Abstract
The WNK-OSR1/SPAK protein kinase pathway regulates ion homeostasis and cell volume, but its other functions are not well understood. To discover undefined signaling functions, we utilized experimentally-derived binding specificity to predict interactions and relative affinities with the conserved C-terminal (CCT) domains of OSR1 and SPAK, which bind short linear motifs. The upstream kinases WNKs 1-4 and their relatives, the pseudokinases NRBP1/2, also contain CCT-like domains which have conserved folds and motif binding pockets. Motifs were scored using peptide arrays, conservation, cytosolic localization, and solvent accessibility. Out of nearly 3700 motifs in the human proteome, 90% of previously published motifs ranked in the top 2% of those predicted. Interactions with selected candidates, including TSC22D1 and CAVIN1, were validated, and their localization and/or modifications were coupled to changes in WNK1 signaling. We also identified additional motif variants and confirmed binding to the NRBP1 CCT-like domain. Our results stress the diverse functionality of CCT/CCT-like domains and implicates unexpected interactions driving WNK biology.
Similar content being viewed by others
Introduction
The WNK [with no lysine (K)] signaling cascade is comprised of the four WNK kinases (WNKs 1–4) and their downstream substrate kinases, oxidative stress responsive 1 (OSR1) and SPS/STE20-related proline-alanine-rich kinase (SPAK). Ancient and pleiotropic kinases, WNKs are so named due to their unique placement of a catalytic lysine required for ATP binding compared to other members of the eukaryotic kinase superfamily1,2. Mutations that cause overexpression of WNK1 and WNK4 cause an inherited form of hypertension, pseudohypoaldosteronism type II, revealing control of blood pressure as an important function of these kinases3. Further demonstrating its importance in the cardiovascular system, disruption of the WNK1 gene in mice showed its requirement in the endothelium for cardiovascular development4.
Subsequent studies found that WNKs interact with and phosphorylate OSR1 and SPAK, enabling OSR1 and SPAK to regulate substrates that control ion transport and cell volume5,6,7,8,9,10,11,12,13,14,15,16,17,18,19. Although the actions of the WNK-OSR1/SPAK pathway as a central regulator of ion homeostasis have been extensively studied, its other functions are less defined mechanistically. Notably, how loss-of-function mutations in WNK1 cause a form of hereditary and sensory neuropathy (HSN2) remains uncertain20. Previous studies have suggested inputs of WNK1 to metabolic pathways, and others have identified single-nucleotide polymorphisms in WNK genes not only in hypertensive disease but also, for example, in neurological syndromes, including autism and schizophrenia21,22,23,24,25,26,27. WNK kinases have been linked to numerous cellular activities, suggesting they are multifaceted in the processes they regulate. Examples include but are not limited to autophagy, cell migration, insulin signaling, fat metabolism, neuronal signaling, and immune system function20,25,28,29,30,31.
Our goal was to deduce understudied signaling functions of the pathway by analyzing the capacity of the kinases OSR1 and SPAK to recognize short sequence motifs in the proteome through their conserved C-terminal (CCT) protein interaction domains. Because WNK kinases also contain two CCT-like domains, with a conserved fold and binding pocket, our findings may also yield insights into direct WNK interactions. We characterized the specificity of CCT domain binding to the core consensus sequence R-F-x-V/I found in WNKs, substrate ion cotransporters, and many other proteins7,8,10,11,32,33,34, as well as motifs with an alternative placement of the arginine (R-x-F-x-V/I), which we previously showed also bind to CCTs in a subset of potassium channels19. After noting that identities of the flanking residues surrounding the core motifs contribute significantly to overall binding, we used specificity data to identify interaction motifs in human proteins, ranked them by predicted interaction strength, and tested selected protein targets.
Our results have deciphered additional features of CCT-motif interactions that expand knowledge of the already large potential of the CCT and CCT-like domains to connect WNK signaling to cellular regulatory events. OSR1 and SPAK protein interactions alone, account for a rich and understudied source of regulatory power for the WNK pathway.
Results
Core motif and flanking residues participate in OSR1 and SPAK CCT interaction specificity
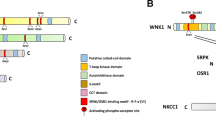
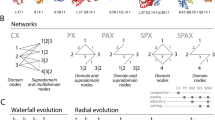
The OSR1 and SPAK CCT domains are ~90 residues at the C-terminus of both kinases, comprised of a β-sheet and two α-helices (Fig. 1A, B). Structural data are available for some of the related domains, two of which are found in each WNK (CCTL1 and CCTL2). The crystal structures of the OSR1 CCT bound to a GRFQVT hexapeptide (PDB ID: 2V3S) and the related domain, WNK2 CCTL1, bound to a longer WNK1-derived RRFxV-containing peptide that engages the residues R-x-F-x-V (PDB ID: 6FBK, Structural Genomics Consortium) show that residues around the core motif binding pocket associate in part through β-strand addition; backbone atoms from a β-strand in one binding partner interact with backbone atoms in a β-strand from the other partner, increasing binding energy (Fig. 1B)33. In this binding mode, adjacent residues within and around the core motif alternate direction.
A Domains of SPAK (STK39) and OSR1 (OXSR1). Conserved C-terminal (CCT) domains interact with R-F-x-V/I and R-x-F-x-V/I motifs. B Crystal structure of the complex of OSR1 CCT and a WNK1-derived hexapeptide, GRFQVT (PDB ID: 2V3S). The peptide interacts partially as a terminal β-strand (β-strand addition), causing sidechains to alternate direction, suggesting adjacent residues do not strongly influence each other’s binding. C Motif numbering scheme. D Peptide array setup. Peptides are covalently linked to the membrane. Each spot represents a single position within the base sequence that was varied to every amino acid. Primary anti-His6 antibody detects bound CCT. Fluorophore-labeled secondary antibody detects primary. E Quantification of peptide array spot intensities. Original array images in Supp. Fig. 1. Asterisk (*) above red residue indicates position varied for every amino acid. Caret (^) below residue indicates site in base peptide that deviates from wild-type sequence. The number to the right indicates the motif position mutated. Top (blue) row: hWNK4 (human WNK4) 1012–1024 G1215A. Middle (orange) rows: wild-type hWNK4 1012–1024. Bottom (purple) rows: hWNK1 1253–1265 R1257A (R-x-F-x-V/I motif; wild-type hWNK1 sequence contains both motifs, R-R-F-x-V/I). See main text for detailed explanations. n = 2 arrays per CCT domain, normalized by the mean intensity of the array. Ranges reported in Supp. Fig. 2. F Affinity determined by fluorescence anisotropy peptide competition with titrated, unlabeled peptides displacing 25 nM labeled peptide (NH3+-NLVGRF-[DAP-FAM]-VSPVPE-COO−] [diaminopropionic acid (DAP)]. SPAK/OSR1 CCTs are constant at 1.5/3.0 μM. Red letters mutated relative to wild-type sequence, and blue letter is position 0. The top group is hWNK4 1012–1024, the bottom is hWNK1 1253–1265. Affinity reported as inhibition dissociation constant (Ki, µM). Upper/lower limits of the 95% confidence interval are indicated. Fold-change decrease in affinity is a fold-change increase in Ki relative to wild type. Goodness of fit reported as R-squared (R2). n = 3. Curves and fluorescent probe binding in Supp. Fig. 3. GraphPad Prism 10, one-site fit to the Ki model used for data analysis. Asterisk (*) indicates previously published19.
We used peptide arrays in which the base peptide was mutated at each position to all other amino acids (Fig. 1D, E). We designated F as the “0” position as it is the most invariant, residues N-terminal as negative (−1, −2, etc…) and residues C-terminal as positive (+1, +2, etc…) (Fig. 1C). The wild-type WNK4 sequence QLVGRFQVTSSKE (1012-1024) was the primary sequence used for analysis. WNK4 QLVARFQVTSSKE (1012–1024 G1215A) was used to interrogate the role of the −3 position in the presence of the less flexible alanine at the −2 position, and WNK1 SAGRAFIVSPVPE (1253-65 R1257A, wild type: SAGRRFIVSPVPE) was used to examine the contribution of residues −1 and −3 around the alternatively placed arginine in the −2 position in R-x-F-x-V/I motifs (Fig. 1C–E, Supp. Figs. 1, 2, and Supp. Data 1). The binding specificities of the OSR1 and SPAK CCT domains are very similar due to conservation of primary motif binding pocket residues32,33,35.
Both OSR1 and SPAK have strong preferences for the core motif residues; the preference for threonine at the −1 position is an artifact as described later (Fig. 1F and Supp. Data 3). Proline is strongly disfavored in and around the core motif, because it would disrupt β-strand addition. In the WNK4 R-F-x-V motif peptide, the strong preference for only glycine at −2 by OSR1 but not by SPAK suggests that OSR1 may preferentially bind −2 glycine motifs, while SPAK is more promiscuous. Although the preference for −2 glycine is more pronounced for OSR1, both kinases still have an increased preference for serine, threonine, histidine, and asparagine, and this matches the residue identity at the −2 position for some previous experimentally validated motifs (Fig. 2B and Supp. Fig. 4A). For most other residues there are more subtle differences that likely have an additive effect on affinity (Supp. Fig. 2). Of the 27 known motifs, proline is present at the +4 position in six while a basic residue is present in five indicating an additional agreement between our peptide array results and previously identified motifs (Figs. 1E, 2B, Supp. Fig. 4A, and Supp. Data 3). Residues around the −2 arginine in the R1257A (R-x-F-x-V) mutant peptide have less pronounced effects (Fig. 1E, Supp Fig. 2, and Supp. Data 3). In the context of the WNK1 but not the WNK4 peptides, arginine is strongly preferred at the −3 position. This result suggests that much like the presence of the −2 arginine in R-x-F-x-V motifs, −3 arginines may also impart additional binding energy or possibly even bypass the requirement for a −1 arginine in certain contexts. Finally, the amino acid identities at adjacent or even distant positions around the core motif may influence the selectivity for certain amino acids either by directly influencing binding at other positions, or indirectly by increasing or decreasing binding affinity such that selectivity at other positions is altered.
A Total motifs in the initial search depending on motif type (R-F-x-V/I or R-x-F-x-V/I). B Multiple sequence alignment of all experimentally identified motifs identified from the literature5,7,8,9,10,11,12,13,16,19,33,34,44,63,76,77,78,79,80,81,82. Literature references for each motif are in the Supp. Fig. 4A. Alignments generated with Clustal Omega (clustal.org). Sequence logo generated with WebLogo (weblogo.berkeley.edu). C Flowcharts for process predicting CCT interactions and with scoring information in Supp. Fig. 4B.
To provide a more quantitative measure of binding affinity, we used fluorescence anisotropy-based peptide competition to quantify binding to a select group of sequences utilizing the same two base peptides derived from WNK4 and WNK1 (Fig. 1F, Supp. Fig. 3A, and Supp. Data 3). After optimization, we chose the probe NH3+-NLVGRF-[DAP-FAM]-VSPVPE-COO-, which combines the N-terminus of the WNK4 peptide and the C-terminus of the WNK1 peptide with an internal diaminopropionic acid conjugated to the fluorophore FAM and the N-terminal glutamine mutated to asparagine for enhanced stability (Supp. Fig. 3B, C and Supp. Data 3). Peptide affinities much higher than the KD of the probe peptide (SPAK: 0.67 µM, OSR1: 1.5 µM) cannot be reliably measured; however, none of the peptides appeared to cross this threshold (no sharper than expected drops in anisotropic signal during peptide titration) (Supp. Fig. 3A and Supp. Data 3).
SPAK CCT has a several-fold higher affinity for peptides compared to the OSR1 CCT, consistent with its stronger binding by peptide array (Fig. 1E, F, Supp. Figs. 1–3, and Supp. Data 3). For the WNK4 peptide, the Ki for OSR1 and SPAK CCT domains were 12.9 and 7.2 µM, respectively. Mutation of the branched chain amino acids at the −4 and −3 positions to alanine, and glycine to alanine or serine at the −2 position resulted in decreased binding. Mutation of the −1 arginine to lysine or threonine effectively prevented the interaction, contrary to what had been observed in the peptide array for threonine, suggesting it was an artifact (Fig. 1E, F and Supp. Data 3). Mutation of the most C-terminal amino acid at the +7 position led to almost no change in OSR1 but a 2.5-fold increase in affinity for the SPAK CCT, suggesting residues that interact outside of the primary binding pocket provide some CCT domain selectivity. The other mutations had a wide range of effects from modest to some with greater than an order of magnitude loss in affinity (Fig. 1F, also see Supp. Fig. 3A and Supp. Data 3). For the WNK1 peptide, the Ki for OSR1 and SPAK CCT domains were 5.1 and 2.5 µM, respectively (Fig. 1F and Supp. Data 3) as noted19. The other mutations had a small but substantial loss in affinity (also see Supp. Fig. 3A and Supp. Data 3). We tested serine phosphorylation at the +3 position since a previous report had indicated that phosphorylation at this position decreased binding affinity by orders of magnitude for a GRFQVpT hexapeptide33. Using the longer WNK1-derived peptide, we observed a modest, approximately 2.5-fold decrease in affinity, indicating that phosphorylation at the +3 position can modulate binding affinity, but to a lesser extent than previously reported.
Altogether, our peptide array and fluorescence anisotropy results suggest that residues outside the core motif contribute significantly to interaction specificity. In some cases, effects on binding affinity may be due to the presence of one or two strongly favored/disfavored amino acids, in other cases modest, additive effects of multiple residues may be required to provide the necessary specificity, and binding may be influenced by nearby residues.
CCT domain specificity predicts potential interactors
We applied these data to sort the >3700 instances of R-F-x-V/I and R-x-F-x-V/I motifs in the human UniProt database (uniprot.org) by predicted relative interaction affinities (Fig. 2A). Multiple sequence alignment of previously validated motifs suggests some conserved motif signatures such as −3 valine, −2 glycine/serine, and +3 serine/threonine, but with only 27 motifs verified, not enough sequence space was covered to reliably rank motifs (Fig. 2B and Supp. Fig. 4A).
We generated scoring matrices from the OSR1 CCT array data (Fig. 1E and Supp. Data 3), using three motif classes. G-R-F-x-V/I, ∆gly-R-F-x-V/I (all residues but G at −2), and R-x-F-x-V/I (Fig. 2C and Supp. Fig. 4B). We utilized the Scansite3 web server (scansite3.mit.edu) to score all identified motifs based on our input scoring matrices (Fig. 2C, Supp. Fig. 4B, and Supp. Data 1). One limitation of our use of Scansite is that motifs with a central motif phenylalanine within six residues of the N- or C-termini of the protein were not included in the final analysis. However, these motifs were still listed for reference (Supp. Data 1). Most previously validated motifs clustered near the top of our lists ranked by predicted relative motif binding affinity, thus providing an internal metric that indicated our initial method of ranking based solely on sequence worked well (Fig. 3A, B).
A Plots comparing scores of all identified motifs within motif classes found using the Scansite3 web server (scansite3.mit.edu). A lower score indicates a higher predicted relative binding affinity. Orange rectangles indicate previous experimentally validated interactions (direct binding or inferred from experimental results (Fig. 2B and Supp. Fig. 4A)). Dashed lines indicate cut-offs used for further analysis (false-positive filtering and subsequent scoring). B Similar to (A) but without scores included on the y-axis. Previously validated motifs cluster near the top of the rankings. C Summary of motifs retained at each step. D Overall scores of all retained motifs subjected to the final round of scoring. Blue line indicates all 501 scored motifs. Red triangles indicate positions of previously validated motifs. Asterisks (*) indicate motifs initially identified in our Scansite3 analysis that led to publication of validated interactions prior to publication of this study19,44. The inset graph depicts motifs ranked 1 to 80 for clarity. E Histogram depicting distribution of scores. Each bin represents the total number of motifs (blue) and previously validated motifs (red) within the bin. Bin size is 5 points. Total motifs per bin listed below. F Gene ontology (GO) analysis of the top 50 motifs.
Predicted interactions following refinement
We analyzed all G-R-F-x-V/I motifs because there were relatively few with glycine at the −2 position. For the ∆gly-R-F-x-V/I and R-x-F-x-V/I, we only analyzed the top 20% of each group. In total, 872 motifs were analyzed (Fig. 3C). To refine the prediction of OSR1 and SPAK CCT interaction motifs, we used a 100-point scale based on solvent accessibility, conservation, cellular localization, and the initial Scansite motif score (Figs. 2C, 3A, B, Supp. Fig. 4B, and Supp. Data 1). Of the 872 motifs analyzed, 371 were discarded due to lack of conservation or to localization outside the cytosol or nucleus, leaving 501 scored (Fig. 3C and Supp. Fig. 4B). 24 of the 27 previously validated motifs were present within the top 78 ranked motifs (top 2.1% of all 3675 motifs), indicating high quality of the scoring method (Fig. 3D, E).
The top 125 motifs are presented in Table 1. Previously validated motifs are indicated to illustrate that most are ranked near the top. We also analyzed the amino acid frequency at each position for each motif class within the top 125 (Supp. Fig. 4C). The presence of some amino acids such as arginine and serine at +1, serine and threonine at +3, and proline at +4 matches what has been previously observed in validated motifs (Fig. 2B and Supp. Fig. 4A). However, much like the previously validated motifs, a variety of amino acids are represented at each position.
Our results also suggest the presence of many additional uncharacterized OSR1 and SPAK CCT binding partners since the majority of motifs clustered with the previously validated motifs have yet to be described (Table 1 and Fig. 3D, E). Gene ontology (GO) analysis of the top 50 genes associated with the highest ranked motifs indicates a range of biological processes (Fig. 3F and Supp. Data 2)36. Some, such as protein phosphorylation, cell volume, and ion homeostasis, are expected. Less anticipated from the WNK literature are categories such as DNA-template transcription initiation and p53 signaling. Therefore, our results imply there are underexplored areas of biology associated with WNK signaling, which we then sought to confirm by experimentally validating some of our predicted interactions.
Validation of a subset of predicted interactions
We validated subsets of interactions biophysically by fluorescence anisotropy and in mammalian cell lysates by co-immunoprecipitation (Figs. 4, 5 and Supp. Data 3). We tested proteins either because of intriguing data in the literature or due to our preliminary data that suggested potential relationships to WNK signaling. Several were a focus because of their connections, like WNK1, to blood pressure regulation. These include NOS3 (eNOS, endothelial nitric oxide synthase), a central regulator of the cardiovascular system and blood pressure; decreased NOS3 activity has been linked to increased risk of developing hypertension and a variety of other pathologies37. The G protein-coupled receptor kinase 4 (GRK4) regulates sodium handling in the kidney via dopaminergic and renin–angiotensin signaling pathways, and mutations in the protein have been linked to hypertension38,39. The K+-Cl- cotransporter 4 (KCC4) is a well-documented substrate of OSR1 and SPAK, but to our knowledge, the motif variant R-x-F-x-V had not been implicated in the interaction of the proteins6,40. A family member (TSC22D3) of the protein TSC22 domain family protein 1 (TSC22D1) has been shown in a mouse knockout model to regulate Na+/K+ balance in the kidney, and WNK1 expression41,42. TSC22D2 was recently found to colocalize in cellular condensates with SPAK and WNKs43. WNK1 also has well-documented actions on protein localization, and two of the predicted interactors were selected due to their known actions on protein trafficking. TBC1 domain family member 4 (TBC1D4) participates in the regulation of surface expression of glucose transporters, and WNK1 has been implicated in TBC1D4 function25,44. Cavin1 is an essential component of caveolae, which are important mediators of protein localization. Rho guanine nucleotide exchange factor 5 (ARHGEF5) was tested because, like WNK1, it is involved in the regulation of cell morphology and the actin cytoskeleton45 and the autophagy-related protein 9A (ATG9A), an essential component of the autophagy machinery, was tested because we previously showed that WNK1 inhibits autophagy31. Glycogen phosphorylase, liver form (PYGL), catalyzes the cleavage of glycogen into glucose monomers, which could be a means to regulate cellular osmolarity. Known interactors WNK1 and NKCC2 (Na+-K+-2Cl- cotransporter) were used as controls.
A AlphaFold structural predictions of motifs in the proteins57,58. Images generated using AlphaFold 3D Viewer. Green sidechains with red lettering indicate core motifs. Structure coloring indicates model confidence. Dark blue: very high, light blue: high, yellow: low, orange: very low. Low model confidence can indicate the region is unstructured in isolation. Model confidence was not taken into account during structural analysis. B Affinity determined by fluorescence anisotropy peptide competition assays (see Fig. 1F and Supp. Fig. 3A). OSR1 CCT WT peptide: solid red line; OSR1 CCT V/I→A: dashed red line; SPAK CCT WT peptide: solid blue line; SPAK CCT V/I→A: dashed blue line. Flanking arginines are added to some peptides to enhance solubility. Error bars represent a 95% confidence interval. Individual data points not included for clarity. Data points can be found in the Supp. Data 3 (n = 3). C Quantitation of results in (B). n = 3 (see Fig. 1F).
A Overexpressed FLAG-hSPAK 50-545 bait protein pre-loaded on beads pulled down overexpressed myc-tagged prey proteins (HA tag for ATG9A) from HEK293T cell lysates. 100 µM CCT blocking peptide (WNK1 1253–1265; NH3+-SAGRRFIVSPVPE-COO−) as control. Supp. Fig. 5 presents the prey input and co-IP bands at identical detection intensities for comparison of the amount of protein remaining after wash steps. B Quantification of (A). n ≥ 3. Error bars indicate 95% confidence intervals. Replicates shown as dots. Green is statistically significant; red is not.
13-mer peptides containing motifs from Cavin1, KCC4, NOS3, TBC1D4, GRK4, and TSC22D1 were interrogated by fluorescence anisotropy peptide competition assays (Fig. 4B, C and Supp. Data 3). Due to peptide solubility issues, not all motifs could be analyzed. From AlphaFold2 models four of the motifs appear in solvent-exposed loops, one was found in a partially exposed loop, and only KCC4 was found to be sandwiched in the middle of a β-sheet (Fig. 4A). Mutation of the V/I to alanine in the core motifs for all six proteins tested led to 1–2 orders of magnitude loss in affinity or no detectable binding for OSR1 and SPAK CCTs, strongly suggesting these peptides are binding specifically to the CCT domains through engagement of the core motif (Fig. 4B, C and Supp. Data 3).
For co-immunoprecipitation (co-IP), anti-FLAG magnetic beads pre-loaded with 3xFLAG-SPAK 50-545 (human, lacking proline-alanine rich N-terminal region) expressed in HEK293T cells were used to co-IP myc or HA-tagged proteins overexpressed in HEK293T cells. As a negative control, we used 100 µM CCT domain blocking peptide, taken from the wild-type WNK1 sequence SAGRRFIVSPVE, to compete with protein binding to the CCT domain (Fig. 5, Supp. Fig. 5, and Supp. Data 3). WNK1 and NKCC2 fragments were positive controls. The majority of the proteins tested showed a statistically significant reduction of binding in the presence of the blocking peptide (TSC22D1, Cavin1, ATG9A, NOS3, and ARHGEF5); two did not (GRK4 and PYGL). The prey input and co-IP blots are shown at different detection intensities for better visualization (prey co-IP higher exposure in most cases), but we also compared them at the same detection intensities (Supp. Fig. 5). We included SNAI2 (aka Slug) although it was not ranked due to the N-terminal position of its motif; we did observe binding sensitive to blocking peptide, but it was not statistically significant (Fig. 5 and Supp. Data 3). Based on these predictions, we also validated the interactions of OSR1 with both Sortilin and TBC1D4 (ranked 28 and 244) in another study44.
WNK signaling influences Cavin1 phosphorylation
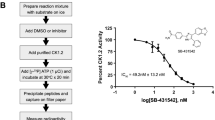
Cavin1, ranked fourth in our prediction analysis, is an essential component of caveolae, which are important mediators of the localization of proteins such as transporters, channels, and other membrane proteins (Table 1)46,47. Cavin1 exhibited strong binding to SPAK and a decrease in interaction following the addition of CCT blocking peptide (Fig. 5, Supp. Fig. 5 and Supp. Data 3). A similar interaction and blocking peptide efficacy was also observed by co-IP of endogenous OSR1 with Cavin1 in MDA-MB231 cells (Supp. Fig. 6B). Following insulin or EGF (epidermal growth factor) stimulation, Cavin1 is phosphorylated on Tyr156, the second most commonly found Cavin1 post-translational modification (PhosphoSitePlus; www.phosphosite.org) and is a few residues away from the R-x-F-x-V binding motif48,49. Inhibition of WNK signaling by either WNK1 siRNA-mediated knockdown or inhibition with the pan-WNK inhibitor WNK463 significantly decreased Tyr156 phosphorylation in human dermal microvascular endothelial cells (HDMEC) with little change in total Cavin1 protein amount (Fig. 6A–D, Supp. Fig. 6A, and Supp. Data 3).
In the experiments above, we noted that phosphorylation of SPAK, unlike OSR1, was resistant to WNK1 knockdown or the pan-WNK inhibitor. This divergence in SPAK and OSR1 activation has been observed previously in other cell lines and experimental conditions (e.g., see ref. 43). WNKs contain primarily G-R-F-x-V/I motifs, and OSR1 has a stronger preference for these types of motifs than SPAK (Fig. 1E and Supp. Data 3), suggesting preferential binding of OSR1 to WNKs as one possible explanation for this discrepancy in phosphorylation. SPAK may also be activated by a WNK-independent mechanism in some settings.
We asked whether pTyr156, which lies at the +5 position of the Cavin1 motif, might alter CCT domain binding (Fig. 6E). Two possible mechanisms for how this might occur are altering intramolecular interactions between Tyr156 and another region of Cavin1, such as adjacent basic residues at the −4, −3, −2, and/or +1 positions, or by altering intermolecular interactions between the Cavin1 motif and SPAK/OSR1 CCT domains. AlphaFold350 models of the complex between SPAK CCT and the Cavin1 13-mer peptide with or without pTyr156 phosphorylation did not indicate major structural shifts, but the Tyr156 is found near a group of charged residues on the SPAK CCT domain, which could affect binding depending on the presence of a negatively charged phosphate (Fig. 6F, left and Supp. Fig. 9A). A model of the 13-mer alone adopts a helix where the +1 lysine interacts with the +5 tyrosine phosphate and could thus provide a means of stabilizing the helical peptide in a binding-deficient conformation (Fig. 6F, right and Supp. Fig. 9A). However, models of such short length are less accurate, and this is primarily shown as an example of one possibility, and the presence of triple arginines at −4, −3, and −2 could also be another source of intramolecular interactions with pTyr156 (Fig. 6G). In our fluorescence anisotropy-based peptide competition assay, the Cavin1 motif peptide exhibited a decrease in binding affinity to both OSR1 and SPAK CCT domains when Tyr156 was phosphorylated, further suggesting phosphorylation inhibits binding (Fig. 6H and Supp. Data 3).
A–D siRNA-mediated knockdown of WNK1 (72 h) or inhibition with the pan-WNK inhibitor, WNK463 (18 h, 1 µM), in human dermal microvascular endothelial cells (HDMEC). Mean ± SD (n = 3; *P < 0.05, **P < 0.01; unpaired two-tailed t-test). E Cavin1 motif. Blue: basic/positively charged amino acids; red: acidic/negative; underlined: R-x-F-x-V; Y156 is the site of phosphorylation. F AlphaFold350 models analyzing Cavin1 pTyr156 effects on intermolecular interactions of SPAK CCT (human 446–545) with Cavin1 13-mer used in experiments and intramolecular interactions of the 13-mer alone50. Model statistics in Supp. Fig. 9A. G Diagram indicating potential intramolecular interactions (dashed lines) between acidic phospho-Tyr156 at +5 position of Cavin1 motif and multiple basic residues at the -4, -3, -2, and +1 positions. Such interactions might reduce CCT domain interactions with the motif. H Fluorescence anisotropy peptide competition (see Fig. 1F). The higher line indicates weaker binding. Quantification unavailable due to Cavin1 p-Tyr peptide instability at higher concentrations. Error bars represent a 95% confidence interval. Individual data points not included for clarity. Data points can be found in the Supp. Data 3 (n = 3).
WNKs, OSR1, and SPAK are serine/threonine kinases; thus, the change in Cavin1 Tyr156 phosphorylation occurs through an effect on a tyrosine kinase such as the EGF receptor. Interestingly, constitutively active mouse SPAK 62-556 T243E phosphorylated MBP-Cavin1 in vitro using bacterially expressed proteins, and addition of the CCT blocking peptide strongly suppressed phosphorylation (Supp. Fig. 6C). Addition of the OSR1/SPAK kinase activity-enhancing protein Mo25α (aka Cab39) also significantly increased phosphorylation, as has been shown for other substrates51. While detailed mechanisms are missing, our findings implicate WNK signaling in aspects of Cavin1 function, OSR1/SPAK as direct interactors of Cavin1, and suggest a possible mechanism for the regulation of the interaction between the kinases and Cavin1 via phosphorylation at Tyr156.
TSC22D1 interacts with OSR1 and SPAK and influences WNK pathway signaling
Several lines of evidence point to a connection between TSC22D1 and WNK signaling. TSC22D proteins were originally identified as TGF-β-regulated transcription factors, and we have previously shown a connection between them and WNK and TGF-β-mediated signaling52,53,54. A recent study found WNKs, TSC22D proteins, and SPAK are all components of phase-separated cellular condensates43. Three of the four TSC22D family members contain strongly-predicted binding motifs with ranks of 3, 8, and 77 for TSC22D1, TSC22D4, and TSC22D2, respectively (Table 1 and Fig. 7A), and all four can heterodimerize through their conserved leucine zippers55. Similar to Cavin1, TSC22D1 was one of the strongest candidates for further analysis based on our co-immunoprecipitation validation assays (Fig. 5 and Supp. Data 3). We found that endogenous TSC22D1 also co-immunoprecipitated with 3xFLAG-OSR1 and the CCT blocking peptide prevented their association in HEK293T cells (Fig. 7B). Endogenous TSC22D1 is also pulled down with GST-OSR1 CCT (Fig. 7C).
A Multiple sequence alignment of TSC22D proteins around the motif. TSC22D3 lacks the region containing the motif. B co-IP of endogenous TSC22D1 with 3xFLAG-OSR1 in HEK293T cells. 50 µM CCT blocking peptide (WNK1 1253–1265; NH3+-SAGRRFIVSPVPE-COO−) used as negative control (n = 3). C Similar to (B) except bacterially expressed GST-OSR1 CCT used for pull-down (n = 3). D Colocalization of endogenous OSR1 and stably-expressed TSC22D1-mNeonGreen in A549 cells ±1 µM pan-WNK inhibitor, WNK463, treated 24 h. Scale bar: 20 µm. (n = 3). E, F Western blot of partial siRNA-mediated knockdown of TSC22D1 in HDMEC after 48 h siRNA treatment. Mean ± SEM (n = 3; *P < 0.05, **P < 0.01, ***P < 0.001; unpaired two-tailed t-test). pSPAK/pOSR1 blot shown twice at varying intensities.
We were unable to validate the TSC22D1 antibodies for immunolocalization, even though they worked well for immunoblotting. As a result, we tagged TSC22D1 with mNeonGreen and made stable A549 lung cancer and HEK293T cell lines expressing the tagged protein (TSC22D1-mNeonGreen). Treatment with the pan-WNK inhibitor, WNK463 (1 µM, 24 h), induced TSC22D1 puncta in both cell types (Fig. 7D and Supp. Fig. 7A). Puncta were smaller and more evenly distributed in A549 cells compared to HEK293T cells. In A549 cells endogenous OSR1 colocalized in puncta with tagged TSC22D1 in the presence of WNK463 (Fig. 7D). In HEK293T cells, treatment with hypotonic media also led to a decrease in TSC22D1 puncta size (Supp. Fig. 7A). These puncta containing TSC22D1 and OSR1 fit well with the findings published by Moffat and colleagues on puncta containing WNKs, TSC22D proteins, and SPAK43.
Partial siRNA-mediated knockdown of TSC22D1 significantly altered amounts of WNK1 and OSR1 in HDMEC (Fig. 7E, F and Supp. Data 3). OSR1 and SPAK activation were also significantly increased, but the increase in phosphorylation is likely related to increased protein expression, as there was no change when pOSR1 was normalized to total OSR1 (Fig. 7E and Supp. Data 3). Treatment of HDMEC with WNK463 also altered mRNA amounts of TSC22D family members (Supp. Fig. 7B and Supp. Data 3). Taken together, these findings point to OSR1 and SPAK directly interacting with TSC22D family members, and a connection between the expression of TSC22D family members and components of the WNK pathway.
CCT-like domains and alternative binding motifs utilize a common binding mode
Each of the four WNKs contains two CCT-like (CCTL) domains (CCTL1 and CCTL2), as was initially reported for WNK1 and later WNK2 and WNK3 (Structural Genomics Consortium)32,56. Structural predictions provided by AlphaFold2 indicate all WNK CCTL domains share a common structure and motif binding pocket with OSR1 and SPAK CCT domains57,58. Using AlphaFold2, we also found that NRBP1 and NRBP2, which are the closest relatives to WNKs in the human kinome tree, also contain a CCTL domain (Fig. 8A, B)59. WNK1 CCTL1 and WNK2 CCTL1 have been shown to interact with motif-containing peptides in an in vitro setting using purified proteins, although no physiological evidence for a function of motif:CCTL interactions has been demonstrated56. While there are slight differences in the topology of CCT and CCT-like domains, the overall folds are strikingly similar; particularly the core motif binding pocket composed of a β-strand followed by a loop and then an α-helix, where all the hydrophobic and acidic residues that bind the core motif are located (Fig. 8C–F). Utilizing AlphaFold350 we generated models of TSC22D1 R-F-x-V motif interactions with OSR1 CCT, WNK1 CCTL1, and NRBP1 CCTL (Fig. 8D)58,60. Previous work studying orthologs of TSC22D and NRBP proteins in Drosophila identified regions within the proteins corresponding to the R-F-x-V motif and CCTL domain that co-immunoprecipitated61. A recent study identified similar interacting regions in human TSC22 and NRBP proteins, but based solely on AlphaFold predictions, suggested the mode of interaction was not via an R-F-x-V motif but by an R-W-x-C motif that binds in a nearly identical conformation43. Interestingly, depending on the version of AlphaFold we used (AlphaFold 2 vs 3), we obtained both types of interactions when using full-length proteins (Supp. Fig. 9B). Our models utilize a fragment of TSC22D1 that only encompasses the region around the R-F-x-V motif in order to obtain the R-F-x-V interaction for NRBP1 using AlphaFold3, but we did obtain similar models using full-length proteins. Additionally, TSC22D2, WNK1, and NRBP1 share the highest degree of codependence in CRISPR and RNAi screens according to DepMap.org. These findings suggested that NRBP CCTL domains might interact with TSC22D R-F-x-V motifs (Fig. 7A) in a manner similar to OSR1 and SPAK CCT domains.
A Region of the human kinome tree containing WNK1-4, OSR1, SPAK, and NRBP1/2; all proteins contain CCT or CCT-like (CCTL) domains59. B Diagram depicting locations of CCT and CCTL domains in proteins. C Multiple sequence alignment of regions of CCT/CCTL domains that comprise motif binding pockets consisting of β-strand–loop–α-helix. Conserved binding residues indicated. D Comparison of crystal structure of OSR1 CCT:GRFQVT hexapeptide complex (left, PDB:2V3S) with AlphaFold3-derived models of OSR1 CCT (430–527), WNK1 CCTL1 (human, 482–597), and NRBP1 CCTL (436–535) bound to TSC22D1 RFxV motif (395–419)50,57,58,60. For NRBP1:TSC22D1, when using full-length proteins, differing models were obtained depending on the AlphaFold modeling server used. Both results can be found in the Supp. Fig. 9B. Identical results were obtained using truncated vs full-length proteins for OSR1 and WNK1. E, F Structural alignment of AlphaFold3 models in (D). Binding residues depicted as sticks. G Model of SPAK CCT:GRFQVT complex made by overlaying SPAK CCT structure (PDB: 7O86) onto the OSR1 CCT:GRFQVT complex structure. D477 (green) was mutated in the present study to inhibit binding while L491 (green) was mutated in previous study62. H, I 3xFLAG-tagged SPAK 50-545 ± D477Q or NRBP1 ± E488Q used to co-IP myc-NKCC2 1-170 and myc-TSC22D1, respectively, in HEK293T cells. Mean ± SEM of mutants relative to WT (n = 3, 95% confidence intervals shown). J Affinity determined by fluorescence anisotropy peptide competition assays (n = 3, see Fig. 1F and Supp. Fig. 3A. Asterisk (*) indicates results previously published19.
We identified a point mutation in the SPAK CCT domain that significantly diminished motif binding. Previous work in mice had identified a point mutation (mouse L502A, human L491A) that drastically reduced motif interactions (Fig. 8G)62. However, because this mutant also impacts hydrophobic interactions required for CCT domain folding, we chose to pursue an alternative mutant, D477Q (human). This point mutant neutralizes a charge and increases side chain length on a residue critical for interactions with the motif arginine (Fig. 8G)33,35. Of several mutants tested by FP with purified SPAK CCT domain, the D477Q mutant led to the largest, ~50-fold, decrease in peptide probe binding affinity (Supp. Fig. 8A and Supp. Data 3). A D477A mutant had been previously shown to diminish interactions with WNK1 and NKCC116. By co-immunoprecipitation of myc-NKCC2 1-170 with 3xFLAG-SPAK 50-545 in HEK293T cells, the mutant led to a near complete loss of interaction (Fig. 8H, I and Supp. Data 3). A similar result was observed for Cavin1 (Supp. Fig. 8B). We then utilized this same strategy in the NRBP1 CCTL domain. NRBP1 E488Q also neutralizes the charge in the position analogous to D477, although it does not alter side chain length. E488Q diminished the interaction between myc-TSC22D1 and 3xFLAG-NRBP1, suggesting that NRBP1 interacts with the TSC22D1 R-F-x-V motif in a manner similar to OSR1 and SPAK.
Beyond alternative CCTL domains, we also investigated whether alternative core binding motif sequences might be able to interact with CCT domains. We analyzed the effects of substitution of arginines in the WNK1-derived peptide to alanine or other basic residues (Fig. 8J, Supp. Fig. 3A, and Supp. Data 3). We previously showed core motif arginine at either −1 or −2 positions can bind with similar affinity19. Replacing the arginines with histidine-histidine led to an approximate sevenfold decrease in affinity, but the affinities were still in the low-mid micromolar range, which could provide enough specificity for signaling, and the same is true for histidine-alanine (11-fold decrease), but the decrease for alanine-histidine was more substantial, leading to Ki values in the high micromolar range. Surprisingly, replacement with lysine-lysine led to Ki of 16.3 and 4.2 µM for OSR1 and SPAK, respectively, which is only a slight loss in affinity compared to wild type, and while both lysine-alanine and alanine-lysine led to a further decrease in binding, all the measured affinities were still in the low-mid micromolar range. These findings suggest our current view of what constitutes a viable motif should be expanded to include these new motif variants, as well as others that may yet be discovered.
Discussion
Protein–protein interactions facilitate the formation of short- and long-lived protein complexes designed to communicate with cellular processes with specificity. WNKs activate OSR1 and SPAK by phosphorylation, enabling the kinases to phosphorylate a variety of substrates such as ion cotransporters, and many of these signaling steps require SPAK/OSR1 CCT domain interactions5,6,7,8,9,10,11,12,13,14,15,16,17,18,19. The capacity of the CCT domains to recognize many protein targets through these common short motifs enables inputs from OSR1, SPAK, and WNKs to extend to many cellular regulatory pathways, as well as providing a means to enhance kinase specificity.
More conventional screening methods used to identify direct protein:protein interactions, such as affinity-purification mass spectrometry (AP-MS), yeast two-hybrid, and BioID, are powerful, but often limited by the inclusion of false-positives (nonspecifically-bound proteins, indirect interactions, irrelevant proteins in close spatial proximity) and false-negatives (weak interactions, low expression, interaction only under certain conditions). This study was designed to minimize these limitations to identify only direct protein-protein interactions that underlie processes related to WNK pathway function, and to accelerate characterization of the essential biochemical mechanisms that encompass both dominant and more modest regulatory control that will help describe a bigger picture of WNK signaling.
We used already identified binding motifs as an internal metric for the success of our ranked predictions. Our results suggest that the method we used to rank and predict interactions worked well since the vast majority of validated interactions found from the literature ranked near the top of our predictions. Additionally, there are many new proteins that have little to no current known connection to WNK signaling. We therefore decided to experimentally validate a subset of these interactions through biochemical interaction studies and found that the majority of those tested bound, and this binding could be inhibited either via mutations of interface residues or competition with a CCT domain binding peptide. Upon more in-depth examination of two of the newly identified interacting proteins' potential roles in WNK signaling, we uncovered potential new biology.
Originally identified as TGF-β-regulated transcription factors, TSC22D proteins are intertwined with the WNK pathway from proteomics and other studies, with biochemical insights only recently being uncovered. The four TSC22D proteins heterodimerize and TSC22D1, 2, and 4 contain binding motifs that are highly ranked, and TSC22D1:CCT interactions by both peptide binding and co-IP are some of the strongest we observed. We show here that TSC22D1 can colocalize with OSR1, and its localization is sensitive to WNK inhibition. In a mouse knockout model, the Pearce lab showed that TSC22D3 (GILZ) regulates Na+/K+ balance in the kidney, and has a strong effect on WNK1 expression42. Transcription of TSC22D family members increases following hypertonic stress41. TSC22D2 and TSC22D4 interact with SPAK as well as WNKs, as deduced from large-scale mass spectrometry-based proteomics studies (BioGRID database, thebiogrid.org), and TSC22D2 was shown to interact with SPAK and WNK3 in a yeast two-hybrid screen63. Surprisingly, TSC22D2 is also the molecule with results most similar to WNK1 in analysis of siRNA and CRISPR screens from data aggregated in Depmap.org. It was from this DepMap data that the recent connection between WNKs, NRBPs, and TSC22D proteins in cellular condensates was uncovered. Although that study was initiated as a result of a different screen compared to ours, the fact that our predictions also led us to begin studying TSC22D proteins in WNK signaling further illustrates the potential of our results to uncover new insights into WNK pathway function43.
A second highly predicted and validated binder, Cavin1, is a compelling target for future investigation because its mechanosensitivity in the endothelium may be related to WNK1 functions in endothelial migration, TGF-β sensitivity54,64, and responses to mechanical stress (Jung et al., in preparation). Cavin1, ranked fourth in our prediction analysis, is an essential component of caveolae, which are important mediators of the localization of proteins such as transporters, channels, and other membrane proteins46,47. Cavin1 (aka PTRF, Polymerase I and transcript release factor), is required for efficient ribosomal biogenesis by dissociating paused RNA polymerase I from transcriptional complexes65,66. In addition to its identification as a pathway target through SPAK/OSR1 binding, our study shows the parallel control of tyrosine phosphorylation of Cavin1 by WNK1. Phosphorylation of Tyr156 has been implicated in ribosomal RNA transcriptional regulation, as a Tyr to Phe mutation reduces ribosomal RNA transcription65,66. Phosphorylation of Tyr156 can be catalyzed by receptors for epidermal growth factor (EGF) and insulin. In previous work, we showed that depletion of WNK1 results in increased degradation of the EGF receptor67. Among possible explanations for the decrease in Cavin1 pY156 occurring as a consequence of WNK1 loss, is a reduction in plasma membrane EGF receptor needed to catalyze the reaction.
In a number of cases, such as Cavin1 Tyr156, phosphorylatable residues lie in close proximity to CCT binding motifs. As noted in Fig. 2B, more than half of the already published motifs are directly followed at the +3 position by Ser or Thr. Work initially by the Alessi/van Aalten labs using short peptides suggested that CCT domain-motif binding is regulated by phosphorylation of such residues33. Although less dramatic, we also see differences in peptide binding as a result of Ser phosphorylation of a WNK1 peptide, as well as phosphorylation of Tyr at the +5 position in the Cavin1 peptide. The common occurrence of Ser/Thr/Tyr near motifs indicates that phosphorylation may be a frequent mechanism controlling CCT domain-target binding.
Some of the proteins found to interact, link the WNK1 pathway mechanistically to previously identified functions; these include sortilin and TBC1D4. We tested TBC1D4, ranked at 246, because of previous work from the Jordan lab indicating an association with WNK1 and glucose transporter trafficking25 and because of earlier studies suggesting effects of WNK1 on glucose homeostasis68. In recent work, we found that TBC1D4, also known as AS160, and a second predicted OSR1 and SPAK interactor, sortilin, ranked at 28, cooperate to maintain insulin-dependent Glut4 trafficking, uncovering one mechanism using two CCT interactions through which WNK1 can influence glucose metabolism44. In work not yet published, we were able to show that OSR1 co-immunoprecipitates with endogenous ATG9A (Kannangara, et al, in preparation), supporting the association observed here. ATG9A is a core autophagy protein that provides another connection to our earlier findings that WNK1 inhibits autophagy31.
In addition to testing highly predicted interactions, we examined binding determinants of the OSR1 and SPAK CCTs in detail to attempt to define limits of CCT-core motif interactions. We also touched upon the relevance of such motifs to interactions with CCT-like domains in WNKs and NRBPs. What we found was that sequence differences in the core motif, for example, ones in which arginine is replaced by other basic residues (e.g., 140 KK motifs in the human proteome), may lead to physiologically important motif variants with only minor decreases in affinity. We also found that sequences outside the core motif can significantly influence binding specificity to the extent that motifs that appear poor may work well because different residues at various positions can provide the requisite binding energy. Competition and multivalency may be features that most complicate what early on appeared to be a simple domain-motif association. More than one CCT or CCT-like domain may interact with the same motifs with varying degrees of specificity, providing an additional layer of complexity to signaling by both providing a means of motif competition and also by bringing different signaling proteins together, such as in the case of motif-containing proteins that oligomerize, e.g., TSC22D proteins and Cavin1, and proteins with CCT/CCT-like domains that themselves oligomerize. WNKs oligomerize, as do OSR1 and SPAK, and use phase separation to collaborate with other proteins55,69,70,71,72,73,74. We conclude that there is a high degree of complexity in how CCT and CCT-like domains contribute to signaling both within and outside the WNK pathway that exceeds a simple 1:1 relationship. As evidenced from this current work, uncovering new CCT/CCT-like interactions will lead to new biological insights related to WNK signaling.
Methods
Protein purification and peptide production
His6-tagged CCT (mouse SPAK 459–556 and human OSR1 433–527) and mouse SPAK 62-556 T243E were expressed in Rosetta (DE3) E. coli (Novagen). Cells grown at 37 °C, 225 rpm until OD600 = 1.0, then induced with 0.5 mM IPTG overnight, 20 °C, 150 rpm. Cells lysed by freeze/thaw (−20 °C), then with 1 mg/ml lysozyme in 50 mM Tris-HCl, pH 8.0, 300 mM NaCl, 10% glycerol, protease inhibitors (1 mM phenylmethanesulfonyl fluoride, 583 µM pepstatin A, 762 µM leupeptin, 10.6 mM Nα-Tosyl-L-arginine methyl ester HCl, 10.8 mM tosyl-lysine-chloromethylketone HCl, 11.3 mM Nα-benzoyl-L-arginine methyl ester carbonate, 200 µM soybean trypsin inhibitor), 30 min, 4 oC, then sonicated. Soluble proteins are applied to the Ni2+-NTA column. Eluted with 500 mM imidazole gradient. Fractions analyzed by SDS-PAGE, then dialyzed 4 °C overnight. CCTs dialyzed into 25 mM Tris-HCl, pH 7.75, 125 mM NaCl, 1 mM DTT. SPAK 62-556 T243E dialyzed into 20 mM HEPES pH 7.5 100 mM NaCl, 1 mM DTT. Proteins run on Superdex200 gel filtration. Fractions pooled. GST-Mo25α 11-333 purified similarly except 5 mM DTT, 1 mM EDTA added to lysis buffer, glutathione beads for affinity-purification followed by on-bead TEV protease cleavage to remove tag and Ni2+-NTA beads to remove TEV, and gel filtration buffer was 20 mM HEPES, pH 7.5, 100 mM NaCl, 1 mM DTT. MBP-Cavin1-His6 required special considerations for purification. Induction in Rosetta2 cells immediately arrests cell growth. Cells were grown to OD600 = 2.0, then 1 mM IPTG. Growth overnight at 16 °C, 150 rpm. Protein affinity purified similarly as His6-tagged proteins but 500 mM NaCl in lysis, then amylose resin followed by on-bead TEV cleavage in 20 mM HEPES pH 7.5, 500 mM NaCl, 1 mM DTT, protease inhibitors/EDTA. The final step was Superdex 200 gel filtration.
CCT domain specificity analysis by peptide array
Immobilized peptide membranes purchased from Kinexus Bioinformatics Corp. (~200 nmol N-terminal acetylated peptides/spot chemically synthesized on TOTD). Membranes rinsed with EtOH, then 30 min with TBST (0.1% Tween). Membranes were blocked for 60 min with 1:1 Odyssey blocking buffer to TBST (blocking solution, BLS). Membrane incubated with 300 µM His6-SPAK CCT (residues 449–545) or His6-OSR1 CCT (residues 433–527) in BLS 60 mins. Membrane washed 2 × 5 min in TBST. Membrane incubated with 1:1000 mouse anti-His6 (Clontech) for 15 min. Membrane washed 2 × 5 min in TBST, then incubated with 1:500 LI-COR IRDye 680RD mouse secondary antibody 15 min. Membrane washed 4 × 5 min in TBST. Quantification by LI-COR Odyssey. Antibody-only (background) spot intensities subtracted from CCT overlay spot intensities. Average spot intensity for each array is used to normalize values between arrays.
Peptide binding affinities by fluorescence polarization
About 1.5 µM His6-SPAK CCT or 3.0 µM His6-OSR1 CCT combined with 25 nM NH3+-NLVGRF[DAP-FAM]VSPVPE-COO- (DAP-FAM: 2,3-diaminopropionic acid conjugated to FAM) in 25 mM Tris-HCl pH 7.75, 125 mM NaCl, 1 mM DTT. Unlabeled competing peptides were then added and subjected to twofold stepwise dilution. Fluorescence polarization measurements on BioTek Synergy H1 multi-mode plate reader equipped with polarizing filter cube (Em: 485 nm, Ex: 528 nm, 510 nm dichroic mirror). Data fit to the model of one-site competitive binding to determine Ki using GraphPad Prism. Measurements of binding affinity for labeled peptides used a similar protocol, but CCT concentration varied. 1 mg/ml BSA was used for all peptides other than WNK1/WNK4 in order to prevent nonspecific binding at higher peptide concentrations. All peptides were diluted in 25 mM Tris-HCl, pH 7.75, 125 mM NaCl to 10 mM. DMSO was added to peptides with solubility issues.
Motif search, ranking, and analysis
Average spot intensities from OSR1 CCT peptide arrays for residues outside core R-F-x-V/I or R-x-F-x-V/I motifs used as input for Scansite3 web server (scansite3.mit.edu)75. The core motif phenylalanine is fixed in the center of the matrix. For positions corresponding to residues R and F in the core motif, no other residues were allowed. For the position corresponding to the V/I ratio of scores for V:I was 2. For the G-R-F-x-V/I motif class, G was also fixed, and rows 2, 3, 6, and 8–10 of the peptide array were used for scoring. For the Δgly-R-F-x-V/I motif class, all residues except G were allowed, and rows 1, 3, 6, and 8–10 were used for scoring. For the R-x-F-x-V/I motif class, rows 6, 8–12 were used for scoring. The UniProt H. sapiens database (uniport.org) was searched for all valid motifs. Motifs scored based on the inputs described above. Scores were then used to rank motifs (lower score = higher probability of interaction).
Cutoffs for additional analysis were the top 20% for Δgly-R-F-x-V/I and R-x-F-x-V/I motifs. No cutoff for G-R-F-x-V/I motifs. Motifs not conserved in Mus musculus (mouse) were discarded. Motifs not annotated in UniProt GO terms as present in either cytosol, nucleus, or within membrane protein region within or partially in the cytosol were discarded. Motifs containing a proline at +1 or +3 positions were discarded. Motifs were scored according to Supp. Fig. 4B. AlphaFold2 was used when possible; otherwise, the protein sequence was used for BLAST search of experimentally derived structures and identical or closest-related protein was used by aligning the motif to the sequence to determine the position in the structure. If a motif was present in a loop or β-hairpin that was >100 residues away from surrounding structural elements, but packed/folded, it was treated as an isolated structural element. If part of motif was present in secondary structure it was treated as that element but assigned partial loop. After total analysis, all three motif classes were combined and ranked by overall score based on a 100-point system (Supp. Fig. 4B).
Gene ontology was performed with the top 50 ranked unique genes using standard GO and GO Slim biological process sets from PantherDB.org. Detailed specifications of the analysis in Supp. Data 2. Top-of-hierarchy terms were used. Terms describing a similar process were removed for clarity.
Co-immunoprecipitation and immunoblots of potential interactors
HEK293T cells were grown in DMEM with 10% FBS. Cells were transfected with pCMV 3xFLAG-SPAK 50-545 (human) or myc-prey (various). Cells were washed and resuspended in 5 ml cold phosphate-buffered saline supplemented with 200 µM PMSF and 1:1000 protease inhibitor cocktail (PBSii). About 1 ml of 3xFLAG-SPAK supernatant was mixed with 30 µl of anti-Flag magnetic agarose beads (Pierce A36797) and incubated at 4 oC for 1 h, washed 3X with 1 ml cold PBSii using magnetic rack, followed by addition of 0.5 ml supernatant myc-tagged prey proteins. 100 µM NH3+-SAGRRFIVSPVPE-COO- blocking peptide diluted in 25 mM Tris-HCl pH 7.75, 125 mM NaCl was added to one of two samples per prey protein and incubated at 4 oC for 1 h. Samples were washed 3X with 1 ml cold PBSii. Myc-tagged proteins were eluted by the addition of 45 ul of 500 µM blocking peptide and 15 µl 5X sample buffer was added to the eluant. Bait and prey proteins were loaded on separate gels (Bio-Rad 4–20% precast gels cat. #4568094). Proteins were transferred to a nitrocellulose membrane and immunoblotted as described above. The mouse monoclonal Myc antibody was from clone 9E10. Mouse anti-FLAG M2 antibody was from Sigma (#F1804). Blots were washed 3 × 20 ml with TBST then incubated with 1:5000 IRDye 680 goat anti-Mouse secondary antibody (LI-COR # 926-68070) and analyzed by LI-COR imaging.
Kinase assay
About 15 µM mouse His6-SPAK 62-556 T243E, 15 µM Mo25α, 5 µM MBP-Cavin1-His6 (or MBP tag alone, maltose binding protein, and 100 µM CCT blocking peptide preincubated at 25 oC 15 min in 20 mM HEPES pH 7.5 150 mM NaCl (or supplemented with NaCl to bring to 300 or 500 mM), 1 mM DTT prior to addition 50 µM ATP, 15 mM MgCl2, [γ-32P]ATP. Reaction at 25 oC, 10 min. Addition of 5X Laemmli buffer and 100 oC 5 min to stop the reaction. 4–20% SDS-PAGE gel.
HDMEC culture
Human dermal microvascular endothelial cells (HDMEC, PCS-110-010) from ATCC (Manassas, VA, USA) maintained in MCDB 131 medium (15100CV, Corning, Manassas, VA, USA) with 10% FBS (F0926, Sigma-Aldrich),10 mM L-glutamine (25005CI, Corning), 10 ng/ml epidermal growth factor (PHG0311, Thermo Fisher Scientific), 1 ug/mL hydrocortisone (H0888, Sigma-Aldrich), and 1% penicillin/streptomycin (SV30010, Thermo Fisher Scientific). All cells were cultured at 37 °C, 95% humidity, 5% CO2.
Antibodies
Unless indicated elsewhere antibodies obtained from following sources: WNK1, Cobb lab Q25612; Cavin1 #69036, OSR1 #3729, and GAPDH #97166 L from Cell Signaling Technology (Beverly, MA, USA); phospho-SPAK Ser373/phospho-OSR1 Ser325 07-2273 from Millipore Sigma (Darmstadt, Germany); phospho-Cavin1 Tyr156 PA5-99575 from Thermo Fisher Scientific (Waltham, MA, USA); Actin sc-8432 from Santa Cruz Biotechnology (Santa Cruz, CA, USA); TSC22D1 A303-582A from Bethyl Labs (Montgomery, TX, USA).
siRNA knockdown
About 20 nM oligonucleotides encoding siRNA for control (Thermo Fisher Scientific, 4390844), TSC22D1 (Thermo Fisher Scientific #4392420 ID #S16891), and WNK1 (5′ CAGACAGUGCAGUAUUCACTT 3′) were transfected into HDMEC using Lipofectamine RNAiMax (Thermo Fisher Scientific, 13778150). After 48 (TSC22D1) or 72 (WNK1) hours transfection, cells were provided with respective treatments and harvested in 0.05% bromophenol blue, 0.1 M DTT, 10% glycerol, 2% SDS, 0.05 M Tris-Cl, and 5% β-mercaptoethanol.
Reverse transcription and qPCR
RNA from HDMEC was extracted using the PureLink RNA kit (Thermo Fisher Scientific, 12183018 A). RNA concentration was measured, and 1 μg was used to synthesize cDNA using iScript™ Reverse Transcription Supermix (Bio-Rad, 1708891) per manufacturer’s instructions. Resultant cDNA diluted 1:10 in nuclease-free water and concentration measured. qPCR reactions with cDNA and appropriate forward/reverse primers set up using iTaq Universal SYBR® Green Supermix (Bio-Rad, 1725121) per manufacturer’s instructions. qPCR cycle: 95 °C for 5 min, then 95 °C for 10 s and 55 °C for 30 s for 40 cycles. Custom oligos were purchased from Thermo Fisher Scientific. TSC22D1-isoform 3 (Forward Primer: 5′-ACTGCAGCCTACTCCTTGCT-3′; Reverse Primer: 5′-GAGTTCTTCTCAAGCAGCTC-3′). TSC22D3 (Forward Primer: 5′-CATGGAGGTGGCGGTCTATC-3′; Reverse Primer: 5′-CACCTCCTCTCTCACAGCGT-3’). TSC22D3- L isoform (Forward Primer: 5′-ACCGCAACATAGACCAGACC-3′; Reverse Primer: 5′-CACAGCGTACATCAGGTGGT-3′). TSC22D2 (Forward Primer: GCAGCGTCCTGACTAGATCC; Reverse Primer: GAGCTGTCAGTTCCCGACTC). 18s (Forward Primer: GTAACCCGTTGAACCCCATT; Reverse Primer: CCATCCAATCGGTAGTAGCG).
Stable cell line generation and immunofluorescence microscopy
Lentiviral vectors containing C-terminal mNeonGreen tag (VectorBuilder), and TSC22D1 (long isoform) inserted to generate TSC22D1-mNeonGreen. A549 and HEK293T cells were transfected using Lipofectamine 3000 using the manufacturer’s instructions. Lipofectamine 3000:P3000:DNA ratio was 2.5 µl:2 µl:1 µg. Cells expressing plasmids selected by growing in media containing 1 or 2 µg/µl puromycin (A549 or HEK293T, respectively). Cells were trypsinized and sorted by fluorescence with flow cytometry. For fluorescence microscopy, cells on glass coverslips were fixed with 4% (vol/vol) paraformaldehyde in PBS for 15 min at room temperature, followed by permeabilization with 0.1% Triton X-100 in PBS for 4.5 min and blocking with 10% (vol/vol) normal goat serum (Invitrogen). Incubation with primary antibodies for 1 h at room temperature or overnight at 4 °C was followed by fluorescently labeled Alexa Fluor secondary antibodies diluted 1:500 for 1 h at room temperature. Coverslips were mounted on DAPI Fluoromount-G (SouthernBiotech) medium. Fluorescent images were obtained with a Zeiss LSM880 laser scanning confocal microscope. The brightness and contrast of each channel of the image panels were adjusted similarly across experimental conditions.
AlphaFold models
Models in Fig. 4 were taken directly from the AlphaFold2 Protein Structure Database (https://alphafold.ebi.ac.uk/). Models in Fig. 6 were generated by the Google DeepMind AlphaFold3 web server (https://alphafoldserver.com). Models in Fig. 8 using full-length proteins were initially generated using the COSMIC2 web server version of AlphaFold2 Multimer (https://cosmic2.sdsc.edu:8443/gateway). Since the binding interface comprises only a small portion of the total protein and because WNK1 and TSC22D1 are primarily unstructured, we generated models using smaller fragments to obtain reliable metrics for the interphase predicted score (ipTM) and predicted TM-score (pTM) (Supp. Fig. 9A) using the AlphaFold3 server. The models generated by both servers arrived at the same solution for the fragment models and when using full-length proteins, except in the case of NRBP1 (Supp. Fig. 9B).
Statistics and reproducibility
All data presented as means ± SEM unless otherwise noted. Data analyzed using GraphPad Prism 10 (GraphPad Software, CA, USA). Statistical significance (P < 0.05) calculated using two-tailed unpaired Student’s t-test unless noted. P values are provided in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Uncropped blot images can be found in Supplementary Fig. 10. Information on protein sequence, cellular localization, and conservation in mouse and frog were obtained from the UniProt database (www.uniprot.org). Most information on protein structure was obtained from the AlphaFold2 web server (alphafold.ebi.ac.uk). Larger proteins or related proteins that are not covered in AlphaFold2 were obtained from the RCSB Protein Databank (www.rcsb.org). Gene Ontology was derived from the Panther Classification System (pantherdb.org). All other data are available in the main text or the supplementary materials.
References
Verissimo, F. & Jordan, P. WNK kinases, a novel protein kinase subfamily in multi-cellular organisms. Oncogene 20, 5562–5569 (2001).
Xu, B. et al. WNK1, a novel mammalian serine/threonine protein kinase lacking the catalytic lysine in subdomain II. J. Biol. Chem. 275, 16795–16801 (2000).
Wilson, F. H. et al. Human hypertension caused by mutations in WNK kinases. Science 293, 1107–1112 (2001).
Xie, J. et al. Endothelial-specific expression of WNK1 kinase is essential for angiogenesis and heart development in mice. Am. J. Pathol. 175, 1315–1327 (2009).
Anselmo, A. N. et al. WNK1 and OSR1 regulate the Na+, K+, 2Cl- cotransporter in HeLa cells. Proc. Natl Acad. Sci. USA 103, 10883–10888 (2006).
de Los Heros, P. et al. The WNK-regulated SPAK/OSR1 kinases directly phosphorylate and inhibit the K+-Cl- co-transporters. Biochem. J. 458, 559–573 (2014).
Gagnon, K. B., England, R. & Delpire, E. A single binding motif is required for SPAK activation of the Na-K-2Cl cotransporter. Cell. Physiol. Biochem. 20, 131–142 (2007).
Moriguchi, T. et al. WNK1 regulates phosphorylation of cation-chloride-coupled cotransporters via the STE20-related kinases, SPAK and OSR1. J. Biol. Chem. 280, 42685–42693 (2005).
Pacheco-Alvarez, D. et al. WNK3-SPAK interaction is required for the modulation of NCC and other members of the SLC12 family. Cell. Physiol. Biochem. 29, 291–302 (2012).
Piechotta, K., Garbarini, N., England, R. & Delpire, E. Characterization of the interaction of the stress kinase SPAK with the Na+-K+-2Cl- cotransporter in the nervous system: evidence for a scaffolding role of the kinase. J. Biol. Chem. 278, 52848–52856 (2003).
Piechotta, K., Lu, J. & Delpire, E. Cation chloride cotransporters interact with the stress-related kinases Ste20-related proline-alanine-rich kinase (SPAK) and oxidative stress response 1 (OSR1). J. Biol. Chem. 277, 50812–50819 (2002).
Ponce-Coria, J. et al. Regulation of NKCC2 by a chloride-sensing mechanism involving the WNK3 and SPAK kinases. Proc. Natl Acad. Sci. USA 105, 8458–8463 (2008).
Richardson, C. et al. Activation of the thiazide-sensitive Na+-Cl- cotransporter by the WNK-regulated kinases SPAK and OSR1. J. Cell Sci. 121, 675–684 (2008).
Richardson, C. et al. Regulation of the NKCC2 ion cotransporter by SPAK-OSR1-dependent and -independent pathways. J. Cell Sci. 124, 789–800 (2011).
Vitari, A. C., Deak, M., Morrice, N. A. & Alessi, D. R. The WNK1 and WNK4 protein kinases that are mutated in Gordon’s hypertension syndrome phosphorylate and activate SPAK and OSR1 protein kinases. Biochem. J. 391, 17–24 (2005).
Vitari, A. C. et al. Functional interactions of the SPAK/OSR1 kinases with their upstream activator WNK1 and downstream substrate NKCC1. Biochem. J. 397, 223–231 (2006).
Darman, R. B. & Forbush, B. A regulatory locus of phosphorylation in the N terminus of the Na-K-Cl cotransporter, NKCC1. J. Biol. Chem. 277, 37542–37550 (2002).
Dowd, B. F. & Forbush, B. PASK (proline-alanine-rich STE20-related kinase), a regulatory kinase of the Na-K-Cl cotransporter (NKCC1). J. Biol. Chem. 278, 27347–27353 (2003).
Taylor, C. A. T. et al. OSR1 regulates a subset of inward rectifier potassium channels via a binding motif variant. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1802339115 (2018).
Shekarabi, M. et al. Mutations in the nervous system-specific HSN2 exon of WNK1 cause hereditary sensory neuropathy type II. J. Clin. Investig. 118, 2496–2505 (2008).
Vavolizza, R. D. et al. The effect of an autism-associated polymorphism in the STK39 gene on the autism symptom domains. J. Autism Dev. Disord. 42, 319–320 (2012).
Siew, K. & O’Shaughnessy, K. M. Extrarenal roles of the with-no-lysine[K] kinases (WNKs). Clin. Exp. Pharmacol. Physiol. 40, 885–894 (2013).
Arion, D. & Lewis, D. A. Altered expression of regulators of the cortical chloride transporters NKCC1 and KCC2 in schizophrenia. Arch. Gen. Psychiatry 68, 21–31 (2011).
Wang, Y. et al. Whole-genome association study identifies STK39 as a hypertension susceptibility gene. Proc. Natl Acad. Sci. USA 106, 226–231 (2009).
Henriques, A. F. A. et al. WNK1 phosphorylation sites in TBC1D1 and TBC1D4 modulate cell surface expression of GLUT1. Arch. Biochem. Biophys. 679, 108223 (2020).
Prisco, S. Z., Eklund, M., Raveendran, R., Thenappan, T. & Prins, K. W. With no lysine kinase 1 promotes metabolic derangements and RV dysfunction in pulmonary arterial hypertension. JACC Basic Transl. Sci. 6, 834–850 (2021).
Shekarabi, M. et al. WNK kinase signaling in ion homeostasis and human disease. Cell Metab. 25, 285–299 (2017).
Jaykumar, A. B. et al. WNK1 enhances migration and invasion in breast cancer models. Mol. Cancer Ther. 20, 1800–1808 (2021).
Köchl, R. et al. WNK1 kinase balances T cell adhesion versus migration in vivo. Nat. Immunol. 17, 1075–1083 (2016).
Sun, S., Cao, X., Castro, L. F. C., Monroig, Ó & Gao, J. A network-based approach to identify protein kinases critical for regulating srebf1 in lipid deposition causing obesity. Funct. Integr. Genomics 21, 557–570 (2021).
Gallolu Kankanamalage, S. et al. Multistep regulation of autophagy by WNK1. Proc. Natl Acad. Sci. USA 113, 14342–14347 (2016).
Taylor, C. A. & Cobb, M. H. CCT and CCT-like modular protein interaction domains in WNK signaling. Mol. Pharmacol. 101, 201 (2022).
Villa, F. et al. Structural insights into the recognition of substrates and activators by the OSR1 kinase. EMBO Rep. 8, 839–845 (2007).
Delpire, E. & Gagnon, K. B. Genome-wide analysis of SPAK/OSR1 binding motifs. Physiol. Genomics 28, 223–231 (2007).
Elvers, K. T., Lipka-Lloyd, M., Trueman, R. C., Bax, B. D. & Mehellou, Y. Structures of the human SPAK and OSR1 conserved C-terminal (CCT) domains. Chembiochem 23, e202100441 (2022).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377–D386 (2013).
Tran, N. et al. Endothelial nitric oxide synthase (eNOS) and the cardiovascular system: in physiology and in disease states. Am. J. Biomed. Sci. Res. 15, 153–177 (2022).
Jose, P. A., Soares-da-Silva, P., Eisner, G. M. & Felder, R. A. Dopamine and G protein-coupled receptor kinase 4 in the kidney: role in blood pressure regulation. Biochim. Biophys. Acta 1802, 1259–1267 (2010).
Wang, Z. et al. Human GRK4gamma142V variant promotes angiotensin II type I receptor-mediated hypertension via renal histone deacetylase type 1 inhibition. Hypertension 67, 325–334 (2016).
Garzon-Muvdi, T. et al. WNK4 kinase is a negative regulator of K+-Cl- cotransporters. Am. J. Physiol. Ren. Physiol. 292, F1197–F1207 (2007).
Fiol, D. F., Mak, S. K. & Kultz, D. Specific TSC22 domain transcripts are hypertonically induced and alternatively spliced to protect mouse kidney cells during osmotic stress. FEBS J. 274, 109–124 (2007).
Rashmi, P. et al. Glucocorticoid-induced leucine zipper protein regulates sodium and potassium balance in the distal nephron. Kidney Int. 91, 1159–1177 (2017).
Xiao, Y. X. et al. The TSC22D, WNK, and NRBP gene families exhibit functional buffering and evolved with Metazoa for cell volume regulation. Cell Rep. 43, 114417 (2024).
Jaykumar, A. B. et al. WNKs regulate mouse behavior and alter central nervous system glucose uptake and insulin signaling. Preprint at bioRxiv https://doi.org/10.1101/2024.06.09.598125 (2024).
Debily, M. A. et al. Expression and molecular characterization of alternative transcripts of the ARHGEF5/TIM oncogene specific for human breast cancer. Hum. Mol. Genet. 13, 323–334 (2004).
Parton, R. G. & del Pozo, M. A. Caveolae as plasma membrane sensors, protectors and organizers. Nat. Rev. Mol. Cell Biol. 14, 98–112 (2013).
Hill, M. M. et al. PTRF-Cavin, a conserved cytoplasmic protein required for caveola formation and function. Cell 132, 113–124 (2008).
Guha, U. et al. Comparisons of tyrosine phosphorylated proteins in cells expressing lung cancer-specific alleles of EGFR and KRAS. Proc. Natl Acad. Sci. USA 105, 14112–14117 (2008).
Humphrey, S. J. et al. Dynamic adipocyte phosphoproteome reveals that Akt directly regulates mTORC2. Cell Metab. 17, 1009–1020 (2013).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Filippi, B. M. et al. MO25 is a master regulator of SPAK/OSR1 and MST3/MST4/YSK1 protein kinases. EMBO J. 30, 1730–1741 (2011).
Shibanuma, M., Kuroki, T. & Nose, K. Isolation of a gene encoding a putative leucine zipper structure that is induced by transforming growth factor beta 1 and other growth factors. J. Biol. Chem. 267, 10219–10224 (1992).
Lee, B. H., Chen, W., Stippec, S. & Cobb, M. H. Biological cross-talk between WNK1 and the transforming growth factor beta-Smad signaling pathway. J. Biol. Chem. 282, 17985–17996 (2007).
Jaykumar, A. B. et al. WNK1 collaborates with TGF-β in endothelial cell junction turnover and angiogenesis. Proc. Natl Acad. Sci. USA 119, e2203743119 (2022).
Kester, H. A., Blanchetot, C., den Hertog, J., van der Saag, P. T. & van der Burg, B. Transforming growth factor-beta-stimulated clone-22 is a member of a family of leucine zipper proteins that can homo- and heterodimerize and has transcriptional repressor activity. J. Biol. Chem. 274, 27439–27447 (1999).
Moon, T. M. et al. Solution structure of the WNK1 autoinhibitory domain, a WNK-specific PF2 domain. J. Mol. Biol. 425, 1245–1252 (2013).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Evans, R. et al. Protein complex prediction with AlphaFold-multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2022).
Gluderer, S. et al. Madm (Mlf1 adapter molecule) cooperates with Bunched A to promote growth in Drosophila. J. Biol. 9, 9 (2010).
Zhang, J., Siew, K., Macartney, T., O’Shaughnessy, K. M. & Alessi, D. R. Critical role of the SPAK protein kinase CCT domain in controlling blood pressure. Hum. Mol. Genet. 24, 4545–4558 (2015).
Li, Q. et al. Yeast two-hybrid screening identified WDR77 as a novel interacting partner of TSC22D2. Tumour Biol. 37, 12503–12512 (2016).
Dbouk, H. A. et al. Actions of the protein kinase WNK1 on endothelial cells are differentially mediated by its substrate kinases OSR1 and SPAK. Proc. Natl Acad. Sci. USA 111, 15999–16004 (2014).
Jansa, P. & Grummt, I. Mechanism of transcription termination: PTRF interacts with the largest subunit of RNA polymerase I and dissociates paused transcription complexes from yeast and mouse. Mol. Gen. Genet. 262, 508–514 (1999).
Liu, L. & Pilch, P. F. PTRF/Cavin-1 promotes efficient ribosomal RNA transcription in response to metabolic challenges. eLife 5, e17508 (2016).
Jung, J. U. & Cobb, M. H. WNK1 controls endosomal trafficking through TRIM27-dependent regulation of actin assembly. Proc. Natl Acad. Sci. USA 120, e2300310120 (2023).
Nishida, H. et al. Phosphatidylinositol 3-kinase/Akt signaling pathway activates the WNK-OSR1/SPAK-NCC phosphorylation cascade in hyperinsulinemic db/db mice. Hypertension 60, 981–990 (2012).
Lenertz, L. Y. et al. Properties of WNK1 and implications for other family members. J. Biol. Chem. 280, 26653–26658 (2005).
Boyd-Shiwarski, C. R. et al. WNK kinases sense molecular crowding and rescue cell volume via phase separation. Cell 185, 4488–4506.e4420 (2022).
Kovtun, O. et al. Structural insights into the organization of the cavin membrane coat complex. Dev. Cell 31, 405–419 (2014).
Lee, S. J., Cobb, M. H. & Goldsmith, E. J. Crystal structure of domain-swapped STE20 OSR1 kinase domain. Protein Sci.18, 304–313 (2009).
Villa, F., Deak, M., Alessi, D. R. & van Aalten, D. M. Structure of the OSR1 kinase, a hypertension drug target. Proteins 73, 1082–1087 (2008).
Taylor, C. A. et al. Domain-swapping switch point in Ste20 protein kinase SPAK. Biochemistry 54, 5063–5071 (2015).
Obenauer, J. C., Cantley, L. C. & Yaffe, M. B. Scansite 2.0: proteome-wide prediction of cell signaling interactions using short sequence motifs. Nucleic Acids Res. 31, 3635–3641 (2003).
Li, Y. et al. SPAK kinase is a substrate and target of PKCtheta in T-cell receptor-induced AP-1 activation pathway. EMBO J. 23, 1112–1122 (2004).
Gagnon, K. B., England, R. & Delpire, E. Volume sensitivity of cation-Cl- cotransporters is modulated by the interaction of two kinases: Ste20-related proline-alanine-rich kinase and WNK4. Am. J. Physiol. Cell Physiol. 290, C134–C142 (2006).
Polek, T. C., Talpaz, M. & Spivak-Kroizman, T. The TNF receptor, RELT, binds SPAK and uses it to mediate p38 and JNK activation. Biochem. Biophys. Res. Commun. 343, 125–134 (2006).
Polek, T. C., Talpaz, M. & Spivak-Kroizman, T. R. TRAIL-induced cleavage and inactivation of SPAK sensitizes cells to apoptosis. Biochem. Biophys. Res. Commun. 349, 1016–1024 (2006).
Gagnon, K. B., England, R., Diehl, L. & Delpire, E. Apoptosis-associated tyrosine kinase scaffolding of protein phosphatase 1 and SPAK reveals a novel pathway for Na-K-2C1 cotransporter regulation. Am. J. Physiol. Cell Physiol. 292, C1809–C1815 (2007).
Smith, L., Smallwood, N., Altman, A. & Liedtke, C. M. PKCdelta acts upstream of SPAK in the activation of NKCC1 by hyperosmotic stress in human airway epithelial cells. J. Biol. Chem. 283, 22147–22156 (2008).
Gagnon, K. B. & Delpire, E. Multiple pathways for protein phosphatase 1 (PP1) regulation of Na-K-2Cl cotransporter (NKCC1) function: the N-terminal tail of the Na-K-2Cl cotransporter serves as a regulatory scaffold for Ste20-related proline/alanine-rich kinase (SPAK) AND PP1. J. Biol. Chem. 285, 14115–14121 (2010).
Acknowledgements
Funding provided by Welch Foundation Grant I1243 (M.H.C.), National Institutes of Health Grant R01HL147661 (M.H.C.), National Cancer Institute Grant P30CA142543 to the Harold C. Simmons Comprehensive Cancer Center for partial support of the Live Cell Imaging Facility, CPRIT training grant RP210041 (J.-U.J.), and NIH K99AG075161-01A1 (A.B.J.). We thank other members of the Cobb laboratory, the Albanesi (Department of Pharmacology) and Goldsmith (Department of Biophysics) labs and Kate Luby-Phelps (Department of Cell Biology and Live Cell Imaging Facility) at the University of Texas Southwestern Medical Center at Dallas (UTSW) for their suggestions about this project and Dionne Ware for administrative support. We thank the lab of Liang Tong (Columbia University) for working with us on one of the predicted hits. We would also like to thank three students who were involved in early stages of this work: Ashesh T. Triveda who was in the UTSW SURF summer program, Sakina Plumber who was in the UTSW-UT Dallas Green Fellows program, and Jonathan Zijiang Yang who was in the UTSW STARS summer program.
Author information
Authors and Affiliations
Contributions
Conceptualization: C.A.T., Methodology: C.A.T., J.-U.J., S.G.K., J.L., A.B.J., M.H.C., Investigation: C.A.T., J.-U.J., S.G.K., J.L., M.G., A.B.J., S.E., S.S., P.S., and E.S., Visualization: C.A.T., J.-U.J., S.G.K., and A.B.J., Supervision: C.A.T., M.H.C., Writing: C.A.T. and M.H.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Taylor, C.A., Jung, JU., Gallolu Kankanamalage, S. et al. Kinase interaction analysis predicts actions of the WNK-OSR1/SPAK pathway. Commun Biol 8, 1335 (2025). https://doi.org/10.1038/s42003-025-08551-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08551-5