Abstract
The effects of therapeutic drugs that inhibit female hormone functions on uterine leiomyomas (ULMs) vary among individuals. ULMs are classified into two subtypes with different histological compositions: MED12 mutation-positive and -negative ULMs. The present study was undertaken to investigate the tumor regression effects of female hormone withdrawal and a selective progesterone (P) receptor modulator (ulipristal acetate, UPA) in the ULM subtypes using a xenograft mouse model. Smooth muscle cells (SMCs) isolated from each ULM subtype were transplanted and treated with estrogen (E) and P for 4 weeks to form the xenograft tumor. They were then divided into four groups: continued EP treatment (E(+)P(+)), EP withdrawal (E(-)P(-)), only P withdrawal (E(+)P(-)), and EP + UPA. Tumors were harvested 8 weeks after transplantation. In both subtypes, they were reduced in size in the E(-)P(-) compared with the E(+)P(+) and further reduced in the EP + UPA. Histological analysis showed that SMCs shrank in the E(-)P(-) and disappeared in the EP + UPA. Gene ontology and pathway analyses of the differentially expressed genes of the xenograft tumors suggested that regression occurred by different mechanisms in the two subtypes. Our results also suggested, for the first time, that UPA caused SMC death by a mechanism other than progesterone receptor inhibition.
Similar content being viewed by others
Introduction
Uterine leiomyomas (ULMs) are tumors derived from uterine smooth muscle cells (SMCs) and are the most common neoplastic disease, occurring in more than 30% of sexually mature women1. ULMs are benign but can cause not only dysmenorrhea, hypermenorrhea, and anemia, but also infertility and miscarriage, severely affecting a woman’s quality of life1,2. Although surgical treatment, such as hysterectomy or myomectomy, is essential to cure ULMs, there is a growing demand for drug therapies that can preserve the uterus because of a trend of women delaying marriage and their childbearing years. The development of effective drugs requires elucidation of the pathophysiology of ULM.
Missense mutations in the mediator complex subunit 12 (MED12) gene have been implicated in the pathogenesis of ULMs, and have been detected in approximately 70% of ULM specimens in multiple ethnic groups3,4,5,6,7,8,9,10,11. Therefore, ULMs have at least two subtypes depending on the presence or absence of the mutations. Recently, it has been reported that the ULM subtypes differ in histological composition, including the amount of collagen fibers and the cellular component of the tumors11,12. We have also shown that ULMs with the MED12 mutation (MED12m(+)-ULM) contain abundant collagen fibers and are composed of approximately 60% SMCs and 40% fibroblasts, whereas ULMs without the mutation (MED12m(-)-ULM) are composed of approximately 80% SMCs and 20% fibroblasts, with low collagen content11. In addition, fibroblasts can proliferate with estrogen (E) only, whereas SMCs require E and progesterone (P) for their growth11, suggesting that the ULM subtypes respond differently to female hormones.
Effective therapeutic drugs for ULMs include gonadotropin-releasing hormone analogs (GnRHa) and selective progesterone receptor modulators (SPRMs), both of which work by inhibiting female hormone function. However, the tumor reduction effects of these drug therapies differ among individuals. Clinically, GnRHa, which inhibits E and P secretion, has been reported to be more effective in tumor regression in MED12m(-)-ULM than in MED12m(+)-ULM13. The effects of SPRMs which inhibit progesterone receptor (PGR) function also differ among individuals14. However, it remains unknown whether the effects of female hormone withdrawal (FHW) or SPRMs are different between the ULM subtypes, and how they are different if so.
Most studies about the effects of female hormones15,16,17,18,19,20,21 and SPRMs22,23,24 on ULM cells have been conducted using 2D monolayer cultures. However, it has been reported that SMCs are selectively lost in the cultures25,26, that the expressions of estrogen receptor 1 (ESR1) and PGR in ULM cells gradually disappear27, and that the genome-wide gene expression profile changes in monolayer culture28, suggesting that it is difficult to maintain the characteristics of the original ULM specimens in such cultures. On the other hand, xenografts are recognized as the only experimental system that can reflect the effects of female hormones in vivo in ULM studies12,29,30,31. A xenograft study showed that SMCs and fibroblasts respond differently to female hormones12.
To elucidate part of the pathophysiology of ULMs, the present study was undertaken to investigate the differences in the tumor regression effects of FHW and an SPRM (ulipristal acetate, UPA) between the ULM subtypes using a xenograft model.
Results
Establishment of a xenograft model using ULM cells
Xenograft tumors were harvested 8 weeks after transplantation and histologically compared with the original ULM specimen (Fig. 1B). Both MED12m(+)- and (-)-ULM xenograft tumors appeared similar to the original ULM specimens in HE staining, trichrome staining, and immunostaining for ACTA2 and VIM, demonstrating that xenografts give accurate characteristics of the original ULMs.
Characterization of established xenograft models using uterine leiomyoma (ULM) cells. (A) Representative immunocytochemistry images for the smooth muscle cell marker ACTA2 (green) and the progesterone receptor PGR (red) in the primary culture of ULM cells used for a xenograft model. The photograph shows the ULM cells of the MED12m(+)-ULM on day 3 of primary culture. Nuclear staining was performed with DAPI. Scale bar = 50 μm. (B) Representative images of HE staining, trichrome staining detecting collagen fibers in blue and cells in dark red, and immunohistochemistry for ACTA2 (green) and mesenchymal cell marker VIM (red) in xenograft tumors and original ULM specimens in MED12m(+)- and (-)-ULMs, respectively. Scale bar = 50 μm. (C,D) Hierarchical clustering of xenograft tumors, primary ULM cells, and original ULM specimens using transcriptome data in MED12m(+)-ULM (n = 3) (C) and MED12m(-)-ULM (n = 3) (D), respectively. (E) Principal component analysis of xenograft tumors, primary ULM cells, and original ULM specimens using transcriptome data in MED12m(+)-ULM (n = 3) and MED12m(-)-ULM (n = 3).
Characterization of the xenograft tumor transcriptomes
Gene expression profiles of the xenograft tumors harvested 6 weeks after transplantation were compared with those of ULM specimens and ULM cells on day 3 of primary culture (Figs. 1C-E). In hierarchical clustering, xenograft tumors and ULM specimens were classified in the same cluster and were distinct from ULM cells in both subtypes (Fig. 1C, D). The PCA was able to distinguish xenograft tumors and ULM specimens from ULM cells (Fig. 1E). This suggests that the gene expression profile of the xenograft tumors is closer to that of the original ULM specimen.
Effects of FHW and UPA on xenograft tumors
The response to FHW and histological staining was similar in all 11 MED12m(+)-ULM samples analyzed (Table 2). The xenograft tumors were reduced in size (24–77% reduction) by EP withdrawal (E(-)P(-)). On the other hand, in MED12m(-)-ULM, 8 of the 11 samples reduced in size (22–51% reduction) by E(-)P(-) (Table 2), but the reductions of the remaining 3 samples were less than 10%. Since one of the goals of this study was to analyze the responsiveness of ULMs to FHW, we excluded the 3 weakly responsive specimens (Fig. S1) from the analysis.
Figure 2B shows photographs and tissue sections of xenograft tumors harvested 8 weeks after transplantation. In the MED12m(+)-ULM, the xenograft tumor size was significantly reduced in the E(+)P(-) group compared to the E(+)P(+) group, and was further reduced in the E(-)P(-) group (Fig. 2C, and Table 2). In the MED12m(-)-ULM, the xenograft tumor size was reduced in the E(+)P(-) group although the difference was not statistically significant, and it was significantly reduced in the E(-)P(-) group compared to the E(+)P(+) group (Fig. 2D, and Table 2). The SMC ratio was reduced in the E(-)P(-) group compared to the E(+)P(+) group (Figs. 2C, D), while the histological finding of HE staining and ACTA2 immunostaining in the E(-)P(-) group showed the presence of SMCs (Fig. 2B). This suggests that FHW caused the SMCs to shrink.
The effect of female hormone withdrawal (FHW) and ulipristal acetate (UPA), a selective progesterone receptor modulator (SPRM), on ULM cells in a xenograft model. (A) Schematic representation of the investigation of the effects of FHW and UPA on ULM cells, using a xenograft model. ULM cells were transplanted under the renal capsule of ovariectomized mice and grown by estrogen (E) and progesterone (P) depot for 4 weeks. Then, transplanted mice were divided into 4 groups: continued with E and P depot (E(+)P(+)), without E and P depot (E(-)P(-)), with E depot only (E(+)P(-)), and with E, P, and UPA depot (EP + UPA). The xenograft tumors were harvested 6 and 8 weeks after transplantation and used for transcriptome and histological analyses, respectively. (B) Representative images of overview, cross-section, and histology of each treated group of the xenograft tumors harvested 8 weeks after transplantation in MED12m(+)- and (-)-ULMs. As histology, images of HE staining, trichrome staining, and immunohistochemistry with ACTA2 (green), VIM (red), and PGR (red) are shown. Scale bar = 50 μm. (C,D) Box and whisker plot of the ratios of tumor volume, smooth muscle cells, collagen fiber, and fibroblasts of each treated group in each xenograft tumor of MED12m(+)-ULM (n = 11) (C) and MED12m(-)-ULM (n = 8) (D), respectively. **; p-value < 0.01, *: p-value < 0.05, (Bonferroni test).
The xenograft tumors in the UPA-treated group (EP + UPA) were expected to shrink to the same extent as in the P withdrawal group (E(+)P(-)). However, they were more regressed than in the E(-)P(-) group in both subtypes (Figs. 2C, D, and Table 2). The SMC ratio was further reduced in the EP + UPA group compared to the E(-)P(-) group in both subtypes (Figs. 2C, D). On the other hand, the fibroblast ratio increased in the EP + UPA group (Fig. 2C, D). Many ACTA2-positive cells (SMCs) disappeared in the EP + UPA group compared to the E(-)P(-) group (Fig. 2B), indicating that SMCs were lost in the EP + UPA group. In both subtypes, the collagen fiber ratio increased in the EP + UPA group compared to the E(-)P(-) group (Fig. 2C, D, and Table 2). Taken together, these histological findings suggest that fibroblasts and collagen fibers were replaced with SMCs due to the cell death of SMCs. Furthermore, the collagen turnover marker, the MMP1/TIMP1 ratio, was calculated from RNA-seq data. In MED12m(+) ULM, it was 0.019 ± 0.009 (mean ± SD) in the E(+)P(+) group and 0.020 ± 0.010 in the EP + UPA group. In MED12m(-) ULM, it was 0.007 ± 0.005 and 0.013 ± 0.014, respectively. Since the values are very low, it is suggested that the MMP1 function is fully suppressed by TIMP1 in both groups. Therefore, the elevated collagen fiber rate observed in the EP + UPA groups may be due to passive accumulation resulting from SMC loss rather than active fibrosis.
Next, we examined cell death. Analysis of apoptotic cells by TUNEL staining revealed that TUNEL-positive cells were most frequently detected in EP + UPA group in MED12m(-)-ULM and were also less frequently detected in EP + UPA group in MED12m(+)-ULM (Fig. 3). In the remaining treatment groups, TUNEL-positive cells were scarcely detected in either ULM subtype.
Detection of apoptotic cells in xenograft tumors by TUNEL staining. The images show the representatives in the xenograft tumors of the E(-)P(-) and EP + UPA groups in both ULM subtypes 8 weeks after transplantation. Rat mammary tissue was used as a positive control to detect the apoptotic cells. A sample of TdT enzyme withdrawal in the EP + UPA group (EP + UPA_TdT enzyme (-)) was used as a negative control to detect the apoptotic cells. This analysis used three specimens in each MED12m (+)- and (-)-ULM. In TUNEL staining, apoptotic cells were detected as the cells that bore dark red-brown nuclei (arrowheads). Scale bar = 50 μm. Because almost no apoptotic cells were detected in E(+)P(+) and E(+)P(-) groups in both ULM subtypes, their images were omitted.
To investigate whether UPA-induced cell death is mediated by host immune response, we performed immunostaining with markers for macrophages (CD68), myeloid cells (CD14), and leucocytes (CD45) because NOD/Scid mice lack almost all T and B cells. The cells positive for all three markers were detected in all treatment groups of both subtypes (Fig. S2).
DEGs in xenograft tumors induced by FHW and UPA
To examine the effects of FHW and UPA on gene expression, we identified differentially expressed genes (DEGs) in E(-)P(-), E(+)P(-), and EP + UPA groups compared to the E(+)P(+) group. In MED12m(+)-ULM, 139 upregulated- and 195 downregulated- DEGs were identified in the E(-)P(-) group by EP withdrawal (E(-)P(-) vs. E(+)P(+)), 85 upregulated- and 202 downregulated- DEGs in the E(+)P(-) group by P withdrawal (E(+)P(-) vs. E(+)P(+)), and 609 upregulated- and 613 downregulated- DEGs in the EP + UPA group by UPA administration (EP + UPA vs. E(+)P(+)) (Fig. 4A and Table S1). In MED12m(-)-ULM, 37 upregulated- and 102 downregulated- DEGs were identified by EP withdrawal, 67 upregulated- and 109 downregulated- DEGs by P withdrawal, and 266 upregulated- and 426 downregulated- DEGs by UPA administration (Fig. 4B and Table S2). In both subtypes, the number of DEGs was more than three times higher in UPA administration than in EP or P withdrawal.
Identification of differentially expressed genes (DEGs) in xenograft tumors by female hormone withdrawal and UPA administration. (A,B) Volcano plot showing the DEGs (red dots) identified in the E(-)P(-), E(+)P(-), and EP + UPA groups compared with the E(+)P(+) group in MED12m(+)-ULM (n = 3) (A) and MED12m(-)-ULM (n = 3) (B). DEGs were defined as the genes whose expression was more than 2-fold or less than 0.5-fold of that of the E(+)P(+) group in each group, and p-value < 0.05, (t-test). The number of the upregulated (Up) and downregulated (Down) DEGs are shown.
GO and pathway analyses of the DEGs induced by FHW and UPA
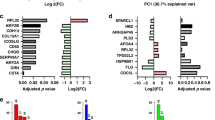
Figure 5 summarizes the effects of EP withdrawal (E(-)P(-)), P withdrawal (E(+)P(-)), and UPA administration (EP + UPA) on the GO terms and KEGG32,33 and IPA pathways. Further details of the GO, KEGG, and IPA analyses are given in Tables S3-S8, S9-S14, and S15-S20, respectively. The functional categories considered to be important for the development and progression of ULMs were reported previously34. Cell proliferation, signal transduction, immune system, angiogenesis, and myocyte function in the functional categories were similarly altered by EP withdrawal and P withdrawal in both ULM subtypes. ECM production, cell adhesion, and inflammation by EP and P withdrawal were prominent in MED12m(+)-ULM. TGFB signaling by P withdrawal and WNT signaling by EP withdrawal were observed in the MED12m(+)-ULM.
Summary of the GO terms, KEGG pathways, and IPA pathways altered by EP withdrawal, P withdrawal, and UPA administration. Gene ontology (GO) enrichment, KEGG pathway, and Ingenuity Pathway Analysis (IPA) pathway analyses were performed using DEGs in each treatment group in each subtype. The GO terms, KEGG pathways, and IPA pathways extracted in EP withdrawal (E(-)P(-)), P withdrawal (E(+)P(-)), and UPA administration (EP + UPA) were divided into functional categories which are considered to be important for the development of ULM, in MED12m(+)- and (-)-ULMs, respectively. The number of GO terms and pathways was manually tabulated and expressed by babble sizes with the numbers.
UPA administration caused remarkable changes in all functions compared with EP or P withdrawal in both ULM subtypes. In particular, the immune system and inflammation were prominent in the MED12m(+)-ULM, while apoptosis was prominent in the MED12m(-)-ULM.
Among the DEGs, the genes involved in apoptosis (TP53 and BCL2L11) and immune response (CD34, CD14, and AKT3) that showed significant differences between E(+)P(+) and EP + UPA groups were selected and validated by real-time RT-PCR (Fig. S3). Significant differences were confirmed between E(+)P(+) and EP + UPA groups for TP53 and CD34. Although no statistically significant differences were observed for BCL2L11, CD14, and AKT3, the expression level trends in the E(+)P(+) and EP + UPA groups were consistent with those observed in the RNA-seq data.
Discussion
Although the ULM subtypes showed no histological differences in the effects of FHW, transcriptome and pathway analyses indicated that FHW caused these subtypes to regress through different signaling pathways, which seems to be consistent with our previous findings that MED12m(+)- and (-)-ULMs develop through different signaling pathways11. In particular, in the MED12m(+)-ULMs, FHW suppressed the pathways involved in fibrosis, such as ECM production and cell adhesion (Tables S3, S4, S9, S10, S15, and S16), and altered the pathways involved in cell proliferation, including TGFB and WNT signaling. This may reflect differences in the pathophysiology of the two ULM subtypes. We previously showed differences in the transcriptome and signaling pathways between ULM subtypes11. Compared to the normal myometrium, ECM production, cell adhesion, and WNT signaling were activated in MED12m(+)-ULM, whereas ECM production, cell adhesion, and TGFB and WNT signaling were inactivated in MED12m(-)-ULM. Thus, each ULM subtype develops and regresses through different signaling pathways.
Since the main action of UPA is the inhibition of PGR function, tumor regression was expected to be comparable to P withdrawal. Interestingly, EP + UPA was more effective than E(-)P(-) at reducing the xenograft tumor size, due to apoptosis of SMCs. This effect of UPA appears to be quite different from that of RU486, which is a blocker of PGR. Studies using the same xenograft model as in the present study showed that tumor regression by RU486 and EP withdrawal was due to the shrinkage and not death of SMCs12,29,30. In general, ULM patients who discontinue GnRHa treatment relapse within several weeks. This is because SMCs resume development in the presence of female hormones. On the other hand, the results from European Tier III and Japanese Tier II studies, which investigated the effects of UPA on ULM, showed no regrowth of ULM in the 25 weeks after cessation of UPA treatment35,36. Taking these results into consideration, it is suggested that UPA can kill SMCs by inducing apoptosis via a mechanism other than inhibiting PGR function. However, further studies are needed to investigate whether UPA-induced SMC death is UPA-specific and whether xenograft tumors regrow after the cessation of UPA treatment.
UPA administration had much greater effects than P withdrawal on the transcriptomes and signaling pathways of xenograft tumors, suggesting that UPA also has direct or indirect effects on SMCs besides inhibition of PGR. In our recent study using ULM spheroid cultures37, UPA shrank the ULM spheroid in a manner similar to P withdrawal but did not kill SMCs. Therefore, the death of SMCs in the xenograft model may have been caused by host-mediated actions of UPA. It is interesting to note that the immune system and inflammation were strongly promoted by UPA administration in the MED12m(+)-ULMs in this study, while apoptosis was activated in the MED12m(-)-ULMs (Tables S5, S8, S11, S14, S17, and S20). We speculated that UPA through host-mediated immunological responses, not only killed SMCs by cytotoxicity in the MED12m(+)-ULMs but also caused apoptosis of SMCs in the MED12m(-)-ULMs. The mechanism of host-mediated UPA action may be similar to the mechanism of immunogenic cell death (ICD) which has been reported in various cancers in recent years38,39. ICD is a mechanism by which the molecules released from cancer cells in response to specific anticancer drugs enhance the host’s immune response against cancer cells, leading to their death. Therefore, to investigate whether UPA-induced cell death is mediated by host immune response, we performed immunostaining with markers for macrophages (CD68), myeloid cells (CD14), and leucocytes (CD45). Contrary to our expectations, the characteristic staining specific to UPA was not found, suggesting that host immune responses with these immune cells are unlikely to be involved in UPA-specific cell death of SMCs. Further studies are needed to clarify how UPA caused remarkable changes in signaling pathways and GO terms, how these different pathways are selectively utilized between the ULM subtypes upon UPA exposure, and how UPA caused cell death of SMCs.
Although all MED12m(+)-ULM samples showed that FHW caused tumor regression, 3 of the 11 MED12m(-)-ULM samples did not show any regression as a result of FHW. In the three FHW weakly responsive ULMs, the expression of PGR in the xenograft tumors of all treatment groups was confirmed to be comparable to that in sensitive ULMs 8 weeks after transplantation (Fig. S1). These ULMs may be female hormone-independent ULMs that can develop even in the absence of female hormones. We will investigate the pathophysiology of these female hormone-independent ULMs using whole-genome sequencing-based mutational profiling in future studies.
In this study, we investigated the effects of FHW and UPA on each ULM subtype using a xenograft model. Although FHW and UPA did not result in any histological differences between the MED12m(+)- and (-)-ULM xenograft tumors, transcriptome and pathway analyses revealed that the xenograft tumors of both subtypes regressed through different signaling pathways. The present results also suggest that mechanisms other than inhibition of PGR function are involved in UPA-induced tumor regression of ULMs.
Methods
Ethical statement
This study was approved by the Institutional Review Board of Yamaguchi University Graduate School of Medicine (No. H27-035). Informed consent was obtained from the patients before any samples were collected. All experiments involving human tissue were performed according to the Declaration of Helsinki. All animal procedures were approved by the Animal Experimentation Ethics Committee of Yamaguchi University Graduate School of Medicine, performed according to the relevant guidelines and regulations, and confirmed compliance with ARRIVE guidelines.
Patient tissue samples
For this study, ULM specimens were obtained from 22 Japanese women who underwent hysterectomy at the Department of Obstetrics and Gynecology at Yamaguchi University Hospital (Table 1). MED12m(+)-ULM specimens were obtained from 11 women (aged 27 to 49 years, mean ± SD; 38.5 ± 7.7). MED12m(-)-ULM specimens were obtained from 11 women (aged 37 to 49 years, mean ± SD; 44.4 ± 3.9). None of the women had received previous treatment with female hormones or gonadotropin-releasing hormone analogs. Dissected specimens were provided for primary culture, formalin-fixed paraffin-embedded blocks for histological analysis, and stored at -80 °C for genomic DNA and total RNA isolation.
Genomic DNA and total RNA isolation
Genomic DNA from ULM specimens and total RNA from ULM specimens, cells, and xenograft tumors were isolated as previously reported11,40,41. In brief, the genomic DNA from ULM specimens was isolated by treatment with Proteinase K (Qiagen, Hilden, Germany), followed by phenol/chloroform extraction and ethanol precipitation. Total RNA from ULM specimens and xenograft tumors was isolated by treatment with ISOGEN reagent (Nippon Gene, Tokyo, Japan), followed by chloroform extraction and 2-propanol precipitation. Total RNA from ULM cells was isolated using an RNeasy mini kit (Qiagen) according to the manufacturer’s instructions.
MED12 exon2 genome or transcript sequencing
The exon 2 region of the MED12 gene in the ULM specimens was sequenced as previously reported3,8,9,10. In brief, genomic PCR or RT-PCR was performed using PrimeSTAR GXL DNA polymerase (Takara, Kyoto, Japan) and primer sets as follows 5’-GCCCTTTCACCTTGTTCCTT-3’ and 5’-TGTCCCTATAAGTCTTCCCAACC-3’ for genomic PCR, or 5’-CAGGACCCCAAACAGAAGGAG-3’ and 5’-CAATAATGCTGCTGAAGTTGGAAC-3’ for RT-PCR, under thermocycling conditions (35 cycles of 98 °C for 10 s, 60 °C for 15 s, and 68 °C for 20 s). The amplified PCR products were purified using the QIAquick PCR Purification Kit (Qiagen) and sequenced using the BigDye Terminator v3-1 Cycle Sequencing Kit (Applied Biosystems, Carlsbad, CA). The resulting sequences were analyzed manually.
Primary culture of ULM cells
ULM cells were isolated from ULM specimens as previously reported31,37. In brief, ULM specimens were cut with a razor into 1–2 mm square pieces and incubated with 5-fold amounts of 1.5 mg/ml collagenase type I (Wako, Osaka, Japan) and 10 µg/ml Dnase I (Roche, Basel, Switzerland) in HBSS at 37 °C for 6 h with shaking at 250 rpm. The ULM cells were passed through a 100 μm cell strainer (Falcon, Corning, NY) to remove the debris, seeded into 10 cm collagen-coated dishes (Corning, Corning, NY), and maintained in DMEM/F12 (Wako) containing 10% FBS (Gibco, Rockville, MD), 2mM L-glutamine (Gibco), and 1% antibiotic-antimycotic (Wako) at 37 °C in a humidified atmosphere of 5% CO2 and 95% air. The percentage of ACTA2 (an SMC marker) positive cells (Fig. 1A) averaged 88.1 ± 6.7% and 85.3 ± 8.4% in MED12m(+)- and (-)-ULMs, respectively, indicating that SMCs are the majority of the ULM cells. Almost all SMCs were confirmed to be PGR-positive (Fig. 1A).
Optimization of the UPA depot
Since there are no reports of UPA depot administration in mice, the optimal dose was investigated using estrous persistence as an indicator of PGR function inhibition. UPA was provided by ASKA Pharmaceutical Co., Ltd. (Tokyo, Japan). Since 0.03–0.15 mg of UPA solution/mouse/day has been reported for daily subcutaneous administration, we tested 0.3 and 1.0 mg of UPA depot/mouse/week42,43,44. UPA at 0 mg (control, sesame oil only), 0.3 mg, or 1.0 mg was mixed with sesame oil and administered subcutaneously to each of three female ICR mice once a week. Daily smear checks were performed on the treated mice along with 3 untreated mice (IDs #9–11). The estrous cycle was observed for 2 weeks (Figs. S4A, B). In control mice (IDs #6–8), estrus lasted 1 or 2 days and transitioned to the metestrus similar to the normal cycle in untreated mice (IDs #9–11). In the 0.3 mg UPA-treated mice, estrus was prolonged in only one of the three mice (ID #4), but not in the other two mice (IDs #3 and 5). In the 1.0 mg UPA-treated mice, estrus was prolonged from 3 to 5 days in all mice (Fig. S4A). Therefore, the UPA depot dose in this study was determined to be 1.0 mg/mouse/week.
Xenograft models
Xenografts were performed according to the protocol of Kurita et al.31,45 with some modifications. In brief, ULM cells on day 3 of the primary culture were mixed with collagen solution [40% rat tail collagen type I (final concentration 1.6 mg/ml; Sigma-Aldrich, Tokyo, Japan), 20% 5 × DMEM, 10% FBS, 10% 0.44 M NaHCO3 and 5% 0.18 N NaOH] at 1 × 107 cells/ml and incubated at 37 °C for 30 min to form cell-collagen pellets. The cell-collagen pellets were cultured for 1 day and transplanted (6 × 105 cells/kidney) under both kidney capsules of ovariectomized severely immunodeficient mice (NOD/ShiJic-scidJcl, Kyudo, Tosu, Japan). To grow the transplanted ULM cells, E (300 µg Progynon/week, Fujipharma, Tokyo, Japan) and P (3.75 mg Progeston/week, Fujipharma) depots were administered subcutaneously to the transplanted mice once a week according to the previous report45.
A schematic representation of the FHW and UPA administration experiments in the xenograft model is shown in Fig. 2A. To evaluate the effects of FHW and UPA, transplanted mice were divided into 4 groups 4 weeks after transplantation: continued with E and P depot (E(+)P(+)), without E and P depot (E(-)P(-)), with E depot only (E(+)P(-)), and with E, P, and UPA depot (EP + UPA). The xenograft tumors were harvested 8 weeks after transplantation (Fig. 2B). All of the xenograft tumors transplanted successfully engrafted in this study. Size was assessed by measuring the diameter of the major and minor axes and the height of the cut surface of the xenograft tumors. Volume was calculated as π/6 x major diameter x minor diameter x height. Samples with an average volume of less than 3.0 mm3 in the E(+)P(+) group were considered to have no proliferative potential and were excluded from this study. 73.3% (11 out of 15) and 57.9% (11 out of 19) of the specimens were 3.0 mm³ or larger in MED12m(+)-ULM and MED12m(-)-ULM, respectively. Eight weeks after transplantation, PGR expression was confirmed in all treatment groups of both ULM subtypes (Fig. 2B).
Immunofluorescent staining
To stain cell cultures, ULM cells on day 3 of the primary culture were double-stained with ACTA2 and PGR as previously reported37. In brief, the cells were fixed with 4% paraformaldehyde (PFA) for 15 min, blocked with a blocking solution (10% fetal bovine serum and 1% bovine serum albumin in PBST) for 60 min, incubated with a mouse anti-ACTA2 monoclonal antibody (Abcam, Tokyo, Japan, RRID: AB_262054) and rabbit anti-PGR monoclonal antibody (Abcam, RRID: AB_443421) as a primary antibody (diluted at 1:500 in the blocking solution) at 4 °C overnight, and incubated with the Alexa Fluor 488 conjugated anti-mouse IgG (Abcam, RRID: AB_2576208) and Alexa Fluor 594 conjugated anti-rabbit IgG (Abcam, RRID: AB_2734147) as a secondary antibody (diluted at 1:1000 in PBS) for 45 min. Nuclei were stained with DAPI (500 ng/ml, Nacalai, Tokyo, Japan).
To stain tissue sections, sections were double-stained with ACTA2 and VIM (a mesenchymal cell marker), PGR, CD14, CD45, or CD68 as previously reported37. In brief, tissue sections (5 μm) of formalin-fixed paraffin-embedded samples were deparaffinized, fixed with 4% PFA for 5 min, blocked with blocking solution for 60 min, incubated with mouse anti-ACTA2 antibody and rabbit anti-VIM antibody (Abcam, RRID: AB_10562134), rabbit anti-PGR antibody, rabbit anti-CD14 antibody (Abcam, RRID: AB_2935854), rabbit anti-CD45 antibody (Abcam, RRID: AB_442810), or rabbit anti-CD68 antibody (Abcam, RRID: AB_10975465) as a primary antibody at 4 °C overnight, incubated with the Alexa Fluor 488 conjugated anti-mouse IgG and Alexa Fluor 594 conjugated anti-rabbit IgG as a secondary antibody for 45 min, and counter-stained with DAPI. The percentage of SMCs (ACTA2 positive cells) (SMC ratio (%)) was calculated using ImageJ for ten randomly chosen areas at x 400 magnification and expressed as mean ± SD.
Trichrome staining
Collagen fibers were visualized using the Trichrome Stain Kit (TRM-1, ScyTec Laboratories Inc., Utah, USA) as previously reported37. Tissue sections (5 μm) of paraffin-embedded samples were deparaffinized, fixed with Bouin’s solution overnight, stained with Weigert’s Iron Hematoxylin for 10 min, stained with Biebrich Scarlet/Acid Fuchsin for 10 min, differentiated in Phosphomolybdic/Phosphotungstic Acid for 15 min, stained with Aniline Blue for 15 min, and immersed in 1% Acetic Acid for 1 min. The area of collagen fibers, which were stained blue, was quantified by ImageJ. The collagen fiber ratio (%) was calculated for 5 randomly chosen areas at x 200 magnification and expressed as mean ± SD.
TUNEL staining
Apoptotic cells were visualized using an In Situ Apoptosis Detection Kit (Takara) according to the manufacturer’s instructions37. The sections (5 μm) of paraffin-embedded samples were deparaffinized, permeabilized with 20 µg/ml proteinase K (Qiagen) for 15 min, intrinsic peroxidase-blocked with 3% H2O2 for 5 min, incubated with terminal deoxynucleotide transferase (TdT enzyme) at 37 °C for 60 min, incubated with an anti-FITC HRP conjugated antibody at 37 °C for 30 min, incubated with diaminobenzidine for 20 min, and counter-stained with methyl green for 20 min. Slides of rat mammary glands included in the kit were used as a positive control of apoptotic cells. Apoptotic cells were checked on 5 randomly chosen areas at x 200 magnification in xenograft tumor of all treatment group in three specimens in each MED12m(+)- and (-)-ULM. Quantitative analysis could not be performed because some samples were not stained by the counterstain.
Transcriptome analysis and identification of the differentially expressed genes (DEGs)
Transcriptomes were obtained as previously reported46. Since xenograft tumors 8 weeks after transplantation have an increased amount of collagen fibers with a decreased number of viable cells, the xenograft tumors 6 weeks after transplantation were used. The RNA-sequencing library was prepared from 100 ng of total RNA using a NEBNext Ultra II RNA Library Prep Kit for Illumina (NEB) and sequenced on an Illumina NextSeq (Illumina, San Diego, CA). Mapping and quantification of the reads were performed by CLC Genomics Workbench (Qiagen).
To identify the differentially expressed genes (DEGs) of the xenograft tumors in each treatment group, the gene expression levels of the three samples in each group were normalized by DESeq2 software47. Then, the genes whose expression was significantly (p-value < 0.05 (t-test)) more than 2-fold or less than 0.5-fold of that of the E(+)P(+) group in each treatment group were defined as DEGs.
Furthermore, mutation analysis of MED12 transcripts confirmed the persistence of MED12-mutant cells in MED12m(+)-ULM xenograft tumors (Fig. S5).
Quantitative real-time RT-PCR
One µg total RNA was reverse-transcribed using a QuantiTect Reverse Transcription Kit (Qiagen) according to the manufacturer’s protocol as previously reported10. A primer pair for Mitochondrial Ribosomal Protein L19 (MRPL19) was used as an internal control. Real-time RT-PCR was performed using Luna Universal qPCR Master Mix (NEB) and a CFX384 Real-time PCR System (BioRad, Hercules, CA). Melting curves of the products were obtained after cycling by a stepwise increase of temperature from 55 to 95 °C. All samples were run in triplicate. The relative quantity of cDNA was calculated with the ∆∆Ct method. The primer sequences used in this analysis are shown in Supplemental Table S21.
Hierarchical clustering, PCA, IPA, GO, and KEGG analysis
Hierarchical clustering was performed by the open resource software MultiExperiment Viewer (MeV; http://www.tm4.org/mev.html) using the data of all 47467 genes. Principle component analysis (PCA) was performed using the prcomp function in an R.
The DEGs from EP withdrawal (E(-)P(-) vs. E(+)P(+)), P withdrawal (E(+)P(-) vs. E(+)P(+)), and UPA (EP + UPA vs. E(+)P(+)) were provided for the canonical pathway analysis using an Ingenuity Pathway Analysis (IPA) software (Ingenuity Systems, Redwood City, CA, USA), and for gene ontology (GO) enrichment analysis and KEGG pathway analyses using a Metascape software48. GO and KEGG analyses were performed separately on the up-and down-regulated DEGs. Furthermore, to summarize the results, the GO terms, KEGG pathways, and IPA pathways extracted in EP withdrawal, P withdrawal, and UPA administration were divided into functional categories which are considered to be important for the development of ULM. The number of GO terms, KEGG pathways, and IPA pathways was manually tabulated (Fig. 5).
Statistical analysis
The significance of differences was analyzed using the Bonferroni, Kruskal-Wallis, and Mann-Whitney tests. P values < 0.05 were considered to indicate statistical significance. All statistical analyses were performed using the R software program (Ver 3.6.1).
Data availability
The datasets generated during and/or analyzed during the current study have been deposited in the DDBJ BioProject database (https://www.ddbj.nig.ac.jp/index.html) with links to BioProject accession number PRJDB23168.
References
Stewart, E. A. Uterine fibroids. Lancet 357, 293–298 (2001).
Bajekal, N. & Li, T. C. Fibroids, infertility and pregnancy wastage. Hum. Reprod. Update. 6, 614–620 (2000).
Mäkinen, N. et al. MED12, the mediator complex subunit 12 gene, is mutated at high frequency in uterine leiomyomas. Science 334, 252–255 (2011).
Mehine, M. et al. Characterization of uterine leiomyomas by whole-genome sequencing. N Engl. J. Med. 369, 43–53 (2013).
Mittal, P. et al. Med12 gain-of-function mutation causes leiomyomas and genomic instability. J. Clin. Invest. 125, 3280–3284 (2015).
McGuire, M. M. et al. Whole exome sequencing in a random sample of North American women with leiomyomas identifies MED12 mutations in majority of uterine leiomyomas. PLoS One. 7, e33251. https://doi.org/10.1371/journal.pone.0033251 (2012).
Bertsch, E. et al. MED12 and HMGA2 mutations: two independent genetic events in uterine leiomyoma and leiomyosarcoma. Mod. Pathol. 27, 1144–1153 (2014).
Tamehisa, T. et al. Establishment of noninvasive prediction models for the diagnosis of uterine leiomyoma subtypes. Obstet. Gynecol. 143, 358–365 (2024).
Sato, S. et al. Identification of uterine leiomyoma-specific marker genes based on DNA methylation and their clinical application. Sci. Rep. 6, 30652. https://doi.org/10.1038/srep30652 (2016).
Sato, S. et al. Potential mechanisms of aberrant DNA hypomethylation on the x chromosome in uterine leiomyomas. J. Reprod. Dev. 60, 47–54 (2014).
Maekawa, R. et al. Different DNA methylome, transcriptome and histological features in uterine fibroids with and without MED12 mutations. Sci. Rep. 12 https://doi.org/10.1038/s41598-022-12899-7 (2022).
Wu, X. et al. Subtype-Specific Tumor-Associated fibroblasts contribute to the pathogenesis of uterine leiomyoma. Cancer Res. 77, 6891–6901 (2017).
Nagai, K. et al. MED12 mutations in uterine leiomyomas: prediction of volume reduction by gonadotropin-releasing hormone agonists. Am. J. Obstet. Gynecol.. 228, 207 e1- e9 https://doi.org/10.1016/j.ajog.2022.09.024 (2023).
Rabe, T. et al. Selective progesterone receptor modulators for the medical treatment of uterine fibroids with a focus on ulipristal acetate. Biomed. Res. Int. 1374821 https://doi.org/10.1155/2018/1374821 (2018).
Shimomura, Y., Matsuo, H., Samoto, T. & Maruo, T. Up-regulation by progesterone of proliferating cell nuclear antigen and epidermal growth factor expression in human uterine leiomyoma. J. Clin. Endocrinol. Metab. 83, 2192–2198 (1998).
Yin, P. et al. Progesterone receptor regulates Bcl-2 gene expression through direct binding to its promoter region in uterine leiomyoma cells. J. Clin. Endocrinol. Metab. 92, 4459–4466 (2007).
Luo, N. et al. Estrogen-mediated activation of fibroblasts and its effects on the fibroid cell proliferation. Transl Res. 163, 232–241 (2014).
Milewska, G. et al. Functional evidence for two distinct mechanisms of action of progesterone and selective progesterone receptor modulator on uterine leiomyomas. Fertil. Steril. 122, 341–351 (2014).
Maruo, T., Ohara, N., Wang, J. & Matsuo, H. Sex steroidal regulation of uterine leiomyoma growth and apoptosis. Hum. Reprod. Update. 10, 207–220 (2004).
Yamada, T. et al. Progesterone down-regulates insulin-like growth factor-I expression in cultured human uterine leiomyoma cells. Hum. Reprod. 19, 815–821 (2004).
Bonazza, C. et al. Primary human uterine leiomyoma cell culture quality control: some properties of myometrial cells cultured under serum deprivation conditions in the presence of ovarian steroids. PLoS One. 11, e0158578. https://doi.org/10.1371/journal.pone.0158578 (2016).
Xu, Q. et al. Progesterone receptor modulator CDB-2914 down-regulates proliferative cell nuclear antigen and Bcl-2 protein expression and up-regulates caspase-3 and Poly (adenosine 5’-diphosphate-ribose) Polymerase expression in cultured human uterine leiomyoma cells. J. Clin. Endocrinol. Metab. 90, 953–961 (2005).
Xu, Q. et al. Progesterone receptor modulator CDB-2914 induces extracellular matrix metalloproteinase inducer in cultured human uterine leiomyoma cells. Mol. Hum. Reprod. 14, 181–191 (2008).
Xu, Q. et al. Progesterone receptor modulator CDB-2914 down-regulates vascular endothelial growth factor, adrenomedullin and their receptors and modulates progesterone receptor content in cultured human uterine leiomyoma cells. Hum. Reprod. 21, 2408–2416 (2006).
Bloch, J., Holzmann, C., Koczan, D., Helmke, B. M. & Bullerdiek, J. Factors affecting the loss of MED12-mutated leiomyoma cells during in vitro growth. Oncotarget 8, 34762–34772 (2017).
Nadine, M. D. et al. Cell cultures in uterine leiomyomas: rapid disappearance of cells carrying MED12 mutations. Genes Chromosomes Cancer. 53, 317–323 (2014).
Severino, M. F. et al. Rapid loss of oestrogen and progesterone receptors in human leiomyoma and myometrial explant cultures. Mol. Hum. Reprod. 2, 823–828 (1996).
Zaitseva, M., Vollenhoven, B. J. & Rogers, P. A. In vitro culture significantly alters gene expression profiles and reduces differences between myometrial and fibroid smooth muscle cells. Mol. Hum. Reprod. 12, 187–207 (2006).
Ishikawa, H. et al. Progesterone is essential for maintenance and growth of uterine leiomyoma. Endocrinology 151, 2433–2442 (2010).
Serna, V. A. et al. Cellular kinetics of MED12-mutant uterine leiomyoma growth and regression in vivo. Endocr. Relat. Cancer. 25, 747–759 (2018).
Serna, V. A. & Kurita, T. Patient-derived xenograft model for uterine leiomyoma by sub-renal capsule grafting. J. Biol. Methods. 5, e91. https://doi.org/10.14440/jbm.2018.243 (2018).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Sugino, N. et al. Uterine fibroids and adenomyosis. (ed. Sugino, N.) 1-173 (Springer, 2018).
Donnez, J. et al. Ulipristal acetate versus leuprolide acetate for uterine fibroids. N Engl. J. Med. 366, 421–432 (2012).
Irahara, M., Maejima, Y., Shinbo, N., Yamauchi, Y. & Mizunuma, H. Ulipristal acetate for Japanese women with symptomatic uterine fibroids: A double-blind, randomized, phase II dose-finding study. Reprod. Med. Biol. 19, 65–74 (2019).
Sakai, T. et al. Establishment of a 3D spheroid culture system to evaluate the responsiveness of uterine leiomyoma cells to female hormones. Reprod. Med. Biol. 24, e12627. https://doi.org/10.1002/rmb2.12627 (2025).
Ahmed, A. & Tait, S. W. G. Targeting Immunogenic cell death in cancer. Mol. Oncol. 2994–3006 (2020).
Meier, P., Legrand, A. J., Adam, D. & Silke, J. Immunogenic cell death in cancer: targeting necroptosis to induce antitumour immunity. Nat. Rev. Cancer. 24, 299–315 (2024).
Sato, S. et al. SATB2 and NGR1: potential upstream regulatory factors in uterine leiomyomas. J. Assist. Reprod. Genet. 36, 2385–2397 (2019).
Maekawa, R. et al. Genome-wide DNA methylation analysis reveals a potential mechanism for the pathogenesis and development of uterine leiomyomas. PLoS One. 8, e66632. https://doi.org/10.1371/journal.pone.0066632 (2013).
Nair, H. B. et al. EC313-a tissue selective SPRM reduces the growth and proliferation of uterine fibroids in a human uterine fibroid tissue xenograft model. Sci. Rep. 9 https://doi.org/10.1038/s41598-019-53467-w (2019).
Liang, B. et al. Efficacy, safety and recurrence of new progestins and selective progesterone receptor modulator for the treatment of endometriosis: a comparison study in mice. Reprod. Biol. Endocrinol. 16 https://doi.org/10.1186/s12958-018-0347-9 (2018).
Small, B. et al. The selective progesterone receptor modulator ulipristal acetate inhibits the activity of the glucocorticoid receptor. J. Clin. Endocrinol. Metab. 105, 716–734 (2010).
Wang, G. et al. Nonobese diabetic/severe combined immunodeficient murine xenograft model for human uterine leiomyoma. Fertil. Steril. 101, 1485–1492 (2014).
Shirafuta, Y. et al. Integrated analysis of transcriptome and histone modifications in granulosa cells during ovulation in female mice. Endocrinology 162, bqab128. https://doi.org/10.1210/endocr/bqab128 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated Estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10 https://doi.org/10.1038/s41467-019-09234-6 (2019).
Acknowledgements
This work was supported in part by ASKA Pharmaceutical Co., Ltd. to Sato S, Sakai T, Tamehisa T, and Sugino N, and JSPS KAKENHI Grants (JP24K12533 to Sato S, JP24K12579 to Tamura I, JP24K12626 to Kajimura T, JP23K27734 and JP22K19603 to Sugino N) for Scientific Research from the Ministry of Education, Science, and Culture, Japan.
Author information
Authors and Affiliations
Contributions
S.S.: Conceptualization, investigation, visualization, funding acquisition, methodology, bioinformatics, writing—original draft, writing–review and editing. T.S.: Resources, investigation, visualization, funding acquisition, statistical analysis. T.T.: Resources, investigation, funding acquisition, statistical analysis. A.S.: RNA-seq analysis, visualization, statistical analysis, bioinformatics. H.T.: Resources, investigation. T.K.: Investigation, funding acquisition. I.T.: Investigation, methodology, funding acquisition. R.M.: RNA-seq analysis, bioinformatics. N.S.: Conceptualization, supervision, funding acquisition, writing–original draft, project administration, writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
Sato S, Sakai T, Tamehisa T, and Sugino N received research funding from ASKA Pharmaceutical Co., Ltd. The other authors declare that there are no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sato, S., Sakai, T., Tamehisa, T. et al. Effects of female hormone withdrawal and ulipristal acetate on MED12 mutation positive and negative uterine leiomyomas using a xenograft model. Sci Rep 15, 40028 (2025). https://doi.org/10.1038/s41598-025-23721-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23721-5