Abstract
Patients with von Hippel-Lindau (VHL) disease develop multiple distinct kidney cysts and clear cell renal cell carcinomas (ccRCCs) throughout their lifetime. Although cysts are typically considered precursors of ccRCC, no molecular evidence of cysts to tumor progression was provided to date, due to the lack of an accurate molecular characterization of these lesions. We performed a comprehensive molecular characterization of four kidney cysts and six ccRCCs obtained from the same VHL-disease patient, thus all sharing the same genetic germline and host environment. We combined whole genome sequencing (WGS) and RNA sequencing (RNA-seq) profiling with pathological examinations. Cysts and tumors exhibited distinct transcriptomic profiles. Tumors showed increased glycolysis and response to hypoxia, while cysts were associated with enrichment in extracellular matrix organization and inflammation and displayed elevated expression of nephron distal portion markers. Deconvolution analysis further revealed different stromal and immune microenvironments with cysts enriched in myofibroblasts and plasma cells signatures whereas tumors were associated with tumor-associated macrophages (TAMs) and tumor vasculature signatures. Genomically, most cysts and tumors were clonally independent, harboring distinct somatic single-nucleotide variants. Despite sharing some somatic mutational signatures, tumors and cysts exhibited divergent somatic copy number alterations. Only tumors displayed chromosome 3p loss, resulting in VHL loss of heterozygosity, without other driver mutations. VHL patients develop multiple kidney cysts and solid tumors throughout their lifetime and their clinical management is challenging. This study presents the first in-depth molecular analysis of kidney cysts and ccRCC in a single VHL patient. We observed distinct molecular profiles between cysts and tumors, suggesting independent origins. While preliminary, these findings challenge the assumption that cysts always serve as precursors of ccRCC in VHL disease and underscore the need for larger studies to improve surveillance and management of renal lesions in patients with VHL disease.
Similar content being viewed by others
Introduction
Von Hippel-Lindau (VHL) disease is an autosomal dominant hereditary cancer syndrome caused by pathogenic germline variants in VHL tumor suppressor gene, located on chromosome 3p251. It has an estimated prevalence of 1 in 47 000 individuals and an incidence of 1 in 27,0002,3. Affected individuals typically inherit a single mutant VHL allele from an affected parent, which increases their susceptibility to developing multiple benign and/or malignant tumors and cysts in selected organs, including the brain, spinal cord, eyes, adrenal glands, pancreas and the kidneys.
The fact that the whole range of renal lesions, from cysts to tumors, may be present in a single kidney in these individuals, as well as the discovery of microscopic malignant foci in the walls of renal cysts, led to the conclusion that cysts were the precursors of clear cell renal cell carcinomas (ccRCC).
Choyke et al. challenged this hypothesis, conducting a longitudinal study using serial CT scans to assess lesion growth rates and patterns4. Their results demonstrated that the transition from cystic to solid lesions is not the primary mechanism of renal cancer development. Instead, Kragel et al. suggested the possibility that the cell of origin may in part determine the type of lesion that develops5. However, these 1990s studies lacked molecular characterization of the renal lesions and clear evidence of cyst-to-tumor progression. In 2003, Kaelin proposed that VHL disease patients, being VHL+/- heterozygotes, develop renal cysts upon somatic inactivation of the remaining VHL allele, which can then progress to tumors after acquiring additional non-VHL mutations. This is supported by findings that Vhl-inactivated mice develop few kidney cysts but not ccRCC6,7, whereas combined inactivation with ccRCC driver genes (Pbrm1, Bap1) drives spontaneous ccRCC formation8,9,10. Advancements in NGS techniques have enabled a comprehensive characterization of tumors, providing deeper and more precise insights into their molecular underpinnings. However, the molecular features of cysts remain largely unexplored. By integrating WGS, RNA-seq, and pathology, we investigated the molecular characteristics of renal tumors and cysts arising within the same genetic and host background, aiming to elucidate their potential molecular relationship.
Results
Case report
A 56-year-old man with a pathogenic germline mutation in the VHL gene (c.262T > G, p. Trp88Gly) presented at our reference center with the unusual clinical condition of having more than 20 distinct tumoral lesions and 81 cysts in both kidneys (Fig. 1A). Baseline eGFR was 43 and was sustained by the right kidney only, with the left silent at the renal scan. The patient underwent radical nephrectomy of the left kidney and targeted tumor ablations in the right kidney.
Cystic and cancer lesions in the kidney of a von Hippel-Lindau disease patient. (a) 3D model of all kidney cysts (purple) and tumors (orange) in the patient and the numerated tumors harvested and analysed in this study. (b) Macroscopic view of the left kidney after nephrectomy showing the multiple tumors and cysts. Sections are representative of ccRCC wiyhout prominent nucleoli at 100X magnification (WHO/ISUP nucleolar grade 2), eosinophilic cytoplasm and conspicuous nucleoli at 100X magnification (WHO/ISUP nucleolar grade 3) and cystic fluid-filled lesions with cuboidal or flat lining epithelium, sometimes characterized by clear cell changes consistent with simple renal cyst.
Histopathological assessment of tumor lesions and cysts
To assess the cellular and genetic landscape of multiple tumors and cysts within this patient, six independent tumors without necrosis or hemorrhage, four cysts and one piece of adjacent normal renal parenchyma were harvested by a dedicated pathologist. Tumors ranged in size from 0.5 to 5.5 centimeters and were predominantly diagnosed as early stage (pT1a) ccRCC, except for the largest lesion, which was classified as locally advanced (pT3a). Most tumors showed nucleolar grade 2 (G2) apart from one which was assigned a nucleolar grade 3 (G3) (Fig. 1B and Supplemental Table 1). A diagnosis of simple renal cysts was made for the four cystic lesions analyzed (Fig. 1B).
Transcription profiles of CcRRCs and kidney cysts showed different cell identities
To investigate transcriptomic differences between kidney cysts and ccRCCs, we performed bulk RNA-sequencing (RNA-seq). Unsupervised PCA analysis showed that the samples clustered based on their histological characteristics (Fig. 2A).
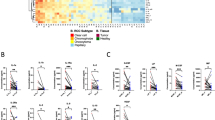
Cystic and cancer lesions show distinct transcriptional patterns. (a) PC1 vs. PC2 score plot produced on the top 5000 most variable genes, on the final high-quality cohort. (b) Illustration of a nephron with the different segment marker genes (c) Volcano plot of tumors versus cysts comparison, with differentially expressed genes (FDR < 0.01) colored in light blue. Marker genes of distal tubules, proximal tubules and ccRCC up and down regulated in tumors versus cysts are indicated in red and green respectively. (d) Representative pathways for cysts or tumor derived from the GO Biological Process. (e) Heatmap of the absolute scores obtained with the deconvolution of the mRNA data with a hierarchical clustering dendrogram.
Differential gene expression analysis (DGE) was performed comparing ccRCCs and cysts (Supplemental Data 1). Cysts expressed markers of Henle’s loop (e.g., KCNJ1, UMOD, SFRP1), distal tubules (e.g., KCNJ1, SLC12A3, SFRP1, AQP2), and collecting duct (e.g., KCNJ1, SLC12A3, SFRP1, AQP2) (Fig. 2B-C, Supplemental Data 1). Tumors, conversely, overexpressed proximal tubule markers (e.g., MIOX, GPX3, NAT8) and SLC17A3, a proximal tubule progenitor-like marker11. Canonical ccRCC markers (e.g., SLC2A1, EGLN3, ESM1, ENPP3, CA9, PFKP, ANGPTL4, VEGFA, HK2, NDUFA4L2) were significantly upregulated in tumors (Fig. 2B-C, Supplemental Data 1) which were also enriched in genes involved in oxygen deprivation response and metabolism (including glycolysis, fatty acid beta-oxidation, lipid and nitrogen compound transport) (Fig. 2D, Supplemental Data 2). Cysts, relative to tumors, showed increased inflammatory response and extracellular matrix organization (Fig. 2D, Supplemental Data 3). RNA-seq deconvolution of the cell type composition revealed an enrichment of gene sets associated with tumor cells, tumor-associated macrophages, and tumor vasculature in tumors. In contrast, cysts showed a higher score of plasma cells and myofibroblasts (Fig. 2E).
Altogether, these findings highlight that cysts and tumors exhibit distinct stromal and myeloid cell compositions and are associated with different nephron segment identities.
WGS depicted independent clonal origins for CcRCC and cysts
WGS was performed with a median coverage of 78X (Supplemental Table 2). Tumors exhibited seven times more single nucleotide variants (SNVs) and indels than cysts (Supplemental Table 3), with around 7800 and 1600 variants, respectively. To evaluate the extent of genomic variability between lesions, we calculated the percentage of unique and shared somatic variants, comparing each sample against all others.
The results revealed that most tumors (L4, L5.1, L7, R10) and cysts (LC1, LC2 and LC4) in this patient were clonally independent, as indicated by their high percentages of unique variants, ranging from 80% to 99% (Fig. 3A). These findings suggest that most tumors and cysts arose independently within this patient. In contrast, tumors L1, L2 and cyst LC3 exhibited significantly lower proportions of unique variants (16%, 26% and 36%, respectively), mutually sharing a total of 2123 variants (accounting for 28%, 24% and 58% of their total variant counts, respectively). This proportion of shared variants provides evidence of a phylogenetic relationship among these lesions, likely attributable to their anatomical proximity.
ccRCCs and cysts have independent clonal origins but share similar mutational signatures. (a) Whole genome variants comparison across all tumors and cysts samples. The percentages of sites unique to each sample is indicated in darker color, the shared proportion in lighter color. (b) SBS-96 activity plot shows the absolute number of mutations in each known COSMIC signature on the y-axis and the sample name on the x-axis. The colours indicate which signature had the mutations and which signatures were found in each sample. (c) SBS-96 activity plot shows the percentage of mutations in each known COSMIC signature on the y-axis and the sample name on the x-axis.
Tumors and cysts show similar mutational signatures profiles
We then characterized the patterns of somatic mutations and the mutational signatures that can provide insights into the underlying mutational processes (Fig. 3B, C). The clock-like signatures SBS512, and SBS1, as well as SBS40b, recently described to be specific of ccRCC 13,14 were present, with similar proportions, in all analyzed samples. SBS40a and SBS8 were instead found only in tumors L5.1 and R10.
Tumors and cysts depict distinct somatic copy number alterations landscapes
To determine if the transcriptional differences observed in the PCA correlated with distinct genomic profiles between tumors and cysts, the somatic copy number alterations (SCNAs) landscape was analyzed. All tumors exhibited an altered SCNA profile with chromosome 3 or 3p loss consistently found as the most frequent alterations, followed by chromosome 5 gain. Loss of chromosome 14 (83% of cases), which has been associated with an increased risk of metastasis15, and gain of chromosomes 12 and 2 (66% and 50%, respectively) were all observed at a considerably higher frequency than that reported in literature16,17 (Fig. 4A). Conversely, most cysts displayed a normal somatic copy number profile, with only LC3 presenting minor alterations, in common with tumors L1 and L2 (Supplementary Fig. 1). These were validated by the IHC anti CAIX that showed that most cysts had negative staining (Supplementary Fig. 2). These observations highlight a significant genomic distinction at the chromosomal copy number level between tumors and cysts, suggesting that these lesions arose independently in the same kidney of this VHL patient.
Tumors show an altered somatic copy number landscape compared to cysts and lack additional driver alterations. (a) Heatmap plot displaying the log2 copy number ratios of tumors and cysts. Each column is a chromosome, each row is a sample. The heatmap represents the consensus variants called with a total of three tools (b) Landscape of driver variants in VHL-related ccRCCs. Heatmap shows the distribution of germline variants in VHL, somatic ccRCC driver variants, and SCNAs.
Lack of canonical somatic CcRCC driver variants in most of the lesions
Additionally, we characterized the landscape of driver variants in these lesions by focusing on canonical ccRCC driver genes. Only tumor L4 harbored a novel pathogenic variant in the BAP1 gene, that was validated by the absence of anti-BAP1 nuclear staining (Supplemental Fig. 3). All other tumors lacked any additional pathogenic or likely pathogenic somatic variants in the canonical ccRCC driver genes (Fig. 4B), only displaying VHL loss of heterozygosity due to chromosome 3p loss. Notably, we could not detect any second hits in VHL gene in cysts, suggesting that most of them have not lost both VHL alleles.
Methods
Patient and samples
The study was approved by the IRCCS San Raffaele Hospital Ethics Committee on 03/03/2014 and registered at ClinicalTrials.gov (NCT06187870) on 28/12/2023. Informed consent was obtained from the patient included in the study. Following radical nephrectomy of the left kidney (with brief hilar clamping to preserve tissue), the right kidney underwent partial nephrectomy and tumor ablation. Cystic tissue samples were collected from the inner surface of the cystic lesions by scraping. In cases where the cyst wall architecture permitted isolation of the parietal layer alone, this component was separately dissected to prevent potential contamination from adjacent normal renal parenchyma or from contiguous neoplastic lesions. All collected tissue samples were washed with calcium/magnesium-free phosphate-buffered saline (PBS), transferred to a pre-chilled, 1.5 mL microcentrifuge tube; excess PBS was removed, and tubes were immediately immersed in liquid nitrogen. For each sample, a piece was snap-frozen samples was transferred to − 80 °C, and another piece was formalin-fixed, paraffin-embedded.
Histology and IHC
ccRCC was diagnosed, staged, and graded per 2022 WHO guidelines (8th TNM, WHO/ISUP nucleolar grade, Fuhrman grade, EAU). Formalin-fixed, paraffin-embedded tissue was sectioned and H&E stained. BAP1 immunohistochemistry was performed using a mouse monoclonal antibody (Santa Cruz Biotechnology, sc-28383, 1:100) on a Leica-Biosystems Bond immunostainer with the Bond Polymer Refine detection system.
RNA-seq
RNA was extracted using miRNeasy (Qiagen) and quality-controlled with NanoDrop and TapeStation 4 100 (RIN ≥ 7). Stranded RNA-seq libraries (Illumina kit) were prepared from 100 ng RNA and sequenced on a NovaSeq 6 000 (100 bp reads, ~ 30 M reads/sample). Data were processed with Trimmomatic (v0.32), aligned to GRCh38 using STAR (v2.5.3a), and quantified with featureCounts (Gencode v31). Quality control was performed with FastQC and MultiQC. Principal Component Analysis (PCA) was performed considering the expression values of the top 5000 most variable genes using the prcomp R package: data were preliminary centered and scaled, to get zero-mean and unit variance on each gene.
Cell types deconvolution
CIBERSORTx was used operating in absolute mode, to deconvolve bulk RNA-seq samples and infer their cell type composition. Cell type expression profiles for deconvolution were derived from single-cell RNA-seq dataset18.
Whole genome sequencing
DNA from 10 ccRCCs (L1-L8, R6/8, R7, R10), 5 cysts (LC1-LC4, RC1), and one blood sample was extracted (Maxwell RSC 48/16, Promega, adapted kit) and quality-controlled (Agilent BioAnalyzer 2100/TapeStation). WGS was performed on six ccRCCs (L1, L2, L4, L5.1, L7, R10), four cysts (LC1-LC4), and blood (Illumina DNA PCR-Free Library Prep, NovaSeq 6 000, 150 × 2 paired-end). Illumina DRAGEN v4.0.3 pipeline was used for alignment and variant calling (GRCh38 refenrence genome). Blood was analyzed with DRAGEN Germline v. 4.0.3 while tumors and cysts with DRAGEN Somatic v. 4.0.3 (tumor/normal pipeline). VCF manipulation and analysis were perfomed with VCFCompare and bcftools.
High-confidence (PASS) SNVs/indels were used for genomic relatedness analysis and signature calling. For driver variant analysis, only non-synonymous variants (coding, splicing, 3’/5’UTR) were considered. Variants were prioritized/classified with eVai enGenome using: (1) in silico gene panel (143 genes from RNA-seq, literature, and TSHC-illumina panel); (2) OMIM association and high computed pathogenicity score (ACMG cl. 4/5); (3) HPO terms (Renal Cell Carcinoma [0005584], ccRCC [0006770]). VCF were then annotated also with ANNOVAR and filtered also for (4) CADD-phred > 25; (5) Phenolyzer disease-specific prioritization. SCNAs were called using DELLY, CNVpytor, and CNVkit. Mutational spectra/signatures were extracted using SigProfiler19.
Statistical analysis
In the RNA-seq analysis, we defined significant genes as those with an adjusted p-value (FDR) of less than 0.01.
All the methods were performed in accordance with the relevant guidelines.
Discussion
Our study offers a unique opportunity to explore tumors and kidney cysts in the same genetic background and host environment.
Our data shows that cysts express loop of Henle, distal tubule and collecting duct markers, whilst tumors express proximal tubule markers. Our findings agree with a previous study in which lectin staining suggests that most simple renal cysts in VHL disease originate from distal tubules, whereas atypical renal cysts and ccRCC originate from proximal tubule cells5. Notably, this is the first study to perform such an in-depth molecular analysis, combining whole-genome sequencing, RNA sequencing, and pathological examination of both cysts and tumors in VHL disease. Previous studies have primarily relied on methods such as loss of heterozygosity (LOH) analysis at the VHL gene locus using polymerase chain reaction single-strand conformation polymorphism20, lectin staining5 and immunohistochemistry for CAIX21 to differentiate between these lesions.
From a transcriptional point of view, we found that tumors showed a proximal tubule identity, consistent with data from scRNA-seq of three sporadic and one hereditary ccRCC showing that tumors most likely originate from a small population of proximal tubule cells11. On the other hand, cysts mostly expressed distal tubules markers, suggesting that kidney cysts and tumors may arise from different nephron segments.
Moreover, in this dataset, kidney cysts and tumors differed at the level of the stromal and immune composition. TAM and tumor vasculature that we found increased in tumors are associated with VHL ccRCC as shown in a scRNA-seq study with two VHL related ccRCC22. Myofibroblasts that were more abundant in the cysts have been described in a single cell analysis of autosomal dominant polycystic kidney disease23. These findings suggest that microenvironment changes may help distinguish cysts from tumors.
In line with published data16,17,24,25, our comparative genomic analysis confirmed the independent clonal origin of tumors, evident from the distinct sets of somatic variants defining each cancer genome. For the first time, this analysis also revealed the independent clonal origin of cysts. At least in this patient, most tumors and cysts appeared to have arisen and evolved as distinct and independent entities. This conclusion is supported by the elevated percentage of unique variants detected in each lesion. Only three lesions (tumors L1 and L2, and cyst LC3) presented a much higher percentage of shared variants. However, given their close anatomical proximity within the kidney, it is likely that these represent a single lesion that appeared macroscopically as three distinct entities.
This conclusion is further supported by the starkly different SCNAs landscapes between tumors and cysts. Only tumors consistently exhibited complete VHL loss of heterozygosity, possibly explaining the difference observed also in hypoxia response and glycolysis between tumors and cysts. Our findings are consistent with Lubensky et al., as they reported that LOH was not uniformly present in all simple cysts they analyzed (detected in 1 case out of 2), whereas it was consistently observed in all the atypical cysts and the tumors20. These results align with our observations, whereby we also detected no VHL LOH in one out of four simple cysts, as opposed to tumors where VHL LOH (through chromosome 3 loss) was always detected.
Interestingly, three somatic mutational signatures SBS40b, SBS1 and SBS5 previously described in ccRCCs13,19,26 were found in both cysts and tumors. These observations are consistent with earlier findings of the same signatures SBS1, 5, and 40 (now split in SBS40a, b and c) 13 in normal kidney cells, which rose with aging27. The presence of signatures found only in ccRCCs might suggest distinct mutational processes may be at play.
Altogether, this comparative analysis revealed that kidney cysts and tumors within this patient displayed distinct nephron segment identities, stromal and immune microenvironment, and genetic alterations, supporting an independent evolution for these lesions.
While we cannot exclude the possibility that kidney cysts may be precursors of tumors, our findings suggest that this is not necessarily the case, given their distinct renal segment of origin and the VHL zygosity. Further studies in additional cases are required to validate and generalize these observations. Nonetheless, these findings may hold important implications for clinical decision-making, particularly regarding the choice between surgical or ablative treatments and active surveillance.
Data availability
RNA-seq data from six ccRCCs, three cysts, and one normal parenchyma sample are available in GEO (GSE254446). WGS data from six ccRCCs, four cysts and one blood DNA sample have been deposited in the European Genome-phenome Archive (EGA) under accession number EGAS50000000295: [https://ega-archive.org/studies/EGAS50000000295] . While the EGA submission is being finalized, a temporary access link to the VCF files is available via the San Raffaele Open Research Data Repository (ORDR) [https://ordr.hsr.it/preview/k3npxv3cw9?a=bd33e184-fba5-4c24-bbe8-6bf1c17102df] . Access to the WGS data can be requested by contacting [pipitone.giovannibattista@hsr.it](mailto: pipitone.giovannibattista@hsr.it) .
References
Maher, E. R. & Kaelin, W. G. von Hippel-Lindau disease. Med. (Baltim). 76, 381–391. https://doi.org/10.1097/00005792-199711000-00001 (1997).
Shepherd, S. T. C., Drake, W. M. & Turajlic, S. The road to systemic therapy in von Hippel-Lindau (VHL) disease: are we there yet? Eur. J. Cancer. 182, 15–22. https://doi.org/10.1016/j.ejca.2022.12.011 (2023).
Larcher, A. et al. Epidemiology of renal cancer: Incidence, Mortality, Survival, genetic Predisposition, and risk factors. Eur. Urol. 88, 341–358. https://doi.org/10.1016/j.eururo.2025.06.005 (2025).
Choyke, P. L. et al. The natural history of renal lesions in von Hippel-Lindau disease: a serial CT study in 28 patients. AJR Am. J. Roentgenol. 159, 1229–1234. https://doi.org/10.2214/ajr.159.6.1442389 (1992).
Kragel, P. J., Walther, M. M., Pestaner, J. P. & Filling-Katz, M. R. Simple renal cysts, atypical renal cysts, and renal cell carcinoma in von Hippel-Lindau disease: a lectin and immunohistochemical study in six patients. Mod. Pathol. 4, 210–214 (1991).
Haase, V. H., Glickman, J. N., Socolovsky, M. & Jaenisch, R. Vascular tumors in livers with targeted inactivation of the von Hippel-Lindau tumor suppressor. Proc. Natl. Acad. Sci. U S A. 98, 1583–1588. https://doi.org/10.1073/pnas.98.4.1583 (2001).
Kleymenova, E. et al. Susceptibility to vascular neoplasms but no increased susceptibility to renal carcinogenesis in Vhl knockout mice. Carcinogenesis 25, 309–315. https://doi.org/10.1093/carcin/bgh017 (2004).
Gu, Y. F. et al. Modeling renal cell carcinoma in mice: Bap1 and Pbrm1 inactivation drive tumor grade. Cancer Discov. 7, 900–917. https://doi.org/10.1158/2159-8290.Cd-17-0292 (2017).
Nargund, A. M. et al. The SWI/SNF protein PBRM1 restrains VHL-Loss-Driven clear cell renal cell carcinoma. Cell. Rep. 18, 2893–2906. https://doi.org/10.1016/j.celrep.2017.02.074 (2017).
Wang, S. S. et al. Bap1 is essential for kidney function and cooperates with Vhl in renal tumorigenesis. Proc. Natl. Acad. Sci. U S A. 111, 16538–16543. https://doi.org/10.1073/pnas.1414789111 (2014).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Sci. (New York N Y). 361, 594–599. https://doi.org/10.1126/science.aat1699 (2018).
Alchahin, A. M. et al. A transcriptional metastatic signature predicts survival in clear cell renal cell carcinoma. Nat. Commun. 13, 5747. https://doi.org/10.1038/s41467-022-33375-w (2022).
Senkin, S. et al. Geographic variation of mutagenic exposures in kidney cancer genomes. Nature https://doi.org/10.1038/s41586-024-07368-2 (2024).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993. https://doi.org/10.1016/j.cell.2012.04.024 (2012).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: tracerx renal. Cell 173, 581–594e512. https://doi.org/10.1016/j.cell.2018.03.057 (2018).
Fei, S. S. et al. Patient-specific factors influence somatic variation patterns in von Hippel-Lindau disease renal tumours. Nat. Commun. 7, 11588. https://doi.org/10.1038/ncomms11588 (2016).
Tabata, M. et al. Inter- and intra-tumor heterogeneity of genetic and immune profiles in inherited renal cell carcinoma. Cell. Rep. 42, 112736. https://doi.org/10.1016/j.celrep.2023.112736 (2023).
Zvirblyte, J. et al. Single-cell transcriptional profiling of clear cell renal cell carcinoma reveals a tumor-associated endothelial tip cell phenotype. Commun. Biology. 7, 780. https://doi.org/10.1038/s42003-024-06478-x (2024).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421. https://doi.org/10.1038/nature12477 (2013).
Lubensky, I. A. et al. Allelic deletions of the VHL gene detected in multiple microscopic clear cell renal lesions in von Hippel-Lindau disease patients. Am. J. Pathol. 149(6), 2089–2094 (1996).
Mandriota, S. J. et al. HIF activation identifies early lesions in VHL kidneys: evidence for site-specific tumor suppressor function in the nephron. Cancer Cell. 1, 459–468. https://doi.org/10.1016/S1535-6108(02)00071-5 (2002).
Jikuya, R. et al. Comparative analyses define differences between BHD-associated renal tumour and sporadic chromophobe renal cell carcinoma. EBioMedicine 92, 104596. https://doi.org/10.1016/j.ebiom.2023.104596 (2023).
Muto, Y. et al. Defining cellular complexity in human autosomal dominant polycystic kidney disease by multimodal single cell analysis. Nat. Commun. 13, 6497. https://doi.org/10.1038/s41467-022-34255-z (2022).
Creighton, C. J. et al. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49. https://doi.org/10.1038/nature12222 (2013).
Turajlic, S. et al. Deterministic Evolutionary Trajectories Influence Primary Tumor Growth: TRACERx Renal. Cell 173, 595–610.e511. https://doi.org/10.1016/j.cell.2018.03.043 (2018).
Culliford, R. et al. Whole genome sequencing refines stratification and therapy of patients with clear cell renal cell carcinoma. Nat. Commun. 15, 5935. https://doi.org/10.1038/s41467-024-49692-1 (2024).
Franco, I. et al. Whole genome DNA sequencing provides an atlas of somatic mutagenesis in healthy human cells and identifies a tumor-prone cell type. Genome Biol. 20, 285. https://doi.org/10.1186/s13059-019-1892-z (2019).
Acknowledgements
We thank the patient enrolled in this study. We acknowledge all the collaborators from the VHL OSR Program. We thank the VHL Alliance which funded this project. F.C. had a co-financed MUR-UniSR fellowship. We thank Alessandra Boletta and Rosa Bernardi for the discussions and advice. We thank the members of the Center for Omics Sciences at the IRCCS Ospedale San Raffaele (COSR), in particular Dejan Lazaveric, Francesca Giannese, Marco Morelli, Lorenza Moronetti for the help in the realization and interpretation of the experiments. We thank Medics3D for providing 3D reconstruction services for Figure 1.
Funding
This study was supported by a research grant from the VHL Alliance.
Author information
Authors and Affiliations
Consortia
Contributions
I.R., A.L., and G.B.P. conceptualized the study. A.L., U.C., A.S., F.M., and F.D.C. contributed patient samples, clinical data, and participated in patient selection and data acquisition. R.L., M.S.A., C.D., N.P.T., and M.P. carried out tissue collection and histopathological/IHC analyses.A.M.F. managed the biobank of specimens. I.R. extracted RNA. G.B.P. and F.C. processed samples for whole-genome sequencing and performed variant calling. G.M.S. conducted RNA-seq bioinformatics analysis. I.R., G.B.P., F.C., I.F., and P.C. performed data analysis and interpretation. I.R. and F.C. wrote the original draft. All authors reviewed, edited, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rowe, I., Corea, F., Pipitone, G.B. et al. Distinct genomic, microenvironmental, and nephron signatures in VHL kidney cysts and tumors. Sci Rep 16, 4394 (2026). https://doi.org/10.1038/s41598-025-34381-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34381-w