Abstract
Utilizing medicinal plants and other natural resources to prevent different types of human cancers is the prime focus of attention. Cervical cancer in women ranks as the fourth most common type of malignancy. The current study used gas chromatography-mass spectrometry (GC–MS) to identify the active phytochemical constituents from Caladium lindenii leaf extracts using ethanol (ECL) and n-hexane (HCL) solvents. Plant extracts were tested for potential cytotoxic effects on HeLa and HEK-293 T cells using the MTT (3-(4,5-Dimethylthiazol-2-yl)-2,5-Diphenyltetrazolium Bromide) and the crystal violet assays. SYBR Green-based real-time PCR was performed to assess the mRNA expression profile of the apoptosis biomarkers (BCL-2 and TP53). The molecular interaction of the compounds with the targeted proteins (TP53, BCL2, EGFR, and HER2) was determined using molecular docking. GC–MS analysis revealed a total of 93 compounds in both extracts. The ECL extract significantly reduced the proliferation of HeLa cervical cancer cells, with an IC50 value of 40 µg/mL, while HEK-293 T cells showed less effect (IC50 = 226 µg/mL). The quantitative RT-PCR gene expression analysis demonstrated the ethanol extract regulated TP53 and BCL2 mRNA expressions in treated cancer cell samples. Heptanediamide, N,Nʹ-di-benzoyloxy-(− 10.1) is the best-docked ligand with a TP53 target found in the molecular docking study, whereas EGFR/Clionasterol had the second highest binding affinity (− 9.7), followed by EGFR/Cycloeucalenol (− 9.6). It is concluded that ECL extract has promising anti-cervical cancer potential and might be valued for developing new plant-derived anticancer agents after further investigations.
Similar content being viewed by others
Introduction
Cancer is one of the most devastating Non-Communicable Diseases (NCD), and its global burden has risen noticeably over the years based on incidence and prevalence data1. Cervical cancer ranks fourth among female cancers globally, with an estimated 341,831 deaths and 604,127 new cases in 2020. It is devastating that 570,000 women have been diagnosed with cervical cancer, of which 311,000 died from the disease in 2018, highlighting the importance of regular screenings and early detection2. It’s important to be aware of the global statistics surrounding cervical cancer that more than 58% of all cases were assessed in Asia, and 20% in Africa, 10% in Europe, and Latin America.
Additionally, more than half of deaths related to cervical cancer were estimated in Asia at 58%, followed by Africa at 22%, and Latin America at 9%. Specifically, 18% of cases and 17% of deaths occur in China, and 21% of cases and 23% of deaths occur in India has been reported. Women should prioritize their health and stay educated about procedures that reduce their risk of developing cervical cancer3. As reported by several studies, many environmental and genetic factors play a role in the development and prognosis of cervical cancer. Nonetheless, HPV (human papillomavirus) is regarded as the prime factor and causative agent. Out of 200 identified papillomaviruses, from which 12 have been identified as carcinogenic, of which hrHPV-16 and hrHPV-18 account for 50% and 10% of the instances, respectively4. Numerous studies across different ethnic groups have demonstrated a correlation between genetic polymorphism and predisposition to cervical cancer. TP53 is an essential tumor suppressor gene that plays a role in angiogenesis suppression, apoptosis, mechanisms involved in DNA repair, as well as the regulation of the cell cycle. Although the results have been inconsistent, studies have shown that SNPs (single nucleotide polymorphisms) in the TP53 gene, particularly in the codons 47 (rs1800371) and 72 (rs1042522), have been strongly associated with a variety of cancers, including cervical cancer5. The BCL-2 protein family, which controls the intrinsic apoptotic pathway, has been a desirable target for anticancer therapy. Interest in the BCL-2 protein family has recently increased owing to the emergence of small chemical inhibitors that specifically and potently block certain family members, causing apoptosis in cancer cells4,6,7. Targeting members of the BCL-2 family and TP53 in cervical cancers has yet to be investigated thoroughly and this could be a novel treatment option.
However, with the advent of novel compounds and metallodrugs, including aryl-substituted derivatives of acetamide, amine derivatives, oxadiazole substituted 2-(4-N, N,-Dimethyl) pyridine, and Schiff base derivatives of 5-(2-phenoxypyridin-3-yl)-1,3,4-thiadiazol-2-amine have been certain advancements in the treatment of cancer. These compounds have demonstrated moderate to potent antitumor activities against several cancer cell lines, such as Caco-2, HeLa, and MCF-78. Due to their complex structures, it is quite challenging to use these chemicals in novel drug development. Furthermore, the molecular mechanisms underlying metallodrug activity are yet to be fully elucidated, and metallodrugs are reported to induce genotoxicity. Moreover, the effectiveness of these drugs is highly dependent upon the cell cycle stages, which may restrict their clinical administration9. It is important to consider more potent natural chemotherapeutics that come from plant sources and have minimal toxicity to normal cells10. Considering herbal medicine as a therapeutic innovation could potentially reduce the cost of cancer therapy. Many people believe that herbal medicines are more natural and may not have as many side effects. It’s important to continue exploring different approaches to cancer treatment to ensure that patients have access to options that work best for them11.
Caladium lindenii (C. lindenii), also known as fancy-leafed or elephant’s ear, is a beautiful flowering plant that is commonly used as an ornamental part. It is topographically found in many parts of the southern USA and is a member of the Araceae family. The authors described some species’ phytochemical contents, characterization, antioxidant, and biological activities12,13,14,15,16,17. From 1998 to 2002 in the Philippines, when the prevalence of lung, breast, and prostate cancer became high, there remained a need to develop a non-toxic and reasonably priced anticancer drug. At that prior time, Caladium bicolor was a particular source of antiangiogenic, antioxidant, and cytotoxic activities. Numerous studies have reported the phytochemical constituents (flavonoids, polyphenols, alkaloids, limonoids, saponins, lactones, terpenes, cyanide, Xanthos, and oxalates) of various species of Caladium18,19,20. Still, no research has been done on the bioactive complexes found in the leaves of C. lindenii.
The current study identified natural compounds from leaf extracts of C. lindenii through GC–MS analysis. It evaluated the impact of these extracts on human-derived 2D cervical cancer and non-cancer (HEK-293 T) cell models. Additionally, the study observed the activity of the leaf extracts on tumor suppressor and apoptosis regulatory genes (TP53 and BCL2). By analyzing the structures and formulae of the bioactive chemicals found in the plant through an in silico molecular docking process, we may be able to discover new anticancer drug formulations. This study might be a crucial step in predicting the potential of these chemicals for cervical cancer treatment and management.
Materials and methods
Plant collection and preservation
Fresh leaf samples of Caladium lindenii were collected from Islamabad city, using the services provided by the local management and botanist. The plant was identified at Government College University, Lahore, Pakistan, and voucher specimens were deposited for future reference (GC. Herb. Bot. 3854). Collected plants were washed with distilled water, dried at 37 °C, stored at 4 °C, and utilized for maceration within 7 days.
Crude extract preparation
For extraction, dried plant leaves were ground into a powder and allowed to air dry at room temperature. After maceration in 1.0 L of ethanol (Sigma-Aldrich, 90%) and n-hexane (Sigma-Aldrich, 95%), the soaked powder (300 g) was left at room temperature for 5 days. Whatman filter paper (Grade 1) was used to filter the mixture and concentrated using a rotary evaporator (Heidolph Hei-Vap, Germany) to get a semi-solid extract. Lyophilization (Freeze drying) yielded dried extracts and kept them in aliquots at 4 °C for additional experiments21. Crude extracts were then subjected to GC–MS study. The equation calculated the percentage yield of extracts22.
Preliminary phytochemical screening
The lyophilized ethanol and n-hexane extracts of plant leaves were sent to the International Center of Chemical and Biological Sciences (ICCBS), HEJ, University of Karachi, Karachi, Pakistan, for gas chromatography–mass spectrometric (GC–MS) analysis for identifying bioactive phytocompounds. The Agilent Technologies (7890A) GC–MS triple quad system with EI and CI ion source was used to analyze the plant extracts. The instrument contains DB 35 MS capillary standard and non-polar column having a 30 mm × 250 µm dimension with ID × 0.25 µm film. Helium gas was used as transporter gas during the 36-min GC–MS experiment and was adjusted to a column velocity flow of 1.0 mL/min. The injector was run at 250 °C, while the oven operated at a holding time of 60 °C, followed by 10 °C per minute to 310 °C per 4 min. The compound identification was based on the HEJ GC–MS library along with their retention indices comparison. The National Institute of Standards and Technology (NIST) database, which contains over 62,000 patterns, was used to examine the mass spectrum. Understanding the name, chemical formula, chemical structure, mass spectrum, and molecular weight of various compounds was made easier by comparing the spectra of known compounds with the NIST library. =
In vitro anticancer evaluation
Cell lines and cell culture
Cervical cancer and non-cancer (HEK-293 T) cell lines were provided by the University of Lahore’s cell BioBank (IMBB/CRiMM). Dulbecco’s Modified Eagle’s Medium (DMEM) (Caisson Lot#02160032), 10% heat-inactivated fetal bovine serum (FBS) (Sigma-Aldrich Lot#BCBS3184V), and 100 U/mL of Penicillin–Streptomycin (Caisson Lot#10201011) were used to culture HeLa cells. RPMI-1640 medium containing 15% inactivated FBS was used to culture non-cancer (HEK-293 T) cells. For the incubation of the cell lines, a temperature of 37 °C was set in a humid environment with 5% CO2, and the cells were sub-cultured two times a week.
Cell viability assay
For the evaluation of cell viability, a crystal violet (0.1%, WV) assay was used; a 96-well microtiter plate was used to seed and obtain 1 × 104 HEK-293 T and HeLa cells per well in 200 µL of complete media. At a temperature of 37 °C, 5% CO2, and humidity, the cells were left for incubation for 72 h. Following the process of fixation using 70% ethanol for 10 min, it is necessary to analyze the results of the assay and determine cell viability at each step. For 30 min, 100 µL of CV solution was added to dye the cells. After washing with PBS, 200 µL of 10% triton X-100 solution (Sigma-Aldrich CAS#9036-195) was used to remove the tint from the cells. After incubating the samples at room temperature for 30 min, with the help of a spectrophotometer, the absorbance of the cells was measured at 570 nm23. Using the following equation, the percentage viability of the cells was calculated24.
MTT cytotoxicity analysis
The MTT assay was employed to assess the activity of the mitochondrial enzyme, succinate dehydrogenase, and quantitatively analyze cell propagation25. In 96-well cell culture plates, cells were seeded in triplicate per biological replicate at a density of 1 × 104 cells/well for anticancer assays, and the plates were incubated for 24 h. The ethanol and n-hexane stock solutions were prepared in DMSO (Invitrogen Inc., USA) at 160 mg/mL concentration. Various concentrations of plant extracts (10, 50, 100, 200, and 400 µg/mL) were prepared and applied to cancer and non-cancer cells and incubated for 72 h. Untreated cells (control) were cultured in plan DMEM media containing 2% FBS and 0.1% DMSO as vehicle control. Cisplatin (10 µg/mL) (INMOL Receipt No, 651278) was used as a positive control. After 72 h of treatment, the media was removed, and the cells were cleaned using a PBS solution. Furthermore, 20 µL of the MTT stock solution (5 mg MTT + 1 mL PBS) (Invitrogen Inc., USA) was added to each well, including the untreated control. In the following step, the 96-well plates were incubated for 2 h at 37 °C and 5% CO2. After the media was discarded from the wells, 150 µL of DMSO was added to all wells, mixed, and placed in an incubator for 10 min at 37 °C25. At 570 nm, the optical density (OD) of cells was calculated using the microplate reader (BIO-RAD). The linear regression method was used to determine IC50, the half-maximal inhibitory concentration.
Morphological assessment
Plant extracts of different concentrations (10, 20, 50, 100, and 400 µg/mL) were added to 96-well microtiter plates containing 1 × 104 cells/mL of cervical cancer and non-cancer (HEK-293 T) and incubated for 72 h. A Floid imaging station was used to observe the morphological variant, and the results were compared with the UT control.
RNA extraction and cDNA synthesis
To measure the expression levels of BCL2 and TP53, appropriate primers were used in Real-time PCR (Table 1). A 6-well culture plate was used to seed cervical cancer cells (3 × 105) for 36 h, and the temperature was maintained at 37 °C to ensure complete adhesion for RNA isolation. The cells were cultivated for another 72 h after exposure to IC50 concentrations of plant ethanol extract26. Following incubation, cells were washed with PBS and removed from the culture plates by adding 1 mL of TRIzole reagent (Sigma-Aldrich). To isolate all three levels: the protein red organic phase, DNA interphase, and the RNA having colorless aqueous phase, 200 μL of chloroform was added, the mixture was vigorously vortexed for 15 s and subjected to centrifugation at 4 °C, and 12,000 rpm for 20 min. The upper aqueous phase was poured into isopropanol-filled equal-volume Eppendorf tubes, and the RNA was pelleted by further centrifuging the samples at 4 °C, and 12,000 rpm for 15 min. After discarding the supernatant, the RNA pellet was cleaned with 1 mL of 75% ethanol, centrifuged at 12,000 rpm for 15 min at 4 °C, and re-pelleted. The RNA pellets were then allowed to dry for 5–10 min in a dry bath before being dissolved in DEPC-treated water. At 260/280 nm, the OD of the RNA (1µL) was measured by using Nanodrop (Thermo scientific ND 2000/2000c Spectrophotometer) and stored at − 80 °C27. cDNA synthesis was performed using 1 µg of extracted RNA following the manufacturer protocol (ABScript II cDNA First-Strand Synthesis Kit, Lot: 962100J09W09).
Real-time PCR analysis
The region to amplify TP53, BCL2, and Hprt1 (reference) genes encompasses a partial 5ʹ upstream region. Primer sequences were retrieved from NCBI and designed through the online software Primer3 (https://primer3.org/) (Table 1)28. SYBR Green master mix (Thermo Scientific, USA) was used in Real-time-PCR (Qiagen Rotor-Gene Q 5plex HRM) to examine the relative expression of genes. A comparative threshold cycle method was used to quantitatively compare the expression of each gene (TP53, BCL2, and Hprt 1). Hprt1 served as a reference gene for each gene of interest, and every Ct value was stabilized to the Ct value of Hprt1 RNA. A single qPCR reaction contains reagents such as SYBR Green Master Mix (12.5 μL), endonuclease-free water (variable), primer forward + reverse, 2 μL), and cDNA (2 μL or 200 ng). The qPCR cycling conditions for TP53 and BCL2 are mentioned in Table 2. The ∆Ct values were calculated by subtracting the endogenous Hprt1 gene’s Ct value from the mean Ct values of the target genes TP53 and BCL2. The ∆∆Ct value was calculated by: ∆Ct of sample-∆Ct of calibrator, and the fold change was measured using the formula 2−∆∆Ct. Real-time PCR was used to evaluate TP53 and BCL2 gene expression. Each reaction was carried out in triplicate28.
Molecular docking
Protein target selection and preparation
The Research Collaboratory for Structural Bioinformatics (RCSB) protein (URL: https://www.rcsb.org) data library was used to retrieve the 3-dimensional (3D) X-ray crystallographic structures of P53, Bcl2, EGFR, and HER2 proteins/receptors with PDB IDs: 2OCJ, 2W3L, 3poz, and 1n8z, respectively. The proteins for molecular docking were prepared by removing water ions, heteroatoms, and inhibitors and adding hydrogen, and the PDBQT format was used to save the proteins29. The docked complexes and potential active site residues were identified using PyMOL (Molecular Graphics, version 2.5.4, 2010, Shrodinger L.L.C.)30. The grid box was placed on the co-crystallized ligand for docking, and the Autodock 4.2.6 program was used to save the data in a config.txt file29. The analysis of various interacting residues and their bond lengths involved in creating stable complexes was conducted using the automated Protein–ligand interaction profiler (PLIP) website.
Ligand selection and preparation
GC–MS identified phytoconstituents structures were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov) as SDF files, and then these ligands were saved into PDB format using the software open babel (http://openbabel.org/wiki/Main_Page)31.
Screening of phytocompounds for drug-likeness
Evaluating a chemical’s drug-likeness, a crucial stage in the drug development process that determines if a particular drug-like compound is either orally active or not, is done using Lipinski’s RO5. Using the SwissADME predictor (http://www.swissadme.ch/index.php), physiochemical parameters were assessed in the ligands; the main goal was to identify the information that would be helpful for the drug development and research process. The structures of the selected phytocompounds were uploaded to the admetSAR (http://lmmd.ecust.edu.cn/admetsar2) website in canonical SMILES format to examine the pharmacokinetics including toxicity, mutagenicity, carcinogenicity, level of solubility, BBB diffusion, cytochrome P450 enzyme inhibition, and human intestinal absorption of compounds29.
Molecular docking analysis using Autodock Vina
AutoDock 4.2.6 was downloaded from ‘The Scripps Research Institute’s official website (http://autodock.scripps.edu/), an automated docking tool. The scientific community regards it as one of the most efficient molecular docking programs, and it is available for free32. The desired folder was set as the starting directory before docking. The protein molecule that had been processed was fetched into the Auto Dock 4.2.6 workspace. After adding the polar hydrogen atoms, the protein’s Kollman and Gasteiger charges were calculated. The protein was subsequently utilized as the target after being saved in PDBQT format33. Each protein’s grid center was selected to match the active sites of the protein (Table 3). The AutoDock 4.2.6 program was utilized to determine the optimal protein–ligand conformations, taking into account their highest binding affinities. Discovery Studio client 2021 was used to see the structures demonstrating how ligands and proteins interact34. The binding mechanisms of complexes were studied using the three-dimensional protein–ligand postures. 2D illustrations of complexes were used to investigate a range of interactions, such as hydrogen bonding and hydrophobic interactions35.
Statistical analysis
Every test was conducted three times, and the mean and standard deviation were used to express the findings. To ascertain how the three variables interacted, a one-way analysis of variance (ANOVA) and Tukey’s multiple comparison test using Graph Pad Prism 8.0 was done36. The R-squared statistics represent the proportion of variance in the results obtained from analyses based upon the general linear model (examples: ANOVA, regression)37. The IC50 values were calculated using a linear regression method to analyze data; a confidence interval of 95% was used, while a p-value of 0.05 or lower was used to determine the statistical significance.
Ethics approval and consent to participate
It has been confirmed that the experimental samples of plants, including the collection of plant material, complied with relevant institutional, national, and international guidelines and legislation with appropriate permissions from Authorities of the Institute of Molecular Biology and Biotechnology, The University of Lahore, 54000, Lahore, Pakistan.
Results
Total yield of crude extracts
It was found that the n-hexane extract of C. lindenii (HCL) produced a percentage yield of 15.2%, while the ethanolic extract of C. lindenii (ECL) produced 16.13%.
Identification of phytoconstituents from C. lindenii extracts by GC–MS
The spectra that were obtained from GC–MS analyses were compared to those present in the reference library at NIST. The phytocompounds were identified based on their retention time (RT), peak value, molecular formula (MF), and their chemical names. A direct correlation has been observed between peak area and the number of components, which can be determined by comparing the standard peak area with the total peak area; compounds are separated based on their chemical natures at different retention times. The obtained mass spectra are patterns of compounds that can be compared to reference spectra present in the data library. The GC–MS spectrum analysis of the ethanol and n-hexane extracts showed the existence of phytocompounds that strongly contribute to the medicinal properties of C. lindenii.
Using GC–MS analysis, 93 chemicals were identified in both ECL and HCL extracts (Table 4), of which four chromatogram peaks from the HCL extract were found to be unknown (Figs. 1 and 2). Our extracts contain natural chemicals, including phenolic compounds, such as 2,4-di-tert-butylphenol. Terpenoids (2,6,10-trimethyltetradecane), terpenes (Dihydroactinidiolide), fatty acid esters (palmitic acid), alkenes (Linolenic acid), and phytol were among the terpenoid derivatives that we obtained in this investigation. Volatile terpene with a potent cytotoxic impact is dihydroactinidiolide. Plant sterols, known as phytosterols, are thought to have several health advantages, such as lowering cholesterol and having antioxidant properties. Phytosterols such as stigmasterol, campesterol, clionasterol, and stigmast-4-en-3-one are present in both extracts. Meanwhile, stigmasterol and Clionasterol are thought to be anti-carcinogenic. This investigation also identified fatty acid methyl esters (Methyl linolenate) and phthalate esters (Dioctyl phthalate). Table 4 lists the discovered phytocompounds of C. lindenii, their retention time (RT), molecular formula, molecular weight, and percentage area; however, Figs. 1 and 2 illustrate the peak spectrum.
GC–MS chromatogram of C. lindenii ethanol (ECL) extract.
GC–MS chromatogram of C. lindenii n-hexane (HCL) extract.
Cytotoxic effect of C. lindenii extracts on HeLa and HEK-293 T cell lines
The compounds were examined to estimate their cytotoxicity on the proliferation of cancer and non-cancer cells using cell-based assays. This technique relies on the enzyme mitochondrial dehydrogenase activity, which catalyzes the conversion of tetrazolium salt to formazan crystals in living cells.
Various ECL and HCL extract concentrations were applied to cancer and non-cancer (HEK-293 T) cell lines to evaluate the antiproliferative effects, as elaborated in Fig. 3. HeLa and HEK-293 T cells were treated with 10, 50, 100, 200, and 400 µg/mL concentrations of ECL and HCL extract for 72 h. ECL concentrations favorably reported the effect on HeLa cells with IC50 = 40 µg/mL. IC50 values of ECL extract showed less antiproliferative activity in HeLa cells compared to healthy cells (IC50 = 226 µg/mL), revealing the existence of organic compounds. However, the effect of HCL extract concentrations on HeLa cells revealed less potential with IC50 = 70 µg/mL and HEK-293 T cells (IC50 = 470 µg/mL). The ECL concentrations (400, 200, 100, and 50 µg/mL) showed more cytotoxicity toward HeLa cells, but the effect of cisplatin (10 µg/mL) was aggressive toward both cell lines. The outcome of all concentrations was compared with the untreated group (UT).
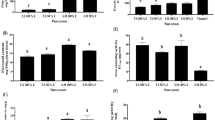
Graphical representation of antiproliferative effects of C. lindenii extracts on HeLa and HEK-293 T cell lines, (A) Analysis of ECL extract on the HeLa (cancer) cell line showed a remarkable decrease in a dose-dependent manner, (B) HCL concentration on HeLa cells also has a good effect but not more than ethanol, (C) HEK-293 T (non-cancer) cells revealed negligible toxicity of ECL extract at all concentrations, (D) HEK-293 T cells exhibited minimal effects when exposed to HCL extract. Cisplatin affects the most on both cell lines. *p ˂ 0.05, **p ˂ 0.001, ***p ˂ 0.0001 denoted as statistically significant and compared with UT control. The data is represented as absorbance mean ± SD of each group. UT untreated; DMSO dimethyl sulfoxide; ECL ethanol extract of C. lindenii; HCL n-hexane extract of C. lindenii.
Cell viability analysis of C. lindenii extracts on HeLa and HEK-293 T cell lines
Crystal violet assay technique was used to quantify the remaining alive cells in treated culture media by calculating the dye absorbance at 570 nm of each well. It enters the cell cytoplasm and nucleus and is bound to only the proteins, RNA, and DNA molecules of alive cells. Cells that lose their permeability and undergo death exhibited less staining dye in the culture medium. For that purpose, the calculated IC50 values from the MTT assay were further evaluated through crystal violet assay to approve the antiproliferative activities of the extracts. Half maximal inhibitory concentration (IC50) is the 50% effect of a drug, toxicant, or antibody that brings a dose–response after a definite exposure of time. The cell counts adherent to the plate surface were calculated through the staining (CV) of cancer and non-cancer (HEK-293 T) cells represented in Fig. 4.
Cell viability analysis of C. lindenii extracts on HeLa and HEK-293 T cells, (A) The percentage assay showed the decreased viability of cells measured at 570 nm. ECL concentrations showed a strong significant effect on HeLa (cancer) cells, (B) The viability percentage showed a strong dose-dependent decrease in HeLa cells with R2 values on treatment with HCL concentrations, (C) HEK-293 T (non-cancer) treated cells with ECL showed less viability at 200, and 400 µg/mL, (D) HCL treatment didn’t show strong activity at any concentration in HEK-293 T. Results are presented as percentage values. *p ˂ 0.05, **p ˂ 0.001, ***p ˂ 0.0001 vs control UT. UT untreated; DMSO dimethyl sulfoxide; ECL ethanol extract of C. lindenii; HCL n-hexane extract of C. lindenii.
The HeLa and HEK-293 T cells were treated with 10, 50, 100, 200, and 400 µg/mL concentrations of ECL and HCL. At 200 and 400 µg/mL, the ECL exhibited low absorbance values as compared to hexane, which showed higher growth inhibition of the HeLa cells recorded as 94%, 72%, 58%, 45%, and 23% in comparison with UT control (100%). HCL showed less percentage viability with 95%, 76%, 65%, 50%, 32%, and 100% of UT. HEK-293 T cells were measured to 95%, 88%, 76%, 65%, and 56% of ECL and 100% of UT. The cell viability of HEK-293 T was 96%, 88%, 78%, 66%, and 56%, and UT was 100% on treatment with HCL extract. However, cisplatin (10 µg/mL) exhibited 42% and 52% viability in HeLa and HEK-293 T cells. The difference in inhibition was not more noticeable at all concentrations of both extracts. R2 represents the coefficient of determination in a regression model and verifies differences among experimental data and standards. R square was calculated as (a) = 0.9931, (b) = 0.9660, (c) = 0.9765, and (d) = 0.9510, and p-value = < 0.05 for both extracts (Fig. 4).
Morphological changes in HeLa and HEK-293 T cells on exposure to C. lindenii ethanol extract
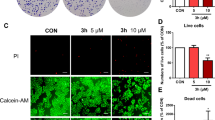
Morphological changes and a reduced number of HeLa cells were observed, starting from lower (10 µg/mL) to higher (400 µg/mL) concentrations and compared with UT and positive (cisplatin 10 µg/mL) controls. Cell detachment, shrinkage, and cytoplasmic condensation were observed with increasing concentrations, as shown in Fig. 5. HeLa cells showed vesicle formation started from 10 µg/mL and worsened with an accelerated dose. However, maximum cell damage and detachment were observed at 200 and 400 µg/mL. HEK-293 T cells remained the same without showing the harmful effect of ECL concentrations. Cisplatin (10 µg/mL) concentration exhibited cytoplasmic condensation and vesicle formation in both cell lines.
Morphological representation of HeLa and HEK-293 T cells exposed to various concentrations of C. lindenii ethanol extract, (A) The effect of various concentrations of ECL extract on HeLa (cancer) cells showed a dose-dependent decrease observed by the Floid cell imaging station, (B) HEK-293 T (non-cancer) cells show minimum degradation at 200, 400 µg/mL. Cisplatin changed the cell morphology of cancer and healthy cell lines. All the outcomes were compared with UT groups. UT untreated; DMSO dimethyl sulfoxide; ECL ethanol extract of C. lindenii.
Morphological changes in HeLa and HEK-293 T cells on exposure to C. lindenii n-hexane extract
Morphological assessment of HCL concentrations of HeLa cells exhibited cytoplasmic condensation starting from 100 µg/mL (Fig. 6). The cells treated with 100, 200, and 400 µg/mL concentrations encouraged nuclear degradation fragmentation, cell rounding, and detachment. HEK-293 T cells showed no possible signs of toxicity. A slightly decreased number of cells was observed in lower (10 and 50 µg/mL) doses in cancer cells. The untreated group showed 100% viability in a cell population. Cisplatin (10 µg/mL) as a chemotherapeutic drug exhibited morphological changes in cancerous and healthy cells.
Morphological representation of HeLa and HEK-293 T cells exposed to various concentrations of C. lindenii n-hexane extract, (A) Morphology of HeLa cells showed the effect of HCL concentrations applied for a period of 72 h, (B) HEK-293 T cells treated with the same concentrations and time duration. The effect of the anticancer drug (Cisplatin) response revealed transformed cellular morphology. All morphological outcomes were compared with the UT group. UT untreated; DMSO dimethyl sulfoxide; HCL n-hexane extract of C. lindenii.
Effect of plant extracts on the mRNA expression levels of cancer-related genes
This study was considered to analyze the differential expression of oncogene (BCL2) and tumor-suppressor (TP53) genes in the cervical cell line. Differences in fold change of the cervical cancer cell expression were observed after being treated with the IC50 concentration of ethanol extract. These values were compared with the untreated (UT) sample and endogenous (Hprt1) control levels, denoted as a housekeeping gene, used to normalize the mRNA levels among samples.
Effect of C. lindenii ethanol extract on the expression level of TP53 in HeLa cell lines
The IC50 concentration of ECL extract has demonstrated TP53 mRNA expression throughout treatment. An upregulation of the TP53 level at 72 h with a fold change of 0.6 was observed. mRNA expression of TP53 decreased with a mean Ct value of 20.67 in control and increased to 16.44 in treated samples in cervical cancer cells. When focusing on mRNA expression of the housekeeping gene (Hprt1), it expresses a possible peak with its Ct value range of 25 cycles in both samples. The Ct values of TP53 and Hprt1 genes have been mentioned in Table 5. The bar graph represents the expression levels of TP53 analyzed by real-time PCR presented in Fig. 7.
Graphical representation of mRNA expression analysis of TP53, BCL2, and HPRT1 genes in the HeLa cell line. The graph shows the regulation of TP53 and BCL2 in HeLa-cells after ECL treatment compared to the control (UT). Results show the statistically significant correlation among groups with p = ˂ 0.0001; TP53 tumor suppressor gene; BCL2 B-cell lymphoma 2 protein; HeL HeLa cell line.
Effect of C. lindenii ethanol extract on the expression levels of BCL2 in HeLa cell line
Figure 7 shows the upregulation of the BCL2 gene in cervical cancer cells in control samples. The IC50 concentration of ECL extract causes apoptosis induction by regulating BCL2 mRNA expression. The expression was observed with the fold change in HeLa cells at 9.40, while the mRNA expression of BCL2 suggested a decreased mean Ct value (19.70) in the treated samples, and the control (UT) sample showed higher expression levels (16.46) in cancer cells. We observe the mRNA expression level of the housekeeping gene (Hprt1) in treated and control samples with a Ct value range of 25 cycles in both samples. The Ct values of BCL2 and Hprt1 genes have been mentioned in Table 5. The bar graph represents the expression levels of BCL2 analyzed by real-time PCR presented in Fig. 7.
Molecular docking
Profiling of compounds for drug-likeness
Drug-likeness features of the ligands are the pivotal characteristics for their practice as remedies. At contemporary, Lipinski’s rule is commonly employed to predict the drug-likeness of small compounds by SwissADME, as presented in Table 6. Six compounds fulfilled Lipinski’s rule, which should be < 500 g/mol, stating that an orally active drug has up to one violation of the standards. Hence, we found that all these potential compounds qualified as oral drugs, approving Lipinski’s rules. The compounds were docked with the targeted apoptotic proteins, following the prediction analyses using molecular docking.
Molecular interaction analysis
The low binding energies specified the pairing’s high binding affinities and strong interactions. We utilized AutoDock 4.2.6 for interaction with targeted proteins (P53, BCL2, EGFR, and HER2) for molecular docking analysis. In this study, we have reported 2 best hit compounds for each target protein. The results demonstrated that Heptanediamide, N,Nʹ-di-benzoyloxy- had a strong binding affinity for P53 (− 10.1 kcal/mol) and Clionasterol with EGFR (− 9.7 kcal/mol). Following others are EGFR/Cycloeucalenol showed − 9.6 kcal/mol, P53/2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde had binding score of − 9.5 kcal/mol, HER2/Heptanediamide, N,Nʹ-di-benzoyloxy (− 9.0 kcal/mol), HER2/Cycloartenol (− 8.5 kcal/mol), BCL2/Obtusifoliol (− 7.6 kcal/mol), and BCL2/Cycloartenol (− 7.7 kcal/mol) showed noteworthy scores. The binding affinities of the docked complexes are presented in Table 6. The RMSD values calculated for all docked complexes were 0.000 Å, and its value ≤ 2 Å is fairly good. The binding affinity of these molecules and their binding energy in kcal/mol are demonstrated in Table 7. The 2D structure of compounds displayed in Fig. 8 and 3D protein–ligand models have been represented in Fig. 9.
2-dimensional configuration of screened compounds. (A) Heptanediamide, N,Nʹ-di-benzoyloxy-; (B) Cycloartenol; (C) Clionasterol; (D) Obtusifoliol; (E) 2-[4-methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde; (F) Cycloeucalenol.
3-dimensional configurations and compounds interaction with cervical cancer-CC target proteins. (A) TP53 with Heptanediamide,N,N’-di-benzoyloxy-; PDB ID: 2OCJ (B) TP53 with 2-[4-methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde; PDBID:2OCJ (C) BCL2 with Cycloartenol; PDB ID: 2W3L (D) BCL2 with Obtusifoliol; PDB ID: 2W3L (E) EGFR with Clionasterol; PDB ID:3poz (F) EGFR with Cycloeucalenol; PDB ID:3poz (G) HER2 with Heptanediamide, N,Nʹ-di-benzoyloxy; PDB ID:1n8z (H) HER2 with Cycloartenol; PDB ID:1n8z.
Amino acid residue interaction analysis
Interactions between amino acid residues and ligands are crucial. BIOVIA Discovery Studio client 2021 was used to analyze and visualize the interaction patterns of the targeted anticancer proteins with ligands (Table 8 and Figs. 10, 11). Drug-protein interaction and the structural integrity of a broad range of biological processes depend on hydrogen bonds. The strength of the H-bond is mostly determined by the donor and acceptor atoms as well as the angle of engagement. The most drug-like compounds for interacting with the targeted tumor suppressor protein P53 were found to be 2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde, and heptanediamide,N,Nʹ-di-benzoyloxy-. Table 8 shows molecular interactions between Heptanediamide,N,Nʹ-di-benzoyloxy- and P53, there are four hydrogen bonds with amino acid residues of Thr68, Pro124, Asn191, and Tyr196, and three hydrophobic connections with Thr68, Ala94, Thr98, Pro123, Asn191, Leu225, Trp262, and Phe293 were formed. Additionally, it exhibited a significant affinity for the HER2 protein, establishing hydrophobic contacts with eight HER2 amino acid residues (Thr5, Leu27, Leu291, Tyr387, Ile413, Leu414, Leu443, and His468) as well as five hydrogen bonds with residues of Thr5, Tyr281, Leu414, Gly417, and Ser441. Similar to how 2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde, and P53 interacted molecularly, one hydrogen (Thr98) and seven hydrophobic interactions (Leu63, Thr68, Thr98, Leu193, Ile195, Trp262, and Phe293) developed between residues. Two hydrogen bonds between Cycloartenol and Thr5 and Ser441 of HER2 were formed. Furthermore, Cycloeucalenol exhibited considerable binding affinity when docked with EGFR, establishing four hydrophobic interactions with Leu718, Val726, Thr790, and Phe997. Also, when docked with Clionasterol, it established nine hydrophobic bonds with residues: Leu718, Ala743, Lys745, Leu777, Leu788, Thr790, Leu844, Thr854, and Phe997. Cycloartenol had a strong binding affinity forming six hydrophobic connections with amino acid residues of Asp62, Phe63, Arg66, Arg105, Tyr161, and Pro163, and a hydrogen bond with Arg105 of BCL2 protein. Obtusifoliol interacted with BCL2 and produced three hydrophobic bonds with residues: Phe63, Tyr67, and Asp70.
2D and 3D-interactions of compounds with targeted proteins. (A) P53/Heptanediamide, N,Nʹ-di-benzoyloxy; (B) p53/2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde; (C) BCL2/Cycloartenol; (D) BCL2/Obtusifoliol.
2D and 3D-interactions of compounds and targeted proteins. (A) EGFR/Clionasterol; (B) EGFR/Cycloeucalenol; (C) HER2/Cycloartenol; (D) HER2/Heptanediamide, N,Nʹ-di-benzoyloxy.
Assessment of in silico ADMET prediction
We determined the compounds’ toxicity profiles using the Admetsar program. It was applied to certain substances to determine whether they presented low, moderate, or high toxicity hazards according to their effects on mutagenicity and irritation. Table 9 displays the comprehensive ADMET investigation of the compounds with the greatest values. Heptanediamide, N,Nʹ-di-benzoyloxy, was shown to be non-mutagenic, non-AMES poisonous, positive BBB permeability, allow HIA and Caco-2 absorbency, and to have non-hERG inhibitor, according to the admetSAR analysis. Out of all the ligands with good BBB penetration, the one with the greatest HIA value was discovered. Two major cytochrome P450 enzymes that are important throughout the liver’s drug metabolism are CYP2D6 and CYP3A4. The investigation results showed that the drug was a CYP2D6 and CYP3A4 non-inhibitor, suggesting that the liver might be involved in drug metabolism. It was also found to be a substrate for P-glycoprotein, which suggests easy transport within the body. The chemical was shown to have only mild hepatotoxicity. No alert was found for Lipinski’s rule, signifying the specificity of the compound. Other substances like 2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde, Obtusifolio, Cycloartenol, Clionasterol, Cycloeucalenol also showed good absorption in the human gut with positive HIA, non-malignant, non-AMES, permeable to BBB and Caco-2 along with non-inhibitor of hERG, while had good water solubility. For the P-gp substrate, 2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde), and Cycloartenol compounds tested negative. Good GI absorption was a feature common among all the chosen compounds and is crucial for medication absorption into the human body. It was discovered that none of the compounds inhibited CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4 except compound I found to be the inhibitor of the CYP2C19 enzyme. No substance was found to have hepatotoxic or Ames toxicity, except compound II, which showed minor hepatotoxicity. The phytochemicals under test had varying total clearance values. Compound III had the lowest value, and the others showed the highest value when removed from the plasma.
Discussion
Many medicinal plant species have been studied due to their bioactive metabolites to develop alternative sources for pharmaceutical sectors and promote health benefits through antioxidant-containing products. Even though most plants’ toxicity profiles have not been completely and thoroughly examined, it is often presumed that medicines derived from plant sources are far safer than their synthetically synthesized counterparts38. This study aimed to investigate the ethanol and n-hexane extracts from the leaves of Caladium lindenii for the first time to identify the therapeutic potential of phytochemicals.
GC–MS analysis of the ECL and HCL extracts identified the presence of 52 and 41 complexes with various chemical groups, including alcohols, aldehydes, esters, fatty acids, hydrocarbons, and ketones (Table 4). We will discuss the biological and pharmaceutical activities of those compounds here identified with high area percentages in extracts. From the ECL extract, the top seven bioactive phytochemicals, including phytol (24.35%), are associated with antibacterial, anticancer, antidiuretic, anti-inflammatory, antioxidant, and neuroprotective properties39. 2-(((2-Ethylhexyl)oxy)carbonyl)benzoic acid covers the total 10.82% area. A naturally occurring fatty acid ester known as ethyl palmitate (10.76%) has been observed to inhibit inflammatory cell growth40. In one study, EP reduces carrageenan-induced rat foot edema to lessen the prostaglandin E2 (PGE2) level in the inflammatory exudates, a fluid that is emitted from blood vessels41. The most prevalent saturated fatty acid in the human body is palmitic acid (8.62%), which is synthesized endogenously from amino acids, carbohydrates, and fatty acids. Additionally, palmitic acid acts as 5-an alpha-reductase inhibitor involved in developing prostate cancer (CaP) and also demonstrates antimicrobial, anti-fibrinolytic, anti-inflammatory, antioxidant, hemolytic, metabolic, and antitumor activities in several types of tumors42. Octacosane (5.07%) may be a potent anti-inflammatory and antioxidant agent43. One study elaborated on the antioxidant and wound-healing effects of octacosane isolated from M. pumilum plant extract using in vitro, in vivo, and in silico techniques44. Ethyl stearate (4.48%) has demonstrated antimicrobial activity45. Ethyl linolenate (3.55%) is an unsaturated fatty acid used in cosmetic formulations due to its clinical benefits, including anti-acne, antibacterial, anti-inflammatory, and anti-melanogenic properties46.
HCL extract identified major phytocompounds with high peak area and biological activities, including vitamin E (α-tocopherol) (16.5%), which is reported as an antioxidant, reduces cancer risks, and protects the skin from UV irradiations47. (E, E, E, E)-Squalene (10.47%) has demonstrated effective inhibitory activity in chemically induced colon, lung, and skin tumorigenesis. The anti-carcinogenic effect of squalene was seen when administered before or during carcinogen treatment48. The chemo-preventive activity of squalene may be attributed to one or many of the following mechanisms: antioxidative activities, modulation of carcinogen activation, and Ras farnesylation49. Methyl palmitate (10.18%) is documented to have properties as a 5-alpha-reductase inhibitor, antioxidative, hemolytic, hypocholesterolemic, and nematicidal agent50. Phytol (6.9%) is associated with antibacterial, anticancer, antidiuretic, anti-inflammatory, antioxidant, and neuroprotective properties. Methyl linolenate (5.88%), α-linolenic acid derived methyl ester, demonstrated antiproliferative activity and is likely to be a potent antifungal agent51. Hentriacontane (3.1%) shows anti-inflammatory and antitumor effects by inhibiting caspase-1 and NF-κB activation52.
Studies have indicated that the antioxidant and antibacterial effects of various Caladium species are found in literature, although its anticancer activity has not been studied earlier. Moreover, this study is the first to report the noteworthy anticancer activity of leaf extracts of C. lindenii. Thus, the existing study was directed to assess the antiproliferative activity of C. lindenii on HeLa and HEK-293 T cell lines using the MTT assay (Fig. 3). A previous study on the plant extracts of Paullinia cupana and Pyracantha angustifolia showed significant inhibition in the two cancer (MDA-MB231 and HepG2) cell line models, evaluated by cell viability assay (MTT). Furthermore, cell cycle arrest was observed in the cancer cell line after treatment with fruits and seed extracts of P. angustifolia and P. cupana. The calculated IC50 values of P. cupana (43.1 ± 10.69 µg/mL) and P. angustifola (21.91 ± 2.10 µg/mL) showed for MDA-MB-231. For HEPG2 cells, the IC50 values were as follows: P. cupana (32.57 ± 7.60 µg/mL), P. angustifolia (28.41 ± 4.53 µg/mL)53. In this study, ECL extract has demonstrated positive effects, such as the suppression of cell viability and the inhibition of the spread of cervical cancer cells, which had an IC50 = 40 µg/mL as opposed to HEK-293 T cell IC50 = 226 µg/mL. A maceration-produced HCL extract, with an estimated IC50 of 70 µg/mL for cervical epithelioid carcinoma and IC50 = 470 µg/mL for HEK-293 T cells, however, had reduced antiproliferative action. HeLa cells were more susceptible to the cytotoxicity of ECL doses (400, 200, 100, and 50 µg/mL), while the effect of cisplatin (10 µg/mL) was significantly more harmful to both cell types (Fig. 3). The assay confirmed that the extracts reduce cell viability dose-dependently and induce apoptotic morphological changes (Figs. 5 and 6).
Among the genetic alterations in various human cancers, one of the most frequent causes is the modulation of the oncogenes. The transcriptional activator TP53 has been shown to play a role in the transcription of many genes, including those linked to apoptosis. That is why it is considered that tumor behavior can be described depending on the alterations of the TP53 gene, and these are considered independent factors in disease evolution prognosis54. It is well-established that the BCL2 gene has a vital regulatory role in apoptosis. The gene encodes a cellular protein that causes inhibition of apoptosis in normal cells55. The current study is the first to reveal that C. lindenii induces apoptosis in cervical cancer cells by regulating TP53 and BCL2 signaling. This study observed the downregulation of BCL2 and the upregulation of TP53 in treated samples of cervical cancer cells (Fig. 7, and Table 5). However, it elucidated the decreased mRNA expression of TP53 and increased expression of the anti-apoptotic gene BCL2 in control samples.
In contrast, P53 is a proapoptotic protein that promotes apoptosis by interaction with BCL2 (antiapoptotic protein). Interaction of BCL2 and P53 causes suppression of the inhibitory effect on BAX, releases cytochrome C, and eventually causes apoptosis56. A mutation in TP53 is closely linked to p53 dysfunction in endometrial cancer. Between 17 and 45 percent of all histological types of endometrial carcinomas are p53-positive. According to Nakamura et al. (2019), there was a high p53 expression in 30–86% of nonestrogen type-II endometrial cancer (highly aggressive), while the p53-positive rate in type-I endometrial cancer was 10–44%57. The stable expression of Hprt1 with Ct value ranges of 25 cycles was observed in both samples (Fig. 7). Previously carried out stability analysis revealed that levels of a particular reference gene depend on the type of tissue. Moreover, when ranked using geNorm, NormFinder, and RefFinder algorithms, in cancer cell lines, including HepG2, Huh7, MCF-7, and MDA-MB-231, the ACTB, B2M, HPRT1, UBC, and YWHAZ were among the five stable genes6,58,59,60.
Roughly 80–90% of cervical carcinomas comprise DNA sequences of specific HPV subtypes, especially those of HPV 16 and 18. Viral infections intervene with the mechanism of cellular growth, immunologic, and DNA restoration, as suggested by the latest molecular data on the natural history of HPV infection and cervical cancer. The connection between p53 expression and cervical cancer progression is ambiguous because of contradictory reports61. It is still unknown how p53 takes care of the various signals to decide the correct cell fate in reaction to stress and how these reactions are linked to tumorigenesis and cancer progression. Depending on the cell context, they can also be linked with response to treatment. Multiple exons and polyadenylation sites are the two promoters of the human p53 gene; hence, it is transcribed into several variants. Examining the expression of p53 isoforms and p53 mutation status can help define various subtypes within a specific cancer type, which would have various reactions to treatment62. The p53 expression and function can be associated with the suppressive effect of many medicinal plants on cell growth and proliferation of cancer cells63.
Scientists may be capable of developing innovative therapies by better understanding the phytochemical’s mechanisms by interacting with targets to inhibit or stimulate the specific proteins and enzymatic pathways for treating a particular disease. Molecular docking aims to contribute to predicting the structures of the ligand-receptor complexes with the aid of computational approaches64. In the present study, the identified phytocompounds were screened and sorted depending on their drug-likeness, molecular properties, physicochemical properties, and Lipinski’s rule of five. These ligands/compounds were screened from C. lindenii and docked against each target protein (p53, BCl2, EGFR, and HER2). As per the selection criteria, only 6 hits of the total compounds fell in the category of a good-docking score. To our knowledge, the literature has not reported the interactions of these selected ligands with the four targets in cancer studies.
Molecular docking analyses were conducted among phytocompounds and proteins that have roles in DNA repair, regulation of transcription, and tumor suppression. In one study, it has been documented that doxorubicin, a commercial drug, does not follow the Lipinski Rule of Five, thereby it can be safely concluded that doxorubicin is a toxic commercial drug64,65,66,67. Two mechanisms have been proposed to understand the mechanism of doxorubicin in cancer cells: (i) modification in the chromatin structure by doxorubicin intercalation into DNA and topoisomerase II inhibition, (ii) the production of free radicals and oxidative damage to biomolecules68. In contrast, the phytochemicals in our study follow the basic criteria of drug-likeness (Lipinski Rule of Five); the phytocompounds discovered from this study are non-toxic and safe for human use (Tables 6 and 8). Accordingly, in the present study, we selected these anticancer targets (P53, BCL2, EGFR, and HER2) as these proteins substantially regulate apoptosis and initiate signaling events in carcinogenesis. Among all compounds, Heptanediamide, N,Nʹ-di-benzoyloxy, 2-[4-Methyl-6-(2,6,6-trimethyl-cyclohex-1-enyl)hexa-1,3,5-trienyl]cyclohex-1-en-1-carboxaldehyde), Clionasterol and Cycloeucalenol exhibited best results with more interactions and docking scores. At the same time, the other compounds also showed acceptable results (Table 7). A hydrogen bond specifies that the ligand had a great affinity for the protein, and a negative score suggests a good binding affinity with the target protein. As a result, the current study concludes that the phytochemicals identified using GC–MS examination of the plant extracts suggest treating cervical cancer69. These compounds have an outstanding pharmacokinetic profile, and the ADMET analysis confirms that they could be a good candidate for anticancer treatment.
Conclusion
Till today, there is no cure for cancer, and research is ongoing in developing lead molecules and precursors that could act as potential anticancer drugs against the disease. The present study aimed to discover natural compounds with potential anticancer effects. In vitro evaluation of ECL extract showed a strong potential to inhibit the proliferation of cervical cancer cells. Our study is a promising addition to the research on the p53 and Bcl2 mRNA expression in cervical cancer cell line proposed genetic modifications. To analyze the association between HPV infection and p53 and bcl2 in cancer samples, a bigger case series is necessary. To explain the variations in TP53 isoform expression in mRNA and protein levels, bigger sample sizes and a wider range of ethnicities are demanded in future research. This will assist in identifying the most appropriate biomarker for clinical use. More SNPs need to be genotyped to determine the proper expression of quantitative trait loci for mRNA and protein expression in cancer tissue and the periphery. Heptanediamide, N,Nʹ-di-benzoyloxy, among the ligands examined, showed an effective chemotherapeutic drug against cervical cancer for the target proteins HER2 and TP53. The lead compounds can be recommended for further medication and research in cancer treatment. Additional research using a variety of methods on cancer genetics, amplifiers, and complex patterns might reveal specific susceptibilities caused by underlying instability but also suggest new treatment plans and cures for the disease.
Data availability
The datasets generated and/or analyzed during the current study are available in the manuscript file.
References
Islam, S. M. S. et al. Non-communicable diseases (NCDs) in developing countries: A symposium report. Glob. Health 10, 1–8 (2014).
Tekalign, T. & Teshome, M. Prevalence and determinants of late-stage presentation among cervical cancer patients, a systematic review and meta-analysis. PLoS ONE 17, e0267571. https://doi.org/10.1371/journal.pone.0267571 (2022).
World Health Organization. Global Strategy to Accelerate the Elimination of Cervical Cancer as a Public Health Problem (World Health Organization, 2020).
Abdul Rahman, S. F., Xiang Lian, B. S. & Mohana-Kumaran, N. Targeting the B-cell lymphoma 2 anti-apoptotic proteins for cervical cancer treatment. Future Oncol. 16, 2235–2249 (2020).
Mostaid, M. S. et al. Elevated serum expression of p53 and association of TP53 codon 72 polymorphisms with risk of cervical cancer in Bangladeshi women. PLoS ONE 16, e0261984 (2021).
Sheema, et al. Molecular identification of hcv genotypes among injecting drug users having HCV and HIV co-infection. Bull. Biol. Allied Sci. Res. 2024, 71. https://doi.org/10.54112/bbasr.v2024i1.71 (2024).
Pervaiz, B. et al. Comparative analysis of different pathogenic microbe collected from mammalian milk. Bull. Biol. Allied Sci. Res. 2021, 32–32 (2021).
Sultana, T., Mitra, A. K. & Das, S. Evaluation of anti-cancer potential of Excoecaria agallocha (L.) leaf extract on human cervical cancer (SiHa) cell line and assessing the underlying mechanism of action. Future J. Pharm. Sci. 8, 3 (2022).
Anthony, E. J. et al. Metallodrugs are unique: Opportunities and challenges of discovery and development. Chem. Sci. 11, 12888–12917 (2020).
Talib, W. H., Alsayed, A. R., Barakat, M., Abu-Taha, M. I. & Mahmod, A. I. Targeting drug chemo-resistance in cancer using natural products. Biomedicines 9, 1353. https://doi.org/10.3390/biomedicines9101353 (2021).
Tejaputri, N. A. et al. Anticancer activity of ruellia britoniana flower on cervical hela cancer cells. Pharmacogn. J. 12, 29–34 (2020).
Uche, F. I., Onuchukwu, D., Ibezim, C. N. & Ogbu, H. I. Methanolic extract of Caladium bicolor leaves against selected clinical isolates. GSC Biol. Pharm. Sci. 6, 098–107 (2019).
Amin, F. et al. Antimicrobial susceptibility profile of various bacteria isolated from respiratory tract infection. Bull. Biol. Allied Sci. Res. 2023, 48–48 (2023).
Din, S., Fazal, M., Ishtiaque, A. & Ullah, A. Antimicrobial activity of lantana camara against pseudomonas aeruginosa, serratia marcescens and staphylococcus aureus to develop ointment based therapy. Bull. Biol. Allied Sci. Res. 2023, 33–33 (2023).
Awan, S. J. et al. Guar gum in therapeutics: A succinct exploration. Bull. Biol. Allied Sci. Res. 2024, 60. https://doi.org/10.54112/bbasr.v2024i1.60 (2024).
Amjad, N., Rehman, M., Khalid, M. & Amjad, I. Construction of the infectious molecule of beta satellite associated with ageratum yellow vein disease of ageratum conyzoides. Bull. Biol. Allied Sci. Res. 2018, 13–13 (2018).
Ullah, W. et al. Microbial profile and nutritional evaluation of broiler and domestic chicken meat from selected districts of Khyber Pakhtunkhwa, Pakistan. Bull. Biol. Allied Sci. Res. 2023, 34–34 (2023).
Zhang, D. et al. Evaluation on antidiabetic properties of medicinal plants from Myanmar. Sci. World J. 2021, 1424675. https://doi.org/10.1155/2021/1424675 (2021).
Ullah, I. et al. Prevalence and risk factors of helicobacter pylori infection among individuals with tobacco consumption habits in district Peshawar: A cross-sectional study. Bull. Biol. Allied Sci. Res. 2023, 42–42 (2023).
Ullah, A. et al. An overview of hepatitis c virus and liver cirrhosis in Pakistan. Bull. Biol. Allied Sci. Res. 2024, 64. https://doi.org/10.54112/bbasr.v2024i1.64 (2024).
Kalsoom, A. et al. In vitro antiproliferative potential of Cassia angustifolia extracts on HepG2 cells to combat liver cancer. Int. J. Appl. Exp. Biol. 3, 113–123 (2024).
Tulashie, S. K., Adjei, F., Abraham, J. & Addo, E. Potential of neem extracts as natural insecticide against fall armyworm (Spodoptera frugiperda (JE Smith)(Lepidoptera: Noctuidae). Case Stud. Chem. Environ. Eng. 4, 100130 (2021).
Nawaz, A., Jamal, A., Arif, A. & Parveen, Z. In vitro cytotoxic potential of Solanum nigrum against human cancer cell lines. Saudi J. Biol. Sci. 28, 4786–4792 (2021).
Larsson, P. et al. Optimization of cell viability assays to improve replicability and reproducibility of cancer drug sensitivity screens. Sci. Rep. 10, 5798 (2020).
Hegazy, M. G., Imam, A. M. & Abdelghany, B. E. Evaluation of cytotoxic and anticancer effect of Orobanche crenata methanolic extract on cancer cell lines. Tumor Biol. 42, 1010428320918685 (2020).
Mbugua, R. W., Njagi, E. M., Ngule, C. M. & Mwitari, P. Gene expression mediated antiproliferative potential and safety of selected medicinal plants against cancerous and normal cells. BioRxiv, 578948 (2019).
Alalwany, R. H. et al. Vascular endothelial growth factor isoforms differentially protect neurons against neurotoxic events associated with Alzheimer’s disease. Front. Mol. Neurosci. 16, 1181626 (2023).
Sattar, H. et al. Genetic association of polymorphism and relative mRNA expression of tumor necrosis factor-alpha gene in mastitis in Sahiwal cow. Int. J. Agric. Biol. 25, 701–708 (2021).
Baeshen, N. A. et al. In silico screening of some compounds derived from the desert medicinal plant Rhazya stricta for the potential treatment of COVID-19. Sci. Rep. 12, 11120 (2022).
Kiran, A. et al. Phytochemical profiling and cytotoxic potential of Arnebia nobilis root extracts against hepatocellular carcinoma using in-vitro and in-silico approaches. Sci. Rep. 13, 11376 (2023).
Upreti, S., Prusty, J. S., Pandey, S. C., Kumar, A. & Samant, M. Identification of novel inhibitors of angiotensin-converting enzyme 2 (ACE-2) receptor from Urtica dioica to combat coronavirus disease 2019 (COVID-19). Mol. Divers. 25, 1795–1809. https://doi.org/10.1007/s11030-020-10159-2 (2021).
Ravindranath, P. A., Forli, S., Goodsell, D. S., Olson, A. J. & Sanner, M. F. AutoDockFR: Advances in protein-ligand docking with explicitly specified binding site flexibility. PLoS Comput. Biol. 11, e1004586 (2015).
Arshad, F. et al. Evaluating the therapeutic efficacy of Swertia chirayita in liver cancer management. J. Biol. Regul. Homeost. Agents 38, 1385–1408 (2024).
Mendie, L. E. & Hemalatha, S. Molecular docking of phytochemicals targeting GFRs as therapeutic sites for cancer: An in silico study. Appl. Biochem. Biotechnol. 194, 215–231 (2022).
Umar, H. I., Siraj, B., Ajayi, A., Jimoh, T. O. & Chukwuemeka, P. O. Molecular docking studies of some selected gallic acid derivatives against five non-structural proteins of novel coronavirus. J. Genet. Eng. Biotechnol. 19, 16. https://doi.org/10.1186/s43141-021-00120-7 (2021).
Belayneh, Y. M., Yoseph, T. & Ahmed, S. A cross-sectional study of herbal medicine use and contributing factors among pregnant women on antenatal care follow-up at Dessie Referral Hospital, Northeast Ethiopia. BMC Complement. Med. Ther. 22, 146 (2022).
Ozili, P. K. in Social Research Methodology and Publishing Results: A Guide to Non-native English Speakers 134–143 (IGI global, 2023).
Naz, R. et al. GC-MS analysis, antimicrobial, antioxidant, antilipoxygenase and cytotoxic activities of Jacaranda mimosifolia methanol leaf extracts and fractions. PLoS One 15, e0236319 (2020).
Kavitha, R. Phytochemical screening and GC-MS analysis of bioactive compounds present in ethanolic extracts of leaf and fruit of Trichosanthesis dioica roxb. Int. J. Pharm. Sci. Res 12, 2755–2764 (2021).
Eo, H. J. et al. GC/MS analysis and anti-inflammatory effect of leaf extracts from Hibiscus syriacus through inhibition of NF-κB and MAPKs signaling in LPS-stimulated RAW264. 7 macrophages. Plant Biotechnol. Rep. 14, 539–546 (2020).
Saeed, N. M. et al. Anti-inflammatory activity of methyl palmitate and ethyl palmitate in different experimental rat models. Toxicol. Appl. Pharmacol. 264, 84–93 (2012).
Zhu, S. et al. Palmitic acid inhibits prostate cancer cell proliferation and metastasis by suppressing the PI3K/Akt pathway. Life Sci. 286, 120046. https://doi.org/10.1016/j.lfs.2021.120046 (2021).
Tanod, W. A., Yanuhar, U., Putra, M. Y. & Risjani, Y. Screening of NO inhibitor release activity from soft coral extracts origin Palu Bay, Central Sulawesi, Indonesia. Antiinflamm. Antiallergy Agents Med. Chem. 18, 126–141. https://doi.org/10.2174/1871523018666190222115034 (2019).
Balachandran, A. et al. Antioxidant, wound healing potential and in silico assessment of naringin, eicosane and octacosane. Molecules 28, 1043 (2023).
Koudehi, M. F., Ardalan, A. A. & Zibaseresht, R. Chemical constituents of an Iranian grown Capsicum annuum and their cytotoxic activities evaluation. Org. Med. Chem. Int. J 9, 148–154 (2020).
Ko, G.-A. & Cho, S. K. Ethyl linoleate inhibits α-MSH-induced melanogenesis through Akt/GSK3β/β-catenin signal pathway. Korean J. Physiol. Pharmacol. Off. J. Korean Physiol. Soc. Korean Soc. Pharmacol. 22, 53 (2018).
Ghaedi, E. et al. Retinol and α-tocopherol levels in the serum and subcutaneous adipose tissue of newly diagnosed basal cell carcinoma patients. Iran. J. Public Health 48, 1838 (2019).
Lozano-Grande, M. A., Gorinstein, S., Espitia-Rangel, E., Dávila-Ortiz, G. & Martínez-Ayala, A. L. Plant sources, extraction methods, and uses of squalene. Int. J. Agron. 2018, 1829160 (2018).
Di Nunzio, M. Hazelnuts as source of bioactive compounds and health value underestimated food. Curr. Res. Nutr. Food Sci. J. 7, 17–28 (2019).
Olivia, N. U., Goodness, U. C. & Obinna, O. M. Phytochemical profiling and GC-MS analysis of aqueous methanol fraction of Hibiscus asper leaves. Future Journal of Pharmaceutical Sciences 7, 1–5 (2021).
Pinto, M. E. et al. Antifungal and antioxidant activity of fatty acid methyl esters from vegetable oils. Anais da Academia Brasileira de Ciências 89, 1671–1681 (2017).
Ryu, J. et al. Comparative analysis of volatile compounds of gamma-irradiated mutants of rose (Rosa hybrida). Plants 9, 1221 (2020).
Kaushik, N. et al. Antiproliferative activity of pyracantha and paullinia plant extracts on aggressive breast and hepatocellular carcinoma cells. Appl. Sci. 10, 7543 (2020).
Graur, F. et al. Analysis of p53 protein expression in hepatocellular carcinoma. J. Gastrointestin Liver Dis. 25, 345–349 (2016).
Entezari, M. & Sheikhan, S. Evaluation of Bcl2 gene expression in MCF-7 human breast cancer cells under treatment of Centaurea behen extract and cisplatin. Arch. Adv. Biosci. 9, 11–16 (2018).
Salam, H. S. et al. Potential apoptotic activities of Hylocereus undatus peel and pulp extracts in MCF-7 and Caco-2 cancer cell lines. Plants (Basel) 11, 2192. https://doi.org/10.3390/plants11172192 (2022).
Nakamura, M., Obata, T., Daikoku, T. & Fujiwara, H. The association and significance of p53 in gynecologic cancers: The potential of targeted therapy. Int. J. Mol. Sci. 20, 5482 (2019).
Gorji-Bahri, G., Moradtabrizi, N. & Hashemi, A. Uncovering the stability status of the reputed reference genes in breast and hepatic cancer cell lines. PLoS One 16, e0259669 (2021).
Fatima, S. et al. The genome-wide bioinformatics analysis of 1-aminocyclopropane-1-carboxylate synthase (acs), 1-aminocyclopropane-1-carboxylate oxidase (aco) and ethylene overproducer 1 (eto1) gene family of fragaria vesca (woodland strawberry). Bull. Biol. Allied Sci. Res. 2023, 38–38 (2023).
Nawaz, K., Khan, S. & Bibi, A. Insights into scabies prevalence and risk factors. Bull. Biol. Allied Sci. Res. 2024, 68. https://doi.org/10.54112/bbasr.v2024i1.68 (2024).
Shukla, S., Dass, J. & Pujani, M. p53 and bcl2 expression in malignant and premalignant lesions of uterine cervix and their correlation with human papilloma virus 16 and 18. South Asian J. Cancer 3, 48–53. https://doi.org/10.4103/2278-330x.126524 (2014).
Laka, K. & Mbita, Z. P53-Related anticancer activities of Drimia calcarata bulb extracts against lung cancer. Front. Mol. Biosci. 9, 876213 (2022).
Bykov, V. J., Eriksson, S. E., Bianchi, J. & Wiman, K. G. Targeting mutant p53 for efficient cancer therapy. Nat. Rev. Cancer 18, 89–102 (2018).
Saravanan, R., Raja, K. & Shanthi, D. GC-MS analysis, molecular docking and pharmacokinetic properties of phytocompounds from Solanum torvum unripe fruits and its effect on breast cancer target protein. Appl. Biochem. Biotechnol. 194, 529–555. https://doi.org/10.1007/s12010-021-03698-3 (2022).
Hassan, N. et al. Antiviral response of drugs used against hbv patients of Khyber Pakhtunkhwa, Pakistan. Bull. Biol. Allied Sci. Res. 2023, 49–49 (2023).
Gohar, M. et al. Prevalence of hepatitis b virus and genotypes in the region of khyber pakhtunkhwa Pakistan. Bull. Biol. Allied Sci. Res. 2023, 53–53 (2023).
Bhatti, M., Ahmad, S., Bilal, S. & Iqbal, M. Evaluation of different strains of entmopathogenic fungi as potential agents for the management of Tribolium castaneum. Bull. Biol. Allied Sci. Res. 2023, 52–52 (2023).
Taymaz-Nikerel, H., Karabekmez, M. E., Eraslan, S. & Kırdar, B. Doxorubicin induces an extensive transcriptional and metabolic rewiring in yeast cells. Sci. Rep. 8, 13672. https://doi.org/10.1038/s41598-018-31939-9 (2018).
Maliehe, T. S., Tsilo, P. H. & Shandu, J. S. Computational evaluation of ADMET properties and bioactive score of compounds from Encephalartos ferox. Pharmacogn. J. 12, 1357–1362 (2020).
Acknowledgements
This work was supported by the Institute of Molecular Biology and Biotechnology, The University of Lahore, Lahore, Pakistan.
Author information
Authors and Affiliations
Contributions
A.K. Prepared original draft, implemented experiments, established cell models, collected data and plotted the graphs; A.A. Conceptualized, supervised and revised the manuscript; M.S. Facilitated in analysis, result interpretation and revising the manuscript; T.M. Guided in the cell culture lab; M.A.B.A. Data interpretation and assisted in revising the manuscript; H.S. Assisted in expression analysis protocols and implementation; G.S. Reviewed and analyzed the data statistically; Q.A. Supervised and validated the data; M.A.J. Edited and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kalsoom, A., Altaf, A., Sarwar, M. et al. GC–MS analysis, molecular docking, and apoptotic-based cytotoxic effect of Caladium lindenii Madison extracts toward the HeLa cervical cancer cell line. Sci Rep 14, 18438 (2024). https://doi.org/10.1038/s41598-024-69582-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69582-2
Keywords
This article is cited by
-
Unveiling the cytotoxic potential of four Callistemon fruit extracts against breast and colon cancer: a combined metabolomic and in silico approach
BMC Complementary Medicine and Therapies (2026)
-
Phytochemical characterization and in vitro assessment of antioxidant, antibacterial, and anticancer activities of Persicaria capitata
Discover Applied Sciences (2025)