Abstract
Peroxisome proliferator-activated receptor gamma (PPARγ) is a master regulator of luminal lineage in urothelial carcinoma. FX-909 is a first-in-class oral small-molecule PPARγ inverse agonist. Here we report the first part of FX-909-CLINPRO-1, a phase 1A 3 + 3 dose-escalation study of FX-909, that enrolled 56 patients with advanced solid tumors, including 46 with urothelial carcinoma. The primary end point was safety and tolerability; secondary end points included recommended phase 2 dose determination, pharmacokinetics and preliminary antitumor activity. FX-909 exhibited an acceptable safety and tolerability profile. Grade ≥3 adverse events included anemia (26.8%), thrombocytopenia (21.4%), fatigue (10.7%) and hyperglycemia (7.1%). Doses of 30 mg and 50 mg daily were selected for recommended phase 2 dose optimization. Objective responses were observed in 17.5% of patients with urothelial carcinoma across all dose levels. Exploratory analyses revealed that tumor responses were enriched in patients with high PPARγ expression. FX-909 demonstrated acceptable safety and tolerability with preliminary antitumor activity, supporting further clinical development in urothelial cancer. ClinicalTrials.gov identifier: NCT05929235.
Similar content being viewed by others
Main
Urothelial carcinoma (UC) represents a substantial clinical burden, with approximately 656,000 new diagnoses and 234,000 deaths projected worldwide in 20251. The landscape of treatment for advanced UC has recently changed with the introduction of immune checkpoint inhibitors and antibody–drug conjugates2. Notwithstanding such advances in systemic therapy, most patients with metastatic UC experience disease progression and a subset exhibit primary refractory disease. These findings highlight an urgent unmet need for novel therapeutic approaches in this disease, particularly those that exploit major advances in our understanding of cancer cell-intrinsic drivers of UC growth.
Comprehensive molecular profiling of UC has revealed distinct molecular subtypes, with the luminal subtype representing approximately 65% of advanced UCs3,4. The luminal subtype is characterized by high expression of peroxisome proliferator-activated receptor gamma (PPARγ), a nuclear hormone receptor and transcription factor. PPARγ plays an essential lineage-determining role in normal urothelial homeostasis and regeneration, and mounting evidence indicates that aberrant PPARγ activation drives tumorigenesis in luminal UC5,6,7.
Genomic evidence for PPARγ activation in UC includes recurrent focal amplifications, activating mutations and hotspot mutations in RXRA (S427F/Y), the obligate heterodimeric partner of PPARγ7,8. The RXRA S427F mutation enhances PPARγ–RXRA heterodimerization, stabilizes the active conformation of PPARγ and confers ligand-independent transcriptional activation7,9. Genome-wide CRISPR screens have consistently identified PPARγ as a top selective dependency in UC cell lines, and PPARγ-activated cell lines demonstrate exquisite sensitivity to genetic ablation of PPARγ7,10,11,12. Cell lines harboring PPARγ gain or amplification exhibit oncogene addiction, demonstrated by dependence on sustained PPARγ signaling for proliferation and survival13.
Targeting lineage-specific nuclear hormone receptors is a clinically validated therapeutic strategy in oncology. The rationale for targeting PPARγ in luminal UC can be conceptualized as similar to androgen receptor inhibition in prostate cancer or estrogen receptor inhibition in breast cancer7. These parallels extend beyond conceptual analogy, as luminal breast and bladder cancers share conserved transcriptional programs, including PPARγ pathway activation and enrichment for FGFR3 alterations, suggesting that PPARγ may represent a therapeutically actionable lineage dependency across luminal solid tumors14,15. As with prostate and breast cancer, the luminal phenotype of UC is also more prevalent in earlier stages of the disease16,17. More recently, luminal transcriptional programs have emerged as potential therapeutic vulnerabilities across a broader spectrum of solid tumors, including lung cancer, head and neck squamous cell carcinoma and pancreatic ductal adenocarcinoma18.
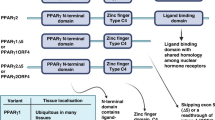
Despite the body of evidence nominating PPARγ as a therapeutic target in UC and other malignancies, attempts to pharmacologically antagonize PPARγ, so far, have demonstrated limited ability in reversing PPARγ activation7. FX-909 is a rationally designed, first-in-class covalent inverse agonist that stably enforces a conformationally repressive state of PPARγ19 (Supplementary Fig. 1). Unlike antagonists that passively block ligand binding, FX-909 actively recruits nuclear corepressors (NCOR1/NCOR2) while simultaneously blocking co-activator binding, thereby achieving potent suppression of both ligand-mediated and basal PPARγ transcriptional activity19. In vitro PPARγ-activated UC cell lines with PPARγ mutations or amplifications or RXRA mutations, showed preferential sensitivity to FX-90919. In xenograft models of PPARγ-activated UC, FX-909 achieved durable tumor regressions19.
Here, we report results from the phase 1 part A dose escalation study of FX-909 (NCT05929235) establishing proof of concept for PPARγ inverse agonism as a therapeutic strategy in patients with locally advanced (unresectable) or metastatic UC. We also report the discovery of a biomarker to enrich for future selection of patients deriving benefit from FX-909.
Results
Phase 1 part A dose escalation study design and patient characteristics
We conducted a first-in-human phase 1 study of FX-909 (FX-909-CLINPRO-1) in patients with advanced solid tumors including advanced UC. The primary objective of the dose-escalation phase (part A) of the study was to assess the safety and tolerability of FX-909, with primary end points of the incidence of dose-limiting toxicities (DLTs) and incidence and severity of adverse events (AEs) and serious AEs (SAEs). The secondary objectives were to: (1) define the preliminary recommended phase 2 dose (RP2D) and maximum tolerated dose of FX-909, (2) characterize the pharmacokinetic profile of FX-909 and (3) evaluate preliminary antitumor activity of FX-909. Exploratory objectives included assessment of the pharmacodynamic effects of FX-909 and potential predictive biomarkers of response to FX-909 that could be used to enrich patients for enrollment in the planned expansion cohort (part B).
The dose-escalation phase (part A), reported here, enrolled 56 patients with advanced or metastatic solid tumors across four dose levels (Fig. 1 and Supplementary Fig. 1) between 24 August 2023 and 8 October 2025. Most treated patients were white men, and the median age of the treated cohort was 70 years (range, 44–86 years) (Table 1). On the basis of the mechanism of action and emerging data, patient accrual subsequently concentrated on UC and more specifically patients with genetic alterations in FGFR3, RXRA and PPARγ that are known to be enriched in luminal UC. Baseline characteristics were generally well balanced across the dose levels with the exception of more women included in the 50 mg cohort, as this cohort accrued while patients with any solid malignancy could be enrolled (Table 1). Each dose level was further enriched for patients with advanced UC via backfill.
A total of 68 patients were screened and 56 patients were enrolled across four doses.
Among the 46 patients with UC treated in part A, the median lines of previous therapy received was 3 (range, 1–8), and previous treatments included enfortumab vedotin + pembrolizumab (34.8%), enfortumab vedotin monotherapy (43.5%), anti-PD(L)1 blockade (95.7%) and FGFR3 inhibition (32.6%) (Table 1). The majority of patients with UC had visceral metastases (72%), including 24% with liver metastases (Table 1).
Dose escalation and DLTs
FX-909 was administered orally once daily (QD) in 28-day cycles. Dose escalation followed a 3 + 3 design (Supplementary Fig. 1). The 56 enrolled patients were assigned to one of four dose levels according to the dose escalation design (30 mg (N = 17), 50 mg (N = 20), 70 mg (N = 13) and 100 mg (N = 6)). The DLT observation period was 28 days or until the first dose of cycle 2 if cycle 2 was delayed. During the DLT observation period, dose interruptions were allowed but dose reductions were not. After the DLT observation period, both dose interruptions and dose reductions were allowed. No DLTs were observed at dose levels of 30 mg or 50 mg (Extended Data Table 1). At the 100-mg dose level, two of five DLT-evaluable patients experienced DLTs: grade 3 proteinuria (nephrotic range) and grade 3 hyperglycemia requiring a brief course of insulin. At the 70 mg dose, 1 of 13 patients experienced a DLT of grade 3 anemia and received less than 75% of the planned dose intensity in cycle 1. Dose modifications, including interruptions and reductions, are presented in Extended Data Table 1.
Treatment-emergent AEs (TEAEs) occurred in 100% of patients, with grade ≥3 TEAEs reported in 75% of patients (Extended Data Table 2). The most common TEAEs of any grade were thrombocytopenia, fatigue, diarrhea, anemia and hyperglycemia, whereas the most common grade ≥3 TEAEs were anemia, thrombocytopenia, fatigue and hyperglycemia (Extended Data Table 2).
Treatment-related AEs (TRAEs) occurred in 95% of patients, with grade ≥3 TRAEs reported in 59% (Table 2). The most common TRAEs of any grade were thrombocytopenia, fatigue, diarrhea, anemia, hyperglycemia, elevated transaminases and hypertriglyceridemia, whereas the most common grade ≥3 TRAEs were anemia, thrombocytopenia, fatigue and hyperglycemia (Table 2). No treatment-related deaths occurred. There were five deaths within 30 days of the last dose of FX-909; four were attributed directly to progression of disease or complications of the underlying urothelial malignancy and one was the result of a specific AE considered unrelated to FX-909. Overall, the median time to onset of the first grade ≥3 TRAEs was 36 days (Extended Data Table 3).
SAEs were experienced by 23 (41.1%) patients during the study, of which 2 cases (3.6%) were deemed possibly related to FX-909; hyperglycemia in one patient and pulmonary embolism and atrial flutter in another patient.
Triplicate electrocardiograms that were time matched to pharmacokinetic collections on cycle 1 days 1 and 2 and on cycle 1 days 15 and 16 did not reveal any obvious prolongation in the QTcF interval with FX-909 treatment.
The median dose intensity was 91% across all dose levels in the first 56 days, including 100% in the 30-mg dose level and 84% in the 50-mg dose level (Extended Data Table 4).
At the time of the data cut, nine patients remained on treatment with FX-909. The most common reason for treatment discontinuation was progressive disease (PD). Permanent discontinuations of FX-909 for AEs or withdrawal of consent were uncommon (Fig. 1). The study met its primary end point, demonstrating acceptable safety and tolerability with FX-909, particularly at the 30-mg daily and 50-mg daily dose levels.
On the basis of the totality of the safety and tolerability, pharmacokinetic and pharmacodynamic data, as described below, Fx-909 30 mg and 50 mg daily were selected as preliminary RP2Ds for further investigation in a randomized phase 1B study.
Pharmacokinetics and target engagement
Pharmacokinetic results were available from 40 patients (Supplementary Table 2). Following oral administration, FX-909 exhibited dose-proportional increases in maximum plasma concentration (Cmax) and the area under the curve (AUC) from 30 mg to 100 mg QD (Extended Data Table 5), with a dose- and time-independent mean steady-state half-life supportive of QD dosing and minimal accumulation (AUC0–24 accumulation ratio ≤1.63) (Extended Data Fig. 1a).
Nonclinical studies in rodents previously demonstrated that the effects of PPARγ inverse agonism on target gene expression in skin were highly correlated with those in tumors20. Therefore, paired skin punch biopsies were collected at screening and cycle 1 day 15 to assess the pharmacodynamic effects of FX-909 (Supplementary Table 3). Robust target engagement was observed across all dose levels. FX-909 induced a >75% reduction in expression of the canonical PPARγ target gene, FABP4, consistent with pharmacological inverse agonism of PPARγ (Extended Data Fig. 1b). An additional nine PPARγ-related target genes (including PLIN4, RBP7 and CD36) were also evaluated and demonstrated similar results (Supplementary Fig. 3a). Exposure–pharmacodynamic analysis revealed no significant correlation between FX-909 plasma concentrations and suppression of the nine-gene composite (Spearman rho −0.09, P = 0.71), reinforcing that target engagement was observed across all dose levels explored (Supplementary Fig. 3b).
Antitumor activity
No antitumor activity was observed among the ten treated patients with non-UC cancers. Among the 46 patients with advanced UC, confirmed objective responses were achieved in 7 of 40 patients with measurable disease (one complete response (CR) and six partial responses (PRs)). An additional 16 patients achieved stable disease (Fig. 2). The DCR and median DOR are presented in Supplementary Table 5. Tumor regressions of ≥30% in the sum of target lesion diameters (SLD) from baseline were observed in five such patients but were not confirmed on a subsequent scan per RECIST v1.1; this included a patient with high expression of PPARγ and mixed responses at different tumor sites who demonstrated an on-treatment decline in circulating tumor DNA (ctDNA), a regression of liver metastases and remained on treatment for 8.4 months (Supplementary Fig. 4a–d).
Waterfall plot of best percentage change from baseline in SLD in measurable radiographic disease for all patients with advanced UC with measurable disease and at least one post-baseline scan (N = 39, excluding one patient with clinical progression during cycle 1 and no post-baseline scan completed).
Across the four dose levels (30 mg (17), 50 mg (16), 70 mg (11) and 100 mg (2)), evidence of antitumor activity in efficacy-evaluable patients with UC was observed at all dose levels. No clear relationship between dose and antitumor activity was observed at the doses investigated.
Post hoc analysis of the prevalence of luminal UC and PPARγ pathway alterations in UC
Interrogation of alterations in the PPARγ signaling axis through molecular characterization of a real-world cohort of 2,685 muscle-invasive patients with UC showed that luminal subtypes accounted for 65% of tumors and were enriched for PPARγ amplifications, RXRA mutations and FGFR3 mutations (Supplementary Fig. 5a and Supplementary Table 1). PPARγ expression was significantly higher in luminal versus nonluminal tumors (Supplementary Fig. 5b), reinforcing PPARγ as a key therapeutic target in luminal UC.
Exploratory analysis of PPARγ protein expression as a biomarker of luminal lineage that enriches for response to FX-909
The early observations of antitumor activity with FX-909 prompted an exploratory effort to evaluate pretreatment biomarkers associated with luminal lineage that might enrich for patient benefit. Archival tumor tissues or fresh biopsies were mandated in all patients with advanced UC enrolled in part A of the study. Among the efficacy-evaluable cohort of patients with UC (N = 40), 35 were deemed biomarker evaluable on the basis of available or adequate tumor specimens (Supplementary Fig. 6a). Tumor tissues were profiled by integrated DNA and RNA sequencing (RNAseq) to characterize the molecular features associated with UC lineage (Supplementary Table 8). Consistent with the real-world cohort (Supplementary Fig. 5b), PPARγ mRNA expression was significantly higher in luminal compared with nonluminal tumors (mean 8.09 versus 5.47 log2(transcripts per million (TPM) + 1), P < 0.0001) (Fig. 3a).
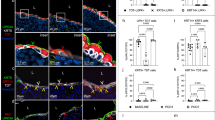
a, PPARγ mRNA expression stratified by lineage (luminal, N = 21; nonluminal N = 5 patients). Box plots display the median and interquartile range; whiskers denote the 5th to 95th percentile range. Two-sided Wilcoxon rank-sum test was used for statistical analysis. b, Nuclear tumor staining of PPARγ by IHC, showing PPARγ-high (100% TPS), PPARγ-expressing (50% TPS) and PPARγ-absent (0% TPS) patterns. Scale bar, 200 μm. For each patient, a hematoxylin and eosin (H&E) stain was performed to assess tissue morphology, and a negative control was included in the same experimental procedure. c, Waterfall plot of the best percentage change in SLD from baseline in patients with advanced UC with PPARγ-high tumors (N = 25; Supplementary Fig. 6a). d, Swimmer plot showing radiographic response over time in PPARγ-high subgroup. Note that, at the time of the data cut, dosing logs were incomplete for some of the patients and the swim lane plots were only plotted over the time that the dosing information was available. e, Spider plot of the percentage change in SLD from baseline over time in the PPARγ-high subgroup. f, Waterfall plot depicting molecular response as percentage change from baseline in ctDNA aVAF at the cycle 2 or 3 time point. Patients with undetectable ctDNA at baseline were excluded (N = 25; Supplementary Table 7). EOT, end of treatment.
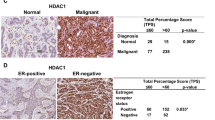
Given that genetic alterations in PPARγ signaling may incompletely capture PPARγ-activated tumors (Supplementary Table 1) and on the basis of pragmatic considerations for companion diagnostic development, an immunohistochemistry (IHC)-based assay for PPARγ (SP500) was performed. PPARγ protein expression was assessed by IHC using a monoclonal antibody, and a tumor proportion score (TPS) was defined as the percentage of tumor cells with nuclear PPARγ staining of any intensity (Fig. 3b and Supplementary Table 4). Among the biomarker-evaluable patients, 15% of tumors were categorized as PPARγ absent/low (≤10% TPS) and 85% as PPARγ-expressing (>10% TPS). Consistent with previous observations21, PPARγ protein expression (TPS%) showed a high degree of concordance with PPARγ mRNA levels (log2(TPM + 1)) (Pearson r = 0.88, P < 0.001) (Extended Data Fig. 2c). PPARγ protein expression, as with mRNA expression, was significantly higher in luminal tumors compared with nonluminal tumors (mean 93.5% versus 30.8% TPS) (Extended Data Fig. 2d).
A PPARγ TPS threshold was defined to optimally identify luminal UCs. The strong concordance between mRNA and protein expression in the phase 1 cohort provided a basis for leveraging PPARγ mRNA expression data from the real-world cohort (n = 2,685)22 to inform protein-based thresholds for identifying tumors of luminal lineage. In the real-world cohort (Supplementary Fig. 5b), a high degree of separation was observed between distributions of PPARγ mRNA values in luminal versus nonluminal lineage UCs across the range of 6.1–7.1 log2(TPM + 1) (Extended Data Fig. 2a). PPARγ mRNA cutoffs within this range were associated with enrichment for luminal lineage, with a sensitivity of >74.5% and a positive predictive value (PPV) of >81% (Extended Data Fig. 2b). Using LM to map mRNA thresholds to protein expression in the phase 1 cohort, a provisional TPS cutoff of ≥60% for defining a PPARγhigh subgroup was identified (Extended Data Fig. 2c). Of note, in the real-world cohort, genetic alterations in PPARγ, RXRA or FGFR3 did not provide additional predictive value for luminal lineage enrichment (Extended Data Table 6).
Having established the PPARγ TPS ≥60% threshold on the basis of biological criteria independent of clinical outcomes with FX-909, the association between PPARγ IHC expression and clinical activity with FX-909 in the phase 1 cohort was explored. Among the 35 biomarker-evaluable patients with advanced UC, 25 composed the PPARγhigh subgroup with a TPS of ≥60% (Supplementary Fig. 6b). On-treatment tumor regression was observed in the majority of this patient subgroup, and confirmed PRs were observed in 5/25 patients (Fig. 3c). PPARγ TPS results for efficacy-evaluable patients with UC (n = 34; Supplementary Fig. 6a) are shown in Supplementary Fig. 7. One additional patient with PPARγhigh UC with a solitary nontarget lesion (mediastinal lymph node) demonstrated a CR and remained on treatment for 8+ months at the time of the data cut. One patient with PPARγhigh UC and a PPARγ mutation (Supplementary Table 10) did not achieve a response to FX-909. This rare PPARγ (C313 missense mutation) is predicted to block the binding of FX-909, with a prevalence of <1% in advanced UCs.
Among the 25 patients in the PPARγhigh subgroup, 5 remained on treatment at the time of data cut: 3 patients at 30 mg QD for 5.3+, 5.8+ and 12.7+ months, 1 patient at 50 mg QD for 5.5+ months and 1 patient at 70 mg QD for 7.6+ months. The durability and depth of responses are further shown in Fig. 3d,e and presented in Supplementary Table 5.
Exploratory analysis of molecular response by ctDNA
On-treatment ctDNA declines as a measure of response, termed liquid biopsy RECIST (LB-RECIST), have been proposed and prospectively validated in solid tumors23,24. We performed an exploratory analysis of ctDNA kinetics and clinical outcomes in patients treated with FX-909. Among 46 patients with advanced UC, 29 had paired baseline and cycle 2 or 3 plasma samples for ctDNA analysis using the CARIS Assure test (Supplementary Table 6). On-treatment quantitative ctDNA changes were classified according to the LB-RECIST framework, and 15/29 patients achieved an LB-RECIST response (Fig. 3f and Supplementary Table 7). LB-RECIST responses correlated with radiographic responses in patients with measurable disease (Supplementary Fig. 8). The vast majority (80%) of those achieving an LB-RECIST response had PPARγhigh tumors (Fig. 3f).
Exploratory analysis of the molecular and immune landscape of PPARγhigh tumors
To further characterize tumors defined as PPARγhigh, an exploratory analysis of transcriptional lineage, genetic alterations and immune features in the phase 1 cohort was performed (Fig. 4, Supplementary Fig. 9 and Supplementary Fig. 6b). PPARγhigh tumors showed strong concordance with luminal biology features (Supplementary Fig. 10). Among patients with luminal-papillary and luminal-infiltrated subtype UCs, 9/19 demonstrated responses to FX-909 compared with 0/6 patients with basal squamous UC (Supplementary Fig. 11a,b). RXRA S427F mutations, which were confined to the luminal lineage, were associated with lower PPARγ protein expression (Supplementary Fig. 11c), consistent with previous reports indicating RXRA mutations confer a >8-fold higher biochemical activation of PPARγ, which we hypothesize provides significant pathway activation independent of PPARγ expression levels19. Two of four patients with RXRA-mutant UC demonstrated responses to FX-909 versus 7/16 patients with RXRA wild type tumors (Supplementary Fig. 11d and Supplementary Table 10). PPARγhigh tumors were enriched for FGFR3 alterations, and 5/11 patients with FGFR3 mutant tumors demonstrated responses with FX-909 versus 4/9 patients with FGFR3 wild type tumors (Supplementary Fig. 11e,f and Supplementary Table 10). Among 6/6 patients whose baseline samples were derived from tumors biopsied after they had received previous FGFR inhibitor therapy, testing revealed PPARγhigh. Notably, a subset of patients with PPARγhigh tumors lacked PPARγ amplifications or RXRA mutations yet demonstrated responses with FX-909 (Fig. 4).
Heat map showing samples grouped by molecular subtype and ordered by decreasing PPARγ protein expression on the basis of TPS (left to right). Molecular features are grouped into lineage-related features and immune-related features ordered by prevalence (for further details, see Supplementary Fig. 11b and Supplementary Table 10). cCR, confirmed complete response; cPR, confirmed partial response; NE, nonevaluable; NM, nonmeasurable disease; SD, stable disease; uPR, unconfirmed partial response; TMB, tumor mutational burden.
PPARγhigh tumors displayed a ‘cold’ immune phenotype, consistent with previous reports of limited responsiveness to anti-PD-1 therapies21,25. Negative correlations were observed between PPARγhigh tumors and immune- or epithelial–mesenchymal transition-related signatures, suggesting mutual exclusivity between differentiated luminal programs and immune-inflamed or mesenchymal tumor states (Supplementary Fig. 10). In addition to PPARγ, RXRA and FGFR3, the cohort harbored recurrent alterations in 29 additional genes mutated in more than 15% of patients (Supplementary Fig. 9 and Supplementary Table 9). Together, these data reinforce that PPARγhigh tumors are associated with other features of luminal lineage UC that may identify PPARγ-activated tumors more broadly than genomic alterations linked to the PPARγ signaling axis. This further supports PPARγ IHC as a potential means of enriching patients with luminal UC for enrollment in the part B dose-expansion cohort.
Discussion
The FX-909-CLINPRO-1 phase 1 part A study establishes proof of concept for pharmacological PPARγ inhibition through inverse agonism as a viable therapeutic strategy in advanced UC. The study met its primary end point demonstrating that FX-909 was safe and tolerable, particularly at 30 mg and 50 mg daily. Despite enrollment of a heavily pretreated patient population, we observed promising preliminary clinical activity across all dose levels explored, with confirmed objective responses and evidence of prolonged disease control. Exploratory analysis revealed that tumor responses were enriched in patients with PPARγhigh tumors (TPS ≥60%), with the majority of these patients experiencing tumor shrinkage and responses observed across multiple dose levels. Furthermore, on-treatment declines in ctDNA were observed.
Our pharmacokinetic and pharmacodynamic data support FX-909 as a pharmacologically active orally bioavailable PPARγ inverse agonist at all doses evaluated. Plasma exposures increased in a roughly dose-proportional manner from 30 mg to 100 mg, with mean steady-state terminal half-lives supporting QD dosing. Pharmacodynamic assessment via suppression of the PPARγ target gene, FABP4, in paired skin biopsies demonstrated target engagement exceeding the predicted 75% threshold for biological activity established in PPARγ-amplified xenograft models. This level of target gene suppression was achieved even at the 30 mg dose, providing mechanistic rationale for the antitumor activity observed across dose levels. No clear dose–pharmacodynamic or dose–response relationships have been observed until now on the basis of the dose levels explored.
The most common TEAEs observed with FX-909 were thrombocytopenia, fatigue, diarrhea, anemia and hyperglycemia. These AEs may be consistent with the known roles of PPARγ in hematopoiesis, energy metabolism and glucose homeostasis26. DLTs occurred at 70 mg and 100 mg, including grade 3 proteinuria, transient hyperglycemia and anemia. The lower incidence of grade ≥3 events, fewer dose modifications and higher relative dose intensity observed at 30 mg and 50 mg daily support these dose levels for further development.
An important component of this study was the parallel development of a PPARγ IHC biomarker to refine selection of patients most likely to benefit from FX-909. The provisional TPS cutoff of ≥60%, determined through integrative biostatistical modeling, accurately stratifies patients by luminal lineage and correlates with clinical benefit. This approach facilitates prospective identification of approximately 65% of patients with advanced luminal subtype UC. Prospective implementation of PPARγ IHC to define eligibility in the ongoing FX-909-CLINPRO-1 phase 1 part B expansion cohort exemplifies precision medicine approaches in a disease that has historically lacked effective biomarker-driven therapies directed at cancer cell-intrinsic drivers of oncogenesis.
In addition to early development of a potential companion diagnostic, the development of FX-909 highlights several other important findings. First, comprehensive molecular characterization of tumors can reveal dependencies on transcription factors previously considered inaccessible to therapeutic modulation by small molecules8. Second, mechanistic understanding of target biology can enable rational drug design to overcome resistance mechanisms19. Third, the potential for combination strategies may emerge from understanding the broader biological context of tumors associated with a given therapeutic target, including the potential for combinations with immunomodulatory therapies in tumors characterized by immune ‘cold’ microenvironments9.
There are potential limitations to this study. The part A cohort is relatively small, reflecting the dose-finding nature of this study, and longer follow-up is needed to assess the durability of responses. Although the PPARγ IHC assay demonstrates separation between responders and nonresponders, the TPS ≥60% cutoff is provisional and will require validation in ongoing studies including the part B cohort. The predictive biomarker analyses were exploratory in nature; however, this reflects the iterative approach inherent to first-in-mechanism clinical development, where emerging pharmacodynamic and efficacy signals necessarily inform biomarker hypotheses that ultimately require prospective validation. The heavily pretreated nature of the study population may underestimate the clinical activity of FX-909 in earlier treatment lines where tumor burden is lower, and tumors may be less molecularly complex and more dependent on luminal lineage for growth. While serial blood specimens were collected, on-treatment biopsies were not collected precluding extensive characterization of the impact of FX-909 on UC cells and the tumor microenvironment.
The development of FX-909 must be contextualized within the rapidly evolving treatment landscape for advanced UC. Although antibody–drug conjugates and immune checkpoint blockade have transformed treatment, resistance mechanisms inevitably emerge in most patients2. FGFR3 alterations are also enriched in luminal UC, with an overall prevalence of ~15–20% of advanced UC4,8. However, FGFR inhibitors benefit only a subset of patients with activating FGFR3 alterations27. The PPARγhigh biomarker identifies approximately 65% of patients with luminal UC, potentially enabling broader clinical applicability. Against this backdrop, FX-909 represents a mechanistically distinct approach targeting a large segment of the UC population. Its potential role may be greatest in the luminal subtype where PPARγ dependency is most pronounced.
In conclusion, the phase 1 part A results establish FX-909 as the first pharmacological agent capable of effectively inhibiting PPARγ in humans, with promising preliminary antitumor activity in heavily pretreated patients with advanced UC, particularly in those with PPARγhigh tumors. The acceptable safety profile, evidence of target engagement and development of a future companion diagnostic strategy support continued development of this first-in-mechanism approach. These findings validate PPARγ as a therapeutic target in luminal UC, positioning it alongside the androgen and estrogen receptors as successfully targeted nuclear receptors that serve as critical oncology targets. The ongoing part B expansion study will further define the role of PPARγ inverse agonism in the treatment of PPARγhigh advanced UC, with the potential for this approach to be extended to other malignancies with PPARγ activation.
Methods
Ethics approval and consent
This study is being conducted in accordance with the following:
-
Consensus ethical principles derived from international guidelines including the Declaration of Helsinki and Council for International Organizations of Medical Sciences (CIOMS) international ethical guidelines
-
Applicable International Council for Harmonisation (ICH) Good Clinical Practice guidelines
All participants provided written informed consent before the performance of any study-related procedures. Each study center received approval for participation from their respective institutional review board (IRB). Local IRB approval was obtained from Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai (Institutional Review Board of the Mount Sinai School of Medicine), Dana-Farber Cancer Institute (Dana-Farber IRB), Memorial Sloan Kettering Cancer Center (Memorial Sloan Kettering IRB), Cleveland Clinic (Cleveland Clinic IRB) and Massachusetts General Hospital Cancer Center (Dana-Farber IRB). Central IRB approval was obtained from Sarah Cannon Research Institute at Tennessee Oncology (Castle IRB), Smilow Cancer Hospital at Yale New Haven Health (Advarra), START Center for Cancer Research (Salus IRB), NEXT Oncology (Salus IRB) and UNC Lineberger Comprehensive Cancer Center (Advarra).
The study was preregistered with ClinicalTrials.gov (NCT05929235) on 3 July 2023 at the following link: https://clinicaltrials.gov/study/NCT05929235.
Protocol amendments
Over the course of the study, changes to the protocol were implemented through amendments that were implemented at the sites and reviewed with IRBs and patients. A summary of the amendments is provided in the Supplementary Information (Summary of FX-909-CLINPRO-1 Protocol Amendments).
Study patients
Approximately 24 patients were to be enrolled in part A (dose escalation) of the study (Supplementary Fig. 2). The Sponsor chose to backfill approximately 30 patients with UC in part A. The study was originally planned to dose participants from 20 mg to 500 mg. The 30 mg and 70 mg dose levels were not specified in the dose escalation table in the protocol. However, in all versions of the protocol, the protocol indicates ‘based on the emerging data and SRC recommendations, the Sponsor may choose to explore an alternative dosing schedule (eg, BID), intermittent dosing (eg, 21 days on, 7 days off), and/or intermediate dose levels.’ Exploring the 30 mg and 70 mg dose levels was approved by the SRC and the Steering Committee during the conduct of part A. As of amendment 6, the protocol indicates that part B will explore dose levels 30 mg and 50 mg (on the basis of the results of part A).
Inclusion criteria
-
(1)
Able to understand and willing to provide informed consent. A legally authorized representative (for example, parent or legal guardian) may consent on behalf of a patient who is otherwise unable to provide informed consent, if acceptable to and approved by the site and/or site’s IRB or ethics committee (EC).
-
(2)
Age ≥18 years.
-
(3)
Eastern Cooperative Oncology Group performance status (ECOG PS) 0, 1 or 2.
-
(4)
An archival, paraffin-embedded, formalin-fixed tumor sample that in part A is no more than 30 months old at the time of screening. If providing formalin-fixed paraffin-embedded (FFPE) slides, all slides (20–30) must come from the same tumor specimen. If an archival tumor sample is not available or is older than 30 months, the patient must consent to provide a fresh biopsy during screening. Biopsies must be collected from a tumor lesion that has not been previously irradiated (or has progressed following radiation therapy), and tumor lesions planned for biopsy must not be followed as target lesions for disease assessments unless the biopsy occurred before the baseline/screening tumor assessment evaluation.
-
(5)
Histologically or cytologically diagnosed, locally advanced (unresectable) or metastatic solid malignancies that have progressed after all available standard therapy for the specific tumor type or for which no standard therapy exists. Patients for whom standard therapies are intolerable or considered inappropriate by the investigator are eligible.
-
(6)
Patients with and without measurable disease (as defined by RECIST version 1.1) will be eligible for enrollment.
-
(7)
Screening laboratory values meet the criteria outlined below.
-
∘ Hematologic criteria may be met with transfusion of blood products or administration of G-CSF, provided that they are not given within 7 days of enrollment. The post-transfusion counts must remain stable without additional transfusion for at least 72 h before C1D1 to confirm sustained hematologic recovery.
-
∘ Hematologic
-
Absolute neutrophil count: ≥1.5 × 109 per liter
-
Hemoglobin: ≥8.5 g dl−1
-
Platelets: ≥150 × 109 per liter
-
-
∘ Clinical chemistry
-
Fasting triglycerides: ≤200 mg dl−1 (may be achieved with lipid lowering medication, such as fibrates)
-
Fasting glucose: ≤140 mg dl−1 (may be achieved with oral hypoglycemic agents (not thiazolidinediones)).
-
Hemoglobin A1c (HbA1c): ≤7.0%
-
-
∘ Renal
-
Creatinine clearance (calculated): ≥60 ml min−1 or serum creatinine: ≤1.5 × upper limit of normal (ULN)
-
-
∘ Hepatic
-
Total bilirubin: ≤1.5 × ULN (except patients with Gilbert’s syndrome who must have total bilirubin ≤3 mg dl−1)
-
AST and ALT: both ≤2.5 × ULN (except patients with liver metastases or tumor infiltration where the limits are ≤5 × ULN)
-
-
∘ Cardiac
-
Ejection fraction: ≥40% by echocardiogram (a multigated acquisition scan is acceptable if echocardiography is not the standard practice at the clinical site)
-
-
Exclusion criteria
-
(1)
Female patients who are pregnant (confirmed with a positive pregnancy test) or breastfeeding. Female patients of childbearing potential who engage in heterosexual intercourse and male patients who are sexually active with female partners of childbearing potential must agree to use a highly effective form of contraception (for example, women: male partner sterilization, estrogen plus progestogen or progestogen-only hormonal contraceptives associated with inhibition of ovulation (oral, intravaginal, transdermal, depot or implant), intrauterine devices or intrauterine hormone-releasing systems; men: male condoms or vasectomy) throughout the study, starting at the time of consent and for at least 90 days after the last dose of the study drug. Female patients of childbearing potential are those who have begun menstruating. To be considered not of childbearing potential, female patients must have had a hysterectomy or bilateral oophorectomy or be 1 year post-menopausal or have had amenorrhea for a period of 12 months or longer in the absence of chemotherapy, anti-estrogens or ovarian suppression. Male patients must not donate sperm throughout the study period and for 90 days after the last dose of FX-909.
-
(2)
Previous anticancer chemotherapy or small-molecule targeted therapy, either investigational or commercially approved and available, within 2 weeks or five half-lives (whichever is shorter) before the start of study drug administration. When the most recent therapy was a biological therapy (including antibody–drug conjugates), an immune checkpoint inhibitor (for example, anti-PD(L)1 or anti-CTLA4) or immune agonist, patients should wait 4 weeks before starting therapy with FX-909 (see exclusion criterion 6 for required radiotherapy windows).
-
(3)
Previous therapy directly inhibiting PPARγ or RXRA. Note that previous treatment with rosiglitazone or pioglitazone for type 2 diabetes would not preclude participation in this study, provided that there is not a need for ongoing treatment with a thiazolidinedione during the study. If given, these agents must have been discontinued at least 2 weeks before the start of study drug administration.
-
(4)
AEs from previous therapy that have not returned to baseline or stabilized at grade 1 (except alopecia, hearing loss, vitiligo, endocrinopathy managed with replacement therapy and grade ≤2 neuropathy) before study drug administration.
-
(5)
Previous major surgery (excluding placement of vascular access) within 4 weeks before study drug administration. Patients must have fully recovered from surgery or its complications before the start of study treatment.
-
(6)
Previous radiation therapy with an inadequate washout between the last dose and the start of the study drug, defined as follows: (1) at least 2 weeks for palliative radiation to the extremities for osseous bone metastases are required and (2) at least 4 weeks for radiation to nonextremity sites are required.
-
(7)
History of another malignancy in the previous 2 years, unless cured by surgery alone and continuously disease free. Exceptions include appropriately treated carcinoma in situ of the cervix, nonmelanoma skin carcinoma, melanoma in situ status post full-thickness resection without recurrence, stage 1 uterine cancer, localized prostate cancer that has been treated surgically with curative intent and presumed cured or other malignancies with an expected curative outcome. Patients requiring adjuvant therapy within the past 2 years for another malignancy will not be considered to have been cured.
-
(8)
QT interval corrected using Fridericia’s formula (QTcF) >470 ms in screening, congenital long QT syndrome, family history of long QT syndrome or unexplained sudden death under 40 years of age in first-degree relatives. QTcF is QT corrected for heart rate according to Fridericia’s correction formula (QTcF = QT/RR0.33) and can be machine calculated or manually overread.
-
(9)
Known active diagnosis of lipodystrophy/lipoatrophy or an ongoing need to receive medications known to cause lipodystrophy/lipoatrophy. Screening for lipodystrophy/lipoatrophy without a known previous diagnosis is not necessary for enrollment in the study.
-
(10)
Any active uncontrolled systemic bacterial, viral or fungal infection requiring treatment.
-
(11)
Known history of human immunodeficiency virus seropositivity. Those who have no detectable viral load on highly active antiretroviral therapy are permitted.
-
(12)
Patients with chronic hepatitis B virus infection (indicated by a positive hepatitis B virus surface antigen and/or hepatitis B core antibody). Patients are permitted with either universal prophylaxis or a pre-emptive treatment approach consistent with regional or national guidelines for patients who receive anticancer therapies.
-
(13)
Active hepatitis C virus infection. Those who have completed curative therapy for hepatitis C virus and have no detectable viral load are permitted.
-
(14)
Previous diagnosis of chronic or recurrent (more than one episode) pancreatitis at any time or a diagnosis of acute pancreatitis within the 6 months before screening (note that elevations of amylase or lipase in the absence of clinical signs and symptoms that are consistent with a diagnosis of pancreatitis do not meet the criteria for exclusion).
-
(15)
Significant impairment of lung function indicated by resting oxygen saturations below 92% on room air or requiring chronic use of ambulatory supplemental oxygen.
-
(16)
Uncontrolled or symptomatic central nervous system metastases, leptomeningeal disease or carcinomatous meningitis. Asymptomatic brain metastasis is allowed if they have been stable after appropriate radiotherapy for 1 month.
-
(17)
Need for treatment with high doses of oral or intravenous steroids (>10 mg per day prednisone or equivalent). Physiologic doses of corticosteroids for treatment of endocrinopathies may be continued if the patient is on a stable dose for at least 1 month.
-
(18)
Need or anticipated need for treatment with a prohibited therapy during the treatment phase of this study.
-
(19)
Concurrent participation in any other investigational therapeutic study.
-
(20)
History of any of the following cardiovascular diseases:
-
Recent history (within the 6 months before screening) of serious uncontrolled cardiac arrhythmia (including atrial fibrillation without adequate rate control) or clinically significant electrocardiogram abnormalities including second-degree (type II) or third-degree atrioventricular node block
-
Documented cerebrovascular event (stroke or transient ischemic attack), cardiomyopathy, myocardial infarction, acute coronary syndromes (including unstable angina pectoris), coronary angioplasty, stenting or bypass grafting within the 6 months before enrollment
-
Congestive heart failure (class III or IV) as defined by the New York Heart Association functional classification system
-
Recent history (within the past 6 months) of symptomatic pericarditis
-
-
(21)
Thromboembolic events and/or bleeding disorders ≤28 days (for example, deep vein thrombosis or pulmonary embolism) before the first dose of the study drug.
-
(22)
Any evidence of severe or uncontrolled systemic diseases, including uncontrolled hypertension (systolic blood pressure ≥160 mmHg and/or diastolic blood pressure ≥100 mmHg) and active bleeding diatheses, which, in the investigator’s opinion, makes it undesirable for the patient to participate in the study or would jeopardize compliance with the protocol. Screening for chronic conditions is not required.
-
(23)
Patients with type 1 diabetes mellitus or type 2 diabetes mellitus that is not adequately controlled with diet, exercise or oral hypoglycemic agents and/or injectable agents other than insulin (as defined by HbA1c and fasting plasma glucose criteria above). Patients taking insulin are excluded from the study. Medication for type 2 diabetes mellitus should have remained stable for the past 14 days before screening.
-
(24)
Known hypersensitivity to FX-909 or any of its excipients.
-
(25)
Patients with gastrointestinal disorders that may interfere with the ability to swallow tablets or absorb study medication.
-
(26)
Patient is or has an immediate family member (for example, spouse, parent or legal guardian, sibling or child) who is a member of the study site or sponsor staff directly involved with this study, unless prospective IRB/EC approval (by chair or designee) is given allowing exception to this criterion for a specific patient.
-
(27)
Patients with any psychological, familial, sociological or geographical condition potentially hampering compliance with the study protocol and follow-up schedule; those conditions should be discussed with the patient before study entry.
-
(28)
Any condition that, in the opinion of the investigator, would interfere with evaluation of the investigational product or interpretation of the patient’s safety or study results.
End points
The primary end points are incidence of DLTs and incidence and severity of AEs and SAEs as reported in this publication.
The secondary end points are: the totality of the safety and tolerability, pharmacokinetic and pharmacodynamic data for FX-909; plasma pharmacokinetic parameters of FX-909, as applicable, including AUC, Cmax, time to maximum plasma concentration (Tmax) and terminal elimination half-life (t½); urine pharmacokinetic parameters of FX-909, as applicable, including renal clearance and percentage of FX-909 in urine; objective response rate (ORR), duration of response (DOR), time to response, disease control rate (DCR) and progression-free survival based on investigator assessment using RECIST version 1.1 criteria; and overall survival. DOR, progression-free survival, overall survival, time to response and urine pharmacokinetics are not reported in this manuscript, as these end points are part of the final analysis at the end of the ongoing phase 1 study.
Exploratory end points reported in this manuscript include: tumor subtyping, PPARγ gene expression, tumor microenvironment and mutations associated with treatment response using genomic and transcriptomic next-generation sequencing (NGS), IHC and/or immunophenotypic assays and methods; longitudinal monitoring of genomic (acquired mutations and tumor burden) changes by ctDNA using a liquid NGS assay; and evaluation of biomarker associations with preliminary antitumor activity.
Treatment
The study drug was administered under a fasted state (that is, no food 2 h before the dose and 1 h after each dose). The cohorts received the study drug QD at doses of 30 mg, 50 mg, 70 mg and 100 mg.
A total of three patients were enrolled into an open cohort. If no DLTs were observed, the next three patients were enrolled at the next highest dose level. If one DLT was observed, three additional patients were entered at the same dose level until six patients were enrolled and completed the DLT observation period. If ≥2 DLTs were observed in any cohort, the next patient was entered at a lower dose level following a review by the safety review committee (SRC) unless six patients had already been dosed at the next lower dose level with ≤1 DLT.
The DLT observation period was 28 days or until the first dose of cycle 2 if cycle 2 was delayed. During the DLT observation period, dose interruptions were allowed but dose reductions were not. After the DLT observation period, both dose interruptions and dose reductions were allowed. The dose modification guidelines and adverse event management algorithms are outlined in the clinical trial protocol.
Safety assessments
All patients underwent safety assessments during the treatment period to include nondirective questioning regarding AEs, physical examinations, vital sign assessments, 12-lead electrocardiograms, clinical laboratory assessments and other protocol-specified tests that are deemed critical to the safety evaluation of the study.
An SRC reviewed available safety, tolerability, pharmacokinetic and pharmacodynamic data for the purposes of study oversight and to make recommendations to the sponsor as it related to dose escalation or de-escalation; choice of schedule; maximum tolerated dose determination; patient, cohort or study termination; changing study eligibility criteria; and adapting dose-modification guidelines.
Pharmacokinetics
Blood samples for FX-909 plasma pharmacokinetics were collected on cycle 1 days 1 and 15, before the dose and at 0.5, 1, 2, 4, 6, 8 and 24 h after the dose (N = 40) (Supplementary Table 2). Additional pre-dose samples were collected on day 1 of the subsequent cycles.
Plasma concentrations of FX-909 were determined, using a validated high-performance liquid chromatographic–tandem mass spectrometry method. In brief, plasma samples were fortified with an internal standard (deuterated analog) and extracted by protein precipitation. Following liquid chromatographic separation, tandem mass spectrometry detection was performed at mass transitions of m/z 361.0 → 282.2 for FX-909 under atmospheric pressure chemical ionization, in positive ion mode. The standard curve had a dynamic range of 2–2,000 ng ml−1.
Key pharmacokinetic parameters evaluated by noncompartmental analysis (Phoenix WinNonlin, Certara) included maximum observed plasma concentration (Cmax), Tmax, area under the plasma concentration–time curve from time zero to 24 h after the dose (AUC0–24 h), t½ and accumulation ratio.
Pharmacodynamics
Paired skin punch biopsies were collected from patients at screening and cycle 1 day 15 (±2 days), after the dose (N = 41) (Supplementary Table 3). Biopsies were taken from the patient’s abdomen, upper arm or shoulder, with baseline and C1D15 taken from the same anatomical location. RNA was isolated from FFPE skin punch biopsies with the RNeasy DSP FFPE Kit (Qiagen, cat. no. 73604) and quantified using the Qubit RNA High Sensitivity Assay Kit (ThermoFisher, cat no. Q32852). Gene expression for nine PPARγ target genes (FABP4, RBP7, APOC1, IGFL3, ISG15, PLIN4, ANGPTL4, CD36 and IL7) was measured on samples with RNA yield >1,000 ng (N = 35) with a custom QuantiGene Plex Gene Expression Assay kit (ThermoFisher, cat no. QGP-180-M23052501) on the Luminex2000 (ThermoFisher). See the “Pharmacodynamic statistical analysis” section for additional details.
Exploratory biomarker assessments
Pre-treatment baseline archival tumor tissue or a fresh tumor biopsy was collected from patients with UC at screening to evaluate potential predictive biomarkers of response to FX-909 and their association with antitumor activity. Molecular characterization of tumors—including lineage classification and subtyping, gene expression, genetic alterations and tumor microenvironment—using whole exome sequencing (WES) and RNAseq was performed. Blood was collected at baseline and on treatment to enable longitudinal circulating tumor DNA analysis, using a liquid biopsy-based assay.
PPARγ IHC
Pre-treatment baseline tumor tissue was obtained from archival specimens or fresh tumor biopsies collected at screening (patients with UC, N = 41) (Supplementary Table 4). FFPE tumor tissue sections of 5-μm thickness were prepared. Following deparaffinization and antigen retrieval, IHC was performed by Roche CDx CAP/CLIA Laboratory (Tucson) using an IHC antibody for PPARγ (SP500; Roche Diagnostics) and standard chromogenic detection. The negative control was performed using an immunoglobulin-matched rabbit monoclonal negative control antibody (Roche Diagnostics, cat. no. 790-4795). Whole-slide images were reviewed by a board-certified pathologist. PPARγ staining was observed in the nucleus of tumor cells. Nuclear staining was assessed specifically in tumor cells, and stromal and immune elements were excluded from tumor scoring. The nuclear tumor positivity was calculated as the percentage of tumor cells staining at any (1+) intensity in the nucleus. This value is referred to as the TPS throughout the report.
Circulating tumor DNA
Blood samples for ctDNA analysis were longitudinally collected from patients at screening, cycle 2 day 1 and cycle 3 day 1, and at the end of treatment. Plasma and buffy coat were isolated from blood for all patients with UC with baseline screening collections (N = 29) (Supplementary Table 6). Somatic mutations in ctDNA were analyzed using the Caris Assure assay, which sequenced plasma cell-free DNA (cfDNA) across the whole exome using a custom hybridization and/or capture methodology28. Matched buffy coat DNA was sequenced to identify and filter out potential germline variants and variants arising from clonal hematopoiesis. WES data were processed through Caris’ proprietary bioinformatics pipeline28.
Aggregate variant allele frequency (aVAF) was calculated by summing the individual VAFs of clinically reported tumor mutations for each sample, excluding genes on the sex chromosomes (chr. X/Y). The list of ~293 clinically reportable genes included PPARγ, RXRA and FGFR3; PPARγ amplification was not among clinically reportable alterations. When buffy coat DNA was ‘quantity not sufficient’ but plasma cfDNA WES data passed quality control, VAFs were reported for such samples using its filtered cfDNA variant call format (VCF) and other samples’ clinically reported mutations and germline VCF collected for the same patient at other time point(s) (Supplementary Table 6). Liquid biopsy response was evaluated using the pre- to on-treatment percentage change in aVAF on the basis of the proposed LB-RECIST criteria25 (Supplementary Table 7).
Efficacy
The antitumor activity of study treatment was evaluated on the basis of the investigator’s assessment, according to RECIST version 1.129. Baseline tumor assessments were performed up to 28 days before cycle 1 day 1 during screening. Post-baseline tumor assessments were performed every 8 weeks ± 7 days using the same imaging techniques as used for the baseline assessment. Confirmatory disease assessments should be done ≥4 weeks after the first partial or CR. PD will include both clinical and radiographical progression.
As this was a phase 1 dose-escalation study and efficacy was only a secondary end point, no formal futility analyses were planned or conducted.
Molecular real-world data
Tumor tissue collected from 2,685 patients with muscle-invasive UC were sequenced using the Tempus xT assay. Molecular classification as luminal (luminal-papillary, luminal and luminal-infiltrated subtypes) or nonluminal (basal squamous and neuronal subtypes) was performed using non-negative matrix factorization with rank 5 following the Robertson method4, excluding PPARγ from the gene set to eliminate inference bias.
Statistics and reproducibility
During the progress of the study, the statistical analysis plan (SAP) was updated to align with protocol amendments and other changes in study procedures. The following is a summary of the updates made to the SAP.
SAP version 2.0 (7 January 2026)
This version replaced version 1.0 (15 May 2024).
Study design changes
-
Study schema updated to reflect part B randomization between doses and patient pre-selection by PPARγ IHC
-
Part B design changed from Simon two-stage to randomized dose-optimization design
-
Sample size increased from 33 to 40 patients
-
Alternative hypothesis updated from ORR ≥35% to ORR ≥40%
-
New appendix added with type I error and power calculations for the revised design
Analysis populations
-
Response-evaluable analysis sets clarified
-
Additional response-evaluable sets defined for reporting purposes
Safety analyses
-
Baseline safety assessment values may be obtained from unscheduled visits before first dose
-
TEAEs of special interest (TEAESIs) defined by combining preferred terms:
-
∘ Thrombocytopenia: thrombocytopenia or platelet count decreased
-
∘ Fatigue: fatigue, lethargy, malaise, or asthenia
-
∘ Hyperglycemia: hyperglycemia, diabetes or glucose intolerance
-
-
TRAE summaries added to reporting
Safety monitoring
-
Toxicity event definition aligned with protocol amendment
As per the SAP, several analysis sets were defined for this study. The safety analysis set included all patients who received at least one dose of FX-909 and was used for all safety end points except DLT evaluation. The DLT-evaluable analysis set included all part A patients who received at least 75% of planned doses during the DLT observation period or who discontinued treatment owing to AEs and was used to assess the maximum tolerated dose and inform dose selection for expansion cohorts.
The response-evaluable analysis sets involving patients enrolled in part A included the following: patients with UC enrolled in part A who have at least one post-baseline response assessment or discontinued the treatment phase owing to disease progression (including death caused by disease progression), tolerability or toxicity before the first post-baseline response assessment without any disqualifying major protocol deviations; part A PPARγhigh UC: subset of the part A patients with UC with PPARγ TPS score ≥60%; part A genetically altered: subset of the part A patients with UC with alterations in PPARγ, RXRA and FGFR3 genes; part A RXRAm: subset of the part A patients with UC with mutations in the RXRA gene; part A FGFR3m: subset of the part A patients with UC with mutations in the FGFR3 gene.
The pharmacokinetic analysis set included all patients who received at least one dose of FX-909 and had at least one post-dose pharmacokinetic measurement.
Summary statistics for continuous variables include n, median, mean, standard deviation, minimum and maximum. For categorical variables, frequencies and percentages are presented. Graphical displays will be provided as appropriate.
DCR is defined as a best confirmed response of stable disease for at least 16 weeks, PR or CR.
Wherever applicable, confidence intervals are reported using the following methods:
-
ORR and DCR: Clopper–Pearson method for binomial proportions
-
DOR median: Kaplan–Meier method
A patient with mixed responses at different tumor sites (Supplementary Fig. 4) is handled differently in the reporting of efficacy summaries, as below:
-
Counted as PD in confirmed ORR calculation but is counted as a responder in unconfirmed response summaries
-
Counted in DCR because that patient had two consecutive assessments of PR after initial progression
-
Counted as responder in TTR and DOR calculation
-
For DOR, the time of progression after assessment of response is used as the event time
The following coding standards will be used in the analysis:
-
AEs and medical histories: MedDRA version 26.0 or higher
-
Previous and concomitant medications: WHODrug B3 Global, March 2023 or higher
Safety and tolerability were assessed by the incidence and severity of AEs as determined by the NCI CTCAE v5.0.
Safety analysis consisted of summaries for AEs, summarized by system organ class and preferred term for all AEs, SAEs, TEAEs, AEs by maximum severity grade and for DLTs.
Sex was self reported during the study. No analysis based on sex or gender was completed given the small size of the cohort.
All statistical analysis outputs were produced using SAS version 9.4 (or a later version) or R version 4.5.0 in a secure and validated environment.
Pharmacodynamic statistical analysis
After background subtraction and normalization with housekeeping genes, fold change in the nine genes was associated with dose and exposure parameters (Cmax, AUC0–24 h) using
-
Linear mixed models for visit and/or replicate-level data
-
Linear regression models (LMs) for fold change from baseline (and adjusting for baseline expression of the gene)
Pre-determined composites of the nine genes were associated with dose or exposure using Spearman and Pearson correlation tests. These pre-determined composite are: mean log fold change, median log fold change, maximum/minimum log fold change and their standardized versions by standardizing individual genes by median absolute deviation after subtracting the dose group median. FDR was used to adjust P values for multiplicity.
WES
Pre-treatment baseline archival tissue or fresh tumor biopsies were collected from patients with UC at screening (N = 41 total; N = 38 with sufficient tissue sections available; Supplementary Table 8). Dual DNA and RNA isolation from patients’ FFPE tissue sections (N = 27, N = 8 pending) was performed using proprietary method (Discovery Life Sciences); sections containing low tumor area were macrodissected (N = 6). For samples with sufficient DNA (N = 25), WES libraries were generated using NEBNext Ultra II FS, hybridized to IDT V2 Exome panel and sequenced to 100× the average coverage using NovaSeq X Plus (Illumina). For a subset of patient samples (N = 3) (Supplementary Table 8), dual DNA and RNA isolation was performed on microdissected sections and libraries were generated using the MI Cancer Seek assay (Caris) and sequenced using NovaSeq 6000 (Illumina).
WES bioinformatics pipelines were run on the Seven Bridges Platform (Velsera). Trimmed WES reads were aligned to the human reference genome (hg38; GENCODE release 27, GRCh38.p10) using BWA-MEM (v0.7.17) with a mismatch penalty of 3, followed by the GATK Pre-Processing workflow (v4.2.0.0) that includes duplicate marking and base recalibration (BQSR). Somatic copy number variants (CNVs) were analyzed using the consensus of the GATK Somatic CNV Pair workflow (v4.2.5.0) and CNVkit (v0.9.9) in tumor-only mode with platform-matched panel-of-normal reference files generated from nine gold-standard samples available through Google Brain Genomics30. Somatic short variant calling was performed using the GATK Somatic single-nucleotide variants (SNVs) and INDELs (Mutect2) workflow (v4.2.5.0) in tumor-only mode with gnomAD as germline resource VCF and a panel of normal VCF generated from the 1000 Genomes Project (GATK Resource Bundle; hg38 data sets), followed by annotation and effect prediction for variants with PASS filter using SnpEff (v5.1 d). One patient sample was excluded from further analysis owing to low mean coverage (<40 reads per base; Supplementary Table 8). For the 81.5% (22/27) of patients with germline WES data available through the Caris Assure blood test (see ‘Circulating tumor DNA’ in Methods), germline short variants were defined as those having VAF ≥0.3 in ≥50% of buffy coat samples profiled from the same patient at different time points. For the 18.5% (5/27) of patients without matched germline data, germline short variants were inferred using PureCN (v2.6.4).
Tumor mutational burden was defined as the number of somatic nonsynonymous SNVs and indels per callable coding territory (Mb). Specifically, germline-subtracted short variants with moderate or high SnpEff impact, variant reads ≥5, total reads ≥30, tumor log odds (TLOD) >6.3 and tumor purity-adjusted VAF ≥0.05 were counted for tumor mutational burden. Callable coding territory was defined as the number of bases (in Mb) within the target exome regions with total reads ≥30.
RNAseq
Pre-treatment baseline archival tissue or fresh tumor biopsies were collected from patients with UC at screening (N = 41 total; N = 38 with sufficient tissue sections available; Supplementary Table 8). Dual DNA and RNA isolation from patients’ FFPE tissue sections (N = 27, N = 8 pending) was performed using the proprietary method (Discovery Life Sciences). RNAseq libraries were generated using KAPA HyperPrep with RiboErase (Roche) and sequenced to obtain >50 M 100-base pair paired-end reads using NovaSeq X Plus (Illumina). For a subset of patient samples (N = 3), dual DNA and RNA isolation and libraries were generated using the MI Cancer Seek assay (Caris) and sequenced using NovaSeq 6000 (Illumina).
RNAseq bioinformatic pipelines were run on the Seven Bridges Platform (Velsera). RNAseq reads were trimmed for adapter, poly-N and low-quality sequences using fastp. Trimmed clean reads were mapped to the human reference genome (hg38; GENCODE Release 27, GRCh38.p10) with STAR (v2.7.10a) using default parameters and gene-level quantification was performed with RSEM (v1.3.3). For the subset of samples analyzed by MI Cancer Seek (N = 3), data were processed through Caris’ proprietary bioinformatics pipeline and gene-level quantification (TPM) was obtained using Salmon31.
We obtained an independent real-world transcriptomic dataset from tumors from patients with advanced UC (N = 29, Tempus xT RNAseq assay) to develop a method to adjust for differences between the Tempus-reported PPARγ TPM values and those obtained using the RSEM pipeline with the DLS assay. For each sample, RSEM TPMs were renormalized for the subset of genes included in TEMPUS xT (20,061 genes). Next, platform-adjusted PPARγ TPM values (Supplementary Table 10) were obtained by applying an LM, which was developed to associate the renormalized RSEM TPMs with TPM values reported by the TEMPUS assay, using the real-world transcriptomic data processed by both RSEM and TEMPUS pipelines for training. Samples analyzed by MI Cancer Seek (N = 3) were excluded from this analysis. PPARγ mRNA expression values of all clinical study samples are being reported using this method.
Gene signature scores were defined as the mean log2(TPM + 1) mRNA expression of genes comprising each published signature: luminal, basal and squamous markers4; epithelial–mesenchymal transition (EMT) and IFNα signatures from the MSigDB Hallmark gene sets32; B cell, T cell, CD8+ T cell, exhausted CD8+ T cell and macrophage markers33; granulocytic myeloid-derived suppressor cell (MDSC) signature34; and immune checkpoint genes (CD274, CTLA4, HAVCR2, LAG3, PDCD1, PDCD1LG2 and TIGIT). The basal/squamous gene expression profile score was defined as the mean of the basal and squamous gene expression profile scores. To obtain expression values comparable across samples, we computed trimmed mean of M-values (TMM)-normalized TPMs used for signature score calculations as follows. RSEM expected counts were divided by effective lengths to obtain length-normalized abundances, which were scaled across samples using TMM normalization (edgeR; v4.0.16). Scaled values were then renormalized to 1 million per sample to generate TMM-normalized TPMs (Supplementary Table 11). Samples analyzed by MI Cancer Seek (N = 3) were excluded from this analysis.
Molecular classification of UC tumor tissues as luminal (luminal papillary, luminal and luminal-infiltrated subtypes) or nonluminal (basal squamous and neuronal subtypes) was performed by applying the BLCAsubtyping classifier to each sample’s UQ-normalized RSEM expected count or Salmon TPM values using the ‘TCGA’ method (https://github.com/cit-bioinfo/BLCAsubtyping) (ref. 3). For samples analyzed by MI Cancer Seek (N = 3), genes boosted by additional baits were removed from the analysis.
Genetic alteration calling for PPARγ, RXRA and FGFR3
PPARγ amplification was reported on the basis of tissue WES data and defined with a tumor purity-adjusted copy number (CN) ≥3. Samples with <50 reads per base mean on-target coverage across chr. 3 (containing PPARγ) were called ‘indeterminate’ for PPARγ amplification. RXRA hotspot mutations (S427F and S427Y) and FGFR3 activating mutations (S249C, Y373C, R248C and G370C) were reported on the basis of both tissue and ctDNA WES data. Any tumors not containing these pre-specified genetic alterations are called ‘wild type’ as shorthand. For tissue WES, mutations supported by SNVs with VAF >0.05 and TLOD >6.3 were called ‘positive’; otherwise, the gene was called ‘negative’ for mutations of interest if its mean coverage was ≥50 reads per base or ‘indeterminate’ if its mean coverage was lower. For ctDNA WES, mutations comprising aVAF at baseline (see ‘Circulating tumor DNA’ in Methods) were called ‘positive’; otherwise, the gene was called ‘negative’ for mutations of interest if the baseline aVAF was >0 and ‘indeterminate’ if the baseline aVAF was 0. Genetic status was determined from the combination of PPARγ amplification, tissue-based and baseline ctDNA-based RXRA/FGFR3 mutations of interest: patients ‘positive’ for any of the alterations were labeled ‘Gen+’; otherwise, they were labeled ‘Gen−’ (Supplementary Table 10).
FX-909 molecular structure
FX-909 is a covalent inverse agonist that was discovered using a ligand-based approach with a sulfone leaving group19. The molecular structure of FX-909 shown in Supplementary Fig. 1 was generated by Flare Therapeutics using ChemDraw Professional 23.1.1.13 software. FX-909 robustly enforces a conformationally ‘repressive’ state of PPARγ, even in highly activated contexts such as RXRA S427F mutation and PPARγ amplification. FX-909 is a potent, highly selective and powerful suppressor of PPARγ transcriptional activity through enhancement of PPARγ nuclear corepressor binding affinity. FX-909 (CAS 2924573-90-8) has been registered on the National Institutes of Health Development Therapeutics Program data warehouse site, which includes the 2D and 3D chemical structures, and with the FDA Global Substance Registration System (https://pubchem.ncbi.nlm.nih.gov/source/FDA%20Global%20Substance%20Registration%20System%20(GSRS)) v3.1.2. Additional details on the compound are available on PubChem (https://pubchem.ncbi.nlm.nih.gov/compound/fx-909).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used in the post hoc exploratory analyses supporting the findings of the ongoing phase 1 study are available within this Article and its Supplementary Information (including genetic data in Supplementary Tables 9–11). Following completion of the phase 1 study and publication of the full study results, including data from part B, raw and processed genomics and transcriptomic data will be deposited in the appropriate public repositories (for example, SRA, GEO and dbGAP), including all accession codes. This adheres to Good Clinical Practice (GCP) and ethical guidelines for next-generation sequencing (NGS) data collected for patients enrolled in an ongoing study and is in consideration for the conduct of additional exploratory biomarker assessments of relevant patient subsets enrolled in part A and part B as part of end-of-phase 1 final reporting. Requests for further data sharing and/or access will be reviewed by Flare Therapeutics to determine whether the request is subject to intellectual property or confidentiality obligations. Further access to de-identified patient-level data, before completion of the phase 1 study, can be requested via email to the corresponding author (matthew.galsky@mssm.edu) but is subject to availability of the data and its intended use. A signed data access agreement with the sponsors is required before data sharing. Real-world data provided by a third-party vendor (Tempus), in formats agreed on per legal contract, are included in the report here; additional queries for academic, nonprofit purposes can be made via email to the corresponding author(s) for consideration on a case-by-case basis. A response will be provided within 90 days of the request depending on the extent of the request.
Change history
11 March 2026
Since the version of the article initially published, ref. 17 has been updated to read “Tate, T. Pparg signaling controls bladder cancer subtype and immune exclusion. Nat. Commun. 12 6160 (2021)” in the HTML and PDF versions of the article.
References
Siegel, R. L. et al. Cancer statistics, 2025. CA Cancer J. Clin. 75, 10–45 (2025.
Powles, T. et al. Enfortumab vedotin and pembrolizumab in untreated advanced urothelial cancer. N. Engl. J. Med. 390, 875–888 (2024).
Kamoun, A. et al. A consensus molecular classification of muscle-invasive bladder cancer. Ero. Urol. 77, 420–433 (2019).
Robertson, A. G. et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 171, 540–556 (2017).
Liu, C. et al. Pparg promotes differentiation and regulates mitochondrial gene expression in bladder epithelial cells. Nat. Commun. 10, 4589 (2019).
Rochel, N. et al. Recurrent activating mutations of PPARɣ associated with luminal bladder tumors. Nat. Commun. 10, 253 (2019).
Goldstein, J. T. et al. Genomic activation of PPARG reveals a candidate therapeutic axis in bladder cancer. Cancer Res. 77, 6987–6998 (2017).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Korpal, M. et al. Evasion of immunosurveillance by genomic alterations of PPARɣ/RXRα in bladder cancer. Nat. Commun. 8, 103 (2017).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Dempster, J. M. et al. Chronos: a cell population dynamics model of CRISPR experiments that improves inference of gene fitness effects. Genome Biol. 22, 343 (2021).
Pacini, C. et al. Integrated cross-study datasets of genetic dependencies in cancer. Nat. Commun. 12, 1661 (2021).
Biton, A. et al. Independent component analysis uncovers the landscape of the bladder tumor transcriptome and reveals insights into luminal and basal subtypes. Cell Rep. 9, 1235–1245 (2014).
Choi, W. et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 25, 152–165 (2014).
Damrauer, J. S. et al. Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proc. Natl Acad. Sci. USA 111, 3110–3115 (2014).
Hedegaard, J. et al. Comprehensive transcriptional analysis of early-stage urothelial carcinoma. Cancer Cell 30, 27–42 (2016).
Tate, T. Pparg signaling controls bladder cancer subtype and immune exclusion. Nat. Commun. 12, 6160 (2021).
Gaona-Romero, C. et al. Luminal and basal subtypes across carcinomas: molecular programs beyond tissue of origin. Cancers 17, 2720 (2025).
Stuckey, J. I. et al. Discovery and characterization of FX-909, a covalent inverse agonist of PPARG rationally designed to impose a powerful repressive bias in PPARG for the treatment of PPARG/RXRA-activated muscle-invasive urothelial cancers. Preprint at bioRxiv https://doi.org/10.1101/2025.05.08.652848 (2025).
Sims, R. et al. Abstract ND08: discovery of FX-909, a first-in-class inverse agonist of the peroxisome proliferator-activated receptor gamma (PPARG) lineage transcription factor, to potentially treat patients with the luminal subtype of advanced urothelial cancer (UC). Cancer Res. 83, ND08 (2023).
Gjini, E. et al. 537 PPARG amplification is associated with lack of response to anti-PD1 in muscle-invasive urothelial cancer. J. Immunotherap. Cancer 11, A611 (2023).
Motley, W. et al. Peroxisome proliferator-activated receptor gamma (PPARG) status defines the luminal lineage in molecular profiles of advanced urothelial cancers (UC). Eur. J. Cancer 174, S119 (2022).
Gouda, M. A. et al. Liquid Biopsy Response Evaluation Criteria in Solid Tumors (LB-RECIST). Ann. Oncol. 35, 267–275 (2024).
Martelli, V. et al. Clinical validation of liquid biopsy-RECIST (LB-RECIST) in metastatic colorectal cancer (mCRC) patients: findings from the PLATFORM-B study. ESMO Open 10, 105760 (2025).
Milowsky, M. et al. 1318 Phase 1 clinical data show FX-909, a first-in-class oral PPARG inhibitor, drives immune modulation and pro-inflammatory cytokine induction in IO-experienced patients with advanced urothelial carcinoma. J. Immunother. Cancer 13, A1556–A1557 (2025).
Gupta, O., Pradhan, T. & Chawla, G. Structural and functional insights into PPAR-ɣ: review of its potential and drug design innovations for the development of antidiabetic agents. Eur. J. Med. Chem. 302, 118264 (2026).
Loriot, Y. et al. Erdafitinib in locally advanced or metastatic urothelial carcinoma. N. Engl. J. Med. 381, 338–348 (2019).
Magee, D. et al. Characterization of plasma cell-free DNA variants as of tumor or clonal hematopoiesis origin in 16,812 advanced cancer patients. Clin. Cancer Res. 31, 2710–2718 (2025).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Baid, G. et al. An extensive sequence dataset of gold-standard samples for benchmarking and development. Preprint at bioRxiv https://doi.org/10.1101/2020.12.11.422022 (2020).
Domenyuk, V. et al. Clinical and analytical validation of MI Cancer Seek®, a companion diagnostic whole exome and whole transcriptome sequencing-based comprehensive molecular profiling assay. Oncotarget 16, 642–659 (2025).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417 (2015).
Danaher, P. et al. Gene expression markers of tumor infiltrating leukocytes. J Immunother. Cancer 5, 18 (2017).
Cristescu, R. et al. Transcriptomic determinants of response to pembrolizumab monotherapy across solid tumor types. Clin. Cancer Res. 28, 1680–1689 (2022).
Acknowledgements
We thank the patients and their families and caregivers who have participated and continue to participate and support this clinical trial. We also thank the investigators and research staff from all study sites. We thank the sponsor, Flare Therapeutics, for their continued hard work. We also thank the medical writer S. Dobney for her editing support.
Author information
Authors and Affiliations
Contributions
M.D.G. contributed to the study design, patient enrollment, data interpretation and manuscript writing. J.B., G.I., M.I.M. and X.G. contributed to the study design, patient enrollment, data interpretation and manuscript editing. C. Mantia, B.G., D.P.P., D.R., S.G. and I.R.-R. contributed to patient enrollment. M.B., A.J., B.K. and E.G. contributed to the study design, assay development, data acquisition, data interpretation, technical support and manuscript writing. A.J. contributed to the study design, statistical analyses, data interpretation and manuscript writing. Y.M. and M.L.M. contributed to the study design, patient enrollment, data interpretation and manuscript editing. J.T. and C. McCrone contributed to the patient enrollment and study execution. P.A.N. contributed to data interpretation, statistical analysis, technical support and manuscript writing. A.M.C. contributed to data acquisition and data review. A.P.R. contributed to data acquisition and technical support. J.A.M. contributed to assay development and data interpretation.
Corresponding author
Ethics declarations
Competing interests
M.D.G. reports consulting fees from Janssen, Merck, Bristol Myers Squibb, Pfizer, EMD Serono, Gilead Sciences and AbbVie and institutional research funding from Janssen Oncology, Bristol Myers Squibb, Merck, AstraZeneca and Genentech/Roche. J.B. reports consulting fees from Pierre Fabre, Pfizer, Merck, Novartis, AstraZeneca/MedImmune, Bristol Myers Squibb and EMD Serono/Merck; stock ownership in Rainier Therapeutics; honoraria from UpToDate; patents/royalties from UpToDate; and research funding from Pfizer/EMD Serono (institutional) and Pfizer/Gilead. M.B. reports employment, leadership role and stock ownership in Flare Therapeutics. C. Mantia reports consulting fees from AADi, Synthekine, Nextech, Cogent Biosciences and EMD Serono; and institutional research funding from Bristol Myers Squibb. B.G. reports institutional consulting fees from AbbVie, Adaptimmune, Adicet Bio, AIQ Global, Amgen, Arcus Biosciences, Arvinas, AstraZeneca, AVEO, Bayer, Bicycle Therapeutics, Bristol Myers Squibb, Eisai, EMD Serono, Exelixis, Genentech/Roche, Janssen, Merck, Monte Rosa Therapeutics, Novartis, Pfizer, Rondo Therapeutics, Seagen, Specialty Networks, Takeda and Xencor; and institutional research funding from multiple pharmaceutical companies including AstraZeneca, AVEO, Exelixis, Janssen Oncology, Merck, Novartis and Pfizer. G.I. reports consulting fees from Janssen, Mirati Therapeutics, Flare Therapeutics, Loxo/Lilly, AstraZeneca, EMD Serono, and Biohaven Pharmaceuticals; other relationships with DAVA Oncology; and institutional research funding from Janssen, Pfizer, AADi, Loxo/Lilly, and Flare Therapeutics. D.P.P. reports consulting fees from Bayer, Exelixis, Pfizer, Roche, Astellas Pharma, AstraZeneca, Lilly, Amgen, Boehringer Ingelheim, Bristol Myers Squibb, Clovis Oncology, Incyte, Janssen, Pharmacyclics, Seagen, Urogen Pharma, Advanced Accelerator Applications, Ipsen, Bicycle Therapeutics, Mirati Therapeutics, Monopteros Therapeutics, Regeneron, and Gilead Sciences; expert testimony for Celgene and Sanofi; and institutional research funding from multiple pharmaceutical companies. D.R. reports institutional research funding from Flare Therapeutics. S.G. reports consulting fees from Merck, Bristol Myers Squibb, AstraZeneca, Novartis, Pfizer, Astellas, Foundation Medicine, Bayer and Johnson & Johnson. I.R.-R. reports an ownership interest or stock in Next Oncology, Texas Oncology and IQVIA and reports consulting fees from Syneos Health, Werewolf Therapeutics and Cardinal Health. Y.M. reports employment and stock ownership at Flare Therapeutics. A.J. reports employment and stock ownership at Flare Therapeutics. P.A.N. reports employment and stock ownership at Flare Therapeutics. B.K. reports employment and stock ownership at Flare Therapeutics. J.T. reports employment and stock ownership at Flare Therapeutics. A.M.C. reports employment and stock ownership at Flare Therapeutics. C. McCrone reports employment and stock ownership at Flare Therapeutics. A.P.R. reports employment at Flare Therapeutics. J.A.M. reports employment and stock ownership at Flare Therapeutics. E.G. reports employment and stock ownership at Flare Therapeutics; an immediate family member is employed by and holds stock in Takeda Pharmaceuticals. M.L.M. reports employment with and leadership roles at Flare Therapeutics, Syndax and Nuvalent; consulting fees from Syndax, Nuvalent and Auron Therapeutics; patents/royalties from Syndax and Nuvalent; and stock ownership in Flare Therapeutics, Syndax, Nuvalent and Auron Therapeutics. M.I.M. reports stock ownership in Pfizer and Gilead Sciences; other relationships with Elsevier, Prime Education, Research to Practice and OncLive/MJH Life Sciences; and institutional research funding from Merck, Bristol Myers Squibb, Alliance for Clinical Trials in Oncology, ALX Oncology, Novartis, Acrivon Therapeutics, OncoC4, Flare Therapeutics, Loxo/Lilly, Astellas Pharma, Pfizer, Amgen, Roche and G1 Therapeutics. X.G. reports consulting fees from Flare Therapeutics, Loxo/Lilly, Bayer, ADC Therapeutics, Hinova Pharmaceuticals, Abeona Therapeutics, Arvinas, 858 Therapeutics, Janssen and Kanaph Therapeutics; and institutional research funding from Arvinas, Exelixis, Pfizer, Harpoon Therapeutics, Aprea Therapeutics, Takeda, Aravive, Merck, Poseida Therapeutics, TopAlliance BioSciences, Novartis, Regeneron, Bayer, ALX Oncology, Nuvation Bio, Loxo/Lilly, Janux Therapeutics and Halda Therapeutics.
Peer review
Peer review information
Nature Medicine thanks Joshua Meeks and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Ranzoni in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Plasma pharmacokinetics (PK) (N = 32 patients, Suppl. Table 3) and skin pharmacodynamics (PD) (N = 34 patients, Suppl. Table 4) support FX-909 is a pharmacologically active drug.
A, Geometric mean steady-state plasma concentrations of FX-909 vs. time following single oral administrations of FX-909 on C1D15. Error bars represent geometric standard error. B, FABP4 expression measured as % change from baseline in paired skin punch biopsies, collected at screening and C1D15. 1 patient had a % change from baseline that was > 250%. The y-axis has been clipped to 100% for presentation purposes.
Extended Data Fig. 2 Provisional PPARG protein cutoff informed by real-world molecular characterization and on-study biomarker data.
A & B, In the Tempus xT real-world cohort (N = 2,685 patients), PPARG is enriched in luminal tumors and a range of PPARG mRNA thresholds identifies luminal lineage with high sensitivity and PPV (Youden’s J stable from 6.1–7.1 log2[TPM + 1]). C, Linear regression line (red solid line) with pointwise 99% confidence band (shaded region) of baseline tumor PPARG mRNA versus protein expression (N = 26 patients; on-study biomarker data) maps a provisional TPS cutoff of ≥60% to an mRNA range of 6.1–7.1, corresponding to >74.5% sensitivity and >81% PPV. Two-sided p value from Pearson’s correlation test is shown. D, PPARG protein expression is significantly higher in luminal (N = 23) versus non-luminal (N = 6) tumors (on-study biomarker data; see Suppl. Table 10). Boxplots display the median and interquartile range; whiskers denote the 5th–95th percentile range; the + symbol denotes the mean (A, D). Two-sided Wilcoxon rank-sum test was used for statistical analysis in (A, D).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–11, Table 1 and Summary of FX-909-CLINPRO-1 Protocol Amendments.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Galsky, M.D., Mantia, C., Bowden, M. et al. A small-molecule inverse agonist of PPARγ for advanced solid tumors: a phase 1 trial. Nat Med (2026). https://doi.org/10.1038/s41591-026-04263-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41591-026-04263-3