Abstract
Type VII secretion system (T7SS) substrate EsxA contributes to the pathogenesis of several Gram-positive bacteria, but its role in viridans streptococci-induced infective endocarditis (IE) remains unclear. Genomic analysis of the Streptococcus gordonii DL1 strain identified a genetic locus encoding five T7SSb core machinery proteins (EsaA, EssA, EsaB, EssB, and EssC) and one substrate, EsxA. T7SSb mediates EsxA secretion, as evidenced by reduced secretion in the essC-deleted mutant and restoration upon complementation. In a rat model of IE, the esxA-deficient strain shows lower survival in circulation and reduced formation of vegetation, suggesting a role for EsxA in IE pathogenesis. Using a transwell system, molecules secreted from wild-type S. gordonii DL1 can induce the release of neutrophil extracellular traps (NETs) by neutrophils, and this activity is lost in the esxA-deficient strain. In addition, recombinant EsxA can dose-dependently induce NET formation, confirming a role for secreted EsxA in the induction of NET formation. Consistently, reduced and restored NET formation is observed in vivo in vegetations caused by esxA-deficient and complemented strains, respectively. Together, these data indicate that EsxA secretion mediated by T7SSb induces NET formation that contributes to bacterial survival in the circulation and vegetation formation in S. gordonii-induced IE.
Similar content being viewed by others
Introduction
Infective endocarditis (IE) is an infectious disease of the heart valves that has high mortality and is usually refractory to antibiotic treatment primarily due to the tendency of causative bacteria to form biofilms1. Blood-borne streptococci from gingival wounds in the oral cavity or staphylococci from contaminated medical devices are the most common pathogens responsible for IE2. These microbes can evade immune surveillance in the bloodstream and colonize injured heart valves, leading to the formation of lesions termed vegetations. Vegetations are characteristic of IE and are thrombi composed of fibrin, aggregated platelets, and bacterial biofilms3. In addition, our previous studies demonstrated that platelets promote biofilm formation on the heart valve endothelium and that bacterial aggregates embedded inside the platelets can recruit neutrophils to promote the release of neutrophil extracellular traps (NETs)4,5.
The formation of NETs, also known as NETosis, is a distinct form of programmed neutrophil cell death first identified in 19966. A series of coordinated steps underlying the release of nuclear DNA during NETosis has been proposed7,8. Briefly, NET-inducing stimuli trigger a burst of reactive oxygen species (ROS) via the MEK/ERK/PKC or JNK signaling pathways9,10. ROS promotes the translocation of neutrophil elastase (NE) to the nucleus to partially cleave histones and also induces peptidylarginine deiminase 4 (PAD4)-mediated histone citrullination, both contributing to chromatin decondensation7,8,11. Eventually, breakdown of the nuclear envelope occur, driven by PKCα-mediated phosphorylation of lamin B and CDK4/6-mediated phosphorylation of lamin A/C12,13, followed by disruption of the plasma membrane, associated with cytoskeletal remodeling, pore formation, and NET extrusion14.
The formation of NETs, along with phagocytosis, constitutes a key innate mechanism by which neutrophils eradicate bacterial infections15,16. NETs were initially reported to trap and kill bacteria by granule enzymes, but results of later studies showed that NETs do not possess direct cytotoxic activity17. Instead, bacteria can evade neutrophil phagocytic killing by inducing NET formation18. Molecules within NETs, including elastase, myeloperoxidase (MPO), cathepsin, and histones can also trigger formation of immunothrombosis in blood vessels, which has been implicated in the pathogenesis of several thrombosis-related diseases, such as atherosclerosis and disseminated intravascular coagulation in sepsis8,19. Our previous study also demonstrated the critical roles of NETs in vegetation expansion in oral streptococci- and S. aureus-induced IE and in central venous thrombosis formation5,20. Although NET formation induced by bacteria contributes to bacterial pathogenesis, the specific bacterial molecules directly responsible for inducing NETs remain unclear.
Bacterial secretion systems play a central role in modulating the secretion of bacterial virulence factors and bacterial pathogenesis21,22,23. Based on the structures and secretion machinery, these bacterial secretion systems can be classified into at least 13 different types (Type I to Type XI, Sec and Tat)24,25,26,27. Most secretion systems are found in Gram-negative bacteria, but the Type VII secretion system (T7SS) was initially discovered in Mycobacterium tuberculosis (Mtb) and was subsequently identified in Firmicutes (designated T7SSa and T7SSb, respectively)28,29. Among Firmicutes, the mechanism and components of T7SSb have been thoroughly investigated in S. aureus. The core machinery of S. aureus T7SSb comprises five proteins including one cytoplasmic protein, EsaB, and four membrane proteins, EsaA, EssA, EssB, and EssC30. EssC encodes an FtsK/SpoIIIE-like ATPase and its C-terminal domain is critical for recognition and export of T7SS substrates31. The best-studied T7SS effector proteins are WXG100 proteins, which are small alpha-helical proteins (approximately 100 amino acids) that contain a central WXG (tryptophan-variable-glycine) motif within a hairpin loop between two alpha-helices32,33,34. The WXG100 protein EsxA was the first T7SS substrate to be identified and is encoded within the T7SS gene locus33. EsxA appears to serve two putative roles: as a secreted effector with cytotoxic activity against host cells and potentially competing bacteria, and as a chaperone-like protein that facilitates the export of other T7SS substrates or effectors30,35,36,37. LXG toxins and orphan WXG100 proteins have since been identified as T7SS substrates or effectors, and additional substrates are likely yet to be discovered29. The extensive diversity of T7SS and its substrates or effectors contributes to species-specific functions. These diverse functions may contribute to the ability of bacteria to rapidly adapt to environmental changes and promote survival or pathogenesis within hosts. In S. aureus, T7SS variants accompanying secreted effector proteins have been shown to contribute to distinct functions in different disease models30,38,39. In addition, T7SS has been shown to play an important role in Group B Streptococcus-induced meningitis40 and in Streptococcus gallolyticus-associated tumor promotion in the colon41.
Despite a growing number of studies highlighting the role of T7SS effector proteins in bacterial pathogenesis, their functions have not yet been characterized in viridans streptococci. Here, we investigated the role of the T7SS substrate EsxA in S. gordonii-induced IE. Using a rat model of IE, we demonstrate that S. gordonii EsxA plays a critical role in disease pathogenesis. We also show that EsxA can induce neutrophils to release NETs, which contribute to bacterial survival in circulation and promote vegetation expansion in IE.
Results
EsxA secretion is mediated by EssC in S. gordonii DL1
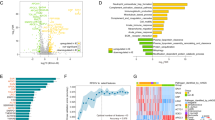
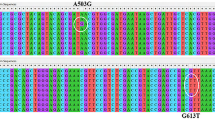
To investigate the role of T7SSb in S gordonii pathogenesis, we compared the S. gordonii T7SSb gene cluster with that of other Gram-positive bacteria, including Streptococcus agalactiae (GBS), Streptococcus gallolyticus, Staphylococcus aureus and Listeria monocytogenes (Fig. 1). The sequences of six genes of T7SSb, including the effector esxA and five core machinery proteins (esaA, esaB, essA, essB and essC) are conserved among these Gram-positive bacteria (Fig. 1a). EssC and EsxA showed high amino acid sequence conservation across these Gram-positive bacteria (Fig. 1b, c and d). These data suggest that EssC and EsxA of these Gram-positive bacteria have similar roles.
EssC mediates EsxA secretion in the S. gordonii DL1. (a) Comparison of the S. gordonii type VII secretion system gene cluster with that of Streptococcus agalactiae, Streptococcus gallolyticus, Staphylococcus aureus and Listeria monocytogenes. (b) Homology tree of EssC amino acid sequences from pathogenic Firmicutes. (c) Alignment of EsxA amino sequences obtained from the NCBI database. (d) Homology tree of EsxA amino sequences from pathogenic Firmicutes. The EssC and EsxA amino acid sequences had a high degree of sequence conservation. (e) Wild-type S. gordonii DL1, ΔessC, ΔesxA, compΔessC and compΔesxA strains were grown in BHI broth and resuspended in sample lysis buffer to obtain a total bacterial protein extract. Supernatant proteins were precipitated with 20% TCA buffer and resuspended in sample buffer. EsxA was detected by western blotting with anti-EsxA antibodies. Uncropped blots are provided in Supplementary Figure S1 and S7. The experiments were performed three times, and representative results from one experiment are shown.
To further confirm that EsxA is the T7SSb substrate in S. gordonii, we constructed a deletion and complementation mutant of essC, a core component of T7SSb34. The essC-deficient mutant showed significantly reduced ability to release EsxA into the culture medium, and EsxA release was restored in the essC-complemented strain (Fig. 1e). These data suggest that T7SSb mediates EsxA secretion.
T7SSb contributes to vegetation formation and bacterial survival in a rat model of S. gordonii-induced IE
To investigate whether EsxA is a T7SSb effector molecule that contributes to S. gordonii pathogenesis in IE, we tested an esxA-deficient mutant strain in a rat model of IE5. Reduced EsxA release in the esxA-deficient mutant was confirmed by western blotting (Fig. 1e, and Supplementary Figure S1), and esxA-deficient mutant exhibited a decreased ability to colonize, form biofilms, and develop vegetations on heart valves compared to the wild-type strain. (Figure 2a, b, c and d). Complementation of esxA in the esxA-deficient mutant (comΔesxA) restored both EsxA secretion (Fig. 1e) and the ability to form biofilms and vegetations in vivo. (Figure 2a, b, c and d). These data suggest that secreted EsxA contributes to S. gordonii-induced IE pathogenesis. Interestingly, esxA-deficient strain also showed reduced survival in circulation in the rat IE model (Fig. 2e). This finding highlights the important role of EsxA in bacterial immune evasion within the bloodstream.
S. gordonii T7SSb contributes to pathogenesis in a rat model of IE. (a) Vegetation formation on the heart valves of rats with IE. A rat model of IE was used to investigate the roles of T7SSb in IE pathogenesis. Catheterized rats were injected with 109 CFU wild-type S. gordonii DL1, ΔesxA, ΔesxA-pDL278 or compΔesxA strains. Black arrows and dashed lines highlight vegetation and valves, respectively. Scale bars correspond to 0.5 mm. (b) Three-dimensional structure of GFP-labeled S. gordonii biofilms within vegetations collected from injured heart valves, visualized by confocal laser scanning microscopy at 630× magnification. The results shown are representative of three independent experiments. Scale bars indicated 50 μm. Vegetation size (c), number of bacteria colonizing the vegetation (d) and survival rate of bacteria in the circulation (e) were measured 24 h post-infection. Data are presented as a scatter plot with mean ± standard error of the mean. **P < 0.001, *P < 0.05 by Kruskal–Wallis test with subsequent Dunn’s test; ns, not significant.
EsxA has cytotoxicity toward neutrophils that contributes to bacterial immune evasion in circulation
To confirm the role of EsxA in the immune evasion of S. gordonii, we intravenously infected rats with wild-type and mutant S. gordonii DL1 via the tail vein and monitored bacterial survival in circulation (Fig. 3a). The EsxA-deficient strain showed lower survival in circulation than the wild-type strain, further suggesting a role for EsxA in bacterial survival. Meanwhile, the comΔesxA bacteria exhibited survival similar to that of the wild-type strain, supporting a role for secreted EsxA in bacterial immune evasion.
EsxA contributes to neutrophil cytotoxicity and phagocytosis escape. (a) Wild-type S. gordonii DL1 (black line), ΔesxA (red line), ΔesxA-pDL278 (green line) or compΔesxA (blue line) strains (109 CFU/mL) were injected into groups of five Wistar rats. Bacteria were collected from the bloodstream 0, 2, 4, 6 and 24 h after injection and cultured on BHI plates to quantify the number of bacteria. Data represent mean ± standard deviation derived from samples for each group of five rats. Statistical significance was assessed using Student’s t-test with ***P < 0.001. (b) Neutrophil cytotoxicity was calculated using an LDH assay. Supernatants were collected from human neutrophils approximately 8 h after infection with wild-type S. gordonii DL1 wild-type and mutant strains at a multiplicity of infection (MOI) of 100. The percentage cytotoxicity was determined and normalized with respect to control levels of 0% for uninfected cells and 100% for cells lysed with lysis buffer. (c) Contact-independent cytotoxicity was evaluated in neutrophils during S. gordonii infection utilizing transwells with polycarbonate membranes having a 0.4 μm pore size that separated S. gordonii and neutrophils. Cytotoxicity measurements were taken as the absorbance at 450 nm after 8 h of incubation. (d) Secreted EsxA was detected in stimulated neutrophils by lysing the neutrophils at the bottom of 24-well plates with SDS sample buffer. Western blotting with anti-EsxA antibodies was performed to detect secreted EsxA with anti-human GAPDH serving as an internal control. Lanes 1–2 and lanes 3–5 were cropped from different parts of the same gel and are separated by a clear white space. Uncropped blots are provided in Supplementary Figure S8. These results presented represent a single representative experiment selected from three independent experiments.
GBS EsxA was previously shown to have cytotoxic activity40. To investigate whether S. gordonii EsxA contributes to bacterial immune evasion via cytotoxic effects on neutrophils, we incubated wild-type and mutant S. gordonii strains with purified human neutrophils. Cell death induced by the esxA-deficient strain was reduced compared to that induced by the wild-type and comΔesxA strains (Fig. 3b). To determine whether the observed cell death was mediated by secreted molecules, we employed a transwell culture system in which neutrophils were cultured in the lower chamber and bacteria were seeded in the upper transwell chamber. Measurements of neutrophil survival consistently showed reduced neutrophil cell death in the presence of secreted molecules from esxA-deficient strains compared to wild-type and comΔesxA strains (Fig. 3c). Additionally, EsxA was detected in neutrophils stimulated with secreted molecules from the wild-type and comΔesxA strains, but not in those stimulated with esxA-deficient strains (Fig. 3d). Taken together, these data indicate that secreted EsxA is important for bacterial cytotoxicity toward neutrophils and contributes to the evasion of bacterial phagocytosis.
EsxA induces NET formation that contributes to immune evasion and vegetation formation in IE
To investigate how EsxA induces neutrophil death, we observed neutrophils cultured in a transwell system and stimulated with molecules secreted by the bacteria (Fig. 4a and b). Interestingly, NET formation was stimulated by molecules secreted from wild-type S. gordonii DL1 and comΔesxA, as evidenced by the colocalization of released DNA, MPO and citrullinated H3 (Fig. 4a and Supplementary Figure S2). In contrast, NET formation was reduced in neutrophils stimulated with the esxA-deficient strain. Consistently, NET formation was detected in neutrophils stimulated with culture supernatants from wild-type S. gordonii DL1 and comΔesxA, but was diminished in samples from the esxA-deficient strain (Supplementary Fig. S3). To further confirm the role of EsxA in NET induction, we purified recombinant EsxA (rEsxA). rEsxA, but not the recombinant AtlA negative control, induced NET release in a dose-dependent manner (Fig. 4c and d; Supplementary Figs. S4 and S5), further supporting the role of EsxA in NET induction.
S. gordonii EsxA triggers NET formation in vitro. (a) Induction of NET formation in a transwell system. Neutrophils collected from human peripheral blood samples were placed on coverslips at the well bottoms of 24-well plates (106 cells/mL) and incubated in RPMI medium for 1 h. After removing non-adherent neutrophils, the neutrophils were infected in transwells in which the S. gordonii bacteria and the neutrophils were separated by a polycarbonate membrane having a 0.4 μm pore size. NETs were visualized by staining with anti-human MPO (diluted 1:200), followed by a Texas red-labelled secondary antibody and observed by confocal microscopy (Leica SP8, magnification 630x). (b) The NET area was quantified using the Volocity program. Co-localization of MPO and DNA was considered to represent NETs. Total DNA was set to 100%. The results were analyzed by analyzed by one-way analysis of variance (ANOVA). **P < 0.01; ***P < 0.001 Experiments were carried out in triplicate and a representative trial is shown. (c) Induction of NET formation in vitro by recombinant EsxA. Isolated neutrophils were stimulated with increasing concentrations (0, 0.25, 0.5, 1 and 10 µg/ml) of recombinant EsxA protein for 4 h. After stimulation, the neutrophils were stained with anti-human MPO, followed by a Texas red-labelled secondary antibody. Extracellular DNA was stained with Hoechst 33,258. (d) The area of NET DNA fibers was quantified for three randomly-selected fields per sample using the Volocity program. Co-localization of MPO and extracellular DNA was considered to be indicative of NETs, with total DNA content normalized to 100%. Data were analyzed using one-way analysis of variance (ANOVA), with significance denoted as *P < 0.05, **P < 0.01 or ***P < 0.001. Experiments were carried out in triplicate and a representative trial is shown.
Our previous study demonstrated that NETs contribute to vegetation expansion in S. gordonii-induced IE pathogenesis5. Therefore, we used confocal laser scanning microscopy to examine NET formation within vegetations induced by S. gordonii. The NET structures were observed in vegetations formed by the wild-type DL1 strain, characterized by the co-localization of extracellular DNA, MPO, and citH3 (Fig. 5 and Supplementary Fig. S6, with examples indicated by arrows). In contrast, the esxA-deficient mutant displayed a markedly reduced NET-positive area, indicating an impaired ability to induce NET formation. This defect was rescued in the complemented strain (comΔesxA), supporting the role of EsxA in promoting NET induction in vivo (Fig. 5a). To further quantify NET formation, we measured the co-localized area of DNA and MPO, as shown in Fig. 5b. These data suggested the esxA-deficient strain showed a reduced ability to induce NET formation within the vegetations compared to the wild-type and comΔesxA strains, which is consistent with our results on vegetation mass (Fig. 2a and b). Taken together, these data suggest that EsxA, secreted by T7SSb, contributes to bacterial immune evasion and vegetation formation in S. gordonii-induced IE.
NET formation induced by S. gordonii strains in situ. In situ NET formation within vegetations induced by wild-type S. gordonii DL1, ΔessC, ΔesxA, and compΔesxA. (a) Vegetations harvested from S. gordonii-infected rats were stained with anti-rat MPO antibody (diluted 1:50) and labeled with Texas Red-conjugated secondary antibody for visualization of NETs using confocal microscopy (magnification 630x). The experiments were repeated three times, and a representative trial is presented. (b) The Volocity program was used to quantify the area of NET DNA fibers from three independent experiments. NETs were defined by co-localization of MPO and extracellular DNA (examples indicated by arrows), with total DNA normalized to 100%. Data are presented as the mean ± standard error of the mean (SEM); differences between two datasets were assessed using ANOVA; **P < 0.01.
Discussion
The roles of T7SSb and its substrates in bacterial pathogenesis has been well-established in various Gram-positive bacteria. However, the significance/contribution of T7SSb in the pathogenesis of viridans streptococci remains unclear. In this study, we provide evidence demonstrating the important role of T7SSb-mediated EsxA secretion in NET induction that contributes to bacterial immune evasion and vegetation formation in S. gordonii-induced IE. Sequence conservation at both the DNA and amino acid levels suggests that T7SSb and EsxA have similar functions among Gram-positive bacteria, especially pathogens that can cause IE. Interestingly, not all viridans streptococci strains possess T7SSb, according to genome database from National Center for Biotechnology Information. In addition, bacteria can be classified based on T7SSb substrate, T7SSb gene cluster or the C-terminal sequence of EssC40,42. These genetic differences might partly explain why not all commensal viridans streptococci strains are capable of causing IE.
NET-related immunothrombosis has been demonstrated to have an important role in several severe infectious diseases, such as COVID-19 and sepsis43. We previously showed the importance of NETs in vegetation expansion in IE5. In the same study, we also found that the ability of bacteria to induce platelet activation contributes to biofilm formation and NET production in IE. Therefore, in addition to directly inducing NET production, EsxA may also promote bacteria-induced platelet activation, which warrants further investigation. NETs were initially reported to have bactericidal activity16. However, studies involving S. aureus and Candida albicans showed that NETs merely trap microbes and do not directly kill them17,44. S. aureus leukocidins can directly induce NET formation to protect bacteria from killing by neutrophil-mediated phagocytosis18. Our previous study also showed that NETs do not affect viridans streptococcal growth5. In the present study, we used a transwell system to show that S. gordonii can secrete ExsA to induce NET formation without direct contact. These findings suggest that S. gordonii, like S. aureus, can induce NET formation at a distance, potentially without being trapped by NETs. These findings suggest that S. gordonii, like S. aureus, can induce NET formation in the circulation from a distance, potentially without being trapped by NETs. S. aureus leukotoxin GH (LukGH; also known as LukAB) has been shown to have pore-forming activity and to mediate NETosis to evade phagocytic killing44,45. Similarly, EsxA in GBS and M. tuberculosis has been reported to form pores40,46. These findings imply that EsxA induction of NET formation, like that by S. aureus LukGH, involves pore-forming activity, and this possibility warrants further investigation.
NETosis has been introduced for almost 30 years, since Takei and colleagues first observed 19966, while the cellular mechanistic understanding of this process still remains incomplete. The underlying molecular mechanisms are highly complex and involve a series of tightly regulated intracellular events7,47,48. Generally, NET-inducing stimuli activate the MEK-ERK signaling pathway, leading to the production of ROS. ROS, in turn, activate MPO and NE, which translocate to the nucleus and promote chromatin decondensation through histone degradation. Concurrently, ROS activate PAD4, which citrullinates histones, further facilitating chromatin relaxation. Eventually, phosphorylation of nuclear lamins drives nuclear envelope breakdown, followed by plasma membrane disruption, pore formation, and NET release. However, the requirement for these known signaling pathways varies depending on the stimulus, and how they influence the key cellular events remains unclear8,47. Previous studies have indicated that pore formation may serve as a critical upstream event in the NETosis pathway by promoting calcium influx, destabilization of cellular membranes, and the release of granule-derived enzymes such as neutrophil elastase and myeloperoxidase49. Each of these processes can contribute to chromatin decondensation and NET formation, depending on the specific cellular context and stimulus. For example, gasdermin D (GSDMD) may contribute to the release of NE from intracellular granules and facilitate its translocation to the nucleus. Conversely, NE can cleave and activate GSDMD, promoting its pore formation in nuclear, organelle, and plasma membranes, thereby enhancing membrane permeability and the redistribution of intracellular enzymes50. This bidirectional regulatory mechanism highlights a potential positive feedback loop between NE and GSDMD during NETosis51. Given the ability of S. gordonii EsxA to act at a distance, its pore-forming activity may directly compromise neutrophil membrane integrity and trigger NETosis without requiring direct bacterial contact. These observations imply that EsxA likely functions similarly to other bacterial pore-forming toxins, serving as an extracellular inducer of neutrophil activation and NET release. Further studies are needed to elucidate whether EsxA-induced NETosis engages canonical mechanisms, such as NOX-dependent ROS generation, phosphorylation and disassembly of nuclear lamins, or GSDMD-mediated membrane permeabilization.
In addition to evading immune surveillance, biofilm formation is a key virulence factor enabling bacteria to cause IE. Our previous studies demonstrated that bacterial extracellular DNA (eDNA) is critical for viridans streptococcal biofilm formation in the pathogenesis of viridans streptococci-induced IE52. Our recent data further showed that the essC-deficient mutant strain released lower levels of eDNA and exhibited impaired biofilm formation. These findings suggest that EsxA or other S. gordonii T7SSb substrates may contribute to the release of eDNA and subsequent biofilm development.
In addition to WXG100 proteins, LXG family proteins are also recognized as T7SSb effectors53. These proteins exhibit diverse toxin activities across different Firmicutes, including NADase activity in Streptococcus intermedius36 and nuclease activity in Bacillus subtilis53. Furthermore, other classes of T7SSb substrates beyond WXG100 and LXG family proteins have also been identified, such as S. aureus EsaD, a non-LXG effector with membrane-depolarizing activity involved in intraspecies competition35. and TslA, a secreted lipase toxin with a reverse domain arrangement54. In S. gordonii DL1, our analysis did not identify a typical LXG family protein near the T7SSb gene locus; however, several small, uncharacterized proteins are located downstream of essC (Fig. 1). We will therefore further investigate the roles of these small proteins in IE and aim to identify additional T7SSb substrates or effectors involved in IE pathogenesis.
In summary, here we demonstrated roles of T7SSb in the pathogenesis of S. gordonii-induced IE (Fig. 6). S. gordonii T7SSb modulates ExsA secretion to induce NET release that inhibits phagocytotic killing and contributes to bacterial survival in the circulation. NETs also promote platelet activation and coagulation by contributing to bacterial biofilm formation and expansion of vegetation on the heart valves in IE5. T7SSb may also mediate biofilm formation through an as-yet unidentified subtract or effector. T7SSb is also found in other IE pathogens, including S. aureus and Enterococci. Therefore, the findings from this study may enhance our understanding of IE pathogenesis caused by other pathogens.
Model for EssC-mediated EsxA secretion that causes IE by promoting NET formation, escape of neutrophil phagocytosis, and vegetation expansion. (a) EssC in T7SSb mediates the transport and secretion of EsxA across the cytoplasmic membrane and thick cell wall in S. gordonii, thereby promoting its release into the extracellular environment. (b) The secreted EsxA binds to neutrophils to trigger NET formation. Secretion of EsxA also inhibits phagocytic killing to enhance bacterial survival in the circulation. NETs facilitate platelet activation and coagulation that promotes bacterial biofilm formation and expansion of vegetation in heart valves in IE.
Materials and methods
Ethics statement
All study participants provided written informed consent allowing use of human blood samples in adherence with guidelines established by the Committee for Regulation of Human Specimens and Volunteers at the National Taiwan University Hospital (NTUH, 202301127RINB).
Bacterial strains and plasmids
Wild-type S. gordonii strains were grown in brain heart infusion broth (BHI, Difco Laboratories Inc., Detroit, MI, USA). For essC (ΔessC) and esxA-deficient (ΔesxA) strains, the BHI contained 500 µg/mL kanamycin. To generate the complementation strain (comΔesxA), bacteria were transformed with the shuttle plasmid pDL278 containing the promoter region and the esxA gene, and grown in BHI supplemented with 500 µg/mL spectinomycin, as previously described52,55. The vector without the promoter and esxA gene regions was also transformed into the esxA-deficient mutant and served as the vector control (ΔesxA-pDL278) to exclude any effect of pDL278. GFP-labeled bacteria were generated by transformed with pPDGFPuv56, which carries the GFPuv sequence and a chloramphenicol resistance cassette. To maintain the GFP-labeled strains, BHI broth was supplemented with 10 µg/mL chloramphenicol to ensure retention of the plasmids during bacterial growth.
Construction of essC and esxA deletion mutants and esxA complementation strains
All methods used in this study were carried out in accordance with relevant guidelines and regulations, and all experimental protocols were approved by Taipei Medical University and National Taiwan University.
Strains having deletion of essC or esxA were generated using a ligation-PCR mutagenesis strategy as described previously that involved insertion of promoter-less kanamycin cassettes to disrupt the corresponding genes in the S. gordonii strain DL152,55. Briefly, gene sequences upstream and downstream of essC and esxA as well as the promoter-less kanamycin resistance gene were amplified from S. gordonii DL1 chromosomal DNA and the pALH124 plasmid, respectively, using primers listed in Table 1. The primers were designed according to the S. gordonii DL1 genomic database (https://www.ncbi.nlm.nih.gov/genome/). The resulting PCR products were digested with EcoRI and ligated with T4 ligase. The ligation products were transformed into wild-type S. gordonii DL1 and mutant strains were verified by PCR amplification and western blotting analysis. To construct the comΔessC and comΔesxA strains, essC and esxA were amplified using the primers listed in Table 1. The PCR products were digested with EcoRI and ligated into the pDL278 plasmid carrying a spectinomycin resistance cassette. The resulting plasmids were then used to transform ΔessC and ΔesxA to generate the comΔessC and comΔesxA strains, respectively. Expression of esxA was further confirmed by western blotting analysis.
Rat model of S. gordonii-induced IE and in vivo bacteria clearance assay
A rat model of streptococcal endocarditis was performed as previously described with the following modifications5. All experiments involving animals were performed according to protocols approved by the National Taiwan University Institutional Animal Care and Use Committee (Taipei, Taiwan; IACUC 20220439). The study was carried out in accordance with the relevant guidelines and regulation with the ARRIVE guidelines 2.0 (https://arriveguidelines.org/arrive-guidelines). All animals were handled in full compliance with the National Institutes of Health guidelines for the care and use of laboratory animals, ensuring ethical and humane treatment throughout the research process. Thirty adult rats (8–10 weeks old, weighing 500–600 g) were obtained from BioLASCO Taiwan Co., Ltd. All animals were housed under standardized laboratory conditions, with ad libitum access to water and food, in an environment maintained at a temperature of 20–24 °C and a relative humidity of 40–75%. For anesthetization, each rat was initially anesthetized with 4% isoflurane for induction and maintained on 2% isoflurane. Additionally, Zoletil (20–40 mg/kg) and xylazine (5–10 mg/kg) were administered intraperitoneally. No repeated anesthesia was performed on the same animal. Briefly, the ventral neck surgical site was shaved and disinfected using an aqueous iodine solution. The aortic valve was injured using a polyethylene tube containing a stainless-steel wire inserted into the left carotid artery. At 24 h post-catheterization, rats were intravenously infected with 109 colony forming units (CFUs) of wild-type or isogenic mutant T7SSb strains. The rats were sacrificed 24 h after infection using 50% CO2 for 10 min, whereupon all cardiac vegetations were harvested and weighed. The vegetations were then homogenized in saline, serially diluted, and plated onto BHI agar to determine bacterial counts, which were normalized according to the vegetation weight. To observe NETs, the vegetations were fixed with 2% paraformaldehyde and stained with an anti-rat myeloperoxidase (MPO) antibody (diluted 1:50; Santa Cruz Biotechnology, California, United States), followed by rhodamine red-conjugated anti-rabbit immunoglobulin G (IgG) antibody (diluted 1:500; Jackson ImmunoResearch Labs, West Grove, Pennsylvania, United States). DNA was stained with Hoechst 33,258 (Sigma-Aldrich, St. Louis, Missouri, United States) and NET structures were observed using a Leica SP8 confocal microscope. For biofilm observation, S. gordonii DL1 wild-type and mutant strains were transformed with pDGFPuv56, and the biofilm structures were monitored using ImarisViewer10.2 Image Analysis Software (https://imaris.oxinst.com/). The bacterial clearance assay was performed as we previously described56. Briefly, 109 CFUs of bacteria were intravenously into groups of 5 Wistar rats for each strain. Each rat represented an independent experiment with distinct bacterial cultures.
Recombinant EsxA purification and anti-EsxA antibody preparation
Purification of DL1 EsxA was performed as previously described with some modifications55. The esxA gene of DL1 was cloned into the BamHI site of the pQE30 expression vector that contains an N-terminal in-frame 6XHis-tag encoding sequence. The resultant EsxA expression plasmid was transformed into M15 Escherichia coli. EsxA expression was induced in M15 E. coli during the exponential growth phase growth by addition of 1 mM isopropyl-β-D-thiogalactopyranoside (IPTG). After 4 h of induction, the bacterial cells were collected, resuspended in binding buffer (30µM imidazole, 0.5µM, 20µM Tris-HCl, pH 7.9) and then disrupted by sonication on ice. The expressed proteins were purified using a nickel chelating affinity chromatography system, dialyzed into 10mM NaPB buffer (1 M NaH2PO4, 1 M Na2HPO4, pH 6.0) and checked by SDS-PAGE. The rat anti-EsxA polyclonal antibody was isolated as previously described56. Briefly, purified rEsxA was mixed and intracutaneously injected into Wistar rats on the back. A second booster injection with 1:1 Freund’s complete adjuvant and rEsxA was given two weeks later and serum was collected two days after the second boost. Anti-EsxA sera titers were analyzed using enzyme-linked immunosorbent assay (ELISA), and the anti-EsxA rat serum specificity was measured by western blotting using recombinant EsxA protein, total protein extracts from wild-type S. gordonii and isogenic mutant T7SSb strains.
Detection of cell wall-associated and secreted S. gordonii EsxA
To detect secreted EsxA in S. gordonii culture supernatants, wild-type and isogenic mutant T7SSb and complementation strains were grown in BHI broth for 16 h at 37 °C. The bacteria were centrifuged at 5000xg for 10 min at 4 °C. EDTA-free protease inhibitor cocktail (1:200 dilution; MCE, catalog: HY-K0010) was added to the supernatants and the flow-through was precipitated with trichloroacetic acid (TCA; final concentration 20%) for 24 h at 4 °C. The precipitated proteins were gently washed with acetone, dried and resuspended in sample buffer (10% SDS, 4% β-mercaptoethanol, 0.5 M Tris, pH 6.8). Total bacterial pellets were gently washed with phosphate buffered saline (PBS) and resuspended in sample buffer. Precipitates and supernatants were boiled for 10 min before SDS-PAGE and western blotting. For western blots, protein samples were transferred to PVDF membranes that were blocked in fast blocking buffer (BioMAN; catalog: FBB01-500) for 30 min at room temperature. After blocking, the membranes were hybridized with an anti-EsxA rat polyclonal antibody in blocking buffer overnight at 4 °C. The membranes were then washed in 1X Tris buffered saline, 0.1% Tween 20 (TBST) and incubated with goat-anti-rat secondary antibodies (1:10,000; Abcam; ab97057) for 2 h at room temperature. After washing three times with TBST, the western blots were visualized using a Muti-function Gel Image system.
Preparation of neutrophils and in vitro induction of NET formation
As mentioned above, this research received approval from the Committee for Regulation of Human Specimens and Volunteers at National Taiwan University Hospital and written informed consent was obtained from all study participants. Volunteers were healthy individuals older than 18 years, regardless of sex, and without prior use of antibacterial or anti-inflammatory drugs. Neutrophils were isolated from heparinized venous blood samples using Ficoll-Histopaque (Histopaque 1119 and Histopaque 1083, Sigma-Aldrich). Neutrophils were collected and seeded onto plasma-coated coverslips (100 µL of the suspension in Roswell Park Memorial Institute (RPMI) medium contains 106 cells/mL), which were placed into the wells of a 24-well plate and incubated in 5% CO2 at 37 °C for 1 h. After removing non-adherent neutrophils, the remaining neutrophils on the coverslip were subsequently stimulated with recombinant S. gordonii EsxA (0.25, 0.5, 1, and 10 µg/mL) in RPMI medium for 4 h. A transwell assay was performed in a 24-well plate using polycarbonate transwell inserts with 8.75 mm diameter membranes having 0.4 µM pores (SPL life science; catalog:35324). The neutrophils were seeded in the lower compartment of the 24-well plate, and wild-type and isogenic S. gordonii strains were added to the transwell bucket at an MOI of 100. The plate was then incubated for 8 h in 5% CO2 at 37 °C. To observe NETs, samples were stained with two primary antibodies: a mouse anti-human MPO antibody (dilution 1:200; Santa Cruz Biotechnology, sc-52707) and a rabbit anti-human citrullinated histone H3 (citH3) antibody (dilution 1:100; Abcam, ab5103, Cambridge, MA). MPO was detected using a FITC- or Texas Red-labeled anti-mouse IgG secondary antibody (Jackson ImmunoResearch Labs, West Grove, PA, USA), while citH3 was followed with an anti-rabbit IgG secondary antibody (Jackson ImmunoResearch Labs, West Grove, PA, USA). DNA was stained with Hoechst 33,258 (Sigma-Aldrich, St. Louis, Missouri, United States). NET formation was observed by confocal microscopy (TCS Leica SP5 or SP8) and analyzed using the Volocity program (PerkinElmer; https://www.volocity4d.com/).
Cell cytotoxicity assay
For neutrophil cytotoxicity assays, bacterial cells were inoculated with neutrophils at a multiplicity of infection (MOI) of 100 and incubated for 8 h. LDH release was then quantified using an LDH assay kit (Abcam ab65393) according to the manufacturer’s instructions. Transwell cytotoxicity assays were performed in 24-well plates with cells seeded in the bottom compartment. Wild-type and isogenic S. gordonii strains were added to the transwell bucket at an MOI of 100. The plate was then incubated for 8 h in 5% CO2 at 37 °C. To achieve maximum LDH release, cell lysis solution was added to each well, followed by addition of 50 µl of the reconstituted substrate mix. The reaction was then stopped, and the absorbance was recorded at 450 nm using an ELISA plate reader. Cytotoxicity was quantified as a percentage relative to an uninfected control that was treated with 1% Triton X-100.
Statistical analysis
Analyses of the statistical significance of differences between two datasets were conducted using an unpaired, two-tailed Student’s t-test. For comparison of more than two datasets, a one-way ANOVA followed by a Bonferroni post-hoc test was utilized. For data with a non-normal distribution, Mann-Whitney U and Kruskal-Wallis tests followed by a Dunn’s post-test were employed. These analyses were performed using GraphPad Prism version 8 (GraphPad Software). A p-value < 0.05 indicated statistical significance.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.
References
Mylonakis, E. & Calderwood, S. B. Infective endocarditis in adults. N Engl. J. Med. 345, 1318–1330 (2001).
Moreillon, P., Que, Y. A. & Infective endocarditis Lancet 363, 139–149, (2004). doi:S0140-6736(03)15266-X [pii]10.1016/S0140-6736(03)15266-X [doi].
Scheld, W. M. Pathogenesis and pathophysiology of infective endocarditis. In Sande, M. A., Kaye, D., Root, R. K. (ed.), 1–32 (Endocarditis. Churchill Livingstone, New York, N.Y., 1984).
Jung, C. J. et al. Platelets enhance biofilm formation and resistance of endocarditis-inducing Streptococci on the injured heart valve. J. Infect. Dis. 205, 1066–1075. https://doi.org/10.1093/infdis/jis021 (2012).
Jung, C. J. et al. Endocarditis pathogen promotes vegetation formation by inducing intravascular neutrophil extracellular traps through activated platelets. Circulation 131, 571–581. https://doi.org/10.1161/CIRCULATIONAHA.114.011432 (2015).
Takei, H., Araki, A., Watanabe, H., Ichinose, A. & Sendo, F. Rapid killing of human neutrophils by the potent activator phorbol 12-myristate 13-acetate (PMA) accompanied by changes different from typical apoptosis or necrosis. J. Leukoc. Biol. 59, 229–240. https://doi.org/10.1002/jlb.59.2.229 (1996).
Wang, H. et al. Neutrophil extracellular traps in homeostasis and disease. Signal. Transduct. Target. Ther. 9, 235. https://doi.org/10.1038/s41392-024-01933-x (2024).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147. https://doi.org/10.1038/nri.2017.105 (2018).
Hakkim, A. et al. Activation of the Raf-MEK-ERK pathway is required for neutrophil extracellular trap formation. Nat. Chem. Biol. 7, 75–77. https://doi.org/10.1038/nchembio.496 (2011).
Lv, X., Liu, Z., Xu, L., Song, E. & Song, Y. Tetrachlorobenzoquinone exhibits immunotoxicity by inducing neutrophil extracellular traps through a mechanism involving ROS-JNK-NOX2 positive feedback loop. Environ. Pollut. 268, 115921. https://doi.org/10.1016/j.envpol.2020.115921 (2021).
Wang, Y. et al. Histone hypercitrullination mediates chromatin decondensation and neutrophil extracellular trap formation. J. Cell. Biol. 184, 205–213. https://doi.org/10.1083/jcb.200806072 (2009).
Li, Y. et al. Nuclear envelope rupture and NET formation is driven by PKCalpha-mediated lamin B disassembly. EMBO Rep. 21, e48779. https://doi.org/10.15252/embr.201948779 (2020).
Amulic, B. et al. Cell-Cycle Proteins Control Production of Neutrophil Extracellular Traps. Dev. Cell 43, 449–462 e445 https://doi.org/10.1016/j.devcel.2017.10.013 (2017).
Li, M., Lyu, X., Liao, J., Werth, V. P. & Liu, M. L. Rho kinase regulates neutrophil NET formation that is involved in UVB-induced skin inflammation. Theranostics 12, 2133–2149. https://doi.org/10.7150/thno.66457 (2022).
Fuchs, T. A. et al. Novel cell death program leads to neutrophil extracellular traps. J. Cell. Biol. 176, 231–241. https://doi.org/10.1083/jcb.200606027 (2007).
Brinkmann, V. Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535. https://doi.org/10.1126/science.1092385 (2004).
Menegazzi, R., Decleva, E. & Dri, P. Killing by neutrophil extracellular traps: fact or folklore? Blood 119, 1214–1216. https://doi.org/10.1182/blood-2011-07-364604 (2012).
Bhattacharya, M. et al. Staphylococcus aureus biofilms release leukocidins to elicit extracellular trap formation and evade neutrophil-mediated killing. Proc. Natl. Acad. Sci. U S A. 115, 7416–7421. https://doi.org/10.1073/pnas.1721949115 (2018).
Engelmann, B. & Massberg, S. Thrombosis as an intravascular effector of innate immunity. Nat. Rev. Immunol. 13, 34–45. https://doi.org/10.1038/nri3345 (2013).
Hsu, C. C. et al. Neutrophil extracellular traps enhance Staphylococcus aureus vegetation formation through interaction with platelets in infective endocarditis. Thromb. Haemost. 119, 786–796. https://doi.org/10.1055/s-0039-1678665 (2019).
Green, E. R. & Mecsas, J. Bacterial secretion systems: an overview. Microbiol. Spectr. 4 https://doi.org/10.1128/microbiolspec.VMBF-0012-2015 (2016).
Costa, T. R. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359. https://doi.org/10.1038/nrmicro3456 (2015).
Lasica, A. M., Ksiazek, M., Madej, M. & Potempa, J. The type IX secretion system (T9SS): highlights and recent insights into its structure and function. Front. Cell. Infect. Microbiol. 7, 215. https://doi.org/10.3389/fcimb.2017.00215 (2017).
Gupta, G. et al. Secretory molecules from secretion systems fine-tune the host-beneficial bacteria (PGPRs) interaction. Front. Microbiol. 15, 1355750. https://doi.org/10.3389/fmicb.2024.1355750 (2024).
Jamali, H., Akrami, F., Layeghkhavidaki, H. & Bouakkaz, S. Bacterial protein secretion systems: Mechanisms, functions, and roles in virulence. Microb. Pathog. 206, 107790. https://doi.org/10.1016/j.micpath.2025.107790 (2025).
Rapisarda, C., Tassinari, M., Gubellini, F. & Fronzes, R. Using Cryo-EM to investigate bacterial secretion systems. Annu. Rev. Microbiol. 72, 231–254. https://doi.org/10.1146/annurev-micro-090817-062702 (2018).
Hayes, C. S., Aoki, S. K. & Low, D. A. Bacterial contact-dependent delivery systems. Annu. Rev. Genet. 44, 71–90. https://doi.org/10.1146/annurev.genet.42.110807.091449 (2010).
Abdallah, A. M. et al. Type VII secretion–mycobacteria show the way. Nat. Rev. Microbiol. 5, 883–891. https://doi.org/10.1038/nrmicro1773 (2007).
Spencer, B. L. & Doran, K. S. Evolving Understanding of the type VII secretion system in Gram-positive bacteria. PLoS Pathog. 18, e1010680. https://doi.org/10.1371/journal.ppat.1010680 (2022).
Bowman, L. & Palmer, T. The type VII secretion system of Staphylococcus. Annu. Rev. Microbiol. 75, 471–494. https://doi.org/10.1146/annurev-micro-012721-123600 (2021).
Rosenberg, O. S. et al. Substrates control multimerization and activation of the Multi-Domain ATPase motor of type VII secretion. Cell 161, 501–512. https://doi.org/10.1016/j.cell.2015.03.040 (2015).
Poulsen, C., Panjikar, S., Holton, S. J., Wilmanns, M. & Song, Y. H. WXG100 protein superfamily consists of three subfamilies and exhibits an alpha-helical C-terminal conserved residue pattern. PLoS One. 9, e89313. https://doi.org/10.1371/journal.pone.0089313 (2014).
Unnikrishnan, M., Constantinidou, C., Palmer, T. & Pallen, M. J. The enigmatic Esx proteins: looking beyond mycobacteria. Trends Microbiol. 25, 192–204. https://doi.org/10.1016/j.tim.2016.11.004 (2017).
Rivera-Calzada, A., Famelis, N., Llorca, O. & Geibel, S. Type VII secretion systems: structure, functions and transport models. Nat. Rev. Microbiol. 19, 567–584. https://doi.org/10.1038/s41579-021-00560-5 (2021).
Cao, Z., Casabona, M. G., Kneuper, H., Chalmers, J. D. & Palmer, T. The type VII secretion system of Staphylococcus aureus secretes a nuclease toxin that targets competitor bacteria. Nat. Microbiol. 2, 16183. https://doi.org/10.1038/nmicrobiol.2016.183 (2016).
Whitney, J. C. et al. A broadly distributed toxin family mediates contact-dependent antagonism between gram-positive bacteria. Elife 6 https://doi.org/10.7554/eLife.26938 (2017).
Sundaramoorthy, R., Fyfe, P. K. & Hunter, W. N. Structure of Staphylococcus aureus EsxA suggests a contribution to virulence by action as a transport chaperone and/or adaptor protein. J. Mol. Biol. 383, 603–614. https://doi.org/10.1016/j.jmb.2008.08.047 (2008).
Lopez, M. S. et al. Host-derived fatty acids activate type VII secretion in Staphylococcus aureus. Proc. Natl. Acad. Sci. U S A. 114, 11223–11228. https://doi.org/10.1073/pnas.1700627114 (2017).
Kengmo Tchoupa, A. et al. The type VII secretion system protects Staphylococcus aureus against antimicrobial host fatty acids. Sci. Rep. 10, 14838. https://doi.org/10.1038/s41598-020-71653-z (2020).
Spencer, B. L. et al. A type VII secretion system in group B Streptococcus mediates cytotoxicity and virulence. PLoS Pathog. 17, e1010121. https://doi.org/10.1371/journal.ppat.1010121 (2021).
Taylor, J. C. et al. A type VII secretion system of Streptococcus Gallolyticus subsp. Gallolyticus contributes to gut colonization and the development of colon tumors. PLoS Pathog. 17, e1009182. https://doi.org/10.1371/journal.ppat.1009182 (2021).
Warne, B. et al. The Ess/Type VII secretion system of Staphylococcus aureus shows unexpected genetic diversity. BMC Genom. 17 https://doi.org/10.1186/s12864-016-2426-7 (2016).
Stark, K. & Massberg, S. Interplay between inflammation and thrombosis in cardiovascular pathology. Nat. Rev. Cardiol. 18, 666–682. https://doi.org/10.1038/s41569-021-00552-1 (2021).
Malachowa, N., Kobayashi, S. D., Freedman, B., Dorward, D. W. & DeLeo, F. R. Staphylococcus aureus leukotoxin GH promotes formation of neutrophil extracellular traps. J. Immunol. 191, 6022–6029. https://doi.org/10.4049/jimmunol.1301821 (2013).
Dumont, A. L. et al. Characterization of a new cytotoxin that contributes to Staphylococcus aureus pathogenesis. Mol. Microbiol. 79, 814–825. https://doi.org/10.1111/j.1365-2958.2010.07490.x (2011).
Ma, Y., Keil, V. & Sun, J. Characterization of Mycobacterium tuberculosis EsxA membrane insertion: roles of N- and C-terminal flexible arms and central helix-turn-helix motif. J. Biol. Chem. 290, 7314–7322. https://doi.org/10.1074/jbc.M114.622076 (2015).
Liu, M. L., Lyu, X. & Werth, V. P. Recent progress in the mechanistic Understanding of NET formation in neutrophils. FEBS J. 289, 3954–3966. https://doi.org/10.1111/febs.16036 (2022).
Singh, J. et al. Moonlighting chromatin: when DNA escapes nuclear control. Cell. Death Differ. 30, 861–875. https://doi.org/10.1038/s41418-023-01124-1 (2023).
Vorobjeva, N. V., Chernyak, B. V. & NETosis Molecular Mechanisms, role in physiology and pathology. Biochem. (Mosc). 85, 1178–1190. https://doi.org/10.1134/S0006297920100065 (2020).
Zhu, C. et al. The gasdermin family: emerging therapeutic targets in diseases. Signal. Transduct. Target. Ther. 9, 87. https://doi.org/10.1038/s41392-024-01801-8 (2024).
Chen, K. W. et al. Noncanonical inflammasome signaling elicits gasdermin D-dependent neutrophil extracellular traps. Sci. Immunol. 3 https://doi.org/10.1126/sciimmunol.aar6676 (2018).
Jung, C. J. et al. PspC domain-containing protein (PCP) determines Streptococcus mutans biofilm formation through bacterial extracellular DNA release and platelet adhesion in experimental endocarditis. PLoS Pathog. 17, e1009289. https://doi.org/10.1371/journal.ppat.1009289 (2021).
Holberger, L. E., Garza-Sanchez, F., Lamoureux, J., Low, D. A. & Hayes, C. S. A novel family of toxin/antitoxin proteins in Bacillus species. FEBS Lett. 586, 132–136. https://doi.org/10.1016/j.febslet.2011.12.020 (2012).
Garrett, S. R. et al. A type VII-secreted lipase toxin with reverse domain arrangement. Nat. Commun. 14, 8438. https://doi.org/10.1038/s41467-023-44221-y (2023).
Hsu, C. C. et al. Streptococcus mutans PrsA mediates AtlA secretion contributing to extracellular DNA release and biofilm formation in the pathogenesis of infective endocarditis. Virulence 13, 1379–1392. https://doi.org/10.1080/21505594.2022.2105351 (2022).
Jung, C. J., Zheng, Q. H., Shieh, Y. H., Lin, C. S. & Chia, J. S. Streptococcus mutans autolysin AtlA is a fibronectin-binding protein and contributes to bacterial survival in the bloodstream and virulence for infective endocarditis. Mol. Microbiol. 74, 888–902. https://doi.org/10.1111/j.1365-2958.2009.06903.x (2009).
Acknowledgements
This study was supported by Taipei Medical University (113-TMU121), National Taiwan University Hospital (113-TMU122 and 113-TMU123), and the National Science and Technology Council (NSTC112-2320-B-038-055-MY3, NSTC113-2320-B-038-015 and NSTC114-2320-B-031-002). We acknowledge the Imaging Core Facility at the First Core Labs, National Taiwan University College of Medicine, for providing technical support in image acquisition and analysis.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design, and reviewed the manuscript. CJ Jung, YC Chung, RB Hsu, and JS Chia conceived the experiments; CC Hsu and JW Chen developed and conducted the experiments; YH Shih and YM Kuo helped analyze the results; and CJ Jung and CC Hsu wrote, revised, and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hsu, CC., Chuang, YC., Hsu, RB. et al. Streptococcus gordonii type VII secretion system substrate EsxA induces neutrophil extracellular trap formation in infective endocarditis. Sci Rep 15, 43708 (2025). https://doi.org/10.1038/s41598-025-27543-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27543-3