Abstract
2D/3D medical image registration is crucial for image-guided surgery and medical research. Single-view methods often lack depth information, while multiview approaches typically rely on large-scale training datasets, which may not always be available in clinical scenarios. This paper explores the feasibility of multiview registration with insufficient data. We propose an innovative multiview 2D/3D image registration model with three key components: a points of interest (POIs) tracking network based on style transfer (POITT), a triangulation layer, and shape alignment based on branch and bound (SA-BnB). POITT constructs a POIs tracking network on Digital Reconstructed Radiographs images (DRRs) and performs points matching on X-ray images via style transfer. The triangulation layer establishes high-precision spatial mapping relationships between X-ray images and the patient. SA-BnB calculates the optimal transformation matrix between the CT volume and the patient. We collected clinical X-ray and CT data from 8 patients and generated 300 pairs of simulated DRRs from two views for each patient to validate the proposed model. The average fiducial registration error was reduced to 6 mm (less than 10 mm is considered successful) with a 96.67\(\%\) average registration success rate on DRRs. The average SSIM reached 0.77 between X-ray images and the aligned DRRs. All experimental results demonstrate significant advantages over traditional 2D/3D registration. This study may provide a reliable solution for minimally invasive pelvic surgery navigation. The key code is available at https://github.com/TUYaYa1/Points-of-Interest-tracking-network.
Similar content being viewed by others
Introduction
Pelvic fractures are serious traumas that involve complex orthopedic surgeries, which have always posed a huge challenge for orthopedic surgeons. Traditional open surgery often results in significant intra-operative trauma and slow recovery. With the development of medical technology, minimally invasive pelvic surgery has gradually emerged as a promising field for the treatment of pelvic fractures due to its low invasiveness and fast recovery1. 2D/3D registration is a crucial technology that provides real-time and accurate 3D positioning for minimally invasive pelvic surgery, aiming to find an optimal 3D transformation that minimizes misalignment between 2D images and 3D volumes. In pelvic surgery, CT is usually the preferred 3D pre-operative data, and DRRs can be generated from CT using ray casting. Therefore, 2D/3D registration can be achieved by directly comparing the X-ray image and the generated DRR.
Traditional 2D/3D registrations are usually solved by optimization-based approaches2. These approaches3,4 have iteratively searched for the 3D pose that best matches the similarity score between DRR and X-ray image. Thus a large number of paired DRR and X-ray images are required during the iterative optimization process, which leads to the approach slow in terms of computation. In addition, the optimization-based methods rely on a good initial posture. Otherwise, the method may be have difficulty converging to the correct position, and resulting in registration failure. With the rapid development of deep learning technology, several learning-based strategies have been developed to 2D/3D registration5,6,7,8,9,10. Learning-based methods utilize the learning ability of convolutional neural networks (CNNs) to directly estimate the pose of 3D data. These methods have the advantages of computational efficiency, large capture range, and good registration performance. However, they still have two limitations. First, a large amount of data is required for training. When the labeled X-ray images used for training are limited, the he model’s generalization ability is significantly reduced. Second, most methods commonly utilize single view images for registration. Due to the inherent limitations of single-view registration, there may be some ambiguities in the registration process.
To improve navigation accuracy, the multiview 2D/3D medical image registration technology has recently emerged. This technology provides more comprehensive and accurate 3D anatomical structure information by aligning medical image data from different views and modalities. Unlike the inherent challenges in single-view registration, this method considers image depth information and integrates image information from different view, providing a potential solution for improving navigation accuracy. However, recent multiview 2D/3D registration methods are limited when the paired DRRs and X-ray images are insufficient. Therefore, the main challenges currently faced are:
-
Instability of single-view registration Single-view registration lacks depth and geometric constraints, leading to unstable results and lower accuracy.
-
Data requirement for multiview registration Multiview registration requires large amounts of labeled data, which is difficult to obtain in clinical settings.
-
Balancing accuracy and computational time There is a trade-off between accuracy and computation time, making it hard to achieve the right balance for real-time clinical applications.
To address the above drawbacks, we propose an innovative multiview 2D/3D image registration method based on style transfer, which includes POITT, triangulation layer, and SA-BnB. POITT establishes point-to-point consistency matching relationships in DRRs and X-ray images by automatically detecting and tracking key interest points. The triangulation layer projects the tracked POIs in X-ray images from multiviews back into the patient’s 3D space. SA-BnB directly aligns 3D CT with patient by searching for the optimal point pair matching. The contributions of this paper are as follows:
-
A novel lightweight POIs tracking network is proposed Combining U-Net and convolutional block attention module (CBAM) for improved accuracy and robustness of POIs tracking. The simple and clear architecture provide an efficient and lightweight solution for POIs tracking for practical application scenarios.
-
An effective solution for addressing insufficient data is proposed based on style transfer Unifying the style between DRRs and X-ray images for effective POIs tracking. This improves model performance on real X-ray images and offers a new method for handling limited data in 2D/3D registration.
-
A more precise BnB shape alignment method is introduced By leveraging a global optimization strategy, an optimal rigid transformation between point sets is efficiently searched, enabling accurate alignment of key structures and improving overall registration accuracy.
Related work
2D/3D registration can be categorized into optimization-based methods and learning-based methods. Optimization-based methods usually use iterative matching strategies to repeatedly sample and generate DRRs, which result in high computational costs and low accuracy. To avoid these problems, several efficient works have been proposed in hardware-level or software-level. Chang et al.3 focused on 2D/3D registration of spinal images and compared the effects of five similarity measures in three optimization algorithms with any combination. Ghafurian et al.4 innovatively used weighted histograms of image gradient directions as image features to calculate image similarity, significantly reducing the number of DRR generation and effectively reducing computational costs. Mathias et al.11 constructed the DeepDRR framework, which can quickly and automatically simulate X-ray images from CT volume data. Meanwhile, DeepDRR as a universal model can be applied to various clinical data. Although these methods have minimized the generation time of DRR as much as possible, the overall registration time was still relatively long, reaching up to 16 seconds in some cases3.
The learning-based methods involve CNNs and deep learning techniques, providing new ideas for image registration. Unberath et al.12 provided a comprehensive review of machine learning in 2D/3D registration for image-guided interventions, highlighting the potential of machine learning-based methods to address some of the notorious challenges in 2D/3D registration. The ReMIND2Reg challenge13 at the MICCAI conference involves 2D-to-3D registration tasks and provides various registration methods. Wu et al.5 used CNN to automatically learn image features and evaluated image similarity by combining subspace parameters to obtain registration parameters. Miao et al.6,7 regarded the registration problem as a regression problem and proposed a real-time 2D/3D medical image registration method based on CNN regression for the first time. They proposed three algorithmic strategies to build regressors and predicted six transformation parameters. However, the model adopted a simple and shallow network structure; therefore, its fitting and generalization abilities can still be improved in the future. Wang et al.8 proposed a self-supervised learning network based on Densnet, which achieved direct regression for 2D/3D registration, significantly improving accuracy and speed. When processing large amounts of data, the network fitting was insufficient. Chen9 regarded the registration problem as a classification problem and constructed a registration model using Google-Net-Inception V3, which can quickly complete registration. While its output transformation parameters were limited with integers, making it more suitable for coarse registration tasks. Gao et al.10 proposed the ProST module for generating DRRs and applied it to 2D/3D medical image registration to achieve end-to-end registration models. However, the translation along the depth direction was less accurate than along other directions. In addition, considering the potential issues associated with single-view registration, Liao et al.14 proposed a multi-view 2D/3D rigid registration model based on interest point tracking and triangulation (Point\(^2\)). This model directly aligned 3D volume by tracking the interest points from DRRs to X-ray images, avoiding out of plane errors. However, the model required numerous data to train, and the insufficient paired DRR and X-ray images would lead to registration failure. Wang et al.15 improved the point\(^2\) by introducing domain adaptation, resulting in slightly better performance on limited data. However, the feature transfer method remained simple, and its generalization ability still needed improvement. At the same time, the style transfer method have handled only for paired DRRs and X-ray images, which limited its application.
Materials and methods
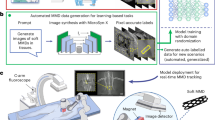
Figure 1 illustrates the overall architecture of the proposed method, which consists of three key modules: POITT, triangulation layer, and SA-BnB. First, the POITT constructs a POIs tracking network on DRRs and leverages style transfer techniques to unify the appearance of X-ray images and DRRs. This enables precise feature point matching in X-ray images. Next, the triangulation layer is introduced after the POITT to establish a high-precision spatial mapping between X-ray images and the patient’s anatomical structures. This mapping allows for the accurate identification of POIs on the patient’s anatomy. Finally, the SA-BnB employs an optimization algorithm to compute the optimal transformation matrix between the CT volume and the patient’s anatomy, ensuring precise image alignment.
The study was a medical imaging investigation approved by the Ethics Committee of the Chinese PLA General Hospital (No.S2022-781-01). Due to its retrospective nature and the fact that the patients’ data were anonymous, a waiver of patients’ informed consent was granted by the Ethics Committee of the Chinese PLA General Hospital. We confirm that all experiments were performed in accordance with relevant guidelines and regulations.
Overview of the proposed method.
The POIs tracking network based on style transfer (POITT)
POIs tracking The primary objective of the POITT is to achieve precise POIs mapping and tracking from DRR to X-ray images. However, acquiring multi-view X-ray data is challenging, and manually annotating POIs is both time-consuming and labor-intensive, resulting in a limited number of X-ray datasets with precise POIs annotations. To address this issue, we utilize DRR as training data and construct the POIs tracking network using synthetic images from different viewpoints. This approach enables the model to learn the feature mapping from DRR to X-ray images, thereby improving the accuracy and robustness of POIs tracking.
Specifically, selecting a set of 3D POIs \(\{X^{ct}_1,X^{ct}_2...X^{ct}_N\}\) on CT volume, where the corresponding DRR(\(I^D\)) and 2D POIs can be obtained through 3D-to-2D projection and we denote the set as \(\varvec{x}^D=\{x^{D}_1,x^{D}_2...x^{D}_N\}\), which can be converted into a sets of heatmaps \(\{M^D_1,M^D_2...M^D_N\}\) using a Gaussian function. The network takes DRRs as input and heatmaps of 2D POIs as label. And the outputs are the tracked POIs on test DRR in the form of heatmaps \(\{ \hat{M}^D_1,\hat{M}^D_2...\hat{M}^D_N\}\).
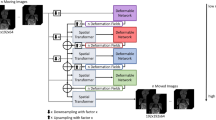
The structure of the POI tracking network is shown in Fig. 2. We use U-Net16 as the basic architecture and integrate CBAM17 to the network. Selecting a view, we take DRRs in this view as input and perform fine-grained feature extraction at the pixel level. To guide the network to focus on key areas of the image, we construct a multi-scale mixed attention mechanisms to reduce interference from irrelevant information. The heatmaps of 2D POIs on DRRs are labels during the training process. The output of the network are estimated heatmaps with the same resolution as the input DRRs. For an image with size \(M \times N\), the size of heatmap is \(M \times N \times C\), where C is the number of channels, equating to the amount of POIs.
Specifically, assuming given transformation matrix R and vector t from m views, we can capture m-view POIs sets in DRR and train m POIs tracking networks from different views. Each of the networks will track POIs for the designed view. Given the sets of X-ray images \(\{I^X_1,I^X_2...I^X_m\}\) from the m views, these networks outputs the tracked X-ray POIs of each view in the form of heatmaps. In general, we can locate the points in three-dimensional space by images from two views. Thus m can be set to 2 in the experiments. As shown in the Fig. 1, two views of anterior-posterior and inlet are selected to construct POI tracking networks. However, the X-ray images obtained in the pelvics surgery usually are significantly different from simulated DRRs, thus we introduce style transfer using CycleGAN18 to reduce the style differences between these two images.
BCELoss has be used as the loss function that regulates the training of POIs tracking network:
where \(M^D_i\) is the ground truth heatmap, BCE is the pixel-wise binary cross entropy function and \(\sigma\) is the sigmoid function.
The architecture of the POI tracking network.
Style transfer Due to significant differences in the generation methods and imaging characteristics of DRR and X-ray images, there is a style discrepancy between them, which often leads to lower accuracy when applying a network trained on DRR to X-ray images. Therefore, CycleGAN is introduced for style transfer, aligning the styles of DRR and X-ray images, thereby improving the accuracy of interest point tracking on X-ray images. Then the POI tracking network takes the two-view X-ray images being consistent with the DRR’s style by style transfer as the input and outputs the heatmaps \(\{\hat{M}^X_1,\hat{M}^X_2...\hat{M}^X_N\}\) in each view.
Triangulation layer
After inputting the two views of X-ray images into the POITT, we can obtain the corresponding heatmaps of the POIs for each image. Then, based on 1, we calculate the POIs coordinates corresponding to the X-ray image.
Where \(\{\hat{M}_{1n}^{X},\hat{M}_{2n}^{X}...\hat{M}_{mn}^{X}\}\) being a set of heatmaps from different views, \(\hat{M}_{mn}^{X}\)is the heatmap of the n-th interest point from the m-th view.
After obtaining the POIs from X-ray images from two views, we use the triangulation layer to locate the corresponding 3D points on CT. Specifically, the triangulation layer leverages the POIs’s coordinates from multiple views and performs 3D reconstruction based on geometric relations. This approach enables accurate conversion from 2D POIs to 3D space points.
Detailed triangulation processes and associated mathematical derivations can be found in the supplementary materials.
Shape alignment based on branch and bound (SA-BnB)
We design the BnB mechanism19 to search for shape alignment. Specifically, assuming \(X^{ct} = \{X_1^{ct},X_2^{ct}...X_N^{ct}\}\) being the initial POIs selected from CT and \(\hat{X}^p = {(\hat{X}_1^p,\hat{X}_2^p...\hat{X}_N^p)}\) being the estimated 3D POIs on the patient, we can calculate the optimal transformation matrix \(T^*\) by aligning these points. Thus, the optimization objective is,
As the transformation matrix \(T^*\) for rigid motion can be parameterized by \(R\in SO(3)\) and \(t \in R^3\), thus we can further derive (2) to the following objective function to ensure that the transformed \(X^{ct}\) is closely aligned with \(\hat{X}^p\). Then, the lower bound can be calculated by the following function,
To speed up the calculation, the upper bound is set to the average value between the centroids distance of the points set and the bounding box distance,
where \(C_{X^{ct}}\) and \(C_{\hat{X}^p}\) are the centers of point sets \(X^{ct}\) and \(\hat{X}^p\). \(B_{X^{ct}}\) and \(B_{\hat{X}^p}\) are the bounding box of point sets \(X^{ct}\) and \(\hat{X}^p\), which are defined as the maximum and minimum coordinates of the points set. The target of search process is to find the optimal transformation matrix of \(X^{ct}\) transfer to \(\hat{X}^p\), where \(X^{ct}\) is the source points set and \(\hat{X}^p\) is the target points set. The specific implementation process of SA-BnB is list in Algorithm 1:
Shape alignment based on branch and bound (SA-BnB)
Experiments and results
Dataset and implementation details
The exprimental dataset is collected from the fourth medical center of Chinese PLA General Hospital, including CT scan data and X-ray images from 8 patients. The two views are acquired from patient’s pelvic anterior-posterior view and inlet view. To verify the performance of the proposed model, we conduct POIs tracking and registration experiments on synthetic DRRs and real X-ray images, respectively. For the experiments on DRRs, we randomly generate 3000 paired DRRs from two views per patient using the DeepDRR. The specific parameter settings can be found in the supplementary materials. For the training of POITT, 2700 DRRs are randomly selected for training, and the remaining 300 images are used for testing. For the experiments on X-ray images, we utilize 8 real intraoperative X-ray images to verify the POIs tracking and registration results.
The detailed software and hardware specifications used for running the experiments are provided in the supplementary materials.
Evaluation metrics
To quantitatively analyze the results of POIs tracking, we use Mean Point Drift (mPD) to measure the tracking error. mPD typically calculates the average Euclidean distance between the tracked POIs and the true points.
To quantitatively analyze the registration performance, we use four evaluation metrics: average registration error of fiducial points (mFRE), gross failure rate (GFR), average transformation error (mTME) and average registration time (mReg.time). mFRE is the average distance between patient 3D POIs of patient and aligned CT. GFR is the percentage of FRE greater than 10mm (Registration fails when FRE is greater than 10 mm14,20). mTME is the errors between predicted transformation and actual transformation parameters. Moreover, in order to present the registration results more intuitively, we use normalized cross correlation (NCC) and structure similarity index measure (SSIM) to evaluate the registration results between aligned DRR and the original X-ray iamge. For registration on real X-ray images, only SSIM and NCC are used as metrics due to the lack of real labels. Additionally, we calculated the 95\(\%\) confidence intervals of the results using a two-sample t-distribution method for the difference of means, and employed the Wilcoxon rank-sum test to compute p-values for further evaluation of the model performance differences.
The results of POIs tracking
Comparison results of 2D POIs tracking. (a) DRRs with ground truth POIs. (b) predicted POIs of POITT. (c) predicted POIs of Landmarks Detect.
To demonstrate the advantages of our POITT, we compared it with a classical X-ray POIs detection method, referred to as “Landmarks Detect”. Fig. 3 and Table 1 present the visual and quantitative results, respectively. (a) shows the DRRs with ground truth POIs, (b) shows the predicted POIs of POITT, and (c) shows the predicted POIs of the Landmarks Detect method. The numbers below (b) and (c) show the mPD between the predicted points and the ground truth points. From the figure, it is clear that both methods demonstrate high accuracy in POIs tracking. Table 1 shows the mPD and tracking time for the results. As seen in the table, the accuracy difference between the two methods is negligible, with our method achieving slightly lower mPD than the comparison method. However, the tracking time for our method is significantly shorter, with an average of only 0.14s, compared to 0.71s for the comparison method, which is nearly 5 times slower. Additionally, POITT achieved a smaller standard deviation and 95% confidence interval compared to Landmarks Detect, indicating that our model is more stable in performance. Therefore, our method shows a distinct advantage in POIs tracking, especially in terms of high tracking efficiency, which is crucial for fast registration.
The results of 2D/3D registration on synthetic DRRs
To validate the effectiveness of our model, we compare it with several representative methods: optimization-based methods21, learning-based methods6, and other multiview registration methods14,15,22. Specfic results are shown in Tables 2 and 4.
Visualization of synthetic DRRs registration comparison.
The registration results are presented in Table 2, which provides a quantitative comparison of different registration methods on synthesized DRRs. The experimental results indicate that optimization-based methods yield the poorest registration accuracy while also incurring the longest computation time. In contrast, learning-based methods achieve improved registration performance with an average processing time of only 1.74s per image. However, these methods suffer from a high registration failure rate of 16.4%, making them unsuitable for clinical applications requiring high precision. As a multi-view registration approach, Point\(^2\) demonstrates notable improvements in registration accuracy. However, this model heavily relies on large-scale training data, and due to the limited dataset used in this study, it fails to reach its optimal performance. The transfer learning-based method incorporates a transfer learning module to mitigate the issue of data inefficiency, leading to better registration performance on the small-scale dataset used in our experiment. Nevertheless, it still falls short when compared to the proposed method, such as on severely misaligned pelvic structures (Case 4 in Table 4). Our method achieves the best registration results across all key metrics, reducing mTRE, GFR, and mTME to 6.19mm, 3.33%, and 0.0655mm, respectively. However, our method only achieved a registration time of 3.12s, which is significantly faster than Opt-based and the BnB methods, but still slower than learning-based methods such as Point\(^2\) (0.78s) and Transfer-learning (0.52s). This indicates that our method achieves a balance between accuracy and efficiency, but there is still room for improvement in runtime performance.
Table 3 presents a comparative analysis of different methods in 2D/3D registration, including the 95% confidence intervals (CIs) for mFRE, mTME, and mReg.time, along with the P-values from Wilcoxon rank-sum tests conducted on the mTME to assess statistical significance between our proposed method and competing approaches. In conjunction with the results from Table 2, it is evident that our method achieves the smallest standard deviation and the narrowest confidence interval for both mFRE and mTME, indicating its superior stability in the registration task. Notably, our method achieves an exceptionally low mTME CI of (0.05, 0.08), significantly outperforming the learning-based method (0.61, 1.13) and the multi-view registration method Point\(^2\) (0.13, 0.27), reflecting a substantial reduction in registration errors and an overall improvement in accuracy. While some overlap in CIs is observed in the mFRE, this does not compromise the statistical significance of our method’s superiority in mTME. Since the Wilcoxon rank-sum test is specifically performed on mTME and all P-values are below 0.05, the performance difference in this key metric is statistically significant, providing strong evidence of the accuracy advantage of our method. In terms of computational efficiency, our method achieves a registration time CI of (2.77, 3.47) seconds, which is slightly higher than that of Point\(^2\) (0.68, 0.88) seconds and the transfer learning-based method (0.45, 0.59) seconds. In future work, we will further explore model lightweighting and efficient inference strategies to improve registration speed while maintaining accuracy.
Figure 4 presents the visualization results of different methods on synthetic DRRs, where each row represents a test instance. The comparison clearly demonstrates that our proposed method achieves the closest alignment with the ground truth, exhibiting the highest accuracy. This strongly validates the effectiveness and superiority of our method in the registration task. In contrast, other methods exhibit varying degrees of misalignment, failing to achieve the same level of precision as our approach, further highlighting its advantages. To more clearly demonstrate the advantages of our method, Supplementary Figure 3 presents the real images and corresponding registration error heatmaps for different approaches. As shown in the figure, our method significantly outperforms the other compared methods in terms of registration accuracy and error heatmap quality. It effectively aligns key regions of the pelvis with a uniform distribution of registration errors, further validating the superiority and robustness of our approach.
In summary, our proposed method significantly outperforms other competing methods in both registration accuracy and error heatmap performance, achieving precise alignment of key pelvic regions with a uniform distribution of registration errors. This further validates the superiority and robustness of our method, demonstrating its high reliability and stability in practical applications.
The results of 2D/3D registration on real X-ray images
POI tracking and 2D/3D registration results. (a) DRR with POI. (b) X-ray image with tracked POI. (c) Aligned DRR.
This section further validates the registration performance of the proposed method on clinical real X-ray images. Due to the significant style differences between DRR and real X-ray images, directly applying a model trained on DRR to X-ray images may lead to a loss in registration accuracy. To address this issue, we introduce CycleGAN for style transfer, aligning the appearance of X-ray images with that of DRRs, thereby improving the model’s adaptability and enhancing registration accuracy.
Table 4 lists the quantitative registration results for five test cases. Since real X-ray data lack ground truth labels, we evaluate registration accuracy using structural similarity index (SSIM) and normalized cross-correlation (NCC) between the registered CT projection and the X-ray image.
The results indicate that both optimization-based and learning-based methods fail to achieve the level of accuracy required for surgical navigation. The Point\(^2\) shows improved registration performance, achieving relatively higher values in SSIM and NCC. However, it heavily relies on large-scale multi-view paired data for training, and due to the limited size of our dataset, its performance does not reach its full potential. Furthermore, while the BnB method supports multi-view registration, it requires 22 sec per case, and its accuracy does not provide a significant advantage. The transfer learning-based approach, which incorporates a VGG-based transfer learning network, successfully mitigates the issue of limited training data and achieves better registration results than our method in Case 3 and Case 5. However, due to the simplicity of the VGG architecture, its performance deteriorates when handling complex misalignment cases (e.g., Case 4), leading to a reduction in registration accuracy. In contrast, our method outperforms the transfer learning-based approach in most test cases. In the Anterior-posterior view, our approach achieves an average NCC of 0.6535 and SSIM of 0.7823, while in the Inlet view, NCC and SSIM reach 0.5841 and 0.7591, respectively, demonstrating a significant improvement in registration accuracy. It is worth noting that the CIs of our method partially overlap with those of the comparison methods. For example, when compared with the transfer-learning method, the NCC intervals are (0.5801,0.7269) vs. (0.5592,0.6738), indicating that in some cases, the performance differences may not be statistically significant. However, based on the average performance and consistency, our method shows a clear advantage in most cases, suggesting stronger robustness and generalization capability.
To provide a more intuitive visualization of the registration performance, Fig. 5 shows the POIs tracking and registration results for five test cases in both anterior-posterior and inlet views. (a) represents the DRRs with annotated points, (b) represents the real X-ray images with the tracked points, and (c) represents the DRRs generated from the registered CT projection, aligned with the X-ray images. The results demonstrate that our method accurately completes the registration task, maintaining high robustness and precision even in cases of severe misalignment (Case4), further validating the accuracy and stability of our approach.
Ablation study
This section discusses an ablation study of the proposed method. We still use mFRE, GFR, mTME, mReg.time, NCC and SSIM to evaluate the performance. The evaluation results are given in Tables 5 and 7.
In this experiment, we explored two POI tracking networks (U-NET and U-NET+CBAM) and two shape alignmets (Procrustes analysis and BnB). Table 5 shows the performance of different cases. We found that the coarse registration can be realized when directly using U-NET for POI tracking, with poor results of 14.4759mm mFRE, 15.8\(\%\) GFR and 0.4375mm mTME. Adding CBAM, the indicators mFRE, GFR and mTME have obviously reduced. As shown in the second and third rows of the Table 5, BnB achieved the best results with 6,1941mm mFRE, 3.33% GFR and 0.0655mm mTME, which have significantly improved compared with other cases. This is because BnB performs better in avoiding local optima and handling larger initial errors. At the same time, from the confidence intervals in Table 6, it can be seen that our model is more stable.
To evaluate the performance of style transfer, we compared the registration results with and without style transfer during the testing stage. Table 7 shows the results of two views. We found that without adding style transfer, the network trained on DRR is difficult to transfer to X-ray image due to the significant discrepancy between DRR and X-ray image, which results in poor registration results. After adding style transfer, the style between DRR and X-ray image is unified, and the registration accuracy is greatly improved. Although the registration time may slow down due to the increased complexity of the network, considering the crucial role of accuracy in surgical navigation, we can choose to lose some time to improve accuracy.
Limitations
To provide a more balanced perspective, we acknowledge the following limitations in our current method:
C-arm relative pose geometry Our method assumes that the relative C-arm pose geometries is known. However, most of the existing C-arm machines in the operating room do not provide such geometry information, which limits the applicability of our method.
Limited multi-view data The limited multi-view data affects the registration accuracy. Although we simulate X-ray images using DRR and unify the style through CycleGAN to reduce accuracy loss, errors in style transfer and fiducial point tracking may further increase the registration error.
Registration timeThe current training strategy trains one patient at a time, which results in longer training times, especially for critically ill patients who cannot afford to wait for the training process. And Our registration time has not reached optimal performance and requires further investigation.
Obstructions in clinical practice This work is based on an ideal dataset, whereas in clinical practice, obstructions such as surgical instruments or implanted screws may affect the model’s accuracy. We are currently investigating how to handle these obstructions and have achieved preliminary results.
Conclusion
This study explores the multi-view 2D/3D image registration problem in minimally invasive pelvic surgery navigation and has achieved the beneficial research results. The proposed model establishes point-to-point correspondence between 2D X-ray images and 3D CT data, and then aligns the shape between matching points to estimate the CT pose. The POI tracking network based on style transfer can accurately track POIs from DRR to X-ray image and establish point-to-point correspondence. The triangulation layers can project the POIs on the X-ray image to the patient. The BnB is employed to shape alignment as it can handle various types of data and perform registration more accurately. The experiment proved feasibility and superiority of the model, and provided a reliable solution for multiview image registration in minimally invasive pelvic surgery navigation.
Data availability
Correspondence and requests for data and materials should be addressed to Jingxin Zhao.
References
Liao, R., Zhang, L., Sun, Y., Miao, S. & Chefd’Hotel, C. A review of recent advances in registration techniques applied to minimally invasive Therap. IEEE Trans. Multimed. 15, 983–1000 (2013).
Markelj, P., Tomaževič, D., Likar, B. & Pernuš, F. A review of 3d/2d registration methods for image-guided interventions. Med. Image Anal. 16, 642–661 (2012).
Chang, C.-J. et al. Registration of 2d c-arm and 3d ct images for a c-arm image-assisted navigation system for spinal surgery. Appl. Bion. Biomech. 2015, 478062 (2015).
Ghafurian, S., Hacihaliloglu, I., Metaxas, D. N., Tan, V. & Li, K. A computationally efficient 3d/2d registration method based on image gradient direction probability density function. Neurocomputing 229, 100–108 (2017).
Wu, G. et al. Unsupervised deep feature learning for deformable registration of mr brain images. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2013: 16th International Conference, Nagoya, September 22–26, 2013, Proceedings, Part II 16 (2013).
Miao, S., Wang, Z. J., Zheng, Y. & Liao, R. Real-time 2d/3d registration via CNN regression. In 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI) (2016).
Miao, S., Wang, Z. J. & Liao, R. A CNN regression approach for real-time 2d/3d registration. IEEE Trans. Med. Imag. 35, 1352–1363 (2016).
Wang, S. et al. Cone beam ct series images rigid registration for temporomandibular joint via self-supervised learning network. In 2021 IEEE International Conference on Medical Imaging Physics and Engineering (ICMIPE) (2021).
Chen, X., Guo, X., Zhou, G., Fan, Y. & Wang, Y. 2d/3d medical image registration using convolutional neural network. Chin. J. Biomed. Eng 39, 394–403 (2020).
Cong, G. et al. Generalizing spatial transformers to projective geometry with applications to 2d/3d registration. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, October 4–8, 2020, Proceedings, Part III 23 (2020).
Unberath, M. et al. Deepdrr–a catalyst for machine learning in fluoroscopy-guided procedures. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2018: 21st International Conference, Granada, September 16-20, 2018, Proceedings, Part IV 11 (2018).
Unberath, M. et al. The impact of machine learning on 2d/3d registration for image-guided interventions: A systematic review and perspective. Front. Robot. AI 8, 716007 (2021).
Ali, S. et al. An objective comparison of methods for augmented reality in laparoscopic liver resection by preoperative-to-intraoperative image fusion from the miccai2022 challenge. Med. Image Anal. 99, 103371 (2025).
Liao, H. et al. Multiview 2d/3d rigid registration via a point-of-interest network for tracking and triangulation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (2019).
Wang, C. et al. Multi-view point-based registration for native knee kinematics measurement with feature transfer learning. Engineering 7, 881–888 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, October 5-9, 2015, Proceedings, Part III 18 (2015).
Woo, S., Park, J., Lee, J.-Y. & Kweon, I. S. Cbam: Convolutional block attention module. In Proceedings of the European conference on computer vision (ECCV) (2018).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the IEEE International Conference on Computer vision (2017).
Olsson, C., Kahl, F. & Oskarsson, M. Branch-and-bound methods for euclidean registration problems. IEEE Trans. Pattern Anal. Mach. Intell. 31, 783–794 (2008).
Miao, S. et al. Dilated fcn for multi-agent 2d/3d medical image registration. In Proceedings of the AAAI Conference on Artificial Intelligence, vol. 32 (2018).
Otake, Y. et al. Robust 3d–2d image registration: application to spine interventions and vertebral labeling in the presence of anatomical deformation. Phys. Med. Biol. 58, 8535 (2013).
Pan, J., Min, Z., Zhang, A., Ma, H. & Meng, M. Q.-H. Multi-view global 2d-3d registration based on branch and bound algorithm. In 2019 IEEE International Conference on Robotics and Biomimetics (ROBIO) (2019).
Acknowledgements
This work was supported in part by the Beijing Municipal Natural Science Foundation (4242016).
Author information
Authors and Affiliations
Contributions
F.J: Writing-review & editing, Supervision, Funding acquisition. Y.W: Conceptualization, Methodology, Software, Writing-original drat. M.D: Reviewing & editing. J.Z: Resources, Investigation. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
Te authors declare no competing interests.
Additional information
Correspondence and requests for materials should be addressed to Jingxin Zhao.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ju, F., Wang, Y., Zhao, J. et al. Multiview 2D/3D image registration in minimally invasive pelvic surgery navigation. Sci Rep 15, 26183 (2025). https://doi.org/10.1038/s41598-025-10884-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-10884-4