Abstract
Valproate (VPA), a drug used to treat neurological disorders such as epilepsy, bipolar disorder, and migraines, induces inositol depletion in the brain by preventing its synthesis, which is a hypothesized therapeutic mechanism for mood stabilizing drugs. However, the mechanism by which this occurs is not known. VPA treatment reduces activity of the enzyme that catalyzes the rate-limiting step of inositol synthesis, myo-inositol-3-phosphate synthase (MIPS). Utilizing the yeast model, we report that VPA induces repression of the MIPS-encoding gene INO1 by increasing nuclear translocation of the transcriptional repressor Opi1. In addition, VPA rapidly inhibits TORC1, an activator of INO1 expression, and increased TORC1 activity partially rescues expression of INO1. VPA also rapidly activates Snf1, a known regulator of INO1. However, neither TORC1 inhibition nor Snf1 activation is required for repression of INO1 by VPA. We postulate that it is likely that the pleiotropic effects of VPA not only bring about the initial decrease in MIPS protein levels by repressing INO1 transcription but also by preventing recovery of inositol synthesis by downregulating MIPS translation. These independent effects of VPA underlie the complex mechanisms that collectively induce inositol depletion.
Similar content being viewed by others
Introduction
Inositol is an essential sugar alcohol required for eukaryotic cell life. It is a key component of numerous signaling molecules. Inositol-containing phospholipids phosphatidylinositol (PI) and phosphatidylinositol phosphates (PIPs) are distributed throughout the intracellular-facing plasma membrane and organelles and serve as hubs for coordinating protein interactions and signal transduction (reviewed in1,2). Inositol phosphates (IPs) serve as essential cofactors in enzymatic reactions or signaling cascades (reviewed in3,4,5,6,7). For example, inositol 1,4,5‐trisphosphate (IP3) mediated Ca2+ release from the endoplasmic reticulum (ER) (reviewed in8,9) has various cell dependent roles, such as inducing smooth and cardiac muscle contraction as well as neuronal depolarization (reviewed in9,10,11,12). IPs give rise to yet another class of inositol compounds, inositol pyrophosphates (IPPs), which are distinguished by their highly energetic diphosphate moieties and are indispensable for diverse cellular functions (reviewed in3,5,7,13,14,15,16,17. At the foundation of these essential signaling compounds and cofactors is the cyclical sugar alcohol, inositol. While many inositol stereoisomers play their own distinctive roles in signaling and human diseases (reviewed in4,16,18,19), myo-inositol (referred to as inositol in this article) is the most abundant isoform, levels of which are tightly regulated.
Inositol and IP6 (phytic acid) are both available in our diets. Cells actively take up inositol from blood plasma via inositol transporters (reviewed in20). Inositol levels are maintained at particularly high levels (mM range) in the brain, only half of which is supplied by blood plasma (reviewed in19); thus, inositol synthesis is necessary to maintain neuronal cell functions. De novo synthesis requires two enzymatic steps: The rate-limiting reaction is the conversion of the glycolytic precursor glucose 6-phosphate to inositol 3-phosphate, catalyzed by myo-inositol 3-phosphate synthase (MIPS); the second step is the dephosphorylation of inositol 3-phosphate by inositol monophosphatases (IMPases) to produce inositol (reviewed in21,22). Interestingly, there is accumulating evidence that inositol synthesis plays a role in tumorigenesis, as transcriptional regulation of these synthesis genes can markedly affect the growth of cancer cells (reviewed in23,24), and both IP6 and inositol may elicit beneficial outcomes for cancer and metabolic disorder treatments4,16,25,26. Thus, it is remarkable that little is known about the cellular mechanisms regulating MIPS and IMPases in mammalian cells (see23 for a summary). Conversely, the regulatory mechanisms of inositol synthesis have been studied for decades in budding yeast. Transcription of yeast MIPS is highly regulated by the Henry regulatory circuit (Ino2-Ino4-Opi1) (reviewed in19 and modeled in Fig. 1A), but the degree of regulation appears less robust in mammalian cells23. Due to the requirement of inositol-derived compounds for cellular function and the particular need to synthesize inositol in neurons (and some cancers), there is merit in treating some diseases by targeting the inositol synthesis pathway.
VPA and TORC1 inhibition reduced INO1 and MIPS induction during the mid-log growth phase. (A) Model of the Henry regulatory circuit during exponential growth in inositol-free medium, medium containing exogenous inositol, and during the diauxic shift/stationary phase. Abbreviations: phosphatidic acid (PA), cytidine diphosphate diacylglycerol (CDP-DAG), inositol (I), phosphatidyl inositol (PI), diacylglycerol (DAG), triacylglycerol (TAG), endoplasmic reticulum (ER), and lipid droplet (LD) (B) Experimental diagram of yeast culture growth before and after treatments. Yeast cells were switched to inositol free medium (I + → I −) just before treatments of VPA (1 mM), rapamycin (400 nM), or vehicles (control) were administered at the early-log growth phase. Samples were collected over an 8 h period. (C) INO1 mRNA and (D) MIPS protein expression assayed by RT-qPCR and SDS-PAGE/WB analyses mRNA expression data is shown relative to 0 h control. Protein levels are reported relative to 4 h control, as this is when we start to see stable levels and no protein was detected at 0 h. Total protein normalization (TPN) was used to normalize WB quantifications. n = 2 biological replicates; error bars are SEM. See Supplementary figure S1 for INO1 and ACT1 transformed ∆Cq data.
Bipolar disorder is a psychiatric illness with an estimated incidence rate of 2.8% among adults in the US; most cases are classified as severe with a high degree of impairment27. While the pathophysiology leading to bipolar disorder is unknown, there is evidence suggesting that underlying neuronal Ca2+ signaling may be aberrant (reviewed in19,28). Although bipolar disorder is a lifelong illness, the symptoms can be managed in part by pharmaceutical mood stabilizers. Lithium was the first treatment for bipolar disorder approved by the FDA in 197029,30. Berridge and colleagues postulated that inositol depletion may be the therapeutic mechanism of mood stabilizers based on findings showing that IMPase is directly inhibited by lithium, leading to lower inositol levels in the brain31. Decreased brain inositol levels was proposed to mitigate the downstream signaling pathways dependent on inositol derived compounds, such as IP3 mediated Ca2+ release from the ER28. Later, the FDA approved the anti-seizure medication valproate (VPA) as well as the antipsychotic drug carbamazepine for the treatment of manic episodes of bipolar disorder (reviewed in29), both of which also reduce intracellular inositol levels (reviewed in19,28). VPA decreases de novo inositol synthesis most likely by reducing MIPS protein levels32,33; however, this mechanism remains to be clarified. The mechanism by which inositol is decreased by carbamazepine is still undetermined, but some evidence suggests it may reduce uptake from the extracellular environment34,35. That each of these structurally dissimilar drugs reduces inositol concentration further supports inositol depletion as an underlying therapeutic mechanism19. Mood stabilizers are not without their side effects and are not universally effective at mitigating patient symptoms. Treating bipolar disorder is especially complicated as both the underlying cause(s) of the illness and therapeutic mechanism(s) of action of the drugs are unknown. Elucidating the biochemical mechanisms whereby mood stabilizers deplete inositol can help clarify both mysteries.
VPA was originally approved for treating epileptic seizures, but its utility has expanded to include mood stabilization and migraine prevention (reviewed in36,37,38). Moreover, it has been under clinical trials as a chemo therapeutic modifier due to its suppressive effects on tumor growth (reviewed in37,39), a neuroprotective agent for nephropathy in diabetes40, as well as a potential type 2 diabetes treatment (reviewed in37,41,42). The variety of conditions treated by VPA may be explained by the pleiotropic and cell-dependent effects of the drug.
There are many hypothesized therapeutic targets of VPA. Histone deacetylase (HDAC) inhibitory properties of VPA have been well-characterized (reviewed in37,38). HDAC inhibition can alter epigenetic regulation and chromatin remodeling, modifying transcription of many genes and, thus, propagating numerous downstream effects. The potent HDAC inhibitory properties of VPA suggested that it may be a viable anticancer drug and offered a likely explanation for the teratogenicity of VPA43,44. Due to the pleiotropic effects of HDAC inhibition, it is tempting to attribute many of the other effects of VPA to this one non-specific activity (i.e., assume HDAC inhibition is the primary target and other documented effects of VPA are secondary/downstream). However, HDAC inhibition dependent vs independent effects of VPA have not been entirely addressed. The anti-epileptic as well as anti-migraine activities are attributed to inhibition of neuronal voltage-gated ion channels, and/or to increasing GABA levels, an inhibitory transmitter of the CNS (due to increasing synthesis and reducing degradation) (reviewed in37,38,45,46,47,48). Proposed therapeutic mechanisms of bipolar disorder include these same effects, but also incorporate inositol depletion19,28, which is likely mediated by GSK3 inhibition19,49. More recently, it has been proposed that AMPK50 and mTOR51, two antagonistic master metabolic regulators, may also be targets for mood stabilization effects of VPA (discussed below). Finally, VPA has been reported to reduce blood glucose concentrations in patients, leading to proposals to expand its utility to manage diabetes mellitus (the mechanisms leading to these effects remain speculative)41,42. These highlighted outcomes of VPA treatment are only a fraction of those reported (see48 for a more complete review). To determine the specific therapeutic targets for these illnesses will require consolidating these outcomes of VPA treatment into cause-and-effect pathways.
Activities of AMPK and TORC1 are altered in response to VPA treatment (discussed below) and both are documented to regulate inositol synthesis33,52,53,54. AMPK and mTOR are master metabolic regulators and have significant influence on many cellular functions. AMPK is activated under stress or nutrient limiting conditions, triggering transcriptional remodeling and altering enzymatic activities of many proteins involved in metabolism (reviewed in55,56,57,58). In general, AMPK activation increases catabolic processes (such as autophagy), reducing cell growth but promoting cell survival. The antagonistic mTOR complexes do the opposite, as activation increases anabolic pathways, promoting cell growth and proliferation (reviewed in55,56,57,59,60,61). The mTOR protein kinase is found among two functionally distinct complexes, mTOR complex 1 (mTORC1) and mTORC2, which localize at different areas of the cell and respond to different stimuli. Transcriptional control by mTOR extends to thousands of genes (> 2/3 of the genome for yeast TORC153). Therefore, in addition to the numerous effects downstream of HDAC inhibition, AMPK and mTOR potentially mediate many of the outcomes of VPA treatment.
While AMPK (Snf1 in yeast) activity is activated by VPA in both mammalian and yeast models33,54,62,63,64,65, the same consensus cannot be reached regarding mTOR activity; some studies report that VPA inhibits mTOR while others describe activation of mTOR by the drug (discussed further below). One of the main differences between these contradictory reports is the types of model organisms used in the experiments. In particular, VPA treatment of cancer cells decreased mTOR activity, whereas activity was increased in non-cancer models (Table 1). Typically, cancer cells undergo aerobic glycolysis (aka fermentative growth, or the Warburg effect) where mTOR activity is high and AMPK activity is low. This metabolic switch facilitates rapid growth and proliferation by producing energy and cellular building blocks. Non-cancerous cells rely more on oxidative phosphorylation to produce energy. This switch from oxidative phosphorylation to aerobic glycolysis is, in part, controlled by AMPK and mTOR activity levels. Thus, the metabolic state of these different models may determine how VPA affects mTOR activity. Yeast cells prefer fermentative over oxidative growth, undergoing aerobic glycolysis like most cancers. Yet, unlike cancer cells, yeast maintain metabolic flexibility, i.e., the ability to downregulate glycolysis and switch to oxidative phosphorylation. Thus, yeast cells are an excellent model organism for discerning the specific effects of VPA on TOR activity.
Yeast is also an ideal model in which to characterize the regulatory role of TORC1 on MIPS expression, as transcriptional regulation of MIPS is well defined. One study concluded that TORC1 increases induction of INO1, the gene encoding MIPS in yeast52. In addition, INO1 mRNA levels were the most downregulated relative to all other genes in a microarray dataset of rapamycin-treated yeast53 (rapamycin specifically inhibits TORC1). Lastly, activation of the yeast AMPK homolog (Snf1) is a known regulator of INO1 expression (reviewed in19), and VPA activates Snf154. Therefore, we hypothesized that VPA-induced inositol depletion is mediated by Snf1 activation and/or TORC1 inhibition. Testing this hypothesis in the current study, we established the following: 1) VPA reduces INO1 and MIPS expression via the Henry regulatory circuit by increasing nuclear localization of Opi1 during fermentative growth, but not during the diauxic shift (which is characterized by reduced glycolysis and increased oxidative phosphorylation). 2) VPA rapidly activates Snf1 and inhibits TORC1 but does not affect TORC2 activity under therapeutically relevant concentrations. 3) Increased TORC1 activity can partially rescue VPA-induced repression of INO1 but repression of INO1 by VPA is not dependent on TORC1 inhibition or Snf1 activation. These findings indicate that the primary mechanism leading to reduced MIPS activity centers on transcriptional down-regulation of INO1 by Opi1 nuclear translocation and preventing recovery of MIPS expression. While increased TORC1 activity can compensate for reduced MIPS expression in the presence of VPA, VPA-induced inhibition of MIPS is not dependent on Snf1 or TORC1. [These data were originally published as part of a PhD dissertation80].
Results
VPA reduces INO1 expression by increasing nuclear localization of Opi1
VPA reduces INO1 induction during the mid-log growth phase
As yeast cells are introduced into fresh glucose-rich medium, they undergo a dynamic metabolic remodeling, transitioning from low to high glycolytic activity (known as the logarithmic/exponential growth phase). As they deplete glucose from the medium, they switch to a more nutritionally conservative metabolism, oxidative phosphorylation (known as the diauxic shift). Utilizing this model organism, we characterized how VPA affects INO1 mRNA and MIPS protein expression, specifically addressing the effects at different growth stages and during conditions conducive to INO1 induction or repression. When yeast cells are grown in inositol-free (I-) medium, INO1 is transcriptionally activated by the Henry regulatory circuit. The transcriptional activators Ino2-Ino4 form a dimer and bind to the upstream activator sequence (UASINO), inducing INO1 expression19. Opi1 is the transcriptional repressor of INO1 and is recruited to the UASINO element by interacting with Ino2. Opi1 is regulated by its localization, either in the nucleus (where it represses INO1 transcription) or the ER (enabling INO1 transcription). Within the nucleus, Opi1 recruits corepressors (such as Sin3, Ssn6-Tup1, and Mot1), which in turn induce HDACs (such as Rpd3 and Hda1) to transcriptionally silence expression (see19 for a review on the Henry regulatory circuit mechanism). The determinate factor directing its localization is the level of phosphatidic acid (PA) in the ER. When PA levels are high (e.g., during the log-growth phase in I- medium), Opi1 is anchored in the ER by an interaction with PA as well as the ER transmembrane protein Scs281. When PA levels decrease (e.g., when cells are grown in the presence of excess inositol, or during the diauxic shift/stationary phase), Opi1 is released from the ER and translocates to the nucleus where it represses UASINO-containing genes, including INO1 (Fig. 1A). When growing yeast in I + medium, INO1 transcription is repressed and MIPS protein is undetectable. Upon switching the yeast cells to I- medium, inositol levels decrease and INO1 transcription is highly induced.
To understand the effects of VPA on INO1 induction, cultures in the early-log growth phase were treated following the switch from I + to I- medium (Fig. 1B). Samples of the cultures were collected every 2 h (up to 8 h) post-induction and INO1 mRNA and MIPS protein levels assayed by RT-qPCR and western blot (WB) analyses (Fig. 1C,D). For untreated control samples, INO1 mRNA increased greatly within 2 h and fell to moderate levels after 4–6 h. After 8 h, control cultures were entering their diauxic shift and INO1 mRNA dropped to levels similar to those observed at 0 h (Fig. 1C). MIPS protein levels rose following INO1 mRNA expression; MIPS levels were low after 2 h but elevated and plateaued after 4 h (Fig. 1D). MIPS protein levels did not decrease after 8 h, although INO1 mRNA expression did. VPA treatment reduced induction of INO1 transcription, particularly diminishing the initial increase observed after 2 h in control samples (Fig. 1C). However, mRNA levels were similar to those of the control after 4–6 h and did not decrease after 8 h (Fig. 1B). There are at least two possible explanations as to why INO1 mRNA levels did not decrease in VPA treated samples after 8 h. First, growth-rate is reduced by VPA, so VPA-treated cells were still in the log growth phase (during which INO1 is transcriptionally active) while control cultures had entered the diauxic shift (in which INO1 is transcriptionally repressed). This is likely as INO1 mRNA levels decreased when VPA treated cultures entered the diauxic shift (Fig. 4). Alternatively, the initial decrease in INO1 mRNA may be followed by a compensatory response due to inositol depletion increasing INO1 transcriptional activation (Figs. 2 and 3). Nonetheless, MIPS protein levels were consistently lower with VPA treatment throughout the duration of the experiment (Fig. 1D), indicating that this altered induction profile results in decreased MIPS protein levels.
VPA reduces INO1 mRNA and MIPS expression during the mid-log growth phase, but rapamycin does not. (A) Experimental diagram of yeast culture growth before and after treatments. Unlike the experiment shown in Fig. 1A, yeast cells were inoculated in inositol-free medium and incubated until the early-log growth phase when treatments of VPA (1 mM), rapamycin (400 nM), or vehicles (control) were administered. Under these conditions, INO1 mRNA and MIPS protein are expressed prior to treatments. Samples were collected over a 4 h period and (B) INO1 mRNA and (C) MIPS protein expression levels assayed by RT-qPCR and SDS-PAGE/WB analyses. Total protein normalization (TPN) was used to normalize WB quantifications. Data is shown relative to 0 h control. n = 4 biological replicates; error bars are SEM; two-way ANOVA * = p < 0.05, **** = p < 0.0001 compared to control at same time point. As ACT1 mRNA was validated as an appropriate normalization reference gene for cells in the mid-log growth phase (Supplementary figure S1C), the transformed ∆Cq data is not shown.
VPA-induced repression of INO1 is mediated by the transcriptional repressor Opi1. (A) WT and opi1∆ were inoculated in inositol-free medium (I-) and incubated until the log growth phase when VPA and/or inositol (I + ; 75 µM) were administered for 2 h. INO1 mRNA was assayed by RT-qPCR analyses. n = 3 biological replicates; error bars are SEM; two-way ANOVA * = p < 0.05, **** = p < 0.0001 compared to WT control I-. (B) A strain endogenously expressing OPI1-mGFP was cultivated to the mid-log growth phase and split into three parts. Each portion was treated with either inositol (75 µM), VPA (1 mM), or vehicle (control). Live cells were then stained with DAPI (5 µg/mL) for 30 min prior to continuously monitoring Opi1-mGFP localization by confocal microscopy imaging up to 2 h after treatment additions. Scale bar in top left most figure represents 2.5 µm.
VPA increases nuclear localization of the Opi1 transcriptional repressor during the mid-log growth phase
To determine if VPA treatment reduces INO1 mRNA and MIPS expression when levels are elevated, yeast cells were grown in I- medium at the onset of inoculation (Fig. 2A). Thus, expression is maintained at a high, steady level prior to administering treatments during the early-log growth phase. VPA induced a significant decrease in INO1 mRNA levels after 2 h (Fig. 2B) and a substantial decrease in MIPS protein after 2–4 h (Fig. 2C). After 4 h of VPA treatment, INO1 expression was elevated to levels similar to those in control samples (Fig. 2B), recapitulating results shown previously32. This increase in INO1 mRNA is likely a response to inositol depletion, which stimulates the Henry regulatory circuit to increase INO1 induction. However, MIPS levels did not follow the same pattern, showing a dissociation of INO1 mRNA and MIPS protein levels with VPA treatment longer than 4 h (similar to the results in Fig. 1C,D), or even after 6 h as previously shown32.
The magnitude of reduced INO1 mRNA levels brought on by VPA treatment suggests that this is facilitated by the INO1 transcriptional repressor, Opi1. To test this, we treated WT and opi1∆ strains with VPA in the presence or absence of exogenous inositol for 2 h and assayed INO1 mRNA levels (Fig. 3A). Exogenous inositol is known to induce nuclear localization of Opi1 and repression of INO1. As expected, exogenous inositol repressed INO1 mRNA expression in WT but not in opi1∆ cells. Interestingly, VPA-induced repression of INO1 mRNA did not occur in opi1∆, suggesting that Opi1 localization to the nucleus and repression of INO1 is required for VPA-induced repression of INO1. A subsequent experiment monitored Opi1-mGFP localization continuously over the course of 2 h following either the addition of exogenous inositol or VPA treatment (Fig. 3B). As expected, Opi1-mGFP remained localized perinuclear in cells grown in I- medium whereas exogenous inositol induced nuclear localization. VPA treatment (in I- medium) altered Opi1-mGFP localization. After 30 min of VPA treatment, Opi1-mGFP was localized both within the nucleus and perinuclear. After an additional 30–45 min (60–75 min in total), the perinuclear ring of mGFP was lost and mGFP mostly co-localized with DAPI in the nucleus (similar to the I + sample). At later times, the Opi1-mGFP signal gradually returned to perinuclear localization, although some remained within the nucleus after 2 h. Thus, VPA reduces INO1 expression by inducing nuclear localization of the Opi1 transcriptional repressor.
VPA does not reduce INO1 expression during the diauxic shift
To determine if VPA affects INO1 mRNA and MIPS expression during the diauxic shift/stationary growth phase, cultures were cultivated to the late-log growth phase and then treated with VPA for up to 24 h (Fig. 4A). VPA-induced repression of INO1 and MIPS levels was less prominent during the diauxic shift (Fig. 4B,C) than that observed in log phase (Fig. 2B,C). Although some INO1 repression occurred in VPA-treated cells during the late-log growth phase, specifically after 1 h, (Fig. 4B), expression was not significantly different from that of untreated cells as cultures transitioned into the diauxic shift (Fig. 4B,C), suggesting that VPA does not affect INO1/MIPS expression levels during the diauxic shift/stationary growth phase.
VPA and rapamycin do not repress INO1 mRNA and MIPS expression during the diauxic shift/stationary phase. (A) Experimental diagram of yeast culture growth before and after treatments. Yeast cells were inoculated in inositol-free medium and incubated until the late-log growth phase when treatments of VPA (1 mM), rapamycin (400 nM), or vehicles (control) were administered. Samples were collected over a 24 h period. (B) INO1 mRNA expression levels, (C) MIPS protein levels, and (D) PP-Rps6 levels (a TORC1 activity biomarker) were assayed by RT-qPCR and SDS-PAGE/WB analyses. mRNA data is shown relative to 24 h control. WB data is shown relative to 0 h control. Total protein normalization (TPN) was used to normalize WB quantifications. n = 2 biological replicates; error bars are SEM.
In summary, VPA reduces INO1 induction and leads to repression of INO1 by increasing nuclear localization of the transcriptional repressor Opi1 during the mid-log growth phase. VPA does not significantly alter expression of INO1 during the diauxic shift/stationary growth phase, when Opi1 is primarily localized to the nucleus.
The regulatory effects of TORC1 on INO1 expression
VPA rapidly inhibits TORC1
As a previous study suggested that TORC1 increases INO1 induction52, we tested the hypothesis that VPA represses INO1 induction by inhibiting TORC1. To do so, we utilized phosphorylation of Rps6 at S232/S233 as detected by antibody PP-Rps6 as a biomarker of TORC1 activity. Phosphorylation at S233 is dependent on TORC1 activity82, whereas S232 phosphorylation is dependent on either TORC1 or TORC2 activity83 (see diagram in Fig. 6B). Antibody PP-Rps6 binds to the target only if both sites are phosphorylated. Therefore, PP-Rps6 antibody measures the activity of TORC1. VPA rapidly inhibited TORC1 in exponentially growing cells, showing a large and statistically significant decrease in PP-Rps6 after 15 min of treatment and a maximum inhibitory effect after 1 h (Fig. 5A). Interestingly, TORC1 activity slightly increased after 3–5 h of sustained treatment but did not return to levels observed in untreated control samples. Inhibition of TORC1 is dose-dependent, with half-maximum inhibition occurring with 0.2 mM VPA and near complete inhibition with 0.6 mM (Fig. 5B). These concentrations are within the effective therapeutic range found in patient cerebral spinal fluid84 and plasma samples85 (0.28–0.98 mM and ≈0.48 mM, respectively). Exogenous inositol did not prevent TORC1 inhibition, suggesting that VPA-induced inhibition of TORC1 is independent of inositol depletion (Fig. 5B).
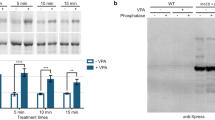
VPA rapidly inhibits TORC1 in a concentration dependent manner. (A) Yeast in the mid-log growth phase were treated with VPA (1 mM), rapamycin (400 nM), or vehicles (control) and cell lysates collected after indicated treatment durations. SDS-PAGE and WB analyses were used to assay TORC1 activity (PP-Rps6) and Snf1 activation (P-Snf1; band indicated with an arrow). WB quantifications are shown below with n = 3 biological replicates. (B) Yeast in the mid-log growth phase were treated with a concentration gradient of VPA (0, 0.1, 0.2, 0.4, 0.6, 0.8, and 1.0 mM). Yeast cultures were also treated with exogenous inositol (I + ; 75 µM) in addition to treating with VPA (V; 1 mM) or vehicle (control; C). Treatment duration lasted 1 h then TORC1 activity was assayed as before. WB quantifications are shown to the right. n = 3 biological replicates for the concentration gradient curve or n = 2 for I + treated cultures. Error bars are SEM and statistical analysis used two-way ANOVA * = p < 0.05, ** = p < 0.01, *** = p < 0.001, **** = p < 0.0001 compared to controls at the same time points. See supplementary figure S4 for uncropped Western blot images.
To determine if VPA has the opposite effect on TORC1 in cells with a lower rate of glycolysis and a higher rate of oxidative phosphorylation (based on the contradictory reports presented in Table 1), we also assayed TORC1 activity in cells during the diauxic shift. The untreated control samples illustrate that TORC1 activity is reduced as cells transition from the late-log growth phase to the diauxic shift/stationary phase (Fig. 4D), a transition period that is defined by decreased fermentation (aerobic glycolysis) and increased oxidative phosphorylation. VPA and rapamycin both inhibited TORC1 activity within 1 h, but all samples had similarly low levels of TORC1 activity after 2 h (Fig. 4D). Thus, we do not see evidence suggesting that VPA alters TORC1 activity in cells during the diauxic shift.
Therapeutically relevant concentrations of VPA do not inhibit TORC2
To determine if VPA inhibits TORC2 as well as TORC1, which have distinct stimulants and downstream targets, we assayed specific downstream effects of VPA for each TOR complex. As mentioned above, Rps6 S232 is phosphorylated indirectly in response to TORC1 or TORC2 activity and S233 is dependent on TORC1 activity (Fig. 6B). Using the PP-Rps6 antibody, which binds to Rps6 only if both S232 & S233 are phosphorylated in combination with an antibody that interacts with phosphorylated S232 independently of S233 (called P-Rps6)83, it is possible to deduce the activity of each TOR complex. Treating cells for 1 h with VPA (1 & 2 mM) reduced PP-Rps6 (S232 & S232) antibody binding to undetectable levels, similar to that observed with treatment with rapamycin (0.4–1.2 µM), which specifically inhibits TORC1 (Fig. 6A), indicating that TORC1 activity is inhibited under these conditions. TORC2 levels are ascertained by P-Rps6 antibody binding when TORC1 is inhibited. In response to 1 mM VPA, P-Rps6 levels are only slightly diminished relative to control, indicating that VPA does not significantly affect TORC2 at therapeutically relevant concentrations. TORC2 activity is reduced with 2 mM VPA; however, this concentration is greater than what has been observed in patients and therefore likely not therapeutically relevant84,85.
VPA inhibits TORC1 but not TORC2 at 1 mM. (A) Cultures were treated with VPA (1 and 2 mM), rapamycin (0.4, 0.8, and 1.2 µM), or vehicles (control; C) for 30 min. Cell lysates were collected and assayed by SDS-PAGE and WB analyses, probing with PP-Rps6 (S232/S233) [TORC1 dependent phosphorylation], P-Rps6 (S232) [either TORC1 or TORC2 dependent phosphorylation], and Rps6 antibodies. Experiment was performed in both WT and npr2∆, the latter has constitutively active TORC1. (B) Diagram of TORC1 and C2 dependent phosphorylation of Rps6. S232 is phosphorylated indirectly in response to either TORC1 or TORC2 activity whereas S233 phosphorylation is dependent on TORC1 activity. The PP-Rps6 antibody binds only if both sites are phosphorylated and the P-Rps6 antibody interacts if either TORC1 or TORC2 is active. (C) Multiple residues on the c-terminus of Sch9 are phosphorylated directly by TORC1. Yeast exogenously expressing c-terminally tagged SCH9-3xHA were treated with cycloheximide (CHX; 100 µM), VPA (V; 1 mM), rapamycin (R; 400 nM), or vehicles (C) for 1 h, using culture medium containing a mixture of 16 amino acids (SC16D) (as our typical SC9D medium did not produce any observable Sch9 c-terminus phosphorylation). The c-terminus of Sch9-3xHA was chemically cleaved in cell lysates, which were then assayed by SDS-PAGE and WB analysis. The membrane was probed with anti-HA antibodies; the multi-phosphorylated Sch9 c-terminus (+ P) migrates slower than its dephosphorylated counterpart. The membrane was reprobed with an antibody that recognizes the TORC1 dependent phosphorylation of T737 on Sch9. (D) Ypk1 is phosphorylated directly by TORC2, as well as other protein kinases. A c-myc tagged YPK1 phosphosite mutant (S51A, S71A, T504A, S644A, T662A) retains only TORC2 specific phosphosites (YPK15A-myc). Cultures exogenously expressing this construct were treated with VPA (V; 1 mM), myriocin (M; 1.25 µM), or vehicles (C) for 0.5–4 h. Cell lysates were assayed by Mn2+ Phos-tag SDS-PAGE and WB analysis and probed with anti-myc antibodies. Phosphorylated proteins migrate slower on Phos-tag gels in a stoichiometric manner compared to their unphosphorylated counterpart. Data shown in A, C, and D is representative of 2 biological replicates which are shown in Supplementary figure S5. See supplementary figure S6 for uncropped Western blot images.
Using a complementary approach, we assayed the effect of VPA on direct targets of TORC1 and TORC2 kinase activities. Sch9 is a primary target of TORC1 and mediates many of its downstream regulatory effects86,87. The c-terminus of Sch9 has multiple TORC1 specific phosphorylation sites. Other protein kinases phosphorylate Sch9 outside of the c-terminus. To assay the phosphorylation status of the TORC1-specific c-terminal phosphosites, we exogenously expressed c-terminally tagged SCH9-3xHA in yeast and treated with VPA or controls rapamycin (a specific TORC1 inhibitor) and cycloheximide (which increases TORC1 activity88). As Sch9 is phosphorylated at other domains, the c-terminus was chemically cleaved off by treating cell lysates with NTCB (2-nitro-5-thiocyanatobenzoic acid). The samples were then assayed by SDS-PAGE and WB. This method produces a c-terminus that exhibits decreased mobility in the SDS-PAGE gel when highly phosphorylated, so that the phosphorylated 3xHA tagged c-terminus is separated from the dephosphorylated counterpart86,89. As expected, phosphorylation was increased with cycloheximide whereas rapamycin reduced phosphorylation (Fig. 6C). Phosphorylation of the Sch9 c-terminus following VPA treatment was similar to that observed in rapamycin treated samples, confirming TORC1 inhibition. To further validate this approach, the membrane was re-probed with an antibody that specifically interacts with phospho-T737 of Sch9 (Fig. 6C), which is directly phosphorylated by TORC187,90,91. These results mirrored the c-terminal results, confirming that VPA inhibits TORC1.
A primary and direct target of TORC2 is Ypk1, which mediates many downstream functions92. We utilized a reporter construct Ypk1(S51A S71A T504A S644A T662A) (called Ypk15A-myc), which is only phosphorylated by TORC2. Yeast cultures exogenously expressing this construct were treated with VPA or myriocin for 0.5–4 h. Myriocin inhibits sphingolipid production, the depletion of which stimulates TORC2 phosphorylation of Ypk192. Cell lysates were assayed by Mn2+ Phos-tag SDS-PAGE and WB. Phosphorylated proteins have reduced mobility in Phos-tag gels; the degree of decreased mobility corresponds to the stoichiometry of phosphorylated residues. Myriocin increased phosphorylation of Ypk15A-myc after 1–4 h, as expected, but VPA did not alter phosphorylation (Fig. 6D), confirming that 1 mM VPA does not decrease TORC2 activity.
To determine the global response of TORC1 and TORC2 targets to VPA, yeast cultures were treated with VPA or rapamycin during the log growth phase (see Supplementary figure S7A for experimental diagram). Samples were collected after 0–4 h and transcriptionally controlled targets of TORC1 and TORC2 were assayed by RT-qPCR (as demonstrated by93). Interestingly, these results revealed distinct differences between inhibition of TORC1 by rapamycin vs. inhibition by VPA. First, expression levels of known downstream transcriptional targets of TORC1 differed in response to VPA vs. rapamycin treatment. In fact, expression of one of the genes (GAP1) was affected in an opposite manner in response to the two drugs (Supplementary figure S7B). Second, VPA had no appreciable effect on TORC2 targets. Prolonged rapamycin treatment has been shown to decrease mTORC2 activity in mammalian cells94,95. After 4 h of rapamycin treatment, we observed transcriptional changes of TORC2 controlled genes (Supplementary figure S7C) consistent with decreased TORC2 activity93. No changes were observed in response to VPA. These data indicate that VPA inhibits TORC1 via a mechanism distinct from rapamycin-specific inhibition and further validated that 1 mM VPA does not affect TORC2 activity.
Increased TORC1 activity partially rescues VPA-induced repression of INO1
As TORC1 has been shown to increase INO1 transcriptional induction (Fig. 1B–D and52,53), we wished to determine if the VPA-mediated reduction in INO1 expression occurs as a result of TORC1 inhibition. VPA causes a robust decrease in TORC1 activity prior to the observed VPA-induced repression of INO1 transcription (Fig. 5A). Although VPA and inhibition of TORC1 by rapamycin both reduced INO1 induction, the expression profiles of the two treatments differed, suggesting that VPA and rapamycin reduce the induction of INO1 potentially by different mechanisms. These differences between VPA and rapamycin treatments are better observed in cells maintaining high expression levels of INO1 and MIPS in the mid-log growth phase. Unlike VPA treatment, rapamycin only slightly decreased INO1 mRNA expression (Fig. 2B) and did not reduce MIPS protein levels (Fig. 2C). Furthermore, TORC1 inhibition by VPA does not have the same effects as rapamycin on the TORC1 transcriptional targets tested (Supplementary figure S7B). During the late-log/diauxic shift growth phase, rapamycin elevated INO1 expression (Fig. 4B), which is also dissimilar to the results of VPA treatment (although MIPS protein levels were unaffected by either treatment (Fig. 4C)). Due to these discrepancies between VPA and rapamycin treatments on INO1 expression, it is unlikely that VPA-induced repression of INO1 is dependent on TORC1 inhibition.
We next sought to determine if increasing TORC1 activity could rescue VPA-mediated inhibition of INO1 expression. We could not stably express TORC1 subunits (TOR1/2) from a plasmid, which was a previously reported approach for rescuing TORC1 activity93. However, the mutant npr2∆ has constitutively active TORC196 and exhibits increased TORC1 activity relative to that of WT yeast. Npr2 is a subunit of the SEACIT complex, which reduces TORC1 signaling by inhibiting the EGO activating complex. In response to 1 mM VPA treatment, npr2∆ retained some TORC1 activity, which is lost in WT; however, 2 mM VPA or 0.4–1.2 µM rapamycin was sufficient to inhibit TORC1 activity in npr2∆ (Fig. 6A). WT and npr2∆ cells in the mid-log growth phase were treated with VPA or vehicle (control) for up to 4 h and INO1 mRNA and MIPS protein levels were then measured. As expected, VPA significantly decreased INO1 mRNA expression after 2 h and MIPS protein levels after 2–4 h in WT cells (Fig. 7). Surprisingly, npr2∆ had less INO1/MIPS expression relative to WT (approximately 50%). This is counterintuitive to what we would expect as TORC1 activity seems necessary for inducing INO1 expression. Treating npr2∆ with VPA led to decreased expression of INO1 mRNA and MIPS, but the magnitude of change was less than that observed in WT (Fig. 7B,D), consistent with the observed partial rescue of TORC1 activity in VPA-treated npr2∆ cells. INO1 mRNA levels in npr2∆ decreased by only 48–51% after 2–4 h of VPA treatment relative to 92% in WT after 2 h (Fig. 7B). MIPS protein levels following VPA treatment were also more stable in npr2∆ relative to WT, which was only decreased by 14–37% compared to 42–58% (Fig. 7D). Therefore, increased activity of TORC1 partially compensates for reduced MIPS expression induced by VPA.
Constitutively active TORC1 mutant (npr2∆) partially rescues VPA-induced MIPS depletion. WT and npr2∆ were inoculated in inositol-free medium and incubated until the log growth phase when cultures were treated with VPA (V; 1 mM) or vehicle (C; control). Samples were collected over a 4 h period. (A and B) INO1 mRNA was assayed by RT-qPCR analyses and plotted either as a function of time (line chart) (A) or as a bar graph to compare treatments/strains (B). (C and D) MIPS protein levels were measured by SDS-PAGE and WB analyses and plotted as a line chart (C) or as a bar graph (D). Percent decrease of (B) INO1 mRNA and (D) MIPS protein expression are shown in their respective bar graphs. Total protein normalization (TPN) was used to normalize WB quantifications. n = 4–5 biological replicates; error bars are SEM; two-way ANOVA, ns = not significant, ** = p < 0.01, *** = p < 0.001 relative to control at same time point. See Supplementary figure S8C-E for relative expression levels of the RT-qPCR data calculated using the transformed ∆Cq method.
The effect of VPA-induced activation of Snf1 on TORC1 activity and MIPS expression
VPA rapidly activates Snf1
Snf1 (yeast AMPK homolog) is a master metabolic regulator antagonistic to the TORC1 regulatory network. Snf1 and TORC1 are regulated by multiple levels of crosstalk (reviewed in56,57,91,97). For instance, activation of Snf1 decreases TORC1 activity directly and by affecting its target mediator proteins (such as Sch9)91,98,99,100. Conversely, TORC1 inhibition by rapamycin was also shown to increase Snf1 protein kinase activity101. VPA has been reported to activate Snf154. Therefore, we wished to determine if activation of Snf1 played a role in inhibiting TORC1 in response to VPA. We first compared the temporal response of VPA activation of Snf1 with that of inhibition of TORC1. An apparent two-fold increase in Snf1 activation (determined by T210 phosphorylation of Snf1102,103) occurred within 5 min of VPA treatment (Fig. 5A; P-Snf1 levels). Although this assay of VPA-treated samples produced high variation, the sustained upward trend of VPA-induced Snf1 activation suggests that activation occurs rapidly. This was accompanied by a significant decrease of TORC1 activity after 15 min (Fig. 5A; PP-Rps6 levels). The temporal response to VPA can be summarized as follows. Snf1 is activated after 5 min and TORC1 activity is reduced within 15 min (Fig. 5A). Opi1 localizes to the nucleus within 30–60 min, then partially returns to the ER after 2 h (Fig. 3B). The timing of Opi1 translocation aligns well with the INO1 mRNA expression profile as we would expect Opi1 nuclear localization to occur prior to the decrease in INO1 mRNA levels at 2 h, followed by a return of Opi1 to the ER, as INO1 mRNA levels are restored after 4 h of VPA treatment (Figs. 2B, 3B). At the same time, TORC1 activity is increased to moderate levels within 3 h of treatment (relative to its mostly inhibited state at 1 h) (Fig. 5A). This timeline indicates that Snf1 activation occurs prior to TORC1 inhibition and INO1 repression. Considering Snf1 activation has been shown to reduce TORC1 activity91,98,99,100 and its role in regulating INO1 expression104,105, we wished to understand how Snf1 activation may lead to inhibition of TORC1 and repression of INO1.
It is likely that inhibition of TORC1 by VPA may be dependent on Snf1 activation. Supporting this possibility, 30 min of VPA treatment did not reduce TORC1 activity in snf1∆ (Fig. 8). This was determined by treating aliquots of WT and snf1∆ cultures with VPA for 30 min at different growth stages (early-log to the diauxic shift) (Fig. 8A). TORC1 activity was measured by assaying PP-Rps6 levels. In WT, TORC1 is active during early to mid-log growth phases and inactive during the late-log/diauxic shift transition (Fig. 8B). In snf1∆, TORC1 is active throughout the log growth phase and even during the diauxic shift. More importantly, 30 min of VPA treatment did not reduce TORC1 activity in snf1∆ at any stage of growth. This suggests that TORC1 inhibition after 30 min of VPA treatment is Snf1-dependent. However, snf1∆ has a much slower growth rate than WT (Fig. 8A) and may require a longer duration of VPA treatment before TORC1 activity is reduced. Additionally, Snf1 dependent inhibition of TORC1 may only be true for the early stages of VPA treatment, as is the case with glucose limitation91,100. Therefore, we wanted to ascertain if TORC1 inhibition by VPA is dependent on Snf1 activation after prolonged treatments.
TORC1 is inhibited by 30 min of VPA treatment only during the mid-log growth phase, and inhibition is absent in snf1∆. (A) Growth curve of WT and snf1∆ on SC9D + inositol (75 µM). OD600 was monitored by a plate reader growth assay. Arrows indicate growth stages where a portion of a culture was divided and treated -/ + VPA (1 mM) for 30 min. (B) Cells were then lysed and assayed by SDS-PAGE and WB. TORC1 activity was determined by probing for PP-Rps6 (a biomarker for activity). Yellow = early-log, blue = mid-log, and green = late-log/diauxic shift. n = 1 biological replicate. See supplementary figure S9 for uncropped Western blot images.
VPA-induced repression of MIPS is not dependent on Snf1 activation
To determine if MIPS-depletion is an outcome of Snf1 activation, and if TORC1 inhibition by VPA is dependent on Snf1, we expressed plasmids containing SNF1 (WT gene) or activation-dead SNF1(T210A) (called SNF1ad) in the snf1∆ mutant. Phosphorylation of T210 in Snf1 (homologous to T172 in AMPK) is required for kinase activation [57,97,102,106]. Thus, the T210A point-mutant retains a functional kinase domain107,108,109 but cannot be activated.
These SNF1 expressing strains (+ SNF1 and + SNF1ad) along with WT and snf1∆ were treated with VPA for 30 min and 2 h. As a positive control for increasing Snf1 activation, we also subjected cultures to low-glucose (0.05%)54,91,110. Cell lysates were assayed for Snf1 activation, TORC1 activity, and MIPS expression. snf1∆ had slightly less MIPS protein content than WT, which was rescued by expressing + SNF1 or + SNF1ad (Supplementary figure S11). SNF1 deletion is known to cause partial inositol auxotrophy105,111,112 which may be explained by decreased INO1 transcription52,105 and decreased MIPS enzyme levels, resulting in decreased production of inositol. As expected, Snf1 was activated with VPA treatment and low-glucose (the latter to a greater degree), and there was no activation of Snf1 in snf1∆ and + SNF1ad strains (as these samples lack P-Snf1) (Fig. 9A). VPA and low-glucose both reduced TORC1 activity even in snf1∆ and + SNF1ad strains (Fig. 9B), suggesting that Snf1 activation by VPA (or low-glucose) is not essential for reducing TORC1 activity. Unlike low-glucose, VPA reduced MIPS expression in all strains, but repression was delayed in strains unable to activate Snf1 (snf1∆ and + SNF1ad) (Fig. 9C). Thus, the early stages may be facilitated by Snf1 activation, but VPA inevitably decreases MIPS expression by another mechanism. Overall, these data suggest that VPA induces Opi1 nuclear localization which represses INO1 transcription, however, repression is not dependent on TORC1 inhibition or Snf1 activation.
Snf1 activity is required for early MIPS repression in response to VPA. WT, snf1∆, snf1∆ + SNF1, and snf1∆ + SNF1ad were inoculated in inositol-free medium and incubated until the log growth phase when cultures were treated with VPA (V; 1 mM), low-glucose (LG; 0.05%), or vehicle (C; control) for 0.5–2 h. Cells were lysed and (A) Snf1 activation (P-Snf1), (B) TORC1 activity (PP-Rps6), and (C) MIPS expression levels measured by SDS-PAGE and WB analyses. Total protein normalization (TPN) was used to normalize WB quantifications. n = 3–4 total biological replicates collected over 3 independent experiments. Data is displayed relative to 0.5 h control treatment for each strain and percent change of MIPS expression levels is displayed above the data bar. Error bars are SEM. Statistical analysis determined for each strain independently relative to controls at the same time point; two-way ANOVA, * = p < 0.05, ** = p < 0.01, *** = p < 0.001, **** = p < 0.0001. See Supplementary figure S11 for WB images and quantifications relative to WT 0.5 h control.
Discussion
The current study utilized the yeast model to determine the mechanism whereby VPA causes inositol depletion. Our findings indicate the following: 1) The mechanism whereby VPA reduces INO1 expression is by increasing nuclear localization of the INO1 repressor Opi1. 2) VPA inhibits activity of TORC1, an activator of INO1 expression. Upregulating TORC1 partially rescues VPA-induced repression of INO1, but TORC1 inhibition is not essential for inhibition of INO1 expression by VPA. 3) Snf1 is activated in response to VPA, but Snf1 activation is not required for VPA-mediated inhibition of TORC1 or INO1 repression. Thus, these are independent outcomes of VPA treatment.
Opi1 was essential for decreasing INO1 expression in response to VPA (Fig. 3A). Opi1 is primarily regulated by its interactions with PA in the ER. Either decreasing pools of PA or increasing the acyl chain lengths of PA can release Opi1, which then translocates to the nucleus81,104. We speculate that VPA modifies the pool of PA in the ER, potentially by increasing triacylglycerol (TAG) synthesis, a process mediated by Snf1 and TORC1, among other factors113,114,115. Accordingly, a quadruple mutant unable to synthesize TAG was resistant to growth suppression by VPA116. VPA is known to alter lipid homeostasis and ER function, inducing stress responses (such as the unfolded protein response)116,117. Although an equivalent to the Henry regulatory circuit has not been identified in mammals, PA plays a key role in regulation of MIPS expression in both yeast and mammalian cells19,33,118,119. In MEF cells, VPA was shown to decrease MIPS expression, levels of which are regulated by PA pools in the plasma membrane33. Alteration to PA homeostasis may be the underlying effect of VPA that links its inositol depletion effect in both yeast and mammals. Accordingly, VPA did not alter INO1 expression during the diauxic shift (Fig. 4B) as Opi1 is natively localized to the nucleus at this growth stage. This is facilitated by decreased PA pools in the ER brought on by increased TAG synthesis during the diauxic shift (Fig. 1A).
While INO1 and MIPS protein expression are reduced in response to VPA during the log growth phase of yeast32, INO1 expression subsequently recovers (Fig. 2B). These responses are dependent on the localization of the Opi1 repressor. In addition, VPA is an inhibitor of HDACs, which are ultimately recruited to reduce INO1 expression by the action of Opi1. Prolonged treatments with VPA may inhibit HDACs, which could also lead to increased INO1 expression (which is what we observe in Fig. 2B, 4hrs). Interestingly, however, MIPS protein levels remain low even after INO1 expression recovers (Fig. 2C). This may be due to a concomitant decrease in translation/protein synthesis following VPA treatment116,117 likely caused by reduced TORC1 activity.
The finding that INO1 and MIPS expression is not significantly altered during the diauxic shift (Fig. 4B,C and S3E) is somewhat contradictory to previous reports, which showed that VPA increases expression during this growth phase120,121. These methods calculated expression levels by normalizing to housekeeping genes, which we found to be an acceptable method during the log growth phase, but problematic during the diauxic shift, leading to overestimating the levels of INO1 expression (see Technical note in Methods section). While we also found that VPA treatment increased INO1 expression during stationary phase, the magnitude of change is much smaller than previously reported (Fig. 4B and S3E). Indeed, these relative expression levels are dwarfed in comparison to those observed during log growth, bringing into question the biological significance of this relatively small increase during the stationary phase. As previously reported, VPA depletes inositol in log-phase and levels remain low throughout the diauxic shift/stationary phase120. As we do not also see a concomitant increase in MIPS protein levels during stationary phase (Fig. 4C), we conclude that this small increase in INO1 expression does not translate into more MIPS enzyme and, therefore, does not lead to increased inositol synthesis during the diauxic shift/stationary phase in VPA treated cells.
Our finding that inhibition of TORC1 by rapamycin decreased INO1 induction (Fig. 1B,D) coincides with the previous report that TORC1 activity enhances INO1 induction52. However, the INO1 expression profiles in response to VPA and rapamycin differed (Fig. 1B), suggesting that VPA and rapamycin reduce the induction of INO1 by different mechanisms. Differences between VPA and rapamycin treatments are better observed in cells maintaining high expression levels of INO1 and MIPS in the mid-log growth phase. Unlike VPA treatment, rapamycin only slightly decreased INO1 mRNA expression and did not reduce MIPS protein levels (Fig. 2B,C). During late-log/diauxic shift, rapamycin, unlike VPA, actually elevated INO1 expression (Fig. 4B). Furthermore, TORC1 transcriptional targets are affected differently by VPA- vs. rapamycin-inhibition of TORC1 (Supplementary Figure S7B). Collectively, these findings suggest that VPA and TORC1 inhibition downregulate INO1 expression by different mechanisms.
VPA rapidly inhibits TORC1 (Figs. 5 and 6), yet numerous downstream effects of TORC1 inhibition were absent, which were otherwise observed with the specific TORC1 inhibitor rapamycin (Figs. 1, 2, 4, S7). There are several possible explanations for the discrepancies between rapamycin and VPA treatments. 1) There are rapamycin resistant targets of TORC1122,123,124,125. Active site inhibitors of TORC1 elicit increased inhibition and more widespread effects on downstream targets, whereas rapamycin allosterically inhibits TORC1, which may retain some residual activity or have compensatory activation following prolonged treatment122. This compensatory activation was also observed in yeast after 3 h of VPA or rapamycin treatment (Fig. 5A), and our data suggest that VPA is not a specific inhibitor or active-site inhibitor of TORC1. 2) Rapamycin is a specific inhibitor of TORC1 whereas VPA is non-specific and causes pleiotropic effects. Thus, VPA inhibits TORC1 but also affects other regulatory networks (e.g., Snf1, GSK3, HDACs, etc.), confounding our ability to interpret cause-and-effect outcomes of treatment. Lastly, 3) TORC1 may not be a mediator of VPA-induced INO1 repression, which we expect is the most likely explanation for these results. However, due to the confounding properties of the npr2∆ experimental data (Fig. 7) and our inability to prevent TORC1 inhibition using SNF1 mutants (Fig. 9), further experimental validation is required to confirm this conclusion.
While increased TORC1 activity in npr2∆ can rescue MIPS expression in response to VPA, it cannot be concluded that TORC1 mediates INO1 repression by VPA, as compensatory mechanisms cannot be ruled out. Furthermore, disrupted cellular homeostasis and distinctive phenotypes of npr2∆ obscure interpretation of the data. Specifically, 1) in VPA treated npr2∆ cells, INO1 mRNA expression decreased to similar levels observed in 2 h VPA treated WT (Fig. 7A,B), which may be the lowest level of repression achievable with VPA treatment. Therefore, the reduced repression observed in npr2∆ (Fig. 7B, WT 92% reduction as opposed to npr2∆ 48% reduction) is due to less INO1 expression occurring prior to VPA treatment. 2) npr2∆ has a slower rate of growth than WT (Supplementary figure S6A). If the rate of decreasing MIPS protein levels is dependent on growth-rate, then slower growing cells will experience MIPS depletion at a slower rate (e.g., as cells divide, the pool of MIPS proteins is distributed to daughter cells and is not replenished by de novo synthesis). As such, VPA treated npr2∆ cells exhibited a slower rate of depletion of MIPS relative to WT (Fig. 7C,D). 3) VPA treated npr2∆ cells only partially rescue TORC1 activity and MIPS depletion. Thus, the rescue of MIPS depletion could be explained by either a dependent mechanism or a compensatory one. We attempted to further increase TORC1 activity in the presence of VPA by additionally treating with CHX (Supplementary figure S6B-E), but CHX treatment (which induced growth arrest) is unsuited for this 4 h long experiment. Lastly, 4) npr2∆ has half the INO1 and MIPS expression levels observed in WT (Fig. 7) and its growth is inhibited by exogenous inositol (Supplementary figure S6A), the latter of which is a very unusual phenotype (discussed further below). Although the VPA-induced decrease in INO1 expression was somewhat attenuated in npr2∆, the mutant did not exhibit recovery of INO1 expression, which was observed in WT after 4 h (Fig. 7A,B). This indicates that npr2∆ has a general defect in regulating INO1 and MIPS expression, implying that TORC1 homeostasis (not just increased activity) is important for MIPS expression. Based on the current evidence, we suspect that TORC1 inhibition is not a dependent mechanism of INO1 repression by VPA, but rather, increased TORC1 activity can compensate for the MIPS-depleting effects of VPA.
Most likely, TORC1 plays two opposing roles in regulating INO1 transcription. TORC1 activity enhances the induction of INO1 (Fig. 1 and52), but surprisingly, the constitutively active TORC1 mutant (npr2∆) has decreased INO1 and MIPS expression (Fig. 7). npr2∆ (untreated) had half the expression levels of INO1 mRNA and MIPS protein compared to WT (untreated) (Fig. 7), which is a counterintuitive result. Strikingly, npr2∆ is not inositol auxotrophic (as this expression profile would imply), but instead grows more slowly when inositol is added to the medium (Figure S6A). Npr2 is part of the SEACIT complex (the mammalian GATOR1 equivalent) that reduces TORC1 activity in yeast, presumably during nutrient-limiting conditions. How can inositol inhibit growth when TORC1 has lost this regulatory input? While this is a rather unusual phenotype, it is reminiscent of how some cancers respond to excess inositol. Growth of some cancers (which notoriously have higher mTORC1 activity) can be inhibited by adding excess inositol, potentially by decreasing activity of PI3K, an activator of mTORC1 and a pathway considered essential for metastasis (reviewed in23). The efficacy of using inositol as a tumor growth suppressor may be linked to the mechanisms inducing mTORC1 hyperactivation in cancers. This link remains to be investigated.
While the specific conditions that lead to inhibition of mTORC1 by VPA are unknown (Table 1), cells undergoing anabolic metabolism (during which activity of mTOR is high and AMPK is low) would be more prone to modulation of these kinases. Snf1 activation inhibits TORC1 during early stages of glucose starvation, but not after prolonged glucose limitation91,100. VPA treatment has a similar outcome, as TORC1 inhibition was reduced in snf1∆ during the early stages of VPA treatments (< 30 min, Fig. 8B), but prolonged treatments (> 30 min) showed similar levels of TORC1 inhibition in both snf1∆ and WT (Fig. 9B). Additionally, exogenous expression of the kinase inactive mutant SNF1ad did not prevent TORC1 inhibition in response to VPA (Fig. 9B). Therefore, the mechanism whereby VPA leads to TORC1 inhibition is ultimately not Snf1 dependent.
INO1 transcription is partially regulated by signaling networks controlled by TORC1 and Snf1. SNF1 loss-of-function mutants are partially inositol auxotrophic due to decreased transcription of INO152,105; exogenous expression of SNF1 restores MIPS expression (Supplementary figure S11D, control treatments). Surprisingly, exogenous expression of kinase inactive mutant SNF1ad also restores INO1 expression to endogenous levels (Supplementary figure S11D, control treatments). This suggests that Snf1 has two opposing roles in regulating INO1: Kinase-inactive Snf1 increases INO1 expression, which is decreased during glucose limitations by kinase-active Snf1, possibly by mediating acetyl-CoA carboxylase (Acc1) activity and PA-Opi1 interactions104,105. Notably, exogenously expressing SNF1 did not rescue the reduced growth of snf1∆ on glucose medium (Supplementary figure S10); expression was from a low-copy number vector and under control of its endogenous promoter and terminator. Thus, expressing SNF1 in snf1∆ can restore some but not all functionality, as snf1∆, snf1∆ + SNF1, and snf1∆ + SNF1ad all grew slower than WT. Oddly, Snf1 activation, which is known to reduce INO1 expression in an Opi1 dependent manner, was not essential for decreasing MIPS expression with 2 h of VPA treatment, although this was the case for low-glucose treatment. Low-glucose reduced MIPS expression in WT and + SNF1 but this effect was lost in snf1∆ and + SNF1ad, suggesting that MIPS repression by glucose limitation is facilitated by Snf1 activation. And while MIPS repression was delayed in strains unable to activate Snf1 (snf1∆ and + SNF1ad), VPA equally reduced expression in all strains after 2 h (Fig. 9C). One difference between low-glucose vs. VPA treatments was that Snf1 activation was much greater with low-glucose as opposed to VPA (Fig. 9A). This degree of activation may explain why Snf1 activation did not play a significant role in MIPS repression following VPA treatment. Therefore, it is most likely that the pleiotropic effects of VPA cause inositol depletion by multiple pathways. While the early response of VPA-induced MIPS repression may be mediated by Snf1, the primary mechanism is Snf1 independent.
A noteworthy target of VPA includes HDACs, which alter histone acetylation and genetic transcription. The general inhibitory effect of VPA on class I and II HDACs can lead to widespread transcriptional changes. The discovery of this activity in efforts to determine teratogenic effects of the drug43,44 led to numerous studies investigating VPA as a potential tumor growth repressor, most of which attribute the tumor growth suppression effects on HDAC inhibition, and assume effects on mTOR activity and autophagy are induced by HDAC inhibition. In light of our new evidence showing that TORC1 inhibition happens within 15 min of treatment (Fig. 5A), we suspect that this assumption is rather incomplete. In addition, the IC50 of VPA for HDAC1 was determined to be 0.4 mM44, well within the clinically relevant concentrations of cerebral spinal fluid (0.28–0.98 mM) and blood plasma samples (≈0.48 mM)84,85. We found that TORC1 inhibition also occurs at similar concentrations in yeast (Fig. 5B). However, TORC1 inhibition and Snf1 activation occur too rapidly to be explained by HDAC inhibition, as this would imply that epigenetic remodeling and expression changes would have to take place prior to influencing the activity of these metabolic pathways. The HDAC-dependent effects induced by VPA remain undetermined (as this hypothesis is based on correlative finding). Nonetheless, we suspect that the pleiotropic effects of VPA on multiple pathways is the reason this drug has such a diverse number of outcomes and underlies its utility as a pharmacological agent.
Snf1 and TORC1 are affected very rapidly by VPA, indicating that this effect is not dependent on VPA-mediated HDAC inhibition. How VPA regulates Snf1 and TORC1 remain important unanswered questions. Potential candidates mediating VPA-induced TORC1 inhibition and AMPK activation include GSK3, which is down-regulated by VPA49,126) or the MAPK stress response pathways (yeast Hog1 and Slt2), which are activated in response to VPA116. The extensive mechanisms of crosstalk among these signaling cascades in addition to the variety of outcomes of VPA treatment convolute our understanding of the mechanism(s) of action of VPA. Determining which of these mechanisms are therapeutically relevant will help to identify better therapeutic targets and contribute to our understanding of the underlying causes of neurological disorders, such as epilepsy, bipolar disorder, and migraines.
Methods
Strains, medium, and growth conditions
Saccharomyces cerevisiae budding yeast strains used in this study include WT BY4741 and derived deletion mutants npr2∆, opi1∆, and snf1∆ isolated from the yeast deletion collection (Invitrogen 95,401.H2). Deletions were confirmed by PCR screening. The Opi1-mGFP expression strain (BY4742 background) was provided to us by another research group127, which was constructed by integrating tagged OPI1 into the chromosome.
YPD medium consisted of 1% yeast extract, 2% bactopeptone, and 2% D-glucose. Inositol-free synthetic complete medium consisted of yeast nitrogen base (with ammonium sulfate, without inositol; MP 4,027,412), 2% D-glucose, and a mixture of 9 amino acids (SC9D) [adenine (4 mg/L), arginine (4 mg/L), histidine (2 mg/L), leucine (12 mg/L), lysine (4 mg/L), methionine (4 mg/L), threonine (60 mg/L), tryptophan (4 mg/L), uracil (8 mg/L)]. Experiments assaying Sch9 phosphorylation required more amino acids and instead utilizing a mixture of 16 amino acids (SC16D) [Adenine (60 mg/L), Uracil (20 mg/L), Histidine (20 mg/L), Leucine (120 mg/L), Tryptophan (80 mg/L), Arginine (20 mg/L), Aspartic acid (100 mg/L), Glutamic acid (100 mg/L), Phenylalanine (50 mg/L), Lysine (30 mg/L), Methionine (20 mg/L), Serine (375 mg/L), Threonine (200 mg/L), Valine (150 mg/L), Tyrosine (30 mg/L), Isoleucine (30 mg/L)]. Inositol (75 μM) was added where indicated (I +).
Cells were maintained on YPD petri dishes (+ 2% agar), or SC9D I + Leu- if housing an expression plasmid (see below). Cells were precultured in 1–5 mL of YPD or SC9D I + Leu- overnight to the stationary growth phase. Precultures were washed in 5 mL sterile water by centrifuging at 3000 × g for 5 min to pellet cells, discarding supernatant, vortexing in water, and centrifugation repeated to collect washed cell pellet. All washed strains (including those housing plasmids) were resuspended in fresh SC9D medium for experimentation, or SC16D for Sch9 phosphorylation analyses. Cell culture ocular densities were measured in a BioMate 160 cuvette spectrophotometer (Thermo Fisher) with a wavelength of 600 nm (OD600). All incubations were at 30 °C. Liquid cultures were incubated with shaking (230 rpm).
Notably, P. Kumar and colleagues used torin 2 to inhibit both TORC1 and TORC2 activity in budding yeast93. Torin 2 is an ATP-analog that very specifically interferes with the ATP binding domain of TOR protein kinases. We did not see inhibition of TORC1 or TORC2 or an effect on growth with torin 2 (data not shown), unlike their results. This may be due to the different yeast S. cerevisiae background strains used in these studies (P. Kumar et al. used JK9-3d whereas I used BY4741).
Plasmid construction and yeast transformation
Plasmids were constructed using the In-Fusion HD cloning kit (Takara) according to the user manual (with minor modifications). Genomic DNA was isolated from BY4741 yeast using a quick lithium acetate extraction method128 and was the template for the following PCR reactions. To construct the pRS415-SCH9-3xHA plasmid, two fragments were amplified by PCR (PrimeSTAR GXL Premix (Takara)) and inserted into pRS415 (linearized by XhoI and NotI). The first fragment included the SCH9 promoter region, SCH9, and half of the 3xHA sequence, and the second fragment included the other half of the 3xHA sequence and the SCH9 terminator region. The linearized vector and fragments were purified from agarose gels with the NucleoSpin Gel and PCR Clean-up kit (Takara). The fragments were subjected to re-amplification up to four times, until enough was obtained for the In-Fusion cloning reaction, where a molar ratio of 2:2:1 (fragment 1:fragment 2:plasmid) was used. The reaction was transformed into Stellar Competent Cells (included with the kit) which were plated on LB ampicillin (100 µg/mL) medium.
Construction of pRS415-SNF1-3xmyc was carried out following a similar method with the following modifications. Two fragments were amplified by PCR (Q5® Hot Start High-Fidelity 2X Master Mix) and inserted into pRS415 (linearized by XbaI and SalI). The first fragment included the SNF1 promoter region, SNF1, and half of the 3xmyc sequence, and the second fragment included the other half of the 3xmyc sequence and the SNF1 terminator region. The fragments and purified vector were combined at a molar ratio of 2:4:1 (fragment 1:fragment 2:plasmid) for the In-Fusion cloning reaction. pRS415-SNF1(T210A)-3xmyc was constructed using the Q5® Site-Directed Mutagenesis Kit according to the user guide.
The pRS315-YPK1(S51A S71A T504A S644A T662A)-myc (aka pRS315-YPK15A-myc) was constructed by another research group92.
All plasmids were isolated from E. coli and sequences confirmed by Sanger sequencing. Yeast cells were transformed with plasmids according to a high efficiency electroporation method as described129. Positive transformations were selected on SC9D I + Leu- medium and protein expression confirmed by SDS-PAGE and WB analyses prior to experimentation. All PCR primers utilized in this section are shown in Table 2.
SDS-PAGE and Western blotting (WB)
Cell lysates for most SDS-PAGE and WB analyses were collected following a boiling technique for yeast lysis as published103, however, we resuspended the cell pellets in 60 µL of 1X Laemmli buffer (62.5 mM tris pH 6.8, 10% (v/v) glycerol, 5% (v/v) 2-mercaptoethanol, 2% sodium dodecyl sulfate, 0.01% bromophenol blue) per 1.0 OD600 x mL cells collected and loaded 2—4 µL of cleared supernatant per well. Hand cast tris–glycine-SDS (-/ + 25% glycerol) gels were used for protein electrophoresis in tris–glycine running buffer (25 mM tris pH 8.5, 190 mM glycine, 0.1% SDS) using the Mini-PROTEAN Tetra cell electrophoresis system (Bio-Rad). Gels were electrophoresed at a constant 120 V (or 160 V when 25% glycerol was included in the separating gel). Proteins were transferred from gels to PVDF membranes using the Mini Trans-Blot system (Bio-Rad) according to the manual. No-Stain Protein Labeling Reagent (Thermo Fisher) was used to label total protein according to the user guide. Membranes were blocked with 3% milk (non-fat) prior to antibody probing. Antibodies used in this study are included in the antibodies table. Membranes were incubated with primary antibodies overnight at 4 °C (unless stated otherwise) followed by washing with TBST (0.1% Tween-20) for 5 min, three times. Secondary antibody incubations lasted for 1 h at room temperature, then washed with TBST for 10 min, three times. SuperSignal West Pico PLUS (Thermo Scientific 34,580) chemiluminescent substrate was incubated with membranes prior to imaging HRP conjugated secondary antibodies (or SuperSignal West Atto Ultimate Sensitivity Substrate (Thermo Scientific A38554) for P-Snf1 imaging). Image capture of No-Stain labeled proteins, Alexa Fluor 800, and HRP used the iBright™ FL1500 Imaging System (Invitrogen). Europium conjugated secondary antibodies were imaged with the SpectraMax i3 ScanLater cartridge (Molecular Devices); we found that dried membranes labeled with No-Stain Labeling Reagent had high levels of background using this imaging technique. Blocking with 3% BSA (bovine serum albumin) rather than milk produced lower background when imaging with the ScanLater cartridge. All antibodies utilized in this section are shown in Table 3.
Sch9-3xHA c-terminus phosphorylation assay
Assaying Sch9 c-terminal phosphorylation was performed according to the original methodology of Jörg Urban and colleagues86 with modifications89. Briefly, cell culture ODs were measured then 5 mL cultures were placed on ice. 500 µL 60% cold TCA (i.e., 1/10 culture volume) was mixed with the culture and incubated on ice for 10 min. Cells were pelleted at 3,000 × g for 5 min at 4 °C. Most of the supernatant was discarded, leaving some behind (200–1000 µL) to resuspend the pellet and transfer to a microcentrifuge tube. Cell pellets were collected by centrifugation at 16,000 × g for 2 min at 4 °C. Supernatant was discarded and cells washed twice with 700 µL cold acetone, pelleting cells at 16,000 × g for 2 min at 4 °C between washing steps. Care was taken to remove most traces of TCA. Pellets were dried by heating to 65 °C for 2 min with the cap open. Pellets were then suspended in urea buffer (3 M urea, 50 mM tris pH 7.5, 1% SDS, 5 mM EDTA), with the addition of Phosphatase Inhibitor Cocktail Set V (Millipore 524,629) (30 µL/mL lysis buffer), ProBlock Gold Yeast/Fungi Protease Inhibitor Cocktail (Gold Biotechnology GB-333–5) (10 µL/mL lysis buffer), 0.5 M EDTA (ProBlock™ Gold kit component) (5 µL/mL lysis buffer), and 1 M 1,10-Phenanthroline (ProBlock™ Gold kit component) (5 µL/mL lysis buffer). Volume of urea buffer used was 33 µL urea buffer per 3.0 OD600 x mL cells collected. Suspended cells were heated to 65 °C for 10 min then allowed to cool to room temperature. Per 3.0 OD600 x mL cells collected, 10 µL 0.5 M CHES [Cyclohexyl-2-aminoethanesulfonic acid] pH 10.5 and 7 µL fresh 5 mM NTCB [2-nitro-5-thiocyanatobenzoic acid] was added and mixed by brief vortexing. Solutions were incubated at room temperature overnight (> 18 h) in the dark. The following day, per 3.0 OD600 x mL cells collected, 16.66 µL 4 × Laemmli buffer (0.25 M Tris pH 6.8, 40% (v/v) glycerol, 20% (v/v) 2-mercaptoethanol, 8% sodium dodecyl sulfate, 0.04% bromophenol blue) was mixed into the solutions, heated to 85 °C for 5 min, then store at -20 °C. Supernatants were clarified by centrifuging at 16,000 × g for 1 min and 10 µL was loaded per lane on SDS-PAGE gels.
To prepare fresh 5 mM NTCB in 20% ethanol, powdered NTCB was dissolved in 100% ethanol (20 µL ethanol/0.1121 mg NTCB) to make 25 mM solution, then diluted with water to 5 mM (80 µL water/0.1121 mg NTCB).
YPK15A-myc phosphorylation assay
Analysis of YPK15A-myc phosphorylation was carried out according to published methodology89,92, with modifications. Briefly, cell culture ODs were measured, 3.0 OD600 x mL cells in culture were transferred to a centrifuge tube, then pelleted at 3,000 × g for 5 min at 4 °C. Pellets were frozen at − 20 °C until all samples were collected. Pellets were resuspended in 0.9 mL 0.255 M NaOH and 1% 2-ME, transferred to a microcentrifuge tube, and incubating on ice for 5 min. 150 µL 60% cold TCA was added, mixed by inversion, and incubate for 15 min on ice. Cell pellets were collected by centrifugation at 16,000 × g for 2 min at 4 °C. Supernatant was discarded and cells washed twice with 700 µL cold acetone, pelleting cells at 16,000 × g for 2 min at 4 °C between washing steps. Care was taken to remove most traces of TCA. Washed pellets were subjected to vacuum centrifugation for 5 min to evaporate all acetone. Pellets were resuspended in 50 µL MURB buffer (100 mM MES pH 7, 3 M urea, 1% SDS, 0.2% bromophenol blue, and 5% 2-ME, plus 1 mM MnCl2) and heated to 100 °C for 5 min then stored at − 20 °C. Supernatant was clarified by centrifugation at 16,000 × g for 1 min prior to loading 5 µL on Mn2+ Phos-tag gels for analysis.
1.0 mm thick Mn2+ Phos-tag protein gels were cast following the Phos-tag SDS-PAGE Guidebook. This gel chemistry uses the standard tris–glycine-SDS page gel formula (see above) but is supplemented with Phos-tag acrylamide (25 µM) and manganese chloride (50 µM). Gel electrophoresis lasted for 3:40 h under constant 120 V. Proteins were transferred from gels to PVDF membranes using a wet-tank method with 1 L transfer buffer (25 mM tris pH 8.5, 190 mM glycine, 10% methanol, 0.1% SDS) under a constant 100 V for 1:30 h.
Real-time quantitative PCR (RT‑qPCR) analyses
Yeast cultures were pelleted by centrifugation at 3000 × g for 5 min. Supernatant was discarded and pellets moved into microcentrifuge tubes by resuspending in 1 mL cold water. Pellet was collected again by centrifugation at 11,000 × g for 1 min, supernatants discarded, and stored at -80 °C. RNA was isolated with the NucleoSpin RNA Plus kit (Takara 740,984) according to the user guide. Yeast cell lysis was achieved by suspending the pellet in 350 µL Buffer LBP (included in the kit) and transferred to a 2 mL screw top microcentrifuge tube filled with 175 µL ZrO2 beads (MP Biomedicals 11–654-0436). Tubes were loaded into the MP FastPrep-24 and cycled at 6.5 M/sec for 60 s followed by resting 2 min in the cold room, repeating a total of four times. Lysates were centrifuged for 2 min at 20,000 × g to collect the clarified supernatant for isolating RNA with the NucleoSpin RNA Plus kit. Concentrations of purified RNA was measured with a NanoDrop 2000c spectrophotometer (Thermo Scientific). cDNA was synthesized using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara RR047A). PowerUp SYBR Green Master Mix (Applied Biosystems A25741) was used for the real-time quantitative PCR (RT‑qPCR) followed the manufacturer guidelines. Specifically, each mix contained 400 nM forward and reverse primers and 1 µL cDNA (diluted 1/10 in water) in a total volume of 10 µL. Three technical replicate reactions were prepared for each sample, which were then averaged to provide the sample Cq value. Real-time fluorescence of the reactions was measured using a QuantStudio 3 Real-Time PCR System (Applied Biosystems).
Primers were designed according to a published set of guidelines130, except for the TAF10 and UBC6 primer pairs which were designed by another research group131 (qPCR primers are shown in Table 4). The efficiencies calculated using standard methodology described elsewhere131. Relative expression fold-changes were calculated using the following methods. For ACT1 housekeeping gene normalizations, Pfaffle’s method132 was used which accounts for primer pair amplification efficiencies. For TAF10 and UBC6 normalizations, a modified Pfaffl methodology was used which calculates their geometric average for normalizing the relative expression data131. Calculating the transformed Cq value used the equation (E-Cq) [E = primer pair amplification efficiency] for each sample. Relative expression was determined by averaging the replicates and normalizing to a sample set designated as the experimental reference (for example, relative expression = average(E-Cq[2 h VPA]) / average(E-Cq[0 h control]), with 0 h control designated as the experimental reference).
Technical note: Expression of housekeeping genes used for normalizing RT-qPCR change during the diauxic shift. During the process of testing INO1 mRNA during the diauxic shift, it became apparent that the housekeeping gene we were using to normalize the RT-qPCR data was decreasing during this growth phase, which was further exacerbated by VPA treatment. This greatly overestimated the calculated expression changes of INO1 mRNA (compare Supplementary Figure S2B vs Fig. 4B and Supplementary Figure S3A vs S3E). This prompted us to find better normalization genes. The publication by Marie-Ange Teste, Manon Duquenne, and colleagues identified a number of genes with stable expression during prolonged growth on glucose (an experimental setup similar to our own). Four of these genes were then identified to remain stable among different strains (e.g., mutants, backgrounds, etc.) and growth conditions (e.g., treatments, carbon source, etc.)131. Furthermore, they strongly discouraged the use of ACT1 as a normalization reference. Our data are in agreement with their assessment as ACT1 expression was diminished during the diauxic shift (Supplementary figures S2C and S3B). To visualize mRNA expression level changes without housekeeping gene normalization, we calculated the transformed Cq value (E-Cq) [E = primer pair amplification efficiency] then normalized to the reference sample (e.g., time 0 h) to determine fold-change. This method is similar to Livak’s 2-∆∆Cq method133 but without normalization to a reference gene, and considers the primer pair efficiency as with Pfaffl’s method132. This transformed Cq method relies on accurately inputting similar amounts of RNA/cDNA per RT-qPCR reaction134, which is a typical normalization step for synthesizing cDNA from RNA. As reported by Teste, Duquenne, et al., we also observed ACT1 mRNA levels decrease during stationary phase and VPA further exacerbated this effect (Supplementary Figures S2C and S3B). However, during the mid-log growth phase, ACT1 expression remained relatively stable (Supplementary Figure S1C), validating its use as a normalization reference gene with our treatment conditions during this growth phase. And while TAF10 and UBC6 expression (two of the four recommended genes mentioned above) are more stable than ACT1, they both decrease by 6- and 3- fold during stationary phase (Supplementary Figures S3D and S3F). This led to overestimating the expression of INO1 mRNA during stationary phase (compare Supplementary Figure S3C vs S3E). VPA did not appear to further reduce expression levels of TAF10 and UBC6, unlike with ACT1. While relying on similar input of RNA/cDNA per reaction does slightly increase replicate sample variation (see error bars in Figures S1C, 6B, S2C and S3B, D-F), we found it to be a necessary compromise for assaying gene expression changes during the diauxic shift. Even cycloheximide (CHX) increased TAF10 and UBC6 expression during the mid-log growth phase (Supplementary Figure S8D and S8E) making these relatively stable reference genes less reliable under this treatment condition. We recommend always calculating the transformed Cq values (E-Cq) and normalizing to the reference sample for each gene tested to confirm that the variation among samples, times, and treatments are acceptable, thus validating the use of the specified reference gene(s) during the experiment. |
Plate reader growth curve assay
Strains were inoculated in 200 µL media in sterile 96-well flat-bottom plates, typically, 3–4 biological replicates with two technical replicates for each strain/condition. ‘Blank’ wells contained media/treatments without yeast cells. Plates were loaded into a SpectraMax i3 multimode plate reader. OD600 measurements were taken every 20 min with continuous shaking, linear (medium intensity for 900 s) then orbital (medium intensity for 300 s) between readings. ‘Blank’ well readings were subtracted from culture readings in excel. Data were then transferred to GraphPad Prism for plotting.
Confocal microscopy
600 µL of yeast culture in the mid-log growth phase was transferred to three microcentrifuge tubes and cells pelleted at 6000 × g for 10 min. 580 µL of the supernatant were discarded. DAPI was added at a final concentration of 5 µg/mL. 75 µM inositol was added to one of the tubes and 1 mM VPA to another, leaving one as the control. Tubes were briefly vortexed to mix. 4 µL was placed on a slide, a coverslip secured atop with coverslip sealant, and left to cure/incubate in the dark for 30 min. Images were captured with a Leica SP8 confocal microscope.
Data plotting and statistical analysis programs
GraphPad Prism was used for plotting graphs and statistical analyses. Affinity Designer (a vector editing program) was used to adjust the labelling of the plots generated with Prism, arrange images (WB and confocal), and create figure models.
Data availability
Additional data will be supplied upon request to the corresponding author, Miriam L. Greenberg.
References
Czech, M. P. Dynamics of phosphoinositides in membrane retrieval and insertion. Annu. Rev. Physiol. 65, 791–815. https://doi.org/10.1146/annurev.physiol.65.092101.142522 (2003).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657. https://doi.org/10.1038/nature05185 (2006).
Kim, S., Bhandari, R., Brearley, C. A. & Saiardi, A. The inositol phosphate signalling network in physiology and disease. Trends Biochem. Sci. 49, 969–985. https://doi.org/10.1016/j.tibs.2024.08.005 (2024).
Chhetri, D. R. Myo-inositol and its derivatives: Their emerging role in the treatment of human diseases. Front. Pharmacol. 10, 1172. https://doi.org/10.3389/fphar.2019.01172 (2019).
Irvine, R. F. & Schell, M. J. Back in the water: The return of the inositol phosphates. Nat. Rev. Mol. Cell Biol. 2, 327–338. https://doi.org/10.1038/35073015 (2001).
Shears, S. B. et al. Defining signal transduction by inositol phosphates. Subcell Biochem. 59, 389–412. https://doi.org/10.1007/978-94-007-3015-1_13 (2012).
Tsui, M. M. & York, J. D. Roles of inositol phosphates and inositol pyrophosphates in development, cell signaling and nuclear processes. Adv. Enzyme Regul. 50, 324–337. https://doi.org/10.1016/j.advenzreg.2009.12.002 (2010).
Mikoshiba, K. IP3 receptor/Ca2+ channel: From discovery to new signaling concepts. J. Neurochem. 102, 1426–1446. https://doi.org/10.1111/j.1471-4159.2007.04825.x (2007).
Mikoshiba, K. Role of IP3 receptor signaling in cell functions and diseases. Adv. Biol. Regul. 57, 217–227. https://doi.org/10.1016/j.jbior.2014.10.001 (2015).
Hill-Eubanks, D. C., Werner, M. E., Heppner, T. J. & Nelson, M. T. Calcium signaling in smooth muscle. Cold Spring Harb. Perspect. Biol. 3, a004549. https://doi.org/10.1101/cshperspect.a004549 (2011).
Fearnley, C. J., Roderick, H. L. & Bootman, M. D. Calcium signaling in cardiac myocytes. Cold Spring Harb. Perspect. Biol. 3, a004242. https://doi.org/10.1101/cshperspect.a004242 (2011).
Berridge, M. J. Neuronal calcium signaling. Neuron 21, 13–26. https://doi.org/10.1016/s0896-6273(00)80510-3 (1998).
Chakkour, M. & Greenberg, M. L. Insights into the roles of inositol hexakisphosphate kinase (IP6K1) in mammalian cellular processes. J. Biol. Chem. 300, 107116. https://doi.org/10.1016/j.jbc.2024.107116 (2024).
Chakraborty, A. The inositol pyrophosphate pathway in health and diseases. Biol. Rev. Camb. Philos. Soc. 93, 1203–1227. https://doi.org/10.1111/brv.12392 (2018).
Chakraborty, A., Kim, S. & Snyder, S. H. Inositol pyrophosphates as mammalian cell signals. Sci. Signal 4, re1. https://doi.org/10.1126/scisignal.2001958 (2011).
Chatree, S., Thongmaen, N., Tantivejkul, K., Sitticharoon, C. & Vucenik, I. Role of inositols and inositol phosphates in energy metabolism. Molecules 25, 5079. https://doi.org/10.3390/molecules25215079 (2020).
Thota, S. G. & Bhandari, R. The emerging roles of inositol pyrophosphates in eukaryotic cell physiology. J. Biosci. 40, 593–605. https://doi.org/10.1007/s12038-015-9549-x (2015).
Thomas, M. P., Mills, S. J. & Potter, B. V. The, “Other” inositols and their phosphates: Synthesis, biology, and medicine (with recent advances in myo-inositol chemistry). Angew. Chem. Int. Ed. Engl. 55, 1614–1650. https://doi.org/10.1002/anie.201502227 (2016).
Case, K. C., Salsaa, M., Yu, W. & Greenberg, M. L. Regulation of Inositol Biosynthesis: Balancing Health and Pathophysiology. Handb. Exp. Pharmacol. 259, 221–260. https://doi.org/10.1007/164_2018_181 (2018).
Schneider, S. Inositol transport proteins. FEBS Lett. 589, 1049–1058. https://doi.org/10.1016/j.febslet.2015.03.012 (2015).
Majumder, A. L., Johnson, M. D. & Henry, S. A. 1L-myo-inositol-1-phosphate synthase. Biochim. Biophys. Acta 1348, 245–256. https://doi.org/10.1016/s0005-2760(97)00122-7 (1997).
Geiger, J. H. & Jin, X. The structure and mechanism of myo-inositol-1-phosphate synthase. Subcell Biochem. 39, 157–180. https://doi.org/10.1007/0-387-27600-9_7 (2006).
Case, K. C., Schmidtke, M. W. & Greenberg, M. L. The paradoxical role of inositol in cancer: A consequence of the metabolic state of a tumor. Cancer Metastasis. Rev. 41, 249–254. https://doi.org/10.1007/s10555-022-10032-8 (2022).
Chen, Q., Shen, L. & Li, S. Emerging role of inositol monophosphatase in cancer. Biomed. Pharmacother. 161, 114442. https://doi.org/10.1016/j.biopha.2023.114442 (2023).
Long, L., Huang, Q., Song, T. & Dai, Z. Myo-inositol rescued insulin resistance and dyslipidemia in db/db mice. J. Appl. Biomed. https://doi.org/10.32725/jab.2024.009 (2024).
DiNicolantonio, J. J. & O'Keefe, J. H. Vol. 9 e001989 (Archives of Disease in childhood, 2022).
Health, N. I. o. M. Bipolar Disorder, <https://www.nimh.nih.gov/health/statistics/bipolar-disorder#part_2605> (2007).
Berridge, M. J. Calcium signalling and psychiatric disease: Bipolar disorder and schizophrenia. Cell Tissue Res. 357, 477–492. https://doi.org/10.1007/s00441-014-1806-z (2014).
Lopez-Munoz, F., Shen, W. W., D’Ocon, P., Romero, A. & Alamo, C. A history of the pharmacological treatment of bipolar disorder. Int. J. Mol. Sci. 19, 2143. https://doi.org/10.3390/ijms19072143 (2018).
Tondo, L. et al. Clinical use of lithium salts: guide for users and prescribers. Int. J. Bipolar Disord. 7, 16. https://doi.org/10.1186/s40345-019-0151-2 (2019).
Berridge, M. J., Downes, C. P. & Hanley, M. R. Lithium amplifies agonist-dependent phosphatidylinositol responses in brain and salivary glands. Biochem. J. 206, 587–595. https://doi.org/10.1042/bj2060587 (1982).
Case, K. C., Beltman, R. J., Pflum, M. K. H. & Greenberg, M. L. Valproate regulates inositol synthesis by reducing expression of myo-inositol-3-phosphate synthase. Sci. Rep. 13, 14844. https://doi.org/10.1038/s41598-023-41936-2 (2023).
Lazcano, P., Schmidtke, M. W., Onu, C. J. & Greenberg, M. L. Phosphatidic acid inhibits inositol synthesis by inducing nuclear translocation of kinase IP6K1 and repression of myo-inositol-3-P synthase. J. Biol. Chem 298, 102363. https://doi.org/10.1016/j.jbc.2022.102363 (2022).
Lubrich, B. & van Calker, D. Inhibition of the high affinity myo-inositol transport system: A common mechanism of action of antibipolar drugs?. Neuropsychopharmacology 21, 519–529. https://doi.org/10.1016/S0893-133X(99)00037-8 (1999).
Wolfson, M. et al. Chronic treatment of human astrocytoma cells with lithium, carbamazepine or valproic acid decreases inositol uptake at high inositol concentrations but increases it at low inositol concentrations. Brain Res. 855, 158–161. https://doi.org/10.1016/s0006-8993(99)02371-9 (2000).
Weant, K. A., Brown, B. L., Craycraft, L. K. & Justice, S. B. Valproic acid in the treatment of migraines. Adv. Emerg. Nurs. J. 42, 243–253 (2020).
Singh, D. et al. Hidden pharmacological activities of valproic acid: A new insight. Biomed. Pharmacother. 142, 112021. https://doi.org/10.1016/j.biopha.2021.112021 (2021).
Rahman, M. & Awosika, A. O. Valproic acid, <https://www.statpearls.com/point-of-care/30956> (2023).
Natale, G. et al. Valproate and lithium: Old drugs for new pharmacological approaches in brain tumors?. Cancer Lett. 560, 216125. https://doi.org/10.1016/j.canlet.2023.216125 (2023).
Rafiullah, M. & Siddiqui, K. Pharmacological treatment of diabetic peripheral neuropathy: An update. CNS Neurol. Disord. Drug Targets 21, 884–900. https://doi.org/10.2174/1871527320666210303111939 (2022).
Makkar, R., Behl, T. & Arora, S. Role of HDAC inhibitors in diabetes mellitus. Curr. Res. Transl. Med. 68, 45–50. https://doi.org/10.1016/j.retram.2019.08.001 (2020).
Rakitin, A. Does valproic acid have potential in the treatment of diabetes mellitus?. Front. Endocrinol. (Lausanne) 8, 147. https://doi.org/10.3389/fendo.2017.00147 (2017).
Gottlicher, M. et al. Valproic acid defines a novel class of HDAC inhibitors inducing differentiation of transformed cells. EMBO J. 20, 6969–6978. https://doi.org/10.1093/emboj/20.24.6969 (2001).
Phiel, C. J. et al. Histone deacetylase is a direct target of valproic acid, a potent anticonvulsant, mood stabilizer, and teratogen. J. Biol. Chem 276, 36734–36741. https://doi.org/10.1074/jbc.M101287200 (2001).
Johannessen, C. U. Mechanisms of action of valproate: A commentatory. Neurochem. Int. 37, 103–110. https://doi.org/10.1016/s0197-0186(00)00013-9 (2000).
Shahien, R., Saleh, S. A. & Bowirrat, A. Intravenous sodium valproate aborts migraine headaches rapidly. Acta Neurol. Scand. 123, 257–265. https://doi.org/10.1111/j.1600-0404.2010.01394.x (2011).
Zanatta, G. et al. Valproic acid interactions with the NavMs voltage-gated sodium channel. Proc. Natl. Acad. Sci. USA 116, 26549–26554. https://doi.org/10.1073/pnas.1909696116 (2019).
Tursunov, A. N., Vasilyev, D. S. & Nalivaeva, N. N. Molecular mechanisms of valproic acid action on signalling systems and brain functions. J. Evol. Biochem. Physiol. 59, 1740–1755. https://doi.org/10.1134/s0022093023050228 (2023).
Yu, W. & Greenberg, M. L. Inositol depletion, GSK3 inhibition and bipolar disorder. Fut. Neurol. 11, 135–148. https://doi.org/10.2217/fnl-2016-0003 (2016).
Morris, G. et al. A model of the mitochondrial basis of bipolar disorder. Neurosci. Biobehav. Rev. 74, 1–20. https://doi.org/10.1016/j.neubiorev.2017.01.014 (2017).
Lima, I. V. A. et al. Antidepressant-like effect of valproic acid-Possible involvement of PI3K/Akt/mTOR pathway. Behav. Brain Res. 329, 166–171. https://doi.org/10.1016/j.bbr.2017.04.015 (2017).
Shirra, M. K., Rogers, S. E., Alexander, D. E. & Arndt, K. M. The Snf1 protein kinase and Sit4 protein phosphatase have opposing functions in regulating TATA-binding protein association with the Saccharomyces cerevisiae INO1 promoter. Genetics 169, 1957–1972. https://doi.org/10.1534/genetics.104.038075 (2005).
Kunkel, J., Luo, X. & Capaldi, A. P. Integrated TORC1 and PKA signaling control the temporal activation of glucose-induced gene expression in yeast. Nat. Commun. 10, 3558. https://doi.org/10.1038/s41467-019-11540-y (2019).
Salsaa, M. et al. Valproate activates the Snf1 kinase in Saccharomyces cerevisiae by decreasing the cytosolic pH. J. Biol. Chem 297, 101110. https://doi.org/10.1016/j.jbc.2021.101110 (2021).
Ewald, J. C. How yeast coordinates metabolism, growth and division. Curr. Opin. Microbiol. 45, 1–7. https://doi.org/10.1016/j.mib.2017.12.012 (2018).
Gonzalez, A. & Hall, M. N. Nutrient sensing and TOR signaling in yeast and mammals. EMBO J. 36, 397–408. https://doi.org/10.15252/embj.201696010 (2017).
Gonzalez, A., Hall, M. N., Lin, S. C. & Hardie, D. G. AMPK and TOR: The yin and yang of cellular nutrient sensing and growth control. Cell Metab. 31, 472–492. https://doi.org/10.1016/j.cmet.2020.01.015 (2020).
Hardie, D. G., Ross, F. A. & Hawley, S. A. AMPK: A nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell Biol. 13, 251–262. https://doi.org/10.1038/nrm3311 (2012).
Inoue, Y. & Nomura, W. TOR signaling in budding yeast. Yeast Role Med. Appl. 1, 55–57. https://doi.org/10.5772/intechopen.70784 (2018).
Loewith, R. & Hall, M. N. Target of rapamycin (TOR) in nutrient signaling and growth control. Genetics 189, 1177–1201. https://doi.org/10.1534/genetics.111.133363 (2011).
Wei, Y. & Zheng, X. F. Nutritional control of cell growth via TOR signaling in budding yeast. Methods Mol. Biol. 759, 307–319. https://doi.org/10.1007/978-1-61779-173-4_18 (2011).
Avery, L. B. & Bumpus, N. N. Valproic acid is a novel activator of AMP-activated protein kinase and decreases liver mass, hepatic fat accumulation, and serum glucose in obese mice. Mol Pharmacol 85, 1–10. https://doi.org/10.1124/mol.113.089755 (2014).
Kim, Y.-J., Lee, J., Song, M.-K., Han, T. & Ryu, J.-C. Valproic acid inhibits cell size and cell proliferation by AMPK-mediated mTOR signaling pathway in JEG-3 cells. BioChip J. 7, 267–277. https://doi.org/10.1007/s13206-013-7310-9 (2013).
Chang, C. Y. et al. Valproic acid sensitizes human glioma cells to gefitinib-induced autophagy. IUBMB Life 67, 869–879. https://doi.org/10.1002/iub.1445 (2015).
Bai, X. et al. Valproate induced hepatic steatosis by enhanced fatty acid uptake and triglyceride synthesis. Toxicol. Appl. Pharmacol. 324, 12–25. https://doi.org/10.1016/j.taap.2017.03.022 (2017).
Xia, Q. et al. Valproic acid induces autophagy by suppressing the Akt/mTOR pathway in human prostate cancer cells. Oncol. Lett. 12, 1826–1832. https://doi.org/10.3892/ol.2016.4880 (2016).
Ji, M. M. et al. Induction of autophagy by valproic acid enhanced lymphoma cell chemosensitivity through HDAC-independent and IP3-mediated PRKAA activation. Autophagy 11, 2160–2171. https://doi.org/10.1080/15548627.2015.1082024 (2015).
Han, W. et al. Valproic acid enhanced apoptosis by promoting autophagy Via Akt/mTOR signaling in glioma. Cell Transplant 29, 963689720981878. https://doi.org/10.1177/0963689720981878 (2020).
Fu, J., Shao, C. J., Chen, F. R., Ng, H. K. & Chen, Z. P. Autophagy induced by valproic acid is associated with oxidative stress in glioma cell lines. Neuro Oncol. 12, 328–340. https://doi.org/10.1093/neuonc/nop005 (2010).
Xu, Y. et al. Induction of apoptosis and autophagy in metastatic thyroid cancer cells by valproic acid (VPA). Int. J. Clin. Exp. Pathol. 8, 8291–8297 (2015).
Proske, J. et al. Adaptive immune response to and survival effect of temozolomide- and valproic acid-induced autophagy in glioblastoma. Anticancer Res. 36, 899–905 (2016).
Kawaoka, K. et al. Valproic acid attenuates renal fibrosis through the induction of autophagy. Clin. Exp. Nephrol. 21, 771–780. https://doi.org/10.1007/s10157-016-1365-6 (2017).
Gilardini Montani, M. S. et al. Histone deacetylase inhibitors VPA and TSA induce apoptosis and autophagy in pancreatic cancer cells. Cell Oncol. (Dordr.) 40, 167–180. https://doi.org/10.1007/s13402-017-0314-z (2017).
Sun, J. et al. Valproic acid targets HDAC1/2 and HDAC1/PTEN/Akt signalling to inhibit cell proliferation via the induction of autophagy in gastric cancer. FEBS J. 287, 2118–2133. https://doi.org/10.1111/febs.15122 (2020).
Wang, Y. et al. Valproic acid increased autophagic flux in human multiple myeloma cells in vitro. Biomed. Pharmacother. 127, 110167. https://doi.org/10.1016/j.biopha.2020.110167 (2020).
Gurpur, P. B., Liu, J., Burkin, D. J. & Kaufman, S. J. Valproic acid activates the PI3K/Akt/mTOR pathway in muscle and ameliorates pathology in a mouse model of Duchenne muscular dystrophy. Am. J. Pathol. 174, 999–1008. https://doi.org/10.2353/ajpath.2009.080537 (2009).
Teng, H. F. et al. Valproic acid enhances Oct4 promoter activity through PI3K/Akt/mTOR pathway activated nuclear receptors. Mol. Cell Endocrinol. 383, 147–158. https://doi.org/10.1016/j.mce.2013.12.008 (2014).
Qin, L., Dai, X. & Yin, Y. Valproic acid exposure sequentially activates Wnt and mTOR pathways in rats. Mol. Cell Neurosci. 75, 27–35. https://doi.org/10.1016/j.mcn.2016.06.004 (2016).
Hao, H. H. et al. Valproic acid reduces autophagy and promotes functional recovery after spinal cord injury in rats. Neurosci. Bull. 29, 484–492. https://doi.org/10.1007/s12264-013-1355-6 (2013).
Case, K. C. Valproate-induced inositol depletion is a consequence of reduced mips expression mediated by the henry regulatory circuit, (Wayne State University, 2024).
Henry, S. A., Kohlwein, S. D. & Carman, G. M. Metabolism and regulation of glycerolipids in the yeast Saccharomyces cerevisiae. Genetics 190, 317–349. https://doi.org/10.1534/genetics.111.130286 (2012).
Gonzalez, A. et al. TORC1 promotes phosphorylation of ribosomal protein S6 via the AGC kinase Ypk3 in Saccharomyces cerevisiae. PLoS ONE 10, e0120250. https://doi.org/10.1371/journal.pone.0120250 (2015).
Yerlikaya, S. et al. TORC1 and TORC2 work together to regulate ribosomal protein S6 phosphorylation in Saccharomyces cerevisiae. Mol. Biol. Cell 27, 397–409. https://doi.org/10.1091/mbc.E15-08-0594 (2016).
Cook, M. et al. Anti-seizure therapy with a long-term, implanted intra-cerebroventricular delivery system for drug-resistant epilepsy: A first-in-man study. EClinicalMedicine 22, 100326. https://doi.org/10.1016/j.eclinm.2020.100326 (2020).
Sun, Y. et al. Early post-traumatic seizures are associated with valproic acid plasma concentrations and UGT1A6/CYP2C9 genetic polymorphisms in patients with severe traumatic brain injury. Scand. J. Trauma Resusc. Emerg. Med. 25, 85. https://doi.org/10.1186/s13049-017-0382-0 (2017).
Urban, J. et al. Sch9 is a major target of TORC1 in Saccharomyces cerevisiae. Mol. Cell 26, 663–674. https://doi.org/10.1016/j.molcel.2007.04.020 (2007).
Caligaris, M. et al. The yeast protein kinase Sch9 functions as a central nutrient-responsive hub that calibrates metabolic and stress-related responses. J. Fungi. (Basel) 9, 787. https://doi.org/10.3390/jof9080787 (2023).
Beugnet, A., Tee, A. R., Taylor, P. M. & Proud, C. G. Regulation of targets of mTOR (mammalian target of rapamycin) signalling by intracellular amino acid availability. Biochem. J. 372, 555–566. https://doi.org/10.1042/BJ20021266 (2003).
Murley, A. et al. Sterol transporters at membrane contact sites regulate TORC1 and TORC2 signaling. J. Cell Biol. 216, 2679–2689. https://doi.org/10.1083/jcb.201610032 (2017).
Caligaris, M. & De Virgilio, C. Proxies introduce bias in decoding TORC1 activity. MicroPubl Biol 2024, https://doi.org/10.17912/micropub.biology.001170 (2024).
Caligaris, M. et al. Snf1/AMPK fine-tunes TORC1 signaling in response to glucose starvation. Elife 12, e84319. https://doi.org/10.7554/eLife.84319 (2023).
Leskoske, K. L., Roelants, F. M., Martinez Marshall, M. N., Hill, J. M. & Thorner, J. The stress-sensing TORC2 complex activates yeast AGC-family protein kinase Ypk1 at multiple novel sites. Genetics 207, 179–195. https://doi.org/10.1534/genetics.117.1124 (2017).
Kumar, P., Awasthi, A., Nain, V., Issac, B. & Puria, R. Novel insights into TOR signalling in Saccharomyces cerevisiae through Torin2. Gene 669, 15–27. https://doi.org/10.1016/j.gene.2018.05.081 (2018).
Sarbassov, D. D. et al. Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB. Mol. Cell 22, 159–168. https://doi.org/10.1016/j.molcel.2006.03.029 (2006).
Schreiber, K. H. et al. Rapamycin-mediated mTORC2 inhibition is determined by the relative expression of FK506-binding proteins. Aging Cell 14, 265–273. https://doi.org/10.1111/acel.12313 (2015).
Kira, S. et al. Dynamic relocation of the TORC1-Gtr1/2-Ego1/2/3 complex is regulated by Gtr1 and Gtr2. Mol. Biol. Cell 27, 382–396. https://doi.org/10.1091/mbc.E15-07-0470 (2016).
Sanz, P., Viana, R. & Garcia-Gimeno, M. A. AMPK in yeast: The SNF1 (Sucrose Non-fermenting 1) protein kinase complex. Exp. Suppl. 107, 353–374. https://doi.org/10.1007/978-3-319-43589-3_14 (2016).
Zhang, J. et al. Mapping the interaction of Snf1 with TORC1 in Saccharomyces cerevisiae. Mol. Syst. Biol. 7, 545. https://doi.org/10.1038/msb.2011.80 (2011).
Hughes Hallett, J. E., Luo, X. & Capaldi, A. P. State transitions in the TORC1 signaling pathway and information processing in Saccharomyces cerevisiae. Genetics 198, 773–786. https://doi.org/10.1534/genetics.114.168369 (2014).
Hughes Hallett, J. E., Luo, X. & Capaldi, A. P. Snf1/AMPK promotes the formation of Kog1/Raptor-bodies to increase the activation threshold of TORC1 in budding yeast. Elife 4, e09181. https://doi.org/10.7554/eLife.09181 (2015).
Orlova, M., Kanter, E., Krakovich, D. & Kuchin, S. Nitrogen availability and TOR regulate the Snf1 protein kinase in Saccharomyces cerevisiae. Eukaryot. Cell 5, 1831–1837 (2006).
McCartney, R. R. & Schmidt, M. C. Regulation of Snf1 kinase. Activation requires phosphorylation of threonine 210 by an upstream kinase as well as a distinct step mediated by the Snf4 subunit. J. Biol. Chem. 276, 36460–36466. https://doi.org/10.1074/jbc.M104418200 (2001).
Orlova, M., Barrett, L. & Kuchin, S. Detection of endogenous Snf1 and its activation state: Application to Saccharomyces and Candida species. Yeast 25, 745–754. https://doi.org/10.1002/yea.1628 (2008).
Hofbauer, H. F. et al. Regulation of gene expression through a transcriptional repressor that senses acyl-chain length in membrane phospholipids. Dev. Cell 29, 729–739. https://doi.org/10.1016/j.devcel.2014.04.025 (2014).
Shirra, M. K. et al. Inhibition of acetyl coenzyme A carboxylase activity restores expression of the INO1 gene in a snf1 mutant strain of Saccharomyces cerevisiae. Mol. Cell Biol. 21, 5710–5722. https://doi.org/10.1128/MCB.21.17.5710-5722.2001 (2001).
Estruch, F., Treitel, M. A., Yang, X. & Carlson, M. N-terminal mutations modulate yeast SNF1 protein kinase function. Genetics 132, 639–650. https://doi.org/10.1093/genetics/132.3.639 (1992).
Portillo, F., Mulet, J. M. & Serrano, R. A role for the non-phosphorylated form of yeast Snf1: Tolerance to toxic cations and activation of potassium transport. FEBS Lett. 579, 512–516. https://doi.org/10.1016/j.febslet.2004.12.019 (2005).
Shinoda, J. & Kikuchi, Y. Rod1, an arrestin-related protein, is phosphorylated by Snf1-kinase in. Biochem. Biophys. Res. Commun. 364, 258–263. https://doi.org/10.1016/j.bbrc.2007.09.134 (2007).
Wade, S. L., Poorey, K., Bekiranov, S. & Auble, D. T. The Snf1 kinase and proteasome-associated Rad23 regulate UV-responsive gene expression. EMBO J. 28, 2919–2931. https://doi.org/10.1038/emboj.2009.229 (2009).
Wilson, W. A., Hawley, S. A. & Hardie, D. G. Glucose repression/derepression in budding yeast: SNF1 protein kinase is activated by phosphorylation under derepressing conditions, and this correlates with a high AMP:ATP ratio. Curr. Biol. 6, 1426–1434. https://doi.org/10.1016/s0960-9822(96)00747-6 (1996).
Hirschhorn, J. N., Brown, S. A., Clark, C. D. & Winston, F. Evidence that SNF2/SWI2 and SNF5 activate transcription in yeast by altering chromatin structure. Genes Dev. 6, 2288–2298. https://doi.org/10.1101/gad.6.12a.2288 (1992).
Villa-Garcia, M. J. et al. Genome-wide screen for inositol auxotrophy in Saccharomyces cerevisiae implicates lipid metabolism in stress response signaling. Mol. Genet. Genom. 285, 125–149. https://doi.org/10.1007/s00438-010-0592-x (2011).
Bozaquel-Morais, B. L., Madeira, J. B., Maya-Monteiro, C. M., Masuda, C. A. & Montero-Lomeli, M. A new fluorescence-based method identifies protein phosphatases regulating lipid droplet metabolism. PLoS ONE 5, e13692. https://doi.org/10.1371/journal.pone.0013692 (2010).
Madeira, J. B. et al. TORC1 inhibition induces lipid droplet replenishment in yeast. Mol. Cell Biol. 35, 737–746. https://doi.org/10.1128/MCB.01314-14 (2015).
Madeira, J. B. et al. Induction of triacylglycerol synthesis in yeast by cell cycle arrest. FEMS Yeast Res. 19, foz030. https://doi.org/10.1093/femsyr/foz030 (2019).
Golla, U., Joseph, D. & Tomar, R. S. Combined transcriptomics and chemical-genetics reveal molecular mode of action of valproic acid, an anticancer molecule using budding yeast model. Sci. Rep. 6, 35322. https://doi.org/10.1038/srep35322 (2016).
Jadhav, S. et al. Valproate induces the unfolded protein response by increasing ceramide levels. J. Biol. Chem. 291, 22253–22261. https://doi.org/10.1074/jbc.M116.752634 (2016).
Yu, W., Ye, C. & Greenberg, M. L. Inositol hexakisphosphate kinase 1 (IP6K1) regulates inositol synthesis in mammalian cells. J. Biol. Chem. 291, 10437–10444. https://doi.org/10.1074/jbc.M116.714816 (2016).
Salsaa, M., Case, K. & Greenberg, M. L. Orchestrating phospholipid biosynthesis: Phosphatidic acid conducts and Opi1p performs. J. Biol. Chem. 292, 18729–18730. https://doi.org/10.1074/jbc.H117.809970 (2017).
Ju, S. & Greenberg, M. L. Valproate disrupts regulation of inositol responsive genes and alters regulation of phospholipid biosynthesis. Mol. Microbiol. 49, 1595–1603. https://doi.org/10.1046/j.1365-2958.2003.03641.x (2003).
Vaden, D. L., Ding, D., Peterson, B. & Greenberg, M. L. Lithium and valproate decrease inositol mass and increase expression of the yeast INO1 and INO2 genes for inositol biosynthesis. J. Biol. Chem. 276, 15466–15471. https://doi.org/10.1074/jbc.M004179200 (2001).
Choo, A. Y. & Blenis, J. Not all substrates are treated equally: Implications for mTOR, rapamycin-resistance and cancer therapy. Cell Cycle 8, 567–572. https://doi.org/10.4161/cc.8.4.7659 (2009).
Evans, S. K., Burgess, K. E. V. & Gray, J. V. Recovery from rapamycin: Drug-insensitive activity of yeast target of rapamycin complex 1 (TORC1) supports residual proliferation that dilutes rapamycin among progeny cells. J. Biol. Chem. 289, 26554–26565. https://doi.org/10.1074/jbc.M114.589754 (2014).
Feldman, M. E. et al. Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2. PLoS Biol. 7, e38. https://doi.org/10.1371/journal.pbio.1000038 (2009).
Montero, J. C., Chen, X., Ocana, A. & Pandiella, A. Predominance of mTORC1 over mTORC2 in the regulation of proliferation of ovarian cancer cells: Therapeutic implications. Mol. Cancer Ther. 11, 1342–1352. https://doi.org/10.1158/1535-7163.MCT-11-0723 (2012).
Azab, A. N., Ishak, J. F., Kaplanski, J., Delbar, V. & Greenberg, M. L. Mechanisms of Action of the Mood Stabilizer Valproate: A Focus on GSK-3 Inhibition. Fut. Neurol. 3, 433–445. https://doi.org/10.2217/14796708.3.4.433 (2008).
Hofbauer, H. F. et al. The molecular recognition of phosphatidic acid by an amphipathic helix in Opi1. J. Cell Biol. 217, 3109–3126. https://doi.org/10.1083/jcb.201802027 (2018).
Looke, M., Kristjuhan, K. & Kristjuhan, A. Extraction of genomic DNA from yeasts for PCR-based applications. Biotechniques 50, 325–328. https://doi.org/10.2144/000113672 (2011).
Becker, D. M. & Guarente, L. Methods in enzymology, Vol. 194 [12] High-efficiency transformation of yeast by electroporation, 182–187 (Elsevier, 1991).
Bustin, S. & Huggett, J. qPCR primer design revisited. Biomol. Detect. Quantif. 14, 19–28. https://doi.org/10.1016/j.bdq.2017.11.001 (2017).
Teste, M. A., Duquenne, M., Francois, J. M. & Parrou, J. L. Validation of reference genes for quantitative expression analysis by real-time RT-PCR in Saccharomyces cerevisiae. BMC Mol. Biol. 10, 99. https://doi.org/10.1186/1471-2199-10-99 (2009).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45. https://doi.org/10.1093/nar/29.9.e45 (2001).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Bio-Rad. BioRadiations (Bio-Rad Laboratories, Inc., 2007).
Acknowledgements
Thank you to Sarah Palm for finding and sending us the pRS315-YPK15A-myc expression plasmid; Dr. Jeremy W. Thorner (UC Berkeley) for graciously providing advice; Dr. Robert Ernst (Universität des Saarlandes) for providing us with the Opi1-mGFP expression strain; Marco Caligaris (Université de Fribourg), for kindly providing guidance and insight for the Sch9 phosphorylation analyses; and Dr. Claudio De Virgilio (Université de Fribourg) for providing us custom made Sch9-pThr737 antibodies. This work was supported by NIH grants R01 GM125082 and R35 GM149271 (to M.L.G.), and T32 HL120822 (providing support to K.C.C. through the Detroit Cardiovascular Training Program at Wayne State University).
Author information
Authors and Affiliations
Contributions
Conceptualization, K.C.C., M.L.G.; methodology, validation, investigation, K.C.C., L.O.; formal analysis, K.C.C.; resources, M.L.G.; data curation, K.C.C.; writing—original draft, K.C.C.; writing—review and editing, K.C.C., M.L.G., L.O.; visualization, K.C.C.; supervision, M.L.G.; funding acquisition, M.L.G., K.C.C.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Case, K.C., Orkun, L. & Greenberg, M.L. Valproate independently activates Snf1, inhibits TORC1, and induces repression of INO1 transcription by increasing nuclear localization of Opi1. Sci Rep 15, 24601 (2025). https://doi.org/10.1038/s41598-025-07540-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07540-2