Abstract
Inflammation is a biological phenomenon beneficial for homeostasis, but it is unfavorable if dysregulated. Although major progress has been made in characterizing inflammation in specific diseases, a global, holistic understanding is still elusive. This is particularly intriguing, considering its function for human health and the potential for modern medicine if fully deciphered. In this study, we leveraged advances in single-cell transcriptomics to delineate inflammatory processes of circulating immune cells during infection, immune-mediated inflammatory diseases and cancer. Our single-cell atlas of more than 6.5 million peripheral blood mononuclear cells from 1,047 patients (56% female, 43% male) and 19 diseases allowed us to learn a comprehensive model of inflammation in circulating immune cells. The atlas expands current knowledge of the biology of inflammation of immune-mediated diseases, acute and chronic inflammatory diseases, infections and solid tumors and lays the foundation to develop a disease classification framework using unsupervised as well as explainable machine learning. Beyond a disease-centered analysis, we charted altered activity of inflammatory molecules in peripheral blood cells, depicting discriminative inflammation-related genes to further understand mechanisms of inflammation. We present a rich resource for the community and lay the groundwork for learning a classifier for inflammatory diseases, presenting cells in circulation as living biomarkers.
Similar content being viewed by others
Main
Inflammation is a state of the immune system that serves to protect the human body from environmental challenges, thereby preserving homeostasis1. Inflammatory processes are activated in response to various triggers, such as infection or injury, and involve a multistep defensive mechanism to eliminate the source of perturbation2. Inflammation represents an altered state within the immune system, which can manifest as either a protective or a pathological response3. The cellular and molecular mediators of inflammation play pivotal roles in nearly every human disease4.
The initiation of inflammatory processes is driven by cellular stimulation, triggered by the release of proinflammatory cytokines5. These cytokines exert autocrine and paracrine effects, activating endothelial cells and subsequently increasing vascular permeability. Chemokines are essential for recruiting additional immune cells for pathogen eradication6. Inflammation is a central driver in cardiovascular7, autoimmune8 and infectious diseases9 and even cancer10. The success of therapies targeting inflammation underscores the importance of understanding the underlying pathways11,12.
Single-cell RNA sequencing (scRNA-seq) is becoming a conventional method for detecting altered cell states, enabling the comparison of transcriptional profiles during inflammation13. A differential analysis of cell states and gene expression programs at the cellular level can guide a more holistic understanding of inflammation in acute and chronic diseases to form the basis for future precision medicine tools. In the present study, we annotated the common immune cell types present in the peripheral blood and identified disease-specific cell states that exhibit functional specialization within the inflammatory landscape. Beyond a disease-centered classification, we modeled the expression profiles of inflammatory molecules to uncover discriminative genes related to immune cell activation, migration, cytotoxic responses and antigen presentation activities. Ultimately, we propose a classifier framework based on peripheral blood mononuclear cells (PBMCs), demonstrating the potential of circulating immune cells to contribute to precision medicine strategies for patients suffering from acute or chronic inflammation.
Results
An inflammation landscape of circulating immune cells
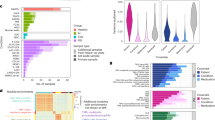
To chart a comprehensive landscape of immune cells in circulation of healthy individuals and patients suffering from inflammatory diseases, we analyzed the transcriptomic profiles of more than 6.5 million PBMCs (6,340,934 after filtering), representing 1,047 patients and 19 diseases, split into a main Inflammation Atlas and two validation datasets (Fig. 1a,b). Diseases were broadly classified into five distinct groups: (1) immune-mediated inflammatory diseases (IMIDs, n = 7) (systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), psoriatic arthritis (PsA), psoriasis (PS), ulcerative colitis (UC), Crohnʼs disease (CD) and multiple sclerosis (MS)); (2) acute (n = 1) (sepsis); (3) chronic inflammation (n = 3) (chronic obstructive pulmonary disease (COPD), asthma and cirrhosis); (4) infection (n = 4) (influenza virus (Flu), SARS-CoV-2 (COVID), hepatitis B virus (HBV) and human immunodeficiency virus (HIV)); and (5) solid tumors (n = 4) (breast cancer (BRCA), colorectal cancer (CRC), nasopharyngeal carcinoma (NPC) and head and neck squamous cell carcinoma (HNSCC)), which were profiled along with healthy donor samples (Fig. 1a and Extended Data Fig. 1a). Our cohort included various scRNA-seq chemistries (10x Genomics 3′ and 5′ mRNA) and experimental designs (CellPlex and genotype multiplexing), as well as individuals of both sexes (56% female, 43% male) and across age groups, to comprehensively capture technical and biological variability (Methods and Supplementary Table 1). To learn a generative model of circulating immune cells of inflammatory diseases, we applied probabilistic modeling of the single-cell data using scVI14 and scANVI15, considering clinical diagnosis, sex and age. Generative probabilistic models proved superior performances in integrating complex datasets compared to other approaches16, particularly if cell annotations are available (Extended Data Fig. 1b,c). Applied here, the resulting cell embedding space was batch effect corrected while preserving biological heterogeneity (that is, previously annotated cell types and states; Supplementary Table 2). From the joint embedding space, we initially assigned cells to major immune cell lineages (Level 1; Fig. 1c and Extended Data Fig. 1d). Then, following a recursive, top-down clustering approach, we obtained a total of 64 immune populations (Level 2), comprehensively resembling immune cell states of the innate and adaptive compartments (Fig. 1d, Supplementary Fig. 1 and Supplementary Table 3). High-level compositional analysis (Level 1) across diseases revealed significant changes of cell type distributions (Extended Data Fig. 1d) and validated previously described alterations in blood cells from patients. For example, we confirmed low levels of unconventional T cells (UTCs), innate lymphoid cells (ILCs) and naive CD4 T cells, together with high proportions of B cells and monocytes, in SLE17. Patients with inflammatory bowel disease (IBD) showed lower levels of UTCs and ILCs18, and we observed lower proportions of UTCs accompanied by a larger fraction of monocytes and B cells in RA19. Lymphopenia, a common event during the development of sepsis20, and lymphocytosis, typical of HIV infection21, were also confirmed.
a, Left, schematic overview illustrating the number of cells, samples and conditions (diseases and disease groups) analyzed. Right, pie charts displaying metadata related to the scRNA-seq chemistry (10x Genomics assay and version) and patient demographics (age and sex). b, Schematic overview of the analysis workflow followed, detailing the division of the overall dataset into Main, unseen patients and unseen studies. The figure illustrates the specific tasks and analyses performed with each dataset. c, Uniform manifold approximation and projection (UMAP) embedding for the scANVI-corrected latent space considering the Main dataset (4,435,922 cells) across patients and diseases colored by the major cell lineages (top, Level 1) and diseases (bottom). d, Sankey diagram showing the Inflammation Atlas cell annotation, considering major cell lineages (Level 1, left) and cell type populations (Level 2, right), along with their correlation to Level 1 cell groups. a,b, Icons were created in BioRender: Aguilar Fernandez, S. (2025): https://biorender.com/h7jfeqm or with Inkscape. CM, central memory; D, disease; DC, dendritic cell; DEG, differentially expressed genes; EM, effector memory; HC, healthy control; HVG, highly variable genes; Mono, monocytes; NK, natural killer; pDC, plasmacytoid dendritic cell; QC, quality control.
Diving deeper into genes and gene programs to characterize inflammatory diseases, our subsequent analysis followed three complementary strategies: (1) to identify disease-driving mechanisms (gene signature and gene regulatory network (GRN) activity); (2) to capture discriminative inflammation-related genes (feature extraction); and (3) to classify patients based on their disease-specific signatures (projection). Therefore, we looked at gene expression profiles holistically but also delineated the inflammatory process by focusing on immune-modulating molecules (Supplementary Table 4).
Inflammation-related signatures across diseases and cell types
We first grouped inflammatory molecules into 21 gene signatures that delineate multiple processes, including immune cell adhesion and activation, cellular migration (chemokines), antigen presentation and cytokine-related signaling (Supplementary Table 4). To tailor these signatures to reflect the inflammation landscape of circulating immune cells, we refined these using Spectra22, yielding a comprehensive set of 119 cell-type-specific factors (Supplementary Table 4). We then ran a univariate linear model (ULM) analysis on the scANVI-corrected gene expression data, providing an inflammation signature activity score for each group. Finally, we ran a linear mixed-effect model (LMEM) between diseased and healthy samples to highlight disease-specific alterations (Supplementary Table 5).
We observed a general trend of increased activity in immune-relevant signatures as compared to healthy donors (>50% increased average signature scores; Fig. 2a). For IMIDs, we found the characteristic upregulation of adhesion molecule signatures, TNF via NFκB signaling, antigen cross-presentation and antigen-presenting signatures23. Interferon (IFN) type 1 and type 2 signatures were significantly downregulated in most IMIDs and cell types, except for non-naive CD8 T cells that showed an upregulation, pointing to a common cell-type-specific mechanism24. Notably, IMIDs showed a strong upregulation of the IFN-induced signature in almost all immune cell types, where SLE was also accompanied by an upregulation of chemokines and chemokine receptors. MS showed a decreased IFN-induced signature and increased chemokine receptor activity, in line with the migratory capacity of blood cells to infiltrate the brain during the course of the disease25. As previously reported, we captured the upregulation of the TNF receptor/ligand signature mainly in non-naive CD8 T cells for sepsis (together with an increase in IFNγ response in monocytes), with a decrease in the other inflammatory signals (adhesion molecules and cytokines)26. By contrast, all chronic inflammatory diseases upregulated the activity of antigen-presenting molecules and increased IFN-induced signaling. This IFN-induced signature was also increased in viral infections, such as Flu and COVID, whereas we found a decreased activity in HIV and HBV. Finally, within solid tumors, CRC and NPC presented a strong upregulation of TNF via NFκB signaling. Intriguingly, only RA, PS, UC and CD showed an enrichment in the T follicular helper (Tfh) signature in non-naive CD4 T cells, highlighting the role of circulating Tfh cells in these diseases. In IMIDs more generally, both naive and non-naive CD4 T cell populations were enriched in T helper signatures, pointing to an early priming of naive T toward helper T cell-driven inflammation27. Finally, to assess the similarity of the inflammatory profiles among diseases, we performed hierarchical clustering of the inflammation signature activity score across all cell types (Level 1; Fig. 2a,b).
a, Heatmap displaying the corrected signature activity score of the 119 cell-type-specific Spectra factors across diseases and cell types (Level 1). Here, the corrected signature activity score represents the coefficient value after running an LMEM comparing diseases versus healthy control (HC) on the ULM estimates computed using the cell type (Level1) and patient pseudobulk on the corrected count matrix. The xaxis represents the Spectra cell-type-specific factor associated with a given function (top annotation). The yaxis represents the diseases grouped by disease group. b, Agglomerative hierarchical clustering with complete linkage, performed considering the Euclidean distance among columns, based on the corrected immune-related signature activity score computed by disease and cell type (Level 1). c, Heatmap displaying the corrected IFN type 1 and type 2 signature activity score across non-naive CD8 T cells (Level 2) and IMIDs. Here, the corrected signature activity score represents the coefficient value after running an LMEM comparing diseases versus HC on the ULM estimates computed using the cell type (Level2) and patient pseudobulk on the corrected count matrix. For a and c, significant signature activity differences between disease and HC are marked with a dot (·) (LMEM, FDR-adjusted P < 0.05). d, Dot plot showing the uncorrected average expression of the FGFBP2 and GZMB genes from IFN type 1 and type 2 signature (xaxis) across different subpopulations of non-naive CD8 T cells (Level 2) on IMIDs and health (SCGT00 study). The dot size reflects the percentage of cells of each disease expressing each gene, and the color represents the average expression level. e, Scaled relative activity of STAT1 and SP1 across cell types (Level 1) and enriched diseases for their transcription factor target genes. Hatched boxes indicate cell types not enriched in the corresponding disease. f, Heatmap representing the average scaled transcription factor activity of STAT1 and SP1 across cell populations (Level 2) for flare and non-flare patients from Perez et al.17. Asterisk (*) indicates statistically significant changes using the two-sided Wilcoxon signed-rank test, FDR-adjusted P < 0.05. CD, Crohnʼs disease; CM, central memory; DC, dendritic cell; MLM, multilevel modeling; MS, multiple sclerosis; EM, effector memory; pDC, plasmacytoid dendritic cell; PS, psoriasis; PSA, psoriatic arthritis; RA, rheumatoid arthritis; TF, transcription factor; UC, ulcerative colitis.
Considering distinct cell types as unique contributors to the inflammatory immune landscape, IFN signatures have been used as a biomarker to define disease activity in autoimmune diseases28. However, it remains elusive which immune subpopulations contribute to these signatures to guide the selection of specific therapeutic interventions. Observing an enriched IFN type 1 and type 2 activity in non-naive CD8 T cells in IMIDs (Fig. 2a), we next sought to discover subpopulations as the signature driver. Here, we observed a significant upregulation across almost all non-naive CD8 T cell populations—however, with a differential pattern across diseases (Fig. 2c). We then decomposed the signal to gene level to identify the most relevant contributors (Supplementary Fig. 2a,b). Intriguingly, FGFBP2 and GZMB showed increased expression levels, with restriction to specific effector memory (EM) CD8 T cell subtypes (EM CX3CR1 high, EM CX3CR1 int, Eff HOBIT and Activated), with a marked increase observed in UC (Fig. 2d). Of note, FGFBP2 and GZMB were recently described as markers of CD8 T cells localized to areas of epithelial damage24. Notably, our blood-based analysis points to their activation in circulating effector CD8 T cell populations even before tissue infiltration.
Expanding on previous observations of increased IFN-induced response across several immune cells and diseases, especially in the myeloid compartment for patients with SLE17 (Fig. 2a), we conducted a GRN analysis to explore the regulatory mechanisms and transcription factors driving the IFN-related activity (Level1; Methods). STAT1 and SP1 were identified as the primary regulators of the IFN-induced signature, with each transcription factor exhibiting cell-type-specific activities (Fig. 2e and Supplementary Table 6). STAT1 primarily regulated canonical IFN signaling genes across multiple lineages, whereas SP1 activated a heterogeneous set of target genes (Extended Data Fig. 2a and Supplementary Table 6)29.
Observing a broad IFN-induced activity across immune cell types, we next investigated whether STAT1 and SP1 regulatory programs were conserved across cell subpopulations (Level 2; Extended Data Fig. 2b,c and Supplementary Table 6). Here, patients with SLE exhibited opposing STAT1 and SP1 activities in monocytes and non-naive CD8 T cells. STAT1 activity was increased in non-classical monocytes, whereas SP1 activity was decreased30. STAT1 was also upregulated in conventional dendritic cells type 2 (cDC2s), whereas SP1 activity was increased across multiple cell types implicated in the pathogenesis of SLE31, including inflammatory and regulatory monocytes, EM CX3CR1 high, CM and activated CD8 T cells as well as adaptive and CD56dimCD16 natural killer cells. Patients with Flu showed a significant increase in STAT1 activity in IFN-response CD8 T cells (Extended Data Fig. 2b). By contrast, patients with cirrhosis presented higher SP1 activity specifically in IFN-response monocytes. In HNSCC, an increased SP1 activity was observed in non-classical monocytes (Extended Data Fig. 2c), a protumoral population related to the suppressive systemic state of monocytes in this cancer type32. Given the cell-type-specific regulatory patterns observed across diseases, we next investigated the contribution of STAT1 and SP1 activity to dynamic changes associated with disease progression. To this end, we assessed their activity in patients with SLE17 experiencing disease exacerbations (flares; Supplementary Table 6). STAT1 activity was elevated during flares, particularly within CD8 T cells, whereas SP1 activity was more prominent in myeloid populations in the absence of flares (Fig. 2f).
Functional gene selection through interpretable modeling
Gene discovery using linear models or standard differential expression analysis suffers from the limitation that genes are considered independently. Thus, we considered the possibility of categorizing cells to their respective disease origin through an interpretable machine learning pipeline, to guide the selection of functional disease discriminatory genes (Methods and Supplementary Table 4). Therefore, we applied a supervised classification approach, together with a post hoc interpretability method, to allow the inference of the gene-wise importance, stratified by disease and cell type (Level 1).
We based our strategy on gradient boosted decision trees (GBDTs), a state-of-the-art machine learning technique proven to be effective in complex tasks with noisy data and nonlinear feature dependencies33 (Methods and Supplementary Table 7). To account for cell-type-specific expression patterns and the differential impact of diseases across immune populations, we trained separate models for each cell type (Level 1). We applied the classification pipeline to the scANVI-corrected gene expression profiles, achieving a balanced accuracy score (BAS) of 0.87 and a weighted F1 (WF1) score of 0.90 on held-out samples (Fig. 3a and Supplementary Table 8). Instead, uncorrected log-normalized counts led to a reduced performance, underscoring the benefits of batch correction (BAS: 0.65 and WF1: 0.78; Fig. 3a). Performances were consistent among cell types, with less abundant cell populations obtaining generally lower scores (for example, plasma cells, BAS: 0.78 and WF1: 0.80; Extended Data Fig. 3 and Supplementary Table 8). We observed that certain diseases exhibited poorer classification performance—for example, the misclassification of patients with severe Flu as COVID (Extended Data Fig. 3). Retraining the GBDT classifier on the Flu and COVID dataset (COMBAT dataset34) and stratifying patients with COVID by their clinical behavior (mild, severe and critical) identified patients with severe Flu to closely resemble severe COVID cases (Extended Data Fig. 4a,b). Similar results were obtained by clustering pseudobulks at the sample level (Extended Data Fig. 4c,d), supporting common inflammatory signatures of patients suffering from these severe respiratory infections. Finally, separating cells from female and male patients yielded similar performances, with no differences between sexes (Extended Data Fig. 5a).
a, Normalized confusion matrices displaying proportion of predictions belonging to each true condition. Diagonal values correspond to the Recall metric. XGBoost was trained on the scANVI batch-corrected (left) or batch-uncorrected (right) log-scaled cell expression profiles. b, Validation of d-SHAP-based gene selection using XGBoost trained with a nested cross-validation on unseen studies’ cells. Each point corresponds to the average left-out fold performance, for each best configuration of each fold combination. The box plots report the WF1 (top) and the BAS (bottom) computed considering top 5, 10 and 20 genes (among the ones expressed in at least 5% of the total cells), for each inflammatory condition present within the unseen studies dataset (that is, healthy, sepsis, CD, SLE, HIV, cirrhosis, RA and COVID) according to the d-SHAP values, across cell types (Level 1). For the same number of genes, we report the performance scores of n = 20 random selected gene sets. The performance of the classifier when trained on the whole gene set, consisting of the genes expressed in at least 5% of the total cells, is also reported. Boxes indicate the interquartile range (IQR) with the median as a center line; whiskers extend to 1.5× IQR; and outliers are shown as individual points. c, Scatter plot of max-normalized gene expression against d-SHAP values computed for CYBA gene on monocyte population (Level 1) and considering the output of disease-XGBoost for a given disease (UC, CD, PS and PSA, from left to right). d, Scatter plot of max-normalized gene expression against d-SHAP values computed for IFITM1 gene on T non-naive CD4 and ILC populations (annotation Level 1) considering the output of the disease-XGBoost for a given disease (asthma and COPD, left and right). In c and d, we limited the visualization to up to 60,000 cells, sampling an equal percentage from each patient corresponding to 5% and 7.5% of monocytes and T non-naive CD4 cells, respectively. Cells belonging to samples with or without the given condition (disease) are marked in orange or blue, respectively. CD, Crohnʼs disease; MS, multiple sclerosis; PS, psoriasis; PSA, psoriatic arthritis; RA, rheumatoid arthritis; UC, ulcerative colitis.
As GBDTs require post hoc interpretability tools, we computed SHapley Additive exPlanation (SHAP)35 values. By combining the two approaches, we obtained a rich resource of gene rankings based on their ability to discriminate inflammatory conditions across different cell types (Methods and Supplementary Table 9). To evaluate the effectiveness of disease-discriminative SHAP (d-SHAP) values, we assessed the classification performance compared to an equal number of randomly selected genes. On unseen studies, d-SHAP genes consistently yielded more accurate predictions (Fig. 3b). Due to the possible collinearity of diseases and studies, d-SHAP values might be affected by batch effects. To disentangle disease-specific from study-specific signals, we trained separate classifiers to predict the study identity (BAS: 0.97 and WF1: 0.99; Supplementary Fig. 4a) and to identify study-associated genes via SHAP values (s-SHAP; Methods). The correlation and overlap between the d-SHAP and s-SHAP values (Supplementary Fig. 4b,c) allowed us to prioritize bona fide disease-discriminative genes for further analysis (Supplementary Table 9).
Ordering genes based on d-SHAP values identified previously described biomarkers, such as STAT3 in CD4 T cells for RA samples36 and IFN genes for SLE samples37 (Extended Data Fig. 6a). The d-SHAP values of CYBA stood out as a strong candidate marker to classify diseases affecting barrier tissue: PSA, PS, UC and CD (Fig. 3c and Extended Data Fig. 6b,c). CYBA encodes the primary component of the microbicidal oxidase system of phagocytes. In line, the importance was seen mainly in monocytes (Extended Data Fig. 6b). Interestingly, high expression of CYBA drove the model to classify intestinal inflammatory diseases (UC and CD), whereas reduced levels were relevant to classify skin-related diseases (PS and PSA) (Fig. 3c). Mutations in CYBA cause chronic granulomatous disease, with patients showing an impaired phagocyte activation and failing to generate superoxide. Consequently, patients show recurrent bacterial and fungal infections in barrier tissues, including the skin38. Thus, we hypothesize that reduction of CYBA in skin-related IMIDs leads to an impaired immune barrier function, causing localized, symptomatic flares of PS and PSA. On the other hand, reactive oxidative species (ROS) produced by mucosa-resident cells or by newly recruited innate immune cells are essential for antimicrobial mucosal immune responses39. In IBDs, an upregulation of CYBA may result in the accumulation of superperoxide and ROS through its oxidase function, a hallmark of these diseases40.
Further exploring d-SHAP value ranks highlighted the importance of IFITM1 across chronic diseases, including COPD and asthma (Extended Data Fig. 6d,e). IFITM1 encodes a protein that inhibits viral entry into host cells by preventing the fusion of the virus with the host cell membrane41. The importance of IFITM1 was mainly observed in lymphoid cells, specifically CD4 non-naive T cells and ILCs (Extended Data Fig. 6d and Supplementary Fig. 3). In both cell types, higher IFITM1 expression drives the model toward classifying COPD, whereas lower expression shifts the classification toward asthma (Fig. 3d). In line, T cell and ILC accumulation is associated with the decline of lung function and severity in patients with COPD42. We hypothesize that chronic inflammation triggers higher expression of IFITM1 in lymphoid cells, thereby facilitating their accumulation43, with further mechanistic validation being needed.
Classifying patients by reference mapping
The ability to accurately classify cells according to their respective diseases prompted us to classify patients based on their disease of origin, creating the basis for a universal classifier as a precision medicine tool for inflammatory diseases. By considering each patient as an ensemble of expression profiles across all circulating immune cells, we learned a generative model while integrating the single-cell reference as a basis to project new patients from a query dataset into the same embedding space. Such strategy allowed us to map unseen and unlabeled query patient data into our reference embedding space, providing a common ground for classification.
Projecting expression data into a lower-dimensional space is a common strategy to reduce noise44 and to map query data into a reference atlas45. Here we introduce a novel computational framework to exploit the cell embeddings for classification, thus turning the single-cell reference into a diagnostic tool (Fig. 4a and Extended Data Fig. 7). Therefore, we first generated the embeddings with scANVI (30 latent embeddings) of both the reference and the unseen query datasets while also transferring the cell labels to the latter (Supplementary Table 7). Then, we defined a cell type pseudobulk profile per patient by averaging the embedded features of the corresponding cells (Level 1; Methods). Next, we trained an independent classifier to assign correct disease labels, considering one cell type at a time. We handled uncertainty at cell type level via a majority voting system to determine most frequent conditions. To assess the performance of our framework, we proposed three scenarios: (1) a five-fold cross-validation splitting the full reference atlas into five balanced sets, (2) a dataset with unseen patients and (3) a dataset with unseen studies (Fig. 4b). We consider these scenarios a representation of the data integration challenges with an increasing degree of complexity.
a, Schematic representation of the patient classifier pipeline. Icons were created with Inkscape. b, Description of the three performance evaluation scenarios. In our datasets, we always have only one sample for each patient. c−e, Performance evaluation in Scenario 1 (five-fold cross-validation, from 817 samples), showing: c, distribution of WF1 scores for each left-out split (boxes indicate the interquartile range (IQR) with the median as a center line, whiskers extend to 1.5× IQR and outliers are shown as individual points (each box includes n = 5 points)); d, F1 score for each combination of cell type and disease, after aggregating all the predictions of the left-out folds; and e, normalized confusion matrices displaying proportion of predictions belonging to each true condition after aggregating all the predictions of the left-out folds. Main diagonal values correspond to the Recall metric. f,g, Performance evaluation in Scenario 2, showing WF1 scores for unseen patients’ observation (f) and F1 score for each combination of cell type and disease (g). h, Normalized confusion matrices displaying proportion of predictions belonging to each true condition. Main diagonal values correspond to the Recall metric. i−k, Performance evaluation in Scenario 3, showing: i, WF1 scores for unseen studies’ observation; j, F1 score for each combination of cell type and disease; and k, normalized confusion matrices displaying proportion of predictions belonging to each true condition. Main diagonal values correspond to the Recall metric. CD, Crohn’s disease; DC, dendritic cell; MS, multiple sclerosis; P, patient; pDC, plasmacytoid dendritic cell; PS, psoriasis; PSA, psoriatic arthritis; QC, quality control; RA, rheumatoid arthritis; UC, ulcerative colitis.
Our classification strategy achieved high performance in the cross-validation scenario (Scenario 1; Supplementary Table 8), resulting in 0.90 ± 0.03 WF1 and 0.85 ± 0.07 BAS (Fig. 4c). Consistent with results obtained from the cell-wise classifier pipeline, Flu was the only disease that failed to be classified (Recall: 0.18) (Extended Data Fig. 8a,b). Training a classifier for each cell type separately allowed us to assess their relevance in distinguishing inflammatory diseases (Fig. 4d,e). Here, plasma and UTC showed the lowest BAS (0.53 and 0.67) and WF1 (0.64 and 0.78), highlighting the strength of our majority voting approach as a robust ensemble (Extended Data Fig. 8b). Although certain diseases (COVID, COPD and asthma) were particularly well classified by lymphoid and myeloid cell types, HIV was best classified by naive lymphoid cells (that is, naive CD4 and CD8 T cells and B cells with F1 of 0.83) in line with the tropism of the virus infecting mainly CD4 T cells46,47 (Fig. 4d and Extended Data Fig. 8c). Increasing the complexity by classifying unseen patient samples (Scenario 2), the performance remained very high, with a BAS of 0.95 and a WF1 of 0.98 (Fig. 4f−h and Supplementary Table 8). However, the classification of samples from unseen studies (Scenario 3) resulted in a strongly decreased BAS of 0.12 and a WF1 of 0.23 (Fig. 4i−k and Supplementary Table 8).
The largest performance drop was observed between Scenario 2 and Scenario 3, the latter classifying patients from unseen studies. We hypothesized that confounding factors, such as variations in assay chemistry or research centers, hindered the classifier’s ability to generalize. To validate our hypothesis and to provide a path toward a generalizable patient classifier, we next considered a Centralized Dataset that includes only data from diseases generated in the same center with a single assay chemistry (SCGT00 data; Supplementary Table 1 and Extended Data Fig. 7). In contrast to Scenario 2, we stratified the samples by sequencing pool and disease, ensuring that reference and query patients belong to distinct cohorts. This new centralized approach included an independent annotation of the reference patients’ cells (Methods and Supplementary Table 3) and new scANVI integration of the reference data, before projecting cells of the query patients. Notably, in this context, WF1 and BAS increased to 0.56 and 0.53, respectively, pointing to a highly improved generalization performance when classifying query patients as compared to Scenario 3 (Fig. 5a−c, Extended Data Fig. 9a,b and Supplementary Table 8). Finally, we evaluated the classifier performance considering male and female patients separately. In Scenario 1, no statistically significant differences were observed between WF1 distributions (Extended Data Fig. 5b), and the majority vote approach also yielded consistent results in the other scenarios (Extended Data Fig. 5c–e).
a−c, Performance evaluation in a Centralized Dataset, showing: a, WF1 scores for left-out pool observation (mean and standard deviation of WF1 score of 100 random condition assignments is reported); b, F1 score for each combination of cell type (Level 1) and disease; and c, normalized confusion matrices displaying proportion of predictions belonging to each true condition (main diagonal values correspond to the Recall metric). d,e, Performance evaluation in Scenario 2 (d) and Scenario 3 (e), showing the distribution of WF1 and BAS for all the configurations of each data integration approach (left) and the mean and standard deviation of each data integration method (right), including random assignment (n = 5 for Harmony&Symphony, scANVI and scGen; n = 10 for scPoli cell, sample and cell&sample embedding; n = 100 for random assignment). Arrows highlight scANVI configuration applied in Scenario 1. CD, Crohn’s disease; DC, dendritic cell; pDC, plasmacytoid dendritic cell; PS, psoriasis; PSA, psoriatic arthritis; RA, rheumatoid arthritis; UC, ulcerative colitis.
Consequently, we hypothesize immune cells in circulation to serve as a source for building a universal classifier for inflammatory diseases. Although the here-used subset of the Inflammation Atlas was limited in cell and patient numbers, future efforts for commercialization are required to develop large single-chemistry training datasets and respective models to further increase the classification accuracy.
We selected scANVI as our Inflammation Atlas integration method for its top-ranked performance in data integration benchmarks. To further assess classification performance for the task at hand, we next compared scANVI against other approaches (that is, Harmony/Symphony, scGen and scPoli) and hyperparameter configurations in Scenario 2 and Scenario 3 (Supplementary Table 7). In concordance with our previous results for scANVI, all newly introduced methods achieve high performance in the dataset with unseen patients (Scenario 2; Fig. 5d, Supplementary Table 8 and Extended Data Figs. 9c and 10a,c,e,g). Although all the approaches lost predictive power on the unseen studies datasets (Scenario 3; Supplementary Table 8), Harmony performed best with a BAS of 0.24 and a WF1 of 0.47 (Fig. 5e and Extended Data Figs. 9d and 10b,d,f,h). Although linear approaches (for example, Harmony) have less representation power than variational autoencoders (VAEs), they are also less prone to overfitting and more robust to the hyperparameter choice. Hence, in settings where hyperparameter tuning and validation are not possible due to the lack of condition labels, tools such as Harmony/Symphony might be preferable to more complex VAEs.
Discussion
Comprehensive mapping of the plasticity of the immune cells in circulation is achieved by single-cell sequencing-based immuno-phenotyping48. Recent technologies enable the sampling of thousands of cells per sample and hundreds of thousands per patient cohort, pushing the resolution toward fine-grained cellular maps and increasing the power to identify disease-specific states49. To date, single-cell sequencing has been applied to a multitude of inflammatory diseases to pinpoint disease-driving mechanisms as potential therapeutic targets13. However, a complete map of immune cell states across diseases, holistically charting immune plasticity in inflammatory diseases, has been elusive.
The concept of using immune cells as a sensor for diseases is highly intriguing and opens the door for the development of future universal diagnostic tools50. For diseases such as rheumatic diseases and IBD, many patients are undiagnosed or diagnosed as false positive, and more accurate universal tools are needed51,52. Our approach using GBDT, together with SHAP-based interpretability and a tailored list of functional immune cell molecules, provided explainable outcomes and serves as a rich resource for identifying disease-discriminative genes33. We further tested the utility of an Inflammation Atlas as a liquid biopsy classification tool by developing a patient classifier based on the latent embeddings after integration. To our knowledge, existing patient classifiers have evaluated settings similar to Scenario 1 and Scenario 2 (scPoli and MultiMIL53). In Scenario 3, we then queried patients belonging to studies excluded from our reference atlas, simulating the application of the Inflammation Atlas as a diagnostic tool. Here, our approach initially failed to generalize to unseen patients, indicating that further optimization was needed to build a generalizable model for more accurate disease diagnostics. To explore the reasons for limited generalization, we performed additional analyses on a Centralized Dataset. Here, the improved performance compared to Scenario 3 highlighted the impact of batch effects introduced through differing assay chemistries and centers.
Although our study provides a comprehensive framework for immune profiling across inflammatory diseases, several aspects warrant further exploration. Most samples in our compendium derive from individuals of European ancestry, and expanding to ancestrally diverse populations will be essential to capture global immune variability and improve model generalizability. Our classifier also requires prospective validation in independent, multicenter cohorts to assess robustness and clinical applicability. Finally, understanding the relationship between circulating and tissue-resident immune cells remains key for diagnostic translation. Circulating cells offer a minimally invasive means to monitor disease activity, yet future studies should validate to what extent their molecular programs reflect tissue-resident inflammatory states across organs and disease contexts.
Bringing reference atlases into clinics remains a complex task, particularly without clear implementation strategies. We contributed to this roadmap by generating a comprehensive landscape of circulating immune cells across inflammatory diseases. Toward leveraging single-cell technologies in diagnostics, we call for the definition of best practices and quality control standards to reduce batch effects, alongside generating large, controlled training datasets. To allow data integration methods to fully generalize, we need to reduce the confounding factors, as demonstrated by our centralized approach, or largely increase training data size and variability. Alternatively, a reference training dataset is generated by multiple centers and diverse chemistries to define a large, heterogeneous atlas, enabling the definition of a foundation model54 to pave the way to a universal disease classifier, robust to batch effects.
Methods
Ethics declaration
Human blood processed in-house for this project was preselected and included within other ongoing studies. All the studies included were conducted in accordance with ethical guidelines, and all patients provided written informed consent. Ethical committees and research project approvals for the different studies included in this paper are detailed in the following text.
SCGT00 and SCGT00val were approved by the Hospital Universitari Vall d’Hebron Research Ethics Committee (PR(AG)144/201). SCGT01 received institutional review board (IRB) approval by the Parc de Salut Mar Ethics Committee (2016/7075/I). SCGT02 received ethics approval by the Medisch-Etische Toetsingscommissie (METc) committee—for patients with asthma (ARMS and ORIENT projects: NL53173.042.15 and NL69765.042.19, respectively), for patients with COPD (SHERLOck project, NL57656.042.16) and for healthy controls (NORM project, NL26187.042.09). SCGT03 was approved by the Comité Ético de Investigación con Medicamentos del Hospital Universitario Vall d’Hebron (654/C/2019). SCGT04 and SCGT06 were approved by the Comitè d’Ètica d’Investigació amb medicaments (CEim) del Hospital de la Santa Creu i Sant Pau (EC/21/373/6616 and EC/23/258/7364). SCGT05 was approved by the IRBs of the Commissie Medische Ethiek UZ KU Leuven/Onderzoek (S66460 and S62294).
Atlas of circulating immune cells
The Inflammation Landscape of Circulating Immune Cells atlas was conceived as a comprehensive resource to expand the current knowledge of physiological and pathological inflammation through the study of circulating immune cells. With this aim, we included data representing both acute and chronic inflammatory processes as well as healthy donors. Further details about the included datasets are available (Supplementary Table 1).
The project includes in-house scRNA-seq data generation from samples shared by our collaborators from several research institutions. Samples were collected with written informed consent obtained from all participants and comply with the ethical guidelines for human samples. Specifically, we generated data from patients suffering from rheumatoid arthritis, psoriatic arthritis, Crohnʼs disease, ulcerative colitis, psoriasis and SLE and from healthy controls in collaboration with the Vall d’Hebron Research Institute within the DoCTIS consortia (https://doctis.eu/) (SCGT00 and SCGT00val). Additionally, we processed and obtained data from healthy controls in collaboration with the Institut Hospital del Mar d’Investigacions Mèdiques (SCGT01); asthma, COPD and healthy control samples in collaboration with the University Medical Center Groningen (SCGT02); BRCA samples in collaboration with the Vall d’Hebron Institute of Oncology (SCGT03); cirrhosis samples in collaboration with the Biomedical Research Institut Sant Pau (SCGT04); CRC samples in collaboration with the Katholieke Universiteit Leuven (SCGT05); and COVID and healthy control samples also in collaboration with the Biomedical Research Institut Sant Pau (SCGT06).
Moreover, we also included publicly available datasets to complete our cohort. Specifically, we considered data from patients suffering from sepsis26,55, HNSCC56, HBV57, multiple sclerosis58, NPC59, HIV60,61, SLE17,62,63, cirrhosis64, Crohnʼs disease65, COVID-Flu-sepsis34 and COVID66 and from healthy controls from Terekhova et al.67 and 10x Genomics, together with the available healthy samples from all the cited studies. The data information and access identifiers for each project can be found in Supplementary Table 1 (Sheet 1). When raw data were available, we downloaded FASTQ files; otherwise, we retrieved the raw count matrices from the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/) or Sequence Read Archieve (SRA) (https://submit.ncbi.nlm.nih.gov/about/sra/), BioStudies Array Express (https://www.ebi.ac.uk/biostudies/arrayexpress), Broad Institute DUOS (https://duos.broadinstitute.org/), Synapse (https://www.synapse.org), Genome Sequence Archive (GSA) (https://ngdc.cncb.ac.cn/gsa-human/), CELLxGENE Data Portal (https://cellxgene.cziscience.com/datasets) and 10x Genomics (https://www.10xgenomics.com/datasets) resources. For all studies, we also collected clinical metadata.
Sample collection
Human blood samples were collected in EDTA tubes (BD Biosciences). PBMCs from the SCGT00, SCGT00val, SCGT02, SCGT04, SCGT05 and SCGT06 datasets were isolated using Ficoll density gradient centrifugation (STEMCELL Technologies, Lymphoprep; GE Healthcare Biosciences AB, Ficoll-Plus). PBMCs belonging to the SCGT01 and SCGT03 datasets were isolated using Vacutainer CPT tubes (BD Biosciences). Subsequently, all aliquots were centrifuged following the manufacturer’s protocol. After centrifugation, PBMCs were washed and resuspended in freezing media. Aliquots were gradually frozen using a commercial freezing box (Thermo Fisher Scientific; Mr. Frosty, Nalgene) at −80 °C for 24 hours before being transferred to liquid nitrogen for long-term storage.
Cell thawing and preprocessing
Cryopreserved PBMCs were thawed in a water bath at 37 °C and transferred to a 15-ml Falcon tube containing 10 ml of prewarmed RPMI media supplemented with 10% FBS (Thermo Fisher Scientific). Samples were centrifuged at 350g for 8 minutes at room temperature, supernatant was removed and pellets were resuspended with 1 ml of cold 1× PBS (Thermo Fisher Scientific) supplemented with 0.05% BSA (Miltenyi Biotec, PN 130-091-376). Samples were incubated during 10 minutes at room temperature with 0.1 mg ml−1 DNAse I (Worthington Biochemical, PN LS002007) to eliminate ambient DNA and favor the resuspension of the pellet. Cells were filtered with a 40-µm strainer (Cell Strainer, PN 43-10040-70) to remove eventual clumps and washed by adding 10 ml of cold PBS + 0.05% BSA. Samples were centrifuged at 350g for 8 minutes at 4 °C and resuspended in an adequate volume of PBS + 0.05% BSA to reach the desired concentration. Cell concentration and viability were verified with a TC20 Automated Cell Counter (Bio-Rad) upon staining of the cells with trypan blue.
Sample multiplexing by genotyping
PBMC samples were evenly mixed in pools of eight donors per library following a multiplexing approach based on the donor’s genotype for a more cost-efficient and time-efficient strategy. Notably, in the case of SCGT00, libraries were designed to pool samples together from the same disease with different response to treatment (not relevant in this study), whereas, in the case of the SCGT02 Asthma+HC cohort, six samples belonging to patients were pooled with two samples derived from non-smoking healthy control individuals. With this approach, we aimed to avoid technical artifacts that could mask subtle biological differences.
3′ CellPlex
PBMC samples belonging to the SCGT01, SCGT02 COPD+HC, SCGT04 and SCGT06 cohorts were multiplexed with 10x Genomics CellPlex Kit following the Cell Multiplexing Oligo Labeling for Single Cell RNA Sequencing Protocol (10x Genomics). Whereas, for SCGT02 COPD+HC and SCGT06 projects, we pooled eight samples from patients with healthy controls together, for SCGT01 and SCGT04 we only included samples from the condition of interest. In brief, 0.2−1 million cells were centrifuged at 350g at room temperature with a swinging bucket rotor, resuspended in 100 µl of Cell Multiplexing Oligo (10x Genomics, 3′ CellPlex Kit Set A, PN-1000261) and incubated at room temperature for 5 minutes. Cells were washed three times with cold 1× PBS (Thermo Fisher Scientific) supplemented with 1% BSA (MACS, Miltenyi Biotec), all centrifugations being performed at 350g at 4 °C. Cells were finally resuspended in an appropriate volume of 1× PBS + 1% BSA to obtain a final cell concentration of approximately 1,600 cells per microliter and counted using a TC20 Automated Cell Counter (Bio-Rad). An equal number of cells of each sample was pooled and filtered with a 40-µm strainer to remove eventual clumps; final cell concentration and viability of the pools were verified before loading onto the Chromium for cell partitioning.
Cell encapsulation and scRNA-seq library preparation
Multiplexed samples were loaded for a target cell recovery between 20,000 and 60,000 cells (corresponding to 5,000−7,500 cells per sample within each plex). More specifically, samples belonging to SCGT00, SCGT00val and SCGT01 cohorts were encapsulated using standard-throughput Chromium Next GEM Single Cell 3′ Reagent Kit version 3.1, whereas multiplex samples belonging to SCGT02 Asthma+HC and COPD+HC, SCGT04 and SCGT06 were encapsulated using the high-throughput Chromium Next GEM Single Cell 3′ HT Reagent Kit version 3.1 in combination with the Chromium X instrument. On the other hand, SCGT03 and SCGT05 cohorts were loaded in a standard assay with a target recovery of 6,000−8,000 cells per sample using Chromium Next GEM Single Cell 5′ Reagent Kit version 2 (10x Genomics, PN-1000263).
Libraries were prepared following the manufacturer’s instructions of protocols CG000315 or CG000390, for the standard assay without and with sample multiplexing, and protocols CG000416 and CG000419, for the high-throughput assay without and with sample multiplexing. Protocol CG000331 was instead followed for the SCGT03 and SCGT05 cohorts. Between 20 ng and 200 ng of cDNA was used for preparing libraries, and final library size distribution and concentration were determined using a Bioanalyzer High Sensitivity chip (Agilent Technologies). Sequencing was carried out on a NovaSeq 6000 system (Illumina) and a NextSeq 500 system (Illumina) using the following sequencing conditions: 28 bp (Read 1) + 10 bp (i7 index) + 10 bp (i5 index) + 90 bp (Read 2), to obtain approximately 40,000 read pairs per cell for the gene expression library and 2,000−4,000 read pairs per cell for the CellPlex library.
Data processing
To profile the cellular transcriptome, we processed the sequencing reads with the 10x Genomics software package Cell Ranger (version 6.1) (https://support.10xgenomics.com/single-cell-gene-expression/software/overview/welcome) and mapped them against the human GRCh38 reference genome (GENCODE version 32/Ensembl 98). This step was applied to the sequencing reads obtained from in-house-processed samples and from published projects, when available.
Genotype processing
Genome-wide genotyping data for patients from the SCGT00, SCGT00val and SCGT02 Asthma+HC studies were generated from PBMC samples. For SCGT00, 184 patients were distributed in four genotyping cohorts (N1 = 64, N2 = 32, N3 = 40 and N4 = 48 samples), and, for SCGT00val, 32 patients were processed with the Illumina Omni2.5-8 and Illumina GSA MG v3-24 arrays, respectively. For SCGT02 Asthma+HC, 16 patients were distributed in two genotyping cohorts (N1 = 8 and N2 = 8 samples) using Infinium Global Screening Array-24 version 3.0 (GSAMD-24v3.0) with the A1 array. Genotyping was done using GRCh37 human genome reference. Data preprocessing and quality control analysis were separately performed for each genotyping batch of samples at IMIDomics, Inc. (Barcelona, Spain) and at Erasmus MC (Rotterdam, Netherlands). Quality control analysis was performed using PLINK software (versions 1.9 and 2). In the SCGT00 and SCGT00val quality control analysis, we identified autosomal single-nucleotide polymorphisms (SNPs), and, using those SNPs from chromosome X, we confirmed the consistency between SNP-estimated and clinically reported genders. Then, we quantified the percentage of SNPs with a minor allele frequency (MAF) higher than 5%. Next, we computed the percentage of missingness both at the SNP-wise and sample-wise levels. Finally, we assessed the heterozygosity rate (F) of each sample in order to evaluate if any of the genotyped samples could be contaminated. In the SCGT02 Asthma+HC quality control analysis, we excluded samples with a SNP calling rate lower than 98%.
Patient genotypes (VCF format) were simplified by removing single-nucleotide variants (SNVs) that were unannotated (chr 0), located in the sexual Y (chr 24), pseudo-autosomal XY (chr 25) or mitochondrial (chr 26) chromosomes. As genotypes were obtained using the human hg19 reference genome, we converted their coordinates to the same reference genome used to mapped the sequencing reads (GRCh38) using the UCSC LiftOver tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver). LiftOver requires an input file in BED format. Thus, we used a Python script (https://github.com/single-cell-genetics/cellsnp-lite/blob/master/scripts/liftOver/liftOver_vcf.py) to convert our VCF file accordingly.
Library demultiplexing
Multiplexed libraries from SCGT00, SCGT00val and SCGT02 Asthma+HC cohorts were demultiplexed with Cellsnp-lite (version 1.2.2) in Mode 1a68, which allows us to genotype single-cell gene expression libraries by piling up the expressed alleles based on a list of given SNPs. To do so, we used a list of 7.4 million common SNPs in the human population (MAF > 5%) published by the 1000 Genomes Project consortium and compiled by the authors (https://sourceforge.net/projects/cellsnp/files/SNPlist/). Then, we performed the donor deconvolution with vireo (version 0.5.6)69, which assigns the deconvoluted samples to its donor identity using known genotypes while detecting doublets and unassigned cells. Finally, we discarded detected doublets and unassigned cells before moving on to the downstream processing steps. For CellPlex libraries, we followed a joint deconvolution strategy combining cell multiplexing oligo (CMO) hashing and genotype-based deconvolution; we generated pools of cells belonging to different samples based on the individual SNPs and traced back to their donor of origin based on the CMO hashing. When no genotype is available, the use of this dual approach minimizes the discarded cells.
Data analysis
All analyses presented in this paper were carried out using mainly Python, unless specified otherwise. In particular, we structured our data in anndata objects70 compatible with SCANPY suite71, which allowed us to apply single-cell data processing and visualization best practices. All experiments and panels are reproducible with the code released in the project’s GitHub repository.
Data standardization
Considering the diversity of the datasets included in the reference of circulating immune cells, a standardization step was needed.
Cell barcodes
The ‘cellID’ barcodes assigned were inspired by The Cancer Genome Atlas (TCGA) project (https://docs.gdc.cancer.gov/Encyclopedia/pages/TCGA_Barcode/). Each barcode unequivocally identifies a cell, and it is composed of the studyID (project), libraryID (10x GEM channel), patientID, chemistry (only when 3′ and 5′ gene expression were available for the same sample), timepoint (if multiple observations were available for a patient) and the 10x Genomics cell barcode, respectively (for example, SCGT00_L046_P006.3P_T0_AAACCCAAGGTGAGAA).
Gene name harmonization
All datasets were mapped using human GRCh38 genome reference, but the annotation file version might differ, resulting in gene names with multiple aliases or deprecated symbols. To avoid gene redundancy or mismatching, we used Ensembl symbols instead of gene names. Then, for datasets without the Ensembl symbols, we compared all gene names with the HUGO Gene Nomenclature Committee (HGNC) database (latest version, February 2024; https://www.genenames.org/), in order to convert them to the latest official HUGO name, merging possible duplicates and retrieving the corresponding Ensembl symbol. For non-official genes, we used the MyGene Python interface (https://mygene.info/) to query the Ensembl symbol. Finally, we removed 16 genes categorized as ‘artifact’ or ‘TEC (To be Experimentally Confirmed)’.
Metadata harmonization
Patient metadata were unified across datasets, using common variable names and values for those present in multiple sources; specifically, we homogenized these variables of interest such as sex, age, disease, diseaseStatus, smokingStatus, ethnicity or institute. For instance, ‘M’, ‘Male’ and ‘Hombre’ entries were replaced with ‘male’. Additionally, we created a new variable, ‘binned_age’, to group patients within a range of 10 years, considering that, for the SCGT01, SCGT04 and SCGT11 datasets, the specific age information was not available. As detailed below, the datasets missing sex and age information were considered as data from unseen studies and used to evaluate the patient classifier.
Data splitting
Some studies in our cohort included patients with samples collected in multiple replicates, different timepoints or using different chemistry protocols. In studies with multiple replicates—that is, Zhang2022 and Terekhova2023—we selected samples with the largest number of cells. When multiple timepoints or disease statuses for the same patient are available—that is, Perez2022, COMBAT2022 and Ren2021—we kept only the samples associated with higher disease severity.
The filtered Inflammation Atlas cohort was then split in two datasets: CORE and unseen studies. Data from unseen studies include 86 samples and are used as an independent validation of our patient classifier pipeline. For this dataset, we selected studies that either involve diseases with a large support in our full cohort or lack metadata on sex and age. These chosen studies are as follows: SCGT00val, SCGT06, Palshikar2022, Ramachandran2019, Martin2019, Savage2021, Jiang2020, Mistry2019 and 10XGenomics.
After performing data quality control (removing low-quality libraries and cells; see section below), the CORE dataset includes 961 samples and was further split into Main and data from unseen patients, with 817 and 144 samples each, respectively. We first stratified samples based on the following metadata: studyID, chemistry and disease. From each of those groups, we randomly selected 20% of samples to be part of the unseen patients, provided that they amounted to at least five samples. In the patient classifier pipeline, the Main dataset is used as a reference, whereas data from unseen patients and unseen studies are used as a query dataset in two independent scenarios.
The Centralized Dataset included samples from SCGT00 and SCGT00val. Because all healthy patients were sequenced in the same pool, we did not take them into account. Then, because multiple samples were multiplexed and sequenced together, we split them, stratifying by sequencing patientPool to generate both the reference and the query datasets that include at least one pool for all the IMID diseases (that is, rheumatoid arthritis, psoriasis, psoriatic arthritis, Crohnʼs disease, ulcerative colitis and SLE). Further information about the samples classified in each group is detailed in Supplementary Table 1.
Data quality control
We performed data quality control on the CORE dataset by computing the main metrics (that is, library size, library complexity and percentage of mitochondrial, ribosomal, hemoglobin and platelet-related gene expression) on the count matrix. Metric distributions were visualized grouping cells by library (10x Genomics) and by considering their chemistry (3′ or 5′ and their version). Consequently, we removed low-quality observations using permissive thresholds, whereas the robust cleaning process was performed during cell annotation tasks. In particular, we initially excluded the low-quality libraries across datasets (<500 cells or <500 median genes recovered). Next, we removed low-quality cells with a very low number of unique molecular identifiers (UMIs) (<500) and genes (<250) or with a high percentage of mitochondrial expression (>25% for 3′ V3 and 20% for 3′ V2 and 5′), as it is indicative of lysed cells. Then, we removed barcodes with a high library size (>50,000 UMIs for 3′ V3 and 5′ V1, >40,000 UMIs for 5′ V2 or >25,000 UMIs for 3′ V2 chemistry) or with a high complexity (>6,000 genes for 3′ V3 and 5′ or >4,000 genes for 3′ V2 chemistry). After cell quality control, we also removed low-quality libraries (<250 cells), low-quality samples (<500 cells or <500 median genes recovered) as well as cells from a library if this patient recovered a low total number of cells (<50 cells). In addition, we eliminated genes that were detected in fewer than 20 cells in fewer than five patients, keeping a total of 22,838 genes. Lastly, we computed the cell cycle score using the gene list provided by the function cc.genes.updated.2019() from the Seurat library72 (version 4.3.0.1) and defined the different cell cycle ‘phases’ (G1, G2M and S). Before the dataset cleanup, we predicted doublets using the function scanpy.external.pp.scrublet() from the SCANPY library (version 1.9.8), which provides a score to flag putative doublets but without filtering them out at this stage. Consequently, during the clustering and annotation step, the clusters co-expressing gene markers from different lineage/population and high doublet score were assessed to determine whether a specific cluster could be classified as a group of doublets and subsequently excluded. After this step, the CORE dataset was split into Main and data from unseen patients, as explained above.
Quality control on the data from unseen studies was performed independently. We applied the same approach described above, but we filtered only poor-quality libraries and cells.
Data processing for annotation
Annotation strategy
To identify all the immune cell types and states present in the human blood, we employed a recursive top-down approach inspired by previous work done by Massoni-Badosa et al.73. Starting with 4,918,140 cells and 817 patients from the Main dataset, we divided the annotation into several stages. In brief, we first grouped all cells into the primary compartments within our study. Subsequently, each compartment was processed aiming to detect potential doublets, low-quality cells and cells resembling platelets or erythrocytes (cells with high expression of hemoglobin genes). Additionally, we also placed back some clusters of cells into their corresponding cell lineages, when wrongly clustered due to similar profiles (for example, T cells found in the natural killer cell group or vice versa). Then, we identified the clusters resembling specific biological cell profiles (cell subtypes), obtaining a final number of 64 different subpopulations, excluding Doublets and LowQuality_cells, that we defined as annotation Level 2. Those cell subtypes were grouped into 15 cell populations that we defined as annotation Level 1. At the end, our Inflammation Atlas contains 4,435,922 cells. For each group identified in the initial stage (cell lineages), we applied the following tasks: normalization, feature selection, integration, clustering and annotation. In the following, we will always refer to the parameters of the initial stage, and the specifics of the subsequent steps (from lineages to cell types), along with the annotation labels and the marker genes used to define them, can be found in Supplementary Table 3.
Data normalization
Following standard practices, filtered cells were normalized by total counts over all genes and multiplied by a scaling factor of 104 (scanpy.pp.normalize_total(target_sum = 104)). Then, the normalized count matrix X was log transformed as loge(X + 1) (scanpy.pp.log1p()).
Feature selection
Gene selection was performed by identifying the highly variable genes (HVGs). Before doing so, we excluded genes related to mitochondrial and ribosomal organules. Also, we skipped T cell and B cell receptor (TCR/BCR) genes, including joining and variable regions, because they are not useful to describe cell identities but, rather, to capture patient-specific clonally expanded cell populations within an inflammatory-related condition. Lastly, we excluded major histocompatibility complex (MHC) genes. To reduce the influence of a study’s specific composition and prevent biases in the gene selection task, we preferred genes that are highly variable in as many studies as possible. Therefore, similar to Sikkema et al.74, we first considered each study independently and computed the HVGs using the Seurat implementation75 (scanpy.pp.highly_variable_genes(min_disp = 0.3, min_mean = 0.01, max_mean = 4)). Then, we ranked genes based on the number of studies in which they are among the highly variable. Finally, for the initial stage, we determined the minimum number of studies required to compose an HVG list of more than 3,000 genes. Applying this strategy, we selected a total of 3,236 genes being highly variable in at least six studies. In the following steps, we requested more than 2,000 HVGs; the minimum number of studies required and the total of selected genes depend on the step and the cells under study. To identify red blood cell (RBCs) and platelets, we kept genes associated with erythrocytes, such as hemoglobin subunits, and pro-platelet basic protein (PPBP) (platelet related) in the HVG list. Because such genes are known to be related to ambient RNA when found in other cell types, we subsequently removed them after having annotated the above cell types.
Data integration
Our dataset includes single-cell data obtained from multiple studies including different chemistry protocols, inflammatory status samples and a broad range of other clinical features (for example, age and sex). Although this is a strength point of our atlas, such high levels of heterogeneity induced by technical confounding factors and unwanted biological variability resulted in challenging integration tasks before clustering and annotating cell populations. Therefore, we employed scVI14, a VAE approach that proves to be one of the most effective integration methods in complex scenarios, particularly when the annotation information is missing16. scVI takes as input the raw count matrix to generate an integrated, low-dimensional embedding space, where the cell states are preserved and the batch effects are reduced. Moreover, scVI’s embedding space can be exploited to cluster and annotate cells based on either known or cluster-specific marker genes. Details on the scVI parameters used in each annotation step can be found in Supplementary Table 7.
Cell clustering
To cluster cells into cell types with the Leiden algorithm, we first built the k-nearest neighbors (KNN) graph using scVI’s latent embeddings and k = 20 as the number of neighbors (scanpy.pp.neighbors(n_neighbors = k)). We then applied the Leiden algorithm using a resolution of r = 0.1 (scanpy.tl.leiden(resolution = r)). The k and r used in every other step for every lineage can be found in Supplementary Table 3.
Cell annotation
Cell clusters were manually annotated by immunology experts by comparing the expression levels of canonical gene markers. Moreover, the final step of annotation was performed using the cluster markers obtained performing a differential expression analysis among clusters (Supplementary Table 3). First, we ranked genes to characterize each cluster (scanpy.tl.rank_genes_groups()), by considering normalized RNA counts with the Wilcoxon rank-sum test. Then, we selected those genes with log2 fold change (log2FC) > 0.25 and false discovery rate (FDR) adjusted P < 0.05 and if they were present in at least 25% of cells. Notably, cells belonging to RBC and platelet populations were excluded from all the downstream analyses, except for label transfer performed as a step during patient classifier tasks (as explained below).
External annotation validation
We compared our independent annotations with the ones available in the largest public datasets. To quantify the overlap of cells among groups, we computed the adjusted Rand index (ARI) to measure the similarity between our label assignments and the ones performed by the original authors. Further details are available in Supplementary Table 2.
Centralized Dataset annotation
All previously described steps were applied to process and annotate the Centralized Dataset (SCGT00), with the following adjustments: (1) standard HVG selection was performed as the dataset included only a single study; (2) the dataset was integrated using ‘patientPool’ as the batch key; and (3) cell annotation was conducted up to Level 1, recovering the same cell types as in the main Inflammation Atlas, as this was necessary for the patient classifier. Here, starting with 855,417 cells and 152 patients included in the reference dataset, we recovered 15 cell populations (Level 1), excluding Doublets and LowQuality_cells. Details on the scVI parameters used in each annotation step can be found in Supplementary Table 7, whereas details on the clustering and annotation steps are provided in Supplementary Table 3.
Feature selection after annotation
Gene selection
To improve the quality of downstream analysis to characterize the inflammation landscape, it is necessary to perform a gene selection in order to remove dataset-specific genes and reduce the batch effect. First, we performed data normalization (as described above), kept only the genes that are expressed (raw count > 0) in at least one cell in each study and removed genes associated with mitochondrial, ribosomal, TCR/BCR, MHC, hemoglobin and platelet cell types. This step retained a total of 14,127 genes. Then, we identified three sets of genes: (1) the HVGs, (2) the differentially expressed genes (DEGs) between healthy and each inflammatory status and (3) Cytopus76, a manually curated immune-specific gene list.
HVGs
Similar to the feature selection approach described in the annotation section, we selected a total of 3,283 HVGs, by using a threshold of at least 3,000 genes. In practice, we first ranked the genes based on the number of studies in which they are concurrently highly variable (scanpy.pp.highly_variable_genes(min_disp = 0.3, min_mean = 0.01, max_mean = 4, batch_key=’libraryID’)) and then chose a minimum number of studies of five.
DEGs between healthy and each disease
We obtained a list of DEGs after grouping single-cell gene expression profiles into pseudobulks. Therefore, we first combined the expression profiles of individual cells to produce pseudobulks for every patient and cell type (Level 1), removing groups with no more than 20 cells, using the Python implementation of decoupleR77 (version 1.6.0) (decoupler.get_pseudobulk(min_cells = 20, sample_col=’sampleID’, groups_col = ‘Level1’, layer = ‘counts’, mode = ‘sum’)). Then, we applied the edgeR (version 4.0.16) quasi-likelihood functions to search for DEGs between healthy patients and each other’s inflammatory conditions, by considering one cell type at a time. Because not all the cell types were detected in each patient, we did not perform the pairwise comparison if one disease had fewer than three pseudobulks. More in detail, for each pairwise comparison, we first removed genes with a low expression value (filterByExpr(y, group=disease)). Second, we normalized by library size the aggregated raw counts (calcNormFactors(y, logratioTrim = 0.3)). Third, we corrected for the main confounding factors—that is, chemistry protocol, sex and binned age—considering an additive model. One patient was excluded from the analysis due to missing age information. We defined the design of our comparison using the following patsy-style (https://patsy.readthedocs.io/en/latest/formulas.html) formula: ‘~0 + C(disease) + C(chemistry) + C(sex) + C(binned_age)’. Fourth, we estimated a negative binomial dispersion for each gene using estimateDisp(), which we fed into a gene-wise negative binomial generalized linear model (glmQLFit(robust = TRUE)) to test for DEGs with a quasi-likelihood F-test (glmQLFTest()). Lastly, results obtained from each comparison were merged together, and the F-test P values were corrected using the Benjamini−Hochberg FDR procedure implemented in R (p.adjust(method = ‘BH’)). Given the corrected P values and the log2FC, we selected 6,868 DEGs with P < 0.01 and absolute log2FC > 1.5.
Curated immune-specific genes
To be able to track the full spectrum of inflammatory processes, including immune activation and progression, we curated nine inflammation-related functions defined in the literature78,79,80,81,82,83,84 (1,364 genes present in our dataset; Supplementary Table 4) and complemented them with a published list of cell-type-specific signatures derived from immunological knowledge based on single-cell studies (Cytopus76). Specifically, we retrieved all global gene sets for the leukocyte category and the following inflammatory-related cell-type-specific factors: naive and non-naive CD4 T cells (CD4T_TFH_UP, CD4T_TH1_UP, CD4T_TH2_UP, CD4T_TH17_UP, Tregs_FOXP3_stabilization); naive and non-naive CD8 T cells (CD8T_exhaustion, CD8T_tcr_activation); B cells (B_effector); monocytes (IFNG response, IL4-IL13 response); and dendritic cells (dendritic cell antigen crosspresentation).
Aggregation of gene sets
We generate the relevant gene set by doing the union of HVGs, DEGs and the manually curated list. The final number of unique genes is 8,253.
Dataset integration and gene expression correction via scANVI
Atlas-level analysis requires a careful preprocessing of the gene expression profiles to deal with the heterogeneity of the studies, the batch effect and the missing or noisy observations. scANVI15 is one of the existing methods capable of addressing these challenges and has been proven effective on atlas-level benchmarks compared to other integration methods. We validated its performance on our data by using the metrics from the scib-metrics package16 (version 0.5) (Extended Data Fig. 1).
scANVI integration
scANVI is an extension of the scVI model, employed previously for data integration, that also leverages the information of the cell type annotation. We first trained an scVI VAE (scvi.model.SCVI) and then trained scANVI (scvi.model.SCANVI) starting from the pretrained scVI model (see parameters in Supplementary Table 7). Both models corrected for the chemistry batch while also considering libraryID, studyID, sex and binned age as covariates. After training, we generated the normalized corrected counts by sampling from scANVI’s negative binomial posterior (SCANVI.get_normalized_expression). The batch effect was mitigated by sampling and averaging each cell’s expression as if it originated from each chemistry protocol by setting the transform_batch parameter to the list of chemistry protocols present in our atlas.
Comparison of cell type composition
To estimate the changes in the proportions of cell populations across conditions, we applied the scCODA package85 (version 0.1.9), a Bayesian modeling tool that takes into account the compositional nature of the data to reduce the risk of false discoveries. scCODA allows us to infer changes between conditions while considering other covariates, corresponding to the disease status in our setting. scCODA searches for changes between a reference cell type, assumed to be constant among different conditions, and the other cell types. We selected as the reference population the one that showed the lower variance across conditions, excluding rare cell populations (that is, progenitors, plasmacytoid dendritic cells and cycling cells). This resulted in the selection of dendritic cell as the reference cell type for all diseases. scCODA takes as input the count of cells belonging to each cell type in each patient and returns the list of cell type proportion changes with the corresponding corrected P values (through the FDR procedure). A patsy-style formula was used to build the covariate matrix, specified with ‘healthy’ as baseline and sex and binned age as covariates (C(disease, Treatment(‘healthy’)) + C(sex) + C(binned_age)), because we are interested in detecting changes between a normal and a diseased status. We reported only changes with a corrected P < 0.05 and a log2FC > 0.2.
Comparison of gene expression profiles
Gene factor inference
To expand the list of curated immune-related genes following a data-driven approach, we employed Spectra22 (version 0.2.0), a matrix factorization algorithm that enables us to identify a minimal set of genes related to specific functions in the data—that is, factors. Spectra takes as input cell type labels to infer global and cell-type-specific factors that decompose the overall gene expression matrix and each cell type submatrices, respectively. Given our list of curated gene sets, we considered the Cytopus ones as global factors, whereas we regarded all the remaining as cell-type-specific factors. The Spectra model was fitted with default parameters with the exception of λ, which was set equal to 0.001. Considering the prohibitive computational resources required for applying Spectra on our single-cell data, we fed the algorithm with the metacell aggregated expression matrix, as described in the paragraph below. Spectra returned a list of 135 factors that are a linear combination of the gene expression from the original matrix. The coefficients included in the matrix can be then used as a proxy of the gene relevance in a given factor.
Metacell generation
We generated metacells using SEACells86 (version 0.3.3), which aggregates cells by exploiting their distances in a low-dimensional embedding space. Starting from the normalizing data, we selected the top 3,000 HVGs using the highly_variable_genes function in SCANPY, with the Seurat flavor. To define SEACells’ input embedding space, we calculated the first 50 principal components and selected those principal components that explain 90% of the total variance observed. To avoid biases due to batch effect and other confounding factors, we executed SEACells for each sample independently. In particular, we generated a number of metacells equal to the number of cells of each patient divided by 50. We further filtered the obtained metacells by computing the proportion of the most abundant annotation label (Level 1) in each SEACells group and then removed the ones with a purity lower than 0.75. Overall, we defined 71,108 metacells. Given the assignment of cells to each metacell, we generated each metacell’s gene expression profile by averaging the corresponding cells’ scANVI normalized and corrected expression profiles. Because scANVI returns counts sampled from a negative binomial distribution, we also log scaled the obtained metacell profiles.
Inflammation-related signature definition
Spectra provided a total of 135 factors that include a refined gene list for each gene set we used as input. Thus, we need to assign those factors to our original gene sets for retrieving the corresponding biological function. For doing so, we performed enrichment analysis with ULMs available in the Python implementation of decoupleR77, to estimate the factors associated with each gene set. The gene coefficients returned by Spectra were considered as the response variable and a vector of weights (1 if the genes were included in the gene set, 0 otherwise) serving as explanatory variables. ULM returns an estimate and a P value for each enrichment. We corrected those P values for multiple comparison by computing the FDR with the Benjamini−Hochberg procedure, implemented in the scipy (version 1.12.0) library. We kept 125 factors with a positive estimate and an adjusted P value < 0.05. Finally, we assign to each factor the biological function that corresponds to the gene set that provided the highest estimated score.
Inflammation-related signature scores
To compare immune-relevant activation profiles across diseases and cell types, we applied an enrichment signature scoring procedure, considering the factors obtained with Spectra22. First, we generated pseudobulks stratified by cell type (Level 1 or Level 2) and patients, discarding groups with fewer than 10 cells. We averaged the scANVI-corrected gene expression matrix of each cell belonging to a given pseudobulk and then log transformed and scaled the expression values to zero mean and unit variance, to reduce the impact of highly expressed genes. We fitted decoupleR’s ULM77 by considering pseudobulk expression profiles as the response variable and the gene coefficient returned by Spectra as the explanatory one. We assessed the scores for the 119 cell-type-specific factors only in their corresponding cell type. The output of the model is a Studentʼs t-statistic for each combination of pseudobulk and factor, which is used as a proxy for the corresponding biological function activity: positive values are associated with more active functions in a given sample and vice versa. To identify the upregulated or downregulated biological functions across inflammatory conditions, we compared the activation score between healthy and each disease, considering only comparisons that include at least three observations in both conditions. To take into account the batch effect induced by studies and chemistry protocols that still affects the data (Extended Data Fig. 1b,c; scbi metrics and principal component analysis (PCA)), we applied a LMEM. In particular, we fitted the function mixedlm() from the statsmodels Python library (version 0.14.1) with the following formula:‘Q(‘{factor}’) ~ C(disease, Treatment(reference = ‘healthy’)) + ‘f’C(chemistry)’, grouping by ‘studyID’. We corrected the P value obtained for multiple testing using FDR considering all the comparisons when tested at Level 1 and within each Level 1 population when tested at Level 2.
GRN analysis
Pseudobulk matrices were calculated by averaging the corrected and standardized count matrices by cell type and sample. We compute differential expression analysis for each cell type in each disease using healthy individuals as reference. LMEMs were used to model the expression levels of each gene independently, considering the disease as a fixed effect while modeling the ID of the study as a random effect. We used the mixedlm() function of the statsmodels (version 0.14.0) Python package to run the analysis. To associate each cell-type-specific ‘IFN-induced’ factor with a given transcription factor regulator, we integrated these signatures with the CollecTRI Gene Regulatory Network87 by matching target genes to identify common genes between transcription factor regulons and Spectra signatures. Therefore, each ‘IFN-induced’ signature was thus linked to a subset of transcription factor regulons. The activity of each transcription factor was calculated using only the common genes between each transcription factor and each signature, employing the UML from decoupleR77 and the z-values obtained from the differential expression analysis. To ensure robustness, only regulons with at least 10 gene targets were considered. This pipeline was applied across ‘IFN-induced’ factors and diseases, focusing on the activity in the cell type where the Spectra signature was identified. Negative activities (t-stat < 0) and non-significant results (P > 0.05) were filtered out. This analysis identified STAT1 and SP1 as the sole transcription factor regulators of the defined cell types. We performed one-versus-all Wilcoxon rank-sum tests to compare transcription factor activity across Level 2 subpopulations within each Level 1 lineage for SLE and Flu. For each transcription factor (SP1 and STAT1), activity within a given Level 2 state was compared to all other states within the same Level 1 compartment. Tests were two-sided and restricted to comparisons with at least three observations per group. P values were adjusted using the Benjamini−Hochberg method. The same approach was applied to monocytes, comparing transcription factor activity across diseases within each Level 2 monocyte subset.
For the comparison of flare and non-flare patients from SLE, non-corrected log1p-normalized single-cell expression matrix from Perez et al.17 was used to further investigate SP1 and STAT1 regulon activities across both categories. Pseudobulk profiles were calculated by averaging by cell type, considering only cell types (Level 2) with a minimum of 10 cells and groups that include at least three patients in both conditions (flare versus non-flare). Prior to calculating transcription factor activities across samples, we standardized the gene expression data on patients with SLE based on healthy individuals. Specifically, for each gene, the mean and standard deviation were calculated from the healthy group, and these statistics were then used to scale the gene expression values across patients with SLE. Only gene targets identified in the previous step were used to calculate enrichment using the ULM method. Finally, the activity of STAT1 and SP1 was calculated at Level 2 using CollecTRI87.
Immune gene importance evaluation
In this section, we introduce our pipeline used to obtain a gene importance metric by interpreting cell-type-specific classifiers for disease prediction. All the steps described below were carried out separately for each cell type (excluding RBCs, platelets, progenitors and cycling cells). Specifically, the classification task was performed with GBDTs implemented in the XGBoost library88 (py-xgboost-gpu: version 2.0.3). Furthermore, interpretability was performed using SHAP values35 (version 0.45.1), a powerful approach assigning an importance to each gene by also taking into account their interactions.
Feature selection
To focus our analysis on cell-type-specific inflammatory-related signatures, we considered only genes relevant in annotated Spectra factors, and we further reduced the list by removing cell identity genes (for example, CD3E and MS4A1) as well as non-protein-coding genes. This filtering gave a final number of 935 genes.
Data processing
We split our data into three parts: the training set and the validation and testing set, used for hyperparameter tuning and performance evaluation, respectively. We balanced the splits by disease, ensuring that each sample’s cells were included in the same set. Initially, we partitioned the data into five splits using the function sklearn.model_selection.StratifiedGroupKFold. Three of these splits were assigned to the training set, and one was designated for validation and one for testing. Accounting for both stratification by disease and patient partitioning might lead to an uneven distribution of cells among diseases. To address this, we assigned splits with a well-balanced distribution of cells to the training and testing sets first.
XGBoost fitting
XGBoost (xgboost.XGBClassifier) hyperparameters were tuned using the Optuna library89 (version 3.6.0). The performance of each model configuration was estimated using the WF1 score on the validation set. To reduce the computational cost, we both pruned unpromising hyperparameters and early stopped the training when no improvement was achieved more than 20 steps before the upper bound of 1,500. We considered 50 configurations of XGBoost, taking into account the hyperparameters detailed in Supplementary Table 7. Using the best configuration and its corresponding number of training steps (equivalent to the number of estimators), we retrained the best model on the union of the training and validation sets. This time, we did not apply early stopping and increased the number of training steps by 20%, to account for the larger number of training samples.
d-SHAP interpretability
To interpret the decision of the selected XGBoost classifier, we employed the widely adopted Shapley values through the SHAP library. SHAP values were computed with shap.TreeExplainer using the observational ‘three_path_dependant’ approach. Given the potential resource-intensive nature of handling all SHAP values for every cell and disease, especially in terms of storage, we computed their mean and variance across all samples in batches using the Weldford online algorithm90. Given a specific cell type ct, we have a SHAP value for every gene in every cell and for each disease: a matrix of real values \(\mathrm{SHAP}^{\mathrm{ct}}(c,g,d)\), where c, g and d identify the cell, gene and disease, respectively. The average contribution of a gene g for a disease d can be computed as \({\mathrm{d{\text-}\mathrm{SHAP}}^{\mathrm{ct}}(g,d)=\mathrm{mean}_{c\in C}{| \mathrm {SHAP}}^{\mathrm{ct}}(c,g,d)|}\), where C is the set of cells. To aggregate the d-SHAP values across multiple diseases—for example, the ones included in the same study—we summed their values across genes.
Gene selection
To validate our ensemble of important genes through d-SHAP values, we tested if our selection generalizes to unseen studies. First, we defined a gene set GS that included genes expressed in at least 5% of the cells. Then, for each of the eight conditions included in unseen studies, we selected from GS the top k ranked genes by d-SHAP importance. We then trained XGBoost in a nested cross-validation setting on data from unseen studies, where we performed both hyperparameter tuning and performance evaluation, using only one of the gene sets as input features. Next, we computed WF1 and BAS to test the performance considering k = 5, k = 10 and k = 20. Given our selection S of genes, with size |S|, we also tested 20 sets of |S| randomly selected genes from GS, excluding the ones in S (that is, not top ranked according to d-SHAP). Lastly, we compared the performances of the models trained on each gene set against XGBoost trained on the whole set of genes GS. The analysis was repeated for each cell type independently.
Study classifier and s-SHAP values
To identify whether the gene importance is driven by the study batch effects, we trained a separate classifier to predict study instead of the disease. Feature selection, data processing and model fitting were done in the same way as explained above for disease classification, apart from the data split where cells were stratified by study instead of disease. SHAP values were computed for each of the cell-type-specific study classifiers, resulting in an average contribution of a gene g for a study s \({\mathrm{s{\text-}\mathrm{SHAP}}^{\mathrm{ct}}(g,s)=\mathrm{mean}_{c\in C}{|\mathrm{SHAP}}^{\mathrm{ct}}(c,g,s)|}\). Because diseases can be associated with multiple studies, we aggregated the s-SHAP values for study prediction by summing them across all studies that include the selected disease. This allowed us to compare the batch-related signal (s-SHAP) with the disease-related signal (d-SHAP).
Patient classifier pipeline
In this section, we describe the pipeline used to validate the Inflammation Atlas as a diagnostic tool. In the following analysis, the terms ‘patient’ and ‘sample’ are equivalent, because, after data splitting, we kept only one sample for each patient. The pipeline consists of (1) integrating an annotated reference dataset with data integration tools that provide batch-corrected embeddings, (2) mapping a query dataset into the reference to obtain its corrected embeddings, (3) transfering the cell annotation labels from the reference, (4) defining a patient embedding space and (5) training a classifier to predict the patient conditions from the embeddings.
Starting from a large annotated reference dataset, we applied four state-of-the-art integration methods, described below, to obtain a batch-corrected embedding. We considered different chemistry protocols as the main source of batch effect; thus, we corrected for the chemistry covariate. Then, an independent query dataset was mapped into the corrected embeddings. This step provides both batch correction and allows us to transfer cell annotation labels from the reference to the query dataset. To define patient-wise embeddings, we averaged each patient’s cell embeddings by cell type, resulting in an embedding for each cell type and each patient (30 embedded dimensions for scANVI main configuration; see Supplementary Table 7 for all the methods and configurations). To predict the inflammatory conditions of the patients in the independent query dataset, we fit one classifier for each cell type on the reference patient embeddings. Then, we predicted the inflammatory condition of the query patients by returning the most frequent condition among the predictions of every cell-type-specific classifier.
We validated our pipeline considering three different settings. In the first one, we performed a cross-validation on the Main dataset, where each left-out split is considered as a query dataset and the remaining as the reference. Moreover, we tested our diagnostic tool on data from unseen patients and unseen studies, this time using the whole Main as a reference.
Integration methods
In this section, we explain each data integration method, and the tested configurations of hyperparameters can be found in Extended Data Fig. 8 and Supplementary Table 7. Note that the scGen and Harmony/Symphony approaches generate one integrated dataset that is independent from the query data, whereas scANVI and scPoli require a fine-tuning of the reference model for a given query dataset.
scGEN
scGen is defined by two main components: a VAE and a latent space arithmetic method. The VAE estimates a posterior distribution of latent variables through the encoder, from which we can reconstruct the expression matrices via the decoder (scGen_model.batch_removal()). Similar to commonly employed VAEs, scGen approximates the posterior through a variational distribution, modeled by the encoder and defined as a multivariate Gaussian. When the scGen’s VAE has been fitted on the reference dataset, latent space arithmetic is employed to correct for the batch effect induced by the chemistry protocol used. Within each cell type, scGen first selects the mean \({\mu }_{\max }\) of the most populated batch and then corrects each batch with mean \({\mu }_{0}\) by adding \({\delta =\mu }_{\max }-{\mu }_{0}\) to each cell’s embedding. Importantly, the cell type has to be inferred when not known. The final corrected count matrix will correspond to the generated count matrix from the arithmetic-corrected embeddings. Following scGen’s tutorials, we will refer as corrected embeddings to the ones obtained given the corrected expression matrix as input. To apply scGen batch correction on the query dataset, we need to also infer the cell types of those cells. This step was performed through label transfer by nearest neighbors, following a similar approach employed in Human Lung Cell Atlas74 and introduced in ref. 45. The idea is to employ (approximate) nearest neighbors through PyNNDescent91 (version 0.5.11) (pynndescent.NNDescent().prepare()) and infer the most probable cell type in the 10 nearest neighbors (pynndescent.NNDescent().query()) from the already annotated cells in the reference dataset. To account for the shape of the distribution of the neighbors, a Gaussian kernel was applied instead of using the Euclidean distance. The most probable nearest neighbor cell type is then assigned to annotate new cells.
scANVI
We first trained scVI and scANVI on the reference dataset, like the dataset integration described before, and then we fine-tuned it to the query dataset. Regarding the label transfer, we employed the scANVI predict() function with default parameters.
Harmony and Symphony
Harmony92 and Symphony93 are two related methods that integrate a reference and map a query dataset to it, respectively. Harmony takes a PCA embedding of cells as input, along with their batch covariates (chemistry). Next, the model represents cell states as soft clusters, where each cell identity is defined as a probabilistic assignment across clusters, with the aim of maximizing diversity among batches within those clusters. Cells are iteratively assigned soft-cluster memberships; those assignments are used as weights in a linear mixture model to remove confounding factors. The result is a new batch-corrected embedded space. The Symphony algorithm starts from the linear model parameters inferred by Harmony to map query cells onto the corresponding embedding space. First, it projects the query gene expression profiles into the same uncorrected low-dimensional space as the reference cells. Next, Symphony computes soft-cluster assignments for the query cells based on their proximity to the reference cluster centroids. Finally, Symphony employs the Harmony mixture model components to estimate and regress out batch effects from the query data. Importantly, the reference cell embedding remains stable during this mapping process. We transferred annotation labels from the reference to the query dataset by exploiting cell proximity in the embedding space using nearest neighbors through sklearn.classifier.KNeighborsClassifier, Symphony default choice (https://github.com/potulabe/symphonypy).
scPoli
In contrast to other integration methods such as scANVI, scPoli94 encodes the condition (chemistry and sampleID) as a learnable conditional embedding and characterizes each cell type as a prototype in the latent embedding to facilitate the label transfer. In the reference building phase, we first pretrained the model given the reference dataset and its conditions and then fine-tuned to optimize the prototypes. In the reference mapping phase, we froze the model and learned the new conditional embeddings belonging to the query dataset. The label transfer is performed by simply assigning the cell type belonging to the closest prototype in the latent embedding space. All the methods belong to the scArches45 class scarches.models.scPoli.
Disease classifiers
Patient embeddings definition
After obtaining the corrected embedding from one of the data integration approaches described previously, we need to aggregate the cell-wise embeddings into patient-wise embeddings. We decided to group at the level of the cell types by computing the mean embedding across cells belonging to the same cell type and sampleID. Only for scPoli, we generated three different types of patient embeddings: the learned patient embeddings (sample), the averaged cell-wise latent embeddings (cell) and the concatenation of the two (cell&sample).
Classifiers definition and hyperparameter tuning
In this phase, the aim is to train a classifier for each cell type on the patient-wise embeddings belonging to the reference dataset. We tested the following classifier types: sklearn.svm.LinearSVC, sklearn.svm.SVC and sklearn.neighbors.KNeighborsClassifier (sklearn version 1.4.1.post1). For each classifier type, we trained different configurations defined in Supplementary Table 7 and evaluated their performance using a five-fold cross-validation on the reference patient embeddings. Similar to what we did to optimize the XGBoost classifier when estimating the immune gene importance, we employed the Optuna library to perform the hyperparameter tuning for each classifier. The best hyperparameter combination was selected according to the WF1 score independently of the cell type.
Majority voting and evaluation
The best classifier type according to the average performance over all cell types is then used to train from scratch the corresponding classifier on the whole reference patient embedding. The predicted condition (disease) for a patient is simply the majority voting among the classifiers. In case of a tie of different conditions, we conservatively rejected the prediction of the classifiers. Then, the overall metrics WF1 score, BAS and Matthews correlation coefficient (MCC) and the disease-wise metrics Precision, Recall, BAS and F1 score were computed by comparing the predicted inflammatory conditions by each classifier type in the query dataset with the available ground truth. All those metrics were computed with the sklearn Python library. When we refer to the weighted version of a given metric, we are using average = ’weighted’ parameter to take into account the unbalance of the inflammatory condition observations.
Note, if a given query patient does not have any cells annotated for a given cell type, the corresponding prediction was set as ‘Not Available’. This label was not taken into account during the majority voting procedure and was considered as a wrong prediction when evaluating the performances of that cell type.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
scRNA-seq in-house-generated raw data and associated processed count matrices are publicly accessible without restrictions at the NCBI GEO database under the GSE248688 SuperSeries, including the following SubSeries accession numbers: GSE248689 (SCGT01), GSE248695 (SCGT02), GSE248685 (SCGT03), GSE248693 (SCGT04) and GSE270165 (SCGT06). To ensure data safety and patient privacy, raw scRNA-seq data from SCGT00, SCGT00val and SCGT05 studies can be downloaded upon reasonable request through the European Genome-phenome Archive (EGA) database using the following access codes: EGAC50000000566 (SCGT00 and SCGT00val) and EGAS50000000590 (SCGT05).
Previously published scRNA-seq data included in this project, either FASTQ files or processed count matrices, were obtained from GEO, BioStudies Array Express, Broad Institute DUOS, Synapse, Genome Sequence Archive, CELLxGENE Data Portal and 10x Genomics. Further details are specified in Supplementary Table 1 (Sheet 1).
The processed scRNA-seq datasets (quality controlled gene expression count matrices) and metadata analyzed in the present study have been deposited at Zenodo: https://doi.org/10.5281/zenodo.14851901.
Code availability
The code to reproduce the full analysis presented in this article is hosted in the GitHub repository: https://github.com/Single-Cell-Genomics-Group-CNAG-CRG/Inflammation-PBMCs-Atlas.
Change history
26 January 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41591-026-04223-x
References
Medzhitov, R. The spectrum of inflammatory responses. Science 374, 1070–1075 (2021).
Casanova, J.-L. & Abel, L. Mechanisms of viral inflammation and disease in humans. Science 374, 1080–1086 (2021).
Medzhitov, R. Origin and physiological roles of inflammation. Nature 454, 428–435 (2008).
Netea, M. G. et al. A guiding map for inflammation. Nat. Immunol. 18, 826–831 (2017).
Roe, K. An inflammation classification system using cytokine parameters. Scand. J. Immunol. 93, e12970 (2021).
Hughes, C. E. & Nibbs, R. J. B. A guide to chemokines and their receptors. FEBS J. 285, 2944–2971 (2018).
Soehnlein, O. & Libby, P. Targeting inflammation in atherosclerosis—from experimental insights to the clinic. Nat. Rev. Drug Discov. 20, 589–610 (2021).
Psarras, A., Wittmann, M. & Vital, E. M. Emerging concepts of type I interferons in SLE pathogenesis and therapy. Nat. Rev. Rheumatol. 18, 575–590 (2022).
Manthiram, K., Zhou, Q., Aksentijevich, I. & Kastner, D. L. The monogenic autoinflammatory diseases define new pathways in human innate immunity and inflammation. Nat. Immunol. 18, 832–842 (2017).
Cao, L. L. & Kagan, J. C. Targeting innate immune pathways for cancer immunotherapy. Immunity 56, 2206–2217 (2023).
Dinarello, C. A., Simon, A. & van der Meer, J. W. M. Treating inflammation by blocking interleukin-1 in a broad spectrum of diseases. Nat. Rev. Drug Discov. 11, 633–652 (2012).
Țiburcă, L. et al. The treatment with interleukin 17 inhibitors and immune-mediated inflammatory diseases. Curr. Issues Mol. Biol. 44, 1851–1866 (2022).
Dann, E. et al. Precise identification of cell states altered in disease using healthy single-cell references. Nat. Genet. 55, 1998–2008 (2023).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Xu, C. et al. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Mol. Syst. Biol. 17, e9620 (2021).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Perez, R. K. et al. Single-cell RNA-seq reveals cell type−specific molecular and genetic associations to lupus. Science 376, eabf1970 (2022).
Ju, J. K. et al. Activation, deficiency, and reduced IFN-γ production of mucosal-associated invariant T cells in patients with inflammatory bowel disease. J. Innate Immun. 12, 422–434 (2020).
Adlowitz, D. G. et al. Expansion of activated peripheral blood memory B cells in rheumatoid arthritis, impact of B cell depletion therapy, and biomarkers of response. PLoS ONE 10, e0128269 (2015).
Drewry, A. M. et al. Persistent lymphopenia after diagnosis of sepsis predicts mortality. Shock 42, 383 (2014).
Moir, S. & Fauci, A. S. B cells in HIV infection and disease. Nat. Rev. Immunol. 9, 235–245 (2009).
Kunes, R. Z., Walle, T., Land, M., Nawy, T. & Pe’er, D. Supervised discovery of interpretable gene programs from single-cell data. Nat. Biotechnol. 42, 1084–1095 (2024).
Ishina, I. A. et al. MHC class II presentation in autoimmunity. Cells 12, 314 (2023).
Thomas, T. et al. A longitudinal single-cell atlas of anti-tumour necrosis factor treatment in inflammatory bowel disease. Nat. Immunol. 25, 2152–2165 (2024).
Feng, X. et al. Interferon-β corrects massive gene dysregulation in multiple sclerosis: short-term and long-term effects on immune regulation and neuroprotection. eBioMedicine 49, 269–283 (2019).
Reyes, M. et al. An immune-cell signature of bacterial sepsis. Nat. Med. 26, 333–340 (2020).
Raphael, I., Joern, R. R. & Forsthuber, T. G. Memory CD4+ T cells in immunity and autoimmune diseases. Cells 9, 531 (2020).
Andreou, N.-P., Legaki, E. & Gazouli, M. Inflammatory bowel disease pathobiology: the role of the interferon signature. Ann. Gastroenterol. 33, 125–133 (2020).
Regis, G., Pensa, S., Boselli, D., Novelli, F. & Poli, V. Ups and downs: the STAT1:STAT3 seesaw of interferon and gp130 receptor signalling. Semin. Cell Dev. Biol. 19, 351–359 (2008).
Aue, A. et al. Elevated STAT1 expression but not phosphorylation in lupus B cells correlates with disease activity and increased plasmablast susceptibility. Rheumatology 59, 3435–3442 (2020).
Szabó, E. et al. Identification of immune subsets with distinct lectin binding signatures using multi-parameter flow cytometry: correlations with disease activity in systemic lupus erythematosus. Front. Immunol. 15, 1380481 (2024).
Olingy, C. E., Dinh, H. Q. & Hedrick, C. C. Monocyte heterogeneity and functions in cancer. J. Leukoc. Biol. 106, 309–322 (2019).
Bentéjac, C., Csörgő, A. & Martínez-Muñoz, G. A comparative analysis of gradient boosting algorithms. Artif. Intell. Rev. 54, 1937–1967 (2021).
Ahern, D. J. et al. A blood atlas of COVID-19 defines hallmarks of disease severity and specificity. Cell 185, 916–938 (2022).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In Proc. 31st International Conference on Neural Information Processing Systems 4768−4777 (Curran Associates, 2017).
Anderson, A. E. et al. Expression of STAT3-regulated genes in circulating CD4+ T cells discriminates rheumatoid arthritis independently of clinical parameters in early arthritis. Rheumatology 58, 1250–1258 (2019).
Rönnblom, L. & Leonard, D. Interferon pathway in SLE: one key to unlocking the mystery of the disease. Lupus Sci. Med. 6, e000270 (2019).
Zhang, L., Yu, L., Li, J., Li, Z. & Zhao, X. Novel compound heterozygous CYBAmutations causing neonatal-onset chronic granulomatous disease. J. Clin. Immunol. 43, 1131–1133 (2023).
Denson, L. A. et al. Clinical and genomic correlates of neutrophil reactive oxygen species production in pediatric patients with Crohn’s disease. Gastroenterology 154, 2097–2110 (2018).
Jarmakiewicz-Czaja, S., Ferenc, K. & Filip, R. Antioxidants as protection against reactive oxidative stress in inflammatory bowel disease. Metabolites 13, 573 (2023).
Diamond, M. S. & Farzan, M. The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nat. Rev. Immunol. 13, 46–57 (2013).
Wen, L., Krauss-Etschmann, S., Petersen, F. & Yu, X. Autoantibodies in chronic obstructive pulmonary disease. Front. Immunol. 9, 66 (2018).
Seguí, J. et al. Superoxide dismutase ameliorates TNBS-induced colitis by reducing oxidative stress, adhesion molecule expression, and leukocyte recruitment into the inflamed intestine. J. Leukoc. Biol. 76, 537–544 (2004).
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat. Rev. Genet. 24, 550–572 (2023).
Lotfollahi, M. et al. Mapping single-cell data to reference atlases by transfer learning. Nat. Biotechnol. 40, 121–130 (2022).
Clark, I. C. et al. HIV silencing and cell survival signatures in infected T cell reservoirs. Nature 614, 318–325 (2023).
Zuroff, L. et al. Immune aging in multiple sclerosis is characterized by abnormal CD4 T cell activation and increased frequencies of cytotoxic CD4 T cells with advancing age. EBioMedicine 82, 104179 (2022).
Edahiro, R. et al. Single-cell analyses and host genetics highlight the role of innate immune cells in COVID-19 severity. Nat. Genet. 55, 753–767 (2023).
Domínguez Conde, C. et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, eabl5197 (2022).
Sanyal, A. J. et al. Diagnostic performance of circulating biomarkers for non-alcoholic steatohepatitis. Nat. Med. 29, 2656–2664 (2023).
van Steenbergen, H. W., Cope, A. P. & van der Helm-van Mil, A. H. M. Rheumatoid arthritis prevention in arthralgia: fantasy or reality? Nat. Rev. Rheumatol. 19, 767−777 (2023).
Feld, L., Glick, L. R. & Cifu, A. S. Diagnosis and management of Crohn disease. JAMA 321, 1822–1823 (2019).
Litinetskaya, A. et al. Multimodal weakly supervised learning to identify disease-specific changes in single-cell atlases. Preprint at bioRxiv https://doi.org/10.1101/2024.07.29.605625 (2024).
Szałata, A. et al. Transformers in single-cell omics: a review and new perspectives. Nat. Methods 21, 1430–1443 (2024).
Jiang, Y. et al. Single cell RNA sequencing identifies an early monocyte gene signature in acute respiratory distress syndrome. JCI Insight 5, e135678 (2020).
Cillo, A. R. et al. Immune landscape of viral- and carcinogen-driven head and neck cancer. Immunity 52, 183–199 (2020).
Zhang, C. et al. Single-cell RNA sequencing reveals intrahepatic and peripheral immune characteristics related to disease phases in HBV-infected patients. Gut 72, 153–167 (2023).
Schafflick, D. et al. Integrated single cell analysis of blood and cerebrospinal fluid leukocytes in multiple sclerosis. Nat. Commun. 11, 247 (2020).
Liu, Y. et al. Tumour heterogeneity and intercellular networks of nasopharyngeal carcinoma at single cell resolution. Nat. Commun. 12, 741 (2021).
Palshikar, M. G. et al. Executable models of immune signaling pathways in HIV-associated atherosclerosis. NPJ Syst. Biol. Appl. 8, 35 (2022).
Wang, S. et al. An atlas of immune cell exhaustion in HIV-infected individuals revealed by single-cell transcriptomics. Emerg. Microbes Infect. 9, 2333–2347 (2020).
Savage, A. K. et al. Multimodal analysis for human ex vivo studies shows extensive molecular changes from delays in blood processing. iScience 24, 102404 (2021).
Mistry, P. et al. Transcriptomic, epigenetic, and functional analyses implicate neutrophil diversity in the pathogenesis of systemic lupus erythematosus. Proc. Natl Acad. Sci. USA 116, 25222–25228 (2019).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Martin, J. C. et al. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell 178, 1493–1508 (2019).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913 (2021).
Terekhova, M. et al. Single-cell atlas of healthy human blood unveils age-related loss of NKG2C+GZMB−CD8+ memory T cells and accumulation of type 2 memory T cells. Immunity 56, 2836–2854 (2023).
Huang, X. & Huang, Y. Cellsnp-lite: an efficient tool for genotyping single cells. Bioinformatics 37, 4569–4571 (2021).
Huang, Y., McCarthy, D. J. & Stegle, O. Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol. 20, 273 (2019).
Virshup, I., Rybakov, S., Theis, F. J., Angerer, P. & Wolf, F. A. anndata: annotated data. Preprint at bioRxiv https://doi.org/10.1101/2021.12.16.473007 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Massoni-Badosa, R. et al. An atlas of cells in the human tonsil. Immunity 57, 379–399.e18 (2024).
Sikkema, L. et al. An integrated cell atlas of the lung in health and disease. Nat. Med. 29, 1563–1577 (2023).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Walle, T. wallet-maker/cytopus: Cytopus v1.30. Zenodo https://doi.org/10.5281/zenodo.8366975 (2023).
Badia-i-Mompel, P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv. 2, vbac016 (2022).
Turner, M. D., Nedjai, B., Hurst, T. & Pennington, D. J. Cytokines and chemokines: at the crossroads of cell signalling and inflammatory disease. Biochim. Biophys. Acta 1843, 2563–2582 (2014).
Pishesha, N., Harmand, T. J. & Ploegh, H. L. A guide to antigen processing and presentation. Nat. Rev. Immunol. 22, 751–764 (2022).
Blum, J. S., Wearsch, P. A. & Cresswell, P. Pathways of antigen processing. Annu. Rev. Immunol. 31, 443–473 (2013).
Bhat, M. Y. et al. Comprehensive network map of interferon gamma signaling. J. Cell Commun. Signal. 12, 745–751 (2018).
Murayama, M. A., Shimizu, J., Miyabe, C., Yudo, K. & Miyabe, Y. Chemokines and chemokine receptors as promising targets in rheumatoid arthritis. Front. Immunol. 14, 1100869 (2023).
Lee, B.-W. & Moon, S.-J. Inflammatory cytokines in psoriatic arthritis: understanding pathogenesis and implications for treatment. Int. J. Mol. Sci. 24, 11662 (2023).
Kany, S., Vollrath, J. T. & Relja, B. Cytokines in inflammatory disease. Int. J. Mol. Sci. 20, 6008 (2019).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 6876 (2021).
Persad, S. et al. SEACells infers transcriptional and epigenomic cellular states from single-cell genomics data. Nat. Biotechnol. 41, 1746–1757 (2023).
Müller-Dott, S. et al. Expanding the coverage of regulons from high-confidence prior knowledge for accurate estimation of transcription factor activities. Nucleic Acids Res. 51, 10934–10949 (2023).
XGBoost: a scalable tree boosting system. in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. https://doi.org/10.1145/2939672.2939785 (Association for Computing Machinery, 2016).
Akiba, T., Sano, S., Yanase, T., Ohta, T. & Koyama, M. Optuna: a next-generation hyperparameter optimization framework. Preprint at arXiv https://doi.org/10.48550/arXiv.1907.10902 (2019).
Welford, B. P. Note on a method for calculating corrected sums of squares and products. Technometrics 4, 419–420 (1962).
Dong, W., Moses, C. & Li, K. Efficient k-nearest neighbor graph construction for generic similarity measures. in Proceedings of the 20th International Conference on World Wide Web 577–586 https://doi.org/10.1145/1963405.1963487 (Association for Computing Machinery, 2011).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
De Donno, C. et al. Population-level integration of single-cell datasets enables multi-scale analysis across samples. Nat. Methods 20, 1683–1692 (2023).
Acknowledgements
The authors would like to thank the helpful support received from publicly available data used in the present study by providing processed data in the optimal format. Additionally, we appreciate the great effort put in creating the DISCO database with multiple-tissue atlases using single-cell datasets. The authors would like to thank the Centro Nacional de Análisis Genómico (CNAG) Scientific IT Unit and the maintainers of the CNAG compute cluster for providing assistance with essential computing resources. Also, the authors would like to thank all the team members, in particular P. Nieto, H. L. Crowell, M. Elosua-Bayes, M. Abdalfatah and R. Massoni-Badosa, for their contributions during brainstorming and discussion sessions. This project has received funding from the European Union’s H2020 Research and Innovation Program under grant agreement number 848028 (DoCTIS; ‘Decision on optimal combinatorial therapies in IMIDs using systems approaches’). L.J.-G. has held an FPU PhD fellowship (FPU19/04886) from the Spanish Ministry of Universities. D. Maspero is supported by the Juan de la Cierva Fellowship (JDC2022-049637-I) from the Spanish Ministry of Science and Innovation and the European Union ‘NextGenerationEU’/PRTR. F.C. is funded by a Swiss National Science Foundation Sinergia grant (CRSII5-205884). M. Boulougouri is funded by the Graph Neural Networks for Explainable Artificial Intelligence ERA-NET+EJP (20CH21_195579) grant. J.L.-B. is supported by the Spanish Ministry of Universities through Margarita Salas fellow (RSUC.UDC.MS06). R.O.R.F. is supported by the Deutsche Forschungsgemeinschaft through CRC/SFB 1550 ‘Molecular circuits of heart disease’. Y.H. is supported by a Junior Postdoctoral fellowship from Research Foundation Flanders (FWO 12D5823N). S.T. is supported by a BOF-Fundamental Clinical Research mandate (FKO) from KU Leuven and by the Belgian Foundation Against Cancer (FAF-C/2018/1301). V.S. is funded by Asociación Española Contra el Cáncer (AECC). M.C.N. acknowledges funding from GlaxoSmithKline, the Netherlands Lung Foundation (project no. 4.1.18.226) and the European Union’s H2020 Research and Innovation Program under grant agreement number 874656 (discovAIR). This collaboration project is co-financed by the Ministry of Economic Affairs and Climate Policy by means of the PPP allowance made available by the Top Sector Life Sciences & Health to stimulate public−private partnerships. A.S. is funded by PID2021-123918OB-I00 from MCIN/AEI/51 10.13039/501100011033 and co-funded by ‘FEDER: a way to make Europe’. Part of the computational analyses was supported by the Google Cloud Research Credits program with award GCP19980904.
Author information
Authors and Affiliations
Contributions
L.J.-G., J.C.N. and H.H. conceived the project. J.C.N. and H.H. supervised the project. L.J.-G., D. Maspero, S.A.-F., F.C., M. Boulougouri and J.L.B. performed the computational and statistical analysis. R.O.R.F., H.A.T., R.N. and J.C.N. provided strategic advice on computational tasks and data interpretation. A.A. and M. Banchero generated VCF files for data demultiplexing. A.S.-M. and A.M.C. provided processed objects for validation analysis. M.R., D. Marchese, G.C., M. Berasategi and Y.H. generated datasets. T.K., P.v.d.V., F.A., F.P., G.S., E.R., M.v.d.B., A.S., J.M.C., A.F.N., E.D., J.D.C., J.T., J.P.G., E.C., G.G., B.S., A.J., V.S., R.E., S.T., S.V., M.C.N. and S.M. provided patient samples. L.J.-G., D. Maspero, S.A.-F., F.C., J.S.-R., A.-C.V., J.C.N. and H.H. interpreted the results. L.J.-G., D. Maspero, S.A.-F., F.C., J.C.N. and H.H. wrote the manuscript, with input from all authors. All authors read and approved the current version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.H. is co-founder and Chief Scientific Officer of Omniscope; a scientific advisory board member of Nanostring, Bruker and MiRXES; and a consultant to Moderna and Singularity. H.H. also received an honorarium from Genentech. J.C.N. is a scientific consultant to Omniscope. V.S. has received research grants from AstraZeneca and honoraria from GlaxoSmithKline unrelated to this study. M.v.d.B. has received research grants (unrestricted) from AstraZeneca, Novartis, GlaxoSmithKline, Roche, Genentech, Chiesi and Sanofi. M.N. has been awarded research grants (unrestricted) from AstraZeneca and GlaxoSmithKline. A.S. is the recipient of research grants from Roche-Genentech, AbbVie, GlaxoSmithKline, Scipher Medicine, Pfizer, Alimentiv, Boehringer Ingelheim and Agomab; receives consulting fees from Genentech, GlaxoSmithKline, Pfizer, HotSpot Therapeutics, Alimentiv, Origo Biopharma, Deep Track Capital, Great Point Partners and Boxer Capital; and is on the advisory boards of BioMAdvanced Diagnostics, Goodgut and Orikine. A.A. is a computational biologist at IMIDomics, Inc. A.J. is the chief data scientist at IMIDomics, Inc. S.M. is the co-founder and chief medical officer at IMIDomics, Inc. J.S.-R. reports funding from GlaxoSmithKline, Pfizer and Sanofi and fees/honoraria from Travere Therapeutics, Stadapharm, Astex, Pfizer, Grunenthal and Owkin. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Composition of the inflammation atlas datasets from the Main Core.
(a) Barplot showing patient count distribution across different diseases, including Healthy condition, stratified by technical variables (studyID and chemistry) and clinical metadata (age and sex). The donor without age information is shown in white. (b) Results from the scib-metrics package computed on five different embedding spaces, ranked by their overall performances. (c) Heatmaps showing the coefficient of determination R2 from a linear regression between each principal component and one of four confounding factors. The Principal Component Analysis was performed on (left) original data (normalized and log-scaled) and (right) from scANVI normalized expression (log-scaled). (d) Cellular proportions (Level 1) across diseases and Healthy donors (Top). Compositional analysis of Level 1 populations (excluding Platelets and RBC) between each disease and Healthy donors (Bottom). The dot size reflects the significance of the result (‘Final parameter’!= 0), and the color represents the log2FC (Disease vs Healthy).
Extended Data Fig. 2 Inflammation-related signatures across diseases and cell types.
(a) Heatmap displaying the transcription factor (TF) specificity of STAT1 and SP1 across different cell types and diseases. The t-statistic represents the relative expression of genes between diseased and Healthy samples, highlighting shared genes between TF target genes and IFN-induced cell type signatures. (b) Boxplot displaying the activity of STAT1 and SP1 across cell types (Level2) in SLE patients and Flu patients. (c) Boxplot displaying the activity of SP1 across monocyte subpopulations (Level 2) in SLE, Flu, Cirrhosis and HNSCC patients. In panels (b) and (c), the pseudobulk value computed for each cell type within each independent patient are presented as median values, with boxes indicating the interquartile range (IQR, 25th–75th percentile) and whiskers extending up to 1.5 × IQR beyond the box boundaries; points outside this range are shown individually as outliers. Statistical significance was assessed using a two-sided Wilcoxon rank-sum test, and P-values were adjusted for multiple comparisons using the Benjamini–Hochberg procedure (adjusted P < 0.05). Asterisks (*) denote significant differences relative to other cell lineages or diseases, with the position of the asterisk indicating the direction of change (above the box: upregulated; below the box: downregulated). Exact P-values, effect sizes, and sample sizes are provided in Supplementary Table 6 (sheets pval_SLE_Level2 and pval_Mono_Level2 for panels b and c, respectively).
Extended Data Fig. 3 Confusion matrices of predicted inflammatory condition by cell type.
Normalized confusion matrices, one for each cell type (Level 1; excluding Cycling cells, Progenitors, Platelets and RBC), displaying proportion of predictions belonging to each True Condition. Diagonal values correspond to the Recall metric. XGBoost was trained on the (a) scANVI batch corrected and log-scaled cell expression profiles, and (b) original normalized and log-scaled cell expression profiles.
Extended Data Fig. 4 Exploring cell misclassification within COMBAT2022 dataset.
(a-b) Normalized confusion matrices, aggregated (left) and one for each cell type (Level 1; excluding Cycling cells, Progenitors, Platelets and RBC) (right), displaying proportion of predictions belonging to each True Condition. Diagonal values correspond to the Recall metric. XGBoost was trained on the original normalized and log-scaled cell expression profiles from (a) whole COMBAT dataset and (b) Healthy, Flu and COVID (stratified by disease severity) samples from COMBAT dataset. (c-d) Agglomerative hierarchical clustering with complete linkage (using the average method and cosine distance) was performed on pseudobulk gene expression at the patient level (c), or at cell type (Level1) and patient level (d), using the log-normalized uncorrected count matrix on the 8,253 gene expression universe. Sample covariates, including sequencing pool, sex, and age, were also incorporated.
Extended Data Fig. 5 Sex-specific classification performances.
(a) Normalized confusion matrices showing the proportion of cell-wise classifier predictions for each true condition. Diagonal values represent the Recall metric. The XGBoost classifier was trained on scANVI batch-corrected data. Values were computed separately for cells from female (n = 448) (left) and male (n = 369) (right) samples. (b–e) Patient-wise classifier performance measured by Weighted F1 scores, stratified by sex. (b) Scenario 1: 5-fold cross-validation distributions for each left-out split. Data are presented as median values, with boxes indicating the interquartile range (IQR, 25th–75th percentile) and whiskers extending up to 1.5 × IQR beyond the box boundaries; points outside this range are shown individually as outliers (n = 5). Non-significant differences (ns, p-value > 0.05) were assessed using a two-sided Mann–Whitney U test for each cell type. (c–e) Scenario 2, Scenario 3, and centralized approach: performance on unseen patients (c), unseen studies (d), and left-out pool observations (e), respectively.
Extended Data Fig. 6 Functional biomarker discovery using interpretable machine learning analysis.
(a) Gene list ranked top-to-bottom by importance (absolute d-SHAP value), coupled with max-normalized expression levels computed per cell type (Level1) and considering selected diseases. From left to right, reporting top ranked genes for n T CD4 Naive cells in RA disease as well as for monocytes and pDC in SLE patients. (b) Rank by importance (absolute d-SHAP value) of the CYBA gene in every combination of cell type (Level1) and disease. (c) Scatter plot of d-SHAP values against the aggregated s-SHAP values on monocyte population and specific diseases (first row: PS, PSA, CD, and second row: UC, Asthma, COPD, from left to right). (d) Rank by importance (absolute d-SHAP value) of IFITM1 gene in every combination of cell type (Level1) and disease. (e) Scatter plot of d-SHAP values against the aggregated s-SHAP values on T CD4 Non-Naive (top) and ILC (bottom) population and specific diseases (Asthma, COPD, and Cirrhosis, from left to right). In Panels (a), (b), and (d) we first dropped the genes expressed in less than 5% of the selected cell population. In Panels (c), and (e), the top 20 genes according to d-SHAP are marked in turquoise; of these, the genes that are also among the top 20 by s-SHAP are marked in purple. The gene of interest is annotated in red.
Extended Data Fig. 7 Extended patient classifier workflow schema.
(a) Definition of reference and query datasets, for Scenarios 1, 2, 3, and centralized dataset (from left to right). (b) Integration of the reference dataset and mapping of the query dataset to define the patient-wise embeddings, stratified by cell type. (c) Patient classifier pipeline composed by the hyperparameter tuning of each classifier family, the selection of the best classifier family and the final evaluation of the left-out query dataset. (d) Schema of the three experiments performed. Icons created with Inkscape.
Extended Data Fig. 8 Additional performance evaluation metrics in Scenario 1.
(a-b) Boxes indicate the interquartile range with the median as a center line; whiskers extend to 1.5×IQR, and outliers are shown as individual points. Each box includes n = 5 points. (a) Boxplots showing the distribution of Balanced Accuracy Score (balanced by true disease support), F1, Precision and Recall computed during 5-fold cross-validation, considering Majority Vote prediction on the left-out split for each inflammatory condition. The average number of samples among 5 splits, with the corresponding ground truth labels, are also reported. (b) Boxplots showing the distribution of Balance Accuracy Score (top), and Matthew Correlation Coefficient (bottom) computed during 5-fold cross-validation, considering Majority Vote and cell type prediction, on the left-out split from 817 samples. (c) Heatmap reporting Recall and Precision computed by aggregating the prediction performed by each cell type on each left-out split during 5-fold cross-validation.
Extended Data Fig. 9 Additional performance evaluation metrics in the Centralized dataset analysis and across state-of-the-art data integration approaches.
(a) Pointplot showing the Balance Accuracy Score (top), and Matthew Correlation Coefficient (bottom) computed, considering Majority Vote, 100 random disease assignments, and cell type prediction, on the samples from left out pools in the Centralized Dataset. (b) Heatmap reporting Recall and Precision obtained on the samples from left out pools by each cell type for each disease included in the centralized dataset. (c-d) Performance evaluation from Scenario 2 (c) and Scenario 3 (d), respectively, showing (left) the distribution of Weighted Recall and Weighted Precision for all the configurations of each data integration approach, and (right) the mean and standard-deviation of each data integration method, including 100 random label assignments. Arrows highlight the scANVI configuration applied in Scenario 1.
Extended Data Fig. 10 Patient classifier performance in Scenarios 2 and 3.
(a, c, e, g) Result obtained with the best parameter configuration for each integration and mapping method, considering Weighted F1 (WF1) score computed on prediction of samples from unseen patients. (b,d,f,h) Result obtained with the best parameter configuration for each integration and mapping method, considering WF1 score computed on prediction of samples from unseen studies. In Panels (a) to (h): (top-left) Pointplot of WF1-scores for Majority vote and each cell type. (bottom-left) F1-score for each combination of cell type and disease, columns ordered for similarities. (right) Normalized confusion matrices displaying proportion of predictions belonging to each true condition. Diagonal values correspond to the Recall metric. Corresponding Majority Vote WF1 score and Balanced Accuracy Score (BAS) were reported. Note, scPoli configurations where embedding space=sample were not considered.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1−4 and supplementary references.
Supplementary Tables (download ZIP )
Compressed archive including Supplementary Tables 1−9 in .xlsx format.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jiménez-Gracia, L., Maspero, D., Aguilar-Fernández, S. et al. Interpretable inflammation landscape of circulating immune cells. Nat Med 32, 633–644 (2026). https://doi.org/10.1038/s41591-025-04126-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-04126-3