Abstract
Highly pathogenic avian influenza (HPAI) poses a major threat to the world economy and human health; hence, rapid and accurate identification of avian influenza virus (AIV) subtypes is required for effective surveillance and outbreak prevention, particularly in remote or resource-limited settings. Here, we present a portable laboratory system designed to streamline the characterisation of AIV subtypes. This system integrates sample processing, rapid sequencing, and easy to use bioinformatics pipelines to enable characterisation of AIV subtypes, directly from the field. The approach also reduces the chances of samples being degraded, delayed or mixed-up during transport to the laboratory. Equipment for the portable system was first assessed in the laboratory, using AIV isolates with known subtypes, then with swabs collected from several locations in New Zealand. The system was then deployed in the field in Antarctica in December 2024, where it identified HPAI H5N1 in a skua, demonstrating its ability to characterise AIV subtypes rapidly and accurately in situ. This highlights the potential of our portable field system to enhance AIV surveillance in challenging environments and to mitigate the impact of its spread.
Similar content being viewed by others
Introduction
Avian influenza virus (AIV) is a global concern due to its significant impact on animal and public health, food security, and the world economy. Recent detections of highly pathogenic avian influenza (HPAI) H5N1 clade 2.3.4.4b in dairy cattle and humans in the United States have raised increasing concerns due to the zoonotic transmission capability of AIV and its continuous spread globally1,2. There have been over 890 confirmed human infections of the H5N1 subtype reported worldwide since 2003, resulting in more than 450 deaths3. Additionally, outbreaks of HPAI in poultry farms have led to the loss of millions of birds, severely impacting national economies. This highlights the need to develop a rapid system that can accurately characterise AIV in poultry farms and wild birds.
AIVs of the genus Alphainfluenzavirus and family Orthymyxoviridae, are negative-sense single-stranded RNA viruses comprising eight genomic segments. Influenza viruses are classified into different subtypes based on the combinations of two surface glycoproteins: haemagglutinin (HA) and neuraminidase (NA)4. The HA protein enables the virus to enter host cells by binding to the host sialic acid receptors, while the NA protein plays a crucial role in releasing viral particles from the infected cells5. Different subtypes of AIV can exhibit varying degrees of virulence. Most subtypes are low pathogenic avian influenza viruses (LPAI), while some are classified as HPAI. The classification is based on the sequence of the HA cleavage site6. There are currently 18 known HA subtypes (H1-18) and 11 known NA subtypes (N1-11) with 16 HA and 9 NA subtypes identified in birds, the remaining subtypes were identified in bats7. HPAI is typically associated with subtype H5 or H7, however, not all H5 and H7 subtypes are HPAI8.
LPAI subtypes often cause minimal or no symptoms in infected poultry and wild birds, whereas HPAI is fatal to domestic poultry and some wild birds6. Wild aquatic birds such as mallard ducks and gulls can be asymptomatic when infected, so they are natural reservoirs for AIV, and are key vectors involved in transmission of the virus to domestic poultry9. When two or more LPAI viruses are circulating, co-infection with multiple LPAI subtypes can occur in a single host. This leads to genetic reassortment and may result in the development of a novel viral strain that can influence viral virulence, host susceptibility, and antigen specificity10.
RNA-dependent RNA polymerases in AIVs lack proofreading exonuclease activity, therefore mutational events are common. Of particular importance are mutations that affect the HA cleavage site, because these can confer increased pathogenicity of the viruses. According to the World Organisation of Animal Health11 and OIE/FAO Network of Expertise on Animal Influenza12, LPAI HA cleavage sites are mono-basic, defined as the presence of a single basic amino acid (usually either lysine or arginine) directly before the amino acid cleavage motif GLF (Glycine-Leucine-Phenylalanine)12. In contrast, HPAI HA cleavage sites are multi-basic, featuring two or more basic amino acids in the critical region12. This arises due to single nucleotide mutational events and/or in-frame insertions of codons that translate to lysine or arginine12. The presence of additional basic amino acids at the HA cleavage site promotes cleavage by furin-like proteases that are ubiquitously expressed in most cells, leading to systemic infection of the disease13,14. In addition, mutations in the polymerase basic 2 (PB2) gene can also enhance host adaptation and increase pathogenicity in mammalian hosts. A common mutation seen in PB2 is the E627K mutation, which facilitates increased replication in mammalian hosts15. For these reasons, early detection of mutational events that could transform LPAI to HPAI is critical for a timely biosecurity response.
Routine laboratory AIV subtyping workflows often require transporting samples from the field to the laboratory, a process that can take days or even weeks, especially when samples come from remote regions. AIV transmission between birds can be rapid, and a delay in diagnosing and characterising the virus can have devastating consequences. A field-deployable system capable of rapid, real-time analysis of influenza subtypes is therefore necessary to overcome these challenges. Such a system would bypass the logistical and temporal constraints of laboratory testing.
We present a portable laboratory system with an easy-to-use bioinformatics pipeline capable of delivering automated AIV characterisation offline and in real-time. The portable equipment was first assessed using AIV viral isolates of known subtypes, then verified in the laboratory using swabs collected across different locations in New Zealand. The system was finally deployed on the Motor Vessel (MV) Argus during a trip to Antarctica in December 2024 for proof of concept that the portable system can characterise HPAI in the field.
Materials and methods
AIV samples used in this study
Oropharyngeal or cloacal swabs were collected from mallard ducks (Anas platyrhynchos) between 2007 and 2025 at selected sites in New Zealand for the New Zealand wild bird Avian Influenza surveillance programme (NZWBAIS)16,17. The sampling sites include Lake Te Rotokare, Parakai Springs, Paeroa, Kaituna, Te Awamutu, Turua, Kerepehi, Lake Aniwhenua, and Gisborne. Swabs were placed in avian transport media (ATM) containing 10,000 units/mL penicillin, 10 mg/mL streptomycin, 250 µg/mL gentamycin, 5000 units/mL myostatin, and 1% bovine serum albumin in isotonic phosphate-buffered saline (PBS) (pH 7.4). Swabs that tested positive by Influenza A matrix gene reverse transcription quantitative polymerase chain reaction (RT-qPCR) assay18,19,20 were inoculated into the allantoic cavity of 9- to 10-day-old embryonated specific-pathogen-free chicken eggs, using established procedures11,19. The sample sets used for this study are summarised in Table 1. AIV swabs from Antarctica (sample set 7), and the Otago Poultry farm (sample set 3) were collected in RNA/DNA shield (Zymo Research, Irvine, CA, USA).
Ethical approval
All procedures and methods carried out from the New Zealand surveillance samples used in this study, including the work of sampling and virus isolation in embryonated pathogen-free chicken eggs were obtained through approved Avian Influenza Surveillance programmes, permitted by the AgResearch Animal Ethics Committee, New Zealand under the permit titled “Avian Influenza in New Zealand Wild Birds -AEC-750”. Samples collected from Antarctica wild birds were collected under the Antarctic Treaty (Environment Protection) Act 1980 (ATEP Permit -Avian Influenza research on MV Argus (2024/2025 season)- reference number: 24/4877) permit, issued by the Australian Government Department of Climate Change, Energy, the Environment and Water, Australian Antarctic Division. All sampling and experimental procedures comply with relevant guidelines and regulations and were reported in accordance with the ARRIVE guidelines.
Laboratory-based nucleic acid extraction using an automated Kingfisher flex purification system
Total nucleic acid was extracted from 200 µL of sample (isolates or swabs) using the Applied Biosystem MagMAX™ CORE Nucleic Acid Purification Kit (Thermo Fisher Scientific, Waltham, MA, USA) on a KingFisher™ Flex Purification System (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol.
RNA extraction using QIAamp viral RNA mini kit on the portable system (Qiagen kit)
RNA was extracted using QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) on the portable Bento Lab Pro (Bento Bioworks Ltd, London, UK). Briefly 140 µL of viral swabs or isolates were lysed and the lysate was mixed with ethanol in QIAamp Mini column. The column was spun down and washed with Buffer AW1 and AW2 at 4000 x g or 8000 x g. RNA was eluted in 30 µL of buffer AVE.
RNA extraction using Zymo Quick-RNA viral kit on the portable system (Zymo kit)
RNA was extracted using Zymo Quick-RNA Viral Kit (Zymo Research, Irvine, CA, USA) on the portable Bento Lab Pro (Bento Bioworks). Viral RNA buffer was added to 400 µL sample in 2:1 ratio, transferred to the Zymo-Spin IC Column and spun down for 2 min. Beta-mercaptoethanol was omitted as recommended by the manufacturer when swabs were used. The column was washed with Viral Wash Buffer and ethanol at maximum speed. RNA was eluted in 15 µL DNase/RNAse-free water.
Influenza A RT-qPCR assay
An influenza A TaqMan-based RT-qPCR assay targeting the Matrix gene was performed using primers from Table 2 and Invitrogen SuperScript™ III Platinum One-Step qRT-PCR System (Thermo Fisher Scientific, Waltham, MA, USA). The assay was developed by Spackman et al. (2002) and modified by Heine et al. (2007). An internal control (18 S rRNA) RT-qPCR was included. All RT-qPCRs were performed either using a CFX Opus Real-Time PCR System (Bio-Rad, Hercules, California, USA) or the portable MIC PCR system (BioMolecular Systems, New South Wales, Australia).
The RT-qPCR assay was performed using the following conditions: for the reverse transcription stage, 1 cycle of 42 °C for 15 min, then 95 °C for 2 min. Thermal cycling was then performed with 45 cycles of 95 °C for 15 s, then 60 °C for 45 s18.
Evaluation of influenza A RT-qPCR assay on MIC PCR system
Analytical sensitivity of the Influenza A RT-qPCR assay19 was determined on the portable MIC PCR system (BioMolecular Systems) and compared with a reference laboratory-based system using the CFX Opus Real-time PCR system (Bio-Rad). RNA from an AIV isolate A/Mallard/NZ/NZ0700316_2/2007(H12N5) was serially diluted from 10− 1 to 10− 8 dilution and analysed in triplicates on both systems. Instrument-specific software was used to analyse Cq values, using an automatic calculation generated from their baseline threshold. A Cq ≤ 45 was considered positive for AIV.
Further Influenza A RT-qPCR assay tests were performed on n = 37 swabs from wild mallard ducks collected from NZWBAIS (sample set 2). The total nucleic acid from these samples was extracted and the assay was performed in duplicate using both MIC PCR (BioMolecular Systems) and CFX Opus Real-time PCR system (Bio-Rad). The relative agreement of results between the two systems were described using Kappa statistics21.
For additional confirmation, RNA of n = 5 swabs from the HPAI H7N6 Otago poultry farm (sample set 3) was tested for the level of Influenza A virus in the sample using the MIC PCR system (BioMolecular Systems).
Whole-genome amplification (WGA) of AIV by RT-PCR
AIV specific WGA by multi-segment RT-PCR was performed using the primers listed in Table 3 (Mitchell et al., 2021, method B)22 These primers target conserved regions across all eight segments of the AIV genome. For both protocols, the primers were added to the Invitrogen SuperScript™ III One-Step RT-PCR System with Platinum™ Taq High Fidelity mix (Thermo Fisher Scientific, Waltham, MA, USA) in a 20 µL total volume according to Method B of Mitchell et al. (2021). Thermal cycling was performed under the following conditions: reverse transcription to cDNA at 55 °C for 30 s, 42 °C for 50 min and 94 °C for 2 min. Following the reverse transcription, 5 cycles of 94 °C for 15 s, 42 °C for 30 s and 68 °C for 3 min and 20 s (4 min when using Bento Lab Pro). A second round of cycling was performed with 35 cycles of 94 °C for 15 s, 57 °C for 30 s and 68 °C for 3 min 20 s (4 min when using Bento Lab Pro), with a final extension of 68 °C for 5 min.
Evaluation of WGA of AIV on portable Bento lab pro
AIV WGA by multi-segment RT-PCR was performed on the total nucleic acid extracted from the AIV isolates (sample set 1) using both the portable Bento Lab Pro System (Bento Bioworks) and the laboratory-based Applied Biosystems Veriti™ thermal cycler (Thermo Fisher Scientific, Waltham, MA, USA).
Following WGA, cDNA amplicons were purified using the Applied Biosystems ExoSAP-IT™ Express PCR Product Cleanup reagent (Thermo Fisher Scientific, Waltham, MA, USA). In the laboratory, the cDNA amplicon sizes were checked using the gDNA ScreenTape Assay (Agilent Technologies, Santa Clara, CA, USA) on the Agilent TapeStation 4200 system (Agilent Technologies, Santa Clara, CA, USA). For the portable system, cDNA amplicons were assessed on Invitrogen E-Gel™ Power Snap Electrophoresis System (Thermo Fisher Scientific, Waltham, MA, USA) using an Invitrogen pre-cast E-Gel EX 1% agarose cassette (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol. 1 µL of each product was quantified using the Invitrogen Qubit™ 1X dsDNA High Sensitivity Quantification Assay kit (Thermo Fisher Scientific, Waltham, MA, USA) on an Invitrogen Qubit™ 4.0 fluorometer (Thermo Fisher Scientific, Waltham, MA, USA).
Library preparation for sequencing using Oxford nanopore technology (ONT)
For both the portable and laboratory-based protocols, purified amplicons were prepared for sequencing using the Rapid Barcoding kit 24 V14 (SQK-RBK114.24) (Oxford Nanopore Technologies, London, UK). Rapid PCR Barcoding Kit 24 V14 (SQK-RPB114.24) (Oxford Nanopore Technologies, London, UK) was used in the laboratory-based protocol, but not for the portable system. For the portable protocol, pooled libraries were loaded onto a R10.4.1 MinION flow cell (Oxford Nanopore Technologies, London, UK) and sequenced using a MinION Mk1B device (Oxford Nanopore Technologies, London, UK) for up to 18 hours. For the laboratory protocol, pooled libraries were loaded onto either a R10.4.1 MinION or PromethION flow cell (Oxford Nanopore Technologies, London, UK), then sequenced using a Mk1B or a P2 Solo instrument (Oxford Nanopore Technologies, London, UK) for up to 18 hours.
Sequencing setup was performed using MinKNOW version 22.10.10 on a laptop with Windows 10 Pro operating system, 16.0 Gb RAM, Intel® Core(TM)-i7-9850 H CPU and 1 TB internal storage as recommended by the manufacturer.
Data analysis and real-time subtyping
Basecalling was performed using Dorado (v7.0.2) with a high-accuracy base calling model v4.3.0. Two methods were used for characterising viral subtypes. The sequences were either manually mapped to a non-redundant reference gene set containing n = 449 sequences from the Global Initiative on Sharing All Influenza Data (GISAID) database (https://gisaid.org/), which is the current standard method for sequence-based subtype assignation, or by using Flui (see https://github.com/dragonfly-science/flui for details on implementation and usage), an automated real-time AIV subtyping tool developed in-house by Dragonfly Data Science. Subtypes generated using Flui were also confirmed by manual mapping to the non-redundant gene set obtained from the GISAID database.
For manual mapping method, raw reads were mapped to the GISAID reference gene set using Minimap2 v2.24 in Geneious Prime v2021.1.1 (https://www.geneious.com)23. HA and NA subtypes were characterised by identifying the references with the greatest number of reads with high uniformity of coverage that mapped to each of the segments. When the average depth of coverage was ≥ 30, consensus sequences were extracted from HA and NA genes and assembled into a single genome. Poor quality sequences were trimmed off. Consensus sequences were deemed to be high quality when the number of nucleotides were called at Q30 + ≥ 95%. For sequences that did not meet these quality thresholds, a “low read” or “—” was noted in the subtyping result.
For automated subtype classification using Flui, HA and NA subtypes were called when both the number of k-mers for the HA and NA genes exceeded 2,000, and the differentiation score exceeded 6.0 with a gap threshold of 2.5. For samples that did not meet these quality thresholds, a “low read” or “—” was noted in the subtyping result.
The pathogenicity status of an influenza virus (specifically, HPAI) was determined by cleavage site analysis of the consensus sequence in Geneious Prime v2021.1.1. A multi-basic motif in the cleavage site classifies the virus as HPAI based on the OFFLU guide Influenza A cleavage site guide12.
All computational analyses and experimental protocols were done using the default parameters unless noted otherwise. The relative agreement of reads passing quality thresholds between manual mapping and Flui were described using Kappa statistics21.
Sample collection and AIV subtyping in the field, in Antarctica
All the essential portable devices in Fig. 1 and consumables were packed into a large black hardshell suitcase (Fig. 2a and b). Swabs were collected from a brown skua (Stercorarius antarcticus), a south polar skua (Stercorarius maccormicki) and a seal pup in various locations in Antarctica following a verbal report of suspicious HPAI mortality from the United States Antarctic Program (Palmer Station, Anvers Island)24. The swabs collected were processed and sequenced using the in-field portable system as in Fig. 3. Influenza A RT-qPCR assay and gel electrophoresis were not performed prior to library preparation. Flui was used to generate the subtype, and cleavage site analysis was performed using Geneious Prime.
Equipment for the AIV portable system. Essential equipment: (i) Bento Lab Pro System (Bento Bioworks Ltd, London, UK), (ii) Qubit 4.0 Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA), (iii) Vortex, (iv) Magnetic rack, (v) Laptop and MinION device (Mk1B) (Oxford Nanopore Technologies, London, UK). Optional equipment: (i) HulaMixer™ Sample Mixer (Thermo Fisher Scientific, Waltham, MA, USA), (ii) MIC PCR system (BioMolecular Systems, New South Wales, Australia), (iii) Invitrogen E-Gel Power Snap Electrophoresis System (Thermo Fisher Scientific, Waltham, MA, USA). Created in BioRender. Wilson, A. (2025) https://BioRender.com/7erudlf.
Portable lab in a suitcase. a Hardshell suitcase that fits all portable field equipment and consumables. b Consumables, reagents and portable field equipment packed in the suitcase.
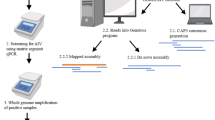
Portable AIV characterisation system. (1) Swabs were collected from mallard ducks in several locations around New Zealand between 2023 and 2024. (2) RNA was extracted using QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) on the Bento Lab Pro (Bento Bioworks Ltd, London, UK) centrifuge component. The Influenza A RT-qPCR assay can be performed on the MIC PCR instrument (BioMolecular Systems, New South Wales, Australia) prior to whole-genome amplification of the virus. (3) Whole-genome amplification by multi-segment AIV-specific RT-PCR was performed on the Bento Lab Pro thermal cycler component. (4) Quantification of purified amplicons was measured using Qubit 4.0 fluorometer (Thermo Fisher Scientific, Waltham, MA, USA). There is an option to run the purified amplicons on a precast electrophoresis gel to determine the size of the amplicons prior to sequencing (not shown). (5) Amplified cDNA products were prepared for sequencing using the Oxford Nanopore Technologies Rapid Barcoding kit V14 (Oxford Nanopore Technologies, London, UK) and sequenced using R10.4.1 MinION flow cells (Oxford Nanopore Technologies, London, UK). (6) AIV subtypes for each sample were identified using the automated subtyping pipeline that we developed, Flui. In blue boxes, are the approximate time needed for each step. Created in BioRender. Wilson, A. (2025) https://BioRender.com/7erudlf.
Results
Evaluation of portable equipment using AIV isolates
Samples with established HA and NA subtypes, which had been previously identified through the laboratory system (Fig. 4), were used to evaluate the portable devices. These were the Bento Lab Pro, the Invitrogen E-Gel Power Snap Electrophoresis System, and the MIC PCR system.
Laboratory-based AIV subtype characterisation workflow. (1) Swabs were collected from mallard ducks at several locations around New Zealand. (2) Swabs were sent to the laboratory for AIV testing. Nucleic acid from the collected swabs or AIV isolates was extracted using the KingFisher MagMAX CORE Total Nucleic Acid Purification Kit (Thermo Fisher Scientific, MA, USA). (3–4) The presence of Influenza A virus was tested using the Influenza A RT-qPCR assay. Swabs that were tested positive by RT-qPCR were inoculated into embryonated chicken eggs. (5) Whole-genome amplification was performed by multi-segment reverse transcription PCR (RT-PCR) using AIV-specific primers. (6–7) The size of purified whole-genome amplified AIV amplicons were assessed on a 4200 TapeStation using Genomic DNA (gDNA) ScreenTape Assay Kit (Agilent Technologies, Santa Clara, CA, USA), and the concentration of the amplicons were quantified using the Invitrogen Qubit™ 1 x dsDNA High Sensitivity Quantification Assay kit (Thermo Fisher Scientific, Waltham, MA, USA) on an Invitrogen Qubit™ 4.0 fluorometer (Thermo Fisher Scientific, Waltham, MA, USA). (8) cDNA was prepared for sequencing using the Oxford Nanopore Technologies (ONT) Rapid PCR Barcoding Kit V14 (Oxford Nanopore Technologies, London, UK) or Rapid Barcoding Kit V14 (Oxford Nanopore Technologies, London, UK) and sequenced using R10.4.1 PromethION or MinION flow cells (Oxford Nanopore Technologies, London, UK). (9) AIV subtypes were identified by manually mapping sequenced reads to a non-redundant Global Initiative on Sharing All Influenza Data (GISAID) dataset. Created in BioRender. Wilson, A. (2025) https://BioRender.com/7erudlf.
To evaluate the performance of WGA on Bento Lab Pro, a mixture of AIV-positive and -negative isolates with known subtypes were used. The samples were processed through the portable system (Fig. 3) to determine influenza subtypes. Results were compared to the known subtypes to assess the veracity of the equipment.
Twelve viral isolates (sample set 1), comprising of 8 AIV-positive and 4 AIV-negative samples were selected to assess the performance of the portable Bento Lab Pro thermal cycler component in performing WGA. All amplicons of AIV-positive samples generated from both the Bento Lab Pro thermal cycler and the laboratory-based Veriti thermal cycler (Fig. 5a) exhibited the expected AIV-specific amplicon profile like in Fig. 5b. Non-specific amplicons were generated from AIV-negative samples (sample 7, 9, 11 and 12) (Fig. 5a). An example of an AIV-specific amplicon profile for AIV-positive sample in the TapeStation ScreenTape image is shown in Fig. 5b.
AIV-specific amplicons profile on TapeStation 4200 gel images using the gDNA ScreenTape assay. a AIV isolates from sample set 1 that have undergone AIV multi-segment RT-PCR whole-genome amplification on Bento Lab Pro thermal cycler (portable system) and the Veriti thermal cycler (laboratory-based system). Sample numbers in black are AIV-positive samples showing AIV-specific amplicon profile. Sample numbers in red are AIV-negative samples with non-specific amplification. b AIV-specific amplicon profile for AIV-positive sample. Lane M, molecular standards. Lane P, amplicons from an AIV-positive sample. The image was cropped to show one example of a positive sample. Full image shown in Supplementary Fig. 4.
To determine whether the Invitrogen E-Gel Power Snap Electrophoresis System can replace the function of the TapeStation system in the field, amplicons for 10 of the 12 samples from sample set 1 were also assessed on a pre-cast 1% E-Gel using the E-Gel EX Power Snap system (Fig. 6). Only ten samples (8 AIV-positive, 2 AIV-negative) were assessed on this system because it is limited to ten samples per run. The amplicons from AIV-positive samples also demonstrated the right AIV amplicon profile as in Fig. 5b when gel electrophoresis was performed using the E-Gel EX Power Snap system (Fig. 6).
1% E-Gel Ex image of AIV-positive and negative multi-segment RT-PCR whole-genome amplified amplicons. A 1 kb plus ladder was used (Lane M). From left, sample 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 as indicated by the lane numbers. Positive samples feature AIV-specific amplicon profile, whereas negative samples have non-specific amplification. Sample 7 and 9 are known AIV-negative samples. The blue arrow marks 2000 bp, orange arrow marks 1500 bp and green arrow marks 1000 bp. Full image shown in Supplementary Fig. 5.
Library preparation was performed on the amplicons from sample set 1 that was generated using Bento Lab Pro and were sequenced. All positive samples were analysed by manual mapping to a non-redundant reference gene set downloaded from GISAID database. The subtypes were identified correctly in all the AIV-positive samples (Table 4).
Evaluation of the influenza A RT-qPCR assay on MIC PCR system
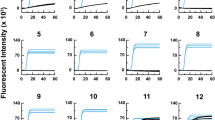
Analysis of a serially diluted AIV RNA sample was performed to assess the analytical sensitivity of the Influenza A RT-qPCR assay on the MIC PCR and compared with a reference system (CFX Opus). Both PCR systems detected the presence of AIV genome across five dilution points (10− 1 to 10− 5) in all replicates (Table 5). Although one replicate was positive at the 10− 7 dilution using the MIC PCR, the limit of detection was comparable at 10− 5 dilution in both systems, where all three replicates were reliably detected with AIV gene fragment (Table 5).
Influenza A RT-qPCR assay was further tested on both the MIC PCR and CFX Opus system using n = 37 swabs from sample set 2. Both systems showed near perfect agreement (Kappa = 0.934, 0.80–1.06 95% CI) in classifying positive or negative samples (36/37) in this panel (Supplementary Table 1). Twenty-six samples were positive for AIV RNA and ten were negative in both systems. One test disagreement observed, (sample 72) had a very late Cq value (39.27) on the MIC PCR system. This is not unexpected, as the stochastic behavior of amplification for samples with very low target concentrations can produce variable results.
The influenza A RT-qPCR assay was also performed on n = 5 swabs from sample set 3 using the MIC PCR system. These samples were known to be HPAI H7N6, based on laboratory system results. RNA from all five samples was processed using the portable system and identified as AIV- positive samples, with an average Cq < 45 (Table 6).
Evaluation of the RNA extraction kit on the portable system using AIV isolates
To determine a more suitable RNA extraction kit for AIV subtyping in the portable system, RNA from an additional 10 isolates with known subtypes (sample set 4 and 5) was extracted using either the QIAamp Viral RNA Mini Kit (Qiagen Kit) or the Zymo Quick-RNA Viral Kit (Zymo Kit) on the Bento Lab Pro instrument. Of the seven samples extracted using the Qiagen Kit, one sample, (sample 21) was omitted from library preparation as it did not show the AIV-specific amplicon profile, consistent with what would be expected for an AIV-negative sample (Table 7). Five of the six sequenced samples resulted in correct assignment of subtype. Sample 19 was partially characterised. The three remaining isolates that were extracted using the Zymo Kit and sequenced using the portable method featured the same subtypes previously generated using the laboratory method (Table 7). Both Qiagen and Zymo kits correctly identified HA and NA subtypes in AIV isolates.
Evaluation of the RNA extraction kits on the portable system using swabs
The portable AIV characterisation system was further tested on five H7N6 HPAI-positive swabs collected from the Otago poultry farm (sample set 3)25. Total RNA was extracted using both Qiagen and Zymo kit and then characterised to evaluate their application for the portable system.
All n = 5/5 swabs extracted using the Qiagen kit were fully characterised as H7N6 HPAI, by manual mapping method (Table 8). For the Zymo kit, n = 2/5 swabs were fully characterised, one was partially characterised and two failed to meet the quality thresholds (Table 8). The Zymo kit had a lower identification rate with 40% (5.27–85.34 95% CI) of samples identified as H7 and 60% (14.66–94.73 95% CI) of samples as N6, compared to 100% (56.55–100 95% CI for HA and 47.82–100 95% CI for NA) samples identified as H7N6 for Qiagen kit. The average depth of coverage in HA and NA segments generated from Qiagen-extracted samples was also generally better than Zymo-extracted samples (Table 8). These findings suggest that the QIAamp Viral RNA Mini Kit outperforms the Zymo Quick-RNA Viral Kit for extracting RNA from swabs.
A consensus sequence of HA regions from the assembled HA gene from Qiagen extracted-sample 13a, revealed a six-nucleotide insertion and a separate single nucleotide variant. This increased the basic amino acids at the cleavage site from one to five- specifically PEGPKRRKR/GLF (Fig. 7). According to WOAH and OFFLU guidelines12, this permits characterisation as HPAI. This result is consistent with the results from the laboratory system25.
Detection of a multi-basic cleavage site in the HA gene from the Qiagen extracted RNA of sample 13a with an accession number W24_2663-1. Nucleotide alignment was generated using MAFFT v7.490. The alignment features the detected H7N6 HPAI HA sequence (first sequence in alignment) with H7N7 LPAI HA sequences isolated through active surveillance in New Zealand (remaining 7 sequences). Alignments were visualised using Geneious Prime® v2021.1.1.
Evaluation of flui as an automated real-time influenza subtyping tool
All sequences generated from sample sets 1, 3, 4 and 5 were also classified using Flui. Subtypes generated by Flui were compared to subtypes identified through manual reference mapping (Table 9). The subtypes generated by both manual and automated methods were comparable to their known subtypes when the reads passed their respective quality thresholds. Although both methods failed to pass HA and NA quality thresholds in two samples (16b and 17b), they had perfect agreement (Kappa = 1.0) in passing the reads of all NA segments (Table 9). Flui and manual mapping methods disagreed in passing HA quality thresholds for sample 17a and 23, resulting in an agreement of 92.59% (25/27). Both methods had substantial agreement (Kappa = 0.76, 0.44–1.07 95% CI) in establishing the passing criteria for HA segment.
HA/NA subtype classification of AIV using the portable system
Using the portable system in Fig. 3, an additional n = 45 swabs collected from mallard ducks at various locations around New Zealand, between 2023 and 2024 (sample set 6), were processed in the laboratory. Three positive controls with known subtypes were added to the cohort. Real-time automated subtyping was performed using Flui and the results were validated by manual mapping. For HA, there is a substantial agreement between the two methods (Kappa = 0.77, 0.58–0.96 95% CI) and a fair agreement for NA (Kappa = 0.33, 0.08–0.59 95% CI). The agreement to pass the NA segment for both methods was lower in this sample set as there were more samples that failed the quality thresholds in either one of the subtyping methods (Supplementary Dataset 1). Most of these samples have lower viral load, Cq > 32. All other swabs were identified and listed in Supplementary Dataset 1.
One sample was artificially mixed in equal volumes of H4N6 and H11N2 samples to mimic a co-infection. Flui identified the mixed sample as H11N2/N6 with the highest score, followed by H4, which had the second-highest score. Manual mapping was able to characterise the sample as H11N6 or N2. H4 did not pass the quality thresholds set for both methods due to the lower number of amplified H4 segments present in the sample. Whilst both methods were able to recognise co-infections in the NA segments, further work is needed to improve upon the accuracy of the output. The three positive control samples were subtyped correctly as H3N8, H11N6 or H11N2 (Supplementary Dataset 1).
Portable in-field system deployment in Antarctica and identification of HPAI
Our portable system was tested in the field, for the first time outside of a laboratory environment, in Antarctica. Five swabs (sample set 7) were collected from carcasses that were suspected of HPAI mortality; a brown skua (Stercorarius antarcticus), a south polar skua (Stercorarius maccormicki) and a seal pup. The swabs were brought back to the MV Argus Ship for sample processing within a few hours. In the ship, the samples were processed using the portable system. RNA was extracted using the Qiagen-portable system, AIV was amplified using whole-genome amplification on Bento lab Pro and the library was prepared for sequencing on the ONT MinION Flow Cell using the Rapid Barcoding Kit (Fig. 3). Two samples from the same skua were identified as H5N1 HPAI after cleavage site analysis24, while the other samples were negative for AIV (Table 10).
Discussion
An easy to perform portable AIV characterisation system is needed for rapid detection of HPAI in the field. Here, we presented a fast, accurate, and highly portable laboratory system capable of rapidly characterising AIV subtypes in real-time and offline for the first time in New Zealand. We have tested the system using AIV isolates and swabs and have successfully demonstrated that HPAI can be identified in the field with our portable system. Our portable system can be deployed anywhere with access to power supply, including in a rural setting, boat, poultry farm or shed.
Many researchers have employed amplicon-based ONT sequencing due to the speed, and capability of real-time analysis26,27,28,29,30,31. The portability of the ONT platform has also enabled rapid identification of AIV subtypes in the field26,28,31. Some groups have used the longer library preparation workflow such as ONT Ligation Sequencing Kit or Native Barcoding Kit26,28,29, which is not ideal for a rapid portable system. We along with other groups30,31 selected the Rapid Barcoding Kit (RBK) as it requires minimal input of DNA and offers shorter library preparation time, making it well-suited for field application.
While our library preparation method was similar to that described by de Vries et al. (2022) and King et al. (2020), our portable system was designed to achieve an optimal balance of affordability, portability, high nucleic acid integrity, and rapidity of workflow. de Vries et al. (2022) used the crude Biomeme M1Bulk Sample Prep Kit and focused solely on classifying the HA subtype; while King et al. (2020) looked at the whole genome of the virus and classified the virus by manual mapping method. Given the high cost of the Biomeme extraction kit and the importance of obtaining complete subtypes or sequence of longer viral segments where possible, a column-based extraction approach was chosen for our system to preserve RNA integrity. An automated real-time subtyping pipeline was also developed in our system to identify the virus full subtypes without the need for internet access. Our portable system was developed so that it is not restricted to AIV field diagnostic testing and can be modified to test for other pathogens of significant interest using the same core equipment and reagent kits. While several studies have used amplicon-based sequencing approach, PCR-free approaches such as direct RNA sequencing have also been explored32,33. However, this method is not suited for rapid field workflow due to RNA instability, and the long library preparation protocol. The ONT direct RNA library preparation kit requires high input of RNA and has a limit of detection at a Cq of 1833. This is much lower than the Cq values we obtained from swabs collected in the field.
We focused on initially evaluating the portable equipment including, the Bento Lab Pro system, the MIC PCR system and the Invitrogen E-Gel Power Snap Electrophoresis System, for generating high-quality whole-genome amplified products and identifying AIV subtypes from AIV isolates with known subtypes. Next, the portable system was tested in the Animal Health Laboratory (AHL) of Ministry for Primary Industries (MPI) in New Zealand using swabs collected from several locations in New Zealand. Our results show that the portable system can generate high-quality products and accurately determine subtypes, like the laboratory systems. Finally, the system was deployed to Antarctica where HPAI H5N1 was identified in a skua24 within two days from sample collection. This would have taken weeks if the samples were to be transported back to a laboratory in AHL MPI New Zealand.
This study has shown that both Zymo and Qiagen extraction kits could be used to successfully identify the correct subtypes in isolates. Although the sample size is limited, experiment conducted shows that the Qiagen kit outperformed the Zymo kit when swabs were used. All samples tested were identified as H7N6 HPAI when using Qiagen kit (Table 8). This may be attributed to exclusion of beta-mercaptoethanol in the Zymo protocol as suggested by the supplier when using swabs. Furthermore, the high viral load and large volume of sample (400 µL) used with the Zymo-Spin IC Column, designed for low viral load may have inhibited the column to perform optimally. Based on these results, the QIAamp Viral RNA Kit is the preferred RNA extraction kit used for AIV characterisation in the field at this stage. Further optimisation of sample input volume and column compatibility is needed for the Zymo Quick-RNA Viral Kit to be used in the field. The multi-basic cleavage site of the H7N6 HPAI HA segment was also successfully identified using the Qiagen in-field method (Fig. 7), suggesting that our data generated from the portable field systems are reliable and accurate when compared to data generated in the laboratory.
We have also demonstrated that the influenza A RT-qPCR assay can be performed using the MIC PCR system to determine the presence of AIV-positive samples. Our study verified that the portable MIC PCR system reliably distinguishes between AIV-positive and -negative (Supplementary Table 1). However, we propose that, for a successful sequencing of both HA and NA subtypes, it is more important to identify the distinct AIV-specific amplicon profile in a sample, rather than relying on positive Cq values. A positive Cq value (≤ 45) does not always guarantee subtype identification, as sample degradation or low viral load can often affect the quality of the sequences (Supplementary Dataset 1). We suggest that visual identification of AIV amplicon sizes in an E-Gel electrophoresis or any other electrophoresis platform is a better indication for assurance of sequencing quality as it shows the quality of the amplified segments prior to library preparation. This is evident in a separate internal quality control study where we saw negative correlation between Cq values and average read depth of HA and NA (Supplementary Fig. 1 and Supplementary Tables 2, 3, 4). Spearman’s rank correlation coefficient test was performed using the cor.test () function from the stats package in R34 with HA segment rho = −1, p = 5.511e-06 and for NA segment rho = −0.98, p-value = 0.0003968. The study also demonstrated that sample with Cq of 32 or higher failed to generate a full subtype in all replicates (Supplementary Tables 2, 3 and 4) and did not present with AIV-specific amplicon profiles (Supplementary Fig. 2). This is consistent with what we concluded from our study from sample set 6 (Supplementary Dataset 1) where most samples with viral load above Cq 32 would not meet the quality thresholds required for either Flui or manual mapping. However, in some instances samples with Cq < 32 can also fail to generate a full subtype (Supplementary Dataset 1). We believe that these may be attributed to sample degradation, contamination or co-infections. Hence, an AIV-specific amplicon profile can be assessed on a portable gel electrophoresis system to avoid sequencing AIV-positive samples that might not pass quality thresholds due to the above reasons.
When assessing AIV-specific amplicon profiles of samples with high integrity RNA, a similar profile should be observed on a TapeStation gDNA ScreenTape assay or any other portable E-gel electrophoresis devices such as Bento-gel (Supplementary Fig. 3). Slight variability in the AIV amplicon profile can be expected when sample RNA is of lower quality. The presence of AIV-specific amplicon profiles in AIV-positive samples was also validated in a separate study (work in progress for publication) where we have successfully sequenced more than 300 samples for developing a New Zealand LPAI genome database. All samples with full genome established also presented with the AIV-specific amplicon profile.
For sequencing reads that passed the quality thresholds in Flui or manual mapping, all results were consistent with their known subtypes. Most samples with Cq < 32 were successfully subtyped in both Flui and manual mapping methods, n = 27/34. Our results also suggested that the two methods can complement each other to fully characterise subtypes. When both methods were used, the number of samples with Cq < 32 and were fully subtyped increased to n = 31/34. Since both Flui and manual mapping methods use different algorithms for AIV subtyping, the quality thresholds are set differently between the two methods. Samples with low viral load (Supplementary Dataset 1) or degraded RNA can result in partially characterised subtypes from either one of the methods. Despite the discrepancy of agreement in passing thresholds when viral load is low, Flui was used to subtype more than 300 LPAI isolates in a separate study as previously mentioned then verified by manual method. All subtypes were consistent in both methods.
Our results also demonstrated that some H4 segments were not amplified as efficiently as other HA subtypes due to slight variation in the sequence, reducing the efficiency of primer binding to the New Zealand H4 sequence. Hence most H4 samples failed to meet the quality thresholds cut-offs in both methods. The failure of H4 not being identified and passing the quality threshold is evident in the sample that was artificially mixed. To address this, we incorporated primers that are specific to the New Zealand H4 segment sequence in the RT-PCR whole-genome amplification assay. The primer sequences are provided in Supplementary Tables 535.
With the portable systems developed, the in-field system was successfully deployed to Antarctica in late 2024, where it enabled the identification of H5N1 HPAI belonging to clade 2.3.4.4b in a skua24. Temperature-sensitive reagents were stored in an insulated box to ensure the reagents were kept within the recommended temperature. Whilst an insulated box was used to keep temperature-sensitive reagents at the appropriate temperature, some reagents were unavoidably brought to ambient temperature during travel yet did not affect sequencing performance or results. All equipment proved durable and performed as expected.
In future, alternative rapid AIV identification methods, such as the reverse transcription loop-mediated isothermal amplification (rRT-LAMP) can be explored. A LAMP method amplifies viral RNA at a constant temperature without complex thermal cycling conditions. In combination with usage of a commercial universal LAMP kit (e.g., WarmStart Colorimetric LAMP kit, BioLabs Inc. New England), AIV can be detected within 30 min of receiving the samples36. This method is cheaper and faster than the Influenza A RT-qPCR assay.
A lyophilised master mix for RT-qPCR can also be used in the field to reduce the number of temperature-sensitive reagents needed. Initial testing conducted at AHL using lyophilised master mix (LyoPrime Luna® Probe One-Step RT-qPCR Mix with UDG, New England Biolabs) on the influenza A RT-qPCR assay showed comparable analytical sensitivity to the SuperScript III RT-qPCR master mix used in this study (Supplementary Table 6). The portable system can also be deployed to the field during the AHL annual AIV surveillance trip to evaluate its performance across a broader range of AIV strains, ensuring its effectiveness across all AIV subtypes.
Our portable system is developed for rapid field diagnostic and surveillance of a HPAI index case or in a response setting where rapid and accurate characterisation of AIV is needed. It is not intended to be used in a large-scale sample testing environment. The portable in-field system is also not restricted to AIV subtyping. It can be adapted for in-field testing for other pathogens of potential economic importance e.g., foot-and-mouth disease virus. In the future, further refinement to the portable system can be made by identifying a suitable extraction kit capable of isolating total nucleic acids, thereby permitting the analysis of both DNA- and RNA-bound pathogen genomes in a single workflow.
Conclusion
For outbreaks of viral disease, the early and accurate identification of the causative virus is critical for guiding containment strategies, and surveillance. For avian influenza, there is a need to develop advanced tools for field-based characterisation. This ensures that authorities can respond swiftly and effectively to reduce the impact of HPAI spread, thereby minimising adverse effects on animal and human health, as well as on the economy. We successfully demonstrate that our portable system can provide rapid and accurate subtyping of AIV in the field confidently, within two days upon sample collection, and can characterise HA cleavage site sequence. Our platform can significantly reduce the delays that are associated with sample transport to a laboratory, and this allows authorities to make quicker and more-informed decisions.
Data availability
The datasets used and/or analysed in this study are available from the corresponding author on reasonable request. All sequence data from sample sets 1, 3, 4, 5 and 6 have been deposited to the NCBI Sequence Read Archive (SRA) under [BioProject ID PRJNA1374278](https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1374278). Samples are accessible under IDs SAMN53683369 to SAMN53683442. Mixed sample in Supplementary Dataset 1 is not deposited as it is an artificially mixed combination of sample W23\_0006-225 (ID SAMN53683408) and W23\_00141-0107 (ID SAMN53683415). Supplementary Dataset 2 contains all relevant metadata, including accession numbers, sample names, and associated URLs. The H5N1 HPAI skua sample from sample set 7 has been deposited under Sequence Read Archive BioProject ID [PRJNA1210083](https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1210083), [GISAID accession EPI_ISL_19645365](https://platform.epicov.org/epi3/frontend), and GenBank accessions [PV163161-8](https://www.ncbi.nlm.nih.gov/nuccore/?term=A%2FBrown+Skua%2FTorgersen+Island%2Fo81-b82%2F2024).
References
Webby, R. J. & Uyeki, T. M. An update on highly pathogenic avian influenza A(H5N1) Virus, clade 2.3.4.4b. J. Infect. Dis. 230, 533–542. https://doi.org/10.1093/infdis/jiae379 (2024).
Peacock, T. P. et al. The global H5N1 influenza panzootic in mammals. Nature 637, 304–313. https://doi.org/10.1038/s41586-024-08054-z (2025).
CDC. Past Reported Global Human Cases with Highly Pathogenic Avian Influenza A (H5N1) (HPAI H5N1) by Country, 1997 2005. https://www.cdc.gov/bird-flu/php/avian-flu-summary/chart-epi-curve-ah5n1.html (2025).
Palese, P. & Young, J. F. Variation of influenza A, B, and C viruses. Science 215, 1468–1474. https://doi.org/10.1126/science.7038875 (1982).
Cheung, T. K. & Poon, L. L. Biology of influenza a virus. Ann. N Y Acad. Sci. 1102, 1–25. https://doi.org/10.1196/annals.1408.001 (2007).
Alexander, D. J. An overview of the epidemiology of avian influenza. Vaccine 25, 5637–5644. https://doi.org/10.1016/j.vaccine.2006.10.051 (2007). https://doi.org:.
Tong, S. et al. New world bats harbor diverse influenza A viruses. PLoS Pathog. 9, e1003657. https://doi.org/10.1371/journal.ppat.1003657 (2013).
Pantin-Jackwood, M. J. et al. Pathogenicity and transmission of H5 and H7 highly pathogenic avian influenza viruses in mallards. J. Virol. 90, 9967–9982. https://doi.org/10.1128/jvi.01165-16 (2016).
Munster, V. J. et al. Spatial, temporal, and species variation in prevalence of influenza A viruses in wild migratory birds. PLoS Pathog. 3, e61. https://doi.org/10.1371/journal.ppat.0030061 (2007).
Lu, L., Lycett, S. J. & Leigh Brown, A. J. Reassortment patterns of avian influenza virus internal segments among different subtypes. BMC Evol. Biol. 14, 16. https://doi.org/10.1186/1471-2148-14-16 (2014).
WOAH. Avian influenza (including infection with high pathogenicity avian influenza viruses) Chap. 3.3.4 in Terrestrial Manual, 1–26 (2025).
OFFLU. Influenza A cleavage sites in H5 and H7 viruses (2022).
de Bruin Anja, C. M., Monique, S., Bestebroer Theo, I., Fouchier Ron, M., Richard, M. & A. M. & Conserved expression and functionality of Furin between chickens and ducks as an activating protease of highly pathogenic avian influenza virus hemagglutinins. Microbiol. Spectr. 11, e04602–04622. https://doi.org/10.1128/spectrum.04602-22 (2023).
Lee, D. H., Lea, K. T. M., Ian, K. M., Swayne, D. E. & B. & and Genome sequences of haemagglutinin cleavage site predict the pathogenicity phenotype of avian influenza virus: statistically validated data for facilitating rapid declarations and reducing reliance on in vivo testing. Avian Pathol. 53, 242–246. https://doi.org/10.1080/03079457.2024.2317430 (2024).
Jonges, M. et al. Emergence of the virulence-associated PB2 E627K substitution in a fatal human case of highly pathogenic avian influenza virus A(H7N7) infection as determined by illumina ultra-deep sequencing. J. Virol. 88, 1694–1702. https://doi.org/10.1128/jvi.02044-13 (2014).
Marquetoux, N., King, S. W. & Mulqueen, C. K. Avian influenza surveillance programme annual report, 36–42 (2024).
Sokolova, M. et al. Avian influenza surveillance programme annual report, 28–34 (Ministry for Primary Industries, Surveillance, 2023).
Spackman, E. et al. Development of real-time RT-PCR for the detection of avian influenza virus. Avian Dis. 47, 1079–1082. https://doi.org/10.1637/0005-2086-47.s3.1079 (2003).
Stanislawek, W. L. et al. Avian influenza viruses in new Zealand wild birds, with an emphasis on subtypes H5 and H7: their distinctive epidemiology and genomic properties. PLOS ONE. 19, e0303756. https://doi.org/10.1371/journal.pone.0303756 (2024).
Heine, H. G., Trinidad, L., Selleck, P. & Lowther, S. Rapid detection of highly pathogenic avian influenza H5N1 virus by TaqMan reverse transcriptase-polymerase chain reaction. Avian Dis. 51, 370–372. https://doi.org/10.1637/7587-040206r.1 (2007).
Sergeant, E. Epitools Epidemiological Calculators (2018). http://epitools.ausvet.com.au
Mitchell, P. K. et al. Method comparison of targeted influenza A virus typing and whole-genome sequencing from respiratory specimens of companion animals. J. Vet. Diagn. Invest. 33, 191–201. https://doi.org/10.1177/1040638720933875 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100. https://doi.org/10.1093/bioinformatics/bty191 (2018).
McCulley, M. et al. High pathogenicity avian influenza (HPAI) H5N1 virus detected in brown Skua using portable laboratory while at sea in Antarctica. Microbiol. Resour. Announc. https://doi.org/10.1128/mra.00041-25.
McCulley, M. et al. High pathogenicity avian influenza (HPAI) H7N6 virus detected in New Zealand poultry. Microbiol. Resour. Announc. https://doi.org/10.1128/mra.00088-25.
Croville, G. et al. An amplicon-based nanopore sequencing workflow for rapid tracking of avian influenza outbreaks, France, 2020–2022. Front. Cell. Infect. Microbiol. 14, 1257586. https://doi.org/10.3389/fcimb.2024.1257586 (2024).
Wang, J., Moore, N. E., Deng, Y. M., Eccles, D. A. & Hall, R. J. MinION nanopore sequencing of an influenza genome. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00766 (2015).
Rambo-Martin, B. L. et al. Influenza A virus field surveillance at a swine-human interface. mSphere https://doi.org/10.1128/msphere.00822-19 (2020).
Crossley, B. M. et al. Nanopore sequencing as a rapid tool for identification and pathotyping of avian influenza A viruses. J. Vet. Diagn. Invest. 33, 253–260. https://doi.org/10.1177/1040638720984114 (2021).
King, J., Harder, T., Beer, M. & Pohlmann, A. Rapid multiplex minion nanopore sequencing workflow for influenza A viruses. BMC Infect. Dis. 20, 648. https://doi.org/10.1186/s12879-020-05367-y (2020).
de Vries, E. M. et al. Rapid, in-field deployable, avian influenza virus haemagglutinin characterisation tool using minion technology. Sci. Rep. 12, 11886. https://doi.org/10.1038/s41598-022-16048-y (2022).
Keller, M. W. et al. Direct RNA sequencing of the coding complete influenza A virus genome. Sci. Rep. 8, 14408. https://doi.org/10.1038/s41598-018-32615-8 (2018).
Perlas, A. et al. Improvements in RNA and DNA nanopore sequencing allow for rapid genetic characterization of avian influenza. Virus Evol. 11, veaf010. https://doi.org/10.1093/ve/veaf010 (2025).
R. A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2024).
McCormick, K. et al. Construction and immunogenicity evaluation of Recombinant influenza A viruses containing chimeric hemagglutinin genes derived from genetically divergent influenza A H1N1 subtype viruses. PLoS One. 10, e0127649. https://doi.org/10.1371/journal.pone.0127649 (2015).
Golabi, M. et al. Development of reverse transcription Loop-Mediated isothermal amplification assay for rapid and On-Site detection of avian influenza virus. Front. Cell. Infect. Microbiol. 11, 652048. https://doi.org/10.3389/fcimb.2021.652048 (2021).
Acknowledgements
We acknowledge contributions from the following individuals at the Ministry for Primary Industries Animal Health Laboratory (MPI AHL) which made this work possible including sampling collection; Wlodek Stanislawek, Della Orr, Maree Joyce, and many others across DRS. We like to thank Paul Maclean from NZ Institute for Bioeconomy Science for his statistical advice in defining the relationship between average read depth and Cq values. We also acknowledge contributions from the following organisations which made this work possible: International Association of Antarctica Tour Operators (IAATO), Federation University Australia, the United States Antarctic Program and field staff at Palmer Station, MV Argus crew, EVOKN Expedition, Consulting Antarctica/Spirit of Sydney, Ministry for Primary Industries (MPI) NZ and the Antarctic Wildlife Health Network. This work is supported by the NZ Food Safety, Ministry for Primary Industries.
Funding
This project was supported through an internal allocation budget from NZ Food Safety, Ministry for Primary Industries, with no funding from external organisations.
Author information
Authors and Affiliations
Contributions
YSL and MM designed and supervised the whole project. YSL performed all the analysis, technical work and prepared the first draft of the manuscript. YSL, HJ and EG performed RNA extraction, RT-PCR and library preparation for ONT sequencing. YSL, AW, RJ provided bioinformatics support on the characterisation of AIV subtypes. RJ and BC supervised the whole Flui set up and performed analyses of AIV subtype using Flui. YSL and RB performed qPCR analyses on the MIC qPCR system and Kappa statistical tests. JO supervised and provided resources for this research. MD and MM carried out sampling work in Antarctica. MM supervised all experiments, carried out the field test in Antarctica, provided critical input and directed the research. All authors reviewed, edited and provided critical commentary to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Low, Y.S., Jauregui, R., Calcott, B. et al. Development of a portable avian influenza virus characterisation system: bringing the inside-out. Sci Rep 16, 2952 (2026). https://doi.org/10.1038/s41598-025-32771-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32771-8