Abstract
Recurrent implantation failure (RIF) remains a major challenge in assisted reproductive technology, and the molecular mechanisms underlying endometrial receptivity are incompletely understood. This study aimed to comprehensively characterize transcriptomic alterations, including alternative splicing events (ASEs), differential gene expression (DEGs), and immune cell dynamics across different phases of endometrial receptivity in women with RIF. Endometrial biopsies were collected from 90 healthy fertile controls and 73 RIF patients during pre-receptive, receptive, and post-receptive phases. High-throughput RNA sequencing was performed, and bioinformatic analyses were conducted to identify ASEs, DEGs, immune cell composition, and RNA-binding protein (RBP) networks. Skipped exons and mutually exclusive exons were the predominant splicing events observed. Both ASEs and DEGs were significantly enriched in pathways regulating cell adhesion, cytoskeletal organization, and immune modulation. KHDRBS3 emerged as a potential key RBP involved in splicing regulation during the window of implantation. Immune profiling revealed dynamic alterations in CD8 + T cells, NK cells, and monocytes between non-receptive and receptive phases, suggesting immune dysregulation associated with implantation failure. Drug repurposing analysis identified several small molecules targeting ASE-related genes, offering promising therapeutic options for RIF. These findings highlight the coordinated changes in alternative splicing, gene expression, and immune cell composition that characterize endometrial receptivity and provide insights that may guide the development of novel diagnostic biomarkers and targeted interventions to improve reproductive outcomes.
Similar content being viewed by others
Introduction
Although assisted reproductive technology (ART) has made significant progress over recent decades, 10% of patients still experience recurrent implantation failure (RIF), which remains a major challenge in clinical practice. RIF has been variably defined in the literature. According to the updated ESHRE guidelines (2023), a comprehensive assessment of contributing factors—including maternal age, embryo ploidy, and uterine pathology—is recommended when establishing a diagnosis of RIF. In the present study, we adhered to these recommendations and adopted an even more stringent definition: women under 40 years of age who failed to achieve a clinical pregnancy despite the transfer of at least four high-quality embryos across a minimum of three embryo transfer cycles1,2. As sequencing technologies evolve, transcriptomic data have become increasingly accessible. Studies have shown that the window of implantation (WOI), a short menstrual period when the endometrium is receptive to blastocyst transfer, is critical for successful implantation3,4. Transcriptomic analyses have revealed distinct genomic characteristics across the pre-receptive (PR), receptive (R), and post-receptive (PS) phases of the endometrium5,6. Based on these findings, transcriptome-based models to predict endometrial receptivity have been developed, leveraging transcript levels to offer personalized assessments of the WOI6,7,8,9,10. This approach has improved the diagnosis and management of RIF patients11,12. However, most existing studies have focused primarily on differential gene expression and have not comprehensively explored alternative splicing events (ASEs) as an additional regulatory layer of endometrial function.
Alternative splicing (AS), which transforms a single mRNA precursor into multiple transcript variants, enhances proteomic diversity and regulates numerous cellular processes13,14. More than 60% of human genes contain multiple exons, many of which give rise to cell-type-specific isoforms15. Studies have demonstrated changes in AS during embryonic development and early pregnancy. By analyzing single-cell RNA-seq data, Tian et al.16 found that AS is widespread in preimplantation embryo development, particularly at the two-cell stage. A homozygous splicing mutation of HFM1 demonstrated the potential risk of chromosomal anomalies under the RIF phenotype17. Similarly, in human embryonic arrest and RIF patients, splicing mutations have been shown to cause abnormal alternative splicing resulting in truncated MEI1 proteins18. Furthermore, Zhao et al. found that increased HOXA11-AS expression caused impaired PKM2 splicing and attenuated decidualization, a change consistently observed in RIF patients19.
RNA-binding proteins (RBPs) are a diverse protein family capable of binding single- or double-stranded RNA20,21,22. RBPs can promote the inclusion or skipping of exons by binding to splicing regulatory elements. Changes in the levels and activity of RBPs can cause dysregulation of AS23,24. Several studies have reported that RBPs play an important role in embryo implantation through the regulation of AS events, particularly in endometrial decidualization19,25,26,27,28. For example, PTBP1 has been shown to regulate PKM1/2 alternative splicing and gene expression, thereby affecting decidualization in RIF patients19. Further analysis of transcriptome data may be essential to understand the distribution, complexity, and regulation of alternative splicing across endometrial receptivity phases and to identify candidate RBPs involved in the control of key splicing events in endometrial function.
Endometrial immune dysfunction is another factor reducing receptivity and contributing to implantation failure. Various immune cells in the endometrium, such as natural killer (NK) cells, macrophages, and T cells, are essential for regulating receptivity and embryo implantation29,30. Uterine NK cells, T regulatory (Treg) cells, dendritic cells (DCs), and macrophages can directly or indirectly influence uterine epithelial adhesion, stromal cell transformation, trophoblast differentiation and invasion, and uterine vascular adaptation31,32,33. These immune cell profiles have been used to assess endometrial receptivity34,35,36. Although earlier studies have suggested that intravenous immunoglobulin therapy might benefit women with RIF37,38, recent ESHRE guidelines have questioned this practice, and its efficacy remains controversial. Therefore, the connection between immune cell composition and endometrial receptivity warrants further investigation with updated clinical evidence.
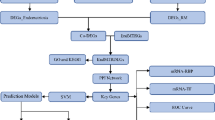
To investigate transcriptome dynamics during embryo implantation, we collected endometrial tissue at pre-receptive, receptive, and post-receptive phases from healthy controls (HC) with proven fertility and patients with RIF. RNA-seq was performed on all samples, followed by an extensive transcriptomic analysis, including DEGs, ASEs, immune cell estimation, and RBP network construction, with the aim of elucidating their regulatory relationships in endometrial function. By comprehensively characterizing the endometrial transcriptome across receptivity phases, this study aimed to clarify the biological significance of widespread alternative splicing and further evaluate immune cell changes and their potential role in implantation failure. In addition, an ASE–RBP correlation network was constructed to explore the dysregulation of splicing by RBPs, and drug repurposing analysis was performed to identify potential therapeutic candidates targeting dysregulated AS. Compared to previous transcriptomic studies, we conducted further analysis of ASEs in endometrial receptivity and attempted to identify key RBPs responsible for splicing regulation. By revealing transcriptional diversity and dysregulation, we aimed to provide a valuable resource for studying splicing and identifying potential biomarkers and therapeutic targets in RIF, as summarized in the study design flowchart (Fig. 1).
Overview of the study design and analytical workflow. Endometrial tissue was collected from pre-receptive (PR), receptive (R), and post-receptive (PS) phases from healthy fertile controls (HC) and patients with recurrent implantation failure (RIF). Bioinformatic analysis included differential gene expression (DEG) and alternative splicing event (ASE) identification, immune cell estimation, RNA-binding protein (RBP) motif enrichment, and drug repurposing analysis. The figure was created with BioRender.com.
Materials and methods
Study design and ethical approval
This prospective observational study was approved by the Ethics Committee of Shanghai Ji Ai Genetics and IVF Institute of Obstetrics and Gynecology, affiliated with Fudan University (JIAI E2019-04; JIAI E2020-015). All participants provided written informed consent prior to sample collection. All methods were carried out in accordance with relevant guidelines and regulations, including the Declaration of Helsinki and its later amendments.
Participant recruitment and inclusion criteria
In total, 163 endometrial biopsy samples were included, comprising 90 samples from fertile healthy controls (HC) and 73 samples from patients with recurrent implantation failure (RIF). The HC cohort (n = 90) was obtained from our previously published endometrial receptivity transcriptome study, in which endometrial biopsies were collected across the natural-cycle luteinizing hormone (LH)–timed phases (LH + 3, LH + 5, LH + 7, and LH + 9) under standardized clinical procedures and subsequently profiled by RNA sequencing as reported previously6. These HC participants were fertile volunteers with proven fertility and no history of infertility, recurrent miscarriage, or known uterine pathology.
The RIF cohort (n = 73) represents newly sequenced samples generated in the present study, collected from 35 patients undergoing a harmonized clinical management protocol at our center. Baseline demographic/clinical characteristics of the newly sequenced RIF cohort are summarized in Table 1, and baseline hormone profiles under the HRT regimen are provided in Table S1.
RIF was defined as unexplained RIF with ≥ 3 failed embryo transfer attempts involving ≥ 4 high-quality embryos, where high-quality embryos were defined as either day-5 blastocysts graded at least 4BB (Gardner’s classification) or cleavage-stage embryos with ≥ 7 cells. Eligible patients were aged 20–40 years, had BMI 19–24 kg/m2, and an endometrial thickness of ≥ 7 mm at the time of endometrial preparation. To minimize confounding, patients with untreated hydrosalpinx, endometrial disease, severe adenomyosis, diminished ovarian reserve, genetic disorders, or immune abnormalities, or those unwilling to participate, were excluded from the study.
For all RIF participants, embryo grading criteria and embryo transfer–related clinical procedures (including cycle management and luteal support strategy) followed uniform institutional standard operating procedures throughout the study period.
Endometrial preparation protocols
For RIF patients, endometrial preparation was performed using artificial hormone replacement therapy (HRT). On menstrual cycle day 2, oral estradiol valerate (Progynova; Bayer, Leverkusen, Germany) was initiated at 4–6 mg/day until endometrial thickness reached ≥ 7 mm. Serum progesterone was then assessed; when progesterone was ≤ 1.5 ng/mL, 90 mg/day sustained-release vaginal progesterone gel (Crinone; Merck-Serono, Darmstadt, Germany) was administered. The start day of progesterone exposure was designated as P + 0, and endometrial biopsies were performed at P + 3, P + 5, and P + 7, corresponding to the pre-receptive, receptive, and post-receptive phases, respectively. Baseline gonadotropin and steroid hormone measurements associated with the HRT protocol are provided in Table S1.
Healthy controls underwent natural cycles with daily urinary LH monitoring (Eupregna, China). The day of the LH surge was defined as LH + 0, and biopsies were performed at LH + 3/LH + 5 (pre-receptive), LH + 7 (receptive), and LH + 9 (post-receptive), consistent with the timing scheme described in the prior publication6.
Endometrial sampling and processing
Endometrial biopsy specimens were collected aseptically using a sterile, single-use endometrial suction catheter (Yudu Medical Apparatus and Instruments Co., Ltd., Suzhou, China). Immediately after collection, tissues were immersed in Allprotect Tissue Reagent (QIAGEN GmbH, Hilden, Germany), transported at 4 °C using standardized cold-chain procedures, and cryopreserved at –80 °C until downstream processing.
RNA extraction, sequencing, and data processing
Total RNA extraction, library preparation, and high-throughput sequencing were performed following the protocols described previously6. Sequencing reads were aligned to the GRCh38 human reference genome. Quality control, read mapping, and transcript quantification were conducted using established pipelines as previously reported6. All RNA-seq datasets generated in this study have been deposited in the Gene Expression Omnibus (GEO) under accession number GSE287072.
Alternative splicing analysis
Quality-controlled data were analyzed using rMATS v4.1.1 to identify differential alternative splicing events (ASEs) across the following comparisons: (1) HC: PR vs. R; (2) HC: PS vs. R; (3) RIF: PR vs. R; and (4) RIF: PS vs. R. Five basic AS types were evaluated: skipped exons (SE), mutually exclusive exons (MXE), alternative 5′ splice sites (A5SS), alternative 3′ splice sites (A3SS), and retained introns (RI). The JCEC model (reads on target and junction counts) was applied for quantification. Significant ASEs were defined as those with an absolute inclusion level difference (ΔPSI) > 0.1 and false discovery rate (FDR) < 0.05, as recommended by Shen et al.39.
Functional enrichment and coding potential assessment
Genes containing significant differential ASEs (ASEGs) were subjected to gene set enrichment analysis using the ClusterProfiler R package40. The MASER Bioconductor package was used to assess coding potential alterations resulting from splicing events41.
Immune cell composition analysis
Immune cell infiltration was estimated using CIBERSORT42, applying a validated signature matrix to deconvolute 22 immune cell types from transcriptomic data. Correlations between ASEs and immune cell proportions were assessed by Pearson correlation.
RBP binding motif and network analysis
The rMAPS2 platform was used to assess enrichment of RBP binding motifs around alternative splicing regions. Correlations between RBP expression and ASE inclusion levels were computed. Networks of RBPs and ASEs with correlation coefficient > 0.5 and p < 0.05 were visualized using Cytoscape v3.9.143.
Drug repurposing analysis
Differentially spliced genes were uploaded to the Connectivity Map platform (https://clue.io), and the Drug Repurposing Hub44, part of the Broad Institute’s Connectivity Map, was used to identify small molecules with transcriptomic signatures inversely correlated to the observed alterations. To prioritize already approved small molecule candidates, overlapping ASEGs from the HC: PR vs. R and RIF: PR vs. R groups were searched in the repurposing database. The same analyses were performed for overlapping ASEGs identified in the HC: PS vs. R and RIF: PS vs. R comparisons. A signature-matching strategy was applied to retrieve compounds potentially capable of reversing disease-associated transcriptomic changes. Finally, the DrugBank database (https://go.drugbank.com)45 was used to confirm compound approval status, pharmacological properties, and known indications.
Statistical analysis
Group differences were assessed using one-way analysis of variance (ANOVA). Results are presented as mean ± standard deviation (SD). A p-value < 0.05 was considered statistically significant. All analyses were performed using R (v4.1.3) and Python (v3.6.10).
Results
Subjects and sequencing quality assessment
The patient recruitment, sample collection, and transcriptomic data analysis of healthy controls have been previously described6. No statistically significant differences were observed in age, body mass index, previous implantation failure, menstrual cycle length, or endometrial thickness among the three groups of endometrial samples collected from RIF patients undergoing HRT (Table 1). The average raw RNA-seq data generated from endometrial tissue samples of RIF patients was approximately 9.70 GB. After quality control and removal of ribosomal RNA (see Materials and Methods), the average clean data size was 8.89 GB. All clean reads were mapped to the human reference genome (GRCh38), and each sample achieved a mapping rate exceeding 80%.
Comprehensive alternative splicing analysis across endometrial receptive phases
To obtain the landscape of alternative splicing variation across human endometrial receptive phases, we performed differential expression and alternative splicing analyses between the non-receptive (PR or PS) and receptive (R) phases in both healthy controls and RIF patients.
In total, 1,240,563 ASEs were identified between PR and R stages in healthy controls (HC: PR vs. R), 981,422 ASEs between PS and R stages in healthy controls (HC: PS vs. R), 985,440 ASEs between PR and R stages in RIF patients (RIF: PR vs. R), and 1,010,588 ASEs between PS and R stages in RIF patients (RIF: PS vs. R), with percentage spliced-in (PSI) scores calculated (Table 2).
After applying stringent criteria (FDR < 0.05 and absolute ΔPSI > 0.1), 1,048 differential ASEs were identified in the HC: PR vs. R comparison, 544 in HC: PS vs. R, 1,744 in RIF: PR vs. R, and 804 in RIF: PS vs. R (Table 2; Figure S1A,B). Consistent across all groups, skipped exons (SE) and mutually exclusive exons (MXE) were the most common splicing events.
Comprehensive pre-receptive and receptive phases transcriptome analysis
Further analysis of the PR and R phases in healthy controls revealed 1,048 differential ASEs, including SE (n = 520, 49.62%), MXE (n = 239, 22.80%), A5SS (n = 88, 8.40%), A3SS (n = 114, 10.88%), and RI (n = 87, 8.30%). In the RIF: PR vs. R group, 1,744 ASEs were identified, comprising 52.24% SE, 21.56% MXE, 7.22% A5SS, 8.89% A3SS, and 10.09% RI (Fig. 2A).
Alternative splicing and immune cell analyses between pre-receptive and receptive phases in RIF patients. (A) Analysis of differential ASEs between endometrial PR vs. R phases of HC and RIF (green) patients (orange). The percentage of each type of ASE within each comparison is represented by a pie chart (right) (SE, skipped exon; MXE, mutually exclusive exons; A3SS, alternative 3’ splice site; A5SS, alternative 5’ splice site; RI, retained intron). (B) Venn diagram showing the genes with alternative splicing events in HC: PR vs. R (blue) and RIF: PR vs. R (orange). (C) The proportions of differential ASEs for each type in in HC: PR vs. R (blue) and RIF: PR vs. R (orange). (D) The frequency of possible changes in coding potential shown for SE, MXE, and RI, respectively. (E) Scatter plot displayed the GO analysis of genes with different alternative splicing events. BP: Biological process, CC: cellular component, MF: molecular function. (F) Comparing immune cell distribution between endometrial PR vs. R phase of RIF patients was shown in the boxplot (ns: p > 0.05, * p < = 0.05, ** p < = 0.01, *** p < = 0.001, **** p < = 0.0001). (G) Heatmap of the distribution of immune cells interacting with different ASEs in the endometrial R phases of RIF patients.
Among the 505 ASEGs detected in the HC: PR vs. R group, only 26 (5.15%) were also DEGs, whereas 479 (94.85%) were not differentially expressed at the gene level. Similarly, in the RIF: PR vs. R group, only 93 (11.74%) of the 792 ASEGs overlapped with DEGs (Figure S2A).
A total of 178 ASEGs were shared between the HC: PR vs. R and RIF: PR vs. R comparisons, of which 11 genes (CKB, FN1, GRAMD1C, KIF12, NNMT, PDGFA, RABGAP1L, RIMKLB, SDCBP2, SYNE2, TYMP) were also differentially expressed (Fig. 2B; Figure S2A). Among these, 31 ASEGs have been previously reported as associated with hormonal regulation, endometrial stromal cell differentiation, trophoblast development, and implantation (Table 3). Five of these genes (EPB41L2, OFD1, POLD4, POSTN, and SH3YL1) have been identified as predictive markers of the endometrial WOI in RIF patients7,12. Six other differentially expressed ASEGs (CKB, GRAMD1C, KIF12, NNMT, PDGFA, and SYNE2) were reported to be related to endometrial receptivity7,12,46,47.
Analysis of the frequency of differential ASEs demonstrated high occurrence rates across samples (Fig. 2C). Evaluation of coding potential indicated that SE events most frequently retained coding capacity, whereas RI events led to frequent switches from coding to non-coding transcripts (Fig. 2D).
Gene Ontology enrichment analyses of ASEGs from both groups showed involvement in actin regulation, cell adhesion, GTPase activity, and RNA splicing (Fig. 2E; Table S2). Corresponding DEGs were also enriched in cell adhesion and immune signaling pathways (Figure S2B; Table S2). These processes have been implicated in embryo implantation48, especially adhesion junction proteins49,50 and immune regulation51,52.
Comprehensive post-receptive and receptive phases transcriptome analysis
Analysis of the PS and R phases in healthy controls identified 544 differential ASEs, including SE (n = 276, 50.73%), MXE (n = 127, 23.35%), A5SS (n = 35, 6.43%), A3SS (n = 52, 9.56%), and RI (n = 54, 9.93%). In the RIF: PS vs. R group, 804 ASEs were found, comprising 53.61% SE, 20.02% MXE, 6.59% A5SS, 9.70% A3SS, and 10.08% RI (Fig. 3A).
Alternative splicing and immune cell analyses between PS and R phases in patients with RIF. (A) Analysis of differential ASEs between endometrial PS vs. R phases (blue) of HC and RIF patients (orange). The percentage of each type of ASEs within each comparison is represented by a pie chart (right) (SE, skipped exon; MXE, mutually exclusive exons; A3SS, alternative 3’ splice site; A5SS, alternative 5’ splice site; RI, retained intron). (B) Venn diagram showing the genes with alternative splicing events in HC: PS vs. R (cyan) and RIF: PS vs. R (blue). (C) The proportions of differential ASEs for each type in HC: PS vs. R (cyan) and RIF: PS vs. R (blue). (D) The frequency of possible changes in coding potential shown for SE, MXE, and RI, respectively. (E) Scatter plot displayed the GO analysis of genes with different alternative splicing events. BP: Biological process, CC: cellular component, MF: molecular function. (F) Comparing immune cell distribution between endometrial PS vs. R phase of RIF patients was shown in the boxplot (ns: p > 0.05, * p < = 0.05, ** p < = 0.01, *** p < = 0.001, **** p < = 0.0001). (G) Heatmap of the distribution of immune cells interacting with different ASEs in the endometrial R phases of RIF patients.
Of the 342 ASEGs identified in the HC: PS vs. R group, only 5 (1.46%) overlapped with DEGs, whereas in the RIF: PS vs. R group, 38 (8.70%) of the 437 ASEGs were also DEGs (Figure S3A). A total of 62 ASEGs were shared between these comparisons (Fig. 3B), with 16 reported to be involved in embryo implantation (Table 3). Among these, APOL2, DUOXA1, SLC37A2, and ELP3 were also DEGs in both groups. AIMP1 and C3 have been associated with endometrial receptivity7,46.
Frequency analysis showed high prevalence of ASEs across samples, with a trend towards higher frequency in RIF patients (Fig. 3C). Coding potential analysis revealed that SE events generally preserved coding capacity, whereas RI events frequently resulted in non-coding isoforms (Fig. 3D).
GO enrichment analyses indicated that ASEGs were mainly related to cell adhesion, substrate junctions, and focal adhesion (Fig. 3E; Table S3). DEGs were enriched in processes such as epidermis development, cytokine activity, and NK cell-mediated immunity (Figure S3B; Table S3).
Immune cell composition and correlation with ASEs
DEGs identified in both healthy controls and RIF patients were enriched in immunomodulatory processes, including immune cell activation and migration. Analysis of immune cell composition revealed that CD8 + T cells, resting NK cells, and resting mast cells were significantly decreased, whereas monocytes and macrophages M0 were significantly increased in the receptive (R) phase endometrium of RIF patients (Fig. 2F). Similarly, an increase in monocytes was observed in the R phase endometrium of healthy controls (Figure S2C).
Heatmap analysis demonstrated that several altered immune cell types, such as resting NK cells and resting mast cells, were closely associated with differential ASEs in both RIF and HC samples (Fig. 2G; Figure S2D).
In the comparison between post-receptive (PS) and receptive (R) phases, immune cell analysis showed a significant increase in CD8 + T cells, M1 macrophages, and M2 macrophages, and a significant decrease in activated NK cells in RIF patients (Fig. 3F). Although no statistically significant differences in immune cell proportions were found between PS and R phases in healthy controls, similar trends in CD8 + T cells and activated NK cells were observed (Figure S3C). Among these immune cell types, activated NK cells displayed high correlation with specific ASEs during the receptive phase (Fig. 3G; Figure S3D).
These findings suggest that dynamic changes in immune cell populations during the window of implantation may be closely linked to recurrent implantation failure.
Dysregulated RNA-binding protein network and ASE regulation
RNA-binding proteins (RBPs) are critical regulators of alternative splicing. Transcriptomic profiling identified seven RBP-related genes (YBX2, CPEB2, IGF2BP2, IGF2BP3, KHDRBS3, BRUNOL5, and RBFOX1) that were differentially expressed in at least one comparison group (Fig. 4A). Among these, CPEB2 has been implicated in trophoblast-related regulatory programs53,54. Notably, KHDRBS3 showed the highest degree of connectivity with differential ASEs in our correlation network (Fig. 4C) and exhibited a consistent decrease in the receptive phase compared with the pre-receptive phase in both HC and RIF cohorts (Fig. 4D), nominating it as a candidate splicing regulator associated with receptivity transitions.
The regulation network of ASEs by differentially expressed RBPs genes. (A) Differentially expressed RBP genes in four subgroups. (B) Significant enrichment of RBP motifs around AS events. On the x-axis, 5’ to 3’, the position (R) of each splicing event is shown relative to the event type. Each region is numbered from R1 to R(n), where n is the total number of regions for each event type. The binding motifs of the RBP are described on the y-axis. Each panel shows a single event type (SE, MXE, A5SS, A3SS, RI). The significance of the RBPs is indicated by the color and size of the circles. (C) Regulatory network of seven differentially expressed RBPs for the five types of alternative splicing (SE, MXE, A5SS, A3SS, RI) of ASEs. (D) Boxplots of KHDRBS3 expression in endometrial PR vs. R phase of HC population and RIF patients. p-values for each dataset are shown with corresponding group names.
Analysis of RBP binding motifs showed significant enrichment in regions flanking SE and MXE events (Fig. 4B). Correlation analysis between expression levels of RBPs and ASE inclusion (PSI) values revealed that KHDRBS3 was associated with the highest number of ASEs, followed by IGF2BP2, YBX2, IGF2BP3, BRUNOL5, RBFOX1, and CPEB2 (Fig. 4C).
Notably, KHDRBS3 expression exhibited a consistent trend of significant reduction in the receptive phase compared to the pre-receptive phase in both HC and RIF groups (Fig. 4D). These results support the hypothesis that RBPs contribute to splicing regulation in endometrial receptivity.
Drug repurposing analysis of ASE-regulated genes
Genes harboring pathogenic or aberrant splice variants are potential therapeutic targets. We collected 178 ASEGs shared between HC: PR vs. R and RIF: PR vs. R groups, and 62 ASEGs shared between HC: PS vs. R and RIF: PS vs. R groups, for drug repurposing analysis.
Differentially spliced genes were uploaded to the Connectivity Map platform (https://clue.io), and the Drug Repurposing Hub44, part of the Connectivity Map, was used to identify small molecules with transcriptomic signatures inversely correlated with observed alterations. A signature-matching strategy was applied to retrieve compounds potentially capable of reversing disease-associated transcriptomic changes. Finally, the DrugBank database45 was used to confirm compound approval status, pharmacological properties, and known indications.
We identified 20 target ASEGs associated with 68 small molecules. Among them, GABRP, MAPK12, and SGK1 were targeted by 26, 13, and 10 compounds, respectively (Table S4). Six ASEGs (CD44, CKB, COX6C, NNMT, NR1H3, and ANXA1) previously reported in embryo implantation were matched with small molecules confirmed in DrugBank. These included hyaluronic acid (CD44), creatine (CKB), cholic acid (COX6C), niacin (NNMT), T-0901317 (NR1H3), and corticosteroids such as amcinonide, dexamethasone, and hydrocortisone phosphate (ANXA1) (Fig. 5A, B).
Drug repurposing analysis of genes with alternative splicing events. (A) Sankey diagram identifying small molecules targeting genes with differentially alternative splicing events. (B) Chemical structures of eight small molecule drugs.
These compounds represent potential candidates for therapeutic intervention in RIF patients.
Discussion
Recurrent implantation failure (RIF) remains a major challenge in assisted reproductive technology, and its pathogenesis is widely considered multifactorial, involving impaired endometrial receptivity, dysregulated immune–stromal crosstalk, and embryo‑related factors. In this study, we profiled bulk endometrial transcriptomes across receptivity phases to characterize coordinated changes in gene expression, alternative splicing (AS), and inferred immune composition in healthy controls (HC) and RIF patients.
Across both cohorts, skipped exons (SE) and mutually exclusive exons (MXE) were the most prevalent AS types, consistent with the predominance of exon-skipping patterns reported in human tissues. We observed a larger shift in both differentially expressed genes (DEGs) and alternative splicing event genes (ASEGs) when comparing the pre-receptive (PR) to receptive (R) phase than when comparing the post-secretory (PS) to R phase, supporting extensive transcriptomic remodeling during establishment of the window of implantation. Functional enrichment of phase-associated ASEGs/DEGs implicated pathways linked to cell adhesion, actin cytoskeleton remodeling, and small-GTPase signaling—processes repeatedly connected to implantation competence through regulation of epithelial–stromal interactions, trophoblast attachment, and endometrial remodeling48,50,51. Together, these data support a model in which receptivity acquisition involves not only transcriptional reprogramming but also broad splicing rewiring affecting adhesion- and cytoskeleton-related networks.
RNA-binding proteins (RBPs) are central regulators of splice-site choice, and their dysregulation can generate coordinated AS programs. By integrating RBP expression with AS patterns, we prioritized several RBPs as candidate upstream regulators of receptivity-associated splicing transitions. Among these, KHDRBS3 emerged as a notable candidate: it exhibited broad connectivity with differential AS events in the RBP–ASE network (Fig. 4C) and showed a consistent decrease from PR to R in both HC and RIF cohorts (Fig. 4D). KHDRBS3 has been implicated in RNA metabolism and signal-transduction-related processes in other biological contexts, suggesting that altered KHDRBS3 activity could plausibly reshape splicing programs relevant to endometrial remodeling. In our dataset, the timing of KHDRBS3 down-regulation coincided with the most pronounced ASEG/DEG transitions, supporting the hypothesis that KHDRBS3-centered splicing regulation participates in the molecular switch into the receptive state. Importantly, because bulk RNA-seq aggregates signals across multiple endometrial cell types, the observed KHDRBS3 dynamics may reflect both cell-intrinsic regulation and compositional shifts; cell type–resolved analyses will be required to localize the dominant cellular source(s) and to establish direct KHDRBS3 targets.
Endometrial immune balance is another determinant of receptivity, and our CIBERSORT-based deconvolution suggested coordinated immune remodeling across phases. Notably, the inferred immune changes were phase-contrast dependent rather than uniform: in PR → R, resting NK cell signatures and several lymphocyte subsets decreased while monocytes and macrophage M0 increased in the receptive-phase endometrium of RIF patients (Fig. 2F), whereas in PS → R, activated NK cell signatures decreased alongside shifts in macrophage subsets (Fig. 3F). Across both comparisons, the most reproducible pattern involved the NK–myeloid axis, indicating an imbalanced remodeling trajectory during the transition toward receptivity in RIF. This pattern is biologically plausible because appropriate implantation requires synchronized immune tolerance and tissue remodeling, processes in which uterine NK cells and myeloid populations play key roles30,31,32,53. In parallel, we observed that altered immune fractions were correlated with specific differential ASEs in the receptive phase (Figs. 2G and 3G), supporting a link between splicing programs and immune microenvironment dynamics.
To place the bulk immune findings into a more granular cellular context, recent single-cell endometrial atlases have mapped dynamic immune and stromal populations across the menstrual cycle, including resident NK and monocyte/macrophage lineages that vary with hormonal state55. In addition, single-cell studies focusing on RIF have reported perturbations in immune and stromal programs compared with fertile controls, including altered NK-cell states and macrophage/monocyte signatures in RIF endometrium56,57. Although single-cell studies differ in cohort characteristics and sampling windows, these reports provide a cell-type–resolved framework that is qualitatively consistent with the immune remodeling patterns inferred from our bulk deconvolution, while highlighting an important limitation of bulk RNA-seq/CIBERSORT: “NK cells” and “macrophages” at the bulk level likely reflect shifts in specific subsets and activation states rather than uniform lineage-wide changes. This is further supported by high-resolution single-cell mapping of the maternal–fetal interface, which has delineated functionally distinct NK subpopulations with divergent cytokine/chemokine and tissue-remodeling programs58.
A salient implication of our integrated results is that AS regulation and immune remodeling may be mechanistically coupled. Splicing changes affecting adhesion molecules, cytoskeletal regulators, or immune signaling mediators could reshape cell–cell interactions and cytokine networks, thereby influencing recruitment or activation of NK cells and monocytes/macrophages. Conversely, inflammatory or stress cues derived from myeloid populations may affect RBP activity and spliceosomal regulation, reinforcing aberrant AS programs. In this framework, the KHDRBS3-associated splicing signature observed here may represent one molecular node at which receptivity-linked remodeling intersects with immune dysregulation in RIF.
Beyond mechanistic inference, we applied a drug repurposing strategy to prioritize compounds targeting genes implicated in the AS/RBP network. Several candidate small molecules were identified, including agents with immunomodulatory or nuclear receptor-related activity, suggesting potential routes to restore receptivity-associated transcriptomic programs. Nevertheless, translating these computational candidates into clinical interventions will require careful preclinical validation, including assessment of endometrial bioavailability, timing relative to the window of implantation, and the directionality of immune effects.
Several limitations warrant consideration. First, embryo aneuploidy cannot be fully excluded because preimplantation genetic testing was not performed; although RIF enrollment required repeated transfers of high-quality embryos, embryo-intrinsic factors remain a potential confounder. Second, bulk RNA-seq cannot definitively assign DEGs/ASEs (including KHDRBS3-linked splicing signals) to specific endometrial cell types, nor can it resolve NK and monocyte/macrophage subpopulations. Third, the RBP–AS associations are correlative; establishing causality and defining direct targets will require orthogonal validation (e.g., perturbation assays coupled with isoform-level readouts and/or RBP–RNA interaction assays).
In summary, our data indicate that acquisition of endometrial receptivity is accompanied by marked phase-dependent remodeling of gene expression and alternative splicing, and that RIF is associated with coordinated perturbations in splicing regulatory RBPs (including KHDRBS3) and dysregulated immune remodeling involving the NK–monocyte/macrophage axis. These integrated observations refine the transcriptomic framework of implantation failure and support the development of receptivity-focused biomarkers and hypothesis-driven functional studies to delineate causal pathways linking splicing regulation, immune–stromal interactions, and implantation outcomes.
Conclusion
This study demonstrates that endometrial receptivity is characterized by phase-dependent alterations in gene expression, alternative splicing events, and immune cell composition. The comprehensive transcriptomic profiling conducted here revealed that alternative splicing, particularly involving KHDRBS3 and other RNA-binding proteins, may play a pivotal role in regulating endometrial function during the window of implantation. Furthermore, the identification of differentially expressed genes and associated immune signatures highlights the complex molecular landscape underlying recurrent implantation failure. Collectively, these findings provide valuable insights into the mechanisms governing embryo implantation and may inform the development of novel diagnostic biomarkers and therapeutic strategies to improve implantation outcomes in assisted reproduction.
Data availability
The data presented in this study are openly available in the Gene Expression Omnibus (GEO) database at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE287072, accession number GSE287072. All other relevant data supporting the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- A3SS:
-
Alternative 3’ splice site
- A5SS:
-
Alternative 5’ splice site
- ART:
-
Assisted reproductive technology
- AS:
-
Alternative splicing
- ASE:
-
Alternative splicing event
- ASEG:
-
Alternative splicing event gene
- DC:
-
Dendritic cell
- DEG:
-
Differentially expressed gene
- dNK:
-
Decidual natural killer
- ERD:
-
Endometrial receptivity diagnosis
- HC:
-
Healthy controls
- LH:
-
Luteinizing hormone
- MXE:
-
Mutually exclusive exons
- NK:
-
Natural killer
- P:
-
Progesterone
- pET:
-
Personalized embryo transfer
- PR:
-
Pre-receptive
- PS:
-
Post-receptive
- R:
-
Receptive
- RBP:
-
RNA-binding protein
- RI:
-
Retained intron
- RIF:
-
Recurrent implantation failure
- SE:
-
Skipped exon
- Tregs:
-
T regulatory cells
- WOI:
-
Window of implantation
References
Simon, A. & Laufer, N. Repeated implantation failure: Clinical approach. Fertil. Steril. 97(5), 1039–1043. https://doi.org/10.1016/j.fertnstert.2012.03.010 (2012).
Griesinger, G. & Larsson, P. Conventional outcome reporting per IVF cycle/embryo transfer may systematically underestimate chances of success for women undergoing ART: Relevant biases in registries, epidemiological studies, and guidelines. Hum. Reprod. Open 2023(2), hoad018. https://doi.org/10.1093/hropen/hoad018 (2023).
Galliano, D., Bellver, J., Diaz-Garcia, C., Simon, C. & Pellicer, A. Art and uterine pathology: How relevant is the maternal side for implantation?. Hum. Reprod. Update 21(1), 13–38. https://doi.org/10.1093/humupd/dmu047 (2015).
Sebastian-Leon, P., Garrido, N., Remohi, J., Pellicer, A. & Diaz-Gimeno, P. Asynchronous and pathological windows of implantation: Two causes of recurrent implantation failure. Hum. Reprod. 33(4), 626–635. https://doi.org/10.1093/humrep/dey023 (2018).
Lipecki, J. et al. EndoTime: Non-categorical timing estimates for luteal endometrium. Hum. Reprod. 37(4), 747–761. https://doi.org/10.1093/humrep/deac006 (2022).
Zhang, W. B. et al. Transcriptomic analysis of endometrial receptivity for a genomic diagnostics model of Chinese women. Fertil. Steril. 116(1), 157–164. https://doi.org/10.1016/j.fertnstert.2020.11.010 (2021).
Diaz-Gimeno, P. et al. A genomic diagnostic tool for human endometrial receptivity based on the transcriptomic signature. Fertil. Steril. 95(1), 50–60. https://doi.org/10.1016/j.fertnstert.2010.04.063 (2011).
Ruiz-Alonso, M. et al. The endometrial receptivity array for diagnosis and personalized embryo transfer as a treatment for patients with repeated implantation failure. Fertil. Steril. 100(3), 818–824. https://doi.org/10.1016/j.fertnstert.2013.05.004 (2013).
Ruiz-Alonso, M., Valbuena, D., Gomez, C., Cuzzi, J. & Simon, C. Endometrial receptivity analysis (ERA): Data versus opinions. Hum. Reprod. Open 2021(2), hoab011. https://doi.org/10.1093/hropen/hoab011 (2021).
Liu, Z. et al. The clinical efficacy of personalized embryo transfer guided by the endometrial receptivity array/analysis on IVF/ICSI outcomes: A systematic review and meta-analysis. Front. Physiol. 13, 841437. https://doi.org/10.3389/fphys.2022.841437 (2022).
Li, N. et al. Personalized embryo transfer guided by rsERT improves pregnancy outcomes in patients with repeated implantation failure. Front. Med. 11, 1369317. https://doi.org/10.3389/fmed.2024.1369317 (2024).
He, A. et al. The role of transcriptomic biomarkers of endometrial receptivity in personalized embryo transfer for patients with repeated implantation failure. J. Transl. Med. 19(1), 176. https://doi.org/10.1186/s12967-021-02837-y (2021).
Graveley, B. R. Alternative splicing: Increasing diversity in the proteomic world. Trends Genet. 17(2), 100–107. https://doi.org/10.1016/s0168-9525(00)02176-4 (2001).
Kwan, T. et al. Heritability of alternative splicing in the human genome. Genome Res. 17(8), 1210–1218. https://doi.org/10.1101/gr.6281007 (2007).
de Klerk, E. & t Hoen, P. A. Alternative mRNA transcription, processing, and translation: Insights from RNA sequencing. Trends Genet. 31(3), 128–139. https://doi.org/10.1016/j.tig.2015.01.001 (2015).
Tian, G. G., Li, J. & Wu, J. Alternative splicing signatures in preimplantation embryo development. Cell Biosci. 10, 33. https://doi.org/10.1186/s13578-020-00399-y (2020).
Tang, F. et al. Novel deleterious splicing variant in HFM1 causes gametogenesis defect and recurrent implantation failure: Concerning the risk of chromosomal abnormalities in embryos. J. Assist. Reprod. Genet. 40(7), 1689–1702. https://doi.org/10.1007/s10815-023-02761-8 (2023).
Dong, J. et al. Novel biallelic mutations in MEI1: Expanding the phenotypic spectrum to human embryonic arrest and recurrent implantation failure. Hum. Reprod. 36(8), 2371–2381. https://doi.org/10.1093/humrep/deab118 (2021).
Zhao, H. et al. Increased expression of HOXA11-AS attenuates endometrial decidualization in recurrent implantation failure patients. Mol. Ther. 30(4), 1706–1720. https://doi.org/10.1016/j.ymthe.2022.01.036 (2022).
Xu, X., Shen, H. R., Zhang, J. R. & Li, X. L. The role of insulin-like growth factor 2 mRNA binding proteins in female reproductive pathophysiology. Reprod. Biol. Endocrinol. 20(1), 89. https://doi.org/10.1186/s12958-022-00960-z (2022).
Downing, P. et al. WD-repeat containing protein-61 regulates endometrial epithelial cell adhesion indicating an important role in receptivity. Mol. Hum. Reprod. https://doi.org/10.1093/molehr/gaae039 (2024).
Potiris, A. et al. The contribution of proteomics in understanding endometrial protein expression in women with recurrent implantation failure. J. Clin. Med. https://doi.org/10.3390/jcm13072145 (2024).
Zhang, J. & Manley, J. L. Misregulation of pre-mRNA alternative splicing in cancer. Cancer Discov. 3(11), 1228–1237. https://doi.org/10.1158/2159-8290.CD-13-0253 (2013).
Witten, J. T. & Ule, J. Understanding splicing regulation through RNA splicing maps. Trends Genet. 27(3), 89–97. https://doi.org/10.1016/j.tig.2010.12.001 (2011).
Zhu, S. et al. GOLPH3 modulates expression and alternative splicing of transcription factors associated with endometrial decidualization in human endometrial stromal cells. PeerJ 11, e15048. https://doi.org/10.7717/peerj.15048 (2023).
Deng, M. et al. YBX1 mediates alternative splicing and maternal mRNA decay during pre-implantation development. Cell Biosci. 12(1), 12. https://doi.org/10.1186/s13578-022-00743-4 (2022).
Goldman-Wohl, D. et al. Trophoblast lineage specific expression of the alternative splicing factor RBFOX2 suggests a role in placental development. Placenta 100, 142–149. https://doi.org/10.1016/j.placenta.2020.07.004 (2020).
Nie, G. Y. et al. A potential molecular mechanism for regulating pre-mRNA splicing of implantation-related genes through unique uterine expression of splicing factor SC35 in women and rhesus monkeys. Reproduction 124(2), 209–217. https://doi.org/10.1530/rep.0.1240209 (2002).
Balasundaram, P., Farhana, A. Immunology at the Maternal-Fetal Interface. StatPearls. (2023).
Robertson, S. A., Moldenhauer, L. M., Green, E. S., Care, A. S. & Hull, M. L. Immune determinants of endometrial receptivity: A biological perspective. Fertil. Steril. 117(6), 1107–1120. https://doi.org/10.1016/j.fertnstert.2022.04.023 (2022).
Aluvihare, V. R., Kallikourdis, M. & Betz, A. G. Regulatory T cells mediate maternal tolerance to the fetus. Nat. Immunol. 5(3), 266–271. https://doi.org/10.1038/ni1037 (2004).
Faas, M. M. & de Vos, P. Uterine NK cells and macrophages in pregnancy. Placenta 56, 44–52. https://doi.org/10.1016/j.placenta.2017.03.001 (2017).
Plaks, V. et al. Uterine DCs are crucial for decidua formation during embryo implantation in mice. J. Clin. Invest. 118(12), 3954–3965. https://doi.org/10.1172/JCI36682 (2008).
Li, B., Duan, H., Wang, S., Wu, J. & Li, Y. Establishment of an artificial neural network model using immune-infiltration related factors for endometrial receptivity assessment. Vaccines (Basel) https://doi.org/10.3390/vaccines10020139 (2022).
Li, B., Duan, H., Wang, S., Wu, J. & Li, Y. Gradient boosting machine learning model for defective endometrial receptivity prediction by macrophage-endometrium interaction modules. Front. Immunol. 13, 842607. https://doi.org/10.3389/fimmu.2022.842607 (2022).
Ledee, N. et al. The uterine immune profile may help women with repeated unexplained embryo implantation failure after In Vitro Fertilization. Am. J. Reprod. Immunol. 75(3), 388–401. https://doi.org/10.1111/aji.12483 (2016).
Ahmadi, M. et al. Regulatory T cells improve pregnancy rate in RIF patients after additional IVIG treatment. Syst. Biol. Reprod. Med. 63(6), 350–359. https://doi.org/10.1080/19396368.2017.1390007 (2017).
Sung, N. et al. Intravenous immunoglobulin G in women with reproductive failure: The Korean Society for Reproductive Immunology practice guidelines. Clin. Exp. Reprod. Med. 44(1), 1–7. https://doi.org/10.5653/cerm.2017.44.1.1 (2017).
Shen, S. et al. rMATS: Robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl. Acad. Sci. U. S. A. 111(51), E5593–E5601. https://doi.org/10.1073/pnas.1419161111 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Aktas Samur, A. et al. In-depth analysis of alternative splicing landscape in multiple myeloma and potential role of dysregulated splicing factors. Blood Cancer J. 12(12), 171. https://doi.org/10.1038/s41408-022-00759-6 (2022).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12(5), 453–457. https://doi.org/10.1038/nmeth.3337 (2015).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Corsello, S. M. et al. The drug repurposing hub: A next-generation drug library and information resource. Nat. Med. 23(4), 405–408. https://doi.org/10.1038/nm.4306 (2017).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 46(D1), D1074–D1082. https://doi.org/10.1093/nar/gkx1037 (2018).
Bhagwat, S. R. et al. Endometrial receptivity: A revisit to functional genomics studies on human endometrium and creation of HGEx-ERdb. PLoS One 8(3), e58419. https://doi.org/10.1371/journal.pone.0058419 (2013).
Paule, S. et al. Proprotein convertase 5/6 cleaves platelet-derived growth factor A in the human endometrium in preparation for embryo implantation. Mol. Hum. Reprod. 21(3), 262–270. https://doi.org/10.1093/molehr/gau109 (2015).
Governini, L., Luongo, F. P., Haxhiu, A., Piomboni, P. & Luddi, A. Main actors behind the endometrial receptivity and successful implantation. Tissue Cell 73, 101656. https://doi.org/10.1016/j.tice.2021.101656 (2021).
Luan, L., Ding, T., Stinnett, A., Reese, J. & Paria, B. C. Adherens junction proteins in the hamster uterus: Their contributions to the success of implantation. Biol. Reprod. 85(5), 996–1004. https://doi.org/10.1095/biolreprod.110.090126 (2011).
Buck, V. U., Windoffer, R., Leube, R. E. & Classen-Linke, I. Redistribution of adhering junctions in human endometrial epithelial cells during the implantation window of the menstrual cycle. Histochem. Cell Biol. 137(6), 777–790. https://doi.org/10.1007/s00418-012-0929-0 (2012).
Mukherjee, N., Sharma, R. & Modi, D. Immune alterations in recurrent implantation failure. Am. J. Reprod. Immunol. 89(2), e13563. https://doi.org/10.1111/aji.13563 (2023).
Stope, M. B., Mustea, A., Sanger, N. & Einenkel, R. Immune cell functionality during decidualization and potential clinical application. Life https://doi.org/10.3390/life13051097 (2023).
Wang, H. et al. A positive feedback self-regulatory loop between miR-210 and HIF-1alpha mediated by CPEB2 is involved in trophoblast syncytialization: implication of trophoblast malfunction in preeclampsiadagger. Biol. Reprod. 102(3), 560–570. https://doi.org/10.1093/biolre/ioz196 (2020).
Jeong, Y., Lee, S. & Choi, I. Regulation of Tjp1 mRNA by CPEB2 for tight junction assembly in mouse blastocyst. Reproduction 163(4), 233–240. https://doi.org/10.1530/REP-21-0227 (2022).
Wang, W. et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle. Nat. Med. 26(10), 1644–1653. https://doi.org/10.1038/s41591-020-1040-z (2020).
Lai, Z. Z. et al. Single-cell transcriptome profiling of the human endometrium of patients with recurrent implantation failure. Theranostics 12(15), 6527–6547. https://doi.org/10.7150/thno.74053 (2022).
Cao, D. et al. Time-series single-cell transcriptomic profiling of luteal-phase endometrium uncovers dynamic characteristics and its dysregulation in recurrent implantation failures. Nat. Commun. 16(1), 137. https://doi.org/10.1038/s41467-024-55419-z (2025).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563(7731), 347–353. https://doi.org/10.1038/s41586-018-0698-6 (2018).
Acknowledgements
The authors thank all study participants and acknowledge the technical support from the laboratory staff.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 82171644).
Author information
Authors and Affiliations
Contributions
Mao-li Wang and Bing-jie Lu designed the research plan, finished experiments and wrote the manuscript. Xiang Lu, Lu Li and Xiao-xi Sun recruited families and collected samples. Hu Liu assisted parts of experiments. Jiucheng- Chen and Chen Chen analyzed datasets. Da-jin Li and Han Wu provided meaningful discussion of key points, gave guidance in whole study and revised the manuscript. Wen-bi Zhang recruited families and collected samples, designed the research plan and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of the Shanghai Ji Ai Genetics and IVF Institute, Obstetrics and Gynecology Hospital of Fudan University (Approval No. JIAI E2019-04; JIAI E2020-015). All participants provided written informed consent prior to sample collection. All procedures involving human participants were performed in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Consent for publication
Written informed consent for publication of potentially identifiable clinical information/images (if any) was obtained from the participant(s).
Capsule
Alternative splicing events and immune cell dynamics are associated with endometrial receptivity in patients with recurrent implantation failure, highlighting potential molecular mechanisms for personalized therapeutic strategies.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Ml., Lu, Bj., Lu, X. et al. Alternative splicing and differential gene expression during changes in endometrial receptivity in patients with recurrent implantation failure. Sci Rep 16, 9754 (2026). https://doi.org/10.1038/s41598-026-40386-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40386-w