Abstract
The raccoon dog (Nyctereutes procyonoides), classified in the order Carnivora within the family Canidae, is native to East Asia and widely distributed throughout Japan due to its adaptability to various environments. Despite the close relationship between raccoon dogs and other animals, viruses infecting raccoon dogs have not been thoroughly investigated in Japan. In this study, we performed metatranscriptomic analyses using fecal samples collected from latrines of wild raccoon dogs in two locations on mainland Japan. Nearly complete viral genomes were identified, including viruses belonging to the genus Kobuvirus (CaKoV), an unclassified canine sapelovirus within the subfamily Ensavirinae (CaSaV), the Genius Mamastrovirus (CaAstV), unclassified hepe-astro-like virus (bastrovirus-like) (Bast-like V), and an unclassified dicistrovirus (DiciV) within the family Dicistroviridae. Phylogenetic analyses revealed that raccoon dog CaKoV, CaSaV, and CaAstV are related to canine strains but form independent clusters specific to raccoon dogs, suggesting they have evolved within this host population. Bast-like V, detected for the first time in raccoon dogs, showed high sequence identity with viruses previously identified in Chinese shrews. The shared insectivorous nature of these hosts and in silico host range predictions suggest that Bast-like Vs may originate from arthropod viruses. Although DiciV is likely of dietary origin due to its arthropod hosts, the large number of sequence reads detected and the phylogenetic clustering of raccoon dog DiciVs with mammalian DiciVs indicate the need to assess their potential infectivity in mammals and the risk of spillover. These findings suggest that raccoon dogs harbor endemic viruses within the canine population and may act as potential vectors for viruses with unknown infectivity in mammals but with spillover risk.
Similar content being viewed by others
Introduction
The raccoon dog (Nyctereutes procyonoides) is a medium-sized carnivore belonging to the order Carnivora, family Canidae. Its native distribution includes East Russia, China, Mongolia, Vietnam, Korean Peninsula and Japan1,2,3. In the middle of the twentieth century, this species was introduced as a fur-bearing animal and has become widespread in northern and eastern Europe, causing problems in local ecosystems and public health4. Globally, they are classified into six subspecies, including N. procyonoides ussuriensis (eastern China and south-eastern Russia), N. procyonoides procyonoides (Indochina and China), N. procyonoides orestes (southwestern China), N. procyonoides koreensis (Korean Peninsula), N. procyonoides viverrinus (Japan, except Hokkaido), and N. procyonoides albus (Hokkaido, Japan)5,6,7. Recently, the Japanese raccoon dog, including the latter two subspecies, has been speculated to be a different species from continental populations through chromosomal, molecular, and morphological studies8,9,10,11,12,13. This species is distributed throughout the country, except on the Tokara, Amami, Okinawa, and Sakishima islands14. Japanese raccoon dogs are opportunistic omnivores with various diets including fruits, vegetables, invertebrates, small vertebrates, carrions, and garbage15,16,17,18,19,20,21,22. This feeding plasticity can also help Japanese raccoon dogs inhabit various habitats, from subalpine mountains16,23 to seashores24,25,26, and from natural forest27,28 to agricultural satoyama region29,30 and urban area31,32. They have a unique defecation behavior using communal latrines, which may serve as olfactory communication to exchange their individual, sexual, reproductive, dietary, and spatial information24,33,34,35.
Although metatranscriptome analysis has facilitated the discovery of RNA viruses, accurately estimating their host range remains crucial for understanding their ecology, evolution, and pathogenicity. Traditional bioinformatics approaches, such as similarity-based methods (e.g., BLAST) and phylogenetic analyses, rely on the availability of genetically close reference sequences in databases. However, these methods often fall short when dealing with novel or highly divergent viruses. To address this gap, sequence composition analysis offers a practical and accessible approach to host prediction. Organisms and their viruses exhibit distinct codon usage biases shaped by evolutionary pressures. For example, bacteriophages often adapt their codon usage to match that of their bacterial hosts closely, likely to enhance replication efficiency within the host’s cellular environment36,37,38. In contrast, viruses infecting eukaryotic hosts may adapt their genome composition to evade immune responses, reflecting a different evolutionary pressure39,40. These host-specific adaptations are reflected in viral genomes and provide a basis for using nucleotide composition in host prediction. Previous studies have demonstrated nucleotide composition for predicting host range, successfully identifying host groups for the picorna-like virus family41,42. This approach is intended as an initial screening step rather than a definitive classification, and the adaptability of this approach depends on the virus. However, narrowing down potential host groups enables researchers to prioritize subsequent analyses, such as phylogenetic inference or experimental validation, facilitating a more efficient workflow for novel virus characterization.
Raccoon dogs harbor various potentially harmful pathogens that can affect animals, including humans. Several viruses, such as rabies virus, Japanese encephalitis virus, influenza A virus, hepatitis E virus, canine distemper virus, and canine parvovirus, have been identified in wild or farmed raccoon dogs43,44,45,46,47,48. However, detailed studies on the viruses carried by raccoon dogs in Japan remain limited, highlighting the need for further research. In this study, we investigated the viruses present in the feces of raccoon dogs by performing metatranscriptomic analyses using fecal samples collected from latrines located in two distinct environments in Japan: a large green area surrounded by urban areas (Site 1) and a small island (Site 2).
Results
Identification of nearly complete viral genomes and confirmation of raccoon dog origin of fecal samples
Because the viral composition in fecal samples may be influenced by the habitat of free-living raccoon dogs, we selected two study areas: a large green space surrounded by urban regions (Site 1) and a small island off a peninsula (Site 2), to investigate raccoon dog fecal viromes. Fecal samples were collected during winter, spring, and fall to represent each season. When an insufficient quantity of fresh feces was obtained in a single survey, additional collection efforts were conducted through repeated visits to latrine sites. Although fecal sampling was also carried out in summer for dietary analysis, these samples were excluded from virome analysis due to the empirically observed low success rate of nucleotide extraction from feces collected during this season. In total, 58 and 61 fecal samples were collected from the latrines of wild raccoon dogs in the large green area surrounded by urban environments (Site 1; 35.562620, 139.490122) between April 9 and November 27, 2023, and from Izushima, an isolated small island off the peninsula (Site 2; 38.456311, 141.524184) between May 5, 2023, and May 2, 2024.
This study focused on vertebrate- and invertebrate-associated viruses recently detected in mammals, with the aim of analyzing the identified viruses in detail. Using a metatranscriptomic approach, RT-PCR, and Sanger sequencing, we identified nearly complete genome sequences of viruses belonging to the genus Kobuvirus and an unclassified genus within the family Picornaviridae, the genus Mamastrovirus within the family Astroviridae, hepe-astroviruses (Bastrovirus, unclassified family), and an unclassified genus within the family Dicistroviridae. These identifications were made using the National Center for Biotechnology Information (NCBI) BLAST program (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Nearly complete genomes of two canine sapeloviruses (CaSaV), one astrovirus (Mamastrovirus canis, CaAstV), and three bastrovirus-like viruses (Bast-like Vs) were identified exclusively in fecal samples from Site 1 raccoon dogs. In contrast, four canine kobuviruses (Kobuvirus aichi, CaKoV) and 11 dicistroviruses (DiciV) were found in samples from both Site 1 and Site 2 (Table 1).
Since fecal samples were collected from latrines, PCR and sequencing of mtDNA were employed to confirm that the samples containing these nearly complete viral genomes originated from raccoon dogs. Based on the nucleotide (nt) sequences of the cytochrome b (Cytb) gene of mitochondrial DNA (mtDNA), all 19 specimens analyzed (seven from Site 1 and 12 from Site 2) were identified as raccoon dogs (Nyctereutes procyonoides viverrinus). Further analysis of partial sequences from the D-loop hypervariable regions of mtDNA revealed that all seven samples from Site 1 belonged to the NproD11 haplotype, whereas the samples from Site 2 consisted of the NproD21 (eight samples) and NproD41 (four samples) haplotypes. These haplotypes were phylogenetically associated with the Japanese raccoon dog (Nyctereutes procyonoides viverrinus) rather than the continental raccoon dog. The results confirmed that the haplotypes differed between samples from Site 1 and those from Site 2, indicating no epidemiological connection between two populations.
Detection of CaKoVs, including a recombinant strain
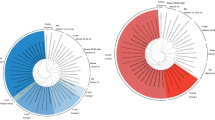
Two partial sequence contigs of the CaKoV genome were detected in raccoon dog samples from both Site 1 and Site 2. Complete genome sequences were determined using RT-PCR with primers designed for full-length genome amplification (Supplementary Table 1) and Sanger sequencing. Initially, these raccoon dog CaKoVs were compared with Japanese CaKoVs; however, only partial 3D sequences from Japanese CaKoVs, including those from canine49 and sewage50 samples, were available. A phylogenetic tree constructed using 390 nt of the 3D region revealed that CaKoVs from Site 1 and Site 2 raccoon dogs clustered separately. Site 1 CaKoVs formed a cluster with CaKoVs from various countries, including China, Brazil, Thailand, South Korea, and Japan, whereas Site 2 CaKoVs clustered with Japanese canine CaKoVs (Fig. 1A). Since the VP1 region is the most variable in picornaviruses and is commonly used for classification, we performed phylogenetic analysis and pairwise sequence identity comparisons using the complete VP1 nt sequences. In the VP1 phylogenetic tree, the Site 1 and Site 2 raccoon dog CaKoVs exhibited significant divergence. However, one of the Site 1 CaKoVs, CaKoV/raccoon dog/Kodo2023-40/2023/JPN (CaKoV_Kodo2023-40), clustered with Site 2 CaKoVs, forming an independent cluster consisting exclusively of raccoon dog-derived CaKoVs. Another Site 1 CaKoV grouped within clusters of CaKoVs from China, Thailand, and Vietnam (Fig. 1B).
Phylogenetic analyses based on partial nucleotide sequences of the 3D region (A) and complete nucleotide sequences of the VP1 region (B) of Japanese raccoon dog CaKoV and kobuvirus strains obtained from the GenBank/EMBL/DDBJ databases. Phylogenetic trees were constructed using the maximum likelihood method in MEGA7 with best-fit models: HKY + G + I model for the 3D region and GTR + G model for the VP1 region. Bootstrap values greater than 70 (1000 replicates) are shown. The scale bars represent corrected genetic distances. CaKoVs from Japanese raccoon dogs and other Japanese CaKoVs, excluding raccoon dog strains, are highlighted in red and green, respectively.
Phylogenetic analyses of the 3D and VP1 nt sequences demonstrated that CaKoV_Kodo2023-40 branched with another Site 1 strain, CaKoV/raccoon dog/2023–5/2023/JPN (CaKoV_Kodo2023-5), in the 3D region, but branched with the Site 2 CaKoVs in the VP1 region. This finding suggests the occurrence of a possible homologous recombination event. Recombination analyses were performed using the sequences of CaKoV_Kodo2023-40, CaKoV_Kodo2023-5, and the Site 2 strain CaKoV/Izus2024-3/2024/JPN (CaKoV_Izus2024-3) with SimPlot v.3.5.151 and the Recombination Detection Program (RDP) v.5.552. A similarity plot analysis revealed that CaKoV_Kodo2023-40 exhibited higher nt similarity to CaKoV_Kodo2023-5 in the 5′ UTR to VP3 and 2A to 3′ UTR regions, while showing lower similarity to CaKoV_Kodo2023-5 than to CaKoV_Izus2024-3 in the VP1 region. These results support the occurrence of a recombination event (Fig. 2A, B). Bootstrap scanning analysis using RDP identified a possible recombination breakpoint near the VP1 region (Fig. 2A, C). CaKoV_Kodo2023-40, CaKoV_Kodo2023-5, and CaKoV_Izus2024-3 were predicted to be the recombinant, major parent, and minor parent, respectively. This recombination event was strongly supported by P-values of 7.515 × 10–22, 3.103 × 10–19, 2.177 × 10–19, 5.842 × 10–18, 2.540 × 10–11, and 2.152 × 10–21 for RDP, BootScan, MaxChi, Chimaera, SiScan, and 3Seq methods, respectively.
(A) Genome structure of CaKoV. (B) Similarity plots of LC849238_CaKoV/Raccoon dog/Kodo2023-5/2023/JPN (red curve) and LC849235_CaKoV/Raccoon dog/Izus2024-3/2024/JPN (blue curve), using LC849239_CaKoV/Raccoon dog/Kodo2023-40/2023/JPN as the query sequence. The analysis was performed with a sliding window of 200 nucleotides and a step size of 20 nucleotides. (C) Recombination analysis of the genome comparing Kodo2023-5/2023 vs Kodo2023-40/2023 (yellow curve), Kodo2023-5/2023 vs Izus2024-3/2024 (blue-green curve), and Kodo2023-40/2023 vs Izus2024-3/2024 (purple curve).
First identification of CaSaV in Japan
CaSaV has primarily been identified in the feces of dogs and red foxes in China, the United Arab Emirates (UAE), and Australia, and more recently in municipal wastewater in the United States. However, CaSaV has not been previously reported in Japan. In this study, two complete CaSaV genomes were detected in raccoon dogs from Site 1. Since Faleye et al. conducted genotyping of CaSaV using the complete capsid region (VP4-VP1)53, a phylogenetic tree was constructed based on the capsid region. The analysis included viruses belonging to the genera Anativirus, Boosepivirus, Diresapivirus, Enterovirus, Felipivirus, Parabovirus, Rabovirus, and Sapelovirus within the subfamily Ensavirinae, as well as a recently reported raccoon dog picornavirus from China54. Phylogenetic analysis revealed that CaSaVs, Rhinolophus picornaviruses, and sapelo-like bat picornaviruses formed a distant branch from other ensaviruses, including the raccoon dog picornavirus, which was more closely related to a felipivirus from Chinese cats. The raccoon dog CaSaV identified in this study was closely related to group C-II, which includes CaSaV strains detected in Chinese dogs and U.S. sewage. Additionally, it formed an independent branch, sharing 79.27–80.17% nt sequence identity and 91.48–91.98% amino acid (aa) sequence identity with the C-II strain (Fig. 3 and Supplementary Table 2).
Phylogenetic analyses based on the complete nucleotide sequences of the P1 region of Japanese raccoon dog CaSaVs and ensavirus strains obtained from the GenBank/EMBL/DDBJ databases. The phylogenetic trees were constructed using the maximum-likelihood method in MEGA7 with best fit models (GTR + G model). Bootstrap values above 70 (1,000 replicates) are indicated. Bars represent corrected genetic distances. CaSaVs from Japanese raccoon dog strains are indicated in red.
Detection of CaAstV closely related to CaAstVs from Korean raccoon dog
Incomplete-length CaAstV sequences were obtained from a single sample collected at Site 1. Sequence-specific primers were designed to amplify the 3’ end of the genome (Supplementary Table 3), and the complete coding sequence (CDS) was determined by RT-PCR and Sanger sequencing. In Japan, only one complete genome sequence for canine CaAstV (Mamastrovirus canis) has been reported55, along with five partial sequences of the ORF1b region detected in sewage56, which are available in the GenBank database. A phylogenetic tree constructed using these short ORF1b sequences revealed that CaAstV/raccoon dog/Kodo2023-51/2023/JPN (CaAstV Kodo2023-51) was distantly related to Japanese CaAstVs detected in sewage. However, it showed a closer relationship to a CaAstV strain from a Japanese dog (LC845131 KU-D4-12) and clustered with European, Chinese, and Korean raccoon dog CaAstVs (Fig. 4A). Since complete ORF2 sequences are required for astrovirus species demarcation57, a phylogenetic analysis based on ORF2 nt sequences was performed. CaAstV Kodo2023-51 was found to be closely related to Korean raccoon dog CaAstVs, exhibiting high sequence identity (95.25–96.55% for nt and 95.85–96.37% for aa). In contrast, it was distantly related to LC845131 KU-D4-12, as well as European and Chinese CaAstVs, with nt and aa sequence identities of 79.08–79.82% and 81.37–81.76%, respectively (Fig. 4B, Table 2).
Phylogenetic analyses based on the complete nucleotide sequences of ORF2 (A) and partial ORF1b (B) of Japanese raccoon dog CaAstV and CaAstV strains obtained from the GenBank/EMBL/DDBJ databases. The phylogenetic trees were constructed using the maximum-likelihood method in MEGA7 with best fit models (GTR + G for ORF2 and T92 + G for ORF1b). Bootstrap values above 70 (1,000 replicates) are indicated. Bars represent corrected genetic distances. CaAstVs from Japanese raccoon dog, raccoon dog CaAstVs from outside Japan, and Japanese CaAstV from dog strains are indicated in red, blue, and green, respectively.
Closely relationship between raccoon dog Bast-like Vs and viruses from Chinese shrew
Through BLAST analysis, contigs showing sequence homology to Bast-like V were identified in the samples from Site 1. This is the first report of Bast-like V detection in raccoon dogs. Initially, these contigs exhibited the highest similarity to a bat Bast-like V (accession no. NC_032426), but the sequence identity was only approximately 50%. To obtain the complete sequence of the raccoon dog Bast-like V, primers were designed based on the sequences of the identified Bast-like V contigs (Supplementary Table 4), and RT-PCR was performed. PCR products covering the Bast-like V genome were successfully amplified from three samples, and full-length CDSs were obtained from two of them through Sanger sequencing. Upon determining the complete raccoon dog Bast-like V sequences, BLAST analysis revealed that their closest homologs were Chinese shrew viruses, with nt sequence identities of 86.22–86.74% and 99–100% coverage. Due to differences in genome organization between Bast-like V, hepeviruses, and astroviruses, as well as the low sequence identities outside the conserved RNA-dependent RNA polymerase (RdRp; ~ 800 nt) and capsid protein (~ 2000 nt) regions, phylogenetic tree analyses and pairwise sequence comparisons were conducted using these conserved regions. The phylogenetic tree constructed using RdRp aa sequences showed that raccoon dog Bast-like Vs were closely related to viruses identified in the lungs of Chinese shrews between 2018 and 202158, forming a well-supported cluster with 100% bootstrap support. These viruses were also related to a virus found in a bird in New Zealand, and together they formed a larger cluster with viruses primarily derived from environmental samples worldwide (Fig. 5A). In the capsid phylogenetic tree, raccoon dog Bast-like Vs also clustered with Chinese shrew viruses; however, the New Zealand bird virus was more distantly related to the raccoon dog and shrew Bast-like Vs (Fig. 5B). Raccoon dog Bast-like Vs shared high nt and aa sequence identities with Chinese shrew strains in the RdRp region (86.69–88.24% and 98.14–98.60%, respectively) and in the capsid region (88.16–90.18% and 92.11–95.53%, respectively) (Table 3).
Phylogenetic analyses based on the amino acid sequences of the capsid (A) and RNA dependent RNA polymerase (RdRp) (B) of Japanese raccoon dog Bast-like Vs and Bast-like V strains obtained from the GenBank/EMBL/DDBJ databases. The phylogenetic trees were constructed using the maximum likelihood method in MEGA7 with best-fit models (LG + G + F for the capsid and LG + G + I model for the RdRp). Bootstrap values above 70 (1,000 replicates) are indicated. Bars represent the corrected genetic distances. Bast-like Vs from Japanese raccoon dog and viruses detected from vertebrates are highlighted in red and green, respectively.
Identification of DiciVs from raccoon dogs in both Site 1 and Site 2 areas
One and ten contigs representing nearly complete genome sequences, including full CDSs, of DiciVs, were identified in raccoon dog samples from Site 1 and Site 2, respectively. Raccoon dog DiciVs were classified into two genotypes, sharing 86.58–86.66% nt sequence identity in ORF1 and 85.98–86.02% in ORF2. DiciVs identified from Site 1 exhibited high similarity to those from Site 2, with 96.97–96.99% nt sequence identity and 98.98–99.05% aa sequence identity (Table 4). Phylogenetic trees based on ORF1 and ORF2 aa sequences revealed that raccoon dog DiciVs were most closely related to squirrel DiciV from the United Kingdom, as well as to DiciVs from red pandas and bats in China. These DiciVs formed a larger cluster predominantly composed of mammalian-host DiciVs (raccoon dogs, squirrels, red pandas, bats, and gorillas) (Fig. 6A, B). Raccoon dog DiciVs shared 52.92–57.50% nt and 47.42–49.66% aa sequence identity in ORF1, and 51.59–52.71% nt and 44.77–45.22% aa sequence identity for ORF2 with squirrel, red panda, and bat DiciVs. In contrast, they exhibited lower identity with insect-associated viruses in the genera Aparavirus, Cripavirus, and Triatovirus (Table 4), with 32.76–36.11% nt and 16.58–17.71% aa sequence identity in ORF1, and 29.04–33.51% nt and 14.18–20.27% aa sequence identity in ORF2.
Phylogenetic analyses based on the amino acid sequences of the ORF1 (A) and ORF2 (B) of Japanese raccoon dog DiciVs and DiciV strains obtained from the GenBank/EMBL/DDBJ databases. The phylogenetic trees were constructed using the maximum-likelihood method in MEGA7 with best fit models (LG + G + I). Bootstrap values above 70 (1,000 replicates) are indicated. Bars represent corrected genetic distances. DiciVs from Japanese raccoon dog and DiciVs detected from vertebrates are indicated in red and green, respectively.
Frequency of CaKoV, CaSaV, CaAstV, Bast-like V, and DiciV in fecal samples
To determine the frequency of CaKoV, CaSaV, CaAstV, Bast-like V, and DiciV in raccoon dog fecal samples, RT-PCR was performed using virus-specific primer pairs (Supplementary Table 5). RT-PCR analysis revealed that 12 (20.7%) and 15 (25.9%) out of 58 samples from Site 1 were positive for CaSaV and Bast-like V, respectively, while no positive samples for these viruses were found at Site 2. CaKoV, CaAstV, and DiciV were detected in 5 (8.6%), 25 (43.1%), and 15 (25.9%) of 58 samples from Site 1, and in 9 (14.8%), 1 (1.6%), and 36 (59.0%) of 61 samples from Site 2, respectively (Table 1). It is important to note that, since these fecal samples were collected from communal latrines, the results may not accurately reflect prevalence rates, as some samples may originate from the same individuals. The potential for overlapping samples differs between Site 1 and Site 2. At Site 1, the bait marking method conducted at the site distinguished individuals into four groups (our unpublished data). Assuming each group consisted of a pair of individuals, at least eight individuals were present. In contrast, at Site 2, analysis of individually identified fecal DNA by Kubo et al. estimated an average of 60 raccoon dogs (21.7–24.3 animals/km2) on the island59. Furthermore, the infection status of the same individuals may vary depending on the sampling periods, and samples from the same individuals could be considered independent. To estimate the proportion of overlapping samples, we conducted a simulation under the assumption that fecal samples were randomly selected. For Site 1, we estimated overlap when one to 38 fecal samples were collected from 8 individuals, while for Site 2, we estimated overlap when 3 to 20 fecal samples were collected from 60 individuals. Using 10,000 iterations of resampling, we calculated the proportion of unique samples and their confidence intervals with the statistical software R. The results indicate that an estimated 79% of the samples from Site 1 and 14% of the samples from Site 2 were duplicates at most. Based on these estimates, the effective sample size, adjusted for unique individual, was calculated to be 75.7 (n = 119) (Supplementary Table 6).
In silico host range prediction
Given that the Bast-like Vs detected in raccoon dogs and shrews clustered with Bast-like Vs from various hosts in the phylogenetic analysis, and considering that DiciVs are typically associated with arthropods, an in silico host range prediction was conducted using established methods41,42. For this analysis, we utilized a sequence dataset in which the infected hosts were reported by Orf et al.42. The results showed that all Bast-like Vs from raccoon dogs and Chinese shrews clustered within the invertebrate group (Fig. 7A). Similarly, raccoon dog DiciVs, along with DiciVs from squirrels, red pandas, gorillas, bats, and birds, fell within the 90% confidence ellipse for the invertebrate group (Fig. 7B).
Canonical score plot of discriminant analysis for Bast-like Vs (A) and DiciVs (B) used to classify viral sequences into host groups by using 4 mononucleotide and 16 dinucleotide frequencies. The graph illustrates the separation of groups using the two most influential factors. Viruses infecting invertebrates are represented by purple circles, those infecting plants by green circles and those infecting vertebrates by red circles. The lines indicate the 90% confidence interval.
Discussion
It has been proposed that the Japanese raccoon dog is genetically and morphologically distinct from the continental raccoon dog12,13. Japanese raccoon dogs have been geographically isolated from their continental counterparts by the sea separating Japan from the mainland. In continental raccoon dogs, including farmed animals, viruses that infect animals have been reported in countries such as China and South Korea. In Japan, aside from Japanese encephalitis virus, distemper virus, and group A rotavirus36,39,60,61,62, virome data from Japanese raccoon dogs are scarce. In this study, no viruses potentially harmful to humans or livestock were detected. However, two picornaviruses and one astrovirus associated with dogs were identified. Picornaviruses are non-enveloped, small icosahedral viruses with positive-sense RNA genomes. The family Picornaviridae currently comprises 68 genera and 159 species (as of March 2024: https://www.picornaviridae.com); however, many viruses remain unclassified. CaKoV belongs to the genus Kobuvirus, while CaSaV, though part of the subfamily Ensavirinae, is not classified within any genus.
CaKoVs have been identified in domestic dogs across Europe, the Americas, and Asia, including Japan, and are suspected to cause gastroenteritis in some dog populations63. Although no diarrheal samples were analyzed in this study, CaKoV was detected in 8.6% and 14.8% of samples from Site 1 and Site 2, respectively. Phylogenetic analysis based on partial sequences of the 3D region, a conserved region in RNA viruses, revealed that the CaKoV strains from Sites 1 and 2 branched separately, although both were homologous to CaKoVs from Japanese dogs and sewage. In contrast, in the VP1 tree, a highly variable region, one CaKoV from Site 1 and all strains from Site 2 branched distinctly from other CaKoVs, forming a single cluster. This suggests that these strains may have evolved within the Japanese raccoon dog population. Furthermore, the CaKoV Kodo2023-40 strain from Site 1 appeared to be a recombinant strain, likely involving recombination between viruses from Sites 1 and Site 2 near the VP1 region. Given that these sites are approximately 400 km apart, direct contact between raccoon dogs is unlikely. However, the presence of CaKoV strains at both sites suggests a potential for recombination. Mixed infections in communal latrines or contaminated feeding sites might facilitate such recombination events.
Another picornavirus, CaSaV, has been detected in the feces, urine, and respiratory swabs of dogs and red foxes in China64,65,66, the UAE65, and Australia67, and more recently in U.S. wastewater53. However, its association with disease remains inconclusive63. This study reports the first detection of CaSaV in Japanese raccoon dogs. The picornaviruses detected in raccoon dogs in China were distantly related to CaSaVs, branching separately among ensaviruses, suggesting they could represent a new genus within the subfamily Ensavirinae. Japanese CaSaVs were related to group C-II, which includes Chinese CaSaVs from dogs and U.S. sewage, but with P1 nt sequence identities of 79.27–80.17%, they formed an independent branch. This suggests that CaSaV is evolving in the raccoon dog population at Site 1.
Astroviruses are small, icosahedral, non-enveloped RNA viruses with a star-like surface structure, transmitted primarily via the fecal–oral route57,68. The astrovirus identified in this study belongs to the species Mamastrovirus canis (CaAstV), which is primarily found in dogs, although other astroviruses have also been detected in canine species69. In raccoons, as in dogs, CaAstV and other astroviruses, including neurotropic astroviruses belonging to the HMO clade and unclassified astroviruses, have been identified70,71,72. The CaAstV detected in this study was distantly related to CaAstVs found in Japanese sewage but closely related to CaAstVs from dogs and Korean raccoon dogs. Phylogenetic analysis of a short sequence (216 nt) in the ORF1b region limited further comparative analysis. However, the full ORF2 tree revealed that Japanese raccoon dog CaAstV clustered with Korean raccoon dog CaAstVs, suggesting a shared common ancestor and parallel evolution in both regions. The mechanism of virus introduction into these geographically distant regions remains unclear. Given the limited data on canine fecal viruses in Japan, further investigations into virus prevalence in dogs and other carnivores are needed. Since RNA viruses mutate rapidly, monitoring potential changes in infectivity and virulence is essential, especially in the raccoon dog population.
Unclassified hepe-astrovirus-like RNA viruses, such as bastroviruses and Bast-like Vs, have been detected in various host species and environmental samples. Bastroviruses have been detected in vertebrates, including humans73, pigs74,75, rats, and bats76. Bast-like Vs have been found in both vertebrates and invertebrates, including humans42, shrews58, bats, birds77, insects78,79, shellfish80,81, and sewage82, as well as in other environmental samples83,84,85,86. In this study, Bast-like Vs from raccoon dogs were closely related to Bast-like Vs from Chinese shrews58, showing high aa identity in RdRp region (98.14–98.60%) and capsid region (92.11–95.79%). The next closest species, based on RdRp aa identity, was a bird Bast-like V from New Zealand77, with 60.0% identity. However, the capsid region showed extremely low homology (7.47–7.68%), suggesting that the bird Bast-like V might be recombinant. Bast-like Vs have been detected in a variety of samples including insects. Since shrews are insectivorous and raccoon dogs are known to consume insects22,28, it is possible that raccoon dogs acquired Bast-like Vs by feeding on insects carrying these viruses or by consuming insectivorous animals such as birds, which inhabit Site 117,30. Bast-like Vs have also been detected in the lungs of shrews, suggesting that the virus may enter the respiratory tract through inhalation or be absorbed via the gastrointestinal tract before subsequently reaching the lungs. However, whether the virus replicates in these organisms remains unclear. Orf et al. detected Bast-like Vs in human sera in Africa using a metagenomic approach. Although virus isolation in cultured cell lines was unsuccessful, in silico evaluation estimated the potential host range of the virus42. Their analysis did not classify the human Bast-like V as one that infects vertebrates. Similarly, in our analysis, Bast-like Vs from raccoon dogs and shrews were also classified into the invertebrate group. Raccoon dog and shrew Bast-like Vs were notably diverse compared to other Bast-like Vs, with aa identities of less than 46.51% (except for the New Zealand bird Bast-like V) in the RdRp region and less than 33.72% in the capsid region. These findings indicate that the raccoon dog and shrew Bast-like Vs form an independent clade. Even if invertebrates were the original hosts of these viruses, the possibility of spillover into vertebrates raises concerns about potential cross-species infection risk. Further investigations into the infectivity of these viruses in mammals and their host range are warranted.
DiciVs are small, non-enveloped viruses with non-segmented RNA genomes approximately 8–10 kilobases in length. All known members of this virus family infect arthropods87. However, DiciVs have recently been detected in human serum and stool samples88,89,90,91,92, as well as in the blood and liver of bats93,94, and in fecal samples from squirrels95, wild gorillas96, red pandas97, and bats98,99. The pathogenicity of DiciVs in mammals remains unexplored. In this study, phylogenetic analysis revealed that DiciVs from raccoon dogs formed a distinct cluster with DiciVs from other mammals, suggesting a shared common ancestor. Despite the geographical separation of the two study sites, DiciV strains from raccoon dogs at Site 1 and Site 2 exhibited high sequence homology, indicating that these strains may be widely distributed across mainland Japan. Although DiciVs are generally considered arthropod-infecting viruses, it is hypothesized that raccoon dogs may acquire them through their diet. However, the widespread detection of DiciVs in raccoon dogs and the high proportion of DiciV-specific reads in individuals with nearly complete DiciV genome sequences (ranging from 0.18 to 23.72% of total reads; Supplementary Table 7) suggest that direct infection may also be possible. While in silico host range evaluation classified DiciVs from raccoon dogs and other mammals into the invertebrate group, the close phylogenetic clustering with mammalian-associated DiciVs raises the possibility of direct mammalian infection. Orf et al. performed an in silico analysis to estimate the zoonotic potential of DiciVs, suggesting that these viruses may have the capacity to infect humans42. These findings underscore the need for further research into the potential infectivity of DiciVs in mammals. Even the most closely related DiciVs, such as those from squirrels, exhibit low aa sequence homology with raccoon dog DiciVs in the ORF1 and ORF2 regions (49.34–49.66% and 44.77–44.88%, respectively). This suggests that DiciVs from different hosts may vary significantly in key virological properties, including host range, pathogenicity, antigenicity, and replication efficiency. Further studies are required to fully understand the epidemiological and pathogenic potential of DiciVs in mammals.
The in silico analysis using nucleotide composition for host estimation has several limitations. This approach is not universally applicable, as different viruses are subject to varying evolutionary pressures. Moreover, the accuracy of this method relies on the availability of unbiased reference data to minimize false estimations. In this study, Linear Discriminant Analysis (LDA) was employed as a preliminary screening tool rather than a definitive classification method. Viruses plotted near the boundary of the discriminant space may be more challenging to classify, resulting in reduced discriminability. Therefore, complementary approaches, such as phylogenetic analysis or experimental validation, are essential for accurately estimating viral hosts.
In summary, this study revealed that raccoon dogs harbor dog-associated viruses, including CaKoV, CaSaV, and CaAstV. Phylogenetic and pairwise identity analyses suggest that these viruses may be maintained and evolving within the raccoon dog population. Additionally, Bast-like Vs and DiciVs, which have recently been suspected of infecting mammals, were identified. Although these viruses were initially believed to be acquired through dietary intake, their widespread presence in raccoon dogs and sequence homology to mammalian-associated strains suggest the possibility of direct infection. This study represents the first metatranscriptomic analysis of raccoon dog feces in Japan. However, further research is required to elucidate the role of raccoon dogs in the ecology and transmission dynamics of these viruses.
Materials and methods
Sample collection
We collected raccoon dog fecal samples from two sites: Site 1 in the Kanto region and Site 2 in the Tohoku region. Site 1 is a relatively large green area (1 km2) surrounded by urban areas spanning Yokohama City, Kanagawa Prefecture, and Machida City, Tokyo, and is used as an amusement park. We collected 10 fecal samples in September and October 2023, and 10 in November 2023 from 10 different latrines of raccoon dogs. Site 2 is a small island (2.63 km2) off the peninsula of Onagawa Town, Miyagi Prefecture, where medium-sized carnivores such as red foxes (Vulpes vulpes), Japanese martens (Martes melampus), Japanese weasels (Mustela itatsi), palm civets (Paguma larvata), and feral cats (Felis catus), in addition to raccoon dogs have been confirmed22. From this location, we collected 15, 20, 22, and 3 fecal samples from up to 16 raccoon dog latrine sites in November 2023 and February, April, and May 2024, respectively. The collected fecal samples were placed in plastic bags at each site, transported to the laboratory, and frozen at − 20 °C until analysis. For processing, the samples were diluted 1:9 (w/v) with sterile phosphate-buffered saline (PBS) and centrifuged at 12,000 × g for 10 min. The resulting supernatants were stored at − 20 °C until use for RNA extraction.
Determination of animal species
DNA was extracted from fecal samples using ISOFECAL (Nippon Gene, Tokyo, Japan) according to the manufacturer’s protocol. To identify the animal species, a primer pair (Fw: 5'-TGAGGACAAATATCMTTYTGAGG-3' and Rv: 5'-GGGTGGAATGGRATTTTRTC-3') which amplified a part of Cytb gene was newly designed using various known mammalian mt DNA Cytb gene sequences. PCR was carried out using TaKaRa Ex Taq® HS DNA polymerase (TaKaRa Bio, Shiga, Japan) under the following conditions: initial denaturation at 95 °C for 3 min, 35 cycles of heat denaturation at 95 °C for 20 s, annealing at 50 °C for 20 s, and extension at 72 °C for 30 s, followed by a final extension at 72 °C for 3 min. The amplified product, with a target size of 263 bp, was confirmed by electrophoresis, and sequenced directly using the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Carlsbad, CA, USA) on the ABI PRISM 3130 Genetic Analyzer (Applied Biosystems). The sequenced product (220 bp, excluding the primer regions) was analyzed using GENETYX-MAC 21.0.1 software (Nihon Server Corporation, Tokyo, Japan). Species identification was conducted by homology searches against international DNA databases (GenBank/EMBL/DDBJ) using NCBI’s BLAST program. In addition, a new primer set (Npro tRNAproF19: 5'-GCACCCAAAGCTGAAATTCTTC-3' and Npro DloopR469: 5'-GGGCTGATTAGTCATTAGTCCA-3') was designed to amplify the mtDNA D-loop hypervariable region. Step-down PCR was performed using TaKaRa Ex Taq® HS DNA polymerase (TaKaRa Bio) under the following conditions: initial denaturation at 95 °C for 3 min, two cycles of denaturation at 95 °C for 20 s, annealing at 59 °C for 20 s, and extension at 72 °C for 30 s, two cycles of denaturation at 95 °C for 20 s, annealing at 57 °C for 20 s, and extension at 72 °C for 30 s, and then 30 cycles of denaturation at 95 °C for 20 s, annealing at 55 °C for 20 s, and extension at 72 °C for 30 s, followed by a final extension at 72 °C for 3 min. The amplified product, with a target size of approximately 450 bp, was confirmed by electrophoresis, and sequenced directly using the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) on the ABI PRISM 3130 Genetic Analyzer. The sequenced product (392 bp, excluding the primer regions), was analyzed using GENETYX-MAC 21.0.1 software. Haplotypes were determined by homology searches of the sequences against GenBank/EMBL/DDBJ (accession numbers AB607907-AB607934, Npro01-28) and undeposited haplotypes Npro29-41.
RNA extraction from fecal samples, cDNA libraries preparation, and deep sequencing
Total RNA was extracted from the samples using TRIzol LS (Life Technologies, Carlsbad, CA, USA), followed by DNase I treatment (Takara Bio). cDNA libraries were constructed for deep sequencing using the NEBNext Ultra II RNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, MA, USA) in accordance with the manufacturer’s guidelines. Library quantities were assessed using a Qubit® 4.0 Fluorometer (Invitrogen, Carlsbad, CA, USA), and paired-end reads of 151 nucleotides were generated using a MiniSeq benchtop sequencer (Illumina, San Diego, CA, USA), as previously described100.
RT-PCR
Total RNA was extracted from the supernatants of 10% fecal samples using TRIzol LS Reagent (Life Technologies) to investigation the frequency of CaKoV, CaSaV, CaAstV, Bast-like V, and DiciV and to determine the complete genomes of CaKoV, CaAstV, and Bast-like V using primers designed for each virus (Supplementary Tables 1 and 2). For both frequency investigation and complete genome determination, PrimeScript™ RT Master Mix (Takara Bio) and TaKaRa Ex Premier™ DNA Polymerase Dye plus (Takara Bio) were used for RT and PCR, respectively. RT was performed at 42 °C for 15 min using the reverse primer, TX30SXN101. PCR conditions generally consisted of an initial denaturation at 94 °C for 1 min, 35 cycles of denaturation at 98 °C for 10 s, annealing at 55 °C 15 s, and extension at 68 °C for 30 s. The extension time was adjusted according to the size of the amplification product. RT-PCR products were resolved by electrophoresis on a 2% agarose gel.
Genome analysis
The FASTQ-formatted sequence data were imported into CLC Genomics Workbench 24.0.2 (CLC bio, Aarhus, Denmark). The sequences were trimmed, and low-quality sequences were omitted. The processed sequence data were then assembled into contigs using the de novo assembly command with default setting in the CLC Genomics Workbench. To identify vertebrate-associated virus sequences, the resulting contigs were analyzed using the BLAST programme. Genome sequences of CaKoV, CaSaV, CaAstV, Bast-like V, and DiciV were aligned with corresponding viral sequences obtained from the GenBank/EMBL/DDBJ database using ClustalW102. Pairwise sequence identity calculations for each gene segment were performed using the CLC Genomics Workbench. Phylogenetic analyses were constructed using nt or aa sequences for all segments, employing the maximum-likelihood method with the best-fit evolutionary models. The models used were as follows: HKY + G + I for the kobuvirus 3D region, HKY + G for the kobuvirus VP1 region, GTR + G + I for the CaSaV P1 region, T92 + G for the CaAstV ORF2 region, GTR + G + I for the CaAstV ORF2 region, LG + G + I for the Bast-like V RdRp and DiciV ORF1 and ORF2 regions, and LG + G + F for the Bast-like V capsid region. Phylogenetic trees were constructed in MEGA7103 and evaluated with 1000 bootstrap replicates104. Recombination analysis was performed using SimPlot42 and RDP543.
Host range prediction analysis
The nucleotide composition of viral genomes was analyzed, and LDA was applied to associate these compositions with specific host groups using a Python script. The training dataset consisted of 945 full-length RefSeq-quality viral genome sequences, primarily from + ssRNA viruses within the phylum Pisuviricota, including classes such as Pisoniviricetes and Stelpaviricetes42. The test dataset included newly identified viral genomes from Bast-like V and DiciV. First, mononucleotide and dinucleotide frequencies were calculated using a custom nucleotide-counting function for each dataset. The frequencies were then normalized to account for variations in sequence length. Next, the normalized frequency data were used as input for LDA, implemented using scikit-learn’s `LinearDiscriminantAnalysis` class105. Using annotated host information from the training dataset, LDA was employed to infer associations between nucleotide compositions and viral host ranges. The LDA results were visualized using Matplotlib106 and Plotly107, providing interactive plots that display the distribution of viruses in the LDA space, colored according to their host ranges. Confidence ellipses, representing the 90% confidence intervals for each host range group, were also included in the plots. The test dataset, consisting of Bast-like V and DiciV genome sequences, was subsequently analyzed using the trained LDA model. The mononucleotide and dinucleotide frequencies of these new viral genome sequences were projected onto the LDA space to compare their nucleotide compositions with those of known viruses. This approach enabled potential host range inference for these newly identified viruses. The script generated separate plots for the test data, allowing for visual comparison with the training data.
Data availability
The GenBank/EMBL/DDBJ accession numbers for the sequences of the raccoon dog virus strains determined in this study are LC849228 to LC849253. Other datasets generated or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Ward, O.G. & Wurster-Hill, D.H. Nyctereutes procyonoides. Mamm. Species 358, 1–5. https://www.science.smith.edu/departments/Biology/VHAYSSEN/msi/pdf/i0076-3519-358-01-0001.pdf (1990).
Kauhala, K. & Saeki, M. Nyctereutes procyonoides. The IUCN Red List of Threatened Species 2016: e.T14925A85658776. https://doi.org/10.2305/IUCN.UK.2016-1.RLTS.T14925A85658776.en (2016).
Van, P. et al. Southern extension of raccoon dog Nyctereutes procyonoides (Mammalia: Carnivora: Canidae) range in Vietnam with comments on its conservation status in the country. Eur J Wildl Res. 69(2), 22. https://doi.org/10.1007/s10344-023-01653-7 (2023).
Kauhala, K. & Kowalczyk, R. Invasion of the raccoon dog Nyctereutes procyonoides in Europe: History of colonization, features behind its success, and threats to native fauna. Curr. Zool. 57, 584–598. https://doi.org/10.1093/czoolo/57.5.584 (2011).
Ellerman, J. R. & Morrison-Scott, T. C. S. Checklist of Palearctic and Indian mammals, British museum (Natural History), 1758–1946. https://biostor.org/reference/127693 (1951).
Wada, M. Y., Lim, Y. & Wurster-Hill, D. H. Banded karyotype of a wild-caught male Korean raccoon dog, Nyctereutes procyonoides koreensis. Genome 34, 302–306. https://doi.org/10.1139/g91-049 (1991).
Kauhala, K. & Saeki, M. Finnish and Japanese raccoon dogs - on the road to speciation? In Biology and Conservation of Wild Canids (eds Macdonald, D. W. & Sillero-Zubiri, C.) 217–226 (Oxford University Press, 2004).
Wada, M. Y. & Imai, T. On the Robertsonian polymorphism found in the Japanese raccoon dog (Nyctereutes procyonoides viverrinus). Jpn. J. Genet. 66(1), 1–11. https://doi.org/10.1266/jjg.66.1 (1991).
Ward, O. G., Wurster-Hill, D. H., Ratty, F. J. & Song, Y. Comparative cytogenetics of Chinese and Japanese raccoon dogs, Nyctereutes procyonoides. Cytogenet. Genome Res. 45(3–4), 177–186. https://doi.org/10.1159/000132451 (1987).
Nie, W., Wang, J., Perelman, P., Graphodatsky, A. S. & Yang, F. Comparative chromosome painting defines the karyotypic relationships among the domestic dog, Chinese raccoon dog and Japanese raccoon dog. Chromosome Res. 11, 735–740. https://doi.org/10.1023/B:CHRO.0000005760.03266.29 (2003).
Kim, S. I. et al. Phylogeography of Korean raccoon dogs: implications of peripheral isolation of a forest mammal in East Asia. J. Zool. 290, 225–235. https://doi.org/10.1111/jzo.12031 (2013).
Kim, S. I., Oshida, T., Lee, H., Min, M. S. & Kimura, J. Evolutionary and biogeographical implications of variation in skull morphology of raccoon dogs (Nyctereutes procyonoides, Mammalia: Carnivora). Biol. J. Linn. Soc. 116, 856–872. https://doi.org/10.1111/bij.12629 (2015).
Hong, Y. et al. Genetic diversity and population structure of East Asian raccoon dog (Ncteryeutes procyonoides): Genetic features in central and marginal populations. Zool. Sci. 35, 249–259. https://doi.org/10.2108/zs170140 (2018).
Saeki, M. Nyctereutes procyonoides Gray. In The wild mammals of Japan 2nd edn (eds Ohdachi, D. et al.) 224–225 (Shoukadoh, D, 2015).
Sasaki, H. & Kawabata, M. Food habits of the raccoon dog Nyctereutes procyonoides viverrinus in a mountainous area of Japan. J. Mamm. Soc. Japan 19, 1–8. https://doi.org/10.11238/jmammsocjapan.19.1 (1994).
Yamamoto, Y. Comparative analyses on food habits of Japanese marten, red fox, badger and raccoon dog in Mt Nyugasa, Nagano Prefecture, Japan. Nat. Environ. Sci. Res. 7, 45–52 (1994).
Sako, T., Kawada, S., Teduka, M., Uesugi, T. & Akihito,. Seasonal food habits of the raccoon dog, Nyctereutes procyonoides, in the Imperial Palace, Tokyo. Bull. Nat. Museum Nat. Sci. Series A (Zool.) 34, 63–75 (2008).
Osaki, A. et al. Comparison of feeding habits and habitat use between invasive raccoons and native raccoon dogs in Hokkaido Japan. BMC Ecol. 19, 35. https://doi.org/10.1186/s12898-019-0249-5 (2019).
Inagaki, A. et al. Vertebrate scavenger guild composition and utilization of carrion in an East Asian temperate forest. Eco. Evo. 10, 1223–1232. https://doi.org/10.1002/ece3.5976 (2020).
Takatsuki, S., Inaba, M., Hashigoe, K. & Matsui, H. Opportunistic food habits of the raccoon dog—a case study on Suwazaki Peninsula, Shikoku, Western Japan. Mamm. Stud. 46, 25–32. https://doi.org/10.3106/ms2020-0061 (2020).
Takatsuki, S. & Kobayashi, K. Seasonal changes in the diet of urban raccoon dogs in Saitama, eastern Japan. Mamm. Stud. 48, 203–213. https://doi.org/10.3106/ms2023-0001 (2023).
Nagasaki, K. et al. A comparison of summer insectivory among four sympatric mesocarnivores on Izushima, a small island in northern Japan. Mammalia 87, 110–121. https://doi.org/10.1515/mammalia-2021-0160 (2022).
Seki, Y. & Koganezawa, M. Factors influencing winter home ranges and activity patterns of raccoon dogs Nyctereutes procyonoides in a high-altitude area of Japan. Acta. Theriologica 56, 171–177. https://doi.org/10.1007/s13364-010-0020-y (2011).
Ikeda, H. Raccoon dog scent marking by scats and its significance in social behaviour. J. Ethol. 2, 77–84. https://doi.org/10.1007/BF02430571 (1984).
Takatsuki, S., Iwata, M., Hiraizumi, H. & Hirabuki, Y. Food habits of raccoon dogs on the Sendai coast of northern Japan: Two populations that returned after the 2011 Tohoku-oki tsunami. Japanese J. Conserv. Ecol. 23, 155–165. https://doi.org/10.18960/hozen.23.1_155 (2018).
Tsukada, H., Abe, K., Takatsuki, S. & Minami, M. How many feces should be sampled from latrines? Spatial sampling biases affecting the dietary analysis of island raccoon dogs. Wildl. Res. 2, 1–9. https://doi.org/10.2981/wlb.00656 (2020).
Okabe, F. & Agetsuma, N. Habitat use by introduced raccoons and native raccoon dogs in a deciduous forest of Japan. J. Mammal. 88, 1090–1097. https://doi.org/10.1644/06-MAMM-A-117R2.1 (2007).
Koike, S., Morimoto, H., Goto, Y., Kozakai, C. & Yamazaki, Y. Insectivory by five sympatric carnivores in cool-temperate deciduous forests. Mamm. Stud. 37, 73–83. https://doi.org/10.3106/041.037.0208 (2012).
Saeki, M., Johnson, P. J. & Macdonald, D. W. Movements and habitat selection of raccoon dogs (Nyctereutes procyonoides) in a mosaic landscape. J. Mammal. 88, 1098–1111. https://doi.org/10.1644/06-MAMM-A-208R1.1 (2007).
Xu, J., Suzuki, K., Kanda, T., Newman, C. & Kaneko, Y. Invasive raccoons (Procyon lotor) have little effect on the food habits of native raccoon dogs (Nyctereutes procyonoides) in a satoyama area of Tokyo. Mamm. Study https://doi.org/10.3106/ms2022-0041 (2023).
Saito, M. & Koike, F. Distribution of wild mammal assemblages along an urban–rural–forest landscape gradient in warm-temperate east Asia. PLoS One 8(5), e65464. https://doi.org/10.1371/journal.pone.0065464 (2013).
Saito, M. & Koike, F. Trait-dependent changes in assemblages of mid-sized and large mammals along an Asian urban gradient. Acta. Oecologica 67, 34–39. https://doi.org/10.1016/j.actao.2015.06.002 (2015).
Yamamoto, I. Latrine utilization and feces recognition in the raccoon dog. Nyctereutes procyonoides. J. Ethol. 2, 47–54. https://doi.org/10.1007/BF02348206 (1984).
Koizumi, R. et al. Individual interaction at latrine sites of raccoon dogs in the Akasaka Imperial Grounds, central Tokyo. J. Field Science. 15, 7–13 (2017).
Watanabe, K., Kumagai, N. & Saito, M. U. Latrine site selection of raccoon dogs in a hilly area in north-eastern Japan. Zool. Ecol. 31, 79–85. https://doi.org/10.35513/21658005.2021.2.2 (2021).
Tian, L., Shen, X., Murphy, R. W. & Shen, Y. The adaptation of codon usage of+ ssRNA viruses to their hosts. Infect, Genet, Evol. 63, 175–179. https://doi.org/10.1016/j.meegid.2018.05.034 (2018).
Gaunt, E. R. & Digard, P. Compositional biases in RNA viruses: Causes, consequences and applications. Interdiscip, Rev, RNA. 13(2), e1679. https://doi.org/10.1002/wrna.1679 (2022).
Simón, D., Cristina, J. & Musto, H. Nucleotide composition and codon usage across viruses and their respective hosts. Front. Microbiol. 12, 646300. https://doi.org/10.3389/fmicb.2021.646300 (2021).
Mordstein, C. et al. Transcription, mRNA export, and immune evasion shape the codon usage of viruses. Genome, Biol, Evol. 13(9), evab106. https://doi.org/10.1093/gbe/evab106 (2021).
Roy, A. et al. Base composition and host adaptation of the SARS-CoV-2: Insight from the codon usage perspective. Front. Microbiol. 12(548275), 2021. https://doi.org/10.3389/fmicb.2021.548275 (2021).
Kapoor, A., Simmonds, P., Lipkin, W. I., Zaidi, S. & Delwart, E. Use of nucleotide composition analysis to infer hosts for three novel picorna-like viruses. J. Virol. 84, 10322–10328. https://doi.org/10.1128/jvi.00601-10 (2010).
Orf, G. S. et al. Metagenomic detection of divergent insect- and bat-associated viruses in plasma from two African individuals enrolled in blood-borne surveillance. Viruses 15(4), 1022. https://doi.org/10.3390/v15041022 (2023).
Ohno, Y. et al. Detection of antibodies against Japanese encephalitis virus in raccoons, raccoon dogs and wild boars in Japan. J. Vet. Med. Sci. 71, 1035–1039. https://doi.org/10.1292/jvms.71.1035 (2009).
Qi, X. et al. Molecular characterization of highly pathogenic H5N1 avian influenza A viruses isolated from raccoon dogs in China. PLoS. One. 4(3), e4682. https://doi.org/10.1371/journal.pone.0004682 (2009).
Liu, Y. et al. Fox- and raccoon-dog-associated rabies outbreaks in northern China. Virol. Sin. 29(5), 308–310. https://doi.org/10.1007/s12250-014-3484-0 (2014).
Suzuki, J. et al. Canine distemper virus infection among wildlife before and after the epidemic. J. Vet. Med. Sci. 77, 1457–1463. https://doi.org/10.1292/jvms.15-0237 (2015).
Zhao, et al. Isolation and genetic characterization of parvoviruses from dogs, cats, minks, and raccoon dogs in the eastern region of Shandong Province China. Front. Microbiol. 13, 862352. https://doi.org/10.3389/fmicb.2022.862352 (2022).
Tian, F. et al. First molecular evidence of hepatitis E virus in farmed raccoon dogs. Emerg. Microbes. Infect. 13(1), 2361025. https://doi.org/10.1080/22221751.2024.2361025 (2024).
Soma, T., Matsubayashi, M. & Sasai, K. Detection of kobuvirus RNA in Japanese domestic dogs. J. Vet. Med. Sci. 78, 1731–1735. https://doi.org/10.1292/jvms.16-0217 (2016).
Yamashita, T. et al. Molecular detection and nucleotide sequence analysis of a new Aichi virus closely related to canine kobuvirus in sewage samples. J. Med. Microbiol. 63, 715–720. https://doi.org/10.1099/jmm.0.070987-0 (2014).
Lole, K. S. et al. Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J. Virol. 73, 152–160. https://doi.org/10.1128/JVI.73.1.152-160.1999 (1999).
Martin, D. P. et al. RDP5: A computer program for analyzing recombination in, and removing signals of recombination from, nucleotide sequence datasets. Virus Evol. 7(1), 1087. https://doi.org/10.1093/ve/veaa087 (2020).
Faleye, T. O. C. et al. Detection of human, porcine and canine picornaviruses in municipal sewage sludge using pan-enterovirus amplicon-based long-read Illumina sequencing. Emerg. Microbes Infect. 11, 1339–1342. https://doi.org/10.1080/22221751.2022.2071173 (2022).
He, W. T. et al. Virome characterization of game animals in China reveals a spectrum of emerging pathogens. Cell 185, 1117–1129. https://doi.org/10.1016/j.cell.2022.02.014 (2022).
Takano, T., Takashina, M., Doki, T. & Hohdatsu, T. Detection of canine astrovirus in dogs with diarrhea in Japan. Arch. Virol. 160, 1549–1553. https://doi.org/10.1007/s00705-015-2405-3 (2015).
Hoque, S. A. et al. Genotype diversity of enteric viruses in wastewater amid the COVID-19 pandemic. Food. Environ. Virol. 15, 176–191. https://doi.org/10.1007/s12560-023-09553-4 (2023).
Guix, S., Bosch, A. & Pintó, R. Astrovirus taxonomy. In Astrovirus research essential ideas, everyday impacts, future directions (ed. Schultz-Cherry, S.) 97–118 (Springer, 2013).
Zhang, D. T. et al. Decoding the RNA viromes in shrew lungs along the eastern coast of China. NPJ. Biofilms. Microbiomes. 10(1), 68. https://doi.org/10.1038/s41522-024-00543-3 (2024).
Kubo, K. et al. Density estimation for an island population of raccoon dogs in Japan using fecal DNA. Wildlife, Biol. 2023(5), e01112. https://doi.org/10.1002/wlb3.01112 (2023).
Ohashi, K. et al. Properties of a new CDV isolate from a raccoon dog (Nyctereutes procyonoides viverrinus) in Japan. Vet. Rec. 148, 148–150. https://doi.org/10.1136/vr.148.5.148 (2001).
Abe, M. et al. Molecular characterization of rotaviruses in a Japanese raccoon dog (Nyctereutes procyonoides) and a masked palm civet (Paguma larvata) in Japan. Vet. Microbiol. 146, 253–259. https://doi.org/10.1016/j.vetmic.2010.05.019 (2010).
Kameo, Y. et al. Epizootic canine distemper virus infection among wild mammals. Vet. Microbiol. 154, 222–229. https://doi.org/10.1016/j.vetmic.2011.07.006 (2012).
Caddy, S. L. New viruses associated with canine gastroenteritis. Vet. J. 232, 57–64. https://doi.org/10.1016/j.tvjl.2017.12.009 (2018).
Woo, P. C. et al. Complete genome sequence of a novel picornavirus, canine picornavirus, discovered in dogs. J. Virol. 86, 3402–3403. https://doi.org/10.1128/JVI.07228-11 (2012).
Woo, P. C. Y. et al. Molecular epidemiology of canine picornavirus in Hong Kong and Dubai and proposal of a novel genus in Picornaviridae. Infect. Genet. Evol. 41, 191–200 (2016).
Li, C. et al. Genomic characterization and phylogenetic analysis of a new canine picornavirus variant in the mainland of China. Virus. Res. 296, 198351. https://doi.org/10.1016/j.virusres.2021.198351 (2021).
Campbell, S. J. et al. Red fox viromes in urban and rural landscapes. Virus. Evol. 6, veaa065. https://doi.org/10.1093/ve/veaa065 (2020).
Donato, C. & Vijaykrishna, D. The broad host range and genetic diversity of mammalian and avian astroviruses. Viruses 9(5), 102. https://doi.org/10.3390/v9050102 (2017).
Mihalov-Kovács, E. et al. Genome analysis of canine astroviruses reveals genetic heterogeneity and suggests possible inter-species transmission. Virus. Res. 232, 162–170. https://doi.org/10.1016/j.virusres.2016.12.005 (2017).
Yang, S. et al. Viral metagenomics reveals diverse viruses in the feces samples of raccoon dogs. Front. Vet. Sci. 8, 693564. https://doi.org/10.3389/fvets.2021.693564 (2021).
Chae, S. B., Jeong, C. G., Park, J. S., Na, E. J. & Oem, J. K. Detection and genetic characterization of astroviruses in brain tissues of wild raccoon dogs. Viruses 15(7), 1488. https://doi.org/10.3390/v15071488 (2023).
Zhao, J. et al. Farmed fur animals harbour viruses with zoonotic spillover potential. Nature https://doi.org/10.1038/s41586-024-07901-3 (2024).
Oude Munnink, B. B. et al. A novel astrovirus-like RNA virus detected in human stool. Virus. Evol. 2(1), vew005. https://doi.org/10.1093/ve/vew005 (2016).
Bauermann, F. V. et al. Identification and genetic characterization of a porcine hepe-astrovirus (bastrovirus) in the United States. Arch. Virol. 164, 2321–2326. https://doi.org/10.1007/s00705-019-04313-x (2019).
Nagai, M. et al. Metagenomic identification, sequencing, and genome analysis of porcine hepe-astroviruses (bastroviruses) in porcine feces in Japan. Infect. Genet. Evol. 88, 104664. https://doi.org/10.1016/j.meegid.2020.104664 (2021).
Yinda, C. K. et al. Cameroonian fruit bats harbor divergent viruses, including rotavirus H, bastroviruses, and picobirnaviruses using an alternative genetic code. Virus. Evol. 4(1), vey008. https://doi.org/10.1093/ve/vey008 (2018).
French, R. K., Filion, A., Niebuhr, C. N. & Holmes, E. C. Metatranscriptomic comparison of viromes in endemic and introduced passerines in New Zealand. Viruses 14(7), 1364. https://doi.org/10.3390/v14071364 (2022).
Sanborn, M. A. et al. Metagenomic analysis reveals Culex mosquito virome diversityand Japanese encephalitis genotype V in the Republic of Korea. Mol. Ecol. 30, 5470–5487. https://doi.org/10.1111/mec.16133 (2021).
Remnant, E. J. et al. A diverse viral community from predatory wasps in their native and invaded range, with a new virus infectious to honey bees. Viruses 13(8), 1431. https://doi.org/10.3390/v13081431 (2021).
Bonny, P. et al. Human and animal RNA virus diversity detected by metagenomics in Cameroonian Clams. Front. Microbiol. 12, 770385. https://doi.org/10.3389/fmicb.2021.770385 (2021).
Richard, J. C., Blevins, E., Dunn, C. D., Leis, E. M. & Goldberg, T. L. Viruses of freshwater mussels during mass mortality events in Oregon and Washington. USA. Viruses. 15(8), 1719. https://doi.org/10.3390/v15081719 (2023).
Dos Anjos, K., Nagata, T. & Melo, F. L. Complete genome sequence of a novel bastrovirus isolated from raw sewage. Genome. Announc. 5(40), e01010–e01017. https://doi.org/10.1128/genomeA.01010-17 (2017).
Chen, Y. M. et al. RNA viromes from terrestrial sites across China expand environmental viral diversity. Nat. Microbiol. 7(8), 1312–1323. https://doi.org/10.1038/s41564-022-01180-2 (2022).
Sadiq, S. et al. Australian terrestrial environments harbour extensive RNA virus diversity. Virology. 593, 110007. https://doi.org/10.1016/j.virol.2024.110007 (2024).
Zell, R., Groth, M., Selinka, L. & Selinka, H.-C. Hepeliviruses in two waterbodies in Berlin. Germany. Arch. Virol. 168(1), 9. https://doi.org/10.1007/s00705-022-05688-0 (2022).
French, R., Charon, J., Lay, C. L., Muller, C. & Holmes, E. C. Human land use impacts viral diversity and abundance in a New Zealand river. Virus. Evol. 8(1), veac032. https://doi.org/10.1093/ve/veac032 (2022).
Bonning, B. C. & Miller, W. A. Dicistroviruses. Annu. Rev. Entomol. 55, 129–150. https://doi.org/10.1146/annurev-ento-112408-085457 (2010).
Phan, T. G. et al. Sera of Peruvians with fever of unknown origins include viral nucleic acids from non-vertebrate hosts. Virus. Genes. 54, 33–40. https://doi.org/10.1007/s11262-017-1514-3 (2018).
Cordey, S. et al. Detection of dicistroviruses RNA in blood of febrile Tanzanian children. Emerg Microbes Infect. 8, 613–623. https://doi.org/10.1080/22221751.2019.1603791 (2019).
Oguzie, J. U. et al. Metagenomic surveillance uncovers diverse and novel viral taxa in febrile patients from Nigeria. Nat. Commun. 14(1), 4693. https://doi.org/10.1038/s41467-023-40247-4 (2023).
Victria, J. G. et al. Metagenomic analyses of viruses in stool samples from children with acute flaccid paralysis. J. Virol. 83, 4642–4651. https://doi.org/10.1128/JVI.02301-08 (2009).
Holtz, L. R. et al. Geographic variation in the eukaryotic virome of human diarrhea. Virology 468–470, 556–564. https://doi.org/10.1016/j.virol.2014.09.012 (2014).
Bennett, A. J. et al. Diverse RNA viruses of arthropod origin in the blood of fruit bats suggest a link between bat and arthropod viromes. Virology 528, 64–72. https://doi.org/10.1016/j.virol.2018.12.009 (2019).
Fumagalli, M. J. et al. Krykféie dicistrovirus: a novel dicistrovirus in velvety free-tailed bats from Brazil. Infect. Genet. Evol. 75, 104036. https://doi.org/10.1016/j.meegid.2019.104036 (2019).
Dastjerdi, A., Everest, D. J., Davies, H., Denk, D. & Zell, R. A novel dicistrovirus in a captive red squirrel (Sciurus vulgaris). J. Gen. Virol 102(3), 001555. https://doi.org/10.1099/jgv.0.001555 (2021).
Duraisamy, R. et al. Detection of novel RNA viruses from free-living gorillas, Republic of the Congo: Genetic diversity of picobirnaviruses. Virus. Genes. 54, 256–271. https://doi.org/10.1007/s11262-018-1543-6 (2018).
Zhao, M. et al. Viral metagenomics unveiled extensive communications of viruses within giant pandas and their associated organisms in the same ecosystem. Sci. Total Environ. 820, 153317. https://doi.org/10.1016/j.scitotenv.2022.153317 (2022).
Reuter, G., Pankovics, P., Gyöngyi, Z., Delwart, E. & Boros, Á. Novel dicistrovirus from bat guano. Arch. Virol. 159(3453–3456), 2014. https://doi.org/10.1007/s00705-014-2212-2 (2014).
Yinda, C. K. et al. Highly diverse population of Picornaviridae and other members of the Picornavirales, in Cameroonian fruit bats. BMC genomics 18, 1–14. https://doi.org/10.1186/s12864-017-3632-7 (2017).
Nagai, M. et al. Full genome analysis of bovine astrovirus from fecal samples of cattle in Japan: Identification of possible interspecies transmission of bovine astrovirus. Arch. Virol 160, 2491–2501. https://doi.org/10.1007/s00705-015-2543-7 (2015).
Oka, T., Doan, Y. H., Shimoike, T., Haga, K. & Takizawa, T. First complete genome sequences of genogroup V, genotype 3 porcine sapoviruses: Common 5’-terminal genomic feature of sapoviruses. Virus Genes 53, 848–855. https://doi.org/10.1007/s11262-017-1481-8 (2017).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic. Acids. Res. 25, 4876–4882. https://doi.org/10.1093/nar/25.24.4876 (1997).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. https://doi.org/10.1093/molbev/msw054 (2016).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x (1985).
Pedregosa, F. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 12, 2825–2830. https://doi.org/10.5555/1953048.2078195 (2011).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95. https://doi.org/10.1109/MCSE.2007.55 (2007).
Plotly Technologies Inc. Title: Collaborative data science Publisher: Plotly Technologies Inc. Place of publication: Montréal, QC Date of publication: 2015 URL: https://plot.ly (2015).
Acknowledgements
This work was supported by Center for Human and Animal Symbiosis Science, Azabu University, and JSPS KAKENHI (Grant No. 21K05947).
Author information
Authors and Affiliations
Contributions
M.O., T.M., H.T. and M.N. conceived the experiments, M.O.,N.T., T.Y., H.T., M.T., F.S., M.N., S.M., H.I., H.M. and T.T. conducted the experiments, M.O., S.S., H.T. and M.N. analyzed the results. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The present study was carried out according to the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology of Japan. This study is reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Oba, M., Sakaguchi, S., Teshima, N. et al. Metatranscriptomic identification of novel RNA viruses from raccoon dog (Nyctereutes procyonoides) feces in Japan. Sci Rep 15, 7100 (2025). https://doi.org/10.1038/s41598-025-90474-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-90474-6