Abstract
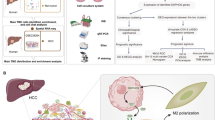
Increased acetyl-coenzyme A (acetyl-CoA) generation facilitates cancer metastasis and represents a critical metabolic characteristic of metastatic cancers. To maintain high acetyl-CoA levels, cancer cells often enhance the uptake of acetate for acetyl-CoA biosynthesis. However, the microenvironmental source of acetate remains largely unknown. Here we demonstrate that acetate is secreted by tumour-associated macrophages (TAMs) and taken up by hepatocellular carcinoma (HCC) cells to support acetate accumulation. Mechanistically, HCC cell-derived lactate activates the lipid peroxidation–aldehyde dehydrogenase 2 (ALDH2) pathway in TAMs, which promotes the TAMs’ acetate production and secretion. Inhibition of ALDH2 or of lipid peroxidation in TAMs abrogates acetate-induced migration of HCC cells in vitro. In an orthotopic HCC model involving male mice, genetic ablation of ALDH2 in TAMs reduces HCC cell acetate levels and HCC lung metastases. Collectively, our findings reveal a metabolic interaction between HCC cells and TAMs—involving lactate, lipid peroxidation and acetate—and position TAMs as an acetate reservoir that drives HCC metastasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw numerical source data are published alongside the paper; the other data that support the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Bergers, G. & Fendt, S. M. The metabolism of cancer cells during metastasis. Nat. Rev. Cancer 21, 162–180 (2021).
Gerstberger, S., Jiang, Q. W. & Ganesh, K. Metastasis. Cell 186, 1564–1579 (2023).
Pietrocola, F., Galluzzi, L., Bravo-San Pedro, J. M., Madeo, F. & Kroemer, G. Acetyl coenzyme A: a central metabolite and second messenger. Cell Metab. 21, 805–821 (2015).
Rios Garcia, M. et al. Acetyl-CoA carboxylase 1-dependent protein acetylation controls breast cancer metastasis and recurrence. Cell Metab. 26, 842–855.e5 (2017).
Lee, J. V. et al. Acetyl-CoA promotes glioblastoma cell adhesion and migration through Ca2+–NFAT signaling. Genes Dev. 32, 497–511 (2018).
Lu, M. et al. ACOT12-dependent alteration of acetyl-CoA drives hepatocellular carcinoma metastasis by epigenetic induction of epithelial–mesenchymal transition. Cell Metab. 29, 886–900.e5 (2019).
Mashimo, T. et al. Acetate is a bioenergetic substrate for human glioblastoma and brain metastases. Cell 159, 1603–1614 (2014).
Gao, X. et al. Acetate functions as an epigenetic metabolite to promote lipid synthesis under hypoxia. Nat. Commun. 7, 11960 (2016).
Park, J. W. et al. A prospective evaluation of 18F-FDG and 11C-acetate PET/CT for detection of primary and metastatic hepatocellular carcinoma. J. Nucl. Med. 49, 1912–1921 (2008).
Tsuchida, T., Takeuchi, H., Okazawa, H., Tsujikawa, T. & Fujibayashi, Y. Grading of brain glioma with 1-11C-acetate PET: comparison with 18F-FDG PET. Nucl. Med. Biol. 35, 171–176 (2008).
Grassi, I. et al. The clinical use of PET with 11C-acetate. Am. J. Nucl. Med. Mol. Imaging 2, 33–47 (2012).
Schug, Z. T., Vande Voorde, J. & Gottlieb, E. The metabolic fate of acetate in cancer. Nat. Rev. Cancer 16, 708–717 (2016).
Ayala, A., Munoz, M. F. & Arguelles, S. Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell. Longev. 2014, 360438 (2014).
Wang, J. et al. Acetate reprogrammes tumour metabolism and promotes PD-L1 expression and immune evasion by upregulating c-Myc. Nat. Metab. 6, 914–932 (2024).
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell 41, 374–403 (2023).
Elia, I. & Haigis, M. C. Metabolites and the tumour microenvironment: from cellular mechanisms to systemic metabolism. Nat. Metab. 3, 21–32 (2021).
Robinson, A., Han, C. Z., Glass, C. K. & Pollard, J. W. Monocyte regulation in homeostasis and malignancy. Trends Immunol. 42, 104–119 (2021).
Zhu, X. D. et al. High expression of macrophage colony-stimulating factor in peritumoral liver tissue is associated with poor survival after curative resection of hepatocellular carcinoma. J. Clin. Oncol. 26, 2707–2716 (2008).
Liu, D. et al. LSECtin on tumor-associated macrophages enhances breast cancer stemness via interaction with its receptor BTN3A3. Cell Res. 29, 365–378 (2019).
Wei, C. et al. Crosstalk between cancer cells and tumor associated macrophages is required for mesenchymal circulating tumor cell-mediated colorectal cancer metastasis. Mol. Cancer 18, 64 (2019).
Christofides, A. et al. The complex role of tumor-infiltrating macrophages. Nat. Immunol. 23, 1148–1156 (2022).
Mehla, K. & Singh, P. K. Metabolic regulation of macrophage polarization in cancer. Trends Cancer 5, 822–834 (2019).
Zhang, X., Ji, L. & Li, M. O. Control of tumor-associated macrophage responses by nutrient acquisition and metabolism. Immunity 56, 14–31 (2023).
Kao, K. C., Vilbois, S., Tsai, C. H. & Ho, P. C. Metabolic communication in the tumour-immune microenvironment. Nat. Cell Biol. 24, 1574–1583 (2022).
Zakhari, S. Overview: How is alcohol metabolized by the body? Alcohol Res. Health. 29, 245–254 (2006).
Chen, C. H., Sun, L. & Mochly-Rosen, D. Mitochondrial aldehyde dehydrogenase and cardiac diseases. Cardiovascular Res. 88, 51–57 (2010).
Su, P. et al. Enhanced lipid accumulation and metabolism are required for the differentiation and activation of tumor-associated macrophages. Cancer Res. 80, 1438–1450 (2020).
Masetti, M. et al. Lipid-loaded tumor-associated macrophages sustain tumor growth and invasiveness in prostate cancer. J. Exp. Med. 219, e20210564 (2022).
Isseroff, R. R., Ziboh, V. A., Chapkin, R. S. & Martinez, D. T. Conversion of linoleic-acid into arachidonic-acid by cultured murine and human keratinocytes. J. Lipid Res. 28, 1342–1349 (1987).
Salem, N., Pawlosky, R., Wegher, B. & Hibbeln, J. In vivo conversion of linoleic acid to arachidonic acid in human adults. Prostag. Leukotr. Ess. 60, 407–410 (1999).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Kagan, V. E. et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat. Chem. Biol. 13, 81–90 (2017).
Singh, S. et al. Aldehyde dehydrogenases in cellular responses to oxidative/electrophilic stress. Free Radic. Biol. Med. 56, 89–101 (2013).
Jeon, J. Y. et al. Regulation of acetate utilization by monocarboxylate transporter 1 (MCT1) in hepatocellular carcinoma (HCC). Oncol. Res. 26, 71–81 (2018).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Endale, H. T., Tesfaye, W. & Mengstie, T. A. ROS induced lipid peroxidation and their role in ferroptosis. Front. Cell Dev. Biol. 11, 1226044 (2023).
Tauffenberger, A., Fiumelli, H., Almustafa, S. & Magistretti, P. J. Lactate and pyruvate promote oxidative stress resistance through hormetic ROS signaling. Cell Death Dis. 10, 653 (2019).
Hayes, J. D. & Dinkova-Kostova, A. T. The Nrf2 regulatory network provides an interface between redox and intermediary metabolism. Trends Biochem. Sci. 39, 199–218 (2014).
Liu, Y. et al. Benzyl isothiocyanate ameliorates acetaldehyde-induced cytotoxicity by enhancing aldehyde dehydrogenase activity in murine hepatoma Hepa1c1c7 cells. Food Chem. Toxicol. 108, 305–313 (2017).
Kitakaze, T., Inoue, M. & Ashida, H. Aged garlic extract prevents alcohol-induced cytotoxicity through induction of aldehyde dehydrogenase 2 in the liver of mice. Mol. Nutr. Food Res. 67, e2200627 (2023).
Murthy, D. et al. Cancer-associated fibroblast-derived acetate promotes pancreatic cancer development by altering polyamine metabolism via the ACSS2–SP1–SAT1 axis. Nat. Cell Biol. 26, 613–627 (2024).
Qiu, J. et al. Acetate promotes T cell effector function during glucose restriction. Cell Rep. 27, 2063–2074.e5 (2019).
Wu, L. et al. An invasive zone in human liver cancer identified by Stereo-seq promotes hepatocyte–tumor cell crosstalk, local immunosuppression and tumor progression. Cell Res. 33, 585–603 (2023).
El-Kenawi, A. et al. Macrophage-derived cholesterol contributes to therapeutic resistance in prostate cancer. Cancer Res. 81, 5477–5490 (2021).
Kloosterman, D. J. et al. Macrophage-mediated myelin recycling fuels brain cancer malignancy. Cell 187, 5336–5356.e30 (2024).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Rabinowitz, J. D. & Enerbäck, S. Lactate: the ugly duckling of energy metabolism. Nat. Metab. 2, 566–571 (2020).
Xu, Q. et al. NADPH oxidases are essential for macrophage differentiation. J. Biol. Chem. 291, 20030–20041 (2016).
Acknowledgements
We thank Z.-S. Zhou, L.-F. Yang, D. Ye, Y. Chen, J. Qin and H.-Y. Yin for their helpful discussions on this project. We thank L. Qiu, Z. Li, J.-Y. Wu and Y.-F. Bu from the Institutional Center for Shared Technologies and Facilities of the Shanghai Institute of Nutrition and Health, Chinese Academy of Sciences, for technical assistance. This work was supported by grants from the National Key R&D Program of China (no. 2022YFA1303000 to M.L.), the National Natural Science Foundation of China (no. 82173132 to M.L., no. 81930074 and no. 91959203 to L.X.Q.; no. 82072696 and no. 82472951 to W.W.Z.).
Author information
Authors and Affiliations
Contributions
M.L. conceived the concept of the study. M.L., L.Q. and W.Z. designed the study. S.W. and C.G. performed the early exploration. L.S., S.F., X.L., Y.B. and Y.C. performed experiments. S.W., C.G., L.S., J.P., H.X. and Y.Z. collected and analysed human HCC samples. Q.L., L.S., S.W. and X.Z. performed the liquid chromatography–mass spectrometry analysis. Q.L. and L.S. performed the stable isotope tracing assay. L.S. conducted the statistical analysis and generated the figures. M.L., L.Q. and W.Z. interpreted the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Evan Lien and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Acetate levels in HCC tissues and the flow cytometry gating strategy of TAMs.
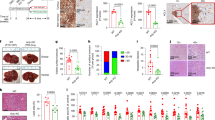
a, b, The levels of acetate in HCC tumor tissues (a) and tumor-derived interstitial fluid samples (b) in comparison to the corresponding adjacent normal liver. Data are shown as mean ± SEM (n = 7 mice per group) (a), or mean ± SEM (n = 5 mice per group) (b). HCC tumor tissues and normal liver tissues were from Hepa1-6 orthotopic HCC mice model. C57BL/6 J mice were orthotopically injected with Hepa1-6-ZsGreen cells. 14 days later, the mice were sacrificed, and the tumor tissues were used for isolation of primary TAMs, TANs and NCCs by flow cytometry. The flow cytometry gating strategy is shown in (c). TAMs were depleted in Hepa1-6-ZsGreen orthotopic HCC mice model by intraperitoneal injection of clodronate liposomes every 2 days for three times. The depletion efficiencies were then determined by measuring the infiltration of CD45+Ly6G−F4/80+CD11bint cells by flow cytometry (d), and primary Hepa1-6-ZsGreen cells were isolated from tumor tissues by flow cytometry. The flow cytometry gating strategy is shown in (e). Representative images result from at least three independent experiments are shown. P values were determined by unpaired two-tailed Student’s t-tests (a, b).
Extended Data Fig. 2 Validation of induction of TAMs and CAFs and TAMs-induced acetate accumulation in other types of cancer cells.
a, The mRNA levels of marker genes in TAMs generated from THP-1 cell-derived MΦs. b, The mRNA levels of CAFs marker genes in LX2 cell-derived CAFs. c, d, The mRNA levels of marker genes in TAMs generated from RAW264.7 cells or from BMDM. e–g, The acH3 and acH3K9 levels in breast cancer cells (MDA231 and AT3), colorectal cancer cells (HCT116 and MC38) and pancreatic cancer cells (PANC-1 and PanC02) after treatment with TAMs-CM. Data are represented as mean ± s.d. of triplicate experiments (a–d), images are representative of three independent experiments (e–g). P values were determined by unpaired two-tailed Student’s t-tests (a–d).
Extended Data Fig. 3 TAMs does not affect the expression of genes involved in acetate metabolism in HCC cells.
a, b, The mRNA levels of genes involved in acetate metabolism in MHCC-97H cells and Hepa1-6 cells after treatment with TAMs-CM, respectively. Data are shown as mean ± s.d. of triplicate experiments. P values were determined by unpaired two-tailed Student’s t-tests.
Extended Data Fig. 4 Lipid peroxidation-ALDH2 pathway is responsible for acetate production in TAMs.
a, b, The protein levels of ALDH2 and ALDH1B1 in TAMs generated from THP-1 cell-derived MΦs or RAW264.7 cells. c, d, Confirmation of ALDH2 or ALDH1B1 knockdown in THP-1 cells stably expressing with shRNA targeting ALDH2 (shALDH2) or ALDH1B1 (shALDH1B1). e, Immunoblots of ALDH2 protein in BMDM or liver isolated from Aldh2-flox or Aldh2-cKO mice. f, g, Confirmation of Aldh2 or Aldh1b1 knockdown in RAW264.7 cells stably expressing shRNA targeting Aldh2 (shAldh2) or Aldh1b1 (shAldh1b1). h, i, Secreted acetate levels in TAMs with silenced Aldh2 or Aldh1b1 (h) or in TAMs treated with ALDH2 inhibitor (i). j, k, Effects of silencing of ALDH2 on the mRNA levels of IL10 and ARG1 in TAMs and corresponding MΦs. l–q, MDA levels (l, m), Cox1 and Cox2 mRNA levels (n, o) and the lipid peroxidation levels (p, q) in TAMs and corresponding MΦs. Quantitation of the lipid peroxidation levels are shown in q. r, s, Effects of Fer-1 (2 μM) or NAC (5 mM) on the lipid peroxidation levels in TAMs (r) and primary TAMs (s). t, Effects of Aspirin (10 mM) on the MDA generation in TAMs. u–z, As depicted in u and v, TAMs generated from RAW264.7 cells were incubated with exogenous [U-13C]-LA for 36 h with or without etomoxir or NAC, and were subjected to determine labeled carbons in LA (w), AA (x) and secreted acetate (y). The proportion of labelled secreted acetate to labelled AA is shown in z. Scale bar, 20 μm (p, r and s). Data are shown as three biologically independent cell cultures (h, i, l, m, t, w and x), four biologically independent cell cultures (y, z), mean ± s.d. of triplicate experiments (j, k, n and o), or mean ± s.d. of 5 representative pictures (q), images are representative of three independent experiments (a–g, q–s). P values were determined by one-way ANOVA with Tukey’s multiple comparisons test (h, i and t) or unpaired two-tailed Student’s t-tests (j–o and q).
Extended Data Fig. 5 Silencing of ALDH2 expression abrogates acetate-induced migration of HCC cells.
a, b, The effect of ALDH2 silencing on pro-migration capacity of TAMs. Hepa1-6 cells were co-cultured with TAMs generated from RAW264.7 cells. Then, cell migration was determined in Hepa1-6 cells (a), and the relative migrated cell numbers are expressed as the fold change to controls (b). c, The effect of ALDH2 silencing in TAMs on growth of Huh7 cells. Huh7 cells were co-cultured with TAMs generated from THP-1 cell-derived MΦs for 24 h. d–g, The effect of ALDH2 silencing on pro-migration capacity of TAMs. Huh7 cells were incubated with TAMs-CM or TAMs-MP generated from THP-1 cell-derived MΦs, Hepa1-6 cells were incubated with TAMs-CM or TAMs-MP generated from RAW264.7 cells. Then, cell migration was determined in these HCC cells (d, f), and the relative migrated cell numbers are expressed as the fold change to controls (e, g). h, i, The effect of ALDH2 inhibitor on the pro-migration capacity of TAMs on Hepa1-6 cells. Hepa1-6 cells were incubated with CM from CVT-10216 (50 nM)-treated TAMs with or without exogenous acetate, the migration of Hepa1-6 cells were then examined (h). The relative migrated cell numbers are expressed as the fold change to controls (i). j, k, The effects of inhibition of lipid peroxidation on pro-migration capacity of TAMs generated from THP-1 cell-derived MΦs. TAMs-CM were prepared by using TAMs treated with the Fer-1 (2 μM), NAC (5 mM) or Aspirin (10 mM), and the migration of Huh7 cells treated with these TAMs-CM were examined (j). The relative migrated cell numbers are expressed as the fold change to controls (k). Scale bar, 100 μm (a, d, f, h, j). Data are shown as mean ± s.d. of 10 (e, g, i and k) or 8 representative pictures (b) from three biologically independent cell cultures, or five biologically independent cell cultures (c). P values were determined by one-way ANOVA with Tukey’s multiple comparisons test (b, e, g, i and k) or two-way ANOVA with Holm-Sidak’s multiple comparisons test (c). ns, not significant.
Extended Data Fig. 6 Silencing of ALDH2 expression does not lead to the accumulation of oxidized lipids and reactive aldehydes in TAMs.
a–h, The effects of ALDH2 silencing on the levels of intracellular ROS (a), lipid peroxidation (b), intracellular MDA and 4-HNE (c, d), secreted MDA, 4-HNE (e, f) and oxidized lipids (g, h) in TAMs generated from THP-1 cell-derived MΦs. Intracellular ROS levels were measured with DCFH-DA, and the relative ROS levels are expressed as the fold change to controls (a). Quantitation of the lipid peroxidation levels are shown in right of b. Oxidized lipids were determined by LC-MS analysis (g, h). i, The effects of ALDH2 silencing on the expressions of other ALDH family genes in TAMs. Scale bar, 20 μm (b). Data are shown as mean ± s.d. of seven biologically independent cell cultures (a), six biologically independent cell cultures (h), 5 representative pictures from three biologically independent cell cultures (right of b), five biologically independent cell cultures (c, e), four biologically independent cell cultures (d, f, g), or triplicate experiments (i). P values were determined by unpaired two-tailed Student’s t-tests. ns, not significant.
Extended Data Fig. 7 TAMs-CM promotes HCC cells migration by increasing acetyl-CoA levels.
a, b, Confirmation of ACSS2 knockdown (shACSS2) and ACSS1 expression in Huh7 cells. c, The acH3 levels of Huh7 shNT cells and Huh7 shACSS2 cells with or without exogenous acetate. d, Confirmation of ACSS1 and ACSS2 knockdown (shACSS1/2) in MHCC-97H cells. e–g, The effect of TAMs-MP treatment on acH3 levels (e) and migration (f, g) of MHCC-97H shNT cells and MHCC-97H shACSS1/2 cells. The relative migrated cell numbers are expressed as the fold change to controls (g). h, i, The effect of MCT1 inhibitor, AZD3965, on the migration of Huh7 cells with or without exogenous acetate. The migration of Huh7 cells were examined (h). The relative migrated cell numbers are expressed as the fold change to controls (i). Scale bar, 100 μm (f, h). Data are shown as mean ± s.d. of 9 (g), or 10 representative pictures (i) from three biologically independent cell cultures, images are representative of three independent experiments (a–e). P values were determined by one-way ANOVA with Tukey’s multiple comparisons test (g, i).
Extended Data Fig. 8 TAMs-derived acetate promotes HCC cells migration by increasing acH3 levels.
a, b, Huh7 cells were treated with CM from shNT TAMs or shALDH2 TAMs and p300 acetyltransferase inhibitor (C646, 5 μM), and the migration (a, b) or acH3 levels (c) of Huh7 cells were examined. The relative migrated cell numbers are expressed as the fold change to controls (b). d–f, Huh7 cells were treated with CM from shNT TAMs or shALDH2 TAMs and Trichostatin A (TSA, 1 μM), and the migration (d, e) or acH3 levels (f) of Huh7 cells were examined. The relative migrated cell numbers are expressed as the fold change to controls (e). g–i, The effect of knockdown of different KATs on migration and acH3 levels. Huh7 shNT cells, Huh7 shKAT2A cells and Huh7 shKAT5 cells were treated with CM from shNT TAMs or shALDH2 TAMs, and the migration (g, h) or acH3 levels (i) of Huh7 cells were examined. The relative migrated cell numbers are expressed as the fold change to controls (h). j, Immunoblots of TWIST and N-cadherin, and E-cadherin in Huh7 cells treated with CM from shNT TAMs or shALDH2 TAMs. Scale bar, 100 μm (a, d and g). Data are shown as mean ± s.d. of 10 representative pictures from three biologically independent cell cultures (b, e and h), images are representative of three independent experiments (c, f, i and j). P values were determined by one-way ANOVA with Tukey’s multiple comparisons test (b, e and h).
Extended Data Fig. 9 Genetic ablation of ALDH2 in TAMs does not affect the immune cells infiltration, and ACSS1/ACSS2 knockdown limits HCC metastasis.
a–h, Flow cytometry analysis of the infiltration of immune cells in primary tumor tissues from Hepa1-6-ZsGreen orthotopic HCC mice model in Aldh2-flox and Aldh2-cKO mice. Data are represented as mean ± SEM (n = 7 mice per group). i, j, Representative mIHC images using human HCC samples showing tumor areas with high infiltration of ALDH2+ CD163+ TAMs. Yellow and white arrows indicate HCC cells and ALDH2+ CD163+ TAMs respectively (Bottom photos in i, j). Scale bars, 3 mm (the left images), 20 μm (the remaining pictures). k–r, Balb/c-nu/nu mice were orthotopically inoculated with Huh7 shNT cells, Huh7 shACSS2 cells, MHCC-97H shNT cells or MHCC-97H shACSS1/2 cells. At the end point (day 21), the primary tumors were harvested (k, o), tumor volumes were calculated (l, p), spontaneous lung metastases were detected by ex vivo bioluminescence imaging, and the endpoint bioluminescence images (m, q) and the quantification of photon flux of lungs (n, r) are shown. Data are represented as mean ± s.e.m. (n = 7 mice per group) (l, n, p, r). P values were determined by unpaired two-tailed t-test (a–h, l, n, p, r). ns, not significant.
Extended Data Fig. 10 HCC cells-derived lactate induces lipid peroxidation-ALDH2 pathway in TAMs.
a, The lactate concentrations in TCM from MHCC-97H cells and Hepa1-6 cells. b, The mRNA levels of ALDH2 in RAW264.7 cells after treatment with lactate (10 mM). c–e, The effect of lactate on the MDA levels (c) and the lipid peroxidation levels (d) in RAW264.7 cells. Quantitation of the lipid peroxidation levels are shown in e. Scale bar, 20 μm. f, The levels of ROS in RAW264.7 cells after treatment with lactate. Intracellular ROS levels were measured with DCFH-DA. g, h, The mRNA levels of ALDH2 (g) and the levels of MDA (h) were determined in RAW264.7 cells after treatment with lactate and NAC (5 mM). i, j, The mRNA levels of NOX1 and NOX2 in THP-1 cell-derived MΦs (i) or RAW264.7 cells (j) after treatment with lactate. Data are shown as mean ± s.d. of triplicate experiments (b, g, i, j), mean ± s.d. of three biologically independent cell cultures (a, c, h), mean ± s.d. of six biologically independent cell cultures (f), or mean ± s.d. of 5 representative pictures from three biologically independent cell cultures (e). P values were determined by one-way ANOVA with Tukey’s multiple comparisons test (g, h) or unpaired two-tailed Student’s t-tests (b, c, e, f, i, j). ns, not significant.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3 and Supplementary Tables 1–4.
Supplementary Data (download XLSX )
Source data for Supplementary Figs. 1–3.
Source data
Source Data Figs. 1–7, Extended Data Figs. 1–10 (download XLSX )
Statistical source data.
Source Data Figs. 2, 5 and 7 and Extended Data Figs. 2, 4, 7 and 8 (download PDF )
Unprocessed western blots and/or gels and statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, L., Wang, S., Gao, C. et al. Tumour-associated macrophages serve as an acetate reservoir to drive hepatocellular carcinoma metastasis. Nat Metab 7, 2268–2283 (2025). https://doi.org/10.1038/s42255-025-01393-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01393-9
This article is cited by
-
Tumor-associated macrophages in cancer: from mechanisms to application
Molecular Biomedicine (2025)