Abstract
The efficacy of automated lesion detection in clinical settings is often hampered by two primary factors: the vast range of pathological scales and the presence of non-target anatomical interference. Standard Mamba-based detectors, while efficient, frequently suffer from fixed receptive fields and background signal leakage. To resolve these challenges, we introduce ScaleMamba-YOLO, an enhanced medical object detection framework that integrates selective state-space modeling with adaptive local feature refinement. Our approach features two innovative structural components: first, a Medical Multi-scale Local Feature Enhancement Block (MMLFE-Block) is positioned at the frontend to diversify the initial receptive field. By utilizing a parallel architecture with heterogeneous convolutional kernels (1\(\times\)1, 3\(\times\)3, and 5\(\times\)5), the model achieves comprehensive hierarchical perception, enabling the concurrent identification of minute calcified spots and extensive diffuse lesions. Second, a Partial-Enhanced C2F (PEC2F) module is designed to refine feature aggregation post-global modeling. This component employs partial convolution (PConv) to selectively process salient feature channels, effectively filtering out irrelevant background noise from normal tissue structures. The robust performance of ScaleMamba-YOLO was validated across three specialized medical datasets (Br35H, BCCD, and PLoPy) and a standard scene dataset (VOC0712). The model recorded Average Precision (AP) scores of 72.7%, 65.0%, 85.7%, and 64.6%, respectively. These metrics represent consistent improvements of 1.7% to 2.3% over the MambaYOLO baseline, underscoring the system’s potential for high-fidelity diagnostic assistance in real-time clinical workflows.
Similar content being viewed by others
Introduction
Deep learning-based object detection and feature extraction technologies have recently witnessed explosive growth and successful applications across a wide range of engineering fields. Notable examples include remote sensing image classification1,2, aerial video tracking3,4, domain adaptation in complex environments5,6, and even signal processing tasks for antenna design7,8,9 and drowsiness detection10.
However, transitioning from these general domains to the contemporary smart healthcare ecosystem presents unique challenges. As a cornerstone of medical image analysis, object detection has been deeply integrated into critical scenarios such as pathological slide analysis, CT/MRI diagnostic imaging, endoscopic surgical navigation, and ultrasound image interpretation. Unlike natural scene images, the inherent complexity of medical images–characterized by vast variations in lesion size, irregular morphologies, and ambiguous boundaries with surrounding normal tissues–poses significant challenges to the precision and robustness of detection algorithms.
Regarding multi-scale feature extraction, the immense disparity in lesion sizes presents a core difficulty in medical image detection. ConvNeXt, proposed by LIU et al.11 in 2022, demonstrated the substantial potential of pure convolutional networks in terms of effective receptive fields and multi-scale capabilities by incorporating designs like large kernel sizes. In the same year, RepLKNet, designed by DING et al.12, and the Visual Attention Network (VAN), proposed by GUO et al.13, explored ultra-large kernel designs from the perspectives of convolution and attention, respectively, to efficiently capture global information. To achieve more dynamic feature extraction, HorNet by RAO et al.14 and MPViT by LEE et al.15implemented parallel processing and interaction of features at different scales by constructing recursive gated convolutions and multi-path Transformer structures. Building on this, in 2023, InternImage by WANG et al.16 combined the concept of deformable convolution with large kernel designs, while the dynamic snake convolution proposed by Qi et al.17 in the same year offered more flexible geometric adaptability for specific medical tasks. Although these cutting-edge designs have significantly enhanced feature representation, they often come with high computational costs, and their generalized designs may not be optimally suited for the specific morphological characteristics of medical lesions. In terms of background interference suppression, lesion targets in medical images are frequently surrounded by morphologically similar normal tissues, making models susceptible to background interference, as illustrated in Fig. 1(b). To address this issue, Transformer-based detectors have made notable progress. DINO, introduced by ZHANG et al.18in 2022, compelled the model to learn more precise foreground-background distinctions by incorporating a contrastive denoising training task. Co-DETR by Zong et al.19 and RT-DETR, designed by LV et al.20 in 2023, improved the convergence efficiency and target focus of DETR-like models by optimizing training strategies and model architectures. Furthermore, advanced label assignment strategies are crucial, such as YOLOX by GE et al.21 and TOOD by FENG et al.22, which mitigate background noise at the source by dynamically assigning high-quality positive samples. While these methods have proven effective, the complex designs of Transformer-based models and dynamic assignment strategies still introduce significant computational overhead, especially in clinical scenarios requiring real-time feedback.
Example comparison of detection results between MambaYOLO and the proposed method with the PEC2F module.
Recently, State Space Models (SSMs), particularly the Mamba architecture .23.24.25, have offered a novel perspective for addressing the aforementioned challenges. By introducing a Selective Scan Mechanism (S2M), Mamba can efficiently model long-range dependencies in sequential data with linear complexity while dynamically focusing on task-relevant information. This characteristic allows it to maintain global context modeling capabilities while being significantly more computationally efficient than Transformers. Inspired by this, researchers have begun to explore its applications in the visual domain (e.g., Vision Mamba .26.27.28.29 ) and have constructed efficient detection frameworks such as MambaYOLO .30. Despite the impressive performance of MambaYOLO, its direct application to medical imaging reveals two key architectural deficiencies: 1) Its local feature extraction relies on standard convolutions, which have limited perceptual capabilities for the extreme scale variations of medical lesions; and 2) At the level of feature fusion and global modeling, the model lacks a targeted mechanism for suppressing background noise. As shown in Fig. 1(b), when a lesion is surrounded by morphologically similar normal tissue, Mamba’s global information aggregation process tends to amplify irrelevant features from background regions, thereby contaminating the final feature representation.
Inspired by the above analysis, this paper proposes ScaleMamba-YOLO, a network deeply optimized based on the MambaYOLO architecture, to solve the specific challenges in medical image detection. The network aims to systematically enhance the model’s capabilities in multi-scale perception and background suppression through the serial integration of specially designed CNN modules with Mamba. Specifically, the data flow progresses through a local feature enhancement stage (MMLFE-Block) to capture multi-scale details, followed by Mamba’s global modeling, and finally a partial-convolution-based aggregation stage (PEC2F) to refine features and suppress noise. This serial design ensures that global dependency modeling is performed on highly representative local features.Its main contributions are as follows:
-
(1)
We propose the ScaleMamba-YOLO framework, which is specifically optimized for medical imaging. This framework effectively combines Mamba’s selective state-space mechanism with specially designed CNN modules to address the dual challenges of long-range dependencies and local feature diversity in medical images.
-
(2)
We design a Medical Multi-scale Local Feature Enhancement Block (MMLFE-Block). Positioned before Mamba’s global modeling, this module enhances the model’s initial local perception of lesions of varying sizes through parallel 1\(\times\)1, 3\(\times\)3, and 5\(\times\)5 heterogeneous convolution kernels, providing a more scale-robust feature input for subsequent processing.
-
(3)
We design a Partial-Enhanced C2f (PEC2F) module. After Mamba completes its global feature aggregation, this module achieves efficient feature refinement by integrating Partial Convolution technology into the classic C2f architecture. As illustrated in Fig. 1(c), PEC2F selectively processes key feature channels, effectively suppressing background noise that may have been amplified during the global modeling process and improving the signal-to-noise ratio of the final features.
-
(4)
The effectiveness of ScaleMamba-YOLO is validated through experiments on four datasets: three medical imaging datasets (Br35H, BCCD, and PLoPy) and one general-purpose dataset (VOC0712). The model achieved Average Precision (AP) scores of 72.7%, 65.0%, 85.7%, and 64.6%, respectively, representing improvements of 2.2%, 1.9%, 1.7%, and 2.3% over the baseline. These results demonstrate that our architectural optimizations are effective not only for medical imaging but also for natural scene object detection.
Related work
MamaYolo
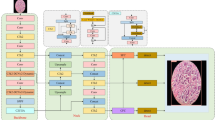
MambaYOLO is a novel object detection network that integrates the speed advantages of the YOLO series models with the global feature modeling capabilities of the Transformer .31 model. Its network architecture, illustrated in Fig. 2, follows the mainstream Backbone-Neck-Head design and is primarily composed of three parts: (a) the ODMamba backbone, (b) the PAFPN neck, and (c) the detection head.
The structure of the MambaYOLO.
PConv
Partial Convolution (PConv).32 is a technique that applies convolutional operations to only a subset of the input feature map channels to optimize feature extraction while preserving critical local information. Its structure is illustrated in Fig. 3. PConv divides the channels of the input feature map into two parts: one part undergoes the convolution operation, while the other is preserved directly. This design simplifies the complexity of feature extraction. Subsequently, a Pointwise Convolution (PWConv) fuses the features from both parts, improving the model’s expressive power. PConv offers both flexibility and efficiency, allowing it to effectively capture local features, reduce redundant operations, and ensure that critical features are fully extracted and utilized. Its mathematical formulation is shown in Equations (1-2).
In Equation (1), a channel splitting operation is performed to divide the input feature map X into a subset for processing, denoted as \(X_{p}\), and an untouched subset, denoted as \(X_{u}\), where \(c_{p}\) represents the number of channels designated for convolution. In this study, following the design principles of FasterNet.32, we set the partial convolution ratio to \(c_p = \frac{c}{4}\). This configuration is chosen because feature maps often contain redundant information; processing only 25% of the channels is sufficient to extract spatial dependencies while significantly reducing computational cost (FLOPs) and memory access latency. For medical images, preserving the remaining 75% of channels without convolution helps retain subtle background textures that might otherwise be lost in aggressive downsampling. In Equation (2), the convolution operation with kernel weights \(W_k\) is applied to \(X_{p}\). The resulting features are then concatenated with \(X_{u}\), and a pointwise convolution parameterized by the weight matrix \(W_{z}\) is applied to fuse the features and produce the final output Z
The structure of the PConv.
Methods
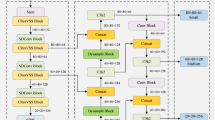
Given that the original ODSSBlock lacks specific optimization for the characteristics of medical images, the ScaleMamba-YOLO proposed in this paper redesigns the fundamental backbone module of MambaYOLO. It is replaced with the SMPBlock, which is better adapted to the significant variations in lesion sizes found in medical images. The structure of the SMPBlock is illustrated in Fig. 4. The SMPBlock module is an organic combination of three functional components: SS2D, MMLFE-Block, and PEC2F. The model retains the original SS2D module to inherit its core advantage in efficiently modeling global long-range dependencies. Secondly, the proposed MMLFE-Block, through its parallel multi-scale convolutional kernel architecture, endows the model with the ability to simultaneously perceive both minute lesions and large, diffuse pathologies, thereby addressing the challenge of drastic variations in lesion size. Finally, the PEC2F module utilizes partial convolution technology to streamline the feature aggregation process, which significantly suppresses background noise interference caused by complex normal tissues. This design closely integrates global context modeling, multi-scale local perception, and high signal-to-noise ratio feature aggregation. As a result, the SMPBlock can generate feature representations that are more adaptive to diverse medical detection tasks, fundamentally enhancing the overall network’s detection accuracy and robustness.
The structure of the ScaleMamba-YOLO.
MMLFE-block
In the task of medical image object detection, lesions exhibit significant scale variations, ranging from tiny calcifications to large, diffuse tumors. Consequently, the single 3\(\times\)3 convolutional kernel employed by MambaYOLO, due to its fixed receptive field, struggles to effectively and simultaneously capture the pathological features of lesions at these different scales. Therefore, to address the unique characteristics of medical images, this paper proposes the Medical Multi-scale Local Feature Enhancement Block (MMLFE-Block). This module operates through a three-stage process. First, in the multi-scale feature co-extraction stage, it employs parallel 1\(\times\)1, 3\(\times\)3, and 5\(\times\)5 depth-wise separable convolutional kernels to achieve hierarchical capture of pixel-level local details through their differentiated receptive fields. Following the aggregation of these multi-branch features, the module enters a feature fusion and enhancement stage, where a secondary 3\(\times\)3 depth-wise separable convolution is introduced. This serves a dual purpose: on one hand, it avoids the semantic gaps caused by direct feature concatenation (e.g., the semantic conflict between a tiny calcification and a large tumor) through progressive receptive field expansion. On the other hand, it leverages the local continuity prior of convolution to suppress feature fragmentation noise that may arise from parallel computation, thereby better aligning with the anatomical continuity of medical images. Finally, the process concludes with a residual-guided feature optimization stage, where a residual connection creates a weighted fusion of the original input with the multi-scale processed features, simultaneously preserving the integrity of underlying anatomical structures while enhancing the saliency of high-level pathological features.
As shown in Fig. 4, this module adopts a three-stage hierarchical design to process the input features. The first stage expands the channel dimension through a linear projection and introduces a non-linear transformation, as expressed in Equation (3):
where x represents the input feature. \(\mathscr {C}(\cdot )\) denotes the linear projection layer (implemented as a \(1\times 1\) convolution) to expand the channel dimension fourfold, and \(\Phi (\cdot )\) represents the GELU activation function.
Multi-scale feature co-extraction stage: This stage employs parallel \(\{1 \times 1, 3 \times 3, 5 \times 5\}\) depth-wise separable convolutional kernels to construct differentiated receptive fields, as expressed in Equation (4):
Where \(\text {DWConv}_{\text {ks}}\) represents a depth-wise separable convolution with a kernel size of ks. KS defines a set of parallel convolutional kernels with values of \(\{1, 3, 5\}\).
Feature fusion and enhancement stage: The multi-scale features are progressively fused through a \(3 \times 3\) depth-wise separable convolution, as expressed in Equation (5):
This design avoids the semantic gaps caused by direct feature concatenation and utilizes the local continuity prior of convolution to suppress feature fragmentation noise.
Residual-guided feature optimization stage: After achieving channel interaction using a \(1 \times 1\) convolution, a residual connection is used to preserve the integrity of the anatomical structure, as expressed in Equation (6):
This design enables the model to understand and integrate features across different dimensions of the image, while also preserving the underlying structure and enhancing the saliency of pathological features, thereby improving its robustness to scale variations.
PEC2F
In medical image detection tasks, the target region is often surrounded by a structurally complex background of normal tissue, which reduces feature discriminability, as illustrated in Fig. 1(b). To precisely isolate target features and suppress background interference, this paper first constructs a novel C2f module that enhances the network’s foundational feature extraction capabilities by optimizing gradient flow paths and cross-layer feature fusion mechanisms. Building on this, to further improve the model’s robustness to redundant information and refine high-level semantic features, we propose a partial convolution enhancement module, namely PEC2F (Partial Enhanced C2f). The core idea of this module is to seamlessly integrate Partial Convolution (PConv) into the feature processing stream to achieve computational suppression of invalid regions and an adaptive focus on key features.
The structural design of PEC2F is shown in Fig. 4. This design strategically embeds PConv operations at both ends of the proposed C2f framework. To facilitate the ablation study discussion, we explicitly define the nomenclature based on the data flow direction: the PConv module located at the input stage (before the channel split) is designated as the “Left” component (P-l), while the PConv module located at the output stage (after feature concatenation) is designated as the “Right” component (P-r).At the frontend of the module (corresponding to P-l), the input feature map is first processed by a PConv layer. This operation can effectively identify and mask irrelevant or redundant activation areas, thereby reducing the complexity of subsequent computations without losing critical information. The feature stream, preliminarily purified by PConv, then enters the core C2f structure, which is composed of multiple Bottleneck units, for deep feature transformation and multi-scale fusion. At the backend of the module, another PConv layer is used for the final calibration and enhancement of the fused features, ensuring that the output feature representation possesses both high discriminability and contextual awareness.
The mathematical expressions for this module are shown in Eqs. (7-9):
The expressions for PCBS, DWCBS, and Bottleneck are shown in Eqs. (10-12), respectively:
Results
Experimental setup
The experiments in this paper were conducted by training the model on three medical imaging datasets (Br35H, BCCD, and PLoPy) and one general-purpose object detection dataset (VOC0712), with model performance measured using COCO evaluation metrics. The hardware configuration consisted of an Intel Core i9-10980XE CPU and an NVIDIA GeForce RTX 4080 GPU. The software environment included the Ubuntu 18.04.6 LTS operating system, the PyTorch 1.13.0 framework, the CUDA 11.7 acceleration library, and the Python 3.11 programming environment. For model training, a fixed input size of 640\(\times\)640 and a batch size of 16 were used, with the SGD optimizer set to a learning rate of 0.001. The number of training epochs was set according to the characteristics of each dataset: 150 epochs for the Br35H dataset, and 300 epochs each for the BCCD, PLoPy, and VOC0712 datasets.
Datasets and evaluation metrics
To ensure robust and reliable evaluation, we conducted experiments on four public datasets using rigorous data splitting protocols. The medical imaging benchmarks included the Br35H brain tumor dataset (701 MRI images: 500 training, 201 validation), the BCCD blood cell dataset (874 microscopy images: 765 training, 73 validation, and 36 testing), and the large-scale PLoPy polyp dataset (27,048 endoscopic images: 21,638 training, 5,410 validation). Furthermore, to verify domain generalization, we utilized the Pascal VOC dataset (21,503 natural images across 20 categories: 16,551 training, 4,952 validation).
Model performance was quantified using standard object detection metrics, with Average Precision (AP) serving as the primary comprehensive index. We specifically examined localization accuracy using \(AP_{50}\) and \(AP_{75}\) at IoU thresholds of 0.5 and 0.75, respectively. Additionally, the model’s adaptability to scale variations was assessed through precision scores for small (\(AP_s\)), medium (\(AP_m\)), and large (\(AP_l\)) objects.
Results
To comprehensively and systematically evaluate the effectiveness and generalization capability of the ScaleMamba-YOLO model, a series of comparative experiments were conducted on four challenging public datasets, covering both medical imaging (Br35H brain tumors, BCCD blood cells, endoscopic polyps) and natural scenes (VOC0712). This section will present the detailed comparative results and provide an in-depth analysis of the mechanisms behind the model’s performance advantages.
In the Br35H brain tumor detection task, ScaleMamba-YOLO’s exceptional performance is immediately apparent. As detailed in Table 1, it achieves a top-tier Average Precision (AP) of 72.7%, creating a significant performance gap by improving upon YOLOv9 by 1.6% and MambaYOLO by 2.2%. Its advantage is further solidified at stricter IoU thresholds, boasting an AP\(_{75}\) score of 89.5%. A key contributor to this success, highlighted by the table, is its powerful multi-scale detection capability. The model scores an impressive 28.3% on small object precision (AP\(_{s}\)) and 77.9% on large object precision (AP\(_{l}\)), representing substantial gains of 3.1% and 3.3% over strong baselines, respectively. This demonstrates the efficacy of its internal mechanisms in capturing nuanced tumor features across different scales.
This trend of superior performance continues in other medical imaging domains. For the BCCD blood cell dataset, Table 2 shows ScaleMamba-YOLO once again leading the field with an AP of 65.0%. Its most significant advantage lies in the identification of small and medium-sized cells–a crucial requirement for accurate clinical diagnosis–where its AP\(_{s}\) (51.8%) and AP\(_{m}\) (68.6%) scores are unmatched, surpassing the Transformer-based DINO by 1.5% and 3.9%, respectively. In the more challenging task of endoscopic polyp detection (PLoPy), where images are often compromised by complex backgrounds, the model’s robustness shines, as shown in Table 3. It achieves the highest AP of 85.7%, and more notably, makes a tremendous leap in detecting easily missed small and medium-sized polyps, with its AP\(_{s}\) (72.2%) and AP\(_{m}\) (79.1%) scores showing massive improvements of 6.9% and 6.1% over DINO.
Finally, to evaluate the model’s versatility, we tested its performance on the general-purpose PASCAL VOC0712 dataset. As shown in Table 4, ScaleMamba-YOLO achieves a leading AP of 64.6%, outperforming comparative models.
This strong performance on natural images suggests that the core challenges addressed by our design—specifically scale variation and background interference—are shared across domains. While the MMLFE-Block was initially developed to handle varying lesion sizes, it proves equally effective at capturing objects of diverse scales in natural scenes. Similarly, the PEC2F module, designed to suppress tissue noise, aids in filtering out cluttered backgrounds common in VOC images. These results indicate that the proposed architectural improvements possess a degree of universality, extending their robustness beyond the medical domain.The consistent performance gains observed across four diverse datasets (medical and natural images) and all evaluation metrics (AP, \(AP_{50}\), \(AP_{75}\)) substantiate the robustness of our method, confirming that these improvements stem directly from the effective architectural optimizations.
Ablation study
The ScaleMamba-YOLO network is primarily composed of two main functional modules: the Medical Multi-scale Local Feature Enhancement Block (MMLFE-Block) and the Partial-Enhanced C2f (PEC2F) feature aggregation module. To systematically validate the independent contributions and synergistic effects of each module, a comprehensive ablation study was conducted on the Br35H dataset. To investigate the optimal configuration of the PEC2F module, this study also included an ablation of its internal structure by introducing partial convolution on the left (P-l) and right (P-r) sides of the C2f structure, respectively.
As shown in Table 5, in the Br35H brain tumor detection task, the baseline model’s Average Precision (AP) was 70.5%. After introducing the MMLFE-Block alone, the AP increased to 72.1%, with significant gains in the AP\(_{m}\) and AP\(_{l}\) metrics. This indicates that the module’s multi-scale feature extraction capability effectively enhanced the model’s ability to recognize and localize medium and large lesions. The internal ablation of the PEC2F module showed that both its P-r and P-l components could independently improve performance, while the complete PEC2F module raised the AP to 72.1% and performed better on the AP\(_{s}\) metric, validating its effectiveness in optimizing feature aggregation and suppressing background noise through partial convolution. Finally, when the two modules worked in synergy, the model’s performance peaked, with the AP increasing to 72.7%. This result confirms that the multi-scale features provided by MMLFE-Block offer higher-quality input for PEC2F’s feature fusion, and their combination produces a significant synergistic effect.
Impact of kernel size and parameter efficiency
To empirically justify the kernel size selection in the MMLFE-Block, we compared four configurations: Baseline, \(\{1, 3, 3\}\), \(\{1, 3, 5\}\), and \(\{1, 3, 5, 7\}\) across all four datasets. The results are visualized in Fig. 5.
Ablation study on kernel size configurations across four datasets. The line chart (left axis) represents Average Precision (AP), while the bar chart (right axis) represents the number of parameters (Params). The \(\{1, 3, 5\}\) configuration achieves the optimal balance, offering significant accuracy gains over the baseline without the excessive parameter overhead seen in the \(\{1, 3, 5, 7\}\) configuration.
As observed in Fig. 5, moving from the baseline to the \(\{1, 3, 5\}\) configuration results in a substantial improvement in AP across all datasets (e.g., Br35H improves from \(72.7\%\) to \(\approx 73.8\%\)) with a moderate increase in parameters (from 5.98M to 6.92M). However, further expanding the kernel set to include \(7 \times 7\) (\(\{1, 3, 5, 7\}\)) increases the parameter count to 7.65M—a significant overhead—while yielding negligible or plateauing performance gains. Consequently, we adopted \(\{1, 3, 5\}\) as the final configuration to prioritize computational efficiency for medical applications.
Model efficiency analysis
As shown in Table 6, we conducted a comprehensive analysis of model complexity and inference speed. It is observed that ScaleMamba-YOLO exhibits a moderate increase in parameters (7.82M) and computational cost (17.0 GFLOPs) compared to the baseline Mamba YOLO (5.98M, 13.6 GFLOPs). This increase is an expected trade-off resulting from the integration of the multi-scale MMLFE-Block and the PEC2F module. However, despite this slight overhead, our model remains significantly more lightweight than Transformer-based detectors like RT-DETR (41.94M parameters, 125.6 GFLOPs) and DINO (47.54M parameters).
Crucially, in terms of inference speed, ScaleMamba-YOLO achieves 110 FPS (9.1 ms per image) on an RTX 4080 GPU. While this is lower than the lightweight YOLOv8 series, it far exceeds the real-time requirement (typically 30 FPS) for clinical video analysis and surgical navigation. This indicates that our architectural improvements successfully strike an optimal balance between high detection accuracy and deployment efficiency, making it suitable for practical medical applications where precision is paramount but real-time feedback is also required.
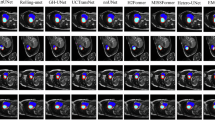
Comparative Visualization between ScaleMamba-YOLO and MambaYOLO.
Visualization
The object detection model proposed in this paper demonstrates excellent performance across a variety of scenarios. As shown in Fig. 6, in the four tasks of brain tumor detection, blood cell detection, polyp detection, and natural image object detection, our model’s detection results maintain a high degree of consistency with the Ground Truth (GT) annotations. Specifically, in the brain tumor detection task, our model accurately identified the tumor region (with an IoU of 0.39 between the detection box and the GT box), whereas the comparative model, MambaYOLO, experienced a missed detection. In the blood cell detection task, our model not only achieved a precise identification of multiple blood cell types but also outputted higher and more stable confidence scores. For both the polyp detection and natural image detection tasks, the overlap of our model’s detection boxes with the GT boxes, as well as its confidence scores, were significantly superior to the baseline model. These experimental results indicate that our model possesses excellent feature extraction capabilities and localization accuracy, demonstrating stable and superior detection performance across different datasets.
Limitations and future work
ScaleMamba-YOLO’s limitations include: its unproven generalization to other imaging modalities like X-ray or ultrasound, as validation was limited to MRI, microscopy, and endoscopy data; its dependency on data size and quality, a challenge given medical data scarcity; and the need for computational efficiency optimization for real-time clinical deployment on resource-constrained devices.
Future research will focus on exploring few-shot learning strategies to address data scarcity and on further improving the model’s computational efficiency, aiming to broaden its applicability in diverse clinical scenarios.
Conclusion
This paper proposes a multi-scale object detection network based on MambaYOLO, named ScaleMamba-YOLO. By introducing the Medical Multi-scale Local Feature Enhancement Block (MMLFE-Block) and the Partial-Enhanced C2f (PEC2F) feature aggregation module, the model’s capability to detect lesions of different scales and its feature fusion efficiency were effectively improved, thereby enhancing the accuracy of precise localization and classification of targets. Experimental results show that ScaleMamba-YOLO performs exceptionally well on the medical datasets Br35H, BCCD, and PLoPy, as well as on the general-purpose dataset VOC0712, validating its effectiveness.
Data availability
The datasets analysed during the current study are publicly available. The Br35H dataset is available from the Baidu AI Studio platform at https://aistudio.baidu.com/datasetdetail/273600. The BCCD dataset is available from the Roboflow repository at https://universe.roboflow.com/joseph-nelson/bccd. The PLoPy dataset is available from the Baidu AI Studio platform at https://aistudio.baidu.com/datasetdetail/37521. The Pascal VOC0712 dataset is available from a PASCAL VOC dataset mirror at https://pjreddie.com/projects/pascal-voc-dataset-mirror/.
References
Gomaa, A. & Saad, O. M. Residual channel-attention (RCA) network for remote sensing image scene classification. Multimed. Tools Appl. 84, 33837–33861. https://doi.org/10.1007/s11042-024-20546-8 (2025).
Salem, M., Gomaa, A. & Tsurusaki, N. Detection of earthquake-induced building damages using remote sensing data and deep learning: A case study of Mashiki Town, Japan. In IGARSS 2023 - 2023 IEEE International Geoscience and Remote Sensing Symposium, 2350–2353. https://doi.org/10.1109/IGARSS52108.2023.10282550 (IEEE, Pasadena, CA, USA, 2023).
Gomaa, A., Abdelwahab, M. M. & Abo-Zahhad, M. Efficient vehicle detection and tracking strategy in aerial videos by employing morphological operations and feature points motion analysis. Multimed. Tools Appl. 79, 26023–26043. https://doi.org/10.1007/s11042-020-09242-5 (2020).
Gomaa, A., Abdelwahab, M. M. & Abo-Zahhad, M. Real-time algorithm for simultaneous vehicle detection and tracking in aerial view videos. In 2018 IEEE 61st International Midwest Symposium on Circuits and Systems (MWSCAS), 222–225. https://doi.org/10.1109/MWSCAS.2018.8624022 (IEEE, Windsor, ON, Canada, 2018).
Gomaa, A. Advanced domain adaptation technique for object detection leveraging semi-automated dataset construction and enhanced YOLOv8. In 2024 6th Novel Intelligent and Leading Emerging Sciences Conference (NILES), 211–214. https://doi.org/10.1109/NILES63360.2024.10753164 (IEEE, Giza, Egypt, 2024).
Gomaa, A. & Abdalrazik, A. Novel deep learning domain adaptation approach for object detection using semi-self building dataset and modified YOLOv4. World Electr. Veh. J. 15, 255. https://doi.org/10.3390/wevj15060255 (2024).
Abdalrazik, A., Gomaa, A. & Afifi, A. Multiband circularly-polarized stacked elliptical patch antenna with eye-shaped slot for GNSS applications. Int. J. Microw. Wirel. Technol. 16, 1229–1235. https://doi.org/10.1017/S175907872400045X (2024).
Abdalrazik, A., Gomaa, A. & Kishk, A. A. A wide axial-ratio beamwidth circularly-polarized oval patch antenna with sunlight-shaped slots for GNSS and WiMAX applications. Wirel. Netw. 28, 3779–3786. https://doi.org/10.1007/s11276-022-03093-8 (2022).
Gomaa, A., Afifi, A. & Abdalrazik, A. A dual-band wide axial-ratio beamwidth circularly-polarized antenna with V-shaped slot for L2/L5 GNSS applications. In 2024 6th Novel Intelligent and Leading Emerging Sciences Conference (NILES), 119–122. https://doi.org/10.1109/NILES63360.2024.10753263 (IEEE, Giza, Egypt, 2024).
Hassan, O. et al. Real-time driver drowsiness detection using transformer architectures: a novel deep learning approach. Sci. Rep. 15, 17493. https://doi.org/10.1038/s41598-025-02111-x (2025).
Liu, Z. et al. A convnet for the 2020s. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 11976–11986 (2022).
Ding, X., Zhang, X., Han, J. & Ding, G. Scaling up your kernels to 31x31: Revisiting large kernel design in cnns. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 11963–11975 (2022).
Guo, M.-H., Lu, C.-Z., Liu, Z.-N., Cheng, M.-M. & Hu, S.-M. Comput. Vis. Media 9, 733–752 (2023).
Rao, Y. et al. Hornet: Efficient high-order spatial interactions with recursive gated convolutions. Adv. Neural Inf. Process. Syst. 35, 10353–10366 (2022).
Lee, Y., Kim, J., Willette, J. & Hwang, S. J. Mpvit: Multi-path vision transformer for dense prediction. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 7287–7296 (2022).
Wang, W. et al. Internimage: Exploring large-scale vision foundation models with deformable convolutions. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 14408–14419 (2023).
Qi, Y., He, Y., Qi, X., Zhang, Y. & Yang, G. Dynamic snake convolution based on topological geometric constraints for tubular structure segmentation. In Proceedings of the IEEE/CVF international conference on computer vision, 6070–6079 (2023).
Zhang, H. et al. Dino: Detr with improved denoising anchor boxes for end-to-end object detection. arXiv preprint arXiv:2203.03605 (2022).
Zong, Z., Song, G. & Liu, Y. Detrs with collaborative hybrid assignments training. In Proceedings of the IEEE/CVF international conference on computer vision, 6748–6758 (2023).
Zhao, Y. et al. Detrs beat yolos on real-time object detection. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 16965–16974 (2024).
Ge, Z., Liu, S., Wang, F., Li, Z. & Sun, J. Yolox: Exceeding yolo series in 2021. arXiv preprint arXiv:2107.08430 (2021).
Feng, C., Zhong, Y., Gao, Y., Scott, M. R. & Huang, W. Tood: Task-aligned one-stage object detection. In 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 3490–3499 (IEEE Computer Society, 2021).
Gu, A., Goel, K. & Ré, C. Efficiently modeling long sequences with structured state spaces. arXiv preprint arXiv:2111.00396 (2021).
Gu, A. et al. Combining recurrent, convolutional, and continuous-time models with linear state space layers. Adv. Neural Inf. Process. Syst. 34, 572–585 (2021).
Smith, J. T., Warrington, A. & Linderman, S. W. Simplified state space layers for sequence modeling. arXiv preprint arXiv:2208.04933 (2022).
Zhu, L. et al. Vision mamba: Efficient visual representation learning with bidirectional state space model. arXiv preprint arXiv:2401.09417 (2024).
Liu, Y. et al. Vmamba: Visual state space model. Adv. Neural Inf. Process. Syst. 37, 103031–103063 (2024).
Huang, T. et al. Localmamba: Visual state space model with windowed selective scan. In European Conference on Computer Vision, 12–22 (Springer, 2024).
Yu, W. & Wang, X. Mambaout: Do we really need mamba for vision? In Proceedings of the Computer Vision and Pattern Recognition Conference, 4484–4496 (2025).
Wang, Z., Li, C., Xu, H., Zhu, X. & Li, H. Mamba yolo: A simple baseline for object detection with state space model. Proc. AAAI Conf. Artif. Intell. 39, 8205–8213 (2025).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 5998–6008 (2017).
Chen, J. et al. Run, don’t walk: chasing higher flops for faster neural networks. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 12021–12031 (2023).
Ren, S., He, K., Girshick, R. & Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. Adv. Neural Inf. Process. Syst. 28, 91–99 (2015).
Jocher, G., Chaurasia, A. & Qiu, J. Ultralytics yolo: Software for object detection. version 8.0. 0, january 2023.
Wang, C.-Y., Yeh, I.-H. & Mark Liao, H.-Y. Yolov9: Learning what you want to learn using programmable gradient information. In European conference on computer vision, 1–21 (Springer, 2024).
Wang, A. et al. Yolov10: Real-time end-to-end object detection. Adv. Neural Inf. Process. Syst. 37, 107984–108011 (2024).
R. K. M. H. YOLOv11: An overview of the key architectural enhancements. arXiv preprint arXiv:2410.17725 (2024).
Funding
This work was supported by the Sci-Tech Innovation 2030–Major Project for ‘Brain Science and Brain-Like Research’ (grant no. 2021ZD0201904) and the Capacity Building Project of Autonomous Region-level Scientific and Technological Innovation Platform (grant no. Gui Ke LT2600640001).
Author information
Authors and Affiliations
Contributions
X.Q. conceived the experiment(s), Q.Q., X.L., C.D., and W.W. conducted the experiment(s), L.P., H.L., Y.G., and J.Z. analysed the results. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qin, X., Qian, Q., Li, X. et al. ScaleMamba-YOLO: a multi-scale MambaYOLO for medical object detection. Sci Rep 16, 10839 (2026). https://doi.org/10.1038/s41598-026-37258-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37258-8