Abstract
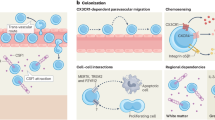
Microglia—resident immune cells in the central nervous system—undergo morphological and functional changes in response to signals from the local environment and mature into various homeostatic states. However, niche signals underlying microglial differentiation and maturation remain unknown. Here, we show that neuronal micronuclei (MN) transfer to microglia, which is followed by changing microglial characteristics during the postnatal period. Neurons passing through a dense region of the developing neocortex give rise to MN and release them into the extracellular space, before being incorporated into microglia and inducing morphological changes. Two-photon imaging analyses have revealed that microglia incorporating MN tend to slowly retract their processes. Loss of the cGAS gene alleviates effects on micronucleus-dependent morphological changes. Neuronal MN-harboring microglia also exhibit unique transcriptome signatures. These results demonstrate that neuronal MN serve as niche signals that transform microglia, and provide a potential mechanism for regulation of microglial characteristics in the early postnatal neocortex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The mass spectrometry proteomics data have been deposited with the ProteomeXchange Consortium via the PRIDE110 partner repository with the dataset identifier PXD056089. The dataset of mouse genome (mm10, GRCm38) is available at http://nov2020.archive.ensembl.org/Mus_musculus/Info/Index. Sequence data have been deposited in the DNA Data Bank of Japan (DDBJ) Sequence Read Archive under the accession code DRA015927. The data in Extended Data Figs. 9e–r and 10l–v can be obtained from the Brain RNA-seq database (https://www.brainrnaseq.org/)58,59. Source data are provided with this paper.
Code availability
The MATLAB CAMDi software is available in Supplementary Software 1.
References
Marquez-Ropero, M., Benito, E., Plaza-Zabala, A. & Sierra, A. Microglial corpse clearance: lessons from macrophages. Front Immunol 11, 506 (2020).
D’Mello, C., Le, T. & Swain, M. G. Cerebral microglia recruit monocytes into the brain in response to tumor necrosis factoralpha signaling during peripheral organ inflammation. J. Neurosci. 29, 2089–2102 (2009).
Chen, X. et al. Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy. Nature 615, 668–677 (2023).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Wolf, S. A., Boddeke, H. W. & Kettenmann, H. Microglia in physiology and disease. Annu Rev Physiol 79, 619–643 (2017).
Ueno, M. et al. Layer V cortical neurons require microglial support for survival during postnatal development. Nat. Neurosci. 16, 543–551 (2013).
Cunningham, C. L., Martinez-Cerdeno, V. & Noctor, S. C. Microglia regulate the number of neural precursor cells in the developing cerebral cortex. J. Neurosci. 33, 4216–4233 (2013).
Aarum, J., Sandberg, K., Haeberlein, S. L. & Persson, M. A. Migration and differentiation of neural precursor cells can be directed by microglia. Proc. Natl Acad. Sci. USA 100, 15983–15988 (2003).
Miyamoto, A. et al. Microglia contact induces synapse formation in developing somatosensory cortex. Nat. Commun. 7, 12540 (2016).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Badimon, A. et al. Negative feedback control of neuronal activity by microglia. Nature 586, 417–423 (2020).
Wake, H., Moorhouse, A. J., Miyamoto, A. & Nabekura, J. Microglia: actively surveying and shaping neuronal circuit structure and function. Trends Neurosci. 36, 209–217 (2013).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Utz, S. G. et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 181, 557–573 e518 (2020).
Hattori, Y. et al. CD206+ macrophages transventricularly infiltrate the early embryonic cerebral wall to differentiate into microglia. Cell Rep. 42, 112092 (2023).
Paolicelli, R. C. et al. Microglia states and nomenclature: a field at its crossroads. Neuron 110, 3458–3483 (2022).
Kierdorf, K., Masuda, T., Jordao, M. J. C. & Prinz, M. Macrophages at CNS interfaces: ontogeny and function in health and disease. Nat. Rev. Neurosci. 20, 547–562 (2019).
Prinz, M., Masuda, T., Wheeler, M. A. & Quintana, F. J. Microglia and central nervous system-associated macrophages-from origin to disease modulation. Annu. Rev. Immunol. 39, 251–277 (2021).
Taketomi, T. & Tsuruta, F. Towards an understanding of microglia and border-associated macrophages. Biology (Basel) 12, 1091 (2023).
Nimmerjahn, A., Kirchhoff, F. & Helmchen, F. Resting microglial cells are highly dynamic surveillants of brain parenchyma in vivo. Science 308, 1314–1318 (2005).
Haynes, S. E. et al. The P2Y12 receptor regulates microglial activation by extracellular nucleotides. Nat. Neurosci. 9, 1512–1519 (2006).
Wake, H., Moorhouse, A. J., Jinno, S., Kohsaka, S. & Nabekura, J. Resting microglia directly monitor the functional state of synapses in vivo and determine the fate of ischemic terminals. J. Neurosci. 29, 3974–3980 (2009).
Mosher, K. I. et al. Neural progenitor cells regulate microglia functions and activity. Nat. Neurosci. 15, 1485–1487 (2012).
Pasciuto, E. et al. Microglia require CD4 T cells to complete the fetal-to-adult transition. Cell 182, 625–640 e624 (2020).
Crasta, K. et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature 482, 53–58 (2012).
Raab, M. et al. ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death. Science 352, 359–362 (2016).
Denais, C. M. et al. Nuclear envelope rupture and repair during cancer cell migration. Science 352, 353–358 (2016).
Fenech, M. et al. Micronuclei and disease—report of HUMN project workshop at Rennes 2019 EEMGS conference. Mutat. Res. 850-851, 503133 (2020).
Shi, L., Qalieh, A., Lam, M. M., Keil, J. M. & Kwan, K. Y. Robust elimination of genome-damaged cells safeguards against brain somatic aneuploidy following Knl1 deletion. Nat. Commun. 10, 2588 (2019).
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Papathanasiou, S. et al. Heritable transcriptional defects from aberrations of nuclear architecture. Nature 619, 184–192 (2023).
Agustinus, A. S. et al. Epigenetic dysregulation from chromosomal transit in micronuclei. Nature 619, 176–183 (2023).
Lin, Y. F. et al. Mitotic clustering of pulverized chromosomes from micronuclei. Nature 618, 1041–1048 (2023).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Gluck, S. et al. Innate immune sensing of cytosolic chromatin fragments through cGAS promotes senescence. Nat. Cell Biol. 19, 1061–1070 (2017).
Yano, S. et al. A MATLAB-based program for three-dimensional quantitative analysis of micronuclei reveals that neuroinflammation induces micronuclei formation in the brain. Sci. Rep. 11, 18360 (2021).
Sekine, K., Honda, T., Kawauchi, T., Kubo, K. & Nakajima, K. The outermost region of the developing cortical plate is crucial for both the switch of the radial migration mode and the Dab1-dependent ‘inside-out’ lamination in the neocortex. J. Neurosci. 31, 9426–9439 (2011).
Tabata, H. & Nakajima, K. Efficient in utero gene transfer system to the developing mouse brain using electroporation: visualization of neuronal migration in the developing cortex. Neuroscience 103, 865–872 (2001).
Kubo, K. et al. Ectopic Reelin induces neuronal aggregation with a normal birthdate-dependent ‘inside-out’ alignment in the developing neocortex. J. Neurosci. 30, 10953–10966 (2010).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Frey, T. et al. Age-associated reduction of nuclear shape dynamics in excitatory neurons of the visual cortex. Aging Cell 22, e13925 (2023).
Rello-Varona, S. et al. Autophagic removal of micronuclei. Cell Cycle 11, 170–176 (2012).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Yamamoto, H., Zhang, S. & Mizushima, N. Autophagy genes in biology and disease. Nat. Rev. Genet. 24, 382–400 (2023).
Ma, X. et al. CCT2 is an aggrephagy receptor for clearance of solid protein aggregates. Cell 185, 1325–1345 e1322 (2022).
Jung, S. et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Abe, T. et al. Establishment of conditional reporter mouse lines at ROSA26 locus for live cell imaging. Genesis 49, 579–590 (2011).
Ginhoux, F., Lim, S., Hoeffel, G., Low, D. & Huber, T. Origin and differentiation of microglia. Front. Cell Neurosci. 7, 45 (2013).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl Acad. Sci. USA 113, E1738–E1746 (2016).
Fritsch, L. E. et al. Type I interferon response is mediated by NLRX1-cGAS-STING signaling in brain injury. Front. Mol. Neurosci. 15, 852243 (2022).
Marsh, S. E. et al. Dissection of artifactual and confounding glial signatures by single-cell sequencing of mouse and human brain. Nat. Neurosci. 25, 306–316 (2022).
Mason, C. E. et al. Standardizing the next generation of bioinformatics software development with BioHDF (HDF5). Adv. Exp. Med. Biol. 680, 693–700 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Li, Q. et al. Developmental heterogeneity of microglia and brain myeloid cells revealed by deep single-cell RNA sequencing. Neuron 101, 207–223 e210 (2019).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Krupina, K., Goginashvili, A. & Cleveland, D. W. Causes and consequences of micronuclei. Curr. Opin. Cell Biol. 70, 91–99 (2021).
Ginhoux, F. & Jung, S. Monocytes and macrophages: developmental pathways and tissue homeostasis. Nat. Rev. Immunol. 14, 392–404 (2014).
Drieu, A. et al. Parenchymal border macrophages regulate the flow dynamics of the cerebrospinal fluid. Nature 611, 585–593 (2022).
Liu, C. et al. Macrophages mediate the repair of brain vascular rupture through direct physical adhesion and mechanical traction. Immunity 44, 1162–1176 (2016).
Masuda, T. et al. Specification of CNS macrophage subsets occurs postnatally in defined niches. Nature 604, 740–748 (2022).
Ponpuak, M. et al. Secretory autophagy. Curr. Opin. Cell Biol. 35, 106–116 (2015).
Ejlerskov, P. et al. Tubulin polymerization-promoting protein (TPPP/p25alpha) promotes unconventional secretion of alpha-synuclein through exophagy by impairing autophagosome-lysosome fusion. J. Biol. Chem. 288, 17313–17335 (2013).
Savina, A., Vidal, M. & Colombo, M. I. The exosome pathway in K562 cells is regulated by Rab11. J. Cell Sci. 115, 2505–2515 (2002).
Hsu, C. et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J. Cell Biol. 189, 223–232 (2010).
Tsuboyama, K. et al. The ATG conjugation systems are important for degradation of the inner autophagosomal membrane. Science 354, 1036–1041 (2016).
Budnik, V., Ruiz-Canada, C. & Wendler, F. Extracellular vesicles round off communication in the nervous system. Nat. Rev. Neurosci. 17, 160–172 (2016).
Christ, L., Raiborg, C., Wenzel, E. M., Campsteijn, C. & Stenmark, H. Cellular functions and molecular mechanisms of the ESCRT membrane-scission machinery. Trends Biochem. Sci 42, 42–56 (2017).
Gong, Y. N. et al. ESCRT-III acts downstream of MLKL to regulate necroptotic cell death and its consequences. Cell 169, 286–300 e216 (2017).
Claude-Taupin, A. et al. ATG9A protects the plasma membrane from programmed and incidental permeabilization. Nat. Cell Biol. 23, 846–858 (2021).
Hayakawa, K. et al. Transfer of mitochondria from astrocytes to neurons after stroke. Nature 535, 551–555 (2016).
Joshi, A. U. et al. Fragmented mitochondria released from microglia trigger A1 astrocytic response and propagate inflammatory neurodegeneration. Nat. Neurosci. 22, 1635–1648 (2019).
Asai, H. et al. Depletion of microglia and inhibition of exosome synthesis halt tau propagation. Nat. Neurosci. 18, 1584–1593 (2015).
Yokoi, A. et al. Mechanisms of nuclear content loading to exosomes. Sci. Adv. 5, eaax8849 (2019).
Decout, A., Katz, J. D., Venkatraman, S. & Ablasser, A. The cGAS-STING pathway as a therapeutic target in inflammatory diseases. Nat. Rev. Immunol. 21, 548–569 (2021).
Gulen, M. F. et al. cGAS-STING drives ageing-related inflammation and neurodegeneration. Nature 620, 374–380 (2023).
Paul, B. D., Snyder, S. H. & Bohr, V. A. Signaling by cGAS-STING in neurodegeneration, neuroinflammation, and aging. Trends Neurosci. 44, 83–96 (2021).
Luteijn, R. D. et al. SLC19A1 transports immunoreactive cyclic dinucleotides. Nature 573, 434–438 (2019).
Ritchie, C., Cordova, A. F., Hess, G. T., Bassik, M. C. & Li, L. SLC19A1 is an importer of the immunotransmitter cGAMP. Mol Cell 75, 372–381.e375 (2019).
Carozza, J. A. et al. Extracellular cGAMP is a cancer cell-produced immunotransmitter involved in radiation-induced anti-cancer immunity. Nat Cancer 1, 184–196 (2020).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
Hattori, Y. et al. Transient microglial absence assists postmigratory cortical neurons in proper differentiation. Nat. Commun. 11, 1631 (2020).
Song, Y. et al. Transient nuclear deformation primes epigenetic state and promotes cell reprogramming. Nat. Mater. 21, 1191–1199 (2022).
Moriyama, M., Koshiba, T. & Ichinohe, T. Influenza A virus M2 protein triggers mitochondrial DNA-mediated antiviral immune responses. Nat. Commun. 10, 4624 (2019).
Sekine, K. et al. Reelin controls neuronal positioning by promoting cell-matrix adhesion via inside-out activation of integrin alpha5beta1. Neuron 76, 353–369 (2012).
Shinoda, T. et al. Elasticity-based boosting of neuroepithelial nucleokinesis via indirect energy transfer from mother to daughter. PLoS Biol. 16, e2004426 (2018).
Goebbels, S. et al. Genetic targeting of principal neurons in neocortex and hippocampus of NEX-Cre mice. Genesis 44, 611–621 (2006).
Okajima, T. et al. Atypical cadherin FAT3 is a novel mediator for morphological changes of microglia. eNEuro 7, ENEURO.0056-20.2020 (2020).
Kim, J. et al. USP15 deubiquitinates TUT1 associated with RNA metabolism and maintains cerebellar homeostasis. Mol. Cell. Biol. 40, e00098-20 (2020).
Tsuruta, F., Okajima, T., Yano, S. & Chiba, T. Quantification of endosome and lysosome motilities in cultured neurons using fluorescent probes. J. Vis. Exp. 123, 55488 (2017).
Tabata, H. & Nakajima, K. Labeling embryonic mouse central nervous system cells by in utero electroporation. Dev. Growth Differ. 50, 507–511 (2008).
Morrison, H., Young, K., Qureshi, M., Rowe, R. K. & Lifshitz, J. Quantitative microglia analyses reveal diverse morphologic responses in the rat cortex after diffuse brain injury. Sci. Rep. 7, 13211 (2017).
Fenech, M. et al. HUMN project: detailed description of the scoring criteria for the cytokinesis-block micronucleus assay using isolated human lymphocyte cultures. Mutat. Res. 534, 65–75 (2003).
Mure, K., Takeshita, T. & Morimoto, K. Categorization of micronuclei by size and measurement of each ratio in cytokinesis-block and conventional cultures of human lymphocytes exposed to mitomycin C and colchicine. Environ. Health Prev. Med. 1, 93–99 (1996).
Kishi, Y. & Gotoh, Y. Isolation of genetically manipulated neural progenitors and immature neurons from embryonic mouse neocortex by FACS. STAR Protoc. 2, 100540 (2021).
Haruwaka, K. et al. Dual microglia effects on blood brain barrier permeability induced by systemic inflammation. Nat. Commun. 10, 5816 (2019).
Takeda, I. et al. Controlled activation of cortical astrocytes modulates neuropathic pain-like behaviour. Nat. Commun. 13, 4100 (2022).
Kim, J. et al. Graph-Based deep learning for prediction of longitudinal infant diffusion MRI data. Comput. Diffus. MRI 2019, 133–141 (2019).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE Blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
ENCODE Project Consortium An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Acknowledgements
We thank laboratory members for helpful discussions and technical support. We thank M Kengaku (iCeMS, Kyoto University) for technical advice of the neuronal migration experiments, M. Urushitani (Shiga University of Medical Science) for distributing BV2 cells and S. Hattori (NIMS) for technical support of mass spectrometry analysis. We thank L.J. Irving (University of Tsukuba) for editing the paper. The mass spectrometry analysis was supported by ‘Nanotechnology Platform Project’ operated by the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (grant no. JPMXP09S19NM0031). We wish to acknowledge the Division for Medical Research Engineering, Nagoya University Graduate School of Medicine, for analysis with IMARIS. This study was supported by the Cooperative Study Program (24NIPS330) of National Institute for Physiological Sciences. This work was supported by Grant-in-Aid from the Ministry of Education, Science, Sports and Culture of Japan JSPS KAKENHI [16KK0158 (F.T.), 20K05951 (F.T.), 23H04214 (Y.K.), 24K02020 (Y.K.), JP20H05688 (K.N.), JP22K19365 (K.N.), JSPS Research Fellowship for Young Scientists (19J20619 (S.Y.), 23KJ0285 (T.T.)], AMED PRIME [24028934 (F.T.)], Kao foundation for health science (F.T.), Gout and uric acid foundation (F.T.), Asahi Glass Foundation (Y.K.), SECOM Science and Technology Foundation (Y.K.) and was partly supported by Center for Quantum and Information Life Sciences (F.T.).
Author information
Authors and Affiliations
Contributions
S.Y. and F.T. designed the research. S.Y., N.A., Y.K., I.T., H.K., Y.H., A.K., K.H., K.-i.K., C.M., C.H., R.i.T., T.T., K.A., T.O-T., B.S. and F.T. performed research. S.Y., N.A., Y.K., I.T., H.K., M.S., C.H., C.M., R.R., T.T., K.A., T.N. and F.T. analyzed the data. T.N developed the MATLAB program. Y.K., Y.H., H.W., Y.G. and T.I. provided materials. S.Y. and F.T. wrote the paper. H.W., Y.G., K.N., T.C. and F.T. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Lucas Cheadle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Physical stress promotes micronuclear formation.
(a) Immunostaining of MAP2+ neurons. Primary cultured neurons (5 DIV) were stimulated with the recombinant 100 ng/ml Reelin for 6 hours. (b) The graph shows the percentage of MN+ neurons. 10 fields, n=3 independent samples, The numbers represent the total counted DAPI+ cells. mean ± SEM, p value was calculated by two-tailed Student's t-test. (c) Schematic illustration for injecting recombinant Reelin. (d) The recombinant Reelin (100 ng) was stimulated into the mice brain (P7) for 6 hours, and the brain sections were stained using DAPI. (e) The graph shows the percentage of MN. 15 images obtained from three independent brain, mean ± SEM, p value was calculated by two-tailed Student's t-test. (f) Schematic illustration for an in vitro migration assay. (g) Immunostaining of migrated neurons at the bottom sides of the trans-well. Nuclear shapes were detected by Lamin B1 staining. Asterisks indicate the marks of the hole. Arrow indicates the micronucleus. Illustration indicates the condition of migration, herniation, and micronuclear formation. (h) Immunostaining of migrated neurons at the bottom side of the trans-well. Yellow arrows indicate MN. (i)(j) The graph shows the population of MN+ neurons.(i) n=6, (j) n=3 trans-wells. More than 200 neurons were analyzed from one trans-well. (i) pore size; 3 μm, (j) pore size 12 μm, mean ± SEM, p value was calculated by two-tailed Student's t-test. (k) Schematic illustration for an in vitro mechanical stress assay. Neuro2A cells received mechanical stresses by pumping. (l) Neuro2A cells were stained with Hoechst. MN were generated by mechanical stress. (m) The graph shows the percentage of population of MN+ cells. Low; 5.0 x 104 cells/300μl, High; 5.0 x 104 cells/150μl, n=9 images, mean ± SEM, p values were calculated by one-way ANOVA Dunnett's multiple comparison test.
Extended Data Fig. 2 The autophagy pathway is involved in micronuclear secretion.
(a) Immunostaining of MAP2+ neurons in the cerebral cortex of either WT or NexCre:Atg7f/f at P21. Yellow arrows indicate MN. (b) The graph shows the population of MN in the cerebral cortex. n=20 images, 10 images obtained from two independent brain; mean ± SEM, p value was calculated by two-tailed Student's t-test. (c) Immunostaining of Iba1+ microglia in the cerebral cortex of either WT or NexCre:Atg7f/f at P21. Yellow arrows indicate MN. (d) The graph shows the percentage of MNs+ microglia in the cerebral cortex. Data were combined from 3 independent brains (8 images per brain, total 24 images), mean ± SEM, p value was calculated by two-tailed Student's t-test. (e) Immunoblotting of the extracellular MN (P15) obtained from primary cortical neurons in the presence or absence of 100 nM BafA treatment. Equal amounts of conditioned medium and total cell lysates were loaded onto separate gels and subjected to SDS-PAGE, followed by western blot analysis using anti-tubulin and anti-Lamin B1 antibodies, respectively. (f) Primary cortical neurons were stimulated with either 300 nM BafA for 3 hours or 50 μM etoposide (Eto, positive control for inducing apoptosis) for 24 hours. Immunostaining of cleaved-caspase 3 and MAP2+ cortical neurons (5 DIV). (g) The graph shows the fluorescence intensity of cleaved-caspase 3 with MAP2. n=4 images (DMSO) and 5 images (BafA and Eto), mean ± SEM, p values were calculated by one-way ANOVA Dunnett's multiple comparison test. (h) Schematic illustration of the collection of extracellular micronuclear for immunocytochemistry. (i) Immunostaining of Tubulin and Rab35 resided in the extracellular MN. (j) Immunostaining of transfected TCP subunits in the migrated neurons at the bottom side of the trans-well (3 DIV). Yellow arrows indicate MN. (k) Immunostaining of GFP+-MN in the cerebral cortex. A small amount of Tubulin resided in GFP+-MN. P14. Yellow allow indicates Tubulin+ signal. (l) Immunostaining of GFP+ MN in the cerebral cortex. MAP2 and LC3 resided in GFP+-MN. P14. (m) Working hypothesis of micronuclear secretion dependent on autophagy-lysosome machinery. TCP subunit is a potential receptor linking the micronucleus to LC3.
Extended Data Fig. 3 Micronuclei are incorporated into microglia.
(a) Schematic illustration for an in vitro micronuclear transfer assay. (b) TEM image of BV2 cells after treatment with conditioned medium. NE: nuclear envelope, MN: micronucleus, Cyto: cytoplasm. (c) Immunostaining of BV2 cells with GFP−-and GFP+-MN. (d) Immunostaining of GFP−- and GFP+-MN in BV2. Yellow arrow indicates micronucleus. (e) The graph shows the percentage of neuronal MN (GFP+ MN) containing BV2 cells after treatment with a conditioned medium. Each spot indicates the combined value of 10 images from one experiment. n=3 independent experiments. NB; fresh neurobasal medium, mean ± SEM, p values were analyzed by one-way ANOVA Dunnett's multiple comparisons test. (f) Schematic illustration for eliminating MN. (g) The graph shows MN+ BV2 cells 6 hours after treatment with a conditioned medium. mean ± SEM, p value was calculated by one-way ANOVA Tukey's multiple comparisons tests. Each spot indicates the combined value of more than 9 images from one experiment. Each column indicates a set of 3 independent experiments. (h) Live imaging of Cx3cr1-EGFP;H2B-mCherry cortical slice culture (P3). The interval of taking images was every 5 min. Yellow arrowheads indicate MN in the microglia. Cyan asterisks indicate MN out of the microglia. No.1-6, 89 z-stack images/field, No. 7-24, 92 z-stack images/field, 1.5 µm pitch.
Extended Data Fig. 4 Microglial morphology differs based on the cortical layer.
(a) Tailing immunostained images of Iba1+ microglia in the cerebral cortex. WT, P14. (b) Representative images of microglia in Layer 1 (L1) and Layers 2-6 (L2-6) from the insets of Extended Data Fig. 4a. (c)(d) The graphs show the analyses of microglial characters. 3 brains. (c) Process length, (d) the number of the process. mean ± SEM, p value was calculated by two-tailed Student's t-test.
Extended Data Fig. 5 Involvement of micronuclei on microglial gene expression.
(a) Immunostaining of Iba1+ and CD68+ microglia with or without MN (MN−; micronuclei-negative, MN+; micronuclei-positive). NexCre:LSL-Sun1GFP mice, P14. (b) The graph shows the expression level of CD68. The numbers counted are shown in the graph. 5 brains. mean ± SEM, p values were analyzed by one-way ANOVA Dunnett's multiple comparisons test. (c) Immunostaining of Iba1+ and Tmem119+ microglia with or without MN. NexCre:LSL-Sun1GFP mice, P14. (d) The graph shows the expression level of Tmem119. The numbers counted are shown in the graph. 3 brains. mean ± SEM, p values were analyzed by one-way ANOVA Dunnett's multiple comparisons test. (e) Immunostaining of Iba1+ and P2ry12+ microglia with or without MNs. The numbers counted are shown in the graph. NexCre:LSL-Sun1GFP mice, P14. (f) The graph shows the expression level of P2ry12. 5 brains, mean ± SEM, p values were analyzed by one-way ANOVA Dunnett's multiple comparisons test.
Extended Data Fig. 6 Two-photon imaging of microglial dynamics obtained from two distinct mice.
(a) to (j'') Quantification of individual microglial morphology. The line graph shows the change in the length of individual processes. Specific processes were measured every minute for one hour. The red outlines present MN+ processes. The violin plot shows the distribution of the lengths of individual processes in the line graph. The red outlines present MN+ processes. The blue and red line indicates the median and quartiles. The stack bars graph indicates the type and the number of individual processes (primary, secondary, tertiary, and quaternary processes) at 0, 30, and 60 minutes. The individual dot plots of stack bar graphs are shown in Supplementary Fig. 1. mean±SEM. Blue title: MN− microglia, n=10 cells, Red title: MN+ microglia, 10 each microglia obtained from 2 individual mice (mice identification number; Mo1 and Mo2).
Extended Data Fig. 7 Two-photon imaging of microglial dynamics obtained from three distinct mice.
(a) to (j'') Quantification of individual microglial morphology. The line graph shows the change in the length of individual processes. Specific processes were measured every minute for one hour. The red outlines present MN+ processes. The violin plot shows the distribution of the lengths of individual processes in the line graph. The red outlines present MN+ processes. The blue and red line indicates the median and quartiles. The stack bars graph indicates the type and the number of individual processes (primary, secondary, tertiary, and quaternary processes) at 0, 30, and 60 minutes. The individual dot plots of stack bar graphs are shown in Supplementary Fig. 2. mean±SEM. Blue title: MN− microglia, n=10 cells, Red title: MN+ microglia, 10 each microglia obtained from 3 individual mice (mice identification number; Mo2, Mo3, and Mo4).
Extended Data Fig. 8 cGAS is involved in morphological changes of microglia.
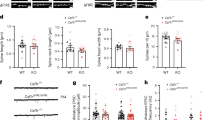
(a) Immunostaining of Iba1+ microglia in the cerebral cortex of either WT or cGAS−/− mice at P6. (b)-(d) The graphs show the analyses of microglial state. 2 brains. The numbers in the graph represent the numbers of cells counted. (b) The number of processes, (c) process length, (d) expression level of CD68. mean ± SEM, p values were analyzed by two-tailed Student's t-test. (e) Immunostaining of CD68 in Iba1+ microglia of either WT or cGAS−/− mice at P6. (f) The graph show the expression level of CD68. The numbers in the graph represent the numbers of cells counted. mean ± SEM, 2 brains. p values were analyzed by one-way ANOVA Tukey's multiple comparisons test. (g) Immunostaining of CD68 in Iba1+ microglia of either WT or cGAS−/− mice, MN−; micronuclei-negative, MN+; micronuclei-positive. P14. (h) The graph show the expression level of CD68. The numbers in the graph represent the numbers of cells counted. mean ± SEM, 2 brains. p values were analyzed by one-way ANOVA Tukey's multiple comparisons test.
Extended Data Fig. 9 Effect of micronuclei on gene expression in microglia.
(a) Representative gating strategy for microglia in NexCre:LSL-Sun1GFP mice. (b) The GO enrichment analysis of DEG in GFPHigh MN+ microglia (1,663 genes). Data was analyzed by Enrichr analysis tool. (c) The scatter plot indicates the ratio and differences of FPKM between GFPNeg MN− microglia and GFPHigh MN+ microglia. FPKM: GFPHigh/GFPNeg>5, GFPHigh-GFPNeg>200, (d) The Heat map indicates the comparison of FPKM among GFPNeg MN− microglia, GFPLow MN+ microglia, and GFPHigh MN+ microglia. FPKM: GFPHigh/GFPNeg>5, GFPHIgh-GFPNeg>200 (e)-(r) The graph shows the expression of each gene obtained from bulk-RNAseq analysis in GFPHigh MN+ microglia [ASC; astrocyte, NEUR; neuron (red), OPC; oligodendrocyte precursor cell, iOLG; immature oligodendrocyte, mOLG; myelinating oligodendrocyte, MG/MAC; microglia/macrophage (blue), EC; Endothelial cell, All others except NEUR and MG/MAC are shown in the black bar. mean, n=2]. This data was obtained from the Brain RNA-Seq database (https://www.brainrnaseq.org/).
Extended Data Fig. 10 Changing microglia-related genes by micronuclei transfer.
(a)-(k) The bar graphs show the value of FPKM obtained from bulk-RNAseq analysis in GFPLow MN+ microglia (P8, 3 mice; P9, 3 mice; P10, 3 mice). Top 11 genes out of 591 genes (Fig. 5c center and Fig. 5f) are represented, FPKM: GFPLow/ GFPNeg>2, GFPLow-GFPNeg>50. n=9, mean ± SEM, p values were analyzed by one-way ANOVA Tukey's multiple comparisons test. (l)-(v) The graph shows the expression of each gene obtained from bulk-RNAseq analysis in GFPHigh MN+ microglia. (ASC; astrocyte, NEUR; neuron (red), OPC; oligodendrocyte precursor cell, iOLG; immature oligodendrocyte, mOLG; myelinating oligodendrocyte, MG/MAC; microglia/macrophage (blue), EC; Endothelial cell, All others except NEUR and MG/MAC are shown in the black bar. mean, n=2). This data was obtained from the Brain RNA-Seq database (https://www.brainrnaseq.org/).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3.
Supplementary Table 1 (download XLSX )
Identification of components of the extracellular micronucleus using LC/MS.
Supplementary Table 2 (download XLSX )
Genes with expression variations in microglia induced by neuronal MN.
Supplementary Table 3 (download XLSX )
Primer sequence for quantitative RT–PCR.
Supplementary Video 1 (download AVI )
Cortical slice culture imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 2 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 3 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 4 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 5 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 6 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 7 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 8 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 9 (download AVI )
in vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Video 10 (download AVI )
In vivo two-photon imaging using Cx3cr1-GFP/H2B-mCherry mice brain.
Supplementary Software (download ZIP )
The package containing the code of CAMDi software.
Source data
Source Data Figs. 1–5 (download XLSX )
Statistical Source Data for main Figs. 1–5.
Source Data Extended Data Figs. 1–10 (download XLSX )
Statistical Source Data for Extended Data Figs. 1–10.
Source Data Fig.2 and Extended Data Fig.2 (download PDF )
Uncropped blot data for Fig. 2 and Extended Data Fig. 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yano, S., Asami, N., Kishi, Y. et al. Propagation of neuronal micronuclei regulates microglial characteristics. Nat Neurosci 28, 487–498 (2025). https://doi.org/10.1038/s41593-024-01863-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-024-01863-5