Abstract
The 4T1 model is extensively employed in murine studies to elucidate the mechanisms underlying the carcinogenesis of triple-negative breast cancer. Molecular biology serves as a cornerstone in these investigations. However, accurate gene expression analyses necessitate data normalization employing housekeeping genes (HKGs) to avert spurious results. Here, we initially delve into the characteristics of the tumor evolution induced by 4T1 in mice, underscoring the imperative for additional tools for tumor monitoring and assessment methods for tracking the animals, thereby facilitating prospective studies employing this methodology. Subsequently, leveraging various software platforms, we assessed ten distinct HKGs (GAPDH, 18 S, ACTB, HPRT1, B2M, GUSB, PGK1, CCSER2, SYMPK, ANKRD17) not hitherto evaluated in the 4T1 breast cancer model, across tumors and diverse tissues afflicted by metastasis. Our principal findings underscore GAPDH as the optimal HKG for gene expression analyses in tumors, while HPRT1 emerged as the most stable in the liver and CCSER2 in the lung. These genes demonstrated consistent expression and minimal variation among experimental groups. Furthermore, employing these HKGs for normalization, we assessed TNF-α and VEGF expression in tissues and discerned significant disparities among groups. We posit that this constitutes the inaugural delineation of an ideal HKG for experiments utilizing the 4T1 model, particularly in vivo settings.
Similar content being viewed by others
Introduction
Breast cancer, excluding cases of non-melanoma skin cancer, stands as the most commonly diagnosed cancer globally, comprising approximately 2.3 million new cases (approximately 11.7% of all cancers or 24.55% solely in women). It also ranks as the leading cause of cancer-related mortality among women in developing countries1. In most cases where, the disease remains localized, whereas the presence of metastases signifies a grim prognosis. Metastases imply a poor prognosis and can manifest in various organs, including but not limited to the liver, lungs, brain, and bones2. For a better understanding of the complex dynamics of cancer, tumor models are necessary to develop new diagnostic and therapeutic strategies in cancer3. In vivo experimental models have been extensively employed in oncology as pivotal tools to enhance our understanding of the mechanisms underlying carcinogenesis and various modalities of cancer treatment4,5,6. The 4T1 murine breast carcinoma model is one of the most extensively studied models in breast cancer research. Its popularity stems from its characteristics as a syngeneic model for human breast cancer, coupled with its notable high invasiveness and metastatic potential. Moreover, it exhibits a clinical course akin to that observed in patients, making it a valuable tool for studying the disease and exploring potential therapeutic interventions7,8,9. Furthermore, the 4T1 model serves as a valuable resource for investigating triple-negative breast cancer (TNBC), a subtype characterized by the absence of estrogen, progesterone, and epidermal growth receptor human 2 (HER-2) receptors. TNBC is characterized by high proliferation, poor differentiation, and a generally unfavorable prognosis10,11. Thus, the use of the 4T1 model enables researchers to explore the TNBC pathophysiology and develop potential therapeutic strategies targeting precisely this aggressive form of breast cancer. We emphasize that the TNBC phenotype accounts for approximately 20% of breast cancers diagnosed annually12,13.
Carcinogenesis has a complex etiology and involves multiple levels of regulation, encompassing endogenous molecular pathways14. Disorders in genes can instigate uncontrolled cell growth, leading to tissue damage. Understanding these alterations can pave the way for developing strategies to prevent dysfunctions in the cell cycle15,16,17. Therefore, the identification of genes and the analysis of their expression are imperative for unraveling the pathogenesis of cancer. Furthermore, they furnish crucial information for diagnosing and prognosticating the disease18,19. Molecular biology is crucial in deepening our comprehension of carcinogenesis, rendering it a valuable tool in cancer prevention efforts20. Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR) stands out as a crucial tool in molecular biology studies, particularly for analyzing gene expression. Its reproducibility, rapid quantification of mRNA transcription, and capacity to furnish vital information regarding gene expression levels make it indispensable. This method enables comparative analysis between control groups and those exposed to specific treatments, thereby facilitating comprehensive insights into molecular responses21,22,23,24,25.
Nevertheless, for accurate gene expression analyses, employing a housekeeping gene (HKG) is crucial. These genes serve as internal controls to normalize differences between samples, ensuring precise and reliable results while mitigating potential errors26,27. On the other hand, the genes commonly employed for this purpose are associated with metabolism, and cancer is a disease that significantly impacts metabolic processes28,29. GAPDH, 18 S, and ACTB are commonly cited for normalization in the literature. However, studies have indicated that these genes’ expression can vary depending on the specific experimental conditions applied30,31,32,33. Therefore, further investigations are necessary to comprehensively elucidate the suitability of these genes as HKGs across diverse experimental scenarios, including their applicability in the 4T1 model.
The characterization of suitable HKGs in oncological research has indeed been extensively explored in the literature34,35,36,37, including in breast cancer model studies38,39and oncology patients samples40,41,42,43. In this regard, studies utilizing the 4T1 in vivo model and conducting gene expression analyses have already been undertaken44,45,46. Nevertheless, despite the existing research, studies specifically evaluating the stability of HKGs in the in vivo 4T1 model were lacking. Hence, we aimed to identify the gene with the most stable expression for data normalization during experiments involving RT-qPCR analysis of breast cancer in mice and metastasis studies.
Materials and methods
Animals and experimental design
For this study were used a total of 36 female BALB/c mice (16–20 g) 10–11 weeks old (Specific pathogen-free) obtained from the Central Animal Facility of the Universidade Federal de Minas Gerais, Brazil and maintained at the Animal Facility of the Department of Parasitology of the Universidade Federal de Minas Gerais under controlled conditions of temperature (24 ± 1˚ C) and lighting (12-hour light-dark cycle). During the experimental period, the mice had access to filtered water and commercial chow (Nuvilab Cr-1, Nuvital Nutrients, Brazil) ad libitum. Mice were randomized and received a subcutaneous injection of 0.1 mL of suspension of RPMI without complements containing 1.0 × 1064T1 cells in the left flank as described by Gao et al. (2011)47. Mammary gland tissues present in the flank region and close to the inner thigh of the control group were extracted and used to compare gene expression with primary tumor.
This study was divided into two stages. First, due to the lack of a standard collection point in metastasis studies in the 4T1 model, we decided to evaluate the evolution of metastases using different techniques and better describe some clinical changes we did not find in other studies in the literature. Thus, mice were euthanized on the 14th, 21st, and 28th days post-injection of cells with 8.5 mg/kg of xylazine and 130 mg/kg of ketamine. These collection points were chosen according to the descriptions of other studies of metastasis48,49,50,51. This step was crucial to identify organ metastasis using scintigraphic images and histopathological analyses for subsequent evaluation of gene expression. Then, we selected the best collection point, and tissues were collected to carry out molecular biology analysis (Fig. 1A).
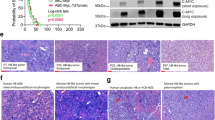
Steps of experimental design and aspects of in vivo development of 4T1 breast carcinoma in BALB/c mice. (A) Experimental design of cancer induction with 1 × 106 4T1 cells representing the experimental design to step 1 to monitor evolution of 4T1 breast carcinoma and metastases and step 2 corresponding to tissue collection to qPCR experiments after determining the best collect point. (B) Clinical aspects of the evolution of 4T1 breast carcinoma in BALB/c mice including survival rate, clinical score over 30 days after injection of cells, assessment of tumor growth (mm3) over 30 days after injection of cells and evolution of ulcerated areas in the primary tumor over 30 days after injection of cells. (C) Scintigraphic images and biodistribution of 99mTc-HYNIC-βAla-Bombesin(7−14) in different tissues. The red arrow indicates renal clearance of peptide radiolabeled. The yellow arrow indicates an exclusion area of intense bladder uptake. The red circle indicates the primary tumor area. The yellow circle indicates an area without a tumor. The orange circle indicates the liver uptake area. (D) Percentage of injected dose per tissue gram in ex vivo experiments in liver, lung, and brain at 14, 21, and 28 days after injection of cells. In B, the log-rank test (Mantel-Cox) was used in the survival curve, and to calculate the Hazard Ratio, the Mantel-Haenszel test was used between the control group (n = 6) and animals with 4T1 breast carcinoma (n = 15); Mann-Whitney test was used in the clinical score, tumor growth and evaluation of ulcerated areas. The one-way ANOVA test, followed by the Tukey test, was used to analyze statistical differences between the control group (n = 6) and 4T1 group (n = 6) in D. Data are represented as mean ± standard error of the mean (SEM). Significant differences are represented by *p < 0.05; **p < 0.01; ***p < 0.001.
Cell culture
Previously, cryopreserved murine breast carcinoma 4T1 cell line aliquots from passage 14 (p14) were utilized. The aliquots were thawed and cultured in RPMI supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin at 37 °C and 5% CO2 in a humidified atmosphere (Water-Jacked Incubator, Thermo Scientific, USA). For cancer induction, three cell passages were conducted after thawing to ensure the malignancy of the cell line, followed by two wash steps with PBS to completely remove fetal bovine serum and then resuspend in RPMI without complements for subsequent inoculation. Subsequently, 1.0 × 106cells were inoculated subcutaneously into the left flank as described by Gao et al. (2011)47.
Survival curve, clinical parameters and score evaluation
On the day of the injection of cells (day 0), mice were monitored every other day. We apply a clinical score adapted from Vandermosten et al. (2018)52. In this study, piloerection (PE), shivering (Sh), abnormal breathing (AB), and trunk curl (TC) were evaluated to calculate a clinical score of disease severity. Scores were given as 0 (absent) or 1 (present) for PE, Sh, AB, and TC. The total clinical score was calculated by the following formula: (PE + Sh + AB + TC). Furthermore, we also evaluated the evolution of the ulcerated areas in the tumors. As we did not find descriptions in previous studies, we created a score and made it available to facilitate other studies using the 4T1 model (Supplementary material 1). Our score uses a scale that varies from 0 to 4 according to the appearance of the ulcer (Supplementary Table S1; Fig. S1). To assess the size of tumors, we measured the tumor dimensions using a caliper, and the tumor volume was calculated using the equation V(mm3) = axb2/2 53, where V is a volume, a is the largest diameter and b is the smallest diameter.
Scintigraphic images
To monitor the evolution of tumors, we implemented a broad and complementary approach using radioisotopes. This technique is already used in other in vivo cancer models5,54,55 but not applied to the 4T1 model. Thus, at 14th, 21st and 28th days post injection of cells, aliquots containing 100µL and 3.7 MBq of HYNIC-βAla-Bombesin(7–14) peptide radiolabeled with 99mTc (Technetium) (See item Radiolabeling of HYNIC-βAla-Bombesin(7−14) and evaluation of radiochemical purity) were administered into the tail vein of mice. Thus, 4 h after injection of 99mTc-bombesin, mice were anesthetized with a solution of ketamine (80 mg/kg) and xylazine (15 mg/kg) and then placed in a prone position under a gamma camera (Mediso, Hungary) employing a low-energy high-resolution collimator. Images were acquired using a 256 × 256 × 16 matrix size with a 20% energy window set at 140 keV for 600 s.
Ex vivo biodistribution experiments
After scintigraphic images, mice were euthanized with 8.5 mg/kg of xylazine and 130 mg/kg of ketamine, and tissues of interest, such as tumor, lungs, liver, and brain, were removed and placed in pre-weighed plastic test tubes. Then, each tissue was weighed, and its associated radioactivity was measured using an automatic gamma counter (Wizard, Finland). Results were expressed as the percentage of injected dose per gram of tissue (% ID/g). A standard dosage containing the same injected amount was counted simultaneously in a separate tube, which was defined as 100% radioactivity. The data were expressed as dose % using the following equation: %dose = cpm in g of tissues/cpm of standard x 100; where cpm represents the counts of radioactivity per minute.
Radiolabeling of HYNIC-βAla-Bombesin(7−14) and evaluation of radiochemical purity
To perform scintigraphic images and ex vivo biodistribution experiments, the HYNIC-βAla-Bombesin(7−14) was radiolabeled as described by Barros et al., 201256. In an amber-sealed vial, we added a 0.9% (w/v) saline containing 20 mg of tricine and 5 mg of EDDA for each 0.5 mL. Then, 10 µg of HYNIC-βAla-Bombesin(7−14) and 10 µL of stannous chloride (SnCl2) 1 mg/mL in 0.25 N/mL HCl were added. We adjusted the pH to 7–8 with 0.1 N NaOH and added 37 MBq of Sodium Pertechnetate (Na99mTcO4). The solution was heated at 100 °C for 15 min and cooled to room temperature.
Radiochemical purity analysis was performed using thin-layer chromatography on silica gel strips (Merck). Methyl-ethyl-ketone was used to determine the amount of free technetium (99mTcO4−), and acetonitrile: water (1:1) was used to quantify hydrolyzed technetium (99mTcO2), as described by Barros et al., 201256. Radioactivity was measured using an automatic gamma counter (Wizard, Finland).
Histopathological analysis
The left lobe of the liver, lungs, and brain were removed from the mice in each group to confirm metastases presence. Samples of these tissues were fixed in 10% buffered formalin solution for three days, then gradually dehydrated in ethanol, diaphanized in xylol, and embedded in paraffin blocks cut at a thickness of 4 μm and fixed on microscope slides. Slides were stained with Hematoxylin and Eosin (H&E) and analyzed by two experienced pathologists who were blinded to the study.
Sample collection, RNA isolation and cDNA synthesis
Tissues are collected, inserted in RNA later solution, and stored at -80 °C for later analysis. Only tumors and samples with confirmed metastases were utilized for total RNA isolation. Samples were processed using NucleoSpin®RNA Plus Kit (Macherey-Nagel, Düren, Germany) according to the manufacturer’s instructions using 30 mg of tissue. The RNA integrity was assayed by 1% agarose gel electrophoresis, and the concentration and purity were measured using NanoDrop 2000 Spectrophotometer (Thermo Fisher Scientific Inc., USA). Samples with a range of 1.8–2.1 to 260/280nm and 260/230nm absorbance ratio were considered pure, protein and ethanol-free RNA, indicating that they were suitable for subsequent cDNA synthesis22,24. cDNA was reverse transcribed from 2 µg of total RNA using the High-Capacity cDNA Reverse Transcriptase Kit (Applied Biosystems Inc., Foster City, CA) in a total volume of 20µL according to the manufacturer’s instructions. The cDNA was diluted 1:10 for subsequent RT-qPCR. Standard curves were generated by diluting the cDNA, without diluting, 1:10, 1:100, and 1:1000.
Selection of candidate housekeeping genes
The HKGs candidates were selected based on those commonly used for RT-qPCR studies in breast cancer. Thus, we selected ten different candidate genes with different cellular functions, such as glyceraldehyde 3-phosphate dehydrogenase (GAPDH), 18s (18 S ribosomal RNA), Actin beta (ACTB), Hypoxanthine phosphoribosyltransferase 1 (HPRT1), Beta-2-Microglobulin (B2M), Glucoronidase Beta (GUSB) and Phosphoglycerate kinase (PGK1)39,40,41,42,57,58,59,60and additionally, we chose to include three other genes Coiled-Coil Serine Rich Protein 2 (CCSER2)42,60, Ankyrin Repeat Domain 17 (ANKRD17), Symplekin Scaffold Protein (SYMPK)42that emerged as promising on the same theme. In this way, we can expand the scope of our analysis, exploring potential genetic markers that can contribute to a more comprehensive and accurate understanding of breast cancer. Furthermore, to measure the impact of HKGs choice on data normalization, Vascular endothelial growth factor (VEGF) and Tumor necrosis factor alpha (TNF-α), previously associated with development of breast tumor51,61were used to take relative expression levels using Livak or Delta-Delta Ct method to validate the choice of HKG62,63.
Primer design, specificity and efficiency
The sequences of GAPDH (F: ACCCAGAAGACTGTGGATGG; R: CACATTGGGGGTAGGAACAC), 18 S (F: GCCGTTCTTAGTTGGTGGAG; R: AACGCCACTTGTCCCTCTAA), ACTB (F: TGTTACCAACTGGGACGACA; R: GGGGTGTTGAAGGTCTCAAA), HPRT1 (F: TTGGGCTTACCTCACTGCTT; R: CTAATCACGACGCTGGGACT), and B2M (F: TCTCACTGACCGGCCTGTAT; R: GTATGTTCGGCTTCCCATTC) were obtained in a recent study25, while the others genes (GUSB, PGK1, CCSER2, ANKRD17, SYMPK, VEGF and TNF-α) were designed through mice nucleotide sequences retrieved from NCBI using the accession numbers specified in Table 1. Subsequently, the web server Primer 3 input (Version 4.1.0) (https://primer3.ut.ee/) was employed for designing all primer sequences based on the following criteria: primer length ~ 18–22 bp, melting temperature 59–61 °C, % of guanine and cytosine content 40–60% and amplicon length of 90–200 bp. To ensure primer quality, we assessed the potential for primer-dimer or secondary structure formation using the RNAfold (http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi) and Autodimer software (National Institute of Standards and Technology, USA). The anticipated amplicon sequences were aligned with those cataloged in the international databases through the Basic Local Alignment Search Tool (BLAST) to confirm specificity. All details regarding the forward and reverse primers, including sequences, are presented in Table 1.
After, serial dilutions of samples were performed for all paired primers designed to determine the amplification efficiency (E%) and coefficient of determination (R2), and the results were used to depict a standard curve. A linear regression was then performed to determine the slope of the line, which is used to calculate the R2 and the efficiency (E%) using the formula: E% = 10 (−1/slope) x 100 64. We considered primer pairs within the range of 90-115% as ideal for proper function25. The specificity of the amplified products was confirmed by visualization of the expected amplicon size on 2% agarose gel electrophoresis.
RT-qPCR
The qPCR was performed using a 7500 Real-Time PCR system (Applied Biosystems™) according to the manufacturer’s instructions with the PowerUP™ Sybr green master mix kit (Applied Biosystems™). We use a total volume of 10µL in reactions containing 5µL from the kit, 2µL cDNA (equivalent to 50 ng of cDNA), 0.5µL of each specific forward and reverse primers (10µM), and 2µL ultrapure nuclease-free water was used for the reaction. Triplicates of each sample and a non-template reaction (negative control) were included. Thermal cycling was performed, starting with an initial step at 50 °C for 2 min, followed by 95 °C for 10 min and 40 cycles of denaturation at 95 °C for 15s and annealing/extension at 60 °C for 60s. Melting curve analysis was performed from 70 °C to 95 °C at 0.1 °C/s for all genes studied. Threshold cycle (Ct) values for each sample were obtained by calculating the arithmetic mean of triplicate values.
Stability analysis of HKGs candidates
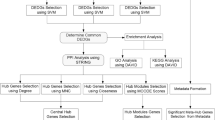
Initially, the distribution of Ct values was analyzed. Then, four different software programs were used to calculate and identify the most stable gene in each tissue: geNorm, which determines the stability expression by the average stability value of M of each gene. The values >1.5 are not suitable, and the lowest M value among genes is high stability65; BestKeeper, an Excel-based tool that performs the classification of HKGs through paired correlation analysis, calculation of Standard Deviation [± Crossing Point] and coefficient of variance (CV) for each gene from the Ct value, Lower Std. Dev. [± CP] values indicate optimal stability, or when the value is more than 1, gene expression is unstable26; NormFinder is also a tool based on Microsoft Excel that provides stability values for each gene inter and intra groups (lower stability values indicate the highest stability)66and RefFinder, a web-based analysis tool which integrates geometric means and a weight is assigned of the stability values showed by these other tools ranked the most stable gene67,68.
Statistical analysis
Grubbs’ test was used to detect possible outliers, and the Shapiro-Wilk test was used to verify data distribution. For survival curve analysis, the log-rank test (Mantel-Cox) was used, and to calculate the Hazard Ratio, the Mantel-Haenszel test was used between the groups. For the comparison between the control group and the 4T1 group on each day, the Mann-Whitney test was used, and to compare groups on the same day and different days, the One-Way ANOVA test, followed by the Tukey test, was used. The GraphPad Prism 8.0.1 software (GraphPad Inc, USA) was used in these analyses to plot values obtained from other programs. In addition, the significance level was set at a p-value equal to or less than 0.05.
Results
Evolution of 4T1 murine breast carcinoma development and confirmation of metastases in target tissues
The initial step in the study was analyzing tumor progression and assessing animal survival, a pivotal procedure for determining the collection points necessary for analysis. Mice were inoculated with 1.0 × 106 cells in the left flank to induce breast cancer, followed by monitoring every two days for clinical signs and tumor evolution. Survival analysis revealed that mice bearing 4T1 cancer exhibited a median survival of 34 days, with a Hazard Ratio value of 17.94 (Fig. 1B). By day 30 post-injection of cells, 30% of the mice succumbed, reaching 50% mortality by day 34 and complete mortality by day 36 (Fig. 1B). Based on these findings, tumor growth, clinical sign assessment, and the evolution of ulcerated areas were documented up to day 30.
We observed the onset of clinical signs starting from the 12th day in a few animals, initially manifesting as piloerection and progressively worsening with the emergence of tremors and abnormal respiration (Fig. 1B). Notably, after the 30th day, some animals presented trunk curvature close to the day of natural death (data not shown). Regarding tumor size, palpation became feasible from the 6th day, followed by exponential growth, reaching a mean size of 3800 mm^3 by the 30th day (Fig. 1B). It is worth mentioning that after the 30th day, the maximum size reached was 4598.20, observed in an animal that succumbed at 34 days (data not shown). Additionally, we devised a scoring system based on the evolution dynamics of 4T1 ulcers in mice to contribute to future studies in the field (Table S1; Fig. S1). Thus, we observed initial appearances in some mice from the 10th day, which worsened over time, with some animals reaching a score of three, indicative of the presence of an ulcerated area occupying more than 50% of the circumference of the primary tumor and infiltrating inwards (intense) (Fig. 1B).
To collect tissues beyond the primary tumor, we meticulously determined the precise timing at which metastases were firmly established in target organs such as the brain, liver, and lungs. Consequently, we selected days 14, 21, and 28 and implemented a comprehensive and complementary approach utilizing radioisotopes and a peptide (HYNIC-βAla-Bombesin (7−14)) to study the 4T1 model in vivo. This multifaceted strategy encompassed scintigraphy and biodistribution studies (ex vivo) to achieve a more dynamic understanding of tumor dissemination. While this technique had been previously employed in other in vivo cancer models5,55, its application to the 4T1 model represented a novel endeavor.
To assess the progression of metastases in different tissues, the animals received injections of 99mTc-HYNIC-βAla-Bombesin (7−14) on days 14, 21, and 28. On the 14th day, we observed radiotracer uptake in the liver, which progressively increased at 21 and 28 days. However, accurate detection of radiation in the lungs and brain proved challenging (Fig. 1C). As these were qualitative data, some animals were euthanized to quantify ex vivo radiation of 99mTc-HYNIC-βAla-Bombesin(7−14) to obtain quantitative data and gain a better understanding of tumor and metastasis evolution (Fig. 1C). The ex vivo results revealed different patterns than those observed in the scintigraphic images, except for the liver, where there was a progressive increase in uptake over time. We observed a significant increase in uptake in the lungs from 21 days, which further increased at 28 days, indicating an escalation in metastases in this organ (Fig. 1D). It is noteworthy that no changes in uptake were observed in the brain, suggesting the absence of metastases in this tissue at the collection points (Fig. 1D).
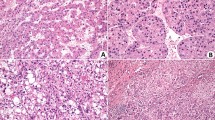
To complement the results obtained from scintigraphy and ex vivo examination, we also collected tissues at 14, 21, and 28 days for histopathological evaluation (Fig. 2). The histopathological findings were consistent with those obtained in the ex vivo experiments using HYNIC-βAla-Bombesin(7–14) radiolabeled with 99mTc. At 14 days, metastases were observed in the liver in all animals (Fig. 2A), while they were not detected in the lungs (Fig. 2B) and brain (Fig. 2C). However, by day 21, metastases were evident in the lungs, and their presence intensified by day 28 lungs (Fig. 2B). Notably, metastases in the brain were not evidenced at any of the collection points through histopathological analysis (Fig. 2C). The number of animals with metastasis and descriptions of the foci can be seen in Table 2.
Representative hematoxylin and eosin staining of the histopathology of the analyzed tissues at 14, 21 and 28 days after injection of cells. (A) Staining of liver sections. (B) Staining of lung sections. (C) Staining of brain sections. In liver (A), symbols represent metastatic foci (arrowhead) and necrosis area (star), while in lung (B), symbols represent alveoli (a), inflammatory infiltration foci (arrow) and metastatic foci in ($). In brain (C) no differences were observed throughout the days in parenchyma. Bar = 50 μm.
Specificity and amplification efficiency of the candidate gene primers
After observing the progression of breast cancer and the development of metastases, we determined that the 21st day was the optimal time point to collect samples for molecular biology analyses. Before conducting gene expression analyses, we assessed the specificity and amplification efficiency of the primers, a critical step that directly influences the accuracy of the results obtained in real-time PCR.
Upon analysis, all designed primers (GUSB, PGK1, CCSER2, SYMPK, ANKRD17, VEGF, and TNF-α) exhibited satisfactory efficiency, with E% values ranging between 95.30% (PGK1) and 111.71% (ANKRD17). Additionally, the R2 value ranged from 0.968 (CCSER2) to 1.000 (PGK1 and SYMPK) (Table 1). Dissociation curves were replicated to confirm the specificity of the real-time PCR assays, as evidenced by the presence of a single peak. Notably, the formation of nonspecific products was not observed in the curves or respective blanks for each gene (Fig. S2).
Distribution of ct values of HKGs in different tissue
Our results reveal that 18 S exhibited the lowest Ct value in all tissues evaluated, while GUSB demonstrated the highest Ct value in the tumor and lung, and SYMPK and CCSER2 in the liver (Fig. 3A). Additionally, we highlight that the expression of the HKGs varied between the cancer group and the control group in some tissues, with the most significant variations observed in the expression of the genes in the primary tumor and the liver. Notably, CCSER2 did not exhibit variation in Ct values in any of the tissues analyzed, while SYMPK consistently varied in all tissues. Specifically, in the primary tumor, Ct values varied in the SYMPK, B2M, HPRT1, ACTB, 18 S, and GAPDH genes (Fig. 3A). In contrast, in the liver, variation was observed in SYMPK, PGK1, GUSB, B2M, HPRT1, and 18 S (Fig. 3A). In the lung, variations were observed only in the ANKRD17, SYMPK, and GAPDH genes (Fig. 3A). These findings underscore that cancer can induce variations in Ct values, indicating potential alterations in gene expression.
Variation of gene expression levels by different methods. (A) Expression levels of different housekeeping genes based on Ct values in different tissues in control and 4T1 groups. (B) Stability test results are calculated using geNorm software in different tissues. (C) Stability test results were calculated using NormFinder software in different tissues. (D) Classification of the most stable genes calculated by RefFinder software in different tissues. In (A), data are represented as mean ± standard error of the mean (SEM). The Mann-Whitney test was used to analyze statistical differences between the control group (n = 6) and the 4T1 group (n = 6). In (B), data are represented as a classification of housekeeping genes according to the M value in each tissue. In (C), data are represented as a classification of housekeeping genes according to the S value in each tissue. Significant differences are represented by *p < 0.05; **p < 0.01; ***p < 0.001.
Stability test and identification of the best HKGs in each tissue
The geNorm software (version 3.5) was initially utilized to calculate the average expression stability (M Value) among genes in different tissues. It is important to note that not all genes presented values below 1.5, which is considered the threshold for considering an excellent HKG. In the tumor, 18 S exhibited the highest M value, while five genes (ACTB, GUSB, ANKRD17, PGK1, and CCSER2) obtained values below 1.5, albeit with significant variability (Fig. 3B). In this context, CCSER2 or the combination of PGK1/CCSER2 emerged as the best options for normalizing expression data in the primary tumor. In other tissues, SYMPK had the highest M value. Additionally, the liver had the lowest number of genes (HPRT1, ANKRD17, GAPDH, and ACTB) that reached values below 1.5 (Fig. 3B). Among the options, ACTB or the combination of GAPDH/ACTB were identified as the best for normalizing data in this organ with metastasis (Fig. 3B). In the lung, there was a greater diversity of genes below 1.5, totaling eight (ANKRD17, GUSB, ACTB, HPRT1, CCSER2, 18 S, B2M, and PGK1). However, 18 S, B2M, PGK1, or the combination of B2M/PGK1 were deemed the best options in the lung (Fig. 3B).
Subsequently, the NormFinder software was utilized to assess the best HKG. The gene with the lowest S value is considered the most stable. In the primary tumor, GAPDH was classified as the most stable gene (Fig. 3C), while in the liver and lung, it was HPRT1 (Fig. 3C). Conversely, 18 S was classified as the least stable in the tumor and lung, while in the liver, the least stable was CCSER2. These results also exhibited some discrepancies with geNorm, with genes classified in different positions, except in the tumor, where 18 S was consistently classified as stable (Fig. 3C). Regarding the analysis carried out in BestKeeper, unlike the programs used previously, ACTB was identified as the gene with the most outstanding stability in all tissues. In contrast, CCSER2 was identified as the gene with the lowest stability in the tumor, and SYMPK recorded the lowest stability in the liver and lung (Table 3).
As gene stability analyses exhibited discrepancies across programs, the RefFinder software was employed to rank the most stable genes by tissue (Fig. 3D). In the tumor, the gene that exhibited the greatest stability, as indicated by NormFinder, was GAPDH, while CCSER2 was identified as the gene with the minor stability, consistent with the results generated by BestKeeper (Fig. 3D; Table 3). In the liver, HPRT1 was identified as the most stable gene, as indicated by NormFinder, while PGK1 was classified as less stable. However, no software categorized it as such (Fig. 3D). In the lung, CCSER2 emerged as the most stable gene, without competition from other software, while SYMPK was identified as less stable, aligning with the values generated by geNorm and BestKeeper (Fig. 3B, D; Table 3).
Validation of HKGs candidates’ accurateness of the selected HKGs
After selecting the most stable genes for each tissue, we assessed the impact of these and other less stable genes on normalizing the expression of VEGF and TNF-α in tissues from control mice and animals with confirmed cancer and metastases, as determined by histopathological analyses and techniques involving radioisotopes. Across all tissues, less stable genes were observed to potentially overestimate expression when normalized with their respective HKG or demonstrate lower expression compared to the control. Conversely, this phenomenon was not observed with the genes identified as more stable.
In the tumor, both VEGF and TNF-α expression, when normalized by GAPDH, exhibited higher levels compared to the respective control (Fig. 4A). In the liver, normalization by HPRT1 effectively increased the expression of VEGF and TNF-α (Fig. 4B). Similarly, in the lung, normalization by CCSER2 was efficient for both VEGF and TNF-α (Fig. 4C). These findings underscore the critical importance of selecting appropriate HKGs for accurate normalization in gene expression studies.
Normalization with the different housekeeping gene candidates on VEGF and TNF-α expression levels in mice’s tumor, liver, and lung with or without 4T1 breast carcinoma. The blue line highlights the gene expression normalized with the most stable gene; the Red line highlights the gene expression normalized with the least stable gene. The Mann-Whitney test was used to analyze statistical differences between the control group (n = 8 and the 4T1 group (n = 8). Significant differences are represented by *p < 0.05; **p < 0.01; ***p < 0.001.
Discussion
The assessment of HKGs has been widely explored in the context of breast cancer, both in human samples40,69and in experimental models27,38,42,60,70, except in the 4T1 model. To the best of our knowledge, this study is the first to compare different HKGs for normalization of expression data in vivo model of 4T1 breast carcinoma, including metastases, which evolve naturally and reach organs such as the liver, lung, and brain71,72,73. Despite the widespread use of the 4T1 breast carcinoma model, a notable observation is the lack of standardization regarding the collection point for experiments involving metastases74. This variability in experimental protocols may introduce inconsistencies and challenges in data interpretation and comparability across studies. Establishing a standardized approach for the timing of sample collection from metastatic sites is crucial for ensuring reproducibility and reliability in research findings using this model. By addressing this issue, future studies can enhance the consistency and robustness of experimental outcomes, ultimately advancing our understanding of breast cancer metastasis and improving the translational relevance of preclinical research.
Thus, a crucial step in our study, preceding the assessment of HKGs, was to identify the ideal point for the collection of organs to ensure the presence of metastases in different organs. Our results of liver metastases differed from prior studies, where evidence of liver metastases at 28 days post-cell inoculation48and in some studies at 20 days50,74, albeit in a limited number of animals, whereas we observed it in all animals of the group at 14 days. In comparison, we came across only one study that reported the presence of lung and liver metastases on the 8th day, with liver metastases being more intense than lung metastases72. We highlight that a scarcity of studies of liver metastasis compared to the lung was found.
On the other hand, numerous studies have reported the presence of metastases using the 4T1 model from 20 days onwards49,50,51,75. The outcomes we observed closely parallel those documented in previous studies, wherein lung metastases were detected 21 days post-cell injection. Moreover, aside from lung involvement, instances of brain metastases have been reported six weeks following inoculation74,76, contrasting with our findings, where we did not detect metastases in the brain. Noteworthy, we noted that the determination of a collection point varies according to the number of cells injected49,77,78,79and area of application50,51,74,80,81. In our study, we injected cells into the flank as done by50,51,82,83, and we used 1 × 106 cells as used by Gao et al. 201147. Our results add to those observed by other studies that employing the same methodology50,51,82,83 suggest that this quantity of cells or area of injection may not be efficient for studying the evolution of brain metastasis in 4T1. Nonetheless, further investigations are necessary to validate this assessment.
Furthermore, besides histopathology, we have also applied a tool using radioisotopes to monitor tumor progression and metastatic areas that have not yet been used in the 4T1 in vivo model. We employed a methodology using the HYNIC-βAla-Bombesin(7–14) peptide radiolabeled with 99mTc to track areas of metastasis, a methodology already used in other cancer models, such as prostate5,54, Ehrlich tumor84and MDA-MB-231 breast cancer model56but not yet reported in the 4T1 model. The diagnostic imaging tools for monitoring metastases in the 4T1 model have been used, mainly using bioluminescent 4T1 cells76,85,86 and with radiolabeling using 99mTc87,88. Our findings underscored the promising efficacy of this peptide, particularly in addressing liver metastases, as evidenced by the successful capture of radiation in scintigraphic images. Furthermore, the ex vivo biodistribution results were consistent with histopathological findings across all organs.
After standardizing of collection point of organs with metastasis, we assessed the expression of seven different genes commonly already evaluated in other breast cancer models, including GAPDH, 18 S, ACTB, B2M, HPRT1, GUSB and PGK139,60,69,70, and we also included three others described by bioinformatics tools as potential targets for use as HKGs (CCSER2, SYMPK and ANKRD17) in the 4T1 in vivo model42. Our results revealed a lack of a shared gene exhibiting consistently stable expression, which could serve as a suitable reference for data normalization across all tissues. This finding contrasts with observations from other studies involving diverse models24,89,90,91,92, including in cancer36,37,38. Therefore, we noted that the 4T1 model significantly impacts the expression of HKGs both within the primary tumor and in organs affected by metastasis.
In breast cancer, information about HKGs is controversial. While some authors suggest the use of commonly used HKGs for normalization, such as GAPDH39,60, 18 S70,93, ACTB40,59,70,93others have reported problems associated with normalizing data using these genes58,69. Indeed, in our work, we observed variability in 18s and ACTB expression between groups, as reported in MCF-7 and MDA-MB231 cells39,60and in samples of fibroadenoma extracted from humans94. Although our findings indicate that ACTB could serve as a secondary option for normalizing data in the tumor and liver, similar consistency was not observed when normalizing the expression of VEGF and TNF-α with ACTB, highlighting its inadequacy for these particular genes. This observation aligns with studies conducted on samples of human invasive carcinoma95, including samples positive for estrogen and progesterone receptors58,69.
In contrast to findings reported in other studies, our results indicate that GAPDH emerged as the optimal choice for data normalization, albeit not in organs exhibiting metastatic presence, such as the liver and lung. The utilization of GAPDH for data normalization has been previously documented in vitro models of triple-negative breast cancer cell lines, such as HCC1806, SUM149PT, HCC1937, and MDA-MB23139,70, and in cell lines mimicking luminal A breast cancer, such as MCF-739. Conversely, studies utilizing human tumor samples emphasize that GAPDH is not a suitable option for normalization, as demonstrated in both triple-negative tumors69and cases of fibroadenoma58,94. In our study utilizing the 4T1 in vivo model, we underscore its safe and effective use solely for studies on the primary tumor.
For the other genes, we observed variability in GUSB among groups across all analyzed organs, consistent with findings by57,58. Similarly, both B2M and PGK1 were ineffective in our study and others39,59,60. Despite the variability observed in HPRT1 expression in lung samples and primary tumors, our results indicate HPRT1 as an optimal choice for normalization in the liver. Previous studies have highlighted the ineffectiveness of HPRT1 in triple-negative cell lines70and human-derived samples57,59. Conversely, in fibroadenoma samples, the use of HPRT1 proved to be the best option94. Therefore, we emphasize HPRT1 as a dependable choice for data normalization in studies focusing on hepatic metastasis. Given that many studies have not explored metastasis across various organs, we believe this to be the first documentation of HPRT1 as a viable option for gene expression analyses in the liver affected by metastases from the 4T1 model. However, further investigations are warranted to evaluate the applicability of HPRT1 in assessing hepatic metastasis using alternative cell models.
Despite the descriptions of CCSER2, ANKRD17, and SYMPK as potential HKGs in breast cancer studies42, our data in the 4T1 experimental model showed that ANKRD17 and SYMPK were the genes with the less stable in all analyzed organs, while CCSER2 was optimal only in the lung. Interestingly, we found only one study evaluating the expression of CCSER2 in MCF-7 cells, which proved effective along with the use of GAPDH60. However, we did not uncover studies evaluating the utilization of ANKRD17 and SYMPK as HKGs, underscoring the necessity for further research to address the variation in existing models for assessing different types of breast cancer. We observed that the significant variability in results concerning HKGs in breast cancer models correlates with the phenotypic diversity of various breast cancer types and cell models38. We consider that the evaluation of HKGs should always be performed and reported according to the different experimental conditions, such as tests of new drugs and the use of different cell quantities, since cancer can influence the expression of different HKGs and compromise gene expression analyses37,46,96. Therefore, the need for future investigations that explore other specific knockout mouse models and relevant human cell lines is crucial to broaden the applicability of our findings. Based on our results, we hope to replicate these tests in other models not yet explored in the literature and validate them for use in studies with human cells. We hope that the results discussed here can provide a solid basis for planning future investigations.
Conclusion
This study revealed that GAPDH was the most stable expressed gene in the tumor, HPRT1 in the liver, and CCSER2 in the lung in the 4T1 in vivo model. Our results demonstrate that there is no standard HKG for use in all tissues, and the 4T1 model can affect the stability of the HKG most commonly used to normalize data in qPCR studies and the importance of carrying out prior validation of HKGs for experiments that use the qPCR technique. These findings underscore the critical importance of selecting appropriate HKGs for accurate normalization in gene expression studies. Using fewer stable genes for normalization can lead to erroneous interpretations of gene expression levels, potentially skewing results and conclusions. In contrast, utilizing stable HKGs enhances the reliability and validity of gene expression analyses, providing a more accurate depiction of biological phenomena. Thus, we strongly encourage using these specific genes in their respective tissues to normalize RT-qPCR data in studies involving the 4T1 model and metastases. Furthermore, we have determined the optimal time point for organ collection in metastasis studies using the 4T1 breast carcinoma model. Finally, when considering incorporating alternative genes, we recommend conducting comparative tests to assess their suitability directly, comparing the results with those from existing studies.
Data availability
All of the data generated or analyzed during this study are available in this manuscript and in supplementary materials section. Further details may obtained from the corresponding author upon reasonable request.
References
Sung, H. et al. Global Cancer statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Wu, Q. et al. Breast cancer subtypes predict the preferential site of distant metastases: a SEER based study. Oncotarget. 8, 27990–27996 (2017).
Schrörs, B. et al. Multi-omics characterization of the 4T1 murine mammary gland Tumor Model. Front. Oncol. 10, 1195 (2020).
Barros, A. C. S. D. et al. Induction of experimental mammary carcinogenesis in rats with 7,12-dimethylbenz(a)anthracene. Rev. Hosp. Clínicas. 59, 257–261 (2004).
Fuscaldi, L. L. et al. Feasibility of the 99mTc-HYNIC-βAla-Bombesin(7–14) for detection of LNCaP prostate tumour in experimental model. J. Radioanal Nucl. Chem. 305, 379–386 (2015).
Fridman, I. A. et al. Morphological Characteristic of Melanoma B16 Progression in C57BL/6 mice with high and low resistance to Hypoxia. Bull. Exp. Biol. Med. 168, 390–394 (2020).
Pulaski, B. A. & Ostrand-Rosenberg, S. Mouse 4T1 breast tumor model. Curr. Protoc. Immunol. Chapter 20, Unit 20.2 (2001).
Gupta, S. K., Tiwari, A. K., Gandham, R. K. & Sahoo, A. P. Combined administration of the apoptin gene and poly (I:C) induces potent anti-tumor immune response and inhibits growth of mouse mammary tumors. Int. Immunopharmacol. 35, 163–173 (2016).
Tiash, S., Chua, M. J. & Chowdhury, E. H. Knockdown of ROS1 gene sensitizes breast tumor growth to doxorubicin in a syngeneic mouse model. Int. J. Oncol. 48, 2359–2366 (2016).
Chiu, H. W. et al. Suberoylanilide Hydroxamic Acid, an inhibitor of histone deacetylase, enhances Radiosensitivity and suppresses lung metastasis in breast Cancer in Vitro and in vivo. PLoS ONE. 8, e76340 (2013).
Lindner, R. et al. Molecular phenotypes in triple negative breast cancer from African American patients suggest targets for therapy. PloS One. 8, e71915 (2013).
Anders, C. K., Carey, L. A. & Biology Metastatic patterns, and treatment of patients with triple-negative breast Cancer. Clin. Breast Cancer. 9, S73–S81 (2009).
Almansour, N. M. Triple-negative breast Cancer: a brief review about Epidemiology, Risk factors, signaling pathways, treatment and role of Artificial Intelligence. Front. Mol. Biosci. 9, 836417 (2022).
Peters, J. M. & Gonzalez, F. J. The evolution of carcinogenesis. Toxicol. Sci. Off J. Soc. Toxicol. 165, 272–276 (2018).
Aizawa, K. et al. Tobacco carcinogen induces both lung cancer and non-alcoholic steatohepatitis and hepatocellular carcinomas in ferrets which can be attenuated by lycopene supplementation. Int. J. Cancer. 139, 1171–1181 (2016).
Alison, M. R. The cellular origins of cancer with particular reference to the gastrointestinal tract. Int. J. Exp. Pathol. 101, 132–151 (2020).
Hahn, W. C. et al. An expanded universe of cancer targets. Cell. 184, 1142–1155 (2021).
Balkwill, F. R., Capasso, M. & Hagemann, T. The tumor microenvironment at a glance. J. Cell. Sci. 125, 5591–5596 (2012).
Wang, R. et al. Invasive cancers are not necessarily from preformed in situ tumours — an alternative way of carcinogenesis from misplaced stem cells. J. Cell. Mol. Med. 17, 921–926 (2013).
Kontomanolis, N. Basic principles of molecular biology of cancer cell-molecular cancer indicators. J. BUON Off J. Balk. Union Oncol. 26, 1723–1734 (2021).
Garcia-Crespo, D., Juste, R. A. & Hurtado, A. Selection of ovine housekeeping genes for normalisation by real-time RT-PCR; analysis of PrPgene expression and genetic susceptibility to scrapie. BMC Vet. Res. 1, 3 (2005).
Zaros, L. G. et al. Evaluation of reference genes for real-time PCR studies of Brazilian somalis sheep infected by gastrointestinal nematodes. Genet. Mol. Biol. 33, 486–490 (2010).
Janik, M. E. et al. RT-qPCR analysis of human melanoma progression-related genes – A novel workflow for selection and validation of candidate reference genes. Int. J. Biochem. Cell. Biol. 101, 12–18 (2018).
Souza, L. N. Analysis of the stability of the reference genes GAPDH, SDHA and RPL-19 in sheep from a semi-arid region infected by gastrointestinal nematodes. BMC Vet. Res. 19, 147 (2023).
Souza, J. L. N. et al. Evaluation of reference genes for gene expression analysis by real-time quantitative PCR (qPCR) in different tissues from mice infected by Ascaris suum. Microb. Pathog. 189, 106567 (2024).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper – Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515 (2004).
Gorji-Bahri, G., Moradtabrizi, N., Vakhshiteh, F. & Hashemi, A. Validation of common reference genes stability in exosomal mRNA‐isolated from liver and breast cancer cell lines. Cell. Biol. Int. 45, 1098–1110 (2021).
Senga, S. S. & Grose, R. P. Hallmarks of cancer—the new testament. Open. Biol. 11, 200358 (2021).
Hanahan, D. Hallmarks of Cancer: New dimensions. Cancer Discov. 12, 31–46 (2022).
De Jonge, H. J. M. et al. Evidence based selection of housekeeping genes. PLoS ONE. 2, e898 (2007).
Zhu, J., He, F., Hu, S. & Yu, J. On the nature of human housekeeping genes. Trends Genet. 24, 481–484 (2008).
Caradec, J. et al. Desperate house genes’: the dramatic example of hypoxia. Br. J. Cancer. 102, 1037–1043 (2010).
Kozera, B. & Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 54, 391–406 (2013).
Bjerregaard, H., Pedersen, S., Kristensen, S. R. & Marcussen, N. Reference genes for gene expression analysis by real-time reverse transcription polymerase chain reaction of renal cell carcinoma. Diagn. Mol. Pathol. 20, 212–217 (2011).
Ayakannu, T. et al. Validation of endogenous control reference genes for normalizing gene expression studies in endometrial carcinoma. Mol. Hum. Reprod. 21, 723–735 (2015).
Zhu, Y. et al. Identification of TMEM208 and PQLC2 as reference genes for normalizing mRNA expression in colorectal cancer treated with aspirin. Oncotarget. 8, 22759–22771 (2017).
Razavi, S. A. et al. Validation of reference genes for normalization of relative qRT-PCR studies in papillary thyroid carcinoma. Sci. Rep. 9, 15241 (2019).
Shah, K. N. & Faridi, J. S. Estrogen, tamoxifen, and akt modulate expression of putative housekeeping genes in breast cancer cells. J. Steroid Biochem. Mol. Biol. 125, 219–225 (2011).
Gorji-Bahri, G., Moradtabrizi, N. & Hashemi, A. Uncovering the stability status of the reputed reference genes in breast and hepatic cancer cell lines. PLOS ONE. 16, e0259669 (2021).
Majidzadeh-A, K., Esmaeili, R. & Abdoli, N. TFRC and ACTB as the best reference genes to quantify urokinase plasminogen activator in breast cancer. BMC Res. Notes. 4, 215 (2011).
Maltseva, D. V. et al. High-throughput identification of reference genes for research and clinical RT-qPCR analysis of breast cancer samples. J. Clin. Bioinforma. 3, 13 (2013).
Tilli, T. M., Castro, C. D. S. & Tuszynski, J. A. Carels, N. A strategy to identify housekeeping genes suitable for analysis in breast cancer diseases. BMC Genom. 17, 639 (2016).
Song, Q. et al. Public transcriptome database-based selection and validation of reliable reference genes for breast cancer research. Biomed. Eng. OnLine. 20, 124 (2021).
Chou, J. et al. GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat. Cell. Biol. 15, 201–213 (2013).
Kaplanov, I. et al. Blocking IL-1β reverses the immunosuppression in mouse breast cancer and synergizes with anti–PD-1 for tumor abrogation. Proc. Natl. Acad. Sci. 116, 1361–1369 (2019).
Hou, M. et al. Icariside I reduces breast cancer proliferation, apoptosis, invasion, and metastasis probably through inhibiting IL-6/STAT3 signaling pathway. J. Pharm. Pharmacol. rgad103https://doi.org/10.1093/jpp/rgad103 (2023).
Gao, Z. G., Tian, L., Hu, J., Park, I. S. & Bae, Y. H. Prevention of metastasis in a 4T1 murine breast cancer model by doxorubicin carried by folate conjugated pH sensitive polymeric micelles. J. Controlled Release. 152, 84–89 (2011).
Aslakson, C. J. & Miller, F. R. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 52, 1399–1405 (1992).
Kim, E. J. et al. Dietary fat increases solid tumor growth and metastasis of 4T1 murine mammary carcinoma cells and mortality in obesity-resistant BALB/c mice. Breast Cancer Res. 13, R78 (2011).
Franco, M. S. et al. Investigation of the antitumor activity and toxicity of long-circulating and fusogenic liposomes co-encapsulating paclitaxel and doxorubicin in a murine breast cancer animal model. Biomed. Pharmacother. 109, 1728–1739 (2019).
Dos Reis, D. C. et al. Versican and Tumor-Associated macrophages promotes Tumor Progression and Metastasis in Canine and murine models of breast carcinoma. Front. Oncol. 9, 577 (2019).
Vandermosten, L. et al. Experimental malaria-associated acute respiratory distress syndrome is dependent on the parasite-host combination and coincides with normocyte invasion. Malar. J. 17, 102 (2018).
Fulzele, S. V., Chatterjee, A., Shaik, M. S., Jackson, T. & Singh, M. Inhalation delivery and anti-tumor activity of Celecoxib in Human Orthotopic Non-small Cell Lung Cancer Xenograft Model. Pharm. Res. 23, 2094–2106 (2006).
Fuscaldi, L. L. et al. Evaluation of the optimal LNCaP prostate tumour developmental stage to be assessed by 99mTc-HYNIC-βAla-Bombesin(7–14) in an experimental model. J. Radioanal Nucl. Chem. 300, 801–807 (2014).
Carlesso, F. N. et al. Evaluation of 99mTc-HYNIC-βAla-Bombesin(7–14) as an agent for pancreas tumor detection in mice. Braz J. Med. Biol. Res. 48, 923–928 (2015).
De Barros, A. L. B., Mota, L. D. G., Ferreira, C. D. A. & Cardoso, V. N. Kit formulation for 99mTc-labeling of HYNIC-βAla-Bombesin(7–14). Appl. Radiat. Isot. 70, 2440–2445 (2012).
McNeill, R. E., Miller, N. & Kerin, M. J. Evaluation and validation of candidate endogenous control genes for real-time quantitative PCR studies of breast cancer. BMC Mol. Biol. 8, 107 (2007).
Lyng, M. B., Laenkholm, A. V., Pallisgaard, N. & Ditzel, H. J. Identification of genes for normalization of real-time RT-PCR data in breast carcinomas. BMC Cancer. 8, 20 (2008).
Gur-Dedeoglu, B. et al. Identification of endogenous reference genes for qRT-PCR analysis in normal matched breast tumor tissues. Oncol. Res. 17, 353–365 (2009).
Jain, N., Nitisa, D., Pirsko, V. & Cakstina, I. Selecting suitable reference genes for qPCR normalization: a comprehensive analysis in MCF-7 breast cancer cell line. BMC Mol. Cell. Biol. 21, 68 (2020).
Ruiz-Manzano, R. A. et al. Potential novel risk factor for breast Cancer: Toxocara canis infection increases tumor size due to modulation of the Tumor Immune Microenvironment. Front. Oncol. 10, 736 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2 – ∆∆CT method. Methods. 25, 402–408 (2001).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, 45e–445 (2001).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, research00341 (2002).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse Transcription-PCR data: a model-based Variance Estimation Approach to identify genes suited for normalization, Applied to bladder and Colon Cancer Data sets. Cancer Res. 64, 5245–5250 (2004).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant. Mol. Biol. 80, 75–84 (2012).
Xie, F., Wang, J. & Zhang, B. RefFinder: a web-based tool for comprehensively analyzing and identifying reference genes. Funct. Integr. Genomics. 23, 125 (2023).
Jo, J. et al. Conventionally used reference genes are not outstanding for normalization of gene expression in human cancer research. BMC Bioinform. 20, 245 (2019).
Liu, L. L. et al. Identification of valid reference genes for the normalization of RT-qPCR expression studies in human breast Cancer cell lines treated with and without transient transfection. PLOS ONE. 10, e0117058 (2015).
Pulaski, B. A. & Ostrand-Rosenberg, S. Mouse 4T1 breast tumor model. Curr. Protoc. Immunol. 39, (2000).
DuPre´, S. A., Redelman, D. & Hunter, K. W. The mouse mammary carcinoma 4T1: characterization of the cellular landscape of primary tumours and metastatic tumour foci. Int. J. Exp. Pathol. 88, 351–360 (2007).
Atiya, H. I. et al. Intraductal adaptation of the 4T1 mouse model of breast Cancer reveals effects of the epithelial microenvironment on Tumor Progression and Metastasis. Anticancer Res. 39, 2277–2287 (2019).
Bailey-Downs, L. C. et al. Development and characterization of a preclinical model of breast Cancer Lung Micrometastatic to Macrometastatic Progression. PLoS ONE. 9, e98624 (2014).
Wenzel, J., Zeisig, R. & Fichtner, I. Inhibition of metastasis in a murine 4T1 breast cancer model by liposomes preventing tumor cell-platelet interactions. Clin. Exp. Metastasis. 27, 25–34 (2010).
Tao, K., Fang, M., Alroy, J. & Sahagian, G. G. Imagable 4T1 model for the study of late stage breast cancer. BMC Cancer. 8, 228 (2008).
Pulaski, B. A. & Ostrand-Rosenberg, S. Reduction of established spontaneous mammary carcinoma metastases following immunotherapy with major histocompatibility complex class II and B7.1 cell-based tumor vaccines. Cancer Res. 58, 1486–1493 (1998).
Muraoka, R. S. et al. Blockade of TGF-β inhibits mammary tumor cell viability, migration, and metastases. J. Clin. Invest. 109, 1551–1559 (2002).
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 28, 341–347 (2010).
Tabariès, S. et al. Granulocytic immune infiltrates are essential for the efficient formation of breast cancer liver metastases. Breast Cancer Res. 17, 45 (2015).
Kocatürk, B. & Versteeg, H. H. Orthotopic injection of breast Cancer cells into the Mammary Fat Pad of mice to study Tumor Growth. J. Vis. Exp. 51967. https://doi.org/10.3791/51967 (2015).
Reis, D. C. et al. Evaluation of the immunomodulatory activity of thalidomide on tumor-associated macrophages in the 4T1 murine metastatic breast cancer model. Arq. Bras. Med. Veterinária E Zootec. 73, 1334–1345 (2021).
Abdel-Salam, M. A. L. et al. LyeTx I-b peptide attenuates Tumor Burden and Metastasis in a mouse 4T1 breast Cancer Model. Antibiotics. 10, 1136 (2021).
De Barros, A. L. B. et al. Bombesin derivative radiolabeled with technetium-99m as agent for tumor identification. Bioorg. Med. Chem. Lett. 20, 6182–6184 (2010).
Hiraga, T., Williams, P. J., Ueda, A., Tamura, D. & Yoneda, T. Zoledronic acid inhibits visceral metastases in the 4T1/luc mouse breast Cancer Model. Clin. Cancer Res. 10, 4559–4567 (2004).
Smith, M. C. P. et al. CXCR4 regulates growth of both primary and metastatic breast Cancer. Cancer Res. 64, 8604–8612 (2004).
Tesan, F. C. et al. 99m Tc-Radiolabeled TPGS Nanomicelles Outperform 99m Tc-Sestamibi as Breast Cancer Imaging Agent. Contrast Media Mol. Imaging 1–9 (2019).
Ekinci, M., Alencar, L. M. R., Lopes, A. M., Santos-Oliveira, R. & İlem-Özdemir, D. Radiolabeled Human serum albumin nanoparticles co-loaded with methotrexate and decorated with trastuzumab for breast Cancer diagnosis. J. Funct. Biomater. 14, 477 (2023).
Stephens, A. S., Stephens, S. R. & Morrison, N. A. Internal control genes for quantitative RT-PCR expression analysis in mouse osteoblasts, osteoclasts and macrophages. BMC Res. Notes. 4, 410 (2011).
Ma, J. et al. Comparison of reference gene expression stability in mouse skeletal muscle via five algorithms. PeerJ. 10, e14221 (2022).
Alieva, A., Kh., Filatova, E. V., Rudenok, M. M. & Slominsky, P. A. Shadrina, M. I. Housekeeping genes for Parkinson’s disease in humans and mice. Cells. 10, 2252 (2021).
Quiñonez-Flores, C. M. et al. Stability of housekeeping genes in inflamed joints of spontaneous and collagen-induced arthritis in DBA/1 mice. Inflamm. Res. 70, 619–632 (2021).
Morse, D. L. et al. Determining suitable internal standards for mRNA quantification of increasing cancer progression in human breast cells by real-time reverse transcriptase polymerase chain reaction. Anal. Biochem. 342, 69–77 (2005).
De Kok, J. B. et al. Normalization of gene expression measurements in tumor tissues: comparison of 13 endogenous control genes. Lab. Invest. 85, 154–159 (2005).
Kılıç, Y., Çelebiler, A. Ç. & Sakızlı, M. Selecting housekeeping genes as references for the normalization of quantitative PCR data in breast cancer. Clin. Transl Oncol. Off Publ Fed. Span. Oncol. Soc. Natl. Cancer Inst. Mex. 16, 184–190 (2014).
Zhao, H. et al. Identification of valid reference genes for mRNA and microRNA normalisation in prostate cancer cell lines. Sci. Rep. 8, 1949 (2018).
Acknowledgements
JLNS is grateful for the PhD fellowship provide by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Post-graduation Program in Parasitology, UFMG.
Funding
This research was financially supported by Pró-reitoria de pesquisa of UFMG, Fundação de Amparo à Pesquisa do Estado de Minas Gerais/FAPEMIG, Brazil (Grant #APQ-4035/17), Rede Mineira de Imunobiológicos (RED-00067-23), and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) Demanda Universal (Grant # 403278/2023-6). RTF (Grant #305514/2022-9), RCR (Grant # 313839/2023-9) and LLB (#310311/2023-3) are research fellows of the CNPq. SOAF, VNC, GDC and RCR are supported by a grant from the FAPEMIG (Rede Mineira de Pesquisa Translacional em Imunobiológicos e Biofármacos no Câncer – REMITRIBIC, RED-00031-21).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: JLNS, RCR, LLB. Performed the experiments: JLNS, ARAP, ISO, CCOA, LOP, IBD, LVBA, EAO, SOAF. Analyzed the data: JLNS, ARAP, LOP, SOAF, FRS, GDC, VNC, RCR. Sample collection: JLNS, ARAP, ISO, CCOA, LOP, IBD, LVBA, EAO, SOAF. Funding acquisition: RTF, RCR and LLB. Supervision: RTF, RCR and LLB. Contributed reagents/materials/analysis tool: GDC, VNC, SOAF, RTF, RCR and LLB. Writing original paper draft: JLNS, ARAP, RCR, and LLB. All other authors revised the data and discussed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The maintenance of animals and all procedures performed during the experiments were conducted according to the Brazilian College of Animal Experimentation (COBEA) guidelines and approved by the local Ethics Committee forAnimal Experimentation (CEUA) of the Universidade Federal de Minas Gerais, Brazil, under protocol number 178/2021. All efforts were made to minimize animal suffering and all methods were performed in accordance with the ARRIVE guidelines (https://arriveguidelines.org).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Souza, J.L.N., Antunes-Porto, A.R., da Silva Oliveira, I. et al. Screening and validating the optimal panel of housekeeping genes for 4T1 breast carcinoma and metastasis studies in mice. Sci Rep 14, 26476 (2024). https://doi.org/10.1038/s41598-024-77126-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-77126-x