Abstract
Colorectal cancer (CRC) is a leading cause of cancer-related deaths globally. Adenomas, precursors to CRC, can be diagnosed early, but the genetic events leading to adenoma–adenocarcinoma conversion remain unclear. This study explored the role of chromosomal instabilities (CINs) in this conversion. Over a 17-year follow-up period, 119 adenomas were analyzed using low-coverage whole-genome sequencing (LC-WGS) and Ultrasensitive Chromosomal Aneuploidy Detector. Risk factors for adenocarcinoma development were identified through logistic regression analysis, and survival was assessed using Kaplan–Meier curves. CIN was found in 32% of adenomas, with a higher incidence in high-grade adenomas (P = 0.0359). Common chromosomal changes included loss of 18q, 1p, and 17p and gain of 8q (MYC), 20q, and 7p (EGFR). During the 17-year follow-up, 88 patients experienced recurrence, including 40 cases of adenomas and 48 cases of progression to adenocarcinoma. CIN was identified in 40% of progression cases, 33.6% of adenoma recurrence cases, and 26% of nonrecurrent cases. A strong genetic linkage was observed before and after tumor transformation, with a high match between the tumors and matched prior adenomas. CIN was significantly associated with disease progression (HR: 2.5, 95% CI: 1.4–4.5, P = 0.00162) and was an independent risk factor. Additionally, MFN2 gene copy number deletion was linked to recurrence and/or progression after resection, with reduced expression in tumor tissues. In conclusion, CIN is a key risk factor for adenoma recurrence and progression, and MFN2 gene copy number deletion is associated with adverse outcomes, providing insights for more accurate clinical prognostication of adenoma-to-adenocarcinoma transformation.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is a multifactorial, heterogeneous disease with a high mortality rate globally, with more than 900,000 deaths from CRC each year1,2. Previous studies have shown that colorectal carcinogenesis and progression are closely associated with the malignant transformation of colorectal polyps (adenomas), and a total of 60–90% of sporadic CRCs develop through the classical adenoma–adenocarcinoma pathway3. Therefore, in CRC screening, it is crucial to determine whether adenoma would progress to adenocarcinoma4. Traditionally, colorectal adenomas have been divided into two categories, namely low-risk and high-risk, based on their number, size, growth pattern, and degree of dysplasia, but this classification does not fully meet clinical expectations5.
Chromosomal instability (CIN) is characterized by chromosomal translocations, allelic imbalances, and aneuploidy6. In solid tumors, CIN is usually found as a cellular copy-number aberration (CNA) accompanied by local amplification of oncogenes or deletion of oncogenes, and thus CIN is widely recognized as a marker of cancer6,7. In a previous study, chromosomal analysis of colorectal adenomas, CRCs, and normal colon tissue samples using low-coverage whole-genome sequencing (LC-WGS) revealed that increased gene copy number was strongly associated with adenoma-to-cancer progression and that POFUT1 and Notch signaling were potential drivers of colorectal tumor development8.
Mitochondria are the primary source of reactive oxygen species (ROS) generation in most mammalian cells, typically at elevated levels in cancer cells. They serve as crucial signaling molecules that regulate autophagy and tumor development9,10. Among these, mitochondrial fusion protein 2 (MFN2) is located on the outer mitochondrial membrane (OMM) and facilitates mitochondrial fusion, thereby influencing ROS generation11. Recent studies have shown that MFN2 is often under expressed in various tumors, including CRC12, bladder cancer13, ovarian cancer14 and other malignancies. Conversely, overexpression of MFN2 can have an anti-carcinogenic role. For instance, MFN2 inhibited cancer progression by suppressing the mTORC2/Akt signaling pathway15. Consequently, MFN2 expression is closely linked to cancer progression. However, it remains unclear whether the absence of MFN2 contributes to the transformation of adenomas into adenocarcinomas.
In this work, we conducted a follow-up of 119 patients with adenomas for up to 17 years by analyzing the patients’ primary and recurrent paired samples by LC-WGS, a technique that allows for the analysis of genome-wide chromosomal instability without the need to have prior knowledge of tumor mutations in the tissues16,17. The aims of the study were as follows: (1) to assess whether CIN is a significant risk factor for recurrence and/or progression of adenomas after initial resection; (2) to identify key genes with copy-number alterations (CNAs) that are critically associated with the progression to adenocarcinoma.
Materials and methods
Ethical statement
This study was approved by the Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (20220305-33). Samples were obtained from the Department of Pathology, Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, and maintained by the research team. These specimens were collected after obtaining informed consent from the participants. All experiments using clinical samples were conducted in accordance with the approved guidelines. This study was conducted in accordance with the Declaration of Helsinki.
Study subjects
We retrospectively enrolled 119 patients who had undergone sigmoidoscopy or colonoscopy and had colorectal adenomatous polyps detected for the first time at the Department of Pathology at Sir Run Run Shaw Hospital Zhejiang University School of Medicine from May 1994 to December 2021. All detected polyps were excised and centrally reviewed against the pathological criteria. Patients with inflammatory bowel disease or known hereditary syndromes (i.e., familial adenomatous polyposis and hereditary nonpolyposis CRC) were excluded.
The patients were followed-up with for up to 17 years. A total of 88 of these patients experienced recurrence, including 48 cases of progression to adenocarcinoma and 40 cases of recurrence of adenoma.
Histological analysis
Adenoma or adenocarcinoma tissues obtained by surgical resection were fixed in 10% formalin buffer and embedded in paraffin. After sectioning, 4-µm-thick sections were stained using hematoxylin and eosin (H&E, Sigma-Adrich, St. Louis, MO, USA). The sections were randomly selected from each tissue sample. We used a light microscope (Olympus, Tokyo, Japan) to examine each section.
DNA extraction
Genomic DNA was extracted from the adenoma or adenocarcinoma tissue samples using the Qiagen QIAamp DNA FFPE Tissue Kit (Qiagen, Courtaboeuf, France) in accordance with the manufacturer’s instructions. Each DNA sample was quantified by nano-spectrophotometry (NanoView, GE Healthcare, Orsay, France) and characterized by 0.8% agarose electrophoresis for identification.
Low-coverage whole-genome sequencing (LC-WGS)
Regarding the operation method of LC-WGS, we refer to previous literature18. In brief, libraries were prepared using Kapa Hyper Prep kits (Roche, CA, USA) and customized adapters (IDT, CA, USA) with starting DNA inputs ranging from 50 to 1,000 ng (median 471 ng), which were used for LC-WGS. The libraries were pooled and sequenced on a HiSeq ×10 (Illumina) using 150-bp double-ended runs sequenced on 1× lanes. Fragment copy numbers were derived using an Ultrasensitive Chromosomal Aneuploidy Detector (UCAD) workflow. If the median absolute deviation of copy ratios (log ratios) between neighboring bins across the genome was greater than 0.38, indicating poor-quality sequence data, the sample was excluded.
Copy-number analysis and CIN score calculation
The cyclic binary segmentation (CBS) algorithm in the R package DNACopy was applied to find the meaningful genomic breakpoints and genomic segments with copy-number changes and analyze the copy-number changes. At least 10 million paired reads were collected for each sample. These reads were localized to the human reference genome hg19. The average coverage of each 200k dataset of the genome was then calculated using the software Samtools mpileup. The average coverage of all datasets was normalized to obtain a Zscore, which was used to assess individual chromosome stability. Copy-number alterations (CNAs) were defined as segments with Z-scores > 3 (amplification) or < − 3 (deletion). A genome-wide CIN score was calculated based on the weighted sum of absolute Z-scores across all segments. The Zscore for each bin was calculated as follows:
where Ctest and Ccontrol are the coverage of the bin.
The degree of variation of the whole genome compared with the human reference genome was summarized as the CIN score, which was used to assess the stability of all chromosomes.
The CIN score was calculated as follows:
where V is the Zscore value of a segment, and L is the length of the segment in base pairs.
The above formula and method refer to Hongnan Mo’s study9. Based on the genomic copy number variations in the resected adenoma tissue samples, the patients were categorized into the positive group (n = 38), which had CIN, and the negative group (n = 81), which was chromosomally stable.
Development of chromosomal variant genes based on LASSO Cox regression
Based on the recurrence status, we divided the follow-up samples into progressed-to-adenocarcinoma group and not-progressed-to-adenocarcinoma group. Chromosomal variant data at the gene level were taken separately, first analyzed by univariate Cox to obtain 2,033 genes as risk factors, and then downscaled by LASSO to finally identify that the MFN2 gene might be associated with adenocarcinoma recurrence.
TCGA data collection
A total of 504 copy number variations (CNVs) and expression data were retrieved from the TCGA database (https://portal.gdc.cancer.gov/) with clinical information from the colon adenocarcinoma (COAD) samples. After filtration, 211 samples remained with information on disease-free interval (DFI).
Statistical analysis
All statistical analyses were performed using R (v.3.6.3). The Wilcoxon rank-sum test was used to analyze the relationship between clinicopathological characteristics and MFN2 expression. The Kaplan–Meier method was used to calculate the DFI for adenocarcinoma recurrence after resection based on TCGA. Univariate logistic regression analysis was used to assess the association of clinical features and CIN with adenocarcinoma recurrence. P values less than 0.05 were considered statistically significant.
Results
Patients’ characteristics
In this study, we recruited 119 patients with adenomas from Sir Run Run Shaw Hospital Zhejiang University School of Medicine. All of the patients were treated with adenoma resection surgery. The clinical and pathological characteristics of the patients are detailed in Table 1. The age of the patients was 65.2 ± 10.6 years in the positive group and 57.2 ± 12.7 years in the negative group (P = 0.0071). In addition, a higher percentage of patients in the positive group had adenomas of high grade (18%) compared with the negative group (4.9%) (P < 0.0359). This finding implies that CIN occurs more frequently in high-grade adenomas.
Adenomas with CIN May develop into adenocarcinomas
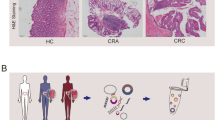
The 119 patients with adenocarcinoma were studied retrospectively. Of these, 88 patients were followed-up with. Forty-eight of them progressed to adenocarcinoma (Fig. 1). Among them, 15 patients (15/38, 39.5%) progressed to adenocarcinoma in the CIN-positive group, and 6 patients (6/81, 7.4%) progressed to adenocarcinoma in the CIN-negative group (Table S1).
Flowchart of study design and patient enrollment.
Genomic copy-number changes were analyzed in the excised tissues from the 88 patients who returned to hospital for treatment with recurrence. Patient 1 was diagnosed with a high-grade adenoma of the rectum in 2005. However, the tumor recurred within 7 years and was diagnosed as well-differentiated adenocarcinoma (Fig. 2A–B). Patient 2 had a low-grade adenoma in the colon, which unfortunately progressed to moderately-to-highly differentiated adenocarcinoma within 2 years (Fig. 2C–D).
Histopathological sections of cases that progressed to adenocarcinoma after adenoma removal surgery. Case 1: (A) The first adenoma removal sample in 2005. Diagnosis of villous tubular adenocarcinoma with high-grade intraepithelial neoplasia was confirmed. (B) The rectal resection specimen 7 years later presented bulging, tubular villous adenoma with carcinoma. Diagnosis of highly differentiated adenocarcinoma, infiltrating into the submucosal layer (immediately adjacent to the muscularis propria), with no lymph node metastasis (0/13) and negative margins, was made. Case 2: (C) The first adenoma resection sample in 2019. The diagnosis was villous tubular adenoma (sigmoid colon) with low-grade intraepithelial neoplasia. (D) Sample resected after recurrence 2 years later. Diagnosis of highly differentiated adenocarcinoma of the colon with reactive hyperplasia of the lymph nodes was confirmed.
Notably, CIN was present in the first resection and recurrent resection tissues of both patients, and the focal CNVs at the lesion site before and after recurrence were consistent. Representative chromosome CNA included 8q gain, 17p loss, 17q gain, and 20q gain in patient 1 (Fig. 3A–B), while 6q loss, 7q gain, 8q loss, 13q gain, 14q gain, 16q gain, and 17p loss were found in patient 2 (Fig. 3C–D). These results indicate that benign adenomas and recurrence-forming malignant adenocarcinomas might originate from the same cell. In addition, we observed that chromosome 17 amplification was significantly elevated in patient 1 seven years after the removal of the adenoma. Patient 2 exhibited a more pronounced downregulation of chromosomes 6 and 9 seven years post-adenoma removal. Alterations in CNA at these sites may be critical in driving the transition from adenoma to adenocarcinoma.
Copy-number plots of tumors and matched prior adenomas from two recurrent patients, where the y-axis represents copy number (normalized log2 ratio) and the x-axis represents chromosomes. (A) Sample of case 1 with first adenoma resection. (B) Sample of case 1 with progression-to-adenocarcinoma resection. (C) Sample of case 2 with first adenoma resection. (D) Sample of case 2 with progression-to-adenocarcinoma resection.
CIN as a risk factor for adenocarcinogenesis
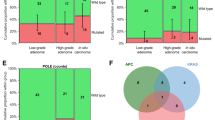
Since high CIN features may be closely associated with recurrent adenocarcinoma, we sought to determine the genomic evidence of CIN in patients with adenocarcinogenesis. As shown in Fig. 4; Table 2, the patients who progressed from adenomas to CIN adenocarcinomas (group 1 and group 2) were found to have frequent alterations in gene copy number. Frequent chr7+ (17.11% of primary adenomas, 22.92% of recurrent adenocarcinomas) and chr8q+ (40.79% of primary adenomas, 50.00% of recurrent adenocarcinomas) gains were found in these patients, where oncogenes EGFR (chr7+) and MYC (chr8q+) were located. In contrast, chr17p- (10.53% of primary adenomas, 41.67% of recurrent adenocarcinomas) was found to be frequently absent in the patients with recurrent adenocarcinomas, and this locus corresponds to the anticancer gene TP53. However, the frequency of genetic alterations was lower in the patients who did not progress to cancer at the follow-up (group 4 and group 5). These results suggest that CIN is frequent in patients whose adenomas have progressed to adenocarcinoma.
Heat map of chromosome copy-number changes in a recurrent sample from an adenoma patient after resection. The yellow color indicates copy-number gain, and blue indicates copy-number loss. Group 1 represents the presence of CIN in the first resected adenoma sample and the presence of CIN in the recurrent adenocarcinoma sample. Group 2 represents the absence of CIN in the first resected adenoma and the presence of CIN in the recurrent adenocarcinoma. Group 3 represents the absence of CIN in the first resected adenoma and the absence of CIN in the recurrent adenocarcinoma (which was not detected in this study). Group 4 represents the presence of CIN in the first resection of the adenoma and the absence of carcinoma in the recurrence. Group 5 represents the first resection of the adenoma without CIN and no carcinoma in the recurrence.
Subsequently, we conducted univariate logistic regression analysis on chromosomal alterations and clinical indicators to explore the risk factors for adenocarcinogenesis. The results showed that CIN was significantly associated with adenocarcinogenesis (HR: 2.5, 95% CI: 1.4–4.5, P = 0.00162) (Fig. 5A). Furthermore, the logistic regression models were adjusted for age, sex, BMI, and adenoma grade. Multicollinearity was assessed using variance inflation factor (VIF), with all variables showing VIF < 5. In addition, patient age was significantly associated with adenocarcinogenesis (HR: 1.8, 95% CI: 1.0–3.2, P = 0.0439) (Fig. 5A). Kaplan–Meier analysis showed that in the follow-up sample, the patients with CIN, with a maximum follow-up of 13 years, had a 90% probability of cancer (Fig. 5B). In summary, the above results demonstrate that CIN is a risk factor for adenocarcinogenesis.
Exploring the risk factors for adenocarcinoma progression after removal of adenomas. (A) Univariate Cox analysis to estimate the risk of adenocarcinoma progression after removal of adenoma. (B) Kaplan–Meier analysis to explore the effect of CIN on the risk of developing cancer. Red represents high risk and blue represents low risk.
Copy-number variation in the MFN2 gene is a risk factor for adenocarcinogenesis
To explore which specific gene with frequent changes promotes progression of adenoma to adenocarcinoma, we grouped the samples from the progressed-to-cancer group and not-progressed-to-cancer group and took the chromosomal variation data at the gene level separately. We obtained 2,033 genes as risk factors by univariate Cox analysis and downscaled by LASSO to obtain MFN2 (Fig. 6A–B). Among them, chr1p (MFN2) had a frequency of 27% in progressing patients; chr17p (TP53) had a frequency of 41.67% in progressing patients; chr8q (MYC) had a frequency of 50% in progressing patients. Among the follow-up patients, 48 patients were found to have progressed to adenocarcinoma, and 13 of them had deletion of the MFN2 gene, which accounted for 27% of the patients diagnosed with adenocarcinoma. In contrast, no deletion of the MFN2 gene was found in the follow-up patients who did not progress to adenocarcinoma. The TCGA database was also used to assess the relationship between MFN2 gene expression status and carcinogenesis. We found that MFN2 expression was reduced in tumor tissues relative to normal tissues (Fig. 6C). In addition, Kaplan–Meier analysis assessed the effect of the MFN2 gene copy number on the DFI, and we found that the MFN2 gene copy-number deletion promoted cancer recurrence after resection (P < 0.037) (Fig. 6D–E). The above results demonstrate that the MFN2 gene copy-number deletion could be a risk factor for the progression of adenoma to adenocarcinoma.
Copy-number variation in the MFN2 gene is a risk factor for adenocarcinogenesis. (A, B) Cvfit and lambda curves showing the least absolute shrinkage and selection operator (LASSO) regression were plotted with the minimum criteria. (C) Analysis of MFN2 expression in tumor and normal tissues based on the TCGA database. (D) Kaplan–Meier analysis to explore the effect of MFN2 expression on disease-free interval (DFI). The red color indicates copy-number deletion, and the blue color indicates copy-number increase or no change. (E) The receiver operating characteristic (ROC) curve shows the potential of the MFN2 copy-number deletion in predicting 1-, 3-, and 5-year DFI.
Discussion
CIN is one of the factors driving the progression of malignant tumors, enhancing intra-tumor heterogeneity, accelerating clonal evolution, driving phenotypic adaptation, and ultimately leading to poor clinical outcomes19,20,21. CRC can develop through adenomas, and thus resection at an early stage can somewhat avert the progression of the deterioration process22. However, the risk of recurrence or even progression to adenocarcinoma remains even after adenoma resection23,24. Therefore, in this study, we investigated the drivers of recurrence or progression to adenocarcinoma after adenoma resection through retrospective and follow-up studies of adenoma patients by LC-WGS.
In this study, 119 patients with adenomas were first recruited and grouped according to genomic CNVs in the excised tissues. We found that in the group with CIN, a higher percentage of patients presented with high-grade adenomas, suggesting that CIN frequently occurs in high-grade adenomas. Similar findings have been reported in previous studies, as Won-Tak Choi et al. found that high-grade atypical hyperplastic adenomas demonstrated aneuploidy, suggesting that they could develop into tumors through the CIN pathway25.
To further analyze the progression of adenomas, these patients were followed-up with for up to 17 years. A total of 88 patients experienced recurrence, including 40 cases of recurrence of adenomas and 48 cases of progression to adenocarcinomas. CIN was often found in tissues that developed into adenocarcinoma. This is consistent with previous reports that the main pathways for colorectal carcinogenesis include CIN and microsatellite instability (MSI)26,27. CIN is also present in CRC primary tumor tissues and distal metastatic cancer tissues28. In the present study, we found that in the patients who progressed to adenocarcinoma, the most significant changes occurred at 8q and 7p (gain) and at 18q, 18p, and 17p (loss). Interestingly, the main regions of gain and loss observed in this study are concordant with previously reported observations on CRC29. These results indicate that CIN in adenomas might lead to adenocarcinomas. Misregulation of a large number of genes is also a putative driver of the progression of epithelial cell adenomas to carcinoma, e.g., POFUT1 is a putative driver of adenoma progression to carcinoma through the Notch signaling pathway8. Unsurprisingly, misregulation of genes was also found in the present study. EGFR and MYC were found to have increased copy numbers, whereas TP53 was found to have decreased copy numbers in the patients who progressed to adenocarcinoma. EGFR and MYC are well-known oncogenes that are often highly expressed in CRC, and they promote CRC cell proliferation and migration30,31,32, while TP53 is mutated in 43% of CRC cases, damaging the function of TP53 33.
To further explore the risk factors for the progression to adenocarcinoma after adenoma resection, we performed univariate analysis of clinical characteristics and chromosomal alterations in the patients who were followed-up with, and we found that CIN was the most significant factor for the progression of adenoma to adenocarcinoma (HR: 2.5, 95% CI: 1.4–4.5, P = 0.00162). Previous studies have also demonstrated that CIN could drive CRC progression34,35. Thus, our conclusions are reasonable, and CIN can be used as one of the indicators to determine the progression of adenoma to adenocarcinoma in clinical practice.
Further analysis revealed that the Mitochondrial fusion protein 2 (MFN2) copy-number deletion was a risk factor for adenocarcinogenesis. MFN2 is involved in a variety of biological processes, including cancer. Cheng’s study showed that MFN2 was downregulated in CRC tissues and was associated with a poor prognosis in CRC patients12. This result is consistent with the results of the present study, in which MFN2 was found to be downregulated in tumors and a low copy number of MFN2 shortened the recurrence period after resection of cancerous tissues. In contrast, increasing MFN2 expression inhibits tumorigenesis15,36. These results show that, on the one hand, the low copy number of MFN2 could be a diagnostic indicator for predicting the progression to adenocarcinoma after initial resection of adenoma, and on the other hand, MFN2 might also be a potential target for the treatment of CRC. Clinically, CIN status could inform surveillance intervals, with CIN-positive adenomas warranting more frequent colonoscopy monitoring. MFN2 status may guide adjuvant therapy decisions.
However, there are still limitations to this study; that is, due to limitations in accessing fresh tissue samples and ethical restrictions on in vivo experimentation, the specific mechanism and function of MFN2 in adenomas and adenocarcinomas were not analyzed. Accordingly, subsequent studies should focus on the mechanism by which MFN2 deficiency drives the progression of adenomas to adenocarcinomas. Currently, there are numerous techniques for detecting CIN, including spatial CITE-seq 37 and spatially resolved gene screening using Perturb-DBiT. In this study, the traditional LC-WGS method was employed, while novel gene screening techniques may be utilized for gene detection in future research. Furthermore, the modest sample size in the MFN2 subgroup warrants validation in larger cohorts. Future studies will focus on expanding these findings.
Conclusions
In this study, we performed a 17-year follow-up of 119 patients with adenomas and analyzed their primary and recurrent paired samples by LC-WGS. We found that CIN was an important risk factor for recurrence and/or progression of adenomas after initial resection. More importantly, the CNA patterns between the tumors and matched previous polyps were highly matched, which further supports a genetic link between adenomas and adenocarcinomas. In addition, the MFN2 gene copy-number deletion was significantly associated with recurrence and/or progression of adenomas after initial resection. This study provides new ideas for predicting the progression of adenomas to adenocarcinomas in the clinical setting.
Data availability
The datasets generated and/or analyzed during the current study are available in the NGDC repository under the query code HRA010671. The data can be accessed at https://ngdc.cncb.ac.cn/gsa/. Further inquiries can be directed to the corresponding author. Additionally, the datasets presented in this study can be found in online repositories, and the names of these repositories are provided in the article.
Abbreviations
- CRC:
-
Colorectal cancer
- CINs:
-
Chromosomal instabilities
- LC-WGS:
-
Low-coverage whole-genome sequencing
- CAN:
-
Copy-number aberration
- CBS:
-
Cyclic binary segmentation
- CNVs:
-
Copy number variations
- COAD:
-
Colon adenocarcinoma
- DFI:
-
Disease-free interval
References
Kuipers, E. J. et al. Colorectal cancer. Nat. Rev. Dis. Primers. 1, 15065 (2015).
Tirendi, S. et al. Colorectal cancer and therapy response: a focus on the main mechanisms involved. Front. Oncol. 13, 1208140 (2023).
Wang, S. et al. NF-κB activator 1 downregulation in macrophages activates STAT3 to promote adenoma-adenocarcinoma transition and immunosuppression in colorectal cancer. BMC Med. 21, 115 (2023).
González, I. A., Bauer, P. S., Liu, J. & Chatterjee, D. Adenoma-like adenocarcinoma: clinicopathologic characterization of a newly recognized subtype of colorectal carcinoma. Hum. Pathol. 107, 9–19 (2021).
Søreide, K., Nedrebø, B. S., Reite, A., Thorsen, K. & Kørner, H. Endoscopy, morphology, morphometry and molecular markers: predicting cancer risk in colorectal adenoma. Expert Rev. Mol. Diagn. 9, 125–137 (2009).
Zheng, S., Guerrero-Haughton, E. & Foijer, F. Chromosomal Instability-Driven cancer progression: interplay with the tumour microenvironment and therapeutic strategies. Cells 12 (23), 2712 (2023).
Chen, X., Agustinus, A. S., Li, J., DiBona, M. & Bakhoum, S. F. Chromosomal instability as a driver of cancer progression. Nat. Rev. Genet. 26 (1), 31–46 (2025).
Komor, M. A. et al. Molecular characterization of colorectal adenomas reveals POFUT1 as a candidate driver of tumor progression. Int. J. Cancer. 146, 1979–1992 (2020).
Garg, A. D. et al. ROS-induced autophagy in cancer cells assists in evasion from determinants of Immunogenic cell death. Autophagy 9 (9), 1292–1307 (2013).
Huang, Q. et al. Increased mitochondrial fission promotes autophagy and hepatocellular carcinoma cell survival through the ROS-modulated coordinated regulation of the NFKB and TP53 pathways. Autophagy 12 (6), 999–1014 (2016).
Luo, L. et al. MFN2 suppresses clear cell renal cell carcinoma progression by modulating mitochondria-dependent dephosphorylation of EGFR. Cancer Commun. (Lond). 43 (7), 808–833 (2023).
Cheng, X., Li, Y. & Liu, F. Prognostic impact of Mitofusin 2 expression in colon cancer. Transl Cancer Res. 11, 3610–3619 (2022).
Cormio, A. et al. Mitofusin-2 Down-Regulation predicts progression of Non-Muscle invasive bladder cancer. Diagnostics (Basel). 11 (8), 1500 (2021).
Ashraf, R. & Kumar, S. Mfn2-mediated mitochondrial fusion promotes autophagy and suppresses ovarian cancer progression by reducing ROS through AMPK/mTOR/ERK signaling. Cell. Mol. Life Sci. 79 (11), 573 (2022).
Xu, K. et al. MFN2 suppresses cancer progression through Inhibition of mTORC2/Akt signaling. Sci. Rep. 7, 41718 (2017).
Mo, H. et al. Genome-wide chromosomal instability by cell-free DNA sequencing predicts survival in patients with metastatic breast cancer. Breast 53, 111–118 (2020).
Guan, Y. et al. Copy number variation of urine exfoliated cells by low-coverage whole genome sequencing for diagnosis of prostate adenocarcinoma: a prospective cohort study. BMC Med. Genomics. 15, 104 (2022).
Ye, L. P. et al. Cost-effective low-coverage whole-genome sequencing assay for the risk stratification of gastric cancer. World J. Gastrointest. Oncol. 14, 690–702 (2022).
Sansregret, L., Vanhaesebroeck, B. & Swanton, C. Determinants and clinical implications of chromosomal instability in cancer. Nat. Rev. Clin. Oncol. 15, 139–150 (2018).
Maleki, S. S. & Röcken, C. Chromosomal instability in gastric cancer biology. Neoplasia 19, 412–420 (2017).
Penner-Goeke, S. et al. The Temporal dynamics of chromosome instability in ovarian cancer cell lines and primary patient samples. PLoS Genet. 13, e1006707 (2017).
Dumoulin, F. L. & Hildenbrand, R. Endoscopic resection techniques for colorectal neoplasia: current developments. World J. Gastroenterol. 25, 300–307 (2019).
Seo, J. Y. et al. Novel risk stratification for recurrence after endoscopic resection of advanced colorectal adenoma. Gastrointest. Endosc. 81, 655–664 (2015).
Tate, D. J. et al. Treatment of adenoma recurrence after endoscopic mucosal resection. Gut 72, 1875–1886 (2023).
Choi, W. T. et al. DNA content analysis of colorectal serrated lesions detects an aneuploid subset of inflammatory bowel disease-associated serrated epithelial change and traditional serrated adenomas. Histopathology 73, 464–472 (2018).
Kim, J. C. & Bodmer, W. F. Genomic landscape of colorectal carcinogenesis. J. Cancer Res. Clin. Oncol. 148, 533–545 (2022).
Pussila, M. et al. Mitotic abnormalities precede microsatellite instability in Lynch syndrome-associated colorectal tumourigenesis. EBioMedicine 103, 105111 (2024).
Golas, M. M. et al. Evolutionary patterns of chromosomal instability and mismatch repair deficiency in proximal and distal colorectal cancer. Colorectal Dis. 24, 157–176 (2022).
Orsetti, B. et al. Impact of chromosomal instability on colorectal cancer progression and outcome. BMC cancer. 14, 121 (2014).
Gamez-Belmonte, R. et al. Epithelial presenilin-1 drives colorectal tumour growth by controlling EGFR-COX2 signalling. Gut 72, 1155–1166 (2023).
Donato, M. D. et al. RIZ2 at the crossroad of the EGF/EGFR signaling in colorectal cancer. J. Transl Med. 21, 736 (2023).
Di, Y. et al. The c-MYC-WDR43 signalling axis promotes chemoresistance and tumour growth in colorectal cancer by inhibiting p53 activity. Drug Resist. Updat. 66, 100909 (2023).
Liebl, M. C. & Hofmann, T. G. The role of p53 signaling in colorectal cancer. Cancers 13, 2125 (2021).
Miao, S. et al. DIAPH1 regulates chromosomal instability of cancer cells by controlling microtubule dynamics. Eur. J. Cell. Biol. 100, 151156 (2021).
Chen, B. et al. The long noncoding RNA CCAT2 induces chromosomal instability through BOP1-AURKB signaling. Gastroenterology 159, 2146–2162e2133 (2020).
Ashraf, R. & Kumar, S. Mfn2-mediated mitochondrial fusion promotes autophagy and suppresses ovarian cancer progression by reducing ROS through AMPK/mTOR/ERK signaling. Cell. Mol. Life Sci. 79, 573 (2022).
Liu, Y. et al. High-plex protein and whole transcriptome co-mapping at cellular resolution with Spatial CITE-seq. Nat. Biotechnol. 41 (10), 1405–1409 (2023).
Acknowledgements
We thank LetPub (www.letpub.com.cn) for its linguistic assistance during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
H.L.: Conceptualization; methodology; investigation; writing—original draft. F.Y.: Investigation; data curation; validation. B.B.: Formal analysis; visualization. Z.J.: Investigation; validation. B.L.: Formal analysis; validation. G.F.: Validation; writing—review and editing. X.H.: Writing—review and editing; resources; supervision; project management. The work reported in the paper has been performed by the authors unless clearly specified in the text.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent for participate
This study was approved by the Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (20220305-33). Samples were obtained from the Department of Pathology, Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, and maintained by the research team. These specimens were collected after obtaining informed consent from the participants. All experiments using clinical samples were conducted in accordance with the approved guidelines. This study was conducted in accordance with the Declaration of Helsinki..
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, H., Yang, F., Bai, B. et al. Tumor associated chromosomal instability drives colorectal adenoma to adenocarcinoma progression based on 17 year follow up evidence. Sci Rep 15, 13733 (2025). https://doi.org/10.1038/s41598-025-96921-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-96921-8