Abstract
Immunotherapy has opened new avenues of treatment for patients with advanced non-small cell lung cancer (NSCLC) without previous hope of survival. Unfortunately, only a small percentage of patients benefit from it and there is not an effective biomarker to predict patients’ response. Since T cells are key effectors of antitumor immunity, T cell receptor (TCR) has emerged as a potential predictive biomarker. Here, we evaluated the potential of baseline TCR repertoire, as a predictive biomarker in advanced NSCLC patients treated with pembrolizumab at first-line. After obtaining peripheral blood and tissue samples at baseline, next-generation sequencing targeting TCRβ/γ was performed. We found an uneven tumor-infiltrating TCRβ repertoire, and the use of various tumor-infiltrating and circulating TRBV/J genes were able of predicting the immunotherapy response. Our results support the potential of evaluating tissue and circulating TCRβ repertoire prior to pembrolizumab, revealing it as a promising immunotherapy response biomarker in NSCLC patients.
Similar content being viewed by others
Introduction
Lung cancer represents the most common and deadliest cancer worldwide1. It can be classified into small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), with the latter accounting for 85% of cases2,3. NSCLC is a highly heterogeneous disease with nonspecific symptoms, often leading to a late diagnosis at advanced stages (III-IV), where surgery is not feasible, and the patient’s survival becomes greatly compromised2,3,4.
Nowadays, targeted therapy against actionable mutations represents the standard treatment for molecularly defined populations2,3. Unfortunately, only 25% of patients carry any of these mutations2. Immunotherapy with Immune Checkpoint Inhibitors (ICIs) has been established as the standard treatment for patients with advanced NSCLC who do not have actionable mutations, either as monotherapy or in combination with chemotherapy, leading to a 5 year survival rate up to 20%2,5,6. Unfortunately, only a small percentage of patients (20–40%) benefit from it2,5,6. Its high cost, the limited time for therapy decision-making, and the possibility of developing autoimmune-related side effects underscore the importance of identifying those patients who can truly benefit from the treatment2,5,6.
PD1/PD-L1 immunostaining is the only approved predictive biomarker for immunotherapy response2,5,6,7. However, its reproducibility has been limited by the heterogeneity of results obtained in subsequent studies, largely due to lack of standardization2,5,6,7. Over time, other alternatives such as tumor mutational burden (TMB)2,5,6,7, tumor-infiltrating lymphocytes (TILs)2,7 and other molecular characteristics have been proposed5,6,7, but none have been able to reliably predict patient response in the real clinical practice2,5,6,7.
Cytotoxic T cells play a key role by recognizing cancer cell-specific antigens, known as neoantigens, and killing cancer cells through their T cell receptors (TCRs)7. TCR is composed of an α and a β chain in Tαβ cells (95%) or by a γ and a δ chain in Tγδ cells (5%)7,8,9,10. Its synthesis occurs through a somatic, stochastic, and imprecise recombination of various non-contiguous homologous gene sets during T cell differentiation in the thymus10. Each chain consists of a variable and a constant region. The variable region of the α and γ chains results from recombination between the V and J genes, while β and δ chains results from recombination between the V, D, and J genes. The result of this V(D)J recombination creates a variable domain that is transcribed and joins one of the constant genes (C), resulting in a functional chain7,8,9,10. Each chain contains three hypervariable loops (CDR1, CDR2, and CDR3) in its structure, with CDR3 mediating peptide recognition and thus determining TCR specificity10. CDR3 is encoded by V(D)J genes and undergoes a series of random nucleotide insertions and deletions (N) during recombination at gene junctions (VN(D)NJ). This entire process ensures the creation of a vast array of unique TCRs capable of recognizing a wide range of antigens7,8,9,10.
Advancements in sequencing technologies and bioinformatics tools have enabled sensitive and precise characterization of TCR repertoire7,8. In recent years, various researches have suggested the potential use of the TCR repertoire as a biomarker for immunotherapy response, both for anti-CTLA4 and anti-PD-L1/PD1 therapies in different types of cancer (such as melanoma, breast and liver), including NSCLC11,12,13,14,15,16,17,18,19,20,21. These analyses have yielded contradictory results but indicate a potential capacity for the TCR repertoire to be used as an immunotherapy response biomarker. Consequently, the main objective of this study is to evaluate the potential of the tumor-infiltrating and circulating TCRβ/γ repertoire at baseline as a biomarker for predicting the response to immunotherapy in a real cohort of advanced NSCLC patients treated with pembrolizumab in the first-line setting.
Materials and methods
Study design and sample collection
This study included 12 patients diagnosed with advanced (IIIA-IVB) NSCLC treated with pembrolizumab, 11 of them in first-line, at Álvaro Cunqueiro Hospital in Vigo. The study, including the experimental protocols, was conducted with appropriate authorization from the Galician Regional Research Ethics Committee (2019/046) following the Helsinki Declaration of 1975 and all patients signed informed consent approving their participation. When available, both circulating and infiltrating TCR repertoire were analysed for each patient (n = 22). Patients were classified as responders (R: complete/partial response or stable disease for each patient) and non-responders (NR: tumor progression or not evaluable)22 according to the computed tomography scan at 3 months. The most relevant clinicopathological characteristics are represented in Table 1. To examine tumor-infiltrating TCR repertoire, formalin-fixed paraffin-embedded (FFPE) tumor tissues were obtained from all patients prior to immunotherapy. To analyse the circulating TCR repertoire, blood samples were drawn from 10 patients before the first ICIs dose. Peripheral blood mononuclear cells (PBMCs) were isolated from blood samples by gradient density centrifugation using Ficoll® (Sigma-Aldrich) and cryopreserved until use. DNA was chosen over RNA for analysis due to its superior stability in FFPE samples.
DNA extraction
DNA from FFPE samples was extracted using the AllPrep® DNA/RNA FFPE Kit (QIAGEN) following the manufacturer’s instructions. DNA extraction from PBMCs was performed using the QIAamp® DNA Blood Mini Kit (QIAGEN) following the manufacturer’s instructions. DNA quantification was carried out using NanoDropTM 2000c Spectrophotometer (Thermo Fisher Scientific).
Library preparation and TCR sequencing
A total of 250 ng of genomic DNA was used to prepare the 22 libraries for TCR CDR3β/γ region sequencing following the Oncomine™ TCR Pan-Clonality Assay (Thermo Fisher Scientific). This targeted next-generation sequencing (NGS) assay has been specifically designed to sequence the FR3-J regions of the TCRβ and TCRγ chains. Libraries were pooled on an Ion 540 chip at 25 pmol/L and sequenced using an Ion GeneStudio S5 Plus Series (Thermo Fisher Scientific).
TCR sequencing data analysis
Raw sequencing data analysis was conducted using Ion Reporter version 5.20.2.0 (Thermo Fisher Scientific). Regarding quality control, off-target and unproductive reads were removed. Read classification is shown in Supp. File S1. Afterwards, the software reported the VDJ rearrangements and the main repertoire metrics, such as richness, convergence, diversity, and evenness. TCR richness represented the total number of clones, defined as unique TCRβ/γ nucleotide sequences. TCR convergence was determined as the aggregate frequency of clones, which shared a variable gene and CDR3 amino acid sequence. Diversity (Shannon’s diversity) and evenness (normalized Shannon’s diversity) were calculated using the equations (1 and 2), in which pi is the frequency of clone i for the sample with n unique clones. Evenness describes how evenly distributed is the TCR repertoire. It ranges from 0, meaning the repertoire is unbalanced by a reduced number of predominant clones, to 1, meaning the repertoire is balanced with similar frequencies of all the clones.
In addition, clones were classified according to their relative abundance in the repertoire, named total clonal space, for each sample. The clonal space was divided into the top 1%, top 3% and top 5%, so the top 1% clonal space is defined as the aggregate frequencies of the top 1% most frequent clones. All the variables were categorized as “high” or “low” based on their median value.
Statistical analysis
Statistical analyses were performed using R version 4.2.1 environment. Statistical significance was considered by an overall p < 0.05. TCR repertoire variables were correlated with clinical-pathological data (age, sex, tumor histology, PD1/PD-L1 expression, and smoking status), clinical response and survival. Overall-survival (OS) was defined as the time from treatment initiation to death or last follow-up, and progression-free survival (PFS) was defined as time from treatment initiation to progression or death (whichever is earlier), or last follow-up. Mann-Whitney U test and Student’s t-test were used to study the relationship between TCR repertoire variables and clinical-pathological data and clinical response. Choice of appropriate test (Mann-Whitney or Student’s t) was based on normality (Shapiro-Wilk test) and variance homogeneity (Levene test). Spearman rank non-parametric test was used for variable correlations. Contingency table analyses when comparing clinical-pathological data with clinical response or clinical-pathological data with TCR repertoire variables were done using Fisher exact test. Kaplan-Meier survival analyses and Log-Rank test were used for time-dependent variables such as OS and PFS. For single-variables, receiver operating characteristics (ROC) curve analyses were performed to evaluate response prediction, and areas under the curve (AUC), sensitivity and specificity values were obtained. The Youden Index method was used to select the best cut-off values for all classification analyses23. Random Forest Analysis (number of trees = 1000) was used to build the combined predictive signatures performances. To enhance model diversity and reduce overfitting, the number of features sampled for splitting at each node was set to the square root of the total number of predictors. Model performance was evaluated by ROC curves derived by leave-one-out cross-validation (LOOCV) by iteratively training the model on all samples except one and testing on the excluded sample, in line with standard practice for a robust model estimation in small datasets24.
Results
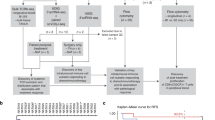
Figure 1 provides a graphical overview of the study design and the analytical workflow with key results (Fig. 1).
Graphical summary of the study design and analytical workflow, including sample collection, analysis steps, and key results. AUC: Area Under the Curve, N: number of patients, *: statistically significance.
Age is associated with immunotherapy response and tumor-infiltrating evenness
This retrospective observational study included a total of 22 samples from 12 patients diagnosed with locally advanced or advanced NSCLC (IIIA-IVB) treated with pembrolizumab (Table 1). Both tissue FFPE (n = 12) and PBMCs (n = 10) samples were collected before treatment and were subjected to TCR sequencing. Among the 12 patients, only two were PD1-PD-L1 negative. We obtained valid TCR data for all samples.
First, we evaluated the effect of immunotherapy response on patient prognosis. R had statistically significant longer PFS (p = 0.015) than NR (Supp. File S2A). We found the same trend regarding OS (p = 0.057) (Supp. File S2B). Interestingly, we detected an association between response and age, where R were statistically significant older than NR (p = 0.008) (Supp. File S3A), but not with the rest of clinicopathological characteristics. All R were above the median age of the cohort (70.5 years) (Supp. File S3B).
Second, we sought for possible associations between the clinicopathological characteristics and TCR repertoire characterization. We detected that squamous tumors presented statistically significant higher tumor-infiltrating TCRβ convergence (p = 0.026) than adenocarcinomas (Supp. File S3C). Furthermore, we found an association between age and tumor-infiltrating evenness in both TCRβ and TCRγ, where younger patients (<71 years old) had statistically significant higher tumor-infiltrating evenness than older patients (≥ 71 years old) (p = 0.022 and p = 0.041, respectively) (Supp. File S3D-E). However, we verified that none of these clinicopathological characteristics have a relevant impact on our metrics regarding immunotherapy response prediction (Supp. File S4).
Tumor-infiltrating TCRβ evenness is associated with immunotherapy response and progression-free survival
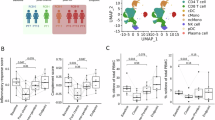
With respect to FFPE tumor samples, we found that R had statistically significant lower tumor-infiltrating TCRβ evenness than NR (p = 0.044) (Fig. 2A). Thus, its ability of potentially predicting the response and survival of these patients was analysed. Tumor-infiltrating TCRβ evenness predicted response with an AUC of 0.86, where a tumor-infiltrating TCRβ evenness lower than 0.795 had a 60% sensitivity and 100% specificity of predicting response (Fig. 2B). Moreover, patients with low tumor-infiltrating TCRβ evenness (< 0.8441) had statistically significant longer PFS than patients with high (≥ 0.8441) tumor-infiltrating TCRβ evenness (p = 0.013) (Fig. 2C). Similarly, we found the same trend regarding circulating TCRβ evenness (p = 0.139) (Fig. 2D). Despite not reaching the statistical significance, the same trends were found regarding TCRγ in both tissue and blood, respectively (p = 0.149 and p = 0.114), showing a lower evenness in R patients (Fig. 2E-F). It is worth to mention that R patients showed a tendency towards a higher tumor-infiltrating and circulating TCRβ top 3% and top 5% clonal space compared with NR patients (Supp. File S5A-D). Accordingly, TCRβ evenness and top 3% and 5% clonal space were always strongly negatively correlated both in tissue (R = −0.81, p = 0.001; R = −0.91, p < 0.001, respectively) and blood (R = −0.99, p < 0.001; R = −0.97, p < 0.001, respectively) (Supp. File S6).
TCRβ/γ evenness as a predictor of clinical response and patient survival. (A) Responders (R) have statistically significant lower tumor-infiltrating TCRβ evenness than non-responders (NR) (p = 0.044). P was obtained using Student’s t test. (B) Tumor-infiltrating TCRβ evenness predicts response with an AUC of 0.86. A lower tumor-infiltrating TCRβ evenness than 0.795 have a 60% sensitivity and 100% specificity predicting response. (C) Patients with low tumor-infiltrating TCRβ evenness have statistically significant longer PFS than patients with high tumor-infiltrating TCRβ evenness (p = 0.013). P was obtained using Log-Rank test. (D) R have a non-statistically significant lower circulating TCRβ evenness than NR (p = 0.139). P was obtained using Student’s t test. (E) R have a non-statistically significant lower tumor-infiltrating TCRγ evenness than NR (p = 0.149). P-value was obtained using Mann-Whitney test. (F) R have a non-statistically significant lower circulating TCRγ evenness than NR (p = 0.114). P was obtained using Mann-Whitney test. AUC: Area Under the Curve, N: number of patients, NE: non-evaluable, PD: progressive disease, PFS: progression-free survival, PR: partial response, SD: stable disease, *: statistical significance.
Tumor-infiltrating and circulating TCRβ, TRBV and TRBJ genes usage is associated with immunotherapy response and progression-free survival
Next, we sought for any relation between TRBV/TRGV and TRBJ/TRGJ genes frequencies and the immunotherapy response. Regarding TCRγ, there was no association between TRGV and TRGJ genes frequency and clinical response, neither in tissue nor in blood. Interestingly, tumor-infiltrating TCRβ TRBV6.5, TRBV11.3 and TRBJ2.1 frequencies were statistically significant lower in R than in NR (Supp. File S7; p = 0.011, p = 0.048 and p = 0.002, respectively). Moreover, circulating TCRβ TRBV5.3 frequency was statistically significant higher in R than in NR (p = 0.019), while TRBV27, TRBV28, TRBJ2.1 and TRBJ2.6 frequencies were statistically significant lower in R than in NR (p = 0.039, p = 0.029, p = 0.017, and p = 0.008, respectively), as shown in Supp. File S8. The distribution of TRBV and TRBJ gene frequencies across the cohort is provided in Supp. File S9.
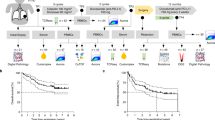
Then, we analysed their ability of potentially predict the PFS and the immunotherapy response in these patients. Patients with low tumor-infiltrating TCRβ TRBV11.3 frequency presented statistically significant longer PFS than patients with high frequency (p = 0.002) (Fig. 3A). Tumor-infiltrating TCRβ TRBV11.3 frequency predicted response with an AUC of 0.86, where a lower frequency than 0.0057 had an 80% sensitivity and 85.7% specificity predicting response (Fig. 3B). In the same line, patients with low tumor-infiltrating TCRβ TRBJ2.1 frequency had longer PFS than patients with high frequency, although not reaching the statistical significance (p = 0.051) (Fig. 3C). Tumor-infiltrating TCRβ TRBJ2.1 frequency predicted response with an AUC of 0.94, where a lower frequency than 0.0460 had 100% sensitivity and 85.7% specificity predicting response (Fig. 3D). Moreover, patients with low circulating TCRβ TRBJ2.6 frequency had statistically significant longer PFS than patients with high frequency (p = 0.003) (Fig. 3E). Circulating TCRβ TRBJ2.6 frequency predicted response with an AUC of 1.00, where a frequency cut-off < 0.0055 had an 100% sensitivity and 100% specificity predicting response (Fig. 3F).
TCRβ gene usage as a predictor of clinical response and patient survival. (A) Patients with low tumor-infiltrating TCRβ TRBV11.3 frequency have statistically significant longer PFS than patients with high tumor-infiltrating TCRβ TRBV11.3 frequency (p = 0.002). P was obtained using Log-Rank test. (B) Tumor-infiltrating TCRβ TRBV11.3 frequency predicts response with an AUC of 0.86. A lower tumor-infiltrating TCRβ TRBV11.3 frequency than 0.0057 have an 80% sensitivity and 85.7% specificity predicting response. (C) Patients with low tumor-infiltrating TCRβ TRBJ2.1 frequency have non-statistically significant longer PFS than patients with high tumor-infiltrating TCRβ TRBJ2.1 frequency (p = 0.051). P was obtained using Log-Rank test. (D) Tumor-infiltrating TCRβ TRBJ2.1 frequency predicts response with an AUC of 0.94. A lower tumor-infiltrating TCRβ TRBJ2.1 frequency than 0.0460 have an 100% sensitivity and 85.7% specificity predicting response. (E) Patients with low circulating TCRβ TRBJ2.6 frequency have statistically significant longer PFS than patients with high circulating TCRβ TRBJ2.6 frequency (p = 0.003). P was obtained using Log-Rank test. F) Circulating TCRβ TRBJ2.6 frequency predicts response with an AUC of 1.00. A lower circulating TCRβ TRBJ2.6 frequency than 0.0055 have an 100% sensitivity and 100% specificity predicting response. AUC: Area Under the Curve, N: number of patients, PFS: progression-free survival, *: statistical significance, **: strongly statistically significant.
Combined TCRβ signatures to predict response
Although some of these TRBV/TRBJ genes did not reach the statistical significance in predicting PFS (Supp. File S10), they were able to accurately predict the clinical response (Supp. File S11). Thus, we decided to develop two Random Forest predictive signatures that combine the previously described tumor-infiltrating and thecirculating TCRβ variables associated with clinical response. For the tissue predictive signature, we included the variables evenness, TRBV6.5, TRBV11.3, and TRBJ2.1. This Random Forest signature was able to predict clinical response with an AUC of 0.83. A positive result from this signature (random forest-derived probability of being classified as responder > 0.230) had a sensitivity of 100% and a specificity of 71.4% in predicting response (Fig. 4A). In contrast, the predictive signature for blood samples included the variables TRBV5.3, TRBV27, TRBV28, TRBJ2.1, and TRBJ2.6. The circulating signature demonstrated a predictive ability with an AUC of 0.92. In this case, a positive result (derived probability > 0.641) had a sensitivity of 75% and a specificity of 100% in predicting response (Fig. 4B).
Tumor-infiltrating and circulating TCRβ predictive signatures for clinical response. (A) A predictive signature including tumor-infiltrating TCRβ variables (evenness, TRBV6.5 frequency, TRBV11.3 frequency and TRBJ2.1 frequency) predicts response with an AUC of 0.83. A positive result has an 100% sensitivity and 71.4% specificity predicting response. (B) A predictive signature including circulating TCRβ variables (TRBV5.3 frequency, TRBV27 frequency, TRBV28 frequency, TRBJ2.1 frequency and TRBJ2.6 frequency) predicts response with an AUC of 0.92. A positive result has an 75% sensitivity and 100% specificity predicting response. AUC: Area Under the Curve.
Discussion
ICIs represent the standard treatment for advanced NSCLC patients who lack of actionable mutations and cannot undergo surgery. Despite ICIs have significantly improved both life expectancy and quality of life for patients who benefit from it, the response rate remains low (20–40%), and there are no reliable predictive response biomarkers2,5,6. Recently, the study of the TCR repertoire has emerged as a potential predictive biomarker for immunotherapy response in various types of cancer but with still uncertain results11,12,13,14,15,16,17,18,19,20,21.
To the best of our knowledge, this is the first study sequencing the tumor-infiltrating and circulating TCRβ/γ repertoire at baseline in advanced NSCLC patients treated with checkpoint inhibition at first-line. Most researches regarding the TCR repertoire have focused solely on Tαβ cells11,14,15,17,18,19,20,21, with very few analysing the Tγδ cells, especially in solid tumors12,13. Although the mechanism of action of Tγδ cells is still not precisely known, it has been shown that they can play an important role in antitumor response in both systemic and local immunosurveillance8,9,12,13,16. For this reason, we decided to include also the γ chain. Moreover, many researches have analysed the TCR repertoire dynamics before and after treatment14,15,17,21, providing valuable insights into the mechanisms behind the antitumor response reactivation. However, most advanced NSCLC patients do not survive long enough to start a second-line therapy. Thus, the study of the TCR repertoire as a biomarker for ICIs response should be applicable before the first-line treatment to have a real impact in diary clinical practice.
We focused on the TCR repertoire at baseline, including both circulating and infiltrating profiles from advanced NSCLC patients treated with pembrolizumab at first-line. Our results confirmed that R achieved greater PFS, with the same trend observed for OS. Interestingly, we found that R were older than NR. On the contrary, a previous comprehensive comparative analysis using a cohort of anti-PD1-treated patients with renal cell carcinoma identified no statistically significant difference in PFS or OS with age 25. Due the limited numbers of older patients treated with ICIs available for their analyses, there is a need for further investigation of this subject in other cancer types, including NSCLC25. We also found older patients had a lower TCRβ/γ evenness in tissue but not in blood. In concordance, Erbe et al. reported a decrease in tissue TCR evenness with age in patients with adenocarcinoma25. Moreover, Dong et al. did not find such an association regarding circulating evenness either15. Our findings support that while TCRβ variables may be influenced by age, they still possess individual predictive capacity. Indeed, the accuracy of the combined signatures in predicting response is equal (blood) or higher (tissue) when age is excluded. However, these findings must be carefully considered, since the observed increase in TCR clonality with age may limit its predictive effectiveness in older patients26. In any case, larger studies are also needed to definitively evaluate how other comorbidities could affect to this evenness.
According to the histological classification and the TCR repertoire, previous studies showed contradictory associations. For instance, some analysis described how patients with squamous cell carcinomas had lower tumor-infiltrating TCRβ evenness11,20, clonal richness20 or diversity11 compared to patients with adenocarcinoma, while others did not find any association between histology and any TCR repertoire metric14,15. In contrast, we observed squamous had higher tumor-infiltrating TCRβ convergence than adenocarcinomas, although our cohort contained only three patients with squamous cell carcinomas, so additional validation in larger cohorts will be needed to confirm these findings.
Our results showed that R exhibited low tumor-infiltrating TCRβ evenness at baseline, and this could predict a longer PFS. The association between low tumor-infiltrating TCRβ evenness and treatment response could be explained by the clonal expansion of specific T cell clones mediating the anti-tumor effect. In other words, certain T cell clones might have recognized tumor-expressed neoantigens but were unable to effectively eliminate the cancer cells until the initiation of immunotherapy in R. Thus, TCRβ evenness could potentially serve as a baseline biomarker for predicting response to immunotherapy. Concordant findings were found in clear cell renal cell carcinoma, where R were observed to have lower tumor-infiltrating TCRβ evenness at baseline21. Moreover, Casarrubios et al. reported a high tumor-infiltrating TCRβ top 1% clonal space and a low evenness at baseline were involved in a pro-inflammatory tumor microenvironment associated with the complete pathological response of partially advanced NSCLC patients treated with neoadjuvant chemo-immunotherapy17. Additionally, they described how these same top 1% clones expanded in peripheral blood after treatment, suggesting a possible role of systemic immunosurveillance in preventing relapse.
We also found that some TCRβ genes were associated with the immunotherapy response either regarding tissue or blood. Interestingly, R had a higher circulating TCRβ TRBV5.3 than NR. The association between treatment response and a higher circulating TCRβ TRBV5-3 frequency might be due to certain tumor neoantigens exhibiting high affinity for TCR sequences encoded by this gene. This could enhance tumor cell recognition and elimination, thereby promoting the clonal expansion of specific T cells harbouring this gene and ultimately mediating the anti-tumor response. Besides, tumor-infiltrating TRBV11.3 and circulating TRBJ2.6 also predicted PFS. Similarly, Dong et al. reported a circulating TCRβ low TRBV6.5 and high TRBV20.1 at baseline showed a longer PFS and OS15. Plus, TRBV20.1 has been reported previously as one of the most used TCRβ V-gene in tumor-infiltrating T cells in several solid27. Therefore, it seems the TRBV/J gene usage is also being able of predicting the response both in tissue and blood. However, it is not clear which genes are more decisive, if there are ones.
Consequently, we decided to combine all the variables associated with response in both a tumor-infiltrating and a circulating predictive signature using a Random Forest classifier. Although both tissue and blood predictive signatures demonstrated high accuracy, they performed differently in terms of sensitivity and specificity. The tissue-based model achieved 100% sensitivity and 71.4% specificity with an AUC of 0.83, meaning that it would correctly identify all R and thus ensure that all patients who could benefit from ICIs treatment receive it. However, it would also misclassify approximately 28.6% of NR, potentially leading to unnecessary treatment in these patients. On the other hand, the blood-based model showed 75% sensitivity and 100% specificity with an AUC of 0.92, implying that any NR would receive ICIs, but 25% of potential R would be missed and left untreated. From a clinical perspective, we believe that the tissue model is more advantageous, as it prioritizes treating all patients who have the potential to benefit from ICIs, ensuring that all likely R are treated with ICIs. However, it is necessary to highlight the potential of identifying potential biomarkers in blood in advanced lung cancer, whose typically small lung tumor biopsies may fail to capture the high intratumoral heterogeneity6. Additionally, finding minimally invasive circulating biomarkers that are easily measurable throughout therapy would facilitate their implementation in daily clinical practice, as the number of recent studies has demonstrated14,15,18.
Regarding TCRγ, we detected the same trend found in TCRβ repertoire without reaching the statistical significance, where R had a lower tumor-infiltrating and circulating TCRγ evenness at baseline than NR. However, we did not find association between TCR variables or V/J genes usage and immunotherapy response. Although a high number of distinct TCRγδ can theoretically be generated, certain rearrangements and chain pairings are substantially over-represented, biasing V(D)J recombination and the selection of functional clones8. This leads to various oligoclonal populations predominating in the circulation and in different tissues. Furthermore, there is minimal knowledge reported to date regarding the TCRγ repertoire as an immunotherapy response biomarker12,13, so further validation will be necessary to verify these findings and draw more definitive conclusions.
In summary, our study suggests a potential of TCRβ repertoire for predicting response to pembrolizumab in advance NSCLC. We have preliminary identified various tumor-infiltrating and circulating TCRβ variables at baseline that correlate with response and progression-free survival. However, given the exploratory nature of this retrospective observational study due to the small cohort size, further research is needed to validate these findings in larger, more diverse populations and to establish causal relationships.
Data availability
Data are available in the NCBI repository under accession number: SUB15462404.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 74, 229–263 (2024).
Wang, M., Herbst, R. S. & Boshoff, C. Toward personalized treatment approaches for non-small-cell lung cancer. Nat. Med. 27, 1345–1356 (2021).
Lahiri, A. et al. Lung cancer immunotherapy: Progress, pitfalls, and promises. Mol. Cancer 22, 40 (2023).
Padinharayil, H. et al. Non-small cell lung carcinoma (NSCLC): Implications on molecular pathology and advances in early diagnostics and therapeutics. Genes Dis. 10, 960–989 (2023).
Dugage, M. R. et al. Biomarkers of response to immunotherapy in early stage non-small cell lung cancer. Eur. J. Cancer 184, 179–196 (2023).
Oitabén, A. et al. Emerging blood-based biomarkers for predicting immunotherapy response in NSCLC. Cancers 14, 2626 (2022).
Luo, H. et al. The nexus of dynamic T cell states and immune checkpoint blockade therapy in the periphery and tumor microenvironment. Front. Immunol. 14, 1267918 (2023).
Mensurado, S., Blanco-Domínguez, R. & Silva-Santos, B. The emerging roles of γδ T cells in cancer immunotherapy. Nat. Rev. Clin. Oncol. 20, 178–191 (2023).
Hu, Y. et al. γδ T cells: Origin and fate, subsets, diseases and immunotherapy. Sig. Transduct. Target Ther. 8, 434 (2023).
Sanromán, Á. F., Joshi, K., Au, L., Chain, B. & Turajlic, S. TCR sequencing: Applications in immuno-oncology research. Immuno Oncol. Technol. 17, 100373 (2023).
Reuben, A. et al. Comprehensive T cell repertoire characterization of non-small cell lung cancer. Nat. Commun. 11, 603 (2020).
Janssen, A. et al. γδ T-cell receptors derived from breast cancer-infiltrating T lymphocytes mediate antitumor reactivity. Cancer Immunol. Res. 8, 530–543 (2020).
Hunter, S. et al. Human liver infiltrating γδ T cells are composed of clonally expanded circulating and tissue-resident populations. J. Hepatol. 69, 654–665 (2018).
Han, J. et al. TCR repertoire diversity of peripheral PD-1+CD8+ T cells predicts clinical outcomes after immunotherapy in patients with non-small cell lung cancer. Cancer Immunol. Res. 8, 146–154 (2020).
Dong, N. et al. Characterization of circulating t cell receptor repertoire provides information about clinical outcome after PD-1 blockade in advanced non-small cell lung cancer patients. Cancers 13, 2950 (2021).
Davey, M. S. et al. Clonal selection in the human Vδ1 T cell repertoire indicates γδ TCR-dependent adaptive immune surveillance. Nat. Commun. 8, 14760 (2017).
Casarrubios, M. et al. Pretreatment tissue tcr repertoire evenness is associated with complete pathologic response in patients with NSCLC receiving neoadjuvant chemoimmunotherapy. Clin. Cancer Res. 27, 5878–5890 (2021).
Abed, A. et al. Circulating pre-treatment T-cell receptor repertoire as a predictive biomarker in advanced or metastatic non-small-cell lung cancer patients treated with pembrolizumab alone or in combination with chemotherapy. ESMO Open 8, 102066 (2023).
Hogan, S. A. et al. Peripheral blood TCR repertoire profiling may facilitate patient stratification for immunotherapy against melanoma. Cancer Immunol. Res. 7, 77–85 (2019).
Kargl, J. et al. Neutrophils dominate the immune cell composition in non-small cell lung cancer. Nat. Commun. 8, 14381 (2017).
Au, L. et al. Determinants of anti-PD-1 response and resistance in clear cell renal cell carcinoma. Cancer Cell 39, 1497-1518.e11 (2021).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: Revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Youden, W. J. Index for rating diagnostic tests. Cancer 3, 32–35 (1950).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning. (Springer, New York, NY, 2009). https://doi.org/10.1007/978-0-387-84858-7.
Erbe, R. et al. Evaluating the impact of age on immune checkpoint therapy biomarkers. Cell Rep. 36, 109599 (2021).
Salih, Z. et al. T cell immune awakening in response to immunotherapy is age-dependent. Eur. J. Cancer 162, 11–21 (2022).
Li, B. et al. Landscape of tumor-infiltrating T cell repertoire of human cancers. Nat. Genet. 48, 725–732 (2016).
Acknowledgements
Authors thank all the enrolled patients and their families. Samples were collected and stored by the Galicia Sur Health Research Institute (IIS Galicia Sur) Biobank (registry B.0000802).
Funding
This work was supported by the Instituto de Salud Carlos III (ISCIII) and cofunded by European Union under Grants PI21/00348 and CP20/00188. MPG is currently supported by the predoctoral fellowship from Ministerio de Ciencia, Innovación y Universidaders (FPU23/00744) and previously supported by the Axencia Galega de Innovación-GAIN (Xunta de Galicia) under Grant for the predoctoral stage IN606A-2024/017. MGG is currently supported by the Axencia Galega de Innovación-GAIN (Xunta de Galicia) under Grant for the postdoctoral stage IN606B-2024/014. MMF is currently supported by the Instituto de Salud Carlos III (ISCIII) under the Miguel Servet program CP20/00188.
Author information
Authors and Affiliations
Contributions
MPG performed formal analytics and statistical analysis and wrote the manuscript. MLQ enrolled patients, reviewed the clinical data annotations and wrote the manuscript. IAA performed and optimized DNA isolation and libraries preparation. MGG gave analytics and statistical analysis support. LJM evaluated tissue availability for FFPE sample selection. AFF reviewed the clinical data annotations. SCF and EJL performed TCR sequencing. MMF conceived and designed the study, provided supervision and oversight for the management and execution of this study, acquired financial support for the project, and correct the manuscript. All authors provided critical review, edited the manuscript, and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pino-González, M., Lázaro-Quintela, M., Alonso-Álvarez, I. et al. Tissue and peripheral T cell receptor repertoire predicts immunotherapy response and progression-free survival in NSCLC patients. Sci Rep 15, 40914 (2025). https://doi.org/10.1038/s41598-025-21612-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-21612-3