression was abrogated in IAV-infected Irf7KO and pDC-less pDC-SBMC mice, compared to CTR, whereas Ifna4 and Ifnb1 were similarly induced in the three types of pDC-SBMC mice (Fig. 5f). Consistently, IFNα protein levels, but not IFNβ, were decreased in the BAL of IAV-infected Irf7KO and pDC-less pDC SBMC mice (Fig. 5g), mirroring the observations in pDC-less mice (Fig. 4 and Extended Data Fig. 4).
a, Scheme illustrating the generation of pDC-SBMC mice. Only the hind limbs of pDC-less mice were irradiated, while the rest of the body was protected by a lead shield (gray oval). Upon local irradiation-induced distress, donor BM cells preferentially reconstituted immune cell types absent in the recipient (pDCs). b, Recipient CD45.1+ (gray) pDC-less mice are used as recipients of pDC-SBMC reconstituted with donor CD45.2+ (black) BM cells. The percentage of donor (black) versus recipient (gray) cells was analyzed in indicated lineages 4 weeks after reconstitution (n = 4). c–e, pDC-SBMC mice reconstituted with CTR, pDC-less or Myd88KO BM (c), with CTR or Tlr7KO BM (d) or with CTR, pDC-less or Irf7KO (e) were infected (i.n.) with 145 p.f.u. of IAV strain and monitored daily to assess their morbidity and survival. Kaplan–Meier survival plots show the percentages of survival of indicated genotypes. c,d, Pool of two independent experiments (8 CTR, 7 Myd88KO and 10 pDC-less SBMC in c; 8 CTR and 9 Tlr7KO SBMC in d). e, Pool of three independent experiments (n = 12 for each SBMC type). A log-rank (Mantel–Cox) test was used for statistical analysis in c–e. f, Expression of indicated genes was analyzed by RT–qPCR in lungs isolated from indicated IAV-infected SBMC 3 days after infection (n = 4 for each strain). A nonparametric one-way ANOVA (Kruskal–Wallis test followed by Dunn’s post hoc test) was used for statistical analysis. g, Cytometric bead arrays was performed to quantify IFNα and IFNβ concentrations in the BAL isolated from the same SBMC mice used in f. Data in f,g are presented as mean ± s.e.m. Each dot represents an individual. n = 4 in f,g.
Thus, selective inactivation of Tlr7/Myd88/Irf7 signaling in pDCs improved mouse tolerance to respiratory IAV infection, demonstrating the detrimental contribution of the IFNs produced by lung pDCs during IAV-induced lung immunopathology.
During IAV infection, lung inflammation and cytokine contents are reduced in the absence of pDCs
During IAV infection, the recruitment of immune cells to the lungs and their local activation are essential to establish protective immunity, but unbridled inflammation leads to severe pneumonia and acute respiratory distress syndrome (ARDS). We performed anatomopathological analysis of lung sections from CTR and pDC-less mice at day 8 p.i., when survival and morbidity start to differ between these two strains (Fig. 3a). The lungs from CTR mice showed a marked broncho-interstitial pneumonia (Fig. 6a), associated with bronchiolar filling by degenerated epithelial cells and with intense alveolar inflammation (Fig. 6b), whereas the lungs isolated from pDC-less mice displayed milder pneumonia and inflammation (Fig. 6c,d). Consistently, semi-quantitative anatomopathological scoring of lung sections (Supplementary Table 1) was significantly higher in CTR than in pDC-less mice (Fig. 6e).
a–d, Representative images of lung hematoxylin and eosin-stained sections of CTR (a,b) and pDC-less (c,d) mice 8 days p.i. (a,b) marked broncho-interstitial pneumonia with terminal bronchiole filled with degenerated neutrophils; (c,d) mild broncho-interstitial pneumonia. Magnification is indicated. e, Semiquantitative scoring of lung lesions according to Supplementary Table 1. The data (mean ± s.e.m.) pooled from two independent experiments (n = 9 CTR, 8 pDC-less) were analyzed using a two-tailed nonparametric Mann–Whitney U-test. f, Absolute numbers of whole and CD45+ lung cells of CTR and pDC-less mice during infection. Data (mean ± s.e.m.) are pooled from two independent experiments (n per strain = 5 at 0, 7 at 3, 8 at 6, 7 CTR and 8 pDC-less at 8 days p.i.). g, Percentages of lung myeloid and lymphoid cells during infection. n = 7 per strain, except cMo, n = 4 per strain and eosinophils, n = 14 CTR and 13 pDC-less at 0, 7 per strain at 3, 8 CTR and 7 pDC-less, except eosinophils, n = 17 CTR and 14 pDC-less, at 6, 7 CTR and 8 pDC-less, except pDCs and other dendritic cells, n = 4 per strain. Data in f,g were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction. h, Survival of IAV-infected CTR and pDC-less mice treated with anti-Ly6G (1A8, triangle) or isotype control (IC; circle) antibodies 5 and 6 days p.i. Individuals pooled of two independent experiments were analyzed using a Kaplan–Meier survival plot with a log-rank (Mantel–Cox) test. IC, n = 8 per strain; 1A8, n = 9 per strain. i, Cytokine concentrations in the BAL of CTR and pDC-less mice. Each dot represents an individual. Data shown (mean + s.e.m.) pooled from two experiments were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction. n = 3 CTR and 4 pDC-less at 0, 3 CTR and 2 pDC-less at 2, 7 per strain at 3, 6 per strain at 6 and 7 per strain at 8 days p.i.
In contrast to what had been previously observed in X31-infected 129S7 mice upon depletion of BST2+ cells37, in C57BL/6 mice infected by Scotland IAV the loss of pDCs affected neither the increase in total CD45+ cells nor the proportion of most myeloid and lymphoid lineages in the lungs (Fig. 6f,g and Extended Data Fig. 6a–d), except for a significant increase in neutrophils and eosinophils at days 6–8 p.i. (Fig. 6g). The role of neutrophils during IAV infection is disputed39,42. In our model, neutrophil depletion at days 5 and 6 p.i. severely decreased mouse survival, irrespective of the presence of pDCs (Fig. 6h), supporting their protective role independent from pDC activity. The absence of pDCs only transiently affected the upregulation on lung myeloid and lymphoid cells of IFN-I-dependent activation molecules such as CD86 and CD69 (Extended Data Fig. 7a,b). In X31-infected 129S7 mice, the depletion of BST2+ cells was previously reported to reduce BAL proinflammatory cytokines and lung immunopathology37. In our model, most of the cytokines and chemokines analyzed were similarly upregulated at messenger RNA and protein levels in infected CTR and pDC-less mice at days 3 and 6 p.i. (Fig. 6i and Extended Data Fig. 7c,d). However, at day 8 p.i., CXCL10, CCL2, IFNγ and IL-10, but not TNF, IL-6 and CCL5, were reduced in the BAL of pDC-less mice (Fig. 6i and Extended Data Fig. 7d). This suggests that pDC-less mice are more tolerant to the late infection phases due to reduced production of cytokines promoting ARDS.
The alterations of lung integrity and permeability induced upon IAV infection are less severe in the absence of pDCs
During IAV infection, unbridled inflammation promotes lung lesions, fluid leakage and respiratory defaults leading to ARDS. We assessed the integrity of bronchi by staining lung sections for the epithelial cell adhesion molecule (EpCAM). Whereas this staining pattern was continuous and homogeneous in uninfected mice of both CTR and pDC-less strains (Fig. 7a), it was heterogeneous in infected animals, identifying disrupted (open) bronchi (Fig. 7a, white arrows). In infected pDC-less mice, the percentages of EpCAM+ areas were higher (Fig. 7b) and of disrupted bronchi lower (Fig. 7c), suggesting reduced breaching of the bronchial tree. Thus, we evaluated pulmonary permeability by measuring the leakage of intratracheally-instilled 10KDa FITC–dextran into the plasma43 (Supplementary Fig. 2). While the FITC–dextran signal was undetectable in the plasma of uninfected mice and of PBS-instilled infected mice, it was clearly detectable in FITC-instilled infected mice but with lower levels in pDC-less animals (Fig. 7d), confirming that pDCs promote lung lesions and permeability during IAV infection.
a, Representative images of histological sections of lungs isolated from CTR and pDC-less mice uninfected (left) or at day 8 after IAV infection (right). The 30-µm-thick sections were stained with anti-CD45 (white) and anti-EpCAM (red) antibodies. EpCAM+ bronchi were classified as closed (red arrow) when EpCAM staining was uninterrupted and as opened (white arrow) when EpCAM was discontinuous. b,c, The percentages of the EpCAM+ area over the entire surface of each lobe (b) and the percentages of closed versus opened bronchi (c) were quantified in lung sections isolated from uninfected versus IAV-infected CTR versus pDC-less mice. The data (mean ± s.e.m.) shown in b,c are from n = 1 at day 0 and n = 4 at day 8 after infection for both mouse strains. We analyzed two lobe sections at day 0 and 9 lobe sections at day 8 for CTR mice, three lobe sections at day 0 and eight lobe sections at day 8 for pDC-less mice. An unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak correction method was used for the statistical analysis. d, 8 days p.i., CTR and pDC-less mice were intratracheally (i.t.) treated with fluorescein isothiocyanate (FITC–dextran) (10 µg per mouse). Barrier permeability was measured as relative fluorescent units (RFUs) of FITC-dextran leaked in the plasma 1 h after i.t. FITC–dextran administration. Each dot represents an individual. Data shown (mean + s.e.m.) are pooled from two experiments (n = 2 for each strain and condition for uninfected mice, as well as for PBS-treated infected mice, 9 for FITC-treated infected CTR and 8 for FITC-treated infected pDC-less mice). An unpaired and nonparametric two-sided multiple t-test (Mann–Whitney) with Holm–Sidak method for correction was used for the statistical analysis.
During mouse SARS-CoV-2 infection, pDCs are dispensable or detrimental for host survival
The increased susceptibility to SARS-CoV-2 infection of patients harboring inborn loss-of-functions mutations in MYD88, IRAK4, IRF7 or TLR7 has been hypothesized to result from impaired pDC-dependent viral control13,14,15,16,17; however, other studies proposed a detrimental role of pDCs in human COVID-19 through IFN-I-dependent activation of macrophages fueling a cytokine storm44,45,46,47. To attempt solving this controversy, we harnessed pDC-less mice. C57BL/6 mice typically do not develop detectable infection or pathology when exposed to most SARS-CoV-2 strains that bind only with low affinity to the mouse receptors for viral entry, including angiotensin-converting enzyme 2 (ACE2). Ectopic expression of human ACE2 under the control of the cytokeratin 18 (K18) promoter renders transgenic K18:hACE C57BL/6 mice highly susceptible to SARS-CoV infections48. Thus, we introduced this transgene into pDC-less mice, by generating K18:hACE × Pacsin1LSL-DTA animals and crossing them with SiglechiCre mice. Offspring triple heterozygous animals (K18:hACE pDC-less) and CTR K18:hACE mice were infected with the SARS-CoV-2 Wuhan/D614 strain. Infected K18:hACE males survived less than females, consistent with the higher morbidity and mortality observed in male patients with COVID-19 (ref. 49) (Fig. 8a and Extended Data Fig. 8a). pDC deficiency increased survival in male but not female mice (Fig. 8a and Extended Data Fig. 8a). Thus, pDCs are dispensable for the resistance of K18:hACE female mice to SARS-CoV-2 infection and are detrimental in males.
a–c, K18:hACE transgenic mice bred or not with indicated mutant mice were infected with 200 p.f.u. (i.n.). of SARS-CoV-2 strain. Survival was evaluated using Kaplan–Meier plots analyzed with a log-rank (Mantel–Cox) test. n = 28 female K18:hACE and K18:hACE pDC-less; 19 male K18:hACE, 26 male K18:hACE pDC-less (a), n = 10 K18:hACE and plain Ifnar1KO, 15 K18:hACE Ifnar1KO (b) and n = 12 female K18:hACE, 9 female K18:hACE Tlr7KO; 11 male K18:hACE, 7 male K18:hACE Tlr7KO (c). d, Normalized expression (log2) of the indicated genes based on bulk RNA-seq samples obtained by whole lung samples isolated from transgenic K18:hACE mice infected with 1.1 × 105 p.f.u. of SARS-CoV-2 at the indicated days p.i. Data (mean ± s.e.m.) shown are from n = 3 at 0 and 1, 5 at 2 and 4, 2 at 6 days p.i. e,f, Percentages of pDCs (e) and YFP+ pDCs (f, IFN-producing pDCs) within lung CD45+ cells of K-18:hACE SCRIPT mice infected with 1.1 × 105 p.f.u. of SARS-CoV-2. n = 10 females (F) at 0, 8F and 4 males (M) at 1, 7F at 2 and 9F and 4M at 4 days p.i. g, Expression of indicated genes were analyzed by RT–qPCR in lungs of SARS-CoV-2-infected CTR, pDC-less and Tlr7KO K18:hACE mice during infection. Data were normalized to Actin expression. Data (mean + s.e.m.) are from two independent experiments (7F and 2M CTR, 5F and 2M pDC-less, 4F and 2M Tlr7KO at 0, 5F and 5M CTR, 7F and 5M pDC-less, 6F and 5M Tlr7KO at 2, 6F and 6M for each strain at 4 days p.i.). A nonparametric one-way ANOVA (Kruskal–Wallis test followed by Dunn’s post hoc test) was used in d–f. A two-way ANOVA with Tukey’s multiple comparison test was used in g.
IFN-I and TLR7 responsiveness are essential for human survival to SARS-CoV-2 infection15,16. Consistently, female K18:hACE Ifnar1KO mice were more susceptible to SARS-CoV-2 than K18:hACE CTR mice, whereas regular Ifnar1KO mice lacking hACE2 were resistant as expected (Fig. 8b and Extended Data Fig. 8b). CTR, pDC-less and Ifnar1KO, K18:hACE mice controlled viral infection similarly (Extended Data Fig. 8c), suggesting that their differential survival was primarily due to alterations in immunopathology. TLR7 deficiency significantly increased the susceptibility of K18:hACE mice to SARS-CoV-2 infection, independently of mouse sex (Fig. 8c and Extended Data Fig. 8d). Thus, the survival of K18-hACE mice to SARS-CoV-2 infection requires Tlr7-dependent but pDC-independent induction of IFN-I responses.
The timing and magnitude of IFN production varies between patients with moderate versus severe COVID-19 (ref. 44). In mice infected with SARS-CoV-1/2 or MERS-CoV, IFNs were beneficial very early after infection, but detrimental later50,51,52. In K18-hACE mice, IFN-I and ISG expression was induced in the lungs 1–2 days p.i., concomitantly with SARS-CoV-2 detection (Fig. 8d and Extended Data Fig. 9a,b). ISG induction persisted after the peak of IFN (day 2), was unaffected in K18:pDC-less mice (Extended Data Fig. 9a,b), but decreased to variable extents in K18:hACE Ifnar1KO mice (Extended Data Fig. 9a,b). Analysis of K18:hACE SCRIPT mice showed that lung pDCs increased starting from day 2 p.i. (Fig. 8e and Extended Data Fig. 9c,d) and that IFN-I+ (YFP+) pDCs were detectable at day 2 and increased at day 4 (Fig. 8f). Thus, the peak of IFN production by pDCs seemed delayed compared to ISG induction and persisted for several days, suggesting that it is dispensable for the early reinforcement of intrinsic immunity but may contribute to sustained, deleterious inflammation.
As IFN-I and Tlr7 both promoted the survival of K18:hACE mice to SARS-CoV-2 infection, whereas pDC were dispensable or detrimental, we investigated the impact of their loss on IFN production and response. Viral titers were comparable in all strains, independently of host sex (Extended Data Fig. 9e). In females, Ifna and Ifnb1 expression was induced at day 2 p.i., but barely detectable at day 4 p.i. (Fig. 8g). In males, the expression of Ifn genes was very low and seemed delayed, becoming detectable only in some individuals and at day 4 p.i. In females, the expression of all the Ifn genes analyzed was Tlr7-dependent, whereas only Ifna1 and Ifna5 were also pDC-dependent. In female mice, at day 2 p.i., the expression of ISGs seemed both Tlr7- and pDC-dependent; in males a similar but not significant trend was observed (Isg15, Irf7 and Rsad2; Extended Data Fig. 9f).
In conclusion, during SARS-CoV-2 infection of K18:hACE mice, pDCs mainly contribute to the production of IFNα, whereas IFNβ is mostly induced in other cells in a Tlr7-dependent manner. pDCs are dispensable in females and detrimental in males, whereas both IFN-I and Tlr7 are protective regardless of host sex.
Discussion
Here, we report the generation of a new mouse model, pDC-less mice, allowing specific and constitutive ablation of pDCs. pDC-less mice developed normally and were healthy in an environment free of specific pathogens, suggesting that pDCs are not essential for homeostatic host fitness. Only CD11clow tDCs were reduced in pDC-less mice, slightly, likely due to shared precursors27. Hence, in pDC-less mice, the specificity of pDC depletion is highly improved compared to previous methods that impacted also DC precursors, tDCs and some monocytes/macrophages or B cells21, including depletion with anti-PDCA1/BST2 antibodies5 or use of the CD11c-Cre;Tcf4−/fl or Siglech-hDTR mice27,53,54,55. pDC-less mice allow evaluating the long-term impact of pDC loss, without the DT-dependent confounding side-effects observed in hBDCA2-DTR mice22,23. Moreover, pDC-less mice can be used as recipients for the generation of SBMC allowing to inactivate candidate genes selectively in pDCs, a solution to overcome the lack of pDC-specific Cre mice.
The molecular make-up and professional production of all subtypes of IFN-I/IIIs by pDCs are evolutionarily conserved across vertebrates1,41,56, implying a contribution to their reproductive fitness. The underlying mechanism is considered to be a critical role of pDCs in promoting protective intrinsic and innate antiviral immunity6,7. Yet, robust experimental data supporting this hypothesis are scarce. In contrast to patients who are defective in IFN-I/III-responses, human individuals whose pDCs are unable to produce IFN-I/IIIs, due to loss-of-function mutations in TLR7, MYD88, IRAK-4 or IRF7, do not show increased susceptibility to most viral infections, except for IAV and SARS-CoV-2 (refs. 11,12,13,14,15,16,17,18,19). Even then, the role of pDCs is unclear, since other cell types and biological processes are affected by these mutations5. In mice, pDC depletion or functional inactivation did not compromise host survival to most viral infections5, except for systemic HSV-2 or ocular HSV-1 infection57,58.
Here, we showed that, in mice, during systemic MCMV and respiratory IAV or SARS-CoV-2 infections, although pDCs were a major IFNα source, they were redundant for ISG induction viral control, and preservation of host health. For IAV infection, this was also true in mice expressing a functional allele of the key antiviral restriction factor Mx1, questioning the mechanism behind the increased viral titers observed previously in Mx1+ mice depleted of Bst2+ cells36. Consistently, during MCMV infection, ISG induction and protective antiviral immune responses are much more profoundly compromised by Ifnar1 inactivation than by Myd88 deficiency20. Thus, contrary to the prevailing dogma, we showed that, in mice, pDCs are redundant with other cellular sources of IFN-I/IIIs for enhancing intrinsic immunity and more generally promoting host resistance to primary acute viral infections, similarly to what has been proposed in humans9,10,59. Our findings illustrate how redundancy contributes to the robustness of immune responses20,60, with multiple cells and pathways contributing to the vital production of IFNs61. Most IFNα subtypes, except IFNα4 and IFNβ, were reduced in the lungs of IAV-infected pDC-less mice, consistent with their dependency on IRF7 that is constitutively expressed rather selectively in pDCs, whereas IFNα4/β can be quickly released by infected cells in an IRF3-dependent IRF7-independent manner62,63. Although all IFN-Is bind the same receptor, they differ in binding affinity and kinetics, resulting in distinct patterns and strengths of ISG induction1. Differently from most infected cells, pDCs are highly motile, especially after activation64. Hence, a division of labor may exist among IFN-producing cells. Virally infected cells may rapidly reinforce local intrinsic immunity to control viral propagation, while pDCs could migrate within infected tissues and across organs to deliver IFNs to specific cells and sites at optimal times25,30, contributing to coordinating antiviral immune responses. This pDC response may be generally beneficial but could become harmful during infections triggering IFN-dependent immunopathology or autoimmunity.
Our study extends to respiratory viral infections the large body of experimental evidence of the harmful proinflammatory functions of pDCs in autoimmune or inflammatory diseases5. This contrasts with the scarcity of definite proof that pDCs can exert critical protective functions in the natural history of viral infections5. This raises the question of the mechanisms behind the evolutionary conservation of pDCs. We need to re-evaluate whether, when and how pDC functions could benefit the host. pDC-less mice, and SBMC mice derived thereof, will be key to address this outstanding question in future studies. Several hypotheses can be formulated in the context of viral infections. First, pDCs might be required to protect immune-privileged organs that are otherwise unable to produce IFNs, such as the cornea during ocular HSV-1 infection58. Second, pDCs may be required to induce a body-wide state of antiviral resistance, including in barrier tissues remote from the initial infection site65, to prevent lethal superinfections by other viruses. Third, pDC IFN production during a primary infection could optimize adaptive immune memory generation to improve protection against secondary infections, explaining in part the enhanced resistance of mice exposed to a normalized microbial environment66. Testing some of these hypotheses will require developing a novel mouse model for conditional pDC depletion free of the confounding factors of existing models23,28 and implementing complex experiments involving sequential exposure to different pathogens67.
We showed that pDCs can impair host survival during respiratory IAV or SARS-CoV-2 infections. This seems contradictory to the increased susceptibility to these infections of patients with loss-of-function mutations compromising the TLR7-MYD88-IRF7 signaling pathway and thus pDC IFN-I/III production, and to the decrease in blood pDC numbers and their ability to produce IFN-Is in patients with severe COVID11,16,68,69,70. However, the role of pDCs in these infections is debated and alternative interpretations should be considered. Here, we showed that, in mice during respiratory viral infections, pDCs increased in the lungs and contributed to local IFNα production, but were redundant or deleterious for host health, whereas Tlr7 was protective in SARS-CoV-2 infection. In murine SARS-CoV-2 infection, Tlr7 deficiency reduced IFNα and IFNβ production in murine lungs more broadly than pDC loss. These findings support a scenario where the pDC-derived IFN-Is are dispensable or detrimental, and the protection conferred by TLR7-MYD88-IRF7 operates via other cell types that also express, and respond to, these molecules such as monocytes/macrophages16,71,72. Future studies are required to test this hypothesis, especially to compare the kinetics and spectrum of the TLR7-dependent production of IFN subtypes between pDCs and other cells in vivo during SARS-CoV-2 infection, an important direction since boosting pDC IFN production has been proposed to treat severe human respiratory infections8. Our results caution that this could rather have detrimental consequences. The decrease in circulating pDCs and their functional exhaustion in severe COVID patients could reflect their recruitment and activation in infected tissues where they could fuel harmful inflammation, as reported during macaque or human infections by immunodeficiency viruses73,74,75. Indeed, during severe respiratory viral infections, lung pDCs have been proposed to exacerbate the pathological activation of monocytes/macrophages44,45,46,47. Better understanding whether and how pDC IFN production can worsen disease could allow selective intervention to inhibit the harmful sources of IFN-I/IIIs while preserving their protective sources. This approach would improve therapy, by avoiding the use of broadly immunosuppressive drugs such as corticosteroids or JAK–STAT inhibitors.
Methods
Mice
All animal experiments were performed in accordance with national and international laws for laboratory animal welfare and experimentation (EEC Council Directive 2010/63/EU, September 2010). Protocols were approved by the Marseille Ethical Committee for Animal Experimentation (registered by the Comité National de Réflexion Ethique sur l’Expérimentation Animale under no. 14; APAFIS no. 21626-2019072606014177 v.4, APAFIS no. 35098-2022020215455814 v.4 and APAFIS no. 26484-2020062213431976 v.6). C57BL/6J (B6) mice were purchased from Janvier Laboratories. SiglechiCre (B6-Siglechtm1(iCre)Ciphe), Ifnar1KO (B6.129S2-Ifnar1tm1agt) and SCRIPT (SiglechiCre;Pacsin1LSL-tdT;IfnbEYFP) mice were previously described25,41. Pacsin1LoxP-STOP-LoxP-DTA (B6-Pacsin1tm1(DTA)Ciphe, Pacsin1LSL-DTA) and Stat1KO (B6-Stat1tm1d(EUCOMM)Ciphe) were generated by the Centre d’ImmunoPhénomique (CIPHE). Homozygous Pacsin1LSL-DTA and SiglechiCre were bred together and their double heterozygous SiglechiCre; Pacsin1LSL-DTA progeny were named pDC-less mice. SiglechiCre and Pacsin1LSL-DTA were also bred with CD45.1 congenic mice, then interbred to generate 5.1 pDC-less mice used as recipient for SMBC generation. Tlr7KO (B6-Tlr7tm1Flv) mice were kindly provided by L. Alexopoulou (CIML). B6.A2G-Mx1+/+ congenic mice were generated and provided by M. Le Bert (Typage et Archivage d’Animaux Modèles, TAAM, Orleans, France). Irf7KO (B6-Irf7tm1Ttg) were kindly provided by C. Svanborg (Lund University). Ifnl2/3KO were kindly provided by A. Broggi (Marseille, France). IfnbEYFP (B6.129-Ifnb1tm1Lky), hBDCA2-DTR (C57BL/6-Tg (CLEC4C-HBEGF)956Cln/J), Ifngr1KO (B6.129S7-Ifngr1tm1Agt/J) and heterozygous K18-hACE C57BL/6J mice (strain 2B6. Cg-Tg (K18-ACE2)2Prlmn/J) were obtained from the Jackson Laboratories. Pacsin1LSL-DTA and B6.A2G-Mx1+/+ mice were intercrossed to generate double homozygous mice, then bred with SiglechiCre to obtain as progeny Mx1+ pDC-less mice. CTR mice used in all experiments were Pacsin1LSL-DTA, SiglechiCre or B6 mice. All mouse strains were on a B6 genetic background and bred at CIPHE or the Centre d’Immunologie de Marseille-Luminy (CIML), under specific-pathogen-free conditions and in accordance with animal care and use regulations. Mice were housed under a 12-h dark–light cycle, with a temperature range of 20–22 °C and a humidity range of 40–70%. All animals used were sex and age matched (8–12 weeks of age for all experiments, except for SBMC). C57BL/6, coming from an external provider, as well as SiglechiCre and Pacsin1LSL-DTA mice, bred in the same mouse house as pDC-less mice, gave similar readouts and were indistinctly used as CTR in experiments with MCMV or IAV.
SMBC generation
The 5-week-old CD45.1+ pDC-less mice were anesthetized, then their hind legs were 9 Gy irradiated, while the rest of the body was protected with a lead shield. At the end of the irradiation, mice were intravenously injected with 15 × 106 cells of BM isolated from indicated CD45.2+ donor mice. At 4 weeks after BM engraft, mice were bled to test the reconstitution with donor BM cells, then used for experimentation.
Viruses, viral infections and mice treatment
MCMV Smith stocks were prepared from salivary gland extracts of 3-week-old MCMV-infected BALB/c mice. Mice were infected intraperitoneally with 105 p.f.u. and killed at the indicated time points. For survival experiments with MCMV, mice were infected with 1.4 × 105 p.f.u. The H3N2 A/Scotland/20/74 IAV strain was produced and titrated in vitro using the Madin–Darby Canine Kidney (MDCK) cell line. Mice were anesthetized with ketamine/xylazine, then intranasally infected with 40 µl DMEM containing 160 p.f.u. IAV strain and killed at the indicated time points. For survival studies, mice were infected with 145 p.f.u. of the same IAV strain. The β CoV/France/IDF0372/2020 SARS-CoV-2 strain was supplied by the National Reference Centre for Respiratory Viruses hosted by the Institut Pasteur (Paris, France). This strain was isolated from a human sample provided by the Bichat Hospital, Paris, France and corresponded to the original Wuhan/D614 SARS-CoV-2 strain detected at the beginning of the COVID-19 pandemic. Infectious stocks were grown by inoculating Vero E6 cells and collecting supernatants upon observation of the cytopathic effect. Debris was removed by centrifugation and passage through a 0.22-mm filter. Supernatants were stored at −80 °C. Mice were anesthetized with 150 μl of ketamine/xylazine, then intranasally infected with 30 µl of medium containing 200 p.f.u. of β CoV/France/IDF0372/2020 SARS-CoV-2. For bulk RNA-seq studies mice were infected with a high dose (1.1 × 105 p.f.u.) of the SARS-CoV-2 strain. IAV-infected and SARS-CoV-2-infected mice were monitored daily for morbidity (body weight) and mortality (survival). During the monitoring period, mice were scored for clinical symptoms (weight loss, eye closure, appearance of the fur, posture and respiration). Mice obtaining a clinical score defined as reaching the experimental end-point were humanely killed, according to the experimental protocols approved by the ethical committee. For pDC depletion in hBDCA2-DTR mice, mice were intraperitoneally injected with 15 ng g−1 of DT (Merck), at day 2 after IAV infection. For neutrophil depletion, mice were intraperitoneally injected at day 5 and 6 after IAV infection with 500 µg per mouse of anti-Ly6G (1A8) or isotype control antibodies, both purchased from BioLegend.
Cell preparation
Mice were killed and perfused with 1× PBS. The spleen and lymph nodes were digested for 25 min at 37 °C with Collagenase IV (Worthington Biochemical) and DNase I (Roche Diagnostics). Organs were then mechanically crushed and filtered over 70-µm cell strainers (Corning). BM cells were flushed from mouse femurs. The livers and lungs were collected, minced and mechanically digested with Collagenase IV and DNase I by using GentleMACS (Miltenyi; program 37_m_LIDK_1 for the liver, 37_m_LDK_1 for the lungs). At the end of the digestion, lung cell suspensions were filtered over 70-µm cell strainers and cells were pelleted by centrifugation. The liver cells were submitted to a 80:40 Percoll gradient. The small intestines were collected, opened longitudinally, cut into 1-cm pieces and washed extensively with 1× PBS, then incubated 3× at 37 °C on shaking (200 r.p.m.) with 1× PBS containing 2% fetal calf serum (FCS) and 5 mM EDTA. At the end of each incubation, supernatants were collected and centrifuged. Pelleted cells from the three incubations were pooled together and submitted to a 67:44 Percoll gradient. BALs were performed upon intratracheal injection of 1 ml cold PBS, repeated once. BALs were then centrifuged, fluids collected and frozen at −80 °C until their use. RBC lysis was performed with RBC lysis buffer (Thermo Fisher). For the lungs, the left lobe was collected for RNA extraction, and the other lobes were used for flow cytometry.
Flow cytometry analysis
Cells were first incubated with 2.4G2 monoclonal antibody for 10 min at 4 °C, then we performed extracellular staining in staining buffer, 1× PBS supplemented with 2 mM EDTA (Sigma-Aldrich) and 2% FCS for 30 min at 4 °C. Dead cell staining (LIVE/DEAD Fixable Aqua Dead Cell Stain, Life Technologies) was performed in 1× PBS according to the manufacturer’s recommendations. All antibodies used and their related dilution are listed in the Reporting Summary. Before acquisition, samples were fixed with Cytofix/Cytoperm BD 1× (eBioscience) or with a 2% formaldehyde solution in 1× PBS when using cells expressing EYFP protein. Samples were acquired with a FACS LSR UV (BD Biosciences) using BD Diva v.9.0. All data were analyzed with FlowJo v.10.8.1 software.
RNA extraction and RT–qPCR
The collected organs were incubated in RNAlater (Thermo Fisher) at 4 °C overnight. Total RNA from lung cells was extracted using RNeasy Plus Mini kit (QIAGEN) following the manufacturer’s protocol. Retrotranscription into complementary DNA was performed using the Quantitect Reverse Transcription kit (QIAGEN). The expression levels of the following murine genes were determined by quantitative PCR (qPCR) using the SYBR Premix Ex TaqTM kit and analyzed using the Prism 7500 Fast PCR System. Relative gene expression was calculated using the ΔΔCt method with Actinb as the housekeeping gene for normalization. The primers used were as as follows: Actinb forward 5′-GGCTGTATTCCCCTCCATCG-3′; reverse 5′-CCAGTTGGTAACAATGCCATGT-3′; Il28b forward 5′-GGAGGCCCAGAGCAAGGA-3′; reverse 5′-TTGAAACAGGTTGGAGGTGACA-3′; Irf7 forward 5′-CCACGCTATACCATCTACCTGG-3′; reverse 5′-GCTGCTATCCAGGGAAGACAC-3′; Isg15 forward 5′-GGTGTCCGTGACTAACTCCAT-3′; reverse 5′-TGGAAAGGGTAAGACCGTCCT-3′; Mx2 forward 5′-AGAGGGAGAATGTCGCCTATT-3′; reverse 5′-CGTCCACGGTACTGCTTTTCA-3′; Rsad2 forward 5′-TGCTGGCTGAGAATAGCATTAGG-3′; reverse 5′-GCTGAGTGCTGTTCCCATCT-3′; Ifna1 forward 5′-GGATGTGACCTTCCTCAGACTC-3′; reverse 5′-ACCTTCTCCTGCGGGAATCCAA-3′; Ifna2 forward 5′-ATCCAGAAGGCTCAAGCCATCC-3′; reverse 5′-GGAGGGTTGTATTCCAAGCAGC-3′; Ifna4 forward 5′-AAGGACAGGAAGGATTTTGGAT-3′; reverse 5′-GAGCCTTCTGGATCTGTTGGTT-3′; Ifna5 forward 5′-GGATGTGACCTTCCTCAGACTC-3′; reverse 5′-CACCTTCTCCTGTGGGAATCCA-3′; Ifnb1 forward 5′-GGTGGTCCGAGCAGAGATCTT-3′; reverse 5′-CAGTTTTGGAAGTTTCTGGTAA-3′. The viral titration of MCMV was performed as previously described20. For the viral titration of IAV, we performed quantitative PCR with reverse transcription (RT–qPCR) by testing the expression of M1 IAV gene, using the following primers: forward 5′-AAGACCAATCCTGTCACCTCTGA-3′; reverse 5′-CAAAGCGTCTACGCTGCAGTCC-3′. For viral titration of SARS-CoV-2, tissues were homogenized with ceramic beads in a tissue homogenizer (Precellys, Bertin Instruments) in 0.5 ml RLT buffer, or lung tissues were collected, minced and mechanically digested with Collagenase IV and DNase I by using GentleMACS (Miltenyi; 37_m_LDK_1 program for the lungs). At the end of the digestion, lung cell suspensions were processed with RTL buffer. RNA was extracted using the RNeasy Mini kit (QIAGEN) or Direct-zol RNA Mini PrepPlus de ZYMO RESEARCH, and reverse transcribed using the High-Capacity cDNA Reverse Transcription kit (Thermo Fisher Scientific). Amplification was carried out using OneGreen Fast qPCR Premix (OZYME) according to the manufacturer’s recommendations. Copy numbers of the RNA-dependent RNA polymerase (RdRp) SARS-CoV-2 gene were determined using the following primers: forward 5′-CATGTGTGGCGGTTCACTAT-3′, reverse 5′-GTTGTGGCATCTCCTGATGA-3′. This region was included in a cDNA standard to determine the copy number determination down to 100 copies per reaction. SARS-CoV-2 copy numbers were compared and quantified using a standard curve and normalized to total RNA levels. An external control (mock-infected wild-type animal) and a positive control (SARS-CoV-2 cDNA containing the targeted region of the RdRp gene at a concentration of 104 copies per µl (1.94 × 104 copies per µl detected in the assay)) were used in the RT–qPCR analysis to validate the assay.
Bulk RNA-seq
RNA was isolated from the lungs of uninfected and IAV-infected CTR and pDC-less mice (160 p.f.u., days 0, 3 and 6 after infection) or of uninfected and SARS-CoV-2-infected K18:hACE2 mice (1.1 × 105 p.f.u., days 0, 1, 2, 4 and 6 after infection), as described in the ‘RNA extraction’ section. For bulk RNA-seq, 500 ng of lung RNA per sample was used to prepare libraries for sequencing by using KAPA RNA HyperPrep kits (Roche). Libraries generated with IAV-infected samples were sequenced by Macrogen, and libraries generated from SARS-CoV-2-infected samples were sequenced at the genomic facility of the CIML. All libraries were sequenced using Illumina platforms. Alignments of fastq files obtained after sequencing were generated with STAR v.2.7.9a software using the GRCm38 reference genome. The number of reads mapped to each gene was determined with featureCounts v.2.0.1 using the ‘Mus_musculus.GRCm38.100.gtf’ annotation file. Gene models (gene symbols of the type ‘Gmxxxx’) were then removed from the feature count file and TPM normalization was applied using the Scuttle package of R v.1.0.34.
FITC–dextran
To assess lung permeability, isoflurane-anesthetized mice were administered intratracheally with 10 μg per mouse (50 µl of a solution at 200 µg ml−1 in 1× PBS) of FITC–dextran (Sigma) at 0 and 8 days after IAV infection. At 1 h later, blood was collected from the anesthetized mice from the retro-orbital sinus and the plasma was separated by centrifugation. Plasma were diluted 1:2 in 1× PBS. Dextran leakage in the bloodstream was measured as FITC fluorescence in the plasma. Fluorescence was read at 520 nm with excitation at 490 nm using a Mithras LB 940 plate reader (Berthold Technologies).
Cytokine quantification
Cytokines and chemokines present in the sera of uninfected versus MCMV-infected mice and in the BAL of uninfected versus IAV-infected mice were analyzed using the LEGENDplex Mouse Anti-Virus Response Panel (BioLegend). Samples and standards were plated in technical duplicates and the assay was executed according to the manufacturer’s protocol. Data were collected using the CANTO II (BD Biosciences) and analyzed using the cloud-specific LEGENDplex Data Analysis Software Qognit (BioLegend).
Histological analysis
Lungs were collected from CTR and pDC-less mice 8 days after IAV infection and kept with 10% buffered formalin (VWR Chemical) for 24 h, then dehydrated and embedded in paraffin (Fischer Histoplast). Then, 3.5-µm-thick sections were cut using the microtome Leica RM2245. Hematoxylin and eosin staining was carried out automatically with a Leica Autostainer XL. Finally, the slides were mounted with Entellan mounting medium (Merck) and kept at room temperature. Tissue sections were blindly assessed by a certified veterinary pathologist. Microscopic examination was conducted using a Nikon Eclipse E400 microscope. Pictures were taken with a Nikon DS-Fi2 camera and NIS Elements imaging software (Nikon, Japan). Histopathological semiquantitative lung inflammation scoring system was performed as shown in Supplementary Table 1.
Immunohistofluorescence, confocal microscopy and image analysis
Mouse lungs were perfused with a mixture of 1:2 of AntigenFix (Diapath, containing 4% paraformaldehyde) and of Optimal Cutting Temperature (OCT; Sakura Fineteck), collected, fixed in AntigenFix for 4 h at 4 °C and then washed several times in phosphate buffer (PB; 0.025 M NaH2PO4 and 0.1 M Na2HPO4). Lungs were incubated in PB solution containing 30% sucrose overnight at 4 °C and then embedded in OCT freezing medium, snap frozen and stored at −80 °C. The 30-μm-thick cryosections were obtained using a microtome (Leica 3050 s Cryostat) at temperatures between −24 °C and −20 °C.
For immunostaining, lung sections were blocked with PB, 0.1% Triton X-100 and 2% BSA for 1 h at room temperature and then stained overnight at 4 °C with primary antibodies diluted in PB, 0.1% Triton X-100 and 2% BSA. After several washings with PB, lung sections were then stained in PB, 0.1% Triton X-100 and 2% BSA with secondary antibodies for 2 h at 4 °C. To detect the YFP signal, after the incubation with secondary antibodies, sections were washed and incubated with PB, 0.1% Triton X-100, 2% BSA and 5% rabbit serum for 45 min at room temperature. Sections were then stained with Alexa 488-conjugated anti-GFP antibodies for 1 h at 4 °C. Finally, lung sections were washed with PB and mounted with a coverslip and Prolong Antifade Gold mounting medium (Life Technologies). Stained lung sections were acquired using spectral confocal microscopes (Zeiss LSM880 or Zeiss LSM980) with ×10/0.45 or ×20/0.8 objectives, respectively. Images were acquired using the ZEN blue software (Carl Zeiss Microscopy), in 16-bit format, in spectral mode to suppress autofluorescence for each fluorochrome used. Images were then analyzed using ImageJ software76. For CD45 quantification, areas expressing low versus high CD45 signal were qualitatively identified in each whole lung section, then a region of interest was created for each area and CD45 mean fluorescence intensity (MFI) was measured. In highly inflamed areas CD45 MFI was at least twice of that detected in the lowly inflamed areas. We then used QuPath v.0.4.3 (ref. 77) for cell quantifications. CD45+ cells, IAV-infected HA+ cells and Tom+ pDCs were quantified using the QuPath cell detection module. Then, using the QuPath classifier module, we quantified among the Tom+ pDCs, the IFN+ (Tom+YFP+) pDCs expressing at least a threshold value of 2,216 in the YFP channel. For lung integrity analysis, EpCAM+ areas were defined as areas expressing at least a 5,000 threshold for the EpCAM channel, then their percentage was calculated on the whole lung surface analyzed. EpCAM+ bronchi were classified as closed when EpCAM staining was continuous and as opened when EpCAM was disrupted. Both types of EpCAM+ bronchi were manually quantified and corresponding percentages were calculated.
Statistical analysis
All statistical analyses were performed using GraphPad Prism v.10 software. All results are expressed as mean ± s.e.m. All quantifications were performed with awareness of experimental groups, meaning not in a blinded fashion. Animals were matched in age and sex between experimental groups and each cage was randomly assigned to a treatment group. No animals or data were excluded under any circumstances. Statistical parameters including the exact value of n (number of biological replicates and total number of experiments) and the types of statistical tests are reported in the corresponding figure legends. Comparisons between two groups were assessed using an unpaired and nonparametric Mann–Whitney U-test. Multiple comparisons were assessed using a Kruskal–Wallis test followed by Dunn’s post hoc test. Survival curves were analyzed using a log-rank (Mantel–Cox) test. Comparisons between groups were planned before statistical testing and target effect sizes were not predetermined. Each dot indicates an individual. P values are specified in each figure panel.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Bulk RNA-seq data have been deposited in the Gene Expression Omnibus repository under accession no. GSE302271 for IAV-infected samples and GSE302124 for SARS-CoV-2-infected samples. Source data are provided with this paper.
References
Tomasello, E., Pollet, E., Vu Manh, T. P., Uze, G. & Dalod, M. Harnessing mechanistic knowledge on beneficial versus deleterious IFN-I effects to design innovative immunotherapies targeting cytokine activity to specific cell types. Front. Immunol. 5, 526 (2014).
Broggi, A. et al. Type III interferons disrupt the lung epithelial barrier upon viral recognition. Science 369, 706–712 (2020).
Major, J. et al. Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science 369, 712–717 (2020).
Reizis, B. Plasmacytoid dendritic cells: development, regulation, and function. Immunity 50, 37–50 (2019).
Ngo, C., Garrec, C., Tomasello, E. & Dalod, M. The role of plasmacytoid dendritic cells (pDCs) in tissue immunity during viral infections and beyond. Cell Mol. Immunol. 21, 1008–1035 (2024).
Bocharov, G. et al. A systems immunology approach to plasmacytoid dendritic cell function in cytopathic virus infections. PLoS Pathog. 6, e1001017 (2010).
Reizis, B., Bunin, A., Ghosh, H. S., Lewis, K. L. & Sisirak, V. Plasmacytoid dendritic cells: recent progress and open questions. Annu. Rev. Immunol. 29, 163–183 (2011).
Sa Ribero, M., Jouvenet, N., Dreux, M. & Nisole, S. Interplay between SARS-CoV-2 and the type I interferon response. PLoS Pathog. 16, e1008737 (2020).
von Bernuth, H. et al. Pyogenic bacterial infections in humans with MyD88 deficiency. Science 321, 691–696 (2008).
Ku, C. L. et al. Selective predisposition to bacterial infections in IRAK-4-deficient children: IRAK-4-dependent TLRs are otherwise redundant in protective immunity. J. Exp. Med. 204, 2407–2422 (2007).
Casanova, J. L. & Abel, L. Mechanisms of viral inflammation and disease in humans. Science 374, 1080–1086 (2021).
Ciancanelli, M. J. et al. Infectious disease. Life-threatening influenza and impaired interferon amplification in human IRF7 deficiency. Science 348, 448–453 (2015).
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370, eabd4570 (2020).
Campbell, T. M. et al. Respiratory viral infections in otherwise healthy humans with inherited IRF7 deficiency. J. Exp. Med. 219, e20220202 (2022).
Zhang, Q. et al. Recessive inborn errors of type I IFN immunity in children with COVID-19 pneumonia. J. Exp. Med. 219, e20220131 (2022).
Asano, T. et al. X-linked recessive TLR7 deficiency in ~1% of men under 60 years old with life-threatening COVID-19. Sci. Immunol. 6, eabl4348 (2021).
Garcia-Garcia, A. et al. Humans with inherited MyD88 and IRAK-4 deficiencies are predisposed to hypoxemic COVID-19 pneumonia. J. Exp. Med. 220, e20220170 (2023).
Mogensen, T. H. Human genetics of SARS-CoV-2 infection and critical COVID-19. Clin. Microbiol Infect. 28, 1417–1421 (2022).
Duncan, C. J. A. et al. Life-threatening viral disease in a novel form of autosomal recessive IFNAR2 deficiency in the Arctic. J. Exp. Med. 219, e20212427 (2022).
Cocita, C. et al. Natural killer cell sensing of infected cells compensates for MyD88 deficiency but not IFN-I activity in resistance to mouse cytomegalovirus. PLoS Pathog. 11, e1004897 (2015).
Dalod, M. & Scheu, S. Dendritic cell functions in vivo: a user’s guide to current and next- generation mutant mouse models. Eur. J. Immunol. 52, 1712–1749 (2022).
Swiecki, M., Gilfillan, S., Vermi, W., Wang, Y. & Colonna, M. Plasmacytoid dendritic cell ablation impacts early interferon responses and antiviral NK and CD8(+) T cell accrual. Immunity 33, 955–966 (2010).
Stutte, S. et al. Type I interferon mediated induction of somatostatin leads to suppression of ghrelin and appetite thereby promoting viral immunity in mice. Brain Behav. Immun. 95, 429–443 (2021).
Le Goffic, R. et al. Detrimental contribution of the Toll-like receptor (TLR)3 to influenza A virus-induced acute pneumonia. PLoS Pathog. 2, e53 (2006).
Valente, M. et al. Novel mouse models based on intersectional genetics to identify and characterize plasmacytoid dendritic cells. Nat. Immunol. 24, 714–728 (2023).
Leylek, R. et al. Integrated cross-species analysis identifies a conserved transitional dendritic cell population. Cell Rep. 29, 3736–3750 e3738 (2019).
Sulczewski, F. B. et al. Transitional dendritic cells are distinct from conventional DC2 precursors and mediate proinflammatory antiviral responses. Nat. Immunol. 24, 1265–1280 (2023).
Dalod, M. et al. Interferon α/β and interleukin 12 responses to viral infections: pathways regulating dendritic cell cytokine expression in vivo. J. Exp. Med. 195, 517–528 (2002).
Zucchini, N. et al. Individual plasmacytoid dendritic cells are major contributors to the production of multiple innate cytokines in an organ-specific manner during viral infection. Int Immunol. 20, 45–56 (2008).
Abbas, A. et al. The activation trajectory of plasmacytoid dendritic cells in vivo during a viral infection. Nat. Immunol. 21, 983–997 (2020).
Dalod, M. et al. Dendritic cell responses to early murine cytomegalovirus infection: subset functional specialization and differential regulation by interferon α/β. J. Exp. Med. 197, 885–898 (2003).
Steinberg, C. et al. The IFN regulatory factor 7-dependent type I IFN response is not essential for early resistance against murine cytomegalovirus infection. Eur. J. Immunol. 39, 1007–1018 (2009).
Baranek, T. et al. Differential responses of immune cells to type I interferon contribute to host resistance to viral infection. Cell Host Microbe 12, 571–584 (2012).
GeurtsvanKessel, C. H. et al. Clearance of influenza virus from the lung depends on migratory langerin+CD11b− but not plasmacytoid dendritic cells. J. Exp. Med. 205, 1621–1634 (2008).
Wolf, A. I. et al. Plasmacytoid dendritic cells are dispensable during primary influenza virus infection. J. Immunol. 182, 871–879 (2009).
Kaminski, M. M., Ohnemus, A., Cornitescu, M. & Staeheli, P. Plasmacytoid dendritic cells and Toll-like receptor 7-dependent signalling promote efficient protection of mice against highly virulent influenza A virus. J. Gen. Virol. 93, 555–559 (2012).
Davidson, S., Crotta, S., McCabe, T. M. & Wack, A. Pathogenic potential of interferon αβ in acute influenza infection. Nat. Commun. 5, 3864 (2014).
Davidson, S. et al. IFNlambda is a potent anti-influenza therapeutic without the inflammatory side effects of IFNα treatment. EMBO Mol. Med. 8, 1099–1112 (2016).
Rappe, J. C. F. et al. A TLR7 antagonist restricts interferon-dependent and -independent immunopathology in a mouse model of severe influenza. J. Exp. Med. 218, e20201631 (2021).
Scheu, S., Dresing, P. & Locksley, R. M. Visualization of IFNβ production by plasmacytoid versus conventional dendritic cells under specific stimulation conditions in vivo. Proc. Natl Acad. Sci. USA 105, 20416–20421 (2008).
Tomasello, E. et al. Molecular dissection of plasmacytoid dendritic cell activation in vivo during a viral infection. EMBO J. 37, e98836 (2018).
Creusat, F. et al. IFN-γ primes bone marrow neutrophils to acquire regulatory functions in severe viral respiratory infections. Sci. Adv. 10, eadn3257 (2024).
Chen, H. et al. Pulmonary permeability assessed by fluorescent-labeled dextran instilled intranasally into mice with LPS-induced acute lung injury. PLoS ONE9, e101925 (2014).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 e1039 (2020).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd1554 (2020).
Laurent, P. et al. Sensing of SARS-CoV-2 by pDCs and their subsequent production of IFN-I contribute to macrophage-induced cytokine storm during COVID-19. Sci. Immunol. 7, eadd4906 (2022).
McCray, P. B. Jr. et al. Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. J. Virol. 81, 813–821 (2007).
Takahashi, T. et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 588, 315–320 (2020).
Channappanavar, R. et al. IFN-I response timing relative to virus replication determines MERS coronavirus infection outcomes. J. Clin. Invest. 129, 3625–3639 (2019).
Channappanavar, R. et al. Dysregulated type I interferon and inflammatory monocyte-macrophage responses cause lethal pneumonia in SARS-CoV-infected mice. Cell Host Microbe 19, 181–193 (2016).
Ullah, T. R. et al. Pharmacological inhibition of TBK1/IKKε blunts immunopathology in a murine model of SARS-CoV-2 infection. Nat. Commun. 14, 5666 (2023).
Swiecki, M. et al. Cell depletion in mice that express diphtheria toxin receptor under the control of SiglecH encompasses more than plasmacytoid dendritic cells. J. Immunol. 192, 4409–4416 (2014).
Takagi, H. et al. Plasmacytoid dendritic cells are crucial for the initiation of inflammation and T cell immunity in vivo. Immunity 35, 958–971 (2011).
Cisse, B. et al. Transcription factor E2-2 is an essential and specific regulator of plasmacytoid dendritic cell development. Cell 135, 37–48 (2008).
Vu Manh, T. P. et al. Defining mononuclear phagocyte subset homology across several distant warm-blooded vertebrates through comparative transcriptomics. Front. Immunol. 6, 299 (2015).
Swiecki, M., Wang, Y., Gilfillan, S. & Colonna, M. Plasmacytoid dendritic cells contribute to systemic but not local antiviral responses to HSV infections. PLoS Pathog. 9, e1003728 (2013).
Jamali, A. et al. Characterization of resident corneal plasmacytoid dendritic cells and their pivotal role in herpes simplex keratitis. Cell Rep. 32, 108099 (2020).
Yang, K. et al. Human TLR-7-, -8-, and -9-mediated induction of IFN-α/β and -lambda Is IRAK-4 dependent and redundant for protective immunity to viruses. Immunity 23, 465–478 (2005).
Zhang, Q. et al. Autoantibodies against type I IFNs in patients with critical influenza pneumonia. J. Exp. Med. 219, e20220514 (2022).
Piersma, S. J. et al. Virus infection is controlled by hematopoietic and stromal cell sensing of murine cytomegalovirus through STING. Elife 9, e56882 (2020).
Marie, I., Durbin, J. E. & Levy, D. E. Differential viral induction of distinct interferon-α genes by positive feedback through interferon regulatory factor-7. EMBO J. 17, 6660–6669 (1998).
Prakash, A. & Levy, D. E. Regulation of IRF7 through cell type-specific protein stability. Biochem. Biophys. Res. Commun. 342, 50–56 (2006).
Brewitz, A. et al. CD8(+) T cells orchestrate pDC-XCR1(+) dendritic cell spatial and functional cooperativity to optimize priming. Immunity 46, 205–219 (2017).
Kadoki, M. et al. Organism-level analysis of vaccination reveals networks of protection across tissues. Cell 171, 398–413 e321 (2017).
Beura, L. K. et al. Normalizing the environment recapitulates adult human immune traits in laboratory mice. Nature 532, 512–516 (2016).
Reese, T. A. et al. Sequential infection with common pathogens promotes human-like immune gene expression and altered vaccine response. Cell Host Microbe 19, 713–719 (2016).
Onodi, F. et al. SARS-CoV-2 induces human plasmacytoid predendritic cell diversification via UNC93B and IRAK4. J. Exp. Med. 218, e20201387 (2021).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Venet, M. et al. Severe COVID-19 patients have impaired plasmacytoid dendritic cell-mediated control of SARS-CoV-2. Nat. Commun. 14, 694 (2023).
Buitendijk, M., Eszterhas, S. K. & Howell, A. L. Gardiquimod: a Toll-like receptor-7 agonist that inhibits HIV type 1 infection of human macrophages and activated T cells. AIDS Res. Hum. Retroviruses 29, 907–918 (2013).
Meng, F. Z. et al. TLR7 activation of macrophages by imiquimod inhibits HIV infection through modulation of viral entry cellular factors. Biology 10, 661 (2021).
Greene, T. T. & Zuniga, E. I. Type I interferon induction and exhaustion during viral infection: plasmacytoid dendritic cells and emerging COVID-19 Findings. Viruses 13, 1839 (2021).
Bruel, T. et al. Plasmacytoid dendritic cell dynamics tune interferon-alfa production in SIV-infected cynomolgus macaques. PLoS Pathog. 10, e1003915 (2014).
Mitchell, J. L. et al. Plasmacytoid dendritic cells sense HIV replication before detectable viremia following treatment interruption. J. Clin. Invest. 130, 2845–2858 (2020).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Acknowledgements
We thank CIPHE for the generation of Pacsin1LSL-DTA mice and their assistance in the breeding of mice, as well as the staff of the CIML mouse houses, flow cytometry, histology, genomics, bioinformatics (CB2M) and imaging (ImagImm) core facilities. We thank R. Akyol for his assistance during the generation of SBMC mice and L. Spinelli for his assistance with statistical analyses. We thank the Centre de Calcul Intensif d’Aix-Marseille for granting access to its high-performance computing resources. We thank C. Svanborg (Lund University, Sweden), for providing Irf7KO mice. We thank M.-L. Dessain (TAAM, Orleans, France), for genotyping Mx1 mice. We also thank C. Lutz and the Jackson Laboratory for providing the K18:hACE2 mice and S. van der Werf, X. Lescure and Y. Yazdanpanah for the β CoV/France/IDF0372/2020 strain. This research was funded by the ‘Fondation pour la Recherche Médicale’ (FRM; grant DEQ20180339172, Equipe Labellisée, to M.D.) and from the French National Research Agency (ANR; no. ANR-21-CO12-0001-01, ‘RIPCOV’ to M.D. and B.M., and no. ANR-23-CE15-0001, ‘RIPIREVI’, to M.D. and A.Z.). This work received funding from the French government, managed by ANRS MIE under France 2030, reference number ‘ANRS-24-PEPRMIE-0009’. The projects that contributed to this publication also received funding from the DCBIOL Labex (ANR-11-LABEX-0043, grant no. ANR-10-IDEX-0001-02 PSL*), the Excellence Initiative of Aix-Marseille Université-A*Midex (grant nos. ANR-11-IDEX-0001-02 and AMX-20-TRA-008) and France-BioImaging (grant no. ANR-10-INBS-04) and PHENOMIN-Infrastructure (no. ANR-10-INBS-07), funded by the French government’s ‘Investissements d’Avenir’ program managed by the ANR. We also acknowledge institutional support from CNRS (French National Centre for Scientific Research), INSERM (French National Institute of Health and Medical Research) and Aix-Marseille Université. C.N. is supported by a fellowship from the French Ministry of Research and Higher Education and by a fourth year stipend from FRM.
Author information
Authors and Affiliations
Contributions
C.N. designed, performed and analyzed most of the experiments, and contributed to writing the manuscript. C.P.M. performed experiments for the manuscript revision. K.R. performed RT–qPCR experiments on MCMV- and IAV-infected samples. M.V. prepared the libraries used for bulk RNA-seq samples and contributed to performing the experiments on SARS-CoV-2-infected samples. N.C. genotyped and stabilized mutant pDC-less mice. G.B. performed viral titration on MCMV-infected samples. M. Fabregue, S.S., S. Mathieu, C.G., S. Mazzoli and A.S. bred K18:hACE and related double-/triple-mutant mice and performed the experiments with SARS-CoV-2-infected mice and samples. F.F. directed, designed and analyzed the generation of Pacsin1LSL-DTA mice. C.L. performed anatomopathological analysis of lung sections from IAV-infected mice. L.A., M.G., C.G., A.B. and S.W. provided advice and shared reagents, mutant mice and experimental protocols. R.R. and N.H., respectively, supervised and performed experiments on the brains of IAV-infected mice. P.M. provided advice and technical support for bulk RNA-seq experiments. B.E. and T.P.V.M. performed the mapping and the first analysis of bulk RNA-seq data. M. Fallet contributed to the setting of the analysis of confocal microscopy images. L.C. performed paraffin-embedding, prepared and stained lung sections used for anatomopathological studies. H.T. genotyped and provided Irf7KO mice. T.B. provided advice and shared experimental protocols. M.L.B. provided B6.A2G-Mx1+/+ mice. B.M. contributed to directing and funding the study. A.Z., M.D. and E.T. contributed to funding the study and directed it, designed experiments, performed some experiments, analyzed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.F., B.M., A.Z., M.D. and E.T. are inventors on a patent application filed on the targeting of pDCs to treat certain respiratory viral infections. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Anne Krug and Meredith O’Keeffe for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: S. Houston, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gating strategy for the identification of myeloid and lymphoid lineages.
a, c, Gating strategy used to identify myeloid and lymphoid immune cell types studied. b, The weight of CTR vs pDC-less was measured along time. Weight mean of the individuals (n = 5) of the same strain was represented at each time point analyzed.
Extended Data Fig. 2 Complementary analysis of MCMV-infected control and pDC-less mice.
a, Cytometric bead arrays was performed to quantify CCL2 concentrations in sera isolated from uninfected (0) and 36 h infected (1.5) CTR and pDC-less mice. Each dot represents an individual. The data shown (mean +/− s.e.m.) are pooled from two independent experiments (n = 3/strain at 0, 7/strain at 1.5 p.i.).). An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction was used for the statistical analysis. b, Weight curves of mice represented in Fig. 2a (n = 10 for each strain). c, Relative expression of Mx2 (data represented as mean +/− s.e.m.) in the spleens of C57BL/6 infected with MCMV at indicated time points (spleen, n = 3, except 2 at 2 days p.i.; lungs= 2, except 3 at day 0; liver=3, except 2 at 0 and 2 days p.i.). d, Expression levels of indicated genes was analyzed by RT-qPCR in indicated organs isolated from uninfected or MCMV-infected CTR, pDC-less and Ifnar1KO mice at indicated days after infection. Data were normalized to expression level of Actin gene. The data (mean + s.e.m.) shown are from two independent experiments (n = 4 for each condition and time point, except 3 pDC-less at 1.5 and 2 days p.i. and 3 Ifnar1KO at 2 days p.i.). Statistical analysis was performed as described in Fig. 2d. e, CD86 MFI values on indicated cDC subsets (mean +/− s.e.m.) were determined in CTR and pDC-less mice at indicated days after MCMV infection (n = 3 at 0, 8 for 1.5, 7 at 2 and 3 days after infection for both strains). Statistical analysis was performed as described in Fig. 2e.
Extended Data Fig. 3 Weight curves of IAV-infected mice deficient in pDCs, in the response to IFN-I or to all IFNs, or in TLR7.
a–c The weight of indicated IAV-infected mice was regularly measured up to 2-3 weeks after infection. Data are represented as percentages of the weight at day 0 of the infection. Mice reaching the end points were euthanized. a, n = 9 CTR and 10 pDC-less, b, 9 for both strains, c, 10 Mx1+, 8 Mx1+ pDC-less, 10 Mx1+;Ifnar1KO mice. d, M1 IAV expression was measured by RT-qPCR in the lungs of indicated mice 2 days after IAV infection. n = 5 for each strain. A nonparametric one-way ANOVA (Kruskal-Wallis test followed by Dunn’s post-hoc test) was used for the statistical analysis. Data are represented as mean +/− s.e.m. e, f Weight loss of indicated IAV-infected mice was measured during infection. Data are represented as percentages of the weight at day 0 of the infection. e, n = 15 CTR, 8 Ifnar1KO, 7 Ifngr1KO, 9 Ifnl2/3KO and 9 Stat1KO mice. f, n = 10 CTR, 9 Tlr7KO mice.
Extended Data Fig. 4 During IAV infection pDCs are increased in the lungs and are a main source of IFN-α.
a, b, Gating strategy for (a) and percentages of (b) cDCs and tDCs isolated from the lungs of IAV-infected Ifnb1EYFP mice. A nonparametric one-way ANOVA (Kruskal-Wallis test followed by Dunn’s post-hoc test) was used for the statistical analysis. n = 2 tDC and 1 cDC at 0, 3 at 2, 6 at 4 and 6, 4 at 8 days after infection. Data are represented as mean +/− s.e.m. c, Normalized expression (log2) of indicated genes based on Bulk RNASeq samples obtained by whole lung samples isolated from CTR and pDC-less mice at day 0 (n = 3/strain), 3 and 6 (n = 4/strain) after IAV infection. Data were analyzed using an unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction. d, Highly vs lowly inflamed areas were defined taking in account the CD45 MFI. For each tissue section CD45 MFI was at least twice in highly inflamed areas, as compared to lowly ones. e–h, The numbers of CD45+ cells (e), IAV+ cells (f), pDCs (g) and YFP+ pDCs (h) per mm² were quantified in highly and lowly inflamed areas in IAV-infected lungs. The data shown (mean +/− s.e.m.) in d–h are the pool of the analysis of two tissue sections/lung lobe/mouse for the following numbers of infected SCRIPT mice: n = 5 at days 2 and 3, n = 10 at day 4 and n = 4 at day 6 after infection. An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction was used for the statistical analysis.
Extended Data Fig. 5 Weight curves of IAV-infected pDC SBMC mice deficient in pDCs or in molecules involved in Tlr7/Myd88/Irf7 signaling pathway.
a–c, The weight of indicated IAV-infected mice was regularly measured up to 3-4 weeks after infection. Data are represented as percentages of the weight at day 0 of the infection. Mice reaching the end points were euthanized. d, Clinical scores of indicated IAV-infected SBMC were evaluated daily based on weight loss, eye closure, appearance of the fur, posture, and respiration. a, b, Pool of two independent experiments (8 CTR, 7 Myd88KO and 10 pDC-less SBMC in a; 8 CTR and 9 Tlr7KO SBMC in b); c, d, Pool of three independent experiments (n = 12 for each strain). e, Histological score was performed on the lungs of indicated SBMC at day 8 after infection. n = 9 for each pDC SBMC type. Data are represented as mean +/− s.e.m.
Extended Data Fig. 6 Gating strategy used to identify immune cell types in cells isolated from the lungs of uninfected vs IAV-infected control and pDC-less mice.
a, b, Cells were isolated after enzymatic digestion of lungs of uninfected or IAV-infected animals and analyzed by flow cytometry. Gating strategy used to define indicated immune cell types has been depicted. Plots shown are from one representative mouse out of 40 analyzed. c, d, Percentages of CD45+ and lymphoid immune cells isolated from the lungs of CTR and pDC-less mice isolated at the indicated day after infection. Data are represented as mean +/− s.e.m. In c, n = 7 per strain at days 0 and 3, 8 CTR and 7 pDC-less at day 6 and 7 CTR and 8 pDC-less at day 8 p.i. In d, n = 7 per strain at days 0 and 3, 8 CTR and 7 pDC-less at day 6 and 4 per strain at day 8 p.i. An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak correction method was used for the statistical analysis.
Extended Data Fig. 7 Activation state of immune cells and cytokine production in the lungs of control vs pDC-less mice during IAV infection.
a, b, Mean Fluorescence intensity (MFI) of CD86 (a) and of CD69 (b) molecules expressed by indicated immune cell types isolated from the lungs of CTR or pDC-less mice at indicated days after IAV infection. n = 7/strain at days 0 and 3, 8 CTR and 7 pDC-less at day 6 p.i. An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction was used for the statistical analysis. c, Normalized expression (log2) of indicated cytokine genes based on Bulk RNASeq samples obtained by whole lung samples isolated from CTR and pDC-less mice at day 0 (n = 3/strain), 3 and 6 (n = 4/strain) after IAV infection. An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction was used for the statistical analysis. d, Cytometric bead arrays was performed to quantify cytokine concentrations in the BAL isolated from CTR and pDC-less mice during IAV infection. Each dot represents an individual. The data shown (mean +/− s.e.m.) are pooled from two independent experiments (n = 3/ strain at day 0, 3 CTR and 2 pDC-less at day 2, 7/strain at day 3, 6/strain at day 6 and 7/strain at day 8 p.i.). An unpaired and nonparametric two-sided multiple t-test (Mann-Whitney) with Holm-Sidak method for correction was used for the statistical analysis.
Extended Data Fig. 8 Impact of the loss of pDCs, Ifnar1 or Tlr7 on the weight curves of SARS-CoV-2-infected K18:hACE mice.
a, b, The weight of indicated SARS-CoV-2-infected mice was regularly measured up to 12 days after infection. Data are represented as percentages of the weight at day 0 of the infection. Mice cohort numbers are indicated in the legend of Fig. 8a, b. c, SARS-CoV-2 viral burdens were measured by RT-qPCR as genome equivalent of RdRp protein/ng of total RNA of lungs isolated from K18:hACE mice at 0, 2 and 5 days p.i. Each plotted dot represents the viral burden for an individual animal and bars represent median +/− s.e.m. (n = 1/strain at day 0; 5F and 5M CTR, 5F and 5M pDC-less, 3F and 3M plain Ifnar1KO, 2F and 3M K18:hACE Ifnar1KO at day 2; 4F and 3M CTR, 3F and 4M pDC-less, 3F and 3M plain Ifnar1KO, 2F and 3M K18:hACE Ifnar1KO at day 5 p.i.). A nonparametric one-way ANOVA (Kruskal-Wallis test followed by Dunn’s post-hoc test) was used for the statistical analysis. d, The weight of indicated SARS-CoV-2-infected mice was regularly measured up to 12 days after infection. Data are represented as percentages of the weight at day 0 of the infection. Mice cohort numbers are indicated in the legend of Fig. 8c.
Extended Data Fig. 9 Interferon response and viral control in SARS-CoV-2-infected mice.
a, Normalized gene expression (log2) in Bulk RNASeq lung samples infected with 1.1 105 PFU of SARS-CoV-2. Data (mean +/− s.e.m.) shown are from n = 3 at 0 and 1, 5 at 2 and 4 and 2 at 6 days p.i. b, ISG expression in uninfected or SARS-CoV-2-infected mice by RT-qPCR. The data (mean + s.e.m.) from two independent experiments (n = 1 for all strains and genders, except 0 for male K18:hACE at 0; 5F and 5M K18:hACE and K18:hACExpDC-less, 3F and 3M K18:hACExIfnar1KO and 2F and 3M Ifnar1KO at 2; 4F and 3M K18:hACE, 3F and 4M K18:hACExpDC-less, 2F and 3M K18:hACExIfnar1KO and 2F and 1M Ifnar1KO at 5 days p.i. c, d, % of pDCs (c) and absolute pDC numbers (d) in indicated mice during infection. n = 7F and 2M K18:hACE, 5F and 2M K18:hACExTlr7KO at 0; 5F and 5M K18:hACE, 6F and 5M K18:hACExTlr7KO at 2; 6 per strain and gender at 4 days p.i. e, SARS-CoV-2 viral burdens were measured by RT-qPCR as genome equivalent of RdRp protein/ng of total lung RNA. Each dot represents viral burden/individual, bars represent median +/− s.e.m. In e, 5F and 2M for each strain at day 0; 5F and 5M K18:hACE, 7F and 5M K18:hACExpDC-less, 6F and 5M K18:hACExTlr7KO at day 2; 6F and 6M for each strain at day 4. f, Lung gene expression was analyzed by RT-qPCR and normalized to Actin. n = 7F and 2M K18:hACE, 5F and 2M K18:hACExpDC-less, 5F and 2M K18:hACExTlr7KO at 0; 5F and 5M K18:hACE, 7F and 5M K18:hACExpDC-less, 6F and 5M K18:hACExTlr7KO at 2; 6 for each strain and gender at 4 p.i.). A nonparametric one-way ANOVA (Kruskal-Wallis test with Dunn’s post-hoc test) in a, b, e, f. A two-way ANOVA with Tukey’s multiple comparison test in c, d.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1 and 2 and Table 1.
Supplementary Table 1 (download XLSX )
Statistical data in Supplementary Fig. 1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ngo, C., Pierini-Malosse, C., Rahmani, K. et al. Plasmacytoid dendritic cells are dispensable or detrimental in murine systemic or respiratory viral infections. Nat Immunol 26, 1962–1976 (2025). https://doi.org/10.1038/s41590-025-02288-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02288-3