Abstract
N6-methyladenosine (m6A) modifications are prevalent across all mammals and represent the most abundant type of epigenetic modification. With advancements in research, new methods for single-cell m6A modification sequencing and prediction have continuously emerged. These methods provide researchers with powerful tools to explore the landscape of epigenetic modifications at the single-cell level. However, challenges such as operational complexity, limited sensitivity, resolution, and consistency across different techniques remain major obstacles for researchers in this field. In this study, we compared four representative single-cell m6A sequencing and prediction methods based on different principles. We also developed a single-cell m6A database, which is freely accessible online. The database allows users to search for the localization and modification levels of single-cell m6A modifications in human and mouse species on the basis of these methods. It also offers cell definitions, data visualization, and data download options. Additionally, we applied Scm6A to single-cell transcriptome data across cancers and spatial transcriptome data from UCEC to predict and visualize m6A modifications, demonstrating its unique superior performance.
Similar content being viewed by others
Background
N6-methyladenosine (m6A) is the most abundant internal modification found in eukaryotic mRNAs and influences a wide range of cellular processes, including mRNA stability, translation efficiency, splicing, and nuclear export1. As a dynamic and reversible modification, m6A is installed by a complex of “writers,” primarily methyltransferase-like 3 (METTL3) and METTL14, which are recognized by “readers” such as YTH domain-containing proteins and removed by “erasers”, such as the demethylases FTO and ALKBH52,3. Given its regulatory roles in diverse biological pathways, m6A has garnered significant attention in the fields of developmental biology, neuroscience, and cancer research4,5,6. Traditionally, m6A modifications have been studied via bulk methods such as MeRIP-seq (methylated RNA immunoprecipitation followed by sequencing) and m6A-seq7,8. These approaches typically involve immunoprecipitating m6A-modified RNA fragments via specific antibodies, followed by high-throughput sequencing to map the m6A peaks across the transcriptome. While these methods have been instrumental in uncovering the global m6A landscape, their application to bulk RNA samples averages the signal across thousands of cells, potentially obscuring cell–to-cell variability and the heterogeneity of m6A modifications9. These limitations are particularly significant in complex tissues or disease contexts where cellular heterogeneity plays a critical role10.
In recent years, the development of single-cell m6A detection methods has transformed our understanding of m6A dynamics at a much finer resolution. Single-cell technologies enable the profiling of m6A modifications in individual cells, revealing the heterogeneity that is often masked in bulk analyses. Several pioneering methods have emerged in this field, each offering unique advantages and challenges. For example, scDART-seq employs an enzymatic approach to convert m6A sites into C-to-T mutations, allowing for precise m6A site identification during sequencing11. sn-m6A-seq, on the other hand, is tailored for detecting m6A in single nuclei, providing insights into nuclear m6A dynamics12. scm6A-seq adapts the traditional m6A-seq protocol to the single-cell level, enabling more direct translation of bulk m6A-seq results into single-cell contexts13. Moreover, Scm6A offers a streamlined and user-friendly protocol for m6A detection, making it accessible for broader applications and facilitating its integration into various research settings14. Despite these advancements, there remains a need for a systematic comparison of these methods to understand their relative strengths and limitations in different experimental contexts. Moreover, the simplicity and versatility of Scm6A present opportunities to extend its application beyond standard single-cell m6A profiling. In this study, we set out with two main objectives. First, we conducted a comprehensive comparison of the available single-cell m6A sequencing and prediction methods, focusing on their underlying principles, advantages and disadvantages, complexity, and cost. To support this, we constructed a dedicated database that collates and organizes data generated by scDART-seq, sn-m6A-seq, scm6A-seq, and Scm6A, providing a valuable resource for researchers in the field. Second, given the practicality of Scm6A, we explore its broader application potential in various research scenarios. One such application is the exploration of m6A heterogeneity across different cancer types via pan-cancer datasets. By applying Scm6A to these datasets, we aimed to uncover potential associations between m6A modification patterns and cancer heterogeneity. Additionally, we extended the use of Scm6A to spatial transcriptomics data, allowing us to predict and visualize the spatial distribution of m6A modifications within tissues. This novel application could offer unprecedented insights into how m6A modifications are correlated with spatial gene expression patterns. Overall, this study aims to advance our understanding of single-cell m6A modifications by providing a critical evaluation of existing methods and demonstrating the potential of Scm6A in novel research contexts. By building a comprehensive database and exploring new applications, we hope to provide the scientific community with tools and insights that will drive further discoveries in the field of m6A biology.
Results
Comparison of the advantages and disadvantages of different single-cell m6A prediction tools
Among the emerging single-cell m6A identification technologies, scDART-seq, scm6A-seq, sn-m6A-CT, and Scm6A stand out as representative methods on the basis of entirely different principles, each with its own advantages and limitations, garnering widespread attention. Unlike the three experimental methods for detecting m6A sites, Scm6A is currently the only available computational method for single-cell m6A prediction; its results remain hypothetical until experimentally validated and are not equivalent to experimentally identified sites. Table 1 lists and compares these four methods. Specifically, the scDART-seq method was initially established using stable HEK293T cell lines. By overexpressing the APOBEC1-YTH protein in cells, it enables C → U conversion within the cell, and through genome alignment, it achieves single-base precision for m6A modification sites. This method uses droplet-based or plate-based single-cell omics platforms for sequencing, enabling high-throughput m6A localization and quantification. However, this method relies on efficient cell transfection, requires stable overexpression of the fusion protein in cells, and is not suitable for samples that are difficult or impossible to transfect. Additionally, it can detect only m6A-containing RNA bound by the YTH domain. The scm6A-seq method was initially developed using cleavage-stage mouse embryo cells. It employs MeRIP and RNA multiple-labeling techniques on plate-based single-cell omics platforms to achieve m6A localization and quantification. Compared with scDART-seq, scDART-seq does not require the overexpression of exogenous genes, but it necessitates single-cell sorting and RNA isolation during sample preparation, with the addition of two barcodes, making the process complex and time-consuming. In addition, its most significant drawback is that it remains a low-throughput sequencing method. The sn-m6A-CT method was used to construct the native population of the cell nucleus. The authors of this method used mouse embryonic stem cells and applied CUT&Tag technology to a droplet-based single-cell omics platform to achieve high-throughput analysis of single-nucleotide m6A methylation and the transcriptome. Notably, most methods use RNA extracted from entire cells for m6A mapping, whereas sn-m6A-CT analyzes RNA from nuclear isolates. The advantage of this technique is that RNA molecules with m6A modifications can be enriched in situ without the need to isolate RNA from cells. However, this method relies on m6A-specific antibodies and has a low resolution for identifying m6A peaks, ranging from 50 to 200 bp, and although it can locate m6A modifications on mRNAs, it cannot quantify them. In contrast to the aforementioned experimental techniques, Scm6A is a machine learning model based on trans-m6A regulatory factors and cis-m6A sequences. This method enables rapid, low-cost, high-throughput m6A prediction without experimental procedures, making it suitable for a broad range of single-cell sequencing data analyses. Its drawback is that it can predict only sites included in the constructed model and requires preexisting single-cell sequencing data as input. Moreover, compared with the results obtained from experimental sequencing, the model’s predictions are considered reference values rather than true values. In summary, each method has distinct characteristics in terms of applicability and precision. Researchers should choose the most appropriate technique on the basis of specific requirements to increase the efficiency and accuracy of single-cell m6A methylation analysis.
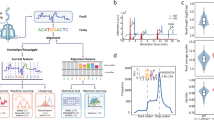
We compared the m6A sites detected via each method with those identified via m6A individual-nucleotide resolution crosslinking and immunoprecipitation (miCLIP). miCLIP is an experimental technique that enables the precise mapping of m6A modifications at single-nucleotide resolution. By combining UV crosslinking, immunoprecipitation with m6A-specific antibodies, and high-throughput sequencing, miCLIP allows for the identification of exact m6A sites on RNA transcripts. Since scm6A-seq and sn-m6A-CT are methods constructed using mouse cells, while scDART-seq and Scm6A are methods constructed using human cell lines, we downloaded BED files for Hek293T (GSE63753) and mESC (GSE86336) CLIPseq datasets for comparison with the aforementioned methods. Then we overlapped the m6A positions of miCLIP with the m6A sites sequenced or calculated by these four methods to verify whether the sites measured by each method were consistent with the miCLIP results. Figure 1A presents a bar graph showing the percentage of m6A-modified genes identified by each method that were validated and not validated by miCLIP. The genes with m6A modifications detected by sn-m6A-CT had the highest proportion validated by miCLIP, reaching 85.7%, whereas scDART-seq had the lowest proportion at 52.8%. However, in terms of quantity, scm6A-seq detected the most m6A-modified genes, identifying a total of 9,052 genes with m6A sites, of which 4,797 were validated by miCLIP. Figure 1B shows the distribution of m6A sites detected by each method on mRNAs. For scDART-seq, the detected m6A sites were more enriched in the 3’UTR, with two distinct peaks near the stop codon and the distal region of the 3’-UTR, and less enrichment in the 5’UTR and CDS. Scm6A detected m6A sites that were concentrated in the CDS and 3’UTR regions, with only one peak near the stop codon. Scm6A-seq revealed two prominent peaks near the start and stop codons. The distribution of sn-m6A-CT was slightly different, with peaks located in the CDS region, which is related to the use of nuclear extracts for sequencing. Figure 1C compares the sequence motif preferences of the m6A sites identified by the four methods. scDART-seq showed a strong preference for the AAACA and AAACT motifs. Scm6A-seq and sn-m6A-CT had overlapping motif preferences and were enriched mainly in AGACA, GGACA, and GGACC motifs. Scm6A showed a preference for motifs such as GGACA and GGACT. This comparison indicates that despite the different methods used, there are common sequence motifs between them, reflecting the common biological characteristics of m6A modification and emphasizing the reliability of these methods in the study of m6A modification at the single-cell level.
A The bar chart visually represents the proportion of miCLIP-validated m6A-modified genes among those identified by each method. Blue indicates m6A sites detected by the method that match miCLIP sequencing results, while yellow indicates those that do not match. B The distribution patterns of m6A sites detected by each method across different regions of the mRNA transcript, including the 5’ UTR, CDS, and 3’ UTR. C Sequence motif analysis of m6A sites identified by the four methods and two miCLIP reference datasets, demonstrating the preference of different detection methods for specific motif sequences.
Pan-cancer analysis of single-cell m6A modifications
Previous studies have shown that m6A RNA methylation is highly heterogeneous across different types of tumors15. In this study, we collected single-cell sequencing data from 29 cancer types using multiple public databases (Supplementary data 1). Then we predicted single-cell m6A levels using Scm6A, merged the prediction results with our custom Python scripts, removed batch effects using the Python package Scanorama, and performed dimensionality reduction and UMAP visualization with the scanpy package to project the data into a two-dimensional space. The results revealed significant heterogeneity in single-cell m6A modification patterns among different cancer types (Fig. 2A). Notably, certain cancer types, such as acute myeloid leukemia (AML) and bladder cancer (BLCA), formed tightly clustered groups, indicating highly specific m6A modification patterns within these groups. In contrast, other cancer types, such as head and neck cancer (HNSC) and thyroid cancer (THCA), exhibited more dispersed clusters, reflecting greater heterogeneity in their m6A modification profiles. Some cancer types, such as prostate adenocarcinoma (PRAD), glioblastoma (GBM), and hepatocellular carcinoma (HCC), display distinct clustering characteristics in terms of m6A modifications, which may be related to their unique RNA methylation regulatory mechanisms. Moreover, biologically similar cancers, such as lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC), which share some similarities in m6A modification patterns, can still be clearly distinguished via visualization, providing clues for further investigation into their molecular differences. The clear separation and distinct clustering of most cancer types at the single-cell level underscore the potential of m6A modifications as biomarkers for cancer classification, and the variability observed in some clusters highlights the complexity of m6A modifications at the single-cell level.
A UMAP projection displaying the clustering of single-cells from various cancer types based on their m6A modification profiles. Each color represents a different cancer type, as indicated by the legend on the right. B, C Heatmaps showing the expression patterns of RBPs and m6A-modified genes in UCEC across different cell types: Epithelial cells, Fibroblasts, T cells, and B cells. The color scale represents the expression z-score, with red indicating higher expression and blue indicating lower expression. D Correlation heatmap illustrating the relationship between m6A modification levels and RBP gene expression in UCEC. The color scale represents the Pearson correlation coefficients, with red indicating a positive correlation, blue indicating a negative correlation, and yellow representing no significant correlation. Genes labeled below the image are some well-known m6A regulators, whereas those labeled on the right are m6A target genes. E Pie charts displaying the distribution of m6A modification sites across different regions of mRNA (5’ UTR, 3’ UTR, start codon, and other exonic regions) in four cell types. The percentage of m6A sites within each region is shown, revealing variation in m6A localization depending on the cell type. F Density plot illustrating the distribution of m6A modification sites along the length of mRNA transcripts (from 5’ UTR to 3’ UTR) in the four cell types. The peak densities indicate the regions where m6A sites are most frequently located, with distinct patterns observed across the different cell types. G Sequence motif analysis showing the top enriched m6A motifs in different cell types, with associated p-values. The motifs are visualized using sequence logos, highlighting the conserved sequences across B cells, T cells, Epithelial cells, and Fibroblasts.
We selected uterine corpus endometrial carcinoma (UCEC) as an example for further studies. By annotating cells within UCEC, we identified five cell types, but the NK cell population contained fewer than five cells and was filtered out, leaving four major cell types: epithelial cells, fibroblasts, B cells, and T cells (Supplementary data 2). We then calculated the gene expression levels of 565 RNA-binding proteins (RBPs) across these different cell types and visualized the data via a heatmap (Fig. 2B, Supplementary data 3). The results revealed significant differences in RBP gene expression levels among the various cell types, with some RBP genes showing high expression in specific cell types and low expression in others, indicating that they regulate different RNAs in various cell types. Simultaneously, we employed Scm6A to assess the modification levels of 4162 single-cell m6A sites, averaging the data by cell type and presenting it in another heatmap (Fig. 2C). The findings indicated marked differences in m6A modification levels among the different cell types, with certain m6A sites exhibiting relatively high modification levels in specific cell types. These findings imply that these modifications may play key regulatory roles in certain cell types. Previous studies have reported that m6A modifications are highly heterogeneous across cells, which may be related to the specific functions and states of different cell types16,17. Our results highlight the significant disparities in gene expression and epigenetic modifications among different cell types in UCEC, providing insights into the relationships between m6A heterogeneity and the distinct functions and states of various cell types in the context of UCEC. In addition, we found obvious correlations between RBP gene expression levels and m6A modification levels in UCEC. In Fig. 2D, the red regions indicate positive correlations, suggesting that the expression of certain RBP genes may affect the modification levels of the corresponding m6A sites. These findings suggest that these genes may actively contribute to the increase in m6A modifications, potentially increasing gene stability or regulating gene expression levels. The blue regions indicate negative correlations, implying that high expression of certain RBP genes is associated with reduced m6A modification levels. These findings might indicate that these genes play a role in inhibiting m6A modifications, affecting mRNA splicing, nuclear export, or degradation processes. Light-colored regions in the heatmap may indicate weak or insignificant correlations between RBPs and m6A sites. This could be due to the functions of these RBPs being relatively independent of m6A modification regulatory mechanisms or because their roles are influenced by other molecular pathways. We found several well-characterized m6A-related genes (e.g., METTL3, FTO, YTHDF1, etc.) and their known target genes or pathways within the heatmap.
In our analysis of m6A modifications in the four main cell types in the UCEC sample, we observed cell type-specific patterns. These patterns revealed clear differences in the distribution, density, and associated motifs of m6A modifications. Figure 2E displays the distribution of m6A modifications predicted by Scm6A across different gene regions. The results show that m6A modifications are most abundant in the 3’ untranslated region (3’ UTR) across all cell types, accounting for nearly half of the modifications (38.73–47.31%). In contrast, the 5’ untranslated region (5’ UTR) has a lower proportion of m6A modifications (9.58–14.35%), while the exon regions (other exons) also have a high level of m6A modifications (35.2–44.6%), indicating a selective preference for m6A modifications in gene expression regulation, which is consistent with current knowledge. Moreover, the distribution varies slightly between cell types. For example, epithelial cells and fibroblasts have a greater proportion of 5’ UTRs (14.35% and 13.07%, respectively) than B and T cells do. Figure 2F shows the density distribution of m6A modifications predicted by Scm6A across the genome. The results show that the distribution patterns in the four cell types are similar, with peaks occurring mainly near the junction of the CDS and 3’-UTR regions, which is related to the role of m6A in regulating mRNA stability and translation efficiency. Figure 2G presents the motif analysis of m6A modifications in different cell types. We observed that while the classic DRACH motif (D = A/G/U, R = A/G, H = A/C/U) is present in all cell types, the composition and frequency of motifs vary between cell types. For example, in B cells and fibroblasts, m6A modifications in the GAACA motif are more prominent. Epithelial cells are rich in the G[G/U]AC motif, whereas T cells exhibit a greater density of modifications in the GAACA motif. These cell-type-specific motif patterns may indicate the biological functions and regulatory mechanisms of m6A modifications in different cell types. In summary, these analyses demonstrate that the global m6A trends captured by Scm6A are reliable. To further visualize the precise level and spatial location of m6A within individual cells, we next performed clustering of single cells based on both their spatial positions and m6A signal intensities.
Single-cell spatial transcriptome analysis
Currently, research on the spatial localization and visualization of RNA methylation is relatively limited. However, by utilizing Scm6A, we can integrate spatial transcriptomics data to conduct spatial m6A analysis. This approach allows for a more comprehensive understanding of m6A distribution within different cellular contexts, enhancing our ability to study the spatial dynamics of RNA methylation. In this study, we employed spatial transcriptomics technology to explore UCEC samples, with a particular focus on several genes closely associated with cancer progression and their m6A modification levels. First, we analyzed the spatial transcriptomics data of UCEC via the standard Seurat workflow. We treated each spot as the same type of cell for a rough analysis, and the results were displayed through UMAP plots and spatial transcriptomics images. Figure 3A shows the UMAP plot of all genes, and the clusters were visualized in different colors on the tissue sections. Under default parameters, the cells were divided into five types, each of which exhibited distinct regionality in the tissue sections (Fig. 3B). We then extracted the gene expression data of the RBPs for reclustering and spatial imaging (Fig. 3C). Similarly, under default parameters, cells were divided into four types, with each type showing the corresponding regionality on the tissue sections (Fig. 3D), indicating that despite the smaller number of RBP genes than all genes, they still show significant representativeness. Next, we used Scm6A to calculate the m6A modification levels of each spot via spatial transcriptomics, clustered them, and mapped them onto the tissue images (Fig. 3E, F). We observed a certain similarity between the m6A and RBP UMAP plots, which possibly suggests a potential functional relationship between m6A and RBPs (Supplementary Fig. 1). Finally, we analyzed the m6A modification levels of the HSP90AB1 and HSP90AA1 genes. These genes encode two isoforms of heat shock protein 90 (HSP90), which play crucial roles in maintaining protein homeostasis, cell signaling, and the response to various stress conditions. Aberrant expression or activity of HSP90 is often related to the survival, proliferation, and drug resistance of tumor cells. As shown in Fig. 4A, B, we observed differences in the m6A modification levels among different transcript variants of HSP90AA1 and HSP90AB1. For example, the transcript variants ENSG00000080824-8 and ENSG00000080824-8 of HSP90AA1 presented relatively high modification levels, whereas ENSG00000080824-5 and ENSG00000080824-7 presented relatively low levels of modification (Wilcoxon rank-sum test, P < 2.2 × 10−16) (Fig. 4A). Similarly, the transcript variants ENSG00000096384-8 and ENSG00000096384-12 of HSP90AB1 presented relatively high and widespread m6A modifications, whereas ENSG00000096384-2, ENSG00000096384-4, and ENSG00000096384-10 presented relatively low levels (Fig. 4B). The differences in m6A modification levels within the HSP90AB1 and HSP90AA1 genes may indicate variations in m6A at different positions within the same gene. High m6A modification levels might be associated with increased mRNA stability and translation efficiency, thereby promoting the expression of HSP90 proteins. This increase in HSP90 protein levels could influence the adaptability and survival capabilities of tumor cells. Future research is needed to further elucidate the specific roles of m6A modifications in the regulation of HSP90 gene expression and how these modifications interact with the tumor microenvironment to promote tumor development.
A, B UMAP and spatial transcriptomic maps generated using the expression profiles of all genes in the sample. These maps show the clustering of cells based on their gene expression patterns, with each cluster represented by different colors corresponding to distinct cell identities or states. C, D UMAP and spatial transcriptomic maps highlighting the expression of RBP genes within the sample. The clustering patterns emphasize the distribution and localization of RBP expression across the tissue. E, F UMAP and spatial transcriptomic maps based on m6A modifications predicted using the Scm6A software. These maps display the distribution of single-cell m6A methylation across different regions of the tissue, illustrating the spatial heterogeneity of m6A modifications.
A, B Spatial transcriptomic images showing the expression levels of two cancer-related genes (HSP90AA1 and HSP90AB1) alongside their corresponding m6A modification levels.
Construction of the SCMD website
On the basis of the four single-cell m6A sequencing and prediction methods, we developed SCMD, a single-cell m6A database. This database focuses on m6A modifications at the single-cell level and is designed for storing, categorizing, and visualizing m6A modification data. Figure 5A shows its construction flowchart. SCMD provides a search function that allows users to select from scDART-seq, scm6A-seq, sn-m6A-CT, and Scm6A methods (Fig. 5B), and users can then search for genes of interest to obtain information on m6A modification sites on those genes. When using the Scm6A method, searching for a gene returns its expression values and m6A modification levels across different diseases, together with a bar chart summarizing the results (Fig. 5C). In contrast, searching for a disease provides gene expression and m6A modification level data from affected individuals, along with corresponding cell annotations and visualizations using t-SNE and UMAP (Fig. 5D). The database includes over 800,000 entries for human and mouse species, and all the data are available for online download (Fig. 5E).
A Schematic workflow illustrating the construction of the SCMD website. The process involves integrating data from multiple single-cell m6A sequencing and prediction methods, including Scm6A, scm6A-seq, sn-m6A-CT, and scDART-seq. The collected data is processed for UMAP visualizations, m6A site identification, and cell annotation, allowing users to explore m6A modifications at the single-cell level. Created in BioRender. Liu, Z. (2025) https://BioRender.com/3poviur. B Screenshot of the SCMD homepage, where users can select different single-cell m6A sequencing and prediction methods to analyze specific genes or datasets. The selection includes options like Scm6A, scm6A-seq, sn-m6A-CT, and scDART-seq. C Example output of the SCMD when using the Scm6A method to search for a specific gene. The table displays the single-cell m6A modification levels across different cancer types, and the bar chart below shows the variation in m6A modification levels across various cell types within those cancers. D Example output of SCMD when using the Scm6A method to search for specific diseases. The output table includes the location information of m6A modification sites associated with the disease and the modification levels in each cell type. t-SNE and UMAP plots display the distribution of m6A modifications across cells (cell types are automatically annotated using SingleR). E Download interface of the SCMD, where users can access datasets generated by different methods and download data for various diseases, such as AML, GBM, and BRCA. The interface allows for easy retrieval of m6A-related data for further analysis.
Discussion
Rapid advancements in single-cell m6A identification techniques have significantly expanded our understanding of epigenetic modifications at an unprecedented resolution. The methods for single-cell m6A detection can be broadly categorized into two types: sequencing-based and computational algorithm-based approaches. Sequencing-based methods, such as scDART-seq, sn-m6A-CT, and scm6A-seq, provide direct measurements of m6A sites but are limited by their reliance on experimental procedures, which can be technically challenging and resource-intensive. On the other hand, computational methods such as Scm6A offer a rapid and cost-effective alternative, leveraging machine learning models to predict m6A sites from existing single-cell sequencing data. However, these predictions are contingent upon the quality and representativeness of the input data and the accuracy of the underlying algorithms. These constraints highlight the need for careful consideration when selecting an appropriate method on the basis of the specific research goals and available resources.
Our pan-cancer analysis using Scm6A revealed significant heterogeneity in m6A modification patterns across different cancer types, underscoring the potential of m6A as a biomarker for cancer classification18. The distinct clustering of m6A modification patterns in certain cancers, such as AML and BLCA, suggests that m6A modifications may play a critical role in defining the molecular identity of these cancers. The variability observed in other cancer types, such as HNSC and THCA, points to the complexity of m6A modifications and the need for further investigation to unravel the underlying mechanisms driving this heterogeneity. In our spatial transcriptomics analysis, we extended the application of Scm6A to explore the spatial distribution of m6A modifications within UCEC tissues. This spatial dimension adds another layer of complexity to our understanding of m6A regulation and suggests that m6A modifications may contribute to the functional diversity of cells within the TME. The field of single-cell epigenetics is poised for further expansion with the integration of other types of modifications and the advent of spatial and long-read single-cell sequencing technologies. The exploration of other RNA modifications, such as m5C, m7G, and pseudouracil, alongside m6A, will provide a more holistic view of the epitranscriptome and its regulatory roles. Spatial transcriptomics, which captures the spatial context of gene expression, can be extended to include m6A, m5C, m7G, etc., offering insights into the spatial heterogeneity of RNA methylation within tissues. Long-read single-cell sequencing technologies have the potential to provide more continuous and accurate genomic information, which could increase the resolution of m6A detection and functional annotation.
The development of the SCMD database represents a significant step forward in making single-cell m6A data accessible to the broader research community. By providing a user-friendly platform for searching, visualizing, and downloading m6A modification data, SCMD facilitates the integration of m6A analysis into a wide range of research projects. Cross-reference m6A data with gene expression and spatial transcriptomics data opens new possibilities for exploring the functional implications of m6A modifications in various biological contexts.
SingleR is one of the most widely used tools for single-cell annotation; however, it relies on reference transcriptomes that may not fully capture the cell-type heterogeneity within our tissue, potentially leading to mis-annotation of closely related subpopulations. While SingleR has certain limitations compared to manual annotation and may not provide highly accurate classification, its overall stability is relatively reliable. Such errors could dilute genuine m6A signal differences or generate spurious ones. Although manual curation of marker genes can mitigate these issues, residual inaccuracies are an inherent limitation. Therefore, we manually inspected our SingleR annotations to ensure their accuracy.
Conclusions
In conclusion, while each of the single-cell m6A identification methods evaluated in this study offers distinct advantages, their limitations must be carefully weighed against the specific requirements of the research at hand. The findings from our pan-cancer and spatial transcriptomics analyses underscore the potential of m6A modifications as biomarkers and regulatory elements in cancer biology. However, the complexity and heterogeneity of m6A modifications necessitate further research to fully elucidate their roles in gene regulation and disease progression. The SCMD database will undoubtedly serve as a valuable resource for researchers aiming to explore these questions, advancing our understanding of m6A biology in the context of single-cell and spatial genomics.
Materials and methods
Data collection
The data sources for this study include scDART-seq, scm6A-seq, and sn-m6A-CT datasets retrieved from their respective publications’ supplementary materials: scDART-seq data from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8857065/, scm6A-seq data from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9852475/, and sn-m6A-CT data from https://www.cell.com/molecular-cell/fulltext/S1097-2765(23)00649-4?_returnURL=https%3 A%2 F%2Flinkinghub.elsevier.com%2Fretrieve%2Fpii%2FS1097276523006494%3Fshowall%3Dtrue#supplementaryMaterial. The Scm6A tool used for single-cell m6A data analysis was sourced from https://github.com/Ansanqi/Scm6A, and additional single-cell data were obtained from repositories such as the Gene Expression Omnibus (GEO) and ArrayExpress, as detailed in Supplementary data 1 of relevant publications. Spatial transcriptomics data downloaded from GSM6177623 in GSE203612 on GEO.
Single-cell transcriptome data analysis
In this study, single-cell analysis was conducted via R software (version 4.4.0). The Seurat package (version 5.1.0)19 was used to analyze the scRNA-seq data. Cells with fewer than 200 or more than 8000 expressed genes, with total molecular counts per cell less than 500 or exceeding 40,000, or with mitochondrial unique molecular identifiers (UMIs) exceeding 20% were filtered out as low-quality cells. The mitochondrial, ribosomal, and hemoglobin genes were subsequently removed from the dataset. The expression levels were normalized via the normalizeData function in the Seurat package, and the top 2500 variable genes were identified via the FindVariableFeatures function. Principal component analysis (PCA) was performed on the variable genes, followed by uniform manifold approximation and projection (UMAP) dimensionality reduction using the top 40 significant principal components (PCs). Cell type annotation was performed automatically via SingleR (version 2.6.0)20.
In-depth single-cell analysis of UCEC was conducted, with a focus on cell types annotated with more than 10 cells to ensure data representativeness and reliability. Furthermore, expression data for 565 RNA-binding proteins (RBPs) were extracted from the gene expression matrix, and their expression levels across different cell types were calculated. Heatmaps were generated to illustrate the relative expression of these RBPs in epithelial cells, fibroblasts, B cells, and T cells. Additionally, using Python (version 3.8.10) and Scm6A14, we predicted the modification levels of 4162 m6A sites in single cells and computed their means by cell type, creating heatmaps via the pheatmap package in R. Visualization of single-cell m6A modifications was also achieved through UMAP plots, following a workflow similar to that for gene expression via Seurat.
Spatial transcriptomic data analysis
Similarly, we utilized the Seurat package (version 5.1.0) to analyze the spatial transcriptomics (ST) data. As the spatial transcriptomic data downloaded from GEO had already been filtered, unlike scRNA-seq, the step of filtering low-quality cells was omitted. We normalized the data via SCTransform, followed by principal component analysis (PCA) of the normalized data. Using the FindNeighbors function based on the top 40 PCA dimensions, we computed a cell adjacency matrix and subsequently performed cell clustering with the FindClusters function. Additionally, we employed the RunUMAP function to perform UMAP dimensionality reduction for better visualization of the clustering results. Finally, we visualized the clustering results by generating a two-dimensional reduction plot in the UMAP space and overlaying the clustering results onto the spatial transcriptomic images.
Database implementation
In this study, we aimed to increase the accessibility and utility of single-cell-level m6A modification data. To achieve this goal, we developed an online query platform called single-cell m6A database (SCMD). SCMD is implemented via the Django (version 3.2.25) web application framework and various common front- and back-end development technologies (HTML, CSS, and JavaScript) to enable functionalities such as web page display, rendering, and data querying. Data visualization is facilitated by bootstrapping (version 3.4.1) and Layui (version 2.6.8). Data processing is performed by Pandas (version 1.3.5) and Numpy (version 1.21.5). SCMD offers a convenient web interface for searching, browsing, and downloading data related to single-cell-level m6A modifications. The core strength of SCMD lies in its robust backend algorithms deployed on the server, enabling rapid response to user queries. Users simply need to input the gene name or disease name of interest, and the system quickly provides relevant gene expression data and m6A modification information. Furthermore, SCMD features an intuitive user interface, making it accessible even to nonspecialist researchers. Our goal is to break down informational barriers, enabling more researchers to easily access and utilize these valuable data resources, thereby advancing research in the fields of single-cell biology and epigenetics. We are also continuously updating and expanding the database content to reflect the latest research findings in the field. The platform is publicly accessible via a website (http://www.splicedb.net:8088/home/).
Using Scm6A to calculate m6A modification levels at each point in spatial transcriptome data
First, we performed a systematic analysis of the spatial transcriptomic data of UCEC (Uterine Corpus Endometrial Carcinoma) using the standard Seurat workflow. This process included data import, quality control filtering, normalization, selection of highly variable genes, principal component analysis (PCA), clustering, and non-linear dimensionality reduction (such as UMAP). We downloaded the BED files for HEK293T (GSE63753) and mESC (GSE86336), and used custom shell scripts to tally the number and genomic positions of m6A sites, as well as the counts of distinct motifs for each dataset. In processing the spatial transcriptomic data, each spot was approximately treated as being composed of a single cell type, although in reality, each spot may contain multiple cells. This simplifying assumption facilitates the analysis of spatial gene expression patterns at the level of overall tissue structure. Upon completing the basic analysis, we generated both UMAP plots and spatial visualizations overlaid with tissue sections.
Next, we input the preprocessed UCEC spatial transcriptomic expression matrix into our self-developed Scm6A computational model. By integrating the gene expression information of each spot, the model predicted the m6A methylation levels at each spot. After obtaining the estimated m6A levels across all spots, we applied the same visualization strategy used in the Seurat analysis to display the variation in m6A levels across spots in a UMAP layout, as well as to intuitively present their spatial distribution on tissue sections. This approach allowed us to reveal potential spatial epigenetic heterogeneity within the tumor tissue.
miCLIP-seq
We overlapped the m6A sites identified by each of the four methods (whether experimentally sequenced or computationally predicted) with the m6A positions reported by miCLIP, to assess the concordance of every method against the miCLIP reference.
Metagene visualization of m6A distribution
For metagene visualization of m6A distribution, we first converted genomic coordinates to transcript coordinates by anchoring each gene to its most highly expressed isoform. Every site was then classified as 5’ UTR, CDS, or 3’ UTR according to its location on the transcript. After rescaling these positions to a relative 0–100% transcript axis, we plotted the normalized density of methylated sites across the transcript.
Network of RNA-binding proteins with m6A
The list of the 565 RBPs is provided in Supplementary data 4. These genes originate from our previous study21, in which we identified 565 m6A-regulated RBPs that exhibit specificity for m6A modification. In our previous work, we established a reliable m6A–RBP interaction network and, based on this correspondence, built predictive models for every m6A site within the network. The total number of such sites—and thus the number of models—is 4162. These modifications are resolved at the single-cell, single-transcript level with a positional accuracy of 100–300 bp.
Data availability
SCMD is available at http://www.splicedb.net:8088/home/. This website is free and open to all users, and there is no login requirement. The source data behind the graphs can be found in Supplementary Data 5 and 6.
Code availability
The code used for part of the visualization and computation has been uploaded to GitHub and is available at: https://github.com/YueqiLi-github/SCMDcode.git.
References
Mukherjee, N. et al. Deciphering human ribonucleoprotein regulatory networks. Nucleic Acids Res. 47, 570–581 (2019).
Meyer, K. D. & Jaffrey, S. R. Rethinking m(6)A Readers, Writers, and Erasers. Annu. Rev. Cell Dev. Biol. 33, 319–342 (2017).
Zaccara, S. & Jaffrey, S. R. A Unified Model for the Function of YTHDF Proteins in Regulating m(6)A-Modified mRNA. Cell 181, 1582–1595.e1518 (2020).
Akhtar, J. et al. m(6)A RNA methylation regulates promoter- proximal pausing of RNA polymerase II. Mol. Cell 81, 3356–3367.e3356. (2021).
An, Y. & Duan, H. The role of m6A RNA methylation in cancer metabolism. Mol. Cancer 21, 14 (2022).
Feng, J. et al. Soot nanoparticles promote ferroptosis in dopaminergic neurons via alteration of m6A RNA methylation in Parkinson’s disease. J. Hazard Mater. 473, 134691 (2024).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206. (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Lang, C. et al. Single-cell sequencing of iPSC-Dopamine neurons reconstructs disease progression and identifies HDAC4 as a regulator of Parkinson cell phenotypes. Cell Stem Cell 24, 93–106.e106. (2019).
Contreras-Trujillo, H. et al. Deciphering intratumoral heterogeneity using integrated clonal tracking and single-cell transcriptome analyses. Nat. Commun. 12, 6522 (2021).
Tegowski, M., Flamand, M. N. & Meyer, K. D. scDART-seq reveals distinct m(6)A signatures and mRNA methylation heterogeneity in single cells. Mol. Cell 82, 868–878.e810 (2022).
Hamashima, K. et al. Single-nucleus multiomic mapping of m(6)A methylomes and transcriptomes in native populations of cells with sn-m6A-CT. Mol. Cell 83, 3205–3216.e5 (2023).
Yao, H. et al. scm(6)A-seq reveals single-cell landscapes of the dynamic m(6)A during oocyte maturation and early embryonic development. Nat. Commun. 14, 315 (2023).
Li, Y. et al. Scm6A: A Fast and Low-cost Method for Quantifying m6A Modifications at the Single-cell Level. Genomics Proteom. Bioinforma. 22, qzae039 (2024).
Lin, Y. et al. Pan-cancer analysis reveals m6A variation and cell-specific regulatory network in different cancer types. Genomics Proteom. Bioinforma. 22, qzae052 (2024).
Wang, B. et al. WTAP/IGF2BP3 mediated m6A modification of the EGR1/PTEN axis regulates the malignant phenotypes of endometrial cancer stem cells. J. Exp. Clin. Cancer Res. 43, 204 (2024).
Zhang, T. et al. m(6)A mRNA modification maintains colonic epithelial cell homeostasis via NF-kappaB-mediated antiapoptotic pathway. Sci. Adv. 8, eabl5723 (2022).
Ponraj, A. et al. A multi-patch-based deep learning model with VGG19 for breast cancer classifications in the pathology images. Digit Health 11, 20552076241313161 (2025).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
An, S. et al. Integrative network analysis identifies cell-specific trans regulators of m6A. Nucleic Acids Res. 48, 1715–1729 (2020).
Acknowledgements
This work was supported by the Guangxi Young Elite Scientist Sponsorship Program (GXYESS2025013), Bagui Outstanding Young Talents Program of Guangxi to Sanqi An, National Natural Science Foundation of China (Grant Nos. 82160389, 8210389), Guangxi Medical University Training Program for Distinguished Young Scholars to Sanqi An, Specific Research Project of Guangxi for Research Bases and Talents, China (Grant no. 2022AC19006), and the Basic Scientific Research Project of the Guangxi Academy of Agricultural Sciences (GNK2025YP135).
Author information
Authors and Affiliations
Contributions
S.A., K.M., H.L., and X.L. conceived and supervised the project. Y.L. and Xinyue Xu developed the Scm6A software and web server, with D.J. and M.C. contributing to key components. Y.L., X.X., D.J., and J.M. performed data analysis and validation. Y.B. and Y.C. performed website feature testing and user feedback optimization. Y.C. and C.H.C. contributed to data curation and technical validation. S.Y.L. and Y.C. provided computational support. Y.C. and J.L. assisted in methodology refinement. S.A. and Y.L. wrote the original manuscript, with significant contributions from Xinyue Xu, and critical revisions from all authors. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Junyao Jiang, Clarence Mah, and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Aylin Bircan and Christina Karlsson Rosenthal. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Xu, X., Chen, M. et al. Systematic evaluation of tools used for single-cell m6A identification. Commun Biol 8, 1841 (2025). https://doi.org/10.1038/s42003-025-09246-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09246-7