Abstract
Glucocorticoid-induced skeletal muscle atrophy severely limits the clinical use of glucocorticoids and occurs in various endocrine and metabolic diseases. However, a detailed understanding of how glucocorticoid receptor (GR) transcriptional responses contribute to muscle atrophy is lacking. Irisin is a myokine induced by exercise and has been shown to exert multiple beneficial effects on muscle mass and metabolism regulation. Here, we show that glucocorticoid genomic effects are the main pathway through which glucocorticoids induce muscle atrophy in mice. Increased GR Ser212 and Ser234 site phosphorylation reduces glucocorticoid-induced muscle atrophy in mice. Irisin ameliorates high-fat diet (HFD) and dexamethasone (Dex)-induced muscle atrophy. Mechanistically, this effect depends on irisin promoting the phosphorylation of ERK and JNK through integrin αVβ5 receptors, which in turn impairs the dephosphorylation of GR Ser212 and Ser234 sites, affecting the GCs genomic effect on the transcription of muscle atrophy-related genes. These findings highlight the genomic effects of GCs as an intervention target to ameliorate GC-induced muscle atrophy and suggest that irisin could be a potential therapeutic target.

Similar content being viewed by others
Introduction
Skeletal muscle is the largest energy metabolic organ and protein reservoir. Muscle mass and function maintenance are crucial for human health and the maintenance of a high standard of quality of life1. Skeletal muscle mass is maintained by a critical balance between protein synthesis and degradation2. Glucocorticoids (GCs) are endogenous steroid hormones secreted by the adrenal cortex under the control of the hypothalamic–pituitary–adrenal (HPA) axis, and their levels fluctuate in response to both physiological and pathological stress3. In clinical practice, GCs are also widely used as exogenous drugs for the treatment of inflammatory and autoimmune diseases4. Persistent elevation of GC levels, whether caused by endogenous overproduction or long-term pharmacological administration, can disrupt muscle protein homeostasis by reducing the rate of protein synthesis and increasing the rate of protein degradation. These changes ultimately lead to skeletal muscle atrophy and weakness5. Previous reports have shown that GCs induce muscle atrophy under conditions of chronic stress, such as aging, cancer, diabetes, and chronic kidney disease (CKD)6,7,8,9. Although GCs may not be the sole factor responsible for muscle atrophy in these disease settings, their role in promoting catabolic pathways in skeletal muscle is well recognized.
In addition to their direct effects on protein synthesis and degradation, GCs also exert systemic regulatory effects on skeletal muscle metabolism. For instance, GCs suppress the hypothalamic–pituitary–growth hormone (GH) axis and reduce circulating levels of GH and insulin-like growth factor‑1 (IGF‑1), both of which are essential for promoting muscle anabolism10,11. GCs are also known to inhibit the synthesis of sex steroid hormones, including androgens and estrogens, which further impairs muscle maintenance12,13. Locally within skeletal muscle, GCs activate proteolytic mechanisms by engaging the ubiquitin proteasome system (UPS) and the autophagy lysosome pathway to promote protein degradation14. Simultaneously, they suppress anabolic signaling through pathways such as PI3K-AKT–mTOR, leading to reduced protein synthesis15. Moreover, GCs impair muscle regenerative processes by inhibiting satellite cell activation, myoblast proliferation, and differentiation16. Together, these systemic and local mechanisms disrupt skeletal muscle homeostasis and contribute to muscle atrophy under a variety of physiological and pathological conditions. Despite extensive knowledge of the pathways affected by GCs, the detailed molecular regulation of gene expression underlying GC-induced muscle atrophy remains to be fully elucidated.
GCs activate GR to promote muscle protein degradation by increasing the transcription of the characterized cyclic E3 ubiquitin ligases, muscle atrophy F-box (Fbxo32)/Atrogin-1, muscle RING finger 1 (Trim63)/MuRF-1, and myostatin (MSTN), a muscle growth negative regulator17. In addition, GCs inhibit the transcription of genes involved in protein synthesis, such as insulin growth factor-1 (IGF-1)18. In addition to ligand binding, several recent studies have provided evidence that crosstalk with other kinases can directly modulate GR transcriptional responses by phosphorylating the GR19. Extensive studies have indicated a link between GR hormone-dependent phosphorylation and transcriptional activity20,21,22,23. However, whether the regulation of GR transcriptional responses affects GC-induced muscle atrophy has not yet been reported.
GR consists of three domains: an N-terminal transcriptional activation function domain (AF-1), a DNA-binding domain (DBD), and a C-terminal ligand-binding domain (LBD)24. Studies have shown that human and mouse GR have highly conserved phosphorylation sites. Mouse GR has been reported to be phosphorylated at Ser122, Ser143, Ser150, Ser212, Ser220, Ser234, and Ser413 (homologous human serines are Ser113, Ser134, Ser141, Ser203, Ser211, Ser226, and Ser404, respectively), which are clustered in the N-terminal transcriptional regulatory region25. Among these, the phosphorylation sites of mouse GR Ser212, Ser220, Ser234, and Ser143 are located in the AF-1 region (which activates or inactivates the transcription of target genes), and the dephosphorylation of GR Ser212, Ser234, and Ser143 has been shown to promote the transcriptional regulation of GC downstream genes26,27,28,29,30.
Furthermore, phosphorylation of GR at these sites is regulated by kinases, such as p38 mitogen-activated protein kinase (p38), cell cycle protein-dependent kinase (CDK), c-Jun N-terminal kinase (JNK), and extracellular signal-regulated kinase (ERK), which can phosphorylate GR at multiple sites and further regulate GR transcriptional activity27,31,32. For example, JNK phosphorylates human GR at Ser226 (homologous to Ser234 in mice), leading to an increase in nuclear export and attenuation of GR transcriptional activity28. Both CDKs and ERK phosphorylate Ser203 (homologous to Ser212 in mice). Interestingly, CDK-mediated phosphorylation of the GR is required for transcriptional activity. Conversely, ERK phosphorylation attenuates GR transcriptional activity, resulting in GR retention in the cytoplasm and its failure to bind to GRE-containing promoters32. P38 phosphorylates Ser134 (homologous to Ser143 in mice), which significantly increases the association of the GR with the zeta isoform of the 14-3-3 class of signaling proteins (14-3-3zeta), leading to an attenuated hormone-dependent transcriptional response33. P38 has also been shown to phosphorylate Ser211(homologous to Ser220 in mice), which increases recruitment to GRE-containing promoters, leading to an increased transcriptional response29. Although the effects of GR phosphorylation on transcriptional function have been demonstrated, the exact mechanisms mediating GR responses to protein synthesis and degradation remain unclear. Further elucidation of effective treatments for GC-induced muscle atrophy is needed.
Irisin, a novel myokine, is secreted by muscle fibers in response to exercise. It promotes skeletal muscle hypertrophy and rescues denervation-induced muscle wasting34. In addition, irisin has versatile beneficial effects on the maintenance of muscle homeostasis in multiple diseases, such as CKD-induced muscle atrophy, age-associated muscle atrophy, and metabolic dysfunction. In addition, it enhances energy expenditure and alleviates high-fat diet (HFD)-induced obesity and insulin resistance35,36,37. The current study found that irisin promotes skeletal muscle hypertrophy by increasing protein kinase B (AKT) and ERK phosphorylation, which are associated with the protein synthesis pathway38. Recent studies have revealed that irisin can stimulate muscle growth-related genes in humans39. Taken together, these studies provide evidence that increased irisin levels may ameliorate skeletal muscle atrophy. However, the molecular mechanism by which irisin mediates these effects is still not well understood. In conditions such as obesity, aging, and CKD, skeletal muscle atrophy occurs and is accompanied by elevated levels of GCs7,40,41,42,43. GCs and irisin, a myokine, have been reported separately to play key roles in regulating a range of metabolic, immunologic, and homeostatic functions in diseases such as diabetes, aging, and inflammation. Uncovering the mechanisms of irisin crosstalk with GC genomic and non-genomic effects could expand our understanding and potentially guide therapeutic strategies for these diseases.
In the present study, we established an HFD and an aging mouse model with elevated circulating GC levels to investigate the role of irisin in muscle atrophy and regeneration. To determine whether irisin acts via the GC pathway, we administered dexamethasone (Dex) both in vivo and in vitro and examined the role of irisin in Dex-induced muscle and myotube atrophy. To further elucidate whether irisin regulates the genomic effects of GCs, we performed RNA-Seq and qRT-PCR on irisin- and Dex-injected mouse muscles to validate the effects of irisin on GC downstream genes related to protein synthesis and degradation. To test whether irisin inhibits the GR transcriptional response by regulating GR phosphorylation, we established C2C12 stable expression cell lines with phosphorylation-deficient (Ser212A, Ser234A, and Ser212/234A) or persistently phosphorylated (Ser212D, Ser234D, and Ser212/234D) variants using lentiviral transduction, and detected the effect of irisin on the transcription of genes downstream of GCs by qRT-PCR. Finally, integrin αvβ5 receptor inhibitors as well as JNK and ERK inhibitors were used to clarify the specific mechanisms by which irisin regulates GR phosphorylation and the actions of the GC genome. Our results demonstrate that phosphorylation of GR at Ser212 and Ser234 is a key mediator of irisin crosstalk with GC genomic effects in mouse muscle atrophy.
Results
Irisin ameliorates HFD-induced muscle atrophy
Elevated serum GC levels are an important contributor to chronic stress-induced muscle wasting44. We analyzed diseases associated with elevated serum GCs using the MalaCards Human Disease Database (http://www.malacards.org/). A composite relatedness score was generated, which displayed the top 200 related diseases (Supplementary Data 2). The top 200 diseases were divided into 14 categories according to the classification section of MalaCards, and the results showed that endocrine and metabolic diseases accounted for the highest proportion (24.0 %) of diseases associated with elevated serum GC levels (Fig. 1A). In terms of endocrine and metabolic diseases associated with elevated serum GC levels (Fig. 1B), conditions such as diabetes, obesity, CKD, and aging have been reported to be associated with skeletal muscle atrophy6,7,8.
A MalaCards human disease database (http://www.malacards.org/) generated the top 200 associated with elevated serum glucocorticoids diseases into 14 categories according to the classifications section of the MalaCards. B Refinement of endocrine and metabolic diseases associated with elevated serum glucocorticoids in Fig. 1A, ranked by relevance score. C The protocol of the HFD mice experiment, n = 5 mice per group. Created in BioRender. J, Y. (2025) https://BioRender.com/4d6mb1h. D Representative images showing wheat germ agglutinin (WGA, green) stained in HFD and irisin administered mice TA. Scale bar = 100 µm. E The myofiber number distribution was calculated as the percentage of the number of myofibers in a designated area divided by the total number of myofibers assessed. F The quantification of mean cross-sectional area (CSA), n = 3 mice per group. G Western blot analyses of Atrogin-1, MuRF-1, p-AKT(Ser473), AKT and GAPDH in Gas muscle, n = 3 mice per group. H The protocol of the young and aging mice muscle injury experiment, n = 3 mice per group. Created in BioRender. J, Y. (2025) https://BioRender.com/xwcfiom. I Representative images showing Laminin (green) stained injury 8d in young and aging mice TA. Scale bar = 100 µm. J The frequency distribution of central nuclei myofibers was calculated as the percentage of the number of central nuclei myofibers in a designated area divided by the total number of myofibers assessed. K The quantification of mean cross-sectional area (CSA), n = 3 mice per group. L Western blot analyses of Atrogin-1, MuRF-1, p-AKT(Ser473), AKT and GAPDH in young and aging mice injury 8d Gas muscle, n = 3 mice per group. Data are presented as means ± SEM. Each symbol in the bar graphs indicates an individual mouse data value. One-way ANOVA was used for statistical analysis (F). Two-way ANOVA was used for statistical analysis (K).
To study the role of irisin in skeletal muscle mass in diseases associated with elevated serum GC levels, we conducted in vivo experiments using recombinant irisin. Initially, we purchased commercial irisin (c-irisin), which is expressed in a prokaryotic system, from Phoenix Pharmaceuticals. As many other published studies also use irisin expressed in a prokaryotic system, we adopted the same approach for our own preparation in order to reduce costs and ensure a sufficient supply for in vivo experiments. Western blot analysis confirmed that recombinant irisin (r-irisin) exhibited band positions consistent with commercial irisin (c-irisin) (Fig. S1A). Moreover, recombinant irisin injection in mice led to a significant increase in UCP1 mRNA levels in white adipose tissue (Fig. S1B). For further exploration, we used humanized Pcsk9 mice (hpcsk9) and fed an HFD to induce obesity (Fig. 1C). After 31 weeks, HFD mice displayed a significant increase in body weight compared with normal-diet (ND) mice (Fig. S1C). A portion of the HFD mice was then treated with irisin for 4 weeks. Next, myofiber areas were evaluated using WGA immunofluorescence-stained TA muscle sections (Fig. 1D). The HFD group shifted the distribution of myofibers in a smaller direction compared to the ND group, whereas the HFD+irisin group shifted the distribution of myofibers in a larger direction compared to the HFD group (Fig. 1E). The mean cross-sectional area (CSA) of the TA muscle was consistently and significantly larger in HFD+irisin mice than in HFD mice (Fig. 1F). The markers of protein degradation and synthesis were also detected by Western bloting, HFD increased MuRF-1 protein levels and decreased phosphorylation of AKT in muscle compared to the ND group, whereas irisin treatment decreased Atrogin-1 and MuRF-1 protein levels in HFD mice (Figs. 1G and S1D).
We investigated the effects of irisin on muscle regeneration in aged mice. Cardiotoxin (CTX)-injected TA muscles were analyzed 8 days after injury (Fig. 1H). The TA muscle sections were immunostained for laminin (Fig. 1I). Aging mice showed a shift in the distribution of myofibers in the central nuclei in a smaller direction, whereas irisin administration markedly promoted a shift toward larger fibers in young mice and elicited only a modest increase in aged mice. (Fig. 1J). Consistently, the central nucleus myofiber CSA of the TA muscle was higher in irisin-injected aging mice (Fig. 1K). Irisin did not significantly decrease the protein levels of Atrogin-1 and MuRF-1 levels in aging mouse muscles, but it increased AKT phosphorylation, suggesting a role in promoting protein synthesis rather than inhibiting degradation (Figs. 1L and S1E). Taken together, these results indicate that irisin alleviates HFD-induced muscle atrophy. In aged mice, irisin promoted an increase in the CSA of regenerating myofibers after injury, but its effect on age-related muscle atrophy was limited.
Irisin prevents GC-induced muscle atrophy
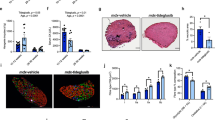
To confirm that irisin impaired muscle atrophy by blocking the effect of GCs, dexamethasone (Dex, a GC analog) was administered for 10 days to induce muscle atrophy in mice, which is a well-established mouse model of muscle wasting45. We simultaneously injected different concentrations of irisin to verify its effects on Dex-induced muscle atrophy (Fig. 2A). Dex injection significantly decreased the body weight and irisin reduced the body weight loss in Dex-injected mice during the later period (Fig. 2B). Notably, Dex treatment resulted in a significant reduction in the muscle weight to tibia length (MW/TL) ratio of the TA, Gas, and EDL muscles, whereas irisin treatment of Dex-injected mice resulted in a significant increase in the MW/TL ratio of the TA, Gas, and EDL muscles compared to Dex treatment alone. These results also showed that there is a dose-dependent increase in muscle weight by irisin in the Dex condition. In Dex-induced mice with 1 mg/kg irisin, muscle weights were almost the same as those in control mice (without Dex and irisin). However, in the Sol muscles, neither Dex nor irisin significantly changed the muscle weight (Fig. 2C). The myofiber CSAs of the TA were quantified by WGA immunofluorescence in Dex-injected and Dex plus different concentrations of irisin-injected mice (Fig. 2D). Dex shifted the distribution of myofibers in a smaller direction, whereas irisin administration shifted it in a larger direction compared to that in control mice. In addition, treatment with irisin plus Dex shifted the distribution of myofibers in a larger direction compared to mice treated with Dex-only (Fig. 2E). In agreement with this, the average myofiber area of irisin plus Dex-treated mice was significantly higher than that of Dex-injected mice (Fig. 2F). The protein levels of Atrogin-1, MuRF-1, P-AKT(Ser473), and AKT were measured in the Gas muscle (Fig. 2G). As expected, we observed a significant increase in Atrogin-1 and MuRF-1 protein levels and a decrease in phosphorylation of AKT in response to Dex treatment. However, irisin plus Dex-treated mice showed a notable reduction in Atrogin-1 and MuRF-1 protein levels and an increase in phosphorylation of AKT, when compared to mice that were only Dex-injected (Fig. 2H). These data suggest that irisin treatment ameliorates Dex-induced muscle atrophy, which is associated with reduced protein levels of key markers of skeletal muscle wasting and increased AKT phosphorylation.
A The protocol of Dex and irisin administered mice. Created in BioRender. J, Y. (2025) https://BioRender.com/pvgf5zf. B The body weight change percentage of mice. n = 6 mice per group. Data are presented as means ± SEM. P values were calculated using two-way ANOVA. * compared Dex(1 mg/kg) and Dex(1 mg/kg)+Irisin (0.5 mg/kg) groups, & compared Dex(1 mg/kg) and Dex(1 mg/kg)+Irisin (1 mg/kg) groups. C The ratio of TA, Gas, EDL and Sol muscle weight to tibia length is shown. n = 6 in each group. D Representative images showing wheat germ agglutinin (WGA, green) stained TA muscle sections in Dex and irisin administered mice. Scale bar = 100 µm. E The quantification of myofiber number distribution. F The quantification of mean cross-sectional area (CSA). n = 5 in each group. G Western blot analyses of Atrogin-1, MuRF-1, p-AKT(Ser473), AKT and GAPDH in Dex and irisin-administered mice Gas muscle. H Protein levels were analyzed, n = 4 mice per group. Data are presented as means ± SEM. Two-way ANOVA was used for statistical analysis.
Irisin attenuates the Dex-induced reduction in myotube size in vitro
To confirm the role of irisin in GC-induced muscle atrophy and its direct action on myofibers, C2C12 myoblasts were incubated with a differentiation medium for 4 days until cell fusion to form myotubes, and treated them with Dex and different concentrations of irisin (Fig. 3A). Immunostaining with embryonic myosin heavy chain (eMyHC) was performed to visualize the myotubes (Fig. 3B). Consistent with the in vivo results, we found that Dex treatment shifted the distribution of myotubes in a smaller direction than in the control, whereas irisin administration shifted the distribution in a larger direction. In addition, treatment with irisin plus Dex shifted the distribution of myotubes in a larger direction compared to Dex-treated myotubes alone (Fig. 3C). We also found that Dex treatment markedly reduced the diameter of myotubes, whereas treatment with irisin plus Dex significantly increased the myotube diameter compared to Dex treatment alone (Fig. 3D). Furthermore, we measured the protein levels of Atrogin-1, MuRF-1, p-AKT(Ser473), and AKT in the myotubes (Fig. 3E). We found that Dex induced an increase in Atrogin-1 and MuRF-1 expression and decreased AKT phosphorylation. In addition, treatment with irisin plus Dex resulted in a notable reduction in Atrogin-1 and MuRF-1 protein levels and increased the phosphorylation of AKT, when compared to Dex treatment alone (Fig. 3F-H). Collectively, these results suggested that irisin attenuated Dex-induced myotube atrophy.
A C2C12 cells were proliferated 3d to 100% and transferred to differentiation medium for 4 days, then treated with or without Dex and irisin for 24h, and cells were collected for WB, qRT-PCR and immunofluorescence. Created in BioRender. J, Y. (2025) https://BioRender.com/apfyv0h. B Immunofluorescence staining of eMyHC (green) and Hoechst (blue) in myotubes, Scale bar = 100 μM. C The frequency distribution of C2C12 myotubes diameter was analyzed. D The mean myotubes diameter was analyzed. E Western blot analyses of Atrogin-1, MuRF-1, p-AKT(Ser473), AKT and GAPDH in Dex and irisin-administered myotubes. F–H Protein levels were analyzed, n = 4 independent replicates. Data are presented as means ± SEM. Two-way ANOVA was used for statistical analysis.
Irisin inhibits nuclear translocation of GC receptor and GC genomic effects
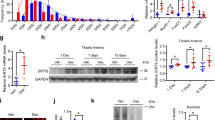
To examine the molecular mechanism by which irisin attenuates Dex-induced muscle atrophy, we analyzed differential gene expression in the Gas muscle of mice injected with Dex and mice injected with both Dex and irisin using RNA-Seq. The volcano map and heatmap results showed that mice treated with irisin plus Dex downregulated GC target genes associated with protein degradation, such as Trim63, Fbxo32, Foxos, and MSTN. Additionally, they upregulated GC target genes associated with protein synthesis, such as IGF-1, compared to mice that were only injected with Dex (Fig. 4A, B). In addition, we further validated these genes in the Gas muscle, consistent with the RNA-Seq results, which showed that treatment with irisin plus Dex decreased Fbxo32, Trim63 and MSTN mRNA expression and increased IGF-1 expression compared to Dex-only injected mice (Fig. 4C, D). We also tested the mRNA levels of IGF-1 in the liver (which mainly produces IGF-1) and found that Dex and irisin did not affect IGF-1 mRNA expression in the liver (Fig. 4E). Likewise, the mRNA levels of these genes in myotubes treated with Dex and irisin were consistent with the in vivo results, compared to Dex treatment alone (Fig. S1F–H, I). These results suggest that irisin can influence the genomic response of GCs and is associated with the degradation of proteins.
A GC and irisin-regulated target gene expression associated with protein synthesis and degradation analysis with FDR-adjusted p value < 0.05 and an absolute log2 fold change >1 are shown in the volcano map, n = 3 mice per group. B Heap map showed the genes expressions of RNA-Seq data, n = 3 mice per group. C, D The mRNA expressions of Fbxo32, Trim63, IGF-1 and MSTN were analyzed by qRT-PCR in Dex and irisin treatment mice Gas muscle, n = 4 mice per group. E The mRNA expressions of IGF-1 were analyzed by qRT-PCR in Dex and irisin treatment mice liver, n = 4 mice per group. F Immunohistochemistry of protein expression of GR in 1 mg/kg Dex treatment and 1 mg/kg Dex and 0.5 mg/kg irisin treatment mice TA muscle sections. G The mean nuclear intensity of GR and GR-positive cells was analyzed. n = 5 mice per group. H The GR protein levels were analyzed in nucleus and cytoplasm by WB after Dex and irisin treatment of 24 h myotubes. I The GR protein levels in the nucleus and cytoplasm were calculated. Histone H3 and GAPDH were used as the nuclear and cytoplasmic control. n = 4 independent replicates. J Western blot analyses of MSTN, P-Smad3, Smad3, Atrogin-1, MuRF-1 and GAPDH in Dex and irisin-administered myotubes. K Western blot analyses of IGF-1, p-AKT(Ser473), AKT and GAPDH in Dex and irisin-administered myotubes. Data are presented as means ± SEM. Two-way ANOVA was used for statistical analysis (C–E), and one-way ANOVA was used for statistical analysis (G, I).
To examine whether irisin affects GR translocation, we performed immunohistochemical staining of GR in the TA muscle and found that the mean nuclear intensity of GR and GR-positive cells increased with Dex treatment, but decreased with irisin plus Dex treatment (Fig. 4F, G). Nuclear and cytoplasmic proteins were separated from C2C12 myotubes for further validation (Fig. 4H). The results showed that GR protein levels decreased in the cytoplasm of Dex-treated C2C12 myotubes but increased in the nucleus. In contrast, treatment with irisin plus Dex resulted in the opposite trend, as the GR in the nucleus was decreased and that in the cytoplasm was increased when compared with the Dex group (Fig. 4I). These results demonstrate that irisin impairs GC genomic effects and inhibits the nuclear translocation of GR.
IGF-I and MSTN are key regulators involved in skeletal muscle anabolism and catabolism18,46. Thus, we further evaluated the effect of irisin on the activation of these two pathways using western blotting (Fig. 4J, K). We found that Dex significantly increased MSTN protein levels and Smad3 phosphorylation as well as Atrogin-1 protein levels, whereas treatment with irisin plus Dex decreased these protein levels and Smad3 phosphorylation when compared to Dex-treated only conditions (Fig. S1I, J). In addition, treatment with irisin and Dex increased IGF-1 protein levels and AKT phosphorylation compared to treatment with Dex alone (Fig. S1K). Taken together, these results indicate that irisin impairs the transcription of GC downstream genes and activates or inhibits the signaling pathways regulated by these genes.
Irisin affects GC genomic effects by regulating GR phosphorylation
Mouse GR is phosphorylated at Ser234, Ser212, and Ser143, which has been shown to attenuate the GC genomic response47. We examined the effect of irisin on these phosphorylation sites and found that phosphorylation at Ser234 and Ser212 in myotubes decreased with Dex treatment alone, whereas irisin treatment impaired Dex-induced Ser234 and Ser212 site dephosphorylation, but did not affect the Ser143 site (Fig. 5A–C). Furthermore, we conducted in vivo experiments involving dexamethasone, irisin, and their combination. As shown in (Fig. S4A–C), irisin prevented the dexamethasone-induced reduction in GR phosphorylation at Ser212 and Ser234 sites, thereby corroborating the mechanistic insights observed in vitro. To determine whether the phosphorylation of GR Ser234 and Ser212 affects the genomic effects of GC on the transcription of genes associated with muscle atrophy, lentiviral constructs of C2C12 cell lines were used, including overexpression of wild-type GR (WT), serine-to-alanine phosphorylation-deficient GR (Ser234A, Ser212A, Ser234/212A) mutants, or serine-to-aspartate persistently phosphorylated GR (Ser234D, Ser212D, Ser234/212D) mutants (Fig. S2A). GR phosphorylation site-specific antibodies were used to verify the successful construction of stably expressing cell lines (Fig. 5D). The results showed that overexpression of GR Ser234D and GR Ser212D significantly increased the phosphorylation levels of GR Ser234 and GR Ser212, respectively, compared to those in the GR WT group. Overexpression of the GR Ser234/212D cell line significantly increased the phosphorylation of both GR Ser234 and GR Ser212. However, overexpression of GR Ser234A, GR Ser212A, and GR Ser234/212A did not affect the phosphorylation levels of GR Ser234 and GR Ser212 compared to those in the GR WT group (Fig. S2B, C). Overall, the overexpressing cell lines showed significantly increased GR protein levels compared to the control group (Fig. S2D). These results indicate that the construction of persistently phosphorylated GR-overexpressing C2C12 cell lines was successful. The constructed cell lines were further treated with or without Dex and it was found that none of overexpressing GR Ser234A, GR Ser212A, and GR Ser234/212A affected the transcription of Trim63, Fbxo32, and IGF-1 compared with the overexpressing GR WT group under the control or Dex condition; however, under the Dex condition, overexpressing GR Ser234D or GR Ser212D significantly decreased Trim63 and Fbxo32 transcription; overexpressing GR Ser234/212D not only significantly decreased Trim63 and Fbxo32 transcription but also increased IGF-1 transcription compared to the GR WT group (Figs. 5E and S2E). These results indicate that regulating the phosphorylation levels of GR Ser234 and GR Ser212 did not affect GC genomic effects without GCs, but in GC conditions, enhancing the phosphorylation levels of GR Ser234 and GR Ser212 could reduce GC genomic effects, whereas raising the dephosphorylation levels of GR Ser234 and GR Ser212 could affect GC genomic effects. To further investigate the role of irisin, we treated cell lines with irisin plus Dex. In the overexpressing GR WT cell line, irisin administration significantly decreased Trim63 and Fbxo32 transcription and increased IGF-1 transcription compared to the Dex group. In the overexpressing GR Ser234D and Ser212D cell lines, irisin administration still decreased the transcription of Trim63 and Fbxo32 and increased the transcription of IGF-1, but in the overexpressing GR Ser234/212D cell line, irisin administration did not affect the transcription of these genes (Figs. 5F and S2F). These results suggest that the phosphorylation of both GR Ser234 and Ser212 mediates the effect of irisin on the transcription of genes downstream of GC. However, in the overexpressing phosphorylation-deficient GR C2C12 cell line, the effect of irisin on the transcription of these genes after irisin administration was consistent with that observed in the GR WT cell line (Figs. 5G and S2G). These results indicate that irisin reduced the genomic effects of GC by enhancing the phosphorylation of both GR Ser234 and Ser212 sites.
A Western blot analyses of P-GR(Ser234), P-GR(Ser212), P-GR(Ser143) and GAPDH in Dex and irisin-administered myotubes. B, C Protein levels were analyzed, n = 4 independent replicates. D Western blot analyses of P-GR(Ser234), P-GR(Ser212), GR and GAPDH in lentiviral constructs of C2C12 cell lines expressing wild-type GR (WT), serine-to-alanine phosphorylation-deficient (S234A, S212A, S234/212A) mutants, or serine-to-aspartate persistently phosphorylated (S234D, S212D, S234/212D) mutants. E The mRNA expressions of Trim63 and Fbxo32 were analyzed by qRT-PCR in Dex treatment lentiviral constructs of overexpression C2C12 cell lines, n = 4 independent replicates. F The mRNA expressions of Fbxo32 were analyzed by qRT-PCR in Dex and irisin treatment lentiviral constructs of overexpression GR S234D, S212D, S234/212D C2C12 cell lines, n = 4 independent replicates. G The mRNA expressions of Trim63 were analyzed by qRT-PCR in Dex and irisin treatment lentiviral constructs of overexpression GR S234A, S212A, S234/212A C2C12 cell lines, n = 4 independent replicates. Data are presented as means ± SEM. One-way ANOVA was used for statistical analysis (B, C), two-way ANOVA was used for statistical analysis (E–G).
Irisin has a mediating role on GR phosphorylation via ERK and JNK phosphorylation
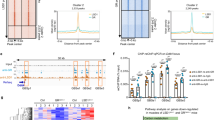
ERK and JNK activity regulates the phosphorylation of GR Ser212 and Ser234 and attenuates the transcriptional activity of GR47. ERK and JNK phosphorylation was assessed by Western blotting (Fig. 6A). The results showed that the phosphorylation levels of ERK and JNK were significantly decreased in myotubes treated with Dex alone, whereas irisin plus Dex treatment was associated with increased ERK and JNK phosphorylation levels (Fig. 6B). Next, we used ERK and JNK inhibitors to block their activities (Fig. 6C, D). The results showed that irisin had no effect on the phosphorylation levels of JNK, GR Ser234, ERK, and GR Ser212 under normal and Dex treatment conditions when administered either as a JNK inhibitor or ERK inhibitor alone or as JNK and ERK inhibitors together (Fig. S3A–H). This suggests that the regulatory effects of irisin on the phosphorylation of these proteins were effectively inhibited by both ERK and JNK inhibitors. Next, we evaluated the effect of inhibiting JNK and ERK activity on the transcription of genes downstream of GCs and found that treatment with irisin plus Dex decreased Fbxo32, Trim63, and MSTN mRNA expression and increased IGF-1 expression compared to treatment with Dex alone. However, when inhibiting JNK and ERK activity, treatment with irisin plus Dex had no effect on the expression of GC target genes compared to the Dex-treated group (Fig. 6E–H). These data suggest that irisin promotes the phosphorylation of GR and genomic effects of GCs by modulating the phosphorylation of ERK and JNK.
A Western blot analyses of P-JNK (Thr183/Tyr185), P-ERK and GAPDH in Dex and irisin-administered myotubes. B Protein levels were analyzed, n = 4 independent replicates. C, D Western blot analyses of P-GR(Ser234) and P-GR(Ser212) in myotubes with JNK inhibitors and combined JNK and ERK inhibitors. E–H The mRNA expressions of Fbxo32, Trim63, IGF-1 and GAPDH were analyzed by qRT-PCR in myotubes treated with JNK inhibitors and ERK inhibitors, alone or in combination, n = 4 independent replicates. Data are presented as means ± SEM. One-way ANOVA was used for statistical analysis (B), two-way ANOVA was used for statistical analysis (E–H).
Irisin effects on GC genomic effects and GR phosphorylation depend on integrin αVβ5 receptor
To further understand how irisin acts to promote JNK and ERK phosphorylation and regulate GC genomic effects, we analyzed integrin αVβ5 receptor protein expression in Dex and irisin treatment myotubes and inhibited integrin αVβ5 receptor with cilengitide. The results showed that treatment with irisin plus Dex increased integrin β5 protein levels compared with treatment with Dex alone (Fig. 7A, B). The action of irisin is mainly dependent on its binding to its receptor, which initiates downstream signaling pathways. We further used cilengitide, an inhibitor of integrin αvβ5, to inhibit the binding of irisin to integrin αvβ5 and its downstream signaling, and found that administration of integrin αvβ5 inhibitor cilengitide significantly decreased the phosphorylation of ERK, JNK, and GR by irisin. In addition, cilengitide abolished the beneficial effects of irisin by decreasing IGF-1 and increasing MSTN protein expression (Fig. 7C–I). Taken together, these results suggest irisin, through the integrin αVβ5 receptor, promotes the phosphorylation of ERK, JNK, GR Ser212, and Ser234 sites, ultimately inhibiting the genomic effects of GCs.
A Western blot analyses of integrin αV, integrin β5 and GAPDH in Dex and irisin-administered myotubes. B Protein levels were analyzed, n = 4 independent replicates. C Western blot analyses of P-ERK, P-GR(Ser212), P-JNK (Thr183/Tyr185), P-GR(Ser234), IGF-1, MSTN and GAPDH in integrin αVβ5 inhibitor cliengitide treatment myotube. D–I Protein levels were analyzed, n = 4 independent replicates. Data are presented as means ± SEM. One-way ANOVA was used for statistical analysis (B), Two-way ANOVA was used for statistical analysis (D–I).
Discussion
GCs are extensively used to treat numerous diseases because of their potent anti-inflammatory and immunosuppressive effects48. However, high doses or sustained use of GCs can cause severe muscular side effects manifesting as skeletal muscle atrophy, which severely limits their clinical application6. Furthermore, GC levels are elevated in conditions such as obesity, aging, and various endocrine and metabolic diseases, and can initially act as a stress response and help the body cope with stress by mobilizing energy reserves49. However, sustained elevation of GC levels can lead to muscle atrophy, not only reducing an individual’s quality of life but also promoting the development of cachexia50,51. In this study, we primarily used exogenous GC administration to induce muscle atrophy in order to establish a clear and controllable experimental system. This allowed us to focus specifically on the direct effects of GCs on skeletal muscle. We fully recognize that in actual pathological conditions such as chronic inflammation, cancer, or aging, endogenous GC levels are regulated by the HPA axis and may present more dynamic and complex changes. Therefore, in this study, we deliberately excluded other disease-related factors and selected an exogenous GC-induced model to better define the direct role of GCs in muscle atrophy and to investigate the regulatory mechanism of irisin under this specific background. In the present study, we demonstrate the critical role of irisin in the modulation of GC-induced skeletal muscle atrophy. Extensive literature indicates that both obesity and aging are associated with elevated GC levels, and we found that irisin ameliorates GC-induced muscle atrophy by modulating GR phosphorylation and GC genomic effects through binding to integrin αVβ5 receptors.
As an exercise-induced myokine, irisin not only induces the browning of white adipocytes, leading to metabolic improvements such as improved adiposity, insulin sensitivity, and hepatic steatosis, but also plays an important role in muscles in an autocrine manner36,38,52. Irisin improves muscle atrophy in different animal models, including muscle denervation, hindlimb suspension, and genetic muscle dystrophic mdx mice, with strong effects on enhancing satellite cell activation, reducing protein degradation, tissue fibrosis, necrosis, and improving sarcolemmal stability34,53,54. Because irisin has been reported to have a very short half-life in vivo, in our previous studies, we administered irisin intraperitoneally at a dose of 0.5 mg/kg every other day. To assess the effectiveness of this dosing regimen, we collected serum on days 8 and 14 and measured irisin levels using ELISA. The results showed that circulating irisin levels remained approximately twice the physiological baseline at both time points, indicating that this dosing strategy was sufficient to maintain a biologically active concentration over time55. In addition, several published studies have employed a similar dosing frequency and reported consistent biological effects36,54. Therefore, we consider the administration schedule of irisin used in this study to be well-supported. However, the role of this myokine in muscle atrophy induced by elevated circulating GCs remains unclear. Our study further showed that irisin ameliorated muscle atrophy and increased muscle mass in mouse disease models characterized by elevated circulating GC levels, such as obesity and direct administration of Dex. In addition, we demonstrated that irisin administration inhibited MuRF-1 and Atrogin-1 protein expression, which are associated with muscle atrophy via the ubiquitin proteasome system, and promoted the phosphorylation of AKT, which is associated with the protein synthesis pathway. Although direct measurement of protein synthesis and degradation rates would provide greater precision in evaluating muscle protein metabolism, we used widely accepted methods to assess skeletal muscle atrophy, including muscle weight measurement, CSA analysis, and the expression levels of atrophy-related markers MuRF-1 and Atrogin-1. In muscle weight measurement, we deliberately selected muscles with distinct and representative fiber-type compositions for muscle mass analysis. Specifically, the tibialis anterior (TA) and gastrocnemius (Gas) are mixed muscles predominantly composed of type IIa and IIb fibers, with a small proportion of type I fibers; the soleus (Sol) consists mainly of oxidative type I fibers; and the extensor digitorum longus (EDL) is primarily composed of fast-twitch type IIb fibers. Our results showed that Dex and irisin treatment significantly influenced the mass of TA, Gas, and EDL, while the mass of Sol remained largely unaffected. These findings suggest that the effects of Dex and irisin are likely more pronounced in muscles enriched in type II fibers. Furthermore, our findings are supported by previous studies reporting that Dex and irisin exert greater effects on type IIb fast-twitch fibers, while their impact on type I fibers appears to be limited[76]. This aligns well with our observations and reinforces our interpretation. Although these E3 ligases, such as MuRF-1 and Atrogin-1, do not fully reflect global protein degradation, they are key components of the UPS and are widely used as markers of muscle atrophy. Likewise, while AKT phosphorylation alone may not directly represent protein synthesis, it is a central node in the PI3K/AKT/mTOR pathway and provides useful insight into anabolic signaling.
GCs regulate skeletal muscle protein synthesis and degradation through both genomic and non-genomic pathways56. The genomic pathway involves the binding of GCs to the GR, which regulates the transcription of genes involved in protein synthesis and degradation. Non-genomic pathways promote proteolysis and inhibit protein synthesis through rapid cellular signaling mechanisms, such as the MAPK and PI3K-AKT pathways, which do not rely on the GR57,58,59. The genomic effects of the GR play a significant role in chronic diseases and long-term GC administration. Although non-genomic effects are rapid, the transcription of both upstream (IGF-1) and downstream (Trim63 and Fbxo32) genes in these pathways is regulated by the GR. Furthermore, experimental evidence suggests that the GR antagonist RU486 significantly inhibits the transcription of Trim63 and Fbxo32, thereby ameliorating the GC-induced muscle mass loss60. Therefore, we hypothesized that the genomic effect of the GR is the primary mechanism underlying the GC-induced imbalance in skeletal muscle protein metabolism. Whether irisin ameliorates GC-induced skeletal muscle dysplasia by modulating the genomic effects of GCs is unknown. Huh et al. showed that in human skeletal muscle cells, administration of irisin significantly promoted the transcription of IGF-1 and inhibited the transcription of MSTN 38. Other studies have demonstrated that irisin significantly inhibits MuRF-1 and Atrogin-1 protein levels in differentiated myotubes34. This suggests that irisin affects these genes; however, its effect on their transcription in the presence of GCs is unknown. In our study, we sequenced the transcriptome of Gas from Dex and irisin-treated mice, although numerous differentially expressed genes were observed in the volcano plot comparing the Dex/Irisin and Dex groups, and this study focused on key genes closely associated with skeletal muscle atrophy. Our transcriptome results showed that irisin could significantly repress protein degradation-related genes and activate protein synthesis-related genes. Furthermore, we also verified the results in vitro and in vivo by qRT-PCR consistent with the transcriptome results. Therefore, irisin might ameliorate GC-induced muscle atrophy by regulating the genomic effects of GCs. It should be noted that the observed transcriptional changes may not be limited to GR-related mechanisms, and that other transcriptional regulatory pathways may also be involved, which offers a valuable direction for future exploration of Irisin’s broader regulatory effect.
It has been reported that the transcription of GC target genes is affected by the phosphorylation of GR and that phosphorylation of Ser226 (orthologous to serine 234 in mice), Ser203 (orthologous to serine 212 in mice), and Ser134 (orthologous to serine 143 in mice) sites of GR can inhibit the genomic response of GCs19,47,61. For example, Reza et al. found that reduced phosphorylation of GR Ser203 increased the nuclear localization of GR and enhanced dexamethasone-mediated regulation of GLUT5 mRNA expression26. JNK-mediated phosphorylation of human GR at Serine 226 has also been found to blunt hormone signaling by enhancing the nuclear export of GR28. However, it is unclear whether alteration of the mouse GR phosphorylation site targets the transcription of genes critical for protein synthesis and degradation and the role of irisin in this process. We found that irisin administration significantly increased the phosphorylation of Ser234 and Ser212, but did not affect Ser143 compared to that in the Dex group. Therefore, we used lentiviruses to construct C2C12 cell lines overexpressing wild-type GR, serine-to-alanine phosphorylation-deficient mutants (Ser234A, Ser212A, and Ser234/212A), and serine-to-aspartate persistently phosphorylated GR mutants (Ser234D, Ser212D, and Ser234/212D). We found that overexpression of Ser234D, Ser212D, and Ser234/212D under GC conditions significantly decreased Trim63 and Fbxo32 transcription and increased IGF-1 transcription; however, irisin administration did not affect the transcription of these genes. These results suggest that increasing the phosphorylation levels of GR Ser234 and Ser212 reduces the genomic effects of GC and that irisin mediates its effects on the transcription of genes downstream of GC by promoting the phosphorylation of GR Ser234 and Ser212. However, in Ser234A, Ser212A, and Ser234/212A cell lines, there was no effect on the transcription of genes downstream of GCs after the administration of Dex and irisin, in contrast to the WT cell lines. This may be due to the fact that in overexpression systems, high levels of GR expression may have saturated the cellular regulatory machinery and may occupy available DNA-binding sites and transcriptional regulatory elements, making it difficult to observe significant changes even when dephosphorylation occurs at specific sites. Although we were unable to conduct experiments using gene knockout mice, the relevant research still provides an important basis for further exploration. This is expected to expand our understanding and potentially guide therapeutic strategies for many diseases.
The phosphorylation of GR at Ser226 (orthologous to serine 234 in mice) and Ser203 (orthologous to serine 234 in mice) is regulated by upstream kinases. An in vitro phosphorylation assay suggested that the CDK complex and ERK phosphorylated GR at Ser20332. Interestingly, phosphorylation of the GR by cyclin CDKs was required for full GR-mediated transcriptional activity, as measured by yeast reporter assays. Conversely, ERK activity attenuates the transcriptional activity of GR Ser203. This suggests that the regulation of GR phosphorylation and GR-mediated transcriptional responses by different kinases is not uniform; different kinases can either positively or negatively modulate GR transcriptional activity32. Furthermore, it has been reported that GR Ser226 is directly phosphorylated by c-Jun N-terminal kinase (JNK), but not by p38, and that JNK phosphorylation of GR directly inhibits GR-mediated activation of transcription28. However, the specific mechanisms and pathways by which irisin regulates GR function through these kinases in the GC signaling pathway are unknown. Studies have shown that irisin promotes ERK phosphorylation, enhancing C2C12 cells34. Additionally, irisin activated ERK1/2 signaling and upregulated IGF-1 expression in Dex-treated myotubes, thereby regulating muscle anabolism62. However, there is no direct evidence suggesting that irisin affects CDK activity. Consistent with this, our study showed that irisin inhibits GR transcriptional activity by promoting ERK phosphorylation, which in turn phosphorylates GR at site 212. Additionally, we used JNK and ERK inhibitors both alone and in combination, followed by the administration of Dex and irisin. We found that irisin inhibited the modulation of GR transcriptional responses in both scenarios. This may be due to the fact that both JNK and ERK are members of the MAPK family, and while SP600125 and U0126 are selective inhibitors of these kinases, respectively, at high concentrations, they may also inhibit the activity of other kinases. SP600125 is primarily an inhibitor of JNK; however, studies have shown that it can also inhibit ERK kinase activity63. Similarly, U0126 is primarily an inhibitor of MEK1/2 and indirectly inhibits ERK activity by targeting MEK1/2; however, it may also affect JNK activity64. In any case, the simultaneous inhibition of both JNK and ERK activities significantly inhibited irisin-mediated regulation of both GR phosphorylation and GC genomic responses. This suggests that irisin inhibits the genomic action of GC by promoting JNK and ERK phosphorylation.
Boström et al. hypothesized that the action of irisin is mediated by cell surface receptors. Recent studies have elucidated the potential receptors and signaling pathways involved in irisin52. Kim et al. have demonstrated that irisin binds to integrin V-like proteins and promotes osteoblast survival and sclerostin secretion. Additionally, FNDC5 knockout in C57BL/6J mice completely prevented ovariectomy-induced bone loss, identifying integrin αVβ5 as a functional receptor for irisin in bone and adipose tissue65. Further studies have shown that irisin acts via integrin receptors in various tissues. For example, irisin significantly inhibits the proliferation, migration, and invasion of osteosarcoma cells via the integrin receptor66. In astrocytes, integrin αVβ5 is essential for irisin-induced release of neprilysin, which facilitates amyloid beta clearance and ameliorates Alzheimer’s disease67. However, whether irisin signaling in skeletal muscles is dependent on integrin receptors remains unclear. In our research, we found that inhibiting the binding of irisin to integrin αVβ5 and its downstream signaling significantly decreased the phosphorylation of ERK, JNK, and GR induced by irisin. In addition, in this case, irisin had no effect on the downstream protein levels of IGF-1 and MSTN. These results suggest that irisin could affect GC signaling through the integrin αVβ5 receptor.
In summary, we have used overexpression systems and inhibitors in vitro to demonstrate that GC genomic effects are the main pathway through which GCs induce muscle atrophy in mice; increased glucocorticoid receptor (GR) Ser212 and Ser234 site phosphorylation reduces GC-induced muscle atrophy in mice; irisin reduces GC genomic effects via GR Ser212/Ser234 phosphorylation in mouse muscle atrophy. Additionally, we have examined key components of this pathway in vivo. This revealed the mechanisms of irisin crosstalk with GC genomic and non-genomic effects.
Methods
MalaCards human disease database
Diseases associated with elevated serum GC levels were searched in the MalaCards database (http://www.malacards.org/)68 using the keyword “elevated glucocorticoids levels.” Relevant diseases were retrieved (Supplementary data 1), and the top 200 diseases were ranked by composite relatedness score. These diseases were subsequently classified into 14 categories according to the MalaCards classification system.
Expression and purification of recombinant irisin
As reported in the study, mouse irisin recombinant protein was expressed and purified using the pET expression system (Novagen)36. Briefly, mouse irisin cDNA was PCR-amplified and cloned into the pET-21a expression vector. The plasmid was transformed into E. coli strain BL21 and cultured in LB medium containing 100 mg/L ampicillin. The BL21 strain was induced to produce His-tagged irisin by the addition of 0.5 mM IPTG at 16 °C for 16 h. The lysates of the BL21 strain were then sonicated using a low temperature ultra-high pressure continuous flow cell disruptor. After sonication, the lysates were centrifuged, and the supernatants were collected and purified using a Ni-NTA agarose affinity column (GE) and size exclusion chromatography column (Superdex 75 pg 16/60, Cytiva). The purity of the recombinant irisin (r-irisin) was compared with commercial irisin (c-irisin, Phoenix Biotech, 067-17) by Western blot analysis (Fig. S1A), and the concentration of irisin was determined using a BCA protein assay kit (Thermo Scientific, 23225). Recombinant irisin was used in both in vitro and in vivo experiments.
HFD mice model
C57BL/6JSmoc-Pcsk9em2(hPCSK9)/Smoc (hPcsk9 mice, Cat. NO. NM-HU-00075) mice were purchased from Shanghai Model Organisms Center. At 6 weeks of age, hPcsk9 mice were fed a regular diet (RD:11% kcal from fat) for 2 weeks. The mice were randomized into two groups, which were fed either a regular diet (RD:11% kcal from fat) (NCD, Charles River Co., Ltd., Beijing, China) or an HFD (60% kcal from fat; Beijing HFK Bioscience Co., Ltd.). After 31 weeks, mice from the HFD group were then split into two groups (HFD and HFD+Irisin). Mice in the HFD+Irisin group were given intraperitoneal (i.p.) injections of irisin (1 mg/kg body weight, dissolved in PBS) every day for 4 weeks, while the mice in the other two groups received the same volume of PBS. Then, mice were killed, and gastrocnemius (Gas), tibialis anterior (TA), soleus (Sol) and extensor digitorum longus (EDL) muscles were dissected and frozen in liquid nitrogen for subsequent studies. A schematic representation of this protocol is presented in Fig. 1C.
Aging mice injury model
Aging C57BL/6J mice were maintained on a standard diet for 18 months. TA muscles of young (8 weeks aged mice) and aging mice (18 months) were injured with 50μL CTX (Sigma, 217503, 10 μM). Before injury, young and aging mice were injected intraperitoneally (i.p.) with irisin (1 mg/kg) every day for 3 days; after injury, young and aging mice were injected intraperitoneally (i.p.) with irisin (1 mg/kg) every other day; control mice were injected with same volume of PBS. At the muscle injury, the 8d TA muscle was dissected and frozen in liquid nitrogen for subsequent studies. A schematic representation of this protocol is presented in Fig. 1H.
Dexamethasone-induced muscle atrophy model
6 weeks of age male C57BL/6J mice were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. After two weeks of accommodation, mice were randomized into 6 groups and injected intraperitoneally daily for 10 days accordingly: control (30% PEG 300 in 0.9% saline), Dex (Sigma, D1756, 1 mg/kg body weight, dissolved in PEG 300 solution), irisin (0.5 mg/kg body weight), Dex (1 mg/kg)+irisin (0.5 mg/kg), irisin (1 mg/kg), and Dex (1 mg/kg)+ irisin (1 mg/kg). Mouse body weight was recorded daily. After the last intraperitoneal injection 24h, mice were sacrificed. Gas, TA, EDL, and Sol muscles were isolated and weighed for further analyses.
Muscle histology and immunohistochemistry
TA muscle was frozen in liquid nitrogen using a slow-freeze procedure for histological analysis. Myofiber areas analysis was performed as described previously, the HFD mice and Dex administered mice tissue sections (6 μm thick) were stained with wheat germ agglutinin (WGA) Alexa Fluor™ 488 conjugate (1:100, Thermo Fisher Scientific, W11261). Fiber sizes were measured using Nikon NIS-Elements Br 3.0 software (Melville, NY). The distribution of myofibers was as previously described16,55,69. In the HFD model, approximately 500 fibers per TA muscle were measured, with 3 mice in total, yielding ~1500 fibers. In the Dex model, ~500 fibers per TA muscle were measured, with 5 mice in total, yielding ~2500 fibers. For the aging mouse TA muscle, sections were incubated with Laminin antibody (1:1000, abcam, ab11575) to calculate the CSAs of the central nuclei myofibers. For each mouse, exactly 100 central nuclei myofibers were measured from 5 randomly selected fields of the TA muscle, and 3 mice were analyzed per group.
Immunohistochemistry was performed. Briefly, the cross sections of TA muscles were fixed in 4% paraformaldehyde (PFA) and permeabilized by incubating in 0.03% Triton X-100 in PBS followed by blocking with serum-free protein block (Solarbio) for 1 h at room temperature and 3% H2O2 (GeneTech (Shanghai) Company Limited) for 15 min in dark. Sections were then incubated with a primary antibody anti-GR (1:300, CST, 12041) overnight at 4 °C. Subsequent antibody incubation and DAB color development were performed with the GTVisionTM+Detection System/Mo&Rb (GeneTech (Shanghai) Company Limited). These sections were visualized with a Nikon 80i microscope (Nikon). Images were analyzed by Nikon NIS-Elements Br 3.0 software (Nikon). The percentage of GR-positive cells is the ratio of the GR-positive cells divided by the total number of cells. The mean nuclear GR intensity was calculated as the ratio of the intensity of GR in the nuclear area to that found over the entire cell. We counted 5 random fields of TA muscle section for each mouse, and five mice were analyzed in each group.
Cell culture and differentiation
C2C12 and HEK293T cell lines were obtained from ATCC and cultured at 37 °C, 5% CO2 in high-glucose DMEM-supplemented media (Gibco, 11965092) containing 10% fetal bovine serum (Gibco, 10099158), 1% penicillin-streptomycin combination (Solarbio, P1400) and 1% L-glutamine (Gibco, A2916801). For the induction of differentiation into myotubes, C2C12 was grown to 100% and transferred to DMEM culture medium with 2% horse serum (Cytiva, SH30074.03), and then cultured for 4 days. The myotubes were treated with 25 μM Dex and different doses of irisin (80 nM and 160 nM) for 24 h. For inhibiting the activity of ERK and JNK, C2C12 myotubes were cultured in a medium containing 20 μM JNK inhibitor SP600125 (MCE, HY-12041), ERK inhibitor 50 μM U0126 (MCE, HY-12031), 25 μM Dex and 80nM irisin for 24 h. An integrin-αVβ5 selective inhibitor 5 μM Cilengitide (MCE, HY-16143) was administered, 25 μM Dex and 80nM irisin for 24h.
Immunofluorescence and myotube diameter measurement
C2C12 cells were placed in 6-well plates with cell slides. When cells grown to 100%, transferred to DMEM culture medium with 2% horse serum, and then cultured for 4 days. Then the cells were treated with both DEX and irisin for 24 hours. C2C12 myotubes were fixed with 4% paraformaldehyde for 10 min, permeabilized with 0.3% Triton X-100 for 10 min, and then blocked with serum-free protein block for 1h at room temperature. Cells were incubated with anti-eMyHC (1:50, DSHB, F1.652) overnight at 4 °C, then incubated with secondary antibody conjugated to Alexa 488 (1:1500, CST, 4408). Images were captured using a fluorescence microscope (Nikon 80i) and measured using Nikon NIS-Elements Br 3.0 software (Melville, NY). Myotube diameter was quantified as follows: 6 fields were chosen randomly from three wells per group, and approximately 600 myotubes were measured70.
Real-time PCR analysis and RNA-Seq
RNA was extracted through the Trizol reagent (Invitrogen, 10296010CN). 1 μg RNA was reverse transcribed to cDNA using HiScript III RT SuperMix (Vazyme, R222-01). Quantitative PCR analysis was performed with UltraSYBR Mixture (Cwbio, CW0957M). The relative gene expression levels were calculated by the 2−ΔΔCt method using GAPDH as an internal control. All data were normalized. The primers used in the experiments were UCP-1 5’-ATGTTTGTGTGCTTTGAA-3’,
5’-CAGTATCTCTTCCTCCAA-3’, IGF-1 5’-TGGATGCTCTTCAGTTCGTG-3’ 5’- CACTCATCCACAATGCCTGT-3’, Fbxo32 5’- TGAATAATCCCAGCACACGA-3’ 5’- ATCGGCAACTGCATCTCTTC-3’, Trim63 5’- TGAATAATCCCAGCACACGA -3’, 5’- ATCGGCAACTGCATCTCTTC -3’, MSTN 5’- TGCTGTAACCTTCCCAGGACCA -3’, 5’- TGGAGTGCTCATCGCAGTCAAG-3’, GAPDH 5’-TGCCCCCATGTTTGTGATG-3’ 5’-TGTGGTCATGAGCCCTTCC-3’.
C57BL/6J mice were injected intraperitoneally daily for 10 days accordingly: control (30% PEG 300 in 0.9% saline), Dex (1 mg/kg), irisin (1mg/kg), and Dex (1 mg/kg)+irisin (1 mg/kg), and Gas was harvested to commission Biomarker Technologies for transcriptome sequencing analysis, 3 mice per group. RNA-Seq data were processed using Omicshare tools. Volcano plots and heat maps were generated using OmicStudio tools.
Western blotting
Total proteins were isolated from C2C12 myotubes with RIPA lysis buffer (Beyotime Biotechnology, P0013B). The cytosolic extract (CE) and nuclear extract (NE) were isolated from treated C2C12 myotubes using the Nuclear and Cytoplasmic Protein Extraction kit (Beyotime Biotechnology, P0027). Western blotting was conducted with equal amounts of protein and separated by SDS-PAGE, then transferred to NC membranes. NC membranes were incubated with the following primary antibodies overnight at 4 °C: anti-irisin (1:1500, Phoenix Biotech, H-067-17), anti-Atrogin-1 (1:1000, abcam, ab168372), anti-MuRF-1 (1:1000, abcam, ab183094), anti-p-AKT(Ser473) (1:1000, CST, 9271S), anti-AKT (1:1000, CST, 9272S), anti-GAPDH(1:1000, ZSGBBIO, TA-08), anti-GR (1:1000, CST, 12041S), anti-Histone H3 (1:1000, CST, 3638S), anti-MSTN (1:250, abcam, ab98337), anti-p-Smad3 (1:1000, CST, 9520S), anti-Smad3 (1:1000, CST, 9523S), anti-IGF-1(1:1000, BOSTER, BA0939), anti-p-GR Ser234 (1:1000, orthologous to serine 226 in human, CST, 97285S), anti-p-GR Ser212 (1:1000, orthologous to serine 203 in human, Thermo Fisher, PA5-104446), anti-p-GR Ser134 (1:1000, orthologous to serine 143 in human, CST, 85060S), anti-p-JNK(Thr183/Tyr185) (1:1000, CST, 9251S), anti-p44/42 MAPK (Erk1/2) (1:1000, CST, 4695T), anti-integrin αV(1:1000, abcam, ab179475), anti-integrin β5(1:1000, CST, 3629S).
Plasmid and lentiviral constructs and stable cell line generation
The plasmid GV657 of wild-type GR and GR Ser212A, GR Ser234A, GR Ser212/234A (replacing serine/threonine with alanine mimics dephosphorylation), GR Ser212D, GR Ser234D, GR Ser212/234D (replacing serine/threonine with aspartic acid mimics phosphorylation) mutants were constructed from GV657 plasmid from Mouse GR (Nr3c1) cDNA construct. Those were obtained from Shanghai GeneChem Co., LTD. M. Lentiviral expression plasmids pLV-EF1a-IRES-Puro (Plasmid #85132), envelope plasmids pMD2.G (Plasmid #12259) and packaging plasmids psPAX2 (Plasmid #12260). BamH1 (Takara, 1605) and EcoRI (Takara, 1612) restriction endonucleases were used to cut the pLV-EF1a-IRES-Puro plasmid and then inserted into wild-type GR and GR Ser212A, GR Ser234A, GR Ser212/234A, GR Ser212D, GR Ser234D, GR Ser212/234D amplified from the GV657 plasmid by homologous recombination, respectively.
To generate stable C2C12 cell lines, HEK293T cells were transfected with plasmids with Lipofectamine 3000 (Thermo Fisher, L3000015) for virus packaging. Briefly, HEK293T cells were placed in 6-well plates co-transfected with 1.6 μg pLV-EF1a-IRES-wild-type GR, GR Ser212A, GR Ser234A, GR Ser212/234A, GR Ser212D, GR Ser234D, GR Ser212/234D constructs and 0.8 μg pMD2G and 0.8 μg psPAX2 packaging plasmids. After 48h, virus-containing supernatants were harvested to infect C2C12 cells. After 48h, all stably transfected cells were selected with 2 μg/ml puromycin (Beyotime Biotechnology, ST551) for 2 days, and after 3 generations of screening, the cells were plated in 6 or 12-well plates for validation by indirect immunoblotting with GR and GR Ser212, GR Ser234 antibodies. The 12-well plates were treated with both Dex and irisin, and the RNA was collected for qRT-PCR experiments.
Statistics and reproducibility
GraphPad Prism 8 was used to conduct all statistical analyses. All values are presented as mean ± standard error of the mean. Statistical analyses used the unpaired Student’s t test, one-way ANOVA and two-way ANOVA. Statistical tests were used as described in the figure legends. A value of p < 0.05 was considered statistically significant. The mice were randomized and assigned to different groups. For all experiments, each group consists of a minimum of three biologically independent samples to ensure reproducibility.
Ethical approval
All animal procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of China Agricultural University (Approval No. AW52302202-2-2), and we have complied with all relevant ethical regulations for animal use. The animals used were Mus musculus (strains: C57BL/6J and C57BL/6JSmoc-Pcsk9em2(hPCSK9)/Smoc); sex: male; ages: 6–8 weeks and 18 months. All mice were maintained under a specific pathogen-free facility with a standard chow diet in a 12 h light/dark cycle at the Laboratory Animal Center of China Agricultural University. All observations were included in the analysis without data exclusion. Mice were anesthetized with intraperitoneal pentobarbital sodium and subsequently euthanized by cervical dislocation performed by trained personnel.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The numerical source data underlying all graphs presented in the manuscript are available in Supplementary Data 1, and the dataset retrieved from the MalaCards human disease database is provided in Supplementary Data 2. The transcriptomic dataset has been deposited at NCBI (SRA: PRJNA1347945), and the plasmid (pET-21a-irisin) has been deposited at Addgene (Plasmid ID: 248349). Before the Addgene record becomes searchable and publicly available, the plasmid is openly available from the corresponding author upon reasonable request. All data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Mukund, K. & Subramaniam, S. Skeletal muscle: a review of molecular structure and function, in health and disease. Wiley Interdiscip. Rev. Syst. Biol. Med. 12, e1462 (2020).
Sartori, R., Romanello, V. & Sandri, M. Mechanisms of muscle atrophy and hypertrophy: implications in health and disease. Nat. Commun. 12, 330 (2021).
Burford, N. G., Webster, N. A. & Cruz-Topete, D. Hypothalamic-pituitary-adrenal axis modulation of glucocorticoids in the cardiovascular system. Int. J. Mol. Sci. 18, 2150 (2017).
Möhlmann, J. E. et al. Pharmacokinetics and pharmacodynamics of systemic corticosteroids in autoimmune and inflammatory diseases: a review of current evidence. Clin. Pharmacokinet. 63, 1251–1270 (2024).
Schakman, O., Kalista, S., Barbé, C., Loumaye, A. & Thissen, J. P. Glucocorticoid-induced skeletal muscle atrophy. Int. J. Biochem. Cell Biol. 45, 2163–2172 (2013).
Li, J. X. & Cummins, C. L. Fresh insights into glucocorticoid-induced diabetes mellitus and new therapeutic directions. Nat. Rev. Endocrinol. 18, 540–557 (2022).
Wang, K. et al. Chronic kidney disease-induced muscle atrophy: Molecular mechanisms and promising therapies. Biochem. Pharm. 208, 115407 (2023).
Gassen, N. C., Chrousos, G. P., Binder, E. B. & Zannas, A. S. Life stress, glucocorticoid signaling, and the aging epigenome: implications for aging-related diseases. Neurosci. Biobehav Rev. 74, 356–365 (2017).
Mayayo-Peralta, I., Zwart, W. & Prekovic, S. Duality of glucocorticoid action in cancer: tumor-suppressor or oncogene. Endocr. Relat. Cancer 28, R157–r171 (2021).
Wong, S. C. et al. Growth and the growth hormone-insulin like growth factor 1 axis in children with chronic inflammation: current evidence, gaps in knowledge, and future directions. Endocr. Rev. 37, 62–110 (2016).
Biagetti, B. & Simó, R. GH/IGF-1 abnormalities and muscle impairment: from basic research to clinical practice. Int. J. Mol. Sci. 22, 415 (2021).
Ruiz, D., Padmanabhan, V. & Sargis, R. M. Stress, sex, and sugar: glucocorticoids and sex-steroid crosstalk in the sex-specific misprogramming of metabolism. J. Endocr. Soc. 4, bvaa087 (2020).
Huang, L. T. & Wang, J. H. The therapeutic intervention of sex steroid hormones for sarcopenia. Front. Med. 8, 739251 (2021).
Sato, A. Y. et al. Glucocorticoids induce bone and muscle atrophy by tissue-specific mechanisms upstream of E3 ubiquitin ligases. Endocrinology 158, 664–677 (2017).
Kim, J., Park, M. Y., Kim, H. K., Park, Y. & Whang, K. Y. Cortisone and dexamethasone inhibit myogenesis by modulating the AKT/mTOR signaling pathway in C2C12. Biosci. Biotechnol. Biochem 80, 2093–2099 (2016).
Dong, Y., Pan, J. S. & Zhang, L. Myostatin suppression of Akirin1 mediates glucocorticoid-induced satellite cell dysfunction. PLoS One 8, e58554 (2013).
Waddell, D. S. et al. The glucocorticoid receptor and FOXO1 synergistically activate the skeletal muscle atrophy-associated MuRF1 gene. Am. J. Physiol. Endocrinol. Metab. 295, E785–E797 (2008).
Ahmad, S. S., Ahmad, K., Lee, E. J., Lee, Y. H. & Choi, I. Implications of insulin-like growth factor-1 in skeletal muscle and various diseases. Cells. 9, 1773 (2020).
Wang, Z., Frederick, J. & Garabedian, M. J. Deciphering the phosphorylation “code” of the glucocorticoid receptor in vivo. J. Biol. Chem. 277, 26573–26580 (2002).
Braun, T. P. et al. Cancer- and endotoxin-induced cachexia require intact glucocorticoid signaling in skeletal muscle. Faseb J. 27, 3572–3582 (2013).
Braun, T. P. et al. Muscle atrophy in response to cytotoxic chemotherapy is dependent on intact glucocorticoid signaling in skeletal muscle. PLoS One 9, e106489 (2014).
Lynch, J. T. et al. The role of glucocorticoid receptor phosphorylation in Mcl-1 and NOXA gene expression. Mol. Cancer 9, 38 (2010).
Wang, Z., Chen, W., Kono, E., Dang, T. & Garabedian, M. J. Modulation of glucocorticoid receptor phosphorylation and transcriptional activity by a C-terminal-associated protein phosphatase. Mol. Endocrinol. 21, 625–634 (2007).
Deploey, N., Van Moortel, L., Rogatsky, I., Peelman, F. & De Bosscher, K. The Biologist’s Guide to the glucocorticoid receptor’s structure. Cells. 12, 1636 (2023).
Oakley, R. H. & Cidlowski, J. A. The biology of the glucocorticoid receptor: new signaling mechanisms in health and disease. J. Allergy Clin. Immunol. 132, 1033–1044 (2013).
Takabe, S., Mochizuki, K. & Goda, T. De-phosphorylation of GR at Ser203 in nuclei associates with GR nuclear translocation and GLUT5 gene expression in Caco-2 cells. Arch. Biochem. Biophys. 475, 1–6 (2008).
Rogatsky, I., Logan, S. K. & Garabedian, M. J. Antagonism of glucocorticoid receptor transcriptional activation by the c-Jun N-terminal kinase. Proc. Natl. Acad. Sci. USA 95, 2050–2055 (1998).
Itoh, M. et al. Nuclear export of glucocorticoid receptor is enhanced by c-Jun N-terminal kinase-mediated phosphorylation. Mol. Endocrinol. 16, 2382–2392 (2002).
Miller, A. L. et al. p38 Mitogen-activated protein kinase (MAPK) is a key mediator in glucocorticoid-induced apoptosis of lymphoid cells: correlation between p38 MAPK activation and site-specific phosphorylation of the human glucocorticoid receptor at serine 211. Mol. Endocrinol. 19, 1569–1583 (2005).
Kino, T. et al. Cyclin-dependent kinase 5 differentially regulates the transcriptional activity of the glucocorticoid receptor through phosphorylation: clinical implications for the nervous system response to glucocorticoids and stress. Mol. Endocrinol. 21, 1552–1568 (2007).
Zeyen, L., Seternes, O. M. & Mikkola, I. Crosstalk between p38 MAPK and GR signaling. Int. J. Mol. Sci. 23, 3322 (2022).
Krstic, M. D., Rogatsky, I., Yamamoto, K. R. & Garabedian, M. J. Mitogen-activated and cyclin-dependent protein kinases selectively and differentially modulate transcriptional enhancement by the glucocorticoid receptor. Mol. Cell Biol. 17, 3947–3954 (1997).
Galliher-Beckley, A. J., Williams, J. G. & Cidlowski, J. A. Ligand-independent phosphorylation of the glucocorticoid receptor integrates cellular stress pathways with nuclear receptor signaling. Mol. Cell Biol. 31, 4663–4675 (2011).
Reza, M. M. et al. Irisin is a pro-myogenic factor that induces skeletal muscle hypertrophy and rescues denervation-induced atrophy. Nat. Commun. 8, 1104 (2017).
Zhou, T. et al. Irisin ameliorated skeletal muscle atrophy by inhibiting fatty acid oxidation and pyroptosis induced by palmitic acid in chronic kidney disease. Kidney Blood Press Res. 48, 628–641 (2023).
Guo, M. et al. Irisin ameliorates age-associated sarcopenia and metabolic dysfunction. J. Cachexia Sarcopenia Muscle 14, 391–405 (2023).
Wang, J. et al. Irisin deficiency exacerbates diet-induced insulin resistance and cardiac dysfunction in type II diabetes in mice. Am. J. Physiol. Cell Physiol. 325, C1085–c1096 (2023).
Huh, J. Y., Dincer, F., Mesfum, E. & Mantzoros, C. S. Irisin stimulates muscle growth-related genes and regulates adipocyte differentiation and metabolism in humans. Int. J. Obes. 38, 1538–1544 (2014).
Shan, T., Liang, X., Bi, P. & Kuang, S. Myostatin knockout drives browning of white adipose tissue through activating the AMPK-PGC1α-Fndc5 pathway in muscle. Faseb J. 27, 1981–1989 (2013).
Adhikary, S., Kothari, P., Choudhary, D., Tripathi, A. K. & Trivedi, R. Glucocorticoid aggravates bone micro-architecture deterioration and skeletal muscle atrophy in mice fed on high-fat diet. Steroids 149, 108416 (2019).
Alev, K. et al. Short-time recovery skeletal muscle from dexamethasone-induced atrophy and weakness in old female rats. Clin. Biomech. 100, 105808 (2022).
Shen, Y. et al. Diabetic muscular atrophy: molecular mechanisms and promising therapies. Front. Endocrinol. 13, 917113 (2022).
Dong, J. et al. Inhibition of myostatin in mice improves insulin sensitivity via irisin-mediated cross talk between muscle and adipose tissues. Int. J. Obes. 40, 434–442 (2016).
Braun, T. P. & Marks, D. L. The regulation of muscle mass by endogenous glucocorticoids. Front. Physiol. 6, 12 (2015).
Wang, B. Y. et al. Is dexamethasone-induced muscle atrophy an alternative model for naturally aged sarcopenia model. J. Orthop. Transl. 39, 12–20 (2023).
Hammers, D. W. et al. Glucocorticoids counteract hypertrophic effects of myostatin inhibition in dystrophic muscle. JCI Insight. 5, https://doi.org/10.1172/jci.insight.133276 (2020).
Galliher-Beckley, A. J. & Cidlowski, J. A. Emerging roles of glucocorticoid receptor phosphorylation in modulating glucocorticoid hormone action in health and disease. IUBMB Life 61, 979–986 (2009).
Ingawale, D. K. & Mandlik, S. K. New insights into the novel anti-inflammatory mode of action of glucocorticoids. Immunopharmacol. Immunotoxicol. 42, 59–73 (2020).
de Guia, R. M. Stress, glucocorticoid signaling pathway, and metabolic disorders. Diab. Metab. Syndr. 14, 1273–1280 (2020).
Yu, J. et al. Bone marrow homeostasis is impaired via JAK/STAT and glucocorticoid signaling in cancer cachexia model. Cancers 13, 1059 (2021).
Martin, A., Gallot, Y. S. & Freyssenet, D. Molecular mechanisms of cancer cachexia-related loss of skeletal muscle mass: data analysis from preclinical and clinical studies. J. Cachexia Sarcopenia Muscle 14, 1150–1167 (2023).
Boström, P. et al. A PGC1-α-dependent myokine that drives brown-fat-like development of white fat and thermogenesis. Nature 481, 463–468 (2012).
Colaianni, G. et al. Irisin prevents and restores bone loss and muscle atrophy in hind-limb suspended mice. Sci. Rep. 7, 2811 (2017).
Reza, M. M. et al. Irisin treatment improves healing of dystrophic skeletal muscle. Oncotarget 8, 98553–98566 (2017).
Yang, J. et al. Mst1/2 is necessary for satellite cell differentiation to promote muscle regeneration. Stem Cells 40, 74–87 (2022).
Vettorazzi, S., Nalbantoglu, D., Gebhardt, J. C. M. & Tuckermann, J. A guide to changing paradigms of glucocorticoid receptor function—a model system for genome regulation and physiology. FEBS J. 289, 5718–5743 (2022).
Panettieri, R. A. et al. Non-genomic effects of glucocorticoids: an updated view. Trends Pharm. Sci. 40, 38–49 (2019).
Gerber, A. N., Newton, R. & Sasse, S. K. Repression of transcription by the glucocorticoid receptor: a parsimonious model for the genomics era. J. Biol. Chem. 296, https://doi.org/10.1016/j.jbc.2021.100687 (2021).
Beaupere, C., Liboz, A., Fève, B., Blondeau, B. & Guillemain, G. Molecular mechanisms of glucocorticoid-induced insulin resistance. Int. J. Mol. Sci. 22, 623 (2021).
Kang, S. H. et al. Forkhead box O3 plays a role in skeletal muscle atrophy through expression of E3 ubiquitin ligases MuRF-1 and atrogin-1 in Cushing’s syndrome. Am. J. Physiol. Endocrinol. Metab. 312, E495–e507 (2017).
Blind, R. D. & Garabedian, M. J. Differential recruitment of glucocorticoid receptor phospho-isoforms to glucocorticoid-induced genes. J. Steroid Biochem. Mol. Biol. 109, 150–157 (2008).
Chang, J. S. & Kong, I. D. Irisin prevents dexamethasone-induced atrophy in C2C12 myotubes. Pflugers Arch 472, 2020–502 (2020).
Doan, T. K. P. et al. Inhibition of JNK and ERK pathways by SP600125-and U0126-enhanced osteogenic differentiation of bone marrow stromal cells. Tissue Eng. Regen. Med. 9, 283–294 (2012).
You, Y. et al. U0126: Not only a MAPK kinase inhibitor. Front. Pharm. 13, 927083 (2022).
Kim, H. et al. Irisin mediates effects on bone and fat via αV integrin receptors. Cell 175, 1756–1768.e1717 (2018).
Kong, G. et al. Irisin reverses the IL-6 induced epithelial-mesenchymal transition in osteosarcoma cell migration and invasion through the STAT3/Snail signaling pathway. Oncol. Rep. 38, 2647–2656 (2017).
Kim, E. et al. Irisin reduces amyloid-β by inducing the release of neprilysin from astrocytes following downregulation of ERK-STAT3 signaling. Neuron 111, 3619–3633.e3618 (2023).
Rappaport, N. et al. MalaCards: an amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res. 45, D877–d887 (2017).
Wang, K. et al. MST1/2 regulates fibro/adipogenic progenitor fate decisions in skeletal muscle regeneration. Stem Cell Rep. 19, 501–514 (2024).
Wang, L. et al. Trimetazidine attenuates dexamethasone-induced muscle atrophy via inhibiting NLRP3/GSDMD pathway-mediated pyroptosis. Cell Death Discov. 7, 251 (2021).
Acknowledgements
This study was supported by the National Natural Science Foundation of China Grants 32072804 and 31772685.
Author information
Authors and Affiliations
Contributions
The conceptualization of the study was completed by J.Y. and Y.D.; the methodology was completed by P.Z., Y.A., J.Y., S.T., S.S., and L.Z.; the experiments performed were completed by P.Z., J.Y., Y.A., Y.L. and S.S.; the results analyzed were completed by Y.A., P.Z., and J.Y.; the writing of the manuscript was completed by J.Y., P.Z., Y.A., and Y.D.; the writing of the review & editing was completed by J.Y., T.G., H.W., and Y.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Rowan Hardy and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Simona Chera and Dario Ummarino. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, J., Zhang, P., An, Y. et al. Irisin regulates the phosphorylation of glucocorticoid receptor Ser212 and Ser234 and mediates glucocorticoid-induced muscle atrophy in mice. Commun Biol 8, 1746 (2025). https://doi.org/10.1038/s42003-025-09203-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09203-4