Abstract
As a crucial lymphocyte subset, γδT cells exhibit tumoricidal properties without the constraint of major histocompatibility complex restriction. In recent years, the in vitro expansion of human peripheral blood-derived γδT cells has emerged as a promising candidate for adoptive immune cell therapy (ACT). However, given the limitations in accessing and sourcing human peripheral blood, it remains challenging to expand γδT cells to the magnitude required for clinical cancer immunotherapy. Therefore, the objective of this study was to establish an innovative approach for the in vitro differentiation of γδT cells from human embryonic stem cells (hESCs). In present study, by simulating the in vivo developmental process of γδT cells, we established stroma cell-free differentiation approach with three stages under 3D suspension culture in conjunction with using hypoxic conditions. Under our protocol, around 39% of CD34+CD31+ hematopoietic endothelial cells were induced in cell spheroids at first stage under hypoxic conditions, and over 99% of suspended/floating cells from cell spheroids were early CD43+ hematopoietic progenitor cells (HPCs), and over 90% of CD45+ HPCs were produced at the second stage, demonstrating the homogeneity of cell population and its high hematopoietic potential. Eventually over 40% of mature CD45+CD3+TCRγδ+ T cells were generated at the final stage, and these γδT cells exerted cytotoxicity against hepatoma cancer cells and simultaneously significantly inhibited their proliferation. RNA sequencing analysis revealed that the differentiation process of hESC-derived γδT cells was transcriptomically similar to those of in vivo courses. Therefore, our findings ultimately offer a novel technique to address the challenge of sourcing γδT cells for ACT therapies.
Similar content being viewed by others
Introduction
Cancer is one of the major fatal diseases worldwide because of its high incidence and mortality1. Therefore, developing new antitumor strategies for reducing the mortality rates and improving patients’ quality of life are urgently needed. Among various emerging therapeutic strategies, allogeneic γδT cell immunotherapy has garnered considerable attention2. γδT cells recognize target cells in an HLA-independent manner, and they release relatively fewer cytokines, resulting in a lower probability of developing cytokine release syndrome (CRS)3,4. Therefore, the transfer therapy of allogeneic γδT cells has good clinical value and advantages in tumor immunotherapy. At present, immunotherapy based on peripheral blood-derived γδT cells has received much attention, and clinical trials have been conducted5. Since the proportion of γδT cells in adult peripheral blood mononuclear cells is only a few percent or less6, the difficulty in expanding cytotoxic γδT cells from peripheral blood to sufficient numbers is a major limiting factor for allogeneic γδT cell therapy in most cases7.
Human pluripotent stem cells (hPSCs), including human embryonic stem cells (hESCs) and human induced pluripotent cells (hiPSCs) have the capacity of self-renewing and the differentiation potential into various cell types8,9. In vitro generation of mature γδT cells from hPSCs can overcome the limitations of their clinical applications, and support large-scale production. Timmermans et al.10 showed that TCRγδ+ T cells could be generated in vitro from hESC-derived CD34highCD43low cells with the co-culture of OP9 stromal cells. Also, Chang et al.11 reported that hiPSCs derived from skin biopsy were capable of producing T lymphocyte populations with a broad TCR repertoire, including hiPSC-derived γδT and αβT cells. Nobuyuki Murai et al.12 induced iPSCs to differentiate into CD34+CD43+ HPCs, and then co-cultured with OP9-DLL1 stromal cells to further induce into γδT cells. Soo-Been Jeon et al.13 investigated whether hPSCs could generate γδT cells with pivotal transcriptional factors via hemogenic Endothelium. While these existing protocols successfully demonstrated the feasibility of in vitro γδT-cell generation, they are collectively constrained by significant technical limitations. These include complex systems reliant on OP9 stromal support, which introduces contamination risk, and prolonged differentiation periods often exceeding 35 days. Furthermore, the use of iPSC-derived starting material in certain protocols raises specific safety and clinical translation risks.
Previously, we differentiated hESCs into CD34+CD31+ hematopoietic endothelial cells (HECs) in vitro using a cytokine cocktail14,15, and these hematopoietic endothelial cells were shown to give rise to CD43-positive HPCs. Therefore, in the present study, a feeder-free three-dimensional differentiation system for hESCs to differentiate towards γδT cells in vitro was established by mimicking the in vivo developmental process of γδT cells. Under our new optimized conditions, hESCs could be induced to differentiate into CD31+CD34+HECs, which further led to the induction of CD43-positive HPCs, and eventually into CD45+CD3+TCRγδ+ T cells. Most of our differentiated cells were CD4−CD8α− double negative (DN) and did not express αβT cell marker TCRαβ, which are phenotypically similar to γδT cells in human peripheral blood. In addition, we investigated the mechanisms regulating the differentiation process and function of γδT cells.
Materials and methods
hESC culture and maintenance
The hESCs, H9 cells (WA09), were obtained from WiCell Research Institute (Madison, WI, USA) with Material Transfer Agreements (No.: 19-W0512, 24-W0162 and 24-W0163), and cultured and passaged on the feeder cells of mouse embryonic fibroblasts (MEF). The culture medium was DMEM-F12 (Gibco) basal medium supplemented with 20% knockout ™ SR(Gibco), 4 ng/ml fibroblast growth factor bFGF (Peprotech), 100 mM L-glutamine (Gibco), 1% MEM non-essential amino acids solution (ThermoFisher), 0.0007% b Mercaptoethanol. The medium was changed daily.
Differentiation to γδT cells from hESCs
Cell aggregates of hESCs were obtained by treating hESC cultures at 70%−80% confluency with Gentle Cell Dissociation Reagent (STEMCELL, 07174). At day 0, the medium was completely replaced by STEMdiff™ APEL™2 Medium (STEMCELL, 05275) supplemented with 3 µM CHIR99021 (Selleckchem, S1263), 10 ng/mL BMP4 (Peprotech, 120-05), 2 ng/mL Activin A (Peprotech, 120 − 14) and 10 µM Rock inhibitor (Y-27632, Selleckchem, S1049). At day 2, the medium was replaced by STEMdiff™ APEL™2 supplemented with 40 ng/mL VEGF (Peprotech, 100 − 20). At day 3, the medium was replaced by STEMdiff™ APEL™2 supplemented with 40 ng/mL VEGF and 20 ng/mL bFGF (Peprotech, 100-18B). At day 6, the medium was replaced by STEMdiff™ APEL™2 supplemented with 50 ng/mL VEGF, 20 ng/mL bFGF, 10 ng/mL Flt3L (Peprotech, 300 − 19), 10 ng/mL IL-7 (Peprotech, 200-07), 20 ng/mL SCF (Peprotech, 300-07) and 4 µM SB431542 (Selleckchem, S1067). At day 10, the medium was replaced by MEM-α (Gibco, 12561049) supplemented with 20% FBS (Excell Bio, FND500), 1% GlutaMAX (Thermo Fisher, 35050061), 30 µM MAP (Sigma, A8960-5G), 10 ng/mL SCF, 5 ng/mL IL-7, 5 ng/mL Flt3L and 10 ng/mL DLL4 (Peprotech, 140-07). Then, half of the medium was changed every other day. Cell differentiation was conducted under hypoxic conditions (5% O2, 5% CO2) for the first six days (Day 0 to Day 6). Subsequently, the cultures were maintained under normoxic conditions (ambient O2, 5% CO2) from Day 6 until the termination of the protocol on Day 25.
qRT-PCR
To test the expression of genes at the different stages during the differentiation of hESC-derived γδT cells, total RNAs were extracted using Trizol reagent (AKARA, 9109). Total RNAs were reverse transcribed to cDNAs using PrimeScript II 1st Strand Synthesis Kit (TAKARA, RR036B) with oligo-dT primers according to the manufacturer’s instructions. qRT-PCR was performed by using ChamQ Universal SYBR qPCR Master Mix (Thermo Fisher, A25742) on QuantStudio (Applied Biosystems Life Technologies, Thermo Fisher Scientific). With GAPDH as internal parameter, the relative mRNA expression of samples was calculated by the 2-ΔΔCt method. The primers used were listed in Table S1.
Immunofluorescence assays
The cells were inoculated and cultured in 12 well plates with cell slides which were placed 1 days before the inoculation, then fixed with 4% paraformaldehyde (PFA, biosharp) at room temperature for at least 15 min, and permeated with 0.5% Triton X-100 (Solarbio) for 20 min. Next the cells were blocked in goat serum (Boster) for 1 h at room temperature, then incubated with primary antibodies at 4 °C overnight. Next day, after rewarming at 37 °C for 1 h and washing, the cells were incubated with secondary antibodies at room temperature in dark for 2 h. The cells were stained with DAPI (Beyotime) to exclude dead cells. The images were collected by Zeiss 880 multiphoton laser scanning microscope or single photon confocal microscope (Nikon). The antibodies used were listed in Table S2.
Flow cytometry analysis
The cells were dissociated into single cells using TrypLE ™ Express Enzyme (1X) (Gibco), and the single cell suspension was collected and washed with PBS once for immunostaining. For the staining of cell surface markers, the cells were directly stained and incubated at 4 °C in dark for 30 min, and the dead cells were eliminated by DAPI or 7AAD (BD) staining. Flow cytometry was performed with BD Celesta and BD Fortessa. FolwJo (V10) software was used for data analysis. The antibodies used were listed in Table S2.
Transcriptome sequencing (RNA-seq) and data analysis
The cells were collected to extract RNAs during the differentiation of hESCs to γδT cells in triplicate for each group. 100 ng of total RNAs was used to prepare sequencing libraries using Ligation Mediated Sequencing (LM-Seq) protocol (21) and quantified with Qubit 2.0 fluorometer (ThermoFisher). Final cDNA libraries were quantitated with Agilent 2100 bioanalyzer, multiplexed and loaded at final concentration of 2.5 nM, and sequenced as single reads on the Illumina NovaSeq 6000 (Illumina).
R software (version 4.1.3; https://www.r-project.org/) and Bioconductor software package (http://www.bioconductor.org/) were used to correct and analyze the original data. RNA-seq data processing and normalization were performed using DESeq2 packages. Principal component analysis (PCA) was used to verify the repeatability of the GSE102485 data. The standard of statistical significance was |log2FC| > 2 and adjusted p < 0.01. DEGs were visualized in Volcano map and Heatmap based on “ggplot2” software package. TBtools software and DAVID database were used to test the statistical enrichment of differential expression genes in KEGG pathways and Gene Ontology. To construct PPI networks for up-regulated DEGs at each time point, we utilized Search Tool for Retrieval of Interacting Genes/Proteins (STRING) database. Up-regulated DEGs were input into STRING database, which provides comprehensive information on known and predicted protein interactions. Original data were uploaded to Gene Expression Omnibus database (GSE301057).
Functional analysis of hESC-derived γδT cells
The γδT cells were stimulated for 6 h in the presence of 2 µL Phorbol 12-myristate 13-acetate (PMA), then the cells were stained with anti-human CD45-APC (BioLegend, 304014) antibody. After staining for surface markers, FOXP3/Transcription factor staining buffer set (Invitrogen) was used to fix and perforate cells. Subsequently, the cells were stained with antibodies of anti-human TNF-α (Biolegend, 502912), anti-human IL-2 (Biolegend, 500347), anti-human IFN-γ (Biolegend, 502509), anti-human Perforin and anti-human Granzyme B (BioLegend, 372205). Finally, the cells were analyzed with BD FACS Verse. Antibodies used were listed in Table S2.
In vitro cytotoxicity assay
To determine the cytotoxicity of γδT cells (effector), several human cancer cell lines (target) were used to perform killing assays, including Jurkat cells (acute T cell leukemia cell line), HepG2 cells (hepatocellular carcinoma cell line), Huh-7 cells (hepatocellular carcinoma cell line), and HeLa cells (cervical cancer cell line). The cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM), formulated with high glucose (Gibco, 11965092), and supplemented with 10% (v/v) Fetal Bovine Serum (Gibco). Cancer cells were pre-stained with 2 µM CFSE. Effector cells and target cells were co-cultured at the ratio of 2:1 at 37 °C for 48 h. The cell apoptosis of cancer cells was analyzed using flow cytometry and DAPI (0.2 µg/mL) staining. As a control, normal 293T cells (human embryonic kidney cells) was used to test the cytotoxicity of γδT cells against normal cells.
Differentiation of human umbilical cord blood into γδT cells using co-culture with OP9 cell
Human umbilical cord blood was collected from donors with normal term deliveries after obtaining informed consent, and CD34+ hematopoietic stem and progenitor cells (HSPCs) were isolated using magnetic bead-based sorting and subsequently utilized for the differentiation of γδT cells. On the day prior to the experiment, OP9 stromal cell line (OP9) cells were seeded in 6-well plate to reach 70–80% confluency and were treated with mitomycin C before use. Expanded HSPCs suspension was then applied onto a 0.4 μm Millicell transwell insert, ensuring even distribution of the cells. The insert was placed into 6-well plate with OP9 cells, forming co-culture system with 1 mL of differentiation medium of γδT cells in the lower chamber. The medium was replaced daily without disturbing the upper chamber, maintaining an air-liquid interface throughout. Every 4–5 days, the insert was transferred to fresh OP9 cell-seeded 6-well plate. After 3 weeks, differentiation medium was replaced with expansion medium of γδT cells, and the culture was continued for an additional week with the same feeding regimen. Differentiation medium consisted of RPMI1640 basal medium supplemented with 4% B-27™ Supplement, 1% GlutaMAX, 30 µM Magnesium Ascorbyl Phosphate (MAP), 10 ng/mL SCF, 5 ng/mL IL-7, 5 ng/mL Flt3L and 10 ng/mL Delta-like 4 (DLL4) (all growth factors from PeproTech). Expansion medium contained the same basal components with additional 10 µM zoledronic acid (ZOL) and 10 ng/mL IL-2.
Statistical analysis
All experimental procedures included triplicate samples and were independently replicated a minimum of three times. Data are presented as mean SEM (Standard Error of the Mean). Statistical significance was determined using a two-tailed Student’s t-test. *P < 0.05, **P < 0.01, and ***P < 0.001 versus the control group, ns, not significant.
Results
Generation of γδT cells using optimized culture conditions
We previously established a protocol for the differentiation of hESCs to hematopoietic endothelial cells (HECs) under hypoxic conditions14,15. Therefore, based on that work, we developed a new three-stage differentiation system to differentiate hESCs into γδT cells by mimicking the in vivo developmental process of γδT cells (Fig. 1A). In the first phase (the differentiation of HECs, from day 0 to day 6) under hypoxic conditions, hESCs were differentiated into the spheroids of mesodermal progenitor cells (MPCs) under the induction with CHIR99021, Activin A and BMP4 for 2 days. After that, we switched to the medium supplemented with VEGF and bFGF for 4 days, and MPCs were further differentiated into the spheroids of HECs (Fig. 1A). In the second phase (the differentiation of HPCs, from day 6 to day 10), the medium was changed by supplementing additional SB43154, stem cell factor (SCF), FLT3 ligand (FLT3L), and interleukin-7 (IL-7) for differentiating into HPCs which are generated by endothelial-to-hematopoietic transition (EHT), and we found that large numbers of individual HPCs were produced from the spheroids of HECs on day 10 (Fig. 1A). In the third phase (the differentiation of T cells, from day 10 to day 25), HPCs were further differentiated with MEMα medium containing fetal bovine serum (FBS), ascorbic acid, SCF, IL-7, FLT3L, and DLL4, and ultimately HPCs were induced to differentiate into mature γδT cells (Fig. 1A).
Stroma-free differentiation of hESCs into HECs under optimized conditions for generating γδT cells. (A) Schematic diagram of the protocol for the differentiation of hESCs into γδT cells (the upper panel), representative phase contrast images of hESCs (day 0) and their derivatives (day 6, day 10, and day 25) (the lower panels). Scale bars, 100 μm. (B) The dynamic changes of cell morphologies from day 1 to day 6 during the differentiation of hESCs. Scale bars, 100 μm. (C, D) Representative immunofluorescence analysis of HECs at day 6 for CD34 (C) and CD31 (D), DAPI was used for nuclear stain. Scale bars, 100 μm (left) and 20 μm (right). (E) The dynamic changes of HEC differentiation from hESCs at different time points. Total live cells were shown in 7AAD-gate, hESCs (H9 cells) were used for the comparison.
Differentiation of HECs from hESCs
The hematopoiesis of human embryos is a complex regulatory process, which undergoes the formation of primitive streak, the induction of mesoderm, and the development of endothelium and hematopoiesis16. Hematoendothelial differentiation could be rapidly induced from MPCs at the stage of HECs derived from hESCs. In our previous study, hESCs were differentiated into HECs by mimicking human embryonic hematopoietic process14,15. hESCs were cultured in 2D system using mouse embryonic fibroblasts (MEF) as feeder layers (Fig. S1A). The hESCs exhibited typical clonal morphology and highly expressed pluripotency-related markers SSEA4, TRA-1-60 and NANOG17 (Fig. S1B,C). The hESCs were observed to form uniformly sized cell spheres under the microscopy during the differentiation process, and vacuolar structures were formed in the spheroids 6 days after the differentiation, and the average diameter of cell spheres was about 500 μm (Fig. 1B). Next, we analyzed whether HECs were produced in the spheroids at day 6 by immunofluorescence staining, and the results showed many CD34+ and CD31+ cells in cell spheres, respectively (Fig. 1C and D). To further determine the proportions of cells, the analysis of flow cytometry was performed during the first 8 days of the differentiation, and the results showed that almost hESCs highly expressed SSEA-4 and OCT4 at day 017 (Fig. S1B), KDR-expressing mesodermal progenitor cells were 22.3% at day 4, decreased to 0.57% at day 8, while CD31+CD34+ HECs were gradually generated and reached to 39.2% (Fig. 1E).
Subsequently, we performed qRT-PCR analysis to determine the dynamic changes of specific genes at the different stages during the first 8 days of the differentiation, and the results showed that pluripotent genes NANOG, SOX2 and OCT4 were highly expressed in hESC, then their expression levels were rapidly decreased at day 2 after the differentiation (Fig. S1D). Mesodermal markers Brachyury and KDR/VEGFR-2 which is associated with embryonic hematopoiesis18 were significantly up-regulated from day 2 to day 4 after the differentiation, and then gradually decreased (Fig. 2A). Mature endothelial junctional markers including VE-Cadherin and CD31 exhibited significant upregulation from days 4 and 6 after the differentiation, confirms the successful transition of the cell population from an early progenitor stage into a definitive endothelial phenotype. Second, and critically, this definitive population simultaneously acquired an arterialized identity, a required step for initiating T-cell fate. This is directly substantiated by the concurrent and significant upregulation of the transcriptional factor SOX1719 and, most importantly, the Notch ligand DLL4 on Day 6 (Fig. 2A). Concurrently, the expression of transcription factors ERG and TIE1, associated with early hematopoiesis20, peaked at day 6.
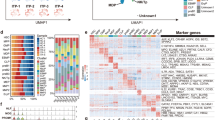
Characterization of HEC differentiation from hESCs. (A) The relative expression levels of key genes were determined by qRT-PCR from day 0 to day 8 during HEC differentiation. (B, C) Volcano maps of DEGs at day 2 (Day2 VS H9 cells) (B) and day 6 (Day6 VS Day2) (C) using differential expression analysis, P < 0.01, |log2 fold change|>2. (D) Heat map showed the transcriptional changes of key DEGs during HEC differentiation. (E, F) The bubble plot showed the significantly enriched GO terms related to biological process identified from upregulated DEGs at days 2 (E) and 6 (F) after the differentiation of hESCs. The y-axis represented GO term, and the x-axis represented gene ratio. The size of each bubble corresponded to the number of DEGs involved in the terms, while the color intensity reflected the -log10 transformed p-value. Terms with p-values less than 0.05 were considered statistically significant. The KEGG software from the Kanehisa laboratory was used to analyze the data (www.kegg.jp/kegg/kegg1.html).
Subsequently, we conducted transcriptome sequencing (RNA-seq) on hESCs, and cell spheres at days 2 and 6 after the differentiation. With the objective of unraveling the molecular mechanisms that govern hematopoietic-endothelial differentiation, we performed differentially expressed genes (DEGs) analysis. This rigorous approach enabled us to identify and quantify the alterations in gene expression that occur during the induction of HECs, providing valuable insights into the underlying transcriptional dynamics of this critical developmental transition. DEGs between the two groups was first performed by R package DESeq2, and the screened DEGs were demonstrated by volcano plots (Fig. 2B and C). Compared with H9 cell group, 2779 significantly different genes were screened in cell sphere group at day 2, and the results showed 1175 genes were significantly up-regulated (Fig. 2B). Whereas, compared with cell spheres at day 2, 3794 significantly different genes were screened in cell sphere group at day 6, and results exhibited that 2174 genes were significantly up-regulated (Fig. 2C). These results indicated that the changes of gene expression were considerable during the differentiation from hESCs to HECs. The heatmap showed that pluripotency genes of hESCs17 NANOG, SOX2, and CDH3 were highly expressed in H9 cell group. Wnt family member WNT5A, mesoderm development-related transcription factors MESP1 and FOXA2, and mesoderm-to-endoderm and hematopoietic development-related genes21 CXCR4, CDX4, and APLNR were expressed at the highest level in cell spheres at the 2nd day of the differentiation (Fig. 2D). Twenty genes related to hematopoietic endothelial development22, including CD34, KLL4, SOX17, ERG, and TIE1, were up-regulated at day 6 after the differentiation (Fig. 2D). This trend further demonstrated that we successfully induced hESCs to differentiate toward mesoderm, and that cells had hematopoietic endothelial properties at day 6.
To further investigate the differentiation mechanism of hESCs into HECs, we performed GO functional annotation of DEGs that were upregulated at days 2 and 6 after the differentiation. Our analysis revealed that DEGs at day 2 were associated with biological processes such as mesoderm formation and development, blood vessel development, and hematopoiesis (Fig. 2E). Furthermore, DEGs identified at day 6 were linked to more processes, including embryonic hematopoiesis, vascular endothelial development, T cell differentiation, platelet activation, erythrocyte differentiation, and endothelial cell migration (Fig. 2F).
Differentiation of HPCs from HECs
The differentiation of HPCs was induced from hESC-derived HECs, and four days after hypoxic hematopoietic induction (i.e., day 10), hollow structure of HEC spheroids became more and more evident, accompanied by the emergence of significant number of individual floating cells and cell clusters coming out from HEC spheroids (Fig. 3A and C). Then we used flow cytometry to determine whether cells within HEC spheroids at day 10 were still HECs. Co-immunostaining results revealed that 39.1% of cells within HEC spheroids were HECs with co-expression of CD31 and CD34, which was comparable to the findings at day 8 (Figs. 1E and 3B). Furthermore, the presence of CD45+, CD43+, and CD7+ cells was also observed (Fig. 3B). These results collectively indicated the existence of early hematopoietic cells at multiple differentiation stages within HEC spheroids.
The generation of HPCs from hESC-derived HECs. (A) Representative images of cell spheroids at day 10 after the differentiation of hESCs. Scale bars, 100 μm. (B) Flow cytometric analysis of the proportions of CD31+CD34+ HECs, CD43+CD45+ HPCs and CD7+ T progenitor cells in cell spheroids at day 10 after the differentiation of hESCs. (C) Representative images of single floating cells at day 10 after the differentiation of hESCs. Scale bars, 100 μm. (D) Flow cytometric analysis of the proportions of CD31+CD34+ HECs, CD43+CD45+ HPCs and CD7+ T progenitor cells in the single floating cells at day 10 after the differentiation of hESCs. (E) Immunofluorescence analysis for CD45 (green) and CD34 (red) of HPCs at day 10 after the differentiation of hESCs. DAPI (blue) was used for nuclear stain. Scale bars, 100 μm. (F) Volcano map showing DEGs in differentiated cells at day 10 (Day10 VS Day6) using differential expression analysis, P < 0.01, |log2 fold change|>2. (G) Heat map showing the transcriptional changes of key DEGs during HPC differentiation. (H) The bubble plot showed the significantly enriched GO terms related to biological process identified from upregulated DEGs at days 10 after the differentiation of hESCs. The KEGG software from the Kanehisa laboratory was used to analyze the data (www.kegg.jp/kegg/kegg1.html).
CD45 and CD43 are markers for HPCs23, while CD7 is a marker for pre-lymphoid and mature T-cells24. Thus, in order to detect the cell type of the round floating cells in the culture medium using flow cytometry the results showed that 22.4% of the floating cells were CD43+CD34+ cells, with a high percentage of 99.4% of CD43+ HPCs (Fig. 3D), which demonstrated the homogeneity of this cell population and its high hematopoietic potential. Meanwhile, we observed 90.1% CD7+ cells and 10.2% CD45+ cells in suspension cell population, but CD5+ cells were not detected (Fig. 3D), indicating that this cell population had the potential to differentiate into T cell lineage. Immunofluorescence staining further showed that two hematopoietic markers, CD34 and CD45, were highly expressed in HPCs (Fig. 3E).
To more clearly elucidate the mechanism underlying the differentiation of HECs into HPCs, we conducted RNA-seq analysis on floating cells at day 10 after the differentiation. Firstly, the differential expression analysis between days 10 and 6 was performed by R package DESeq2, and DEGs were presented by volcano plot (Fig. 3F). Compared with Day 6 group, there were 5218 genes with significantly differential expression in floating cells in Day 10 group, this result suggested that gene expression profile of cells was significantly changed during the differentiation of HECs to HPCs. Subsequently, in order to explore the gene expression alterations during the differentiation process of HPCs, based on the differential expression analysis, we screened the genes related to endothelium, hematopoiesis and immunity that have been extensively reported in the literature, and the heatmap showed that the expression of HEC-related DLL4, CD34, SOX17, ERG and CDH5 genes was significantly down-regulated in suspension cells at day 10 (Fig. 3G), and the expression of markers specifically associated with HPCs, such as CD43/SPN and CD45/PTPRC, was significantly up-regulated in suspension cells at day10 (Fig. 3G). Next, to explore the mechanism underlying the differentiation of HECs into HPCs, we conducted GO functional annotation on DEGs that were upregulated at day 10. DEGs at day 10 were associated with platelet aggregation, inflammatory response, immune response, and T cell proliferation and differentiation. We also found that these up-regulated genes were significantly enriched in Type I interferon signaling pathway, which can stimulate T cell differentiation and functional development, and play a role in both primitive and adaptive immune responses25 (Fig. 3H).
Differentiation of HPCs into γδT cells
We next examined whether HPCs could be induced to differentiate into γδT cells, RPMI1640 medium or MEM-α medium in combination with cytokines were used to induce hematopoietic cells to differentiate into T cells24,26. We transferred floating cells and cell spheres at day 10 after the differentiation into new cell culture plate with new culture medium for 15 days, then collected floating cells, and performed precise phenotypic analysis of these cells by flow cytometry (Fig. 4A and D). 15 days after the induction, we clearly observed that the proportions of CD45+ cells produced in RPMI1640 medium groups with floating cells (R-HECs) or with both floating cells and HEC spheres (R+HECs) were 47.9% and 66.5%, respectively (Fig. 4A and B), and the use of MEM-α medium with floating cells (M-HECs) or with both floating cells and HEC spheres (M+HECs) produced higher proportions of CD45+ cells with 93.8% and 87.3, respectively (Fig. 4A and B). These results indicated that MEM-α medium was more beneficial to promote the differentiation towards hematopoietic cells in our differentiation system. In MEM-α group, when HPCs were co-cultured with both floating cells and HEC spheres (M+HECs), more CD3+TCRγδ+ cells could be induced with 42.4% in the proportion, which was significantly better than the other three groups (Fig. 4C and D). Based on the above results, we determined that co-culture of HPCs with both floating cells and HEC spheres in conjunction with the use of MEM-α culture medium was better strategy to induce the differentiation of hESC-derived HPCs into γδT cells. Therefore, in subsequent experiments, this differentiation method was used to ensure the efficient production of γδT cells.
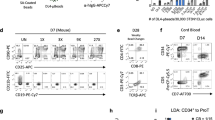
The generation of γδT cells from HPCs. (A, B) Flow cytometric analysis of the frequency of CD45+ T cells at day 15 under different combinations of RPMI1640 medium (R) or MEM-α medium (M) with floating cells (R/M-HECs) or with both floating cells and HEC spheres (R/M+HECs), gated on 7AAD− cells. (C, D) Flow cytometric analysis of the frequency of CD3+TCRγδ+ T cells at day 15 under different combinations of RPMI1640 medium (R) or MEM-α medium (M) with floating cells (R/M-HECs) or with both floating cells and HEC spheres (R/M+HECs), gated on CD45+ cells. (E) The dynamic changes of cell morphologies from day 15 to day 25 during the differentiation of hESCs. Scale bars, 100 μm. (F, G) Flow cytometric analysis of the frequency of CD45+ cells during the differentiation of HPCs towards T cells at the time points, gated on 7AAD− cells. (H, I) Flow cytometric analysis of the frequency of CD3+TCRγδ+ cells during the differentiation of HPCs towards T cells at different time points, gated on CD45+ cells.
Therefore, from day 10, floating HPCs and HEC spheres were transferred to MEM-α culture medium for further differentiation for additional 25 days. We found that cells displayed the rounded morphologies and had cell aggregations by phase-contrast microscopy (Fig. 4E), indicating that lymphoid cells were undergoing active proliferation at this time. Floating cells at days 15, 20 and 25 after the differentiation were detected by flow cytometry (Fig. 4F and I), and the results of co-staining with CD45, CD3 and TCRγδ showed that the proportions of CD45+ cells (i.e., lymphocytes) were significantly increased with the progression of the differentiation, reaching about 74.2% at day 25 (Fig. 4F and G). CD3+TCRγδ+ cells (i.e. mature γδT cells) began to appear at day 20 and reached 37.4% by day 25 (Fig. 4H and I). These results confirmed that HPCs could be successfully induced to differentiate into γδT cells. Remarkably, almost all of the detected CD3+TCRγδ+ cells were derived from CD45+ cell population, further demonstrating that hESC-derived HPCs could be differentiated into mature γδT cells.
In order to explore whether the expression of transcription factors and specific markers related to the differentiation of γδT cells was increased under specific conditions, qRT-PCR analysis was performed using single floating cells at days 10, 15, 20 and 25 after the differentiation (Fig. 5A). The expressions of megakaryocytic and erythroid transcription factors FLI1 and KLF1 were gradually decreased, and hematopoietic marker genes CD43 and RUNX1 were gradually downregulated during the differentiation process, and pro-T marker gene CD7 was also gradually downregulated. Meanwhile, the expressions of T cell development-related transcription factors27 NOTCH1, IL7R, TCF7, and BCL11B were increased at days 20 and 25 after the differentiation. Pan-hematopoietic marker CD45 was also gradually upregulated. We also detected high expression of NK cell marker CD56 gene at day 25 (Fig. 5A). These results indicated the proper differentiation of HPCs into γδT cells.
Characterization of γδT cell differentiation. (A) The relative expression levels of key genes were determined by qRT-PCR from day 10 to day 25 during the differentiation of γδT cells. (B) Flow cytometric analysis of the proportions of TCRγδ+TCRαβ+ cells, CD8a+CD4+ cells and CD56+CD41a+ cells in the floating cell at day 25 after the differentiation of hESCs. (C) Volcano map showed DEGs in differentiated cells at day 25 (Day25 VS Day10) using differential expression analysis, P < 0.01, |log2 fold change|>2. (D) The bubble plot showed the significantly enriched GO terms related to biological process identified from upregulated DEGs at days 25 after the differentiation of hESCs. (E) Heat map showed the transcriptional changes of key DEGs during the differentiation of hESCs towards T cells. The KEGG software from the Kanehisa laboratory was used to analyze the data (www.kegg.jp/kegg/kegg1.html).
Next, we further characterized the fully differentiated γδT cells at day 25 by flow cytometry, and the results showed that TCRαβ+ cells were only 2.79%, TCRγδ+ cells were 45.53% and CD8a−CD4− were 88.6%, and 20.2% of cells expressed CD56. Notably, megakaryocyte marker CD41a was barely expressed in cells at day 25 (Fig. 5B), these results are consistent with the phenotype of natural γδT cells28.
To more clearly delineate gene expression changes during the differentiation of HPCs into γδT cells, we collected single suspension cells at days 10, 15, 20, and 25 during the differentiation for RNA-seqs. Firstly, the differential expression analysis between days 25 and 10 was performed by R package DESeq2, and DEGs were presented by volcano plot (Fig. 3F). Compared with Day 10 group, there were 3764 genes with significantly differential expression in floating cells in Day 25 group, this result suggested that gene expression profile of cells was significantly changed during the differentiation of HPCs to γδT cells (Fig. 5C). Afterwards, we performed GO functional annotation of DEGs that were upregulated at day 25, and these DEGs were related to inflammatory response, immune response, T cell proliferation and differentiation, leukocyte migration and activation and other biological processes (Fig. 5D). We also enriched the positive regulation of cytokine production of IL-1B, IL-2, IL-6, IL-8, TNF-α, type I interferon and other processes (Fig. 5D). These cytokines play an important role in the activation and proliferation of γδT cells27, anti-tumor response, and inflammatory response. Subsequently, DEG at each stage were screened, and Heat map exhibited that the 10th day was the stage of pro-T development of HPCs, with CD34, SPN (CD43), CD7, RUNXI and other early hematopoietic and lymphoid lineage genes were significantly overexpressed, and then significantly down-regulated later (Fig. 5E), and that the 15th day was the transition stage from pro-T to mature γδT cells, with genes related to the activation and function of TNF, IFNB1, CD28 and other T cells were significantly overexpressed (Fig. 5E), and that the 20th day was the mature stage of γδT cells, with CD3E, PTPRC (CD45), IL6 and other mature γδT cell specific genes were highly expressed, and that the 25th day was the proliferation and activation stage of γδT cells, with transcription factors EGR3 and SOX13 related to the development of γδT cells, and genes of CD80, CD86, IL7R and IL15RA related to the proliferation and activation of T cells continued to be highly expressed (Fig. 5E). We also found that CD4, CD8A, CD5, ICAM1 (CD56) and other mature lymphoid lineage genes29 continued to be highly expressed from day 20 to day 25. These results demonstrated that we successfully generated γδT cells from hESC-derived HPC under our optimized culture conditions.
Next, we validated the above differentiation protocol using umbilical cord blood-derived HSPCs. Unexpectedly, only in the presence of feeder cells were CD34⁺CD43⁺CD45⁺ HSPCs able to differentiate into CD3⁺TCRγδ⁺ γδT cells (Figs. S2–S4). In in vitro differentiation of T cells, OP9 cells have been widely used due to their unique biological characteristics24. Briefly, CD34+ HSPCs were isolated from human umbilical cord blood by magnetic bead sorting, of which 65.0% were CD34 positive and almost all expressed CD45 and CD43 (Fig. S2). Next, we established an air-liquid culture system to induce γδT cell development by co-culturing HSPCs with OP9 cells via transwell culture (Fig. S3A,B). Specifically, when the confluence of OP9 cells reached 50% to 70% (Fig. S3D), we placed transwell inserts as up chamber into the well of OP9 cells as low chamber and added HSPCs uniformly over the inserts (Fig. S3C), thereby creating a well-defined air-liquid interface between the inserts and the medium (Fig. S3B). Consequently, by co-culturing umbilical cord blood-derived HSPCs with OP9 cells in combination with the induction of ZOL and IL-2, we successfully generated 72.1% CD3+TCRγδ+ γδT cells within 4 weeks (Fig. S4A), and cells were TCRαβ negative (Fig. S4B).
Functional validation of γδT cells
During the differentiation of HPCs into γδT cells (days 10–25), the expression changes of genes related to the function of γδT cells were determined by qRT-PCR (Fig. 6A). EGR1, EGR2, EGR3, SOX13 and TBET are the key transcription factors in the development of γδT cells30, and their expressions were significantly high at day 25 after the differentiation (Fig. 6A), indicating that our differentiation protocol could effectively promote the maturation of γδT cells. In addition, the expression of TNFSF11 (tumor necrosis factor ligand superfamily member 11), which is closely related to cytotoxic potential, was gradually up-regulated, and the expression level of ZAP70, a key gene for T cell activation31, was at the highest level at day 25. These results further supported the effectiveness of this differentiation protocol in promoting the maturation and activation of γδT cells.
Functional analysis of hESC-derived γδT cells. (A) The relative expression levels of key genes were determined by qRT-PCR from day 10 to day 25 during the differentiation of hESCs towards T cells. (B) Flow cytometry to detect cytokines TNF-α and IL-2 levels released by differentiated cells at 25 days following the stimulation with PMA and ionomycin for 6 h. (C, D) Phase-contrast images and flow cytometry for CFSE and DAPI staining of 293T cells after the co-culture for 48 h without effector cells (C) or with hESC-derived γδT cells (D). Scale bars, 100 mm in left panels. Flow cytometry was performed to determine the percentage of live (CFSE+ and DAPI−) or dead (CFSE+ and DAPI+) cells in target 293T cells. (E, F) Phase-contrast images and flow cytometry for CFSE and DAPI staining of HepG2 cells after the co-culture for 48 h without effector cells (E) or with hESC-derived γδT cells (F). Scale bars, 100 mm in left panels. Flow cytometry was performed to determine the percentage of live (CFSE+ and DAPI−) or dead (CFSE+ and DAPI+) cells in target HepG2 cells. (G, H) Phase-contrast images and flow cytometry for CFSE and DAPI staining of Huh-7 cells after the co-culture for 48 h without effector cells (G) ot with hESC-derived γδT cells (H). Scale bars, 100 mm in left panels. Flow cytometry was performed to determine the percentage of live (CFSE+ and DAPI−) or dead (CFSE+ and DAPI+) cells in target Huh-7 cells. (I, J) This figure displays the percentage of dead target cells (CFSE+ and DAPI+) and the percentage of overall target cells (CFSE+) following co-culture with γδT cells. The γδT effector cells were co-cultured with CFSE-labeled target cells (293T, HepG2, Huh7, HeLa, and Jurkat) at an Effector-to-Target (E: T) ratio of 2:1 for 48 h. Dead target cells were identified by DAPI positivity among the CFSE-positive population via flow cytometry.
It has been reported that γδT cells can directly kill tumor cells through Perforin and Granzyme B32, and also kill and inhibit tumor growth by secreting a variety of cytokines such as tumor necrosis factor alpha (TNF-α), interferon gamma (IFN-γ) and Interleukin-2 (IL2), or regulate the activity of other immune cells. To determine whether hESC-derived γδT cells released these factors, we exposed differentiated cells at day 25 to both PMA and ionomycin for 6 h to simulate signalings during the activation of T cells33. Subsequently, we collected these stimulated cells and used flow cytometry to detect cytokines TNF-α, IL-2, granzyme B, perforin, and IFN-γ. As shown in Fig. 6B and 30.4% of these CD45+ cells expressed TNF-α and 13.4% expressed IL-2. These results indicated that hESC-derived γδT cells were functionally similar to natural γδT cells. They could not only directly kill or inhibit tumors by secreting TNF-α, but also activate and enhance the activity of immune cells by releasing IL-2. However, the expressions of granzyme B, perforin and IFN-γ in hESCs-derived γδT cells were at low level (Fig. S5A), indicating that hESC-derived γδT cells had different characteristics than natural γδT cells or not fully mature.
We then investigated the ability of γδT cells to kill four different types of cancer cells in vitro, including Jurkat cells (an acute T cell leukemia cell line), HepG2 cells (a hepatocellular carcinoma cell line), Huh-7 cells (a hepatocellular carcinoma cell line), and HeLa cells (a cervical cancer cell line). These cell lines have different HLA types than hESCs used in this study. We used carboxyfluorescein diacetate succinimidyl ester (CFSE) to label cancer cells as target cells (T), and hESC-derived γδT cells as effector cells (E), then co-cultured with E and T at the ratio of 2:1. All cells were collected 48 h after the co-culture for flow cytometry, and dead cells were excluded by DAPI (4’,6-diamidino-2-phenylindole)staining. The results showed that there was little cell death in normal human 293T cells with or without effector cells, indicating that hESC-derived γδT cells had no obvious killing effect on human normal cells (Fig. 6C and D). In contrast, there were 26.1% and 23.4% of cell deaths in HepG2 cells and Huh-7 cells respectively under the influence of effector cells (Fig. 6E and H), indicating that γδT cells can specifically induce the death of these two liver cancer cells (Fig. 6I). Second, γδT cells significantly suppressed the proliferation of both liver cancer cell lines compared with the control (Fig. 6J). However, hESC-derived γδT cells demonstrated significant inhibition of proliferation yet only weak direct cytotoxicity against Jurkat and HeLa cells (Fig. S5B–E).
Transcriptome sequencing revealed the differentiation mechanism of hESC-derived γδT cells
To further validate the differentiation approach, transcriptome sequencing was performed to reveal the expression profiles of cells at day 0 (H9 cells), 2, 6, 10,15, 20 and 25 during the differentiation. For the analysis of the transcriptome data, Principal Component Analysis (PCA) was performed to examine the overall gene expression profiles at different differentiation stages (days 0, 2, 10, 15, 20, and 25). PCA results revealed distinct clustering of the samples according to the differentiation time points, with clear separation between early and late stages of the differentiation (Fig. 7A). Specifically, the samples at days 0 and 2 clustered closely together, suggesting a similarity in gene expression patterns during the initial differentiation phases. As the differentiation progressed, samples from days 6, 10, 15, 20, and 25 showed progressive shifts in gene expression profile (Fig. 7A), reflecting the dynamic nature of T cell differentiation. These results indicated a clear temporal progression in the molecular events underlying the differentiation process, with significant changes in gene expression as cells maturated into T cells. These findings highlighted the robustness of our differentiation method and the distinct molecular transitions at each stage of the differentiation (Fig. 7A).
Transcriptome analysis for the mechanism of the differentiation of hESCs into mature γδT cells in vitro. (A) Principal component analysis of γδT cell differentiation. The scatter plot showed the projection of the first two principal components (PC1 and PC2) at different time points. These points represented individual samples and were colored by time points. (B) The bubble plot showed the significantly enriched KEGG pathways identified from upregulated DEGs at days 2, 6, 10, and 25 after the differentiation of hESCs. The y-axis represented KEGG pathway, and the x-axis represented gene ratio. Pathways with p-values < 0.05 were considered significantly enriched. (C–F) PPI networks of up-regulated DEGs among cells from H9 cells to differentiated cells at day 2 (C), day 6 (D), day 10 (E) and day 25 (F) analyzed in STRING database. The KEGG software from the Kanehisa laboratory was used to analyze the data (www.kegg.jp/kegg/kegg1.html).
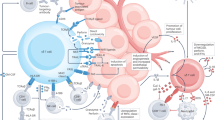
In order to investigate whether the differentiation mechanism of hESC-derived γδT cells was similar to that of human γδT cells, we performed KEGG pathway analysis for the following four stages of differential genes during the differentiation, compared significantly upregulated differential genes at day 2 with those at day 0, at day 6 with those at day 2, at day 10 with those at day 6, at day 25 with those at day 10 (Fig. 7B). KEGG pathway analysis showed that DEGs at day 6 were significantly enriched in several key signaling pathways closely related to stem cell proliferation and differentiation, such as PI3K-AKT, MAPK, and Hippo signaling pathways, and also enriched in the core pathways of mesoderm development, such as WNT and TGF-beta, which play a crucial role in embryo formation and organ development34. In addition, these DEGs were involved in signaling pathways that promote mesoderm development into blood and blood vessels, including BMP4 and Notch, which play an integral role in blood system generation and vascular network construction35. Notably, these genes were also enriched in inflammation-related TNF signaling pathway and the immune system-related Th17 and IL-17 pathways, suggesting that these differential genes also had a potential impact on inflammatory and immune responses (Fig. 7B). DEGs at day 10 were significantly enriched in calcium signaling pathway, which can regulate the proliferation, differentiation and maturation of stem cells during hematopoietic development and plays an important role in maintaining the normal function of the hematopoietic system19. We also concentrated on NOD-like receptors, leukocyte transendothelial migration (TEM), FC-γR-mediated phagocytosis, and inflammatory mediator regulation of TRP channels which is a kind of calcium channel and related to immune cell migration and cytokine release, and can promote the functional development of immune cells36. DEGs at day 25 were significantly enriched in signaling pathways related to cell proliferation and differentiation, such as PI3K-AKT, MAPK, Jak-STAT, Hippo and Ras. Jak-STAT has been reported to be involved in T cell development and immunomodulatory responses. Nod-like receptor, TEM, FC-γR-mediated phagocytosis, TNF, NF-κB, toll-like receptor and T cell receptor are all involved in immune response, lymphocyte differentiation and functional development37.
Furthermore, we analyzed the PPI networks of up-regulated DEGs in cells at different induction stages using the STRING database, and further analysis validated the associations between genes and biological functions (Fig. 7C and F). The results of PPI analysis showed that genes such as WNT3A, MSGN1, and TBXT were closely linked at day 2 of the induction (Fig. 7C), and were Hub Genes (Highly Connected Genes). WNT3A is essential for normal embryonic mesoderm development and the formation of caudal somites38, while MSGN1 regulates the expression of T-box transcription factors required for mesoderm formation and differentiation39, and TBXT is involved in transcriptional regulation of genes required for mesoderm formation and differentiation40. At day 6, PPI analysis showed that ESAM, CDH5 and CLDN5 were closely linked as Hub Genes at this stage (Fig. 7D). ESAM can mediate cell aggregation, most likely through homophilic molecular interactions. CDH5 associated with alpha-catenin to form a link to the cytoskeleton and works in conjunction with KRIT1 and MPP5 to establish and maintain proper endothelial cell polarity and vascular lumen integrity. CLDN5 plays a crucial role in the tight junction-specific sealing of the intercellular space. At day 10, PPI analysis showed that genes such as CCR2, CSF2 and PTPRC were closely linked and were Hub Genes at this stage (Fig. 7E). CCR2 regulates the expression of T-cell inflammatory cytokines and influences T-cell differentiation, specifically promoting the differentiation of T-cells into T-helper 17 (Th17) cells during the inflammation41. CSF2 is a cytokine that promotes the growth and differentiation of hematopoietic precursor cells from multiple lineages, including granulocytes, macrophages, eosinophils, and erythrocytes42. PTPRC is a protein tyrosine phosphatase essential for T-cell activation through the antigen receptor, and functions as a positive regulator of T-cell coactivation upon binding to DPP443. At day 25, PPI analysis showed that genes such as FCGR3A, PDCD1LG2 and TLR8 were closely linked and were Hub Genes at this stage (Fig. 7F). FCGR3A binds both complexed or aggregated IgG as well as monomeric IgG, and is essential for facilitating antibody-dependent cellular cytotoxicity (ADCC) and other antibody-dependent responses, such as phagocytosis44. PDCD1LG2 plays a role in the costimulatory signaling pathway that is essential for T-cell proliferation and IFNG production45. TLR8 is a key component of both innate and adaptive immunity, functioning through MYD88 and TRAF6 to activate NF-kappa-B, which subsequently triggers cytokine secretion and the inflammatory response46.
Discussion
In this study, we successfully established a differentiation protocol to generate mature γδT cells from hESCs and our differentiation approach offers significant advantages. Firstly, the differentiation timeline was substantially shortened. Under our optimized conditions, we were able to generate a high proportion of γδT cells within 25 days, which is considerably faster than over 35 days in existing protocols. Secondly, our approach did not require foreign stromal cells, which are commonly used in previous differentiation protocols to support the development of γδT cells, consequently introducing additional complexity and increasing potential safety risks. By optimizing cytokine combinations and culture conditions, we successfully achieved the differentiation of γδT cells without the need for exogenous stromal cells, simplifying the procedure and reducing operational complexity.
Interestingly, αβT cells are predominantly generated in in vivo environments, however, we did not observe the presence of αβT cells in our differentiation system. Similarly, when human umbilical blood-derived CD34+ cells were used in the same differentiation protocol, the differentiation of αβT cell was also significantly suppressed, while γδT cells were predominant. These results indicated that our differentiation approach was highly directional and selective toward the γδ lineage. This selectivity suggests that inducing αβT cell differentiation requires more complex conditions or additional factors. Overall, these findings highlighted the strengths and efficacy of our differentiation system, while also shedding light on the complex regulatory mechanisms underlying the differentiation of γδT cells.
During the 25-day culture period, mature and tumor-reactive γδT cells were successfully differentiated from hESCs, these cells displayed normal developmental gene signatures and expressed markers of characteristics of γδT cells. Under the stimulation with PMA and ionomycin, hESC-derived γδT cells produced TNF-α and IL-2, confirming their functional potential. Importantly, these γδT cells exhibited significant cytotoxicity against the hepatocellular carcinoma lines HepG2 and Huh-7, achieving 20% to 30% killing efficiency, which is comparable to the 20% to 60% cytotoxicity reported for γδT cells isolated from human peripheral blood47. Furthermore, they effectively suppressed the proliferation of multiple tumor cell lines while showing negligible impact on normal cells. These findings not only validate the effectiveness of our differentiation strategy, but also further confirm the functional capacity of γδT cells in vitro. However, the relatively lower expression and secretion of perforin and granzymes is a critical issue that warrants attention. Given the high heterogeneity of γδT cells, where different subsets exhibit significant variations in tissue distribution and function, our next essential step is to utilize TCR sequencing to accurately determine the specific γδT cell subtype generated by our protocol, thereby exploring the relationship between the subset identity and its observed functional profile.
Taken together, our differentiation method offers notable advantages in terms of production potential, the ease of operation, and the safety. This approach potentially provides a new source of cells for γδT cell-based immunotherapy and drug development.
Moving forward, to fully realize the clinical potential of our hESC-derived γδT cells, we must address several key challenges associated with the current differentiation approach. Firstly, while our final product yielded approximately 40% γδT cells, the identity of the remaining 60% of cells remains uncharacterized. Furthermore, the detection of abnormal γδT subsets, such as the TCRγδ+CD3− population, presents a potential hurdle for the clinical application of these cells. Consequently, our immediate next step involves a thorough analysis of the final cell product using advanced techniques, such as single-cell sequencing, to accurately define the composition and heterogeneity of the output. Secondly, achieving large-scale expansion is mandatory for clinical translation. We will therefore need to further optimize our culture conditions to support the long-term, sustained expansion of γδT cells in vitro, or alternatively, focus on the robust expansion of the intermediate hematopoietic stem and progenitor cells HSPCs. Finally, to ensure clinical applicability, it is essential to develop a differentiation protocol that is entirely serum-free or uses defined serum substitutes. These concerted efforts are critical for refining the current protocol, enhancing the reliability of our results, and increasing the clinical relevance of our hESC-derived γδT cell product.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71 (3), 209–249 (2021).
Mikkilineni, L. & Kochenderfer, J. N. Chimeric antigen receptor T-cell therapies for multiple myeloma. Blood 130 (24), 2594–2602 (2017).
Mensurado, S., Blanco-Domínguez, R. & Silva-Santos, B. The emerging roles of γδ T cells in cancer immunotherapy. Nat. Rev. Clin. Oncol. 20 (3), 178–191 (2023).
Neelapu, S. S. et al. Chimeric antigen receptor T-cell therapy - assessment and management of toxicities. Nat. Rev. Clin. Oncol. 15 (1), 47–62 (2018).
Kobayashi, H. et al. Complete remission of lung metastasis following adoptive immunotherapy using activated autologous gammadelta T-cells in a patient with renal cell carcinoma. Anticancer Res. 30 (2), 575–579 (2010).
Aljurf, M. & Ezzat, A. Emerging role of gammadelta T-cells in health and disease. Blood Rev. 16 (4), 203–206 (2002).
Silva-Santos, B., Mensurado, S. & Coffelt, S. B. γδ T cells: pleiotropic immune effectors with therapeutic potential in cancer. Nat. Rev. Cancer. 19 (7), 392–404 (2019).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318 (5858), 1917–1920 (2007).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126 (4), 663–676 (2006).
Timmermans, F. et al. Generation of T cells from human embryonic stem Cell-Derived hematopoietic Zones1. J. Immunol. 182 (11), 6879–6888 (2009).
Chang, C. W. et al. Broad T-cell receptor repertoire in T-lymphocytes derived from human induced pluripotent stem cells. PLoS One. 9 (5), e97335 (2014).
Murai, N. et al. Re-generation of cytotoxic gammadeltat cells with distinctive signatures from human gammadeltat-derived iPSCs. Stem Cell. Rep. 18 (4), 853–868 (2023).
Jeon, S. B. et al. Lymphoid lineage γδ T cells were successfully generated from human pluripotent stem cells via hemogenic endothelium. Int. J. Stem Cells. 16 (1), 108–116 (2023).
Wang, N. et al. Hypoxia drives hematopoiesis with the enhancement of T lineage through eliciting arterial specification of hematopoietic endothelial progenitors from hESC. Stem Cell. Res. Ther. 13 (1), 282 (2022).
Chen, C. et al. Highly efficient generation of mature megakaryocytes and functional platelets from human embryonic stem cells. Stem Cell. Res. Ther. 15 (1), 454 (2024).
Bai, H. et al. Definitive hematopoietic multipotent progenitor cells are transiently generated from hemogenic endothelial cells in human pluripotent stem cells. J. Cell. Physiol. 231 (5), 1065–1076 (2016).
Suresh Babu, S. et al. Characterization of human induced pluripotent stems cells: current approaches, challenges, and future solutions. Biotechnol. Rep. (Amst). 37, e00784 (2023).
Bernardo, A. S. et al. BRACHYURY and CDX2 mediate BMP-induced differentiation of human and mouse pluripotent stem cells into embryonic and extraembryonic lineages. Cell. Stem Cell. 9 (2), 144–155 (2011).
Dzierzak, E. & Bigas, A. Blood development: hematopoietic stem cell dependence and independence. Cell. Stem Cell. 22 (5), 639–651 (2018).
Li, Y. et al. Induced expression of VEGFC, ANGPT, and EFNB2 and their receptors characterizes neovascularization in proliferative diabetic retinopathy. Invest. Ophthalmol. Vis. Sci. 60 (13), 4084–4096 (2019).
Zheng, Z. et al. Uncovering the emergence of HSCs in the human fetal bone marrow by single-cell RNA-seq analysis. Cell. Stem Cell. 29 (11), 1562–1579e7 (2022).
Ranzoni, A. M. et al. Integrative Single-Cell RNA-Seq and ATAC-Seq analysis of human developmental hematopoiesis. Cell. Stem Cell. 28 (3), 472–487e7 (2021).
Shim, S. H. et al. SAHA enhances differentiation of CD34 + CD45+ hematopoietic stem and progenitor cells from pluripotent stem cells concomitant with an increase in hemogenic endothelium. Stem Cells Transl Med. 11 (5), 513–526 (2022).
Murai, N. et al. Re-generation of cytotoxic γδT cells with distinctive signatures from human γδT-derived iPSCs. Stem Cell. Rep. 18 (4), 853–868 (2023).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14 (1), 36–49 (2014).
Montel-Hagen, A. et al. Organoid-Induced differentiation of conventional T cells from human pluripotent stem cells. Cell. Stem Cell. 24 (3), 376–389e8 (2019).
Li, Z. et al. Single-cell RNA-seq and chromatin accessibility profiling Decipher the heterogeneity of mouse γδ T cells. Sci. Bull. (Beijing). 67 (4), 408–426 (2022).
Belghali, M. Y. et al. Phenotypic characterization of human peripheral γδT-cell subsets in glioblastoma. Microbiol. Immunol. 66 (10), 465–476 (2022).
Shifrut, E. et al. Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175 (7), 1958–1971 (2018). e15.
Mathewson, N. D. et al. Inhibitory CD161 receptor identified in glioma-infiltrating T cells by single-cell analysis. Cell 184 (5), 1281–1298e26 (2021).
Song, Z. et al. Human γδ T cell identification from single-cell RNA sequencing datasets by modular TCR expression. J. Leukoc. Biol. 114 (6), 630–638 (2023).
O’Neill, K. et al. Perforins expression by cutaneous gamma delta T cells. Front. Immunol. 11, 1839 (2020).
Brandes, M. et al. Cross-presenting human gammadelta T cells induce robust CD8 + alphabeta T cell responses. Proc. Natl. Acad. Sci. U S A. 106 (7), 2307–2312 (2009).
Shen, J. et al. Defining early hematopoietic-fated primitive streak specification of human pluripotent stem cells by the orchestrated balance of Wnt, activin, and BMP signaling. J. Cell. Physiol. 234 (9), 16136–16147 (2019).
Gering, M. & Patient, R. Notch signalling and Haematopoietic stem cell formation during embryogenesis. J. Cell. Physiol. 222 (1), 11–16 (2010).
Feske, S., Wulff, H. & Skolnik, E. Y. Ion channels in innate and adaptive immunity. Annu. Rev. Immunol. 33, 291–353 (2015).
Banerjee, S. et al. JAK-STAT signaling as a target for inflammatory and autoimmune diseases: current and future prospects. Drugs 77 (5), 521–546 (2017).
Dunty, W. C. Jr. et al. Wnt3a/beta-catenin signaling controls posterior body development by coordinating mesoderm formation and segmentation. Development 135 (1), 85–94 (2008).
Wittler, L. et al. Expression of Msgn1 in the presomitic mesoderm is controlled by synergism of WNT signalling and Tbx6. EMBO Rep. 8 (8), 784–789 (2007).
Bulger, E. A. et al. TBXT dose sensitivity and the decoupling of nascent mesoderm specification from EMT progression in 2D human gastruloids. Development 151 (6). (2024).
Li, L. et al. M2 macrophage-Derived sEV regulate Pro-Inflammatory CCR2(+) macrophage subpopulations to favor Post-AMI cardiac repair. Adv. Sci. (Weinh). 10 (14), e2202964 (2023).
Komuczki, J. et al. Fate-Mapping of GM-CSF expression identifies a discrete subset of Inflammation-Driving T helper cells regulated by cytokines IL-23 and IL-1β. Immunity 50 (5), 1289–1304e6 (2019).
Williamson, A. J. et al. A specific PTPRC/CD45 phosphorylation event governed by stem cell chemokine CXCL12 regulates primitive hematopoietic cell motility. Mol. Cell. Proteom. 12 (11), 3319–3329 (2013).
Arce Vargas, F. et al. Fc effector function contributes to the activity of human Anti-CTLA-4 antibodies. Cancer Cell. 33 (4), 649–663e4 (2018).
Latchman, Y. et al. PD-L2 is a second ligand for PD-1 and inhibits T cell activation. Nat. Immunol. 2 (3), 261–268 (2001).
Xu, X. et al. CD8 T Cell-Derived Exosomal miR-186-5p elicits renal inflammation via activating tubular TLR7/8 signal axis. Adv. Sci. (Weinh). 10 (25), e2301492 (2023).
Xu, Y. et al. Allogeneic Vgamma9Vdelta2 T-cell immunotherapy exhibits promising clinical safety and prolongs the survival of patients with late-stage lung or liver cancer. Cell. Mol. Immunol. 18 (2), 427–439 (2021).
Acknowledgements
The authors declare that they have not used Artificial Intelligence in this study.
Funding
This work was supported in part by the National Key Research and Development Program of China (2018YFA0108201), by Research Starting Funding of South China University of Technology (D6201880, K5180910, K5204120, and D6212440), by Research Starting Funding of the Second Affiliated Hospital of South China University of Technology (KY09060026), by Research Agreement between South China University of Technology and Guangzhou First People’s Hospital (D9194290), by Foundation of Guangzhou Key Clinical Specialties (Institute of Clinical Medicine) (2024–2026), by Project of Key Medical Discipline in Guangzhou (2025–2027).
Author information
Authors and Affiliations
Contributions
YD, XZ, YF and YZ conceived the project and designed the experiments. XZ and CC performed the experiments, data analysis, and bioinformatics analysis. HW and YF cultured hESCs. NW, XS and YF contributed expertise. XZ and YD wrote and revised the manuscript. YD and YZ provided financial supports. All authors contributed to the manuscript and approved the version as submitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Guangzhou First people’s Hospital (Approval No.: K-2021–200, and approval date: December 12, 2021.) to use umbilical cord blood from healthy volunteers for Developing Differentiation System of Hematopoietic Stem/Progenitor Cells from Human Pluripotent Stem Cells and Function Validation of their Derivatives. The patients or their guardians/legally authorized representatives provided written informed consent for participation in the study with the use of umbilical cord blood. All mouse experiments were performed according to our experimental protocols approved by the Animal Ethics Committee of South China University of Technology University (Approval number: 2019073, and approval date: December 12, 2019.). WiCell Research Institute has confirmed that there was the ethical approval for collection of human embryonic stem cells (WA01(H1) and WA09(H9)) with NIH Approval NIHhESC-10–0043 and NIH Approval NIHhESC-10–0062, and that the donors had signed informed consent (www.wicell. org). We have confirmed that this study is reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Consent for publication
All authors consent for publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Chen, C., Fu, Y. et al. Generation of TCRγδ + T cells from human embryonic stem cells. Sci Rep 16, 6762 (2026). https://doi.org/10.1038/s41598-026-37941-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37941-w