Abstract
Brown adipose tissue (BAT) contributes to thermoregulation and glucose metabolism, but how these functions are coordinated remains unclear. While thermogenesis in the activated BAT typically coincides with increased blood flow and glucose uptake1,2,3,4,5, several pathophysiological and nutritional states dissociate these processes6,7, suggesting they are governed by distinct sympathetic circuits. Here we identify subpopulations of sympathetic neurons in the stellate ganglion that mediate distinct functions of intrascapular BAT (iBAT) in mice. Two main types of sympathetic neurons project to iBAT: those that innervate the organ parenchyma and those that innervate the large blood vessels feeding the depot8,9,10,11,12. Here we develop a toolkit to parse the functions of these neuronal subclasses through targeted chemogenetic activation of projections to iBAT, while sparing other organs, and single-cell transcriptomics coupled to retrograde tracing from iBAT to the stellate ganglion. We find that stimulation of the parenchymal projections increases blood flow and thermogenesis in iBAT, without affecting circulating glucose levels. Conversely, stimulation of the vascular projections improves glucose tolerance but does not alter blood flow or thermogenesis in iBAT. These data provide a mechanistic explanation for the dissociation between the thermogenic and glycaemic effects of BAT activation13,14,15,16.
Similar content being viewed by others
Main
Cold-induced thermogenesis in brown adipose tissue (BAT) is an energy-demanding process that relies on enhanced blood flow as well as uptake of fuels, processes that are all triggered by sympathetic nervous system (SNS) signalling17. Thus, these parameters are often considered interchangeably when assessing the physiological consequences of stimulating BAT in humans. However, glucose uptake in BAT does not require cold-induced thermogenesis in humans18 or rodents13,14,15, and the metabolically beneficial effects of glucose uptake in BAT are dependent on adrenergic signalling but are independent of cold ambient temperature16. Conversely, diabetes in humans and mouse models is associated with impaired glucose uptake in BAT19,20, while its metabolic activity is normal6. These observations raise the possibility that thermogenic and glucoregulatory functions of BAT are controlled by distinct sympathetic circuits, although the exact neuronal substrates are unknown.
Cold responses in the skin are mediated through the coordinated regulation of functionally distinct populations of sympathetic neurons defined by the expression of different combinations of neurotransmitters and neuropeptides21,22. Neurons that co-express norepinephrine and neuropeptide Y (NPY) project to the vasculature and promote vasoconstriction23,24, while norepinephrine neurons with low levels of NPY promote piloerection (goosebumps)25. This neuroanatomical organization of sympathetic fibres is preserved in BAT8,9,10,11,12. Gain and loss of Npy expression in the SNS impacts both cold tolerance and glucose metabolism11,12,26,27. Since NPY is also expressed in neurons that innervate visceral organs11,28, the function of sympathetic neurons that innervate intrascapular BAT (iBAT) per se cannot be evaluated in these genetic models. Here, we developed novel mouse genetic tools and in vivo assays to parse the functions of molecularly and neuroanatomically distinct sympathetic projections to iBAT, without affecting sympathetic projections to other visceral organs or sensory projections to iBAT that also affect metabolism.
Retrograde adeno-associated viruses (AAVrg) are routinely used to mark and manipulate neural projections to a target of interest within the brain29. We assessed the efficacy of retrograde labelling from iBAT to its largest source of sympathetic innervation, the stellate ganglion (SG)30,31. Despite using viral titres tenfold higher than commonly used in the brain (>1013 vg ml−1), SG labelling following intra-iBAT injections of AAVrg::CAG-GFP was much less efficient than tracing with cholera toxin subunit B (CTB; Extended Data Fig. 1a,b). Intersectional approaches involving direct viral injections into the nodose ganglion can effectively target sensory projections to adipose tissues32. However, this approach is not feasible in the SG owing to its physical inaccessibility. Thus, the dearth of tools to robustly modulate SG projections to peripheral organs in the thoracic cavity is a major obstacle to studying their impact on physiology.
Intra-iBAT injections of the MaCPNS1 serotype, which was engineered for use in the periphery33, achieved robust labelling in the SG but also labelled sensory projections from the dorsal root ganglia (DRG) (Fig.1a,b and Extended Data Fig. 1c). Injecting a Cre-dependent MaCPNS1 virus into iBAT of Th-Cre mice robustly labelled sympathetic projections to iBAT (SGBAT) from the SG through T4, with a few neurons labelled between T5 and T7 (Extended Data Fig. 1e). Since iBAT-projecting neurons are distinct from the subpopulation of tyrosine hydroxylase-expressing neurons in the DRG (Extended Data Fig. 2), this strategy avoided labelling sensory neurons that could provide negative feedback to the SNS32 (Fig. 1c and Extended Data Fig. 1d).
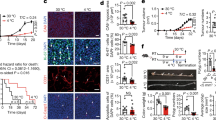
a, A schematic illustration of retrograde labelling to sympathetic (yellow) and/or sensory (blue) ganglia with AAV injections into iBAT. b, Injections of GFP-expressing MaCPNS1 virus into iBAT of WT mice labelled neurons in the SG and DRG at C4, C6 and T1. Scale bar, 200 µm. c, Injections of Cre-dependent mCherry-expressing MaCPNS1 into iBAT of Th-Cre mice labelled neurons in the SG but not DRGs. Scale bar, 200 µm. d,e, Representative traces of thermogenesis (ΔiBAT − lower back (LB), top) and blood flow (bottom) in iBAT in a Th-Cre mouse injected with a control Cre-dependent mCherry-expressing MaCPNS1 virus (d) or with a Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus (e). f,g, Change in temperature (f) and blood flow (g) in iBAT following SAL or CNO injection (n = 16 mice, tested in both conditions, P < 0.001). h, A schematic illustration of the random crossover experimental design to assess the impact of DCZ versus SAL treatment in awake Th-Cre mice that received Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT (data shown in i–m). i, The average traces of energy expenditure (EE). The solid lines represent means, shaded areas represent ± s.e.m. (n = 8 mice). j,k, Changes in EE (j) and RER (k) following SAL or DCZ injection (n = 16 mice each tested in both conditions, P < 0.001). l, The effect of DCZ or SAL delivered immediately before an oral glucose tolerance test (OGTT) on circulating glucose levels. Data presented as the mean ± s.e.m. (n = 19 mice, tested in both conditions, P = 0.0016 at 15 min). m, Within-subject effects (DCZ minus SAL) on blood glucose levels from the same mice shown in l. n, A schematic illustration of the random crossover experimental design to assess the impact of DCZ versus SAL treatment in awake Th-Cre mice that received Cre-dependent hM4Di–mCherry-expressing MaCPNS1 virus in iBAT (data shown in o and p). o, The effect of DCZ or SAL delivered immediately before a glucose gavage on circulating glucose levels. Data presented as the mean ± s.e.m. (n = 5 mice, P = 0.04271 at 15 min). p, Within-subject effects (DCZ minus SAL) on blood glucose levels from the same mice shown in o. In f, g, j and k, the dots represent average values and lines represent individual mice. Paired t-test, two tailed, no adjustments. **P < 0.01 and ***P < 0.001. Panel a created with BioRender.com.
We used this approach to investigate the physiological outcomes of acutely activating SGBAT neurons with Cre-dependent MaCPNS1 viruses expressing either mCherry (control) or an excitatory designer receptor exclusively activated by designer drugs (DREADD, hM3Dq; Extended Data Fig. 1f). Three weeks after the viral injection, we assessed the effect of intraperitoneal (i.p.) injections of saline (SAL) and the DREADD agonist clozapine-N-oxide (CNO, 10 mg kg−1) on thermogenesis and blood flow in iBAT in mice under constant anaesthesia. We used the difference between the temperature of iBAT and the lower back as an estimate of iBAT thermogenesis. CNO injections into hM3D-expressing mice increased iBAT thermogenesis and blood flow, without effects on core body temperature, while mice injected with the control virus did not respond (Fig. 1d,e). Within 5 min of CNO treatment, iBAT temperature increased by ~0.8 °C and blood flow increased by ~25% (Fig. 1f,g). The number of neurons expressing the DREADD virus in this cohort varied from 2,283 to 3,595, but there was no correlation with changes in thermogenesis or blood flow (Extended Data Fig. 3). Thus, when >2,000 neurons are infected, the effect is largely binary. The number of labelled SG neurons and the magnitude of responses were similar in males and females.
Delivery of the β-adrenergic receptor antagonist propranolol rapidly reversed the effects of chemogenetic stimulation on thermogenesis and blood flow in both sexes, demonstrating the requirement for SNS signals (Fig. 1e). Treatment with the α-adrenergic receptor antagonist phentolamine acutely reduced blood flow by 60 ± 24.3% (mean ± s.e.m.) but did not affect thermogenesis (Extended Data Fig. 4). While it is possible that increased blood flow contributes to elevated iBAT temperature, the two processes can be decoupled. The experiments presented here do not distinguish between UCP1-dependent and UCP1-independent thermogenic mechanisms.
Next, we explored the effects of chemogenetic manipulation of SGBAT neurons on systemic metabolism in freely moving, awake mice. To minimize possible off-target effects of CNO, we switched to a higher-affinity DREADD ligand, deschloroclozapine6 (DCZ, 200 µg kg−1). Since activated BAT in humans is associated with improved glucose homeostasis16 and β3-adrenergic stimulation improves glucose tolerance34,35, we assessed the effect SAL versus DCZ injections immediately before an oral gavage of glucose (2 g kg−1 at time 0) in a random crossover design (Fig. 1h). DCZ treatment lowered blood glucose levels after 15 min in male and female mice expressing a Th-Cre-driven excitatory DREADD (Fig. 1l,m) but had no effect in wild type (WT) mice (Extended Data Fig. 5a–c). The glucose-lowering effect of chemogenetic stimulation was not observed in the absence of oral glucose gavage (Extended Data Fig. 6a–c), raising the possibility that SNS projections to iBAT are endogenously activated by glucose. To test this, we performed complementary intra-iBAT injections of a MaCPNS1 virus expressing a Cre-dependent inhibitory DREADD (hM4Di) in Th-Cre mice. Chemogenetic inhibition impaired glucose tolerance at 30 min after gavage in both sexes (Fig. 1n–p).
Mice expressing the excitatory DREADD were transferred to metabolic cages where they were acclimated to i.p. SAL injections for 5 days before receiving either SAL or DCZ in a random crossover design. While the acute injection stress caused an increase in energy expenditure in all mice, this effect was significantly greater in DCZ-treated mice in both sexes (Fig. 1i,j). DCZ and CNO did not affect energy expenditure when injected into WT mice (Extended Data Fig. 5d,e). Increased energy expenditure following chemogenetic stimulation was not accompanied by a compensatory increase in food intake (Supplementary Fig. 1). The average respiratory exchange ratio (RER) in the 30 min post-injection decreased with DCZ but not SAL (Fig. 1k), but some of this effect may be attributable to DCZ alone, as it also lowers RER in WT mice (Extended Data Fig. 5f).
To further investigate the effects of acute chemogenetic stimulation, we performed bulk RNA-sequencing and lipidomics in iBAT in a separate cohort of female and male mice 30 min after SAL or DCZ treatment. We identified 87 (56 upregulated and 31 downregulated) differentially expressed genes with DCZ treatment after controlling for sex (Fig. 2a). Gene set enrichment analysis (GSEA) identified ‘cell activation’ as the top enriched term in upregulated genes (Fig. 2b). This category includes immediate early genes, such as Jun and Egr3, and genes encoding downstream components of adrenergic signalling (that is, Adrb2) and regulators of thermogenesis, metabolism and angiogenesis (Irf4, Irs2 and Thbs1)36,37,38. Notably, Ucp1 expression was not increased at this timepoint. Other enriched GSEA terms are consistent with effects on metabolism in iBAT. Many of the most upregulated and downregulated genes are implicated in lipid homeostasis. Genes encoding proteins that promote lipid biosynthesis (for example Fasn and Acsl5) and lipid transport and signalling (Ffar4/Gpr120, Trim67 and Fabp5) were downregulated, while those that suppress it were upregulated (for example, Sik1 and Dusp1). Together, these results indicate that acute stimulation of SNS projections to iBAT regulates thermogenesis and fuel disposal at the local (tissue) level. In addition, we observed marked sex differences in the transcriptional profile of iBAT, independent of chemogenetic stimulation (Fig. 2c). Females preferentially expressed genes related to amino acid catabolism, glycolysis and triglyceride (TG) biosynthesis (Fig. 2d), whereas males preferentially expressed genes related to lipid and glutathione catabolism (Fig. 2e). The idea that sex differences exist in the metabolic properties of iBAT was also reflected in how stimulating SGBAT neuronal impacts the lipid profiles of both plasma (Fig. 2f,g,j,k) and iBAT (Fig. 2h,i,l,m). Chemogenetic stimulation of SGBAT neurons reduced circulating TG species to a similar degree in females and males (43.1% and 46.4%, respectively) (compare Fig. 2f,g with Fig. 2j,k). Within iBAT of females, chemogenetic stimulation increased the abundance of TGs, although this did not reach significance (Fig. 2i). However, a number of specific lipid species such as TG 58:6-FA20:4 or TG 56:4-FA18:1 that were found significantly lower in plasma accumulate in iBAT of females, suggesting that chemogenetic stimulation mediates uptake and storage of TG species. In males, however, lipids did not accumulate in iBAT (Fig. 2l,m), which is consistent with the idea that the activated iBAT of males is primed to metabolize the TGs taken from the circulation. Uncovering sex differences in the metabolism of different macronutrients that underlie these results is an important area for future research.
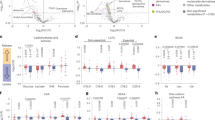
a, A heat map of significantly differentially expressed genes between DCZ- and SAL-treated anaesthetized Th-Cre mice that received Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT, controlling for sex (F, female; M, male). b, Statistically significant GO terms from GSEA of genes from a. c, A heat map of significantly differentially expressed genes between males and females from a, controlling for treatment. d, Selected GO terms from GO-ORA of upregulated genes in males from c. e, Selected GO terms from GO-ORA of upregulated genes in females from c. f,g, A volcano plot of lipid species (f) and lipid class concentrations (g) in plasma of female mice from a. Data presented as mean ± s.e.m. (n = 5 SAL-treated and n = 4 DCZ-treated mice, P = 0.0113 for TG). h,i, A volcano plot of lipid species (h) and lipid class concentrations (i) in iBAT of female mice from a. Data presented as mean ± s.e.m. (n = 5 SAL-treated and n = 4 DCZ-treated mice). j,k, A volcano plot of lipid species (j) and lipid class concentrations (k) in plasma of male mice from a. Data presented as mean ± s.e.m. (n = 3 SAL-treated and n = 4 DCZ-treated mice, P = 0.0011). l,m, A volcano plot of lipid species (l) and lipid class concentrations (m) in iBAT of male mice from a. Data presented as mean ± s.e.m. (n = 3 SAL-treated and n = 4 DCZ-treated mice). Dots represent individual lipid species. Paired t-test, two tailed, no adjustment. *P < 0.05 and **P < 0.01. CE, cholesterol ester; Cer d18:1, ceramide on a sphingosine backbone; DG, diacylglycerol; FFA, free fatty acid; HexCER, hexosylceramide; LPC, lysophosphatidylcholine; LPE, lysophosphatidylethanolamine; PA, phosphatidic acid; PC, phosphatidylcholine; PE, phosphatidylethanolamine; PG, phosphatidylglycerol; PI, phosphatidylinositol; PS, phosphatidylserine; SM, sphingomyelin.
To explore the molecular identity of post-ganglionic sympathetic neurons innervating iBAT, we performed single-cell RNA-sequencing (scRNA-seq) coupled with retrograde tracing from distinct peripheral targets in the mouse SG. We focused on SG neurons innervating iBAT (SGBAT) and forelimb (SGFL), because they are both involved in thermoregulation and the projections share a similar anatomical trajectory via the dorsal rami. We performed simultaneous retrograde tracing from iBAT and forelimb by injecting CTB conjugated to Alexa555 (CTB–A555) into iBAT and CTB–A488 into the forelimbs. Three days later, we generated z-stacked images of intact cleared ganglia. SGBAT and SGFL neurons were intermingled in the lateral SG but were not overlapping (Fig. 3a).
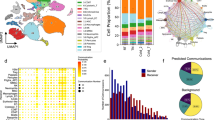
a, Retrograde labelling of SG neurons that project to iBAT (red) and forelimb (green) with cholera toxin B. Scale bar, 100 µm. b, A heat map of the expression of cluster-enriched genes in cells sorted by cluster. c, Violin plots showing the expression of sympathetic and cluster-specific marker genes. d, Validation of cluster marker genes in the SG using smFISH in n = 4 mice. The SG1 markers assessed were Npy, Fst, Sctr and Rfxp1. The markers for other SG populations were Slc18a1 (SG2), Colq (SG2–4) and Aqp1 (SG3). Scale bar, 20 µm. e,f, Single-cell transcriptomes visualized with t-distributed stochastic neighbour embedding (t-SNE), coloured by cluster (e) and by projection target (f). g, The percentage of cells of each SG subtype within cells retrogradely labelled from iBAT (red), forelimb (green) or other targets (grey, non-labelled cells). h, Whole-tissue imaging of iBAT from a Th-Cre::tdTomato mouse injected with IL-647. Scale bar, 100 µm. i, Whole-tissue imaging of iBAT from an Rxfp1-Cre::tdTomato mouse injected with IL-647. Scale bar, 100 µm. j,k, mCherry and NPY immunofluorescence staining in the SG of Rxfp1-Cre (j) and Vmat1-Cre (k) mice injected with MaCPNS1:CAG-DIO-mCherry in iBAT. Scale bars, 100 µm. l, A schematic illustrating the breeding strategy to obtain Rxfp1-Flp::tdTOM; Vmat1-Cre mice (left) and representative images of tdTomato, NPY and EGFP immunofluorescence staining in the SG of these mice injected with AAVrg:CAG-DIO-EGFP in iBAT (right). Scale bar, 100 µm Panel l created with BioRender.com.
Next, we performed scRNA-seq in conjunction with CTB retrograde tracing from iBAT and forelimb (Extended Data Fig. 7). We collected a total of 1,039 single cells by fluorescence-activated cell sorting (FACS) (433 SGBAT (A555+) and 220 SGFL (A488+) and 375 non-labelled neurons) from three independent sorts of n = 12 SG in 8-week-old mice. The A555+ and A488+ cells were distinct, confirming that SGBAT and SGFL neurons do not overlap (Extended Data Fig. 7d). Based on an unsupervised analysis of single-cell transcriptomic signatures, we divided SG neurons into four clusters, SG1 to SG4 (SG1–4; Fig. 3b,e). Projecting the CTB labelling onto the unsupervised clustering revealed that each cluster contains neurons projecting to iBAT (A555+, red), forelimb (A488+, green) and other organs such as the heart (grey) (Fig. 3f,g). However, >85% of SNS inputs to iBAT derived from the SG1 and SG2 subtypes, with most of the remaining innervation coming from SG3 (Fig. 3g).
Neurons in all clusters expressed Th, Dbh and Slc18a2/Vmat2, consistent with a sympathetic neuronal identity39, but expression was highest in SG1 and SG2 neurons (Fig. 3c). Similarly, Npy was expressed in all clusters, but SG1 neurons expressed it at levels that were 7–50-fold higher than the other clusters (Fig. 3c). While NPY immunoreactivity has been used to mark SG1 neurons (Fig. 3d), its low expression levels in SG2–4 disqualifies its use as the basis for a Cre driver. SG2 neurons expressed Slc18a1/Vmat1 in addition to Slc18a2/Vmat2. This was unexpected as VMAT1 expression is thought to be restricted to neuroendocrine cells in the adrenal gland and gut. SG4 neurons expressed the cholinergic markers Chat and Slc18a3/VAChT (Fig. 3c). We confirmed the specificity of the expression patterns of the marker genes identified in the bioinformatic analyses by multiplex single-molecule fluorescent in situ hybridization (smFISH) on SG tissue sections (Fig. 3d). We confirmed that the SG1 markers Fst, Sctr and Rfxp1 were co-expressed with Npy, but not with Slc18a1 (SG2), Colq (G2–4) or Aqp1 (SG3). Nearly all Slc18a1+ neurons (SG2) expressed Colq (SG2–4), but little to no Aqp1 (SG3). The transcriptional profiles within each cluster were shared by SGBAT, SGFL and unlabelled neurons. However, the proportion of each of the four subclusters differed according to the projection target. The most notable difference was in the cholinergic sudomotor neurons (SG4), which preferentially projected to forelimb, consistent with the paws serving as the major site of sweat release in rodents.
Next, we explored why the unique markers of SG subpopulations identified here were not revealed by earlier analyses. A major difference between the approaches is our exclusive focus on the SG, while earlier studies pooled sympathetic ganglia from SG through T13 (refs. 40,41). We evaluated the expression of the markers for SG1–3 in ganglia at more rostral (superior cervical ganglion; SCG) and caudal (celiac ganglion) positions in the chain. We performed multiplex smFISH in Npy-GFP reporter mice to map the expression of SG markers genes relative to vasoconstrictor neurons in the SCG previously reported by Lundberg et al. and Pernow et al.23,24. These Npy-GFPHI neurons co-expressed the SG1 markers Rxfp1, Fst and Sctr, and not markers for SG2–4 neurons such as Cckar, Colq or Aqp1 (Extended Data Fig. 8a). Based on these results, we conclude that the gene expression pattern in the SCG parallels our observations in the SG.
Gene expression in celiac ganglion, which innervates the gastrointestinal tract and liver, diverged substantially from the pattern observed ganglia of the rostral sympathetic chain (SCG and SG). Npy-GFPHI neurons co-expressed the SG1 marker Rfxp1, but not the other SG1 markers such as Fst and Sctr (Extended Data Fig. 8b). In addition, the SG2–3 markers Cckar, Colq and Aqp1 were preferentially expressed in Npy-GFPHI neurons in the celiac ganglion (Extended Data Fig. 8b), consistent with prior reports that Npy is broadly expressed by sympathetic neurons in the celiac ganglion11. The only SG markers that maintained the same pattern of relative expression in Npy-GFPHI neurons in the SCG, SG and celiac ganglion were Rfxp1 (SG1) and Slc18a1 (SG2). The differences in the combinations of genes expressed by subpopulations of neurons in the rostral and caudal sympathetic chain explain why pooled sympathetic ganglia analyses across the entire chain did not identify subpopulation-specific markers.
To selectively manipulate SG1/Rxfp1-Cre and SG2/Vmat1-Cre populations, we generated Rxfp1-Cre and Vmat1-Cre lines and validated them using a variety of approaches. We crossed Rxfp1-Cre, Vmat1-Cre and Th-Cre lines to the tdTomato reporter strain (Ai9) and validated the expression of the Tomato lineage trace in the SG by performing smFISH with Rxfp1 and Tomato probes and by visualizing the pattern of fibre projections in the intact iBAT depot. The Rxfp1-Cre lineage trace exhibited the expected pattern: reporter expression confined to Rxfp1 neurons in the SG but absent in Colq-expressing neurons (SG2–4)(Extended Data Fig. 9a–c). We optimized the Fast 3D Clear system42 to visualize axonal projections in iBAT from Th-Cre::tdTOM and Rxfp1-Cre::tdTOM mice that were retroorbitally injected with isolectin (IL)–A647 to mark the vasculature. Pan-sympathetic fibres labelled with the Th-Cre lineage trace were detected around large blood vessels and in the parenchyma (Fig. 3h and Supplementary Videos 1–3). Fibres labelled with the Rxfp1-Cre lineage trace were confined to blood vessels with a diameter of ~10–11 µm (Fig. 3i, Supplementary Videos 4–6 and Extended Data Fig. 10a), consistent with reports that NPY+ fibres exclusively project to arterioles in iBAT12. To quantify this relationship, we calculated the average distance between each 647-nm-positive pixel (blood vessels) and its nearest 550-nm-positive pixel (nerves) and compared the proportional distribution of nerve vessel distances. Rxfp1-Cre::tdTOM fibres had a higher proportion of contacts in close proximity to blood vessels (0.41–2 µm) than the pan-sympathetic Th-Cre::tdTOM fibres, which were the predominant fibres found at greater distances (>5 µm) from vessels (Extended Data Fig. 10c). This further supports our observation that Rxfp1-Cre::tdTOM projections are preferentially associated with blood vessels. It should be noted that Rxfp1-Cre::tdTOM fibres are a subset of the Th-Cre::tdTOM fibres.
When Vmat1-Cre mice were crossed to the Ai9 reporter strain, Tomato was detected in all iBAT-projecting neurons in the SG, including those that expressed Rxfp1 (Extended Data Fig. 9d–f) owing to transient Vmat1 expression in the embryonic peripheral nervous system43,44. To overcome this limitation, we injected Cre-dependent mCherry retrograde viruses into iBAT of Rxfp1-Cre or Vmat1-Cre strains in adult mice and assessed reporter expression in conjunction with NPY immunostaining in the SG and in the DRG. In Rxfp1-Cre mice, most of the neurons expressing the viral mCherry reporter co-expressed NPY (Fig. 3j). NPY+ neurons that did not express mCherry presumably represent SG1 neurons that project to other targets, such as the forelimbs. By contrast, in Vmat1-Cre mice, little overlap between the mCherry reporter and NPY was detected (Fig. 3k). The viral reporter was not detected in the DRGs (Extended Data Fig. 1g,h), confirming the specificity of the SNS targeting.
To further confirm that Rxfp1- and Vmat1-based driver lines target distinct subpopulations in the adult SG, we generated an Rxfp1-Flp driver line and crossed it to Flp-dependent Tomato reporter and Vmat1-Cre lines. The resulting Rxfp1-Flp::FSF-Tomato;Vmat1-Cre mice were injected with a Cre-dependent EGFP-expressing AAVrg in iBAT (Fig. 3l). Notably, most tdTomato+ SG neurons co-expressed NPY, while EGFP+ SG2 neurons did not express NPY or tdTomato (Fig. 3l), demonstrating the existence of two molecularly distinct neuronal populations in the SG projecting to iBAT. Given that Rxfp1+ and Vmat1+ neurons account for >85% of sympathetic innervation of iBAT (Fig. 3g), it is likely that the parenchyma-projecting fibres observed in Th-Cre::tdTOM mice originate from Vmat1+ neurons. However, we cannot exclude the possibility that Vmat1+ neurons also project to blood vessels.
We compared the functional consequences of acute selective stimulation of either Rxfp1+ (vasomotor) or Vmat1+ (parenchymal) SGBAT neurons using the chemogenetic tools and paradigms described above (Fig. 4a,e). Of note, the specific stimulation of SG2/Vmat1+ neurons innervating brown adipocytes in the parenchyma induced thermogenesis (Fig. 4b,c), while activation of SG1/Rxfp1+ neurons projecting exclusively to the blood vessels did not (Fig. 4f,g). Contrary to the dogma that NPY+ neurons are vasoregulators8,23,24, blood flow was only increased by stimulating parenchymal SG2 projections (Fig. 4b,d,f,h). Increases in iBAT thermogenesis and blood flow by stimulating parenchymal SG2 projections to iBAT (Fig. 4c,d) were comparable to those achieved by stimulating all iBAT-projecting neurons in Th-Cre mice (Fig. 1f,g).
a–d, The effects of chemogenetic activation of parenchyma-projecting SG2 neurons in anaesthetized mice: the experimental strategy to record from Vmat1-Cre mice injected with Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT (a); average traces of thermogenesis (ΔiBAT − LB, top) and blood flow (bottom) in iBAT recorded from mice injected with SAL and CNO (n = 14 mice) (b); and changes in temperature (c) and in blood flow (d) in iBAT following SAL or CNO treatment in mice shown in b. P < 0.001 for both measurements. e–h, The effects of chemogenetic activation of blood vessel-projecting SG1 neurons in anaesthetized mice: the experimental strategy to record from Rxfp1-Cre mice injected with Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT (e); average traces of thermogenesis (ΔiBAT − LB, top) and blood flow (bottom) in iBAT recorded from mice injected with SAL and CNO (n = 10 mice) (f); and changes in temperature (g) and in blood flow (h) in iBAT following SAL or CNO in mice shown in f. i–l, The effects of chemogenetic activation of parenchyma-projecting SG2 neurons in awake mice: a schematic of the random crossover experimental design to assess the impact of DCZ versus SAL treatment in awake Vmat1-Cre mice that received Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT (i); average traces of EE (n = 6 mice) (j); and changes in EE (k) and RER (l) following SAL or DCZ injection (n = 9, P = 0.04591 for k and P = 0.0031 for l). m–p, The effects of chemogenetic activation of arteriole-projecting SG1 neurons in awake mice: a schematic of the random crossover experimental design to assess the impact of DCZ versus SAL treatment in awake Rxfp1-Cre mice that received Cre-dependent hM3D–mCherry-expressing MaCPNS1 virus in iBAT (m); the average traces of EE (n = 8 mice) (n); and changes in EE (o) and RER (p) following SAL or DCZ injection in mice shown in n (P = 0.0188 for p). q,r, The effects of chemogenetic activation of parenchyma-projecting SG2 neurons in awake mice. The effect of DCZ or SAL delivered immediately before a glucose gavage on circulating glucose levels (q) and within-subject effects (DCZ minus SAL) on blood glucose levels (r) from the same mice shown in q. Data presented as the mean ± s.e.m. (n = 6 mice, tested in both conditions). s,t, Effects of chemogenetic activation of arteriole-projecting SG1 neurons in awake mice. The effect of DCZ or SAL delivered immediately before a glucose gavage on circulating glucose levels (s) and within-subject effects (DCZ minus SAL) on blood glucose levels (t) from the same mice shown in s (n = 8, tested in both conditions, P = 0.01377 at 15 min). u–w, The effects of chemogenetic inhibition of arteriole-projecting SG1 neurons in awake mice: a schematic of the random crossover experimental design to assess the impact of DCZ versus SAL treatment in awake Rxfp1-Cre mice that received Cre-dependent hM4Di–mCherry-expressing MaCPNS1 virus in iBAT (u); the effect of DCZ or SAL delivered immediately before a glucose gavage on circulating glucose levels (v) (n = 6 mice, tested in both conditions, P = 0.02997 at 15 min and P = 0.04776 at 30 min); and within-subject effects (DCZ minus SAL) on blood glucose levels (w) from the same mice shown in v. In b, f, j and n, the lines represent the means and shaded areas represent ± s.e.m. In c, d, g, h, k, l, o and p, dots represent average values and lines represent individual mice. Paired t-test, two tailed *P < 0.05, **P < 0.01 and ***P < 0.001. NS, not significant.
Next, we examined the impact of stimulating each subpopulation on systemic metabolism in awake animals (Fig. 4i,m). Consistent with observations in anaesthetized mice, activation of parenchyma-projecting SG2/Vmat1+ neurons resulted in higher average energy expenditure across the study in both sexes (Fig. 4j,k), whereas stimulation of vasomotor SG1/Rxfp1+ neurons did not (Fig. 4n,o). RER was decreased in both Cre lines (Fig. 4l,p). The magnitude of the response from SG2/Vmat1+ stimulation (−0.038 ± 0.006, mean ± s.e.m.) was similar to pan-sympathetic stimulation in Th-Cre mice (−0.045 ± 0.007; Fig. 1k), and the RER response to SG1/Rxfp1+ stimulation (−0.022 ± 0.006) was comparable to that in WT mice (−0.028 ± 0.007; Extended Data Fig. 5f). However, we cannot definitively parse the circuits regulating fat oxidation.
In the context of an oral glucose challenge, chemogenetic activation of vasomotor SG1/Rxfp1+ projections to iBAT produced a marked reduction in blood glucose levels at 15 min (Fig. 4s,t), but stimulation without glucose gavage had no effect on baseline glucose levels (Extended Data Fig. 6d–f). Chemogenetic inhibition resulted in increased circulating glucose at 30 and 45 min (Fig. 4u–w), fully recapitulating the effects observed when all iBAT-projecting sympathetic neurons were activated or inhibited in Th-Cre mice (Fig. 1l–p). Notably, stimulating parenchyma-projecting SG2/Vmat1+ neurons had no effect on glycaemia (Fig. 4q,r and Extended Data Fig. 6g–i). In summary, we demonstrated that thermogenesis and blood flow in iBAT are regulated by parenchyma-projecting SG2/Vmat1+ neurons, while systemic glucose is modulated by vasomotor SG1/Rxfp1+ neurons.
By combining retrograde injections with Cre-dependent viral constructs, we targeted manipulations to subpopulations of BAT-projecting sympathetic neurons without confounding effects on the brain, sensory nervous system or other peripheral organs. Acute stimulation of SG2 projections to the iBAT parenchyma is sufficient to increase blood flow and heat production in iBAT and to promote fat utilization, whereas effects on total energy expenditure are modest and variable, consistent with observations in humans45. Our findings do not exclude the possibility that sympathetic stimulation of white adipose tissue lipolysis is needed to fully support energy combustion by iBAT20,46.
NPY co-transmission in the SNS has been implicated in regulating a wide range of physiological functions. In contrast to the dogma that NPYergic sympathetic projections are vasoconstrictors23,24, chemogenetic activation of SG1 projections did not affect blood flow or thermogenesis in iBAT. Since SG1 neurons in the SCG and SG are molecularly distinct from NPYergic neurons in the celiac ganglion, exploring whether other NPYergic neurons in the rostral sympathetic chain share projection targets and functionality is an important area for future research. Another important question raised by our results is whether activation of SG1 and SG2 neurons is sensed by molecularly and functionally distinct populations of DRG neurons that innervate iBAT, as shown for cutaneous-projecting DRG neurons47. For example, it is possible that subpopulations of thermosensors provide negative feedback to prevent overheating in response to activation of SG2 neurons, but not SG1 neurons.
Our data support the idea that effects of SNS-derived NPY on total energy expenditure, cold tolerance and adiposity11,12 reflect direct actions in other tissues and/or chronic adaptations in the context of a congenital knockout driven by a Th-Cre reporter. Since this driver line is also expressed in the brain and DRG, it is possible that central and sensory circuits contribute to these effects. On the other hand, some of the effects of SNS-derived NPY on glucose tolerance that were attributed to actions in the pancreas, liver and white adipose tissue could stem from signalling in iBAT11. Consistent with this idea, glucose uptake in BAT is associated with lower glycaemia and risk of diabetes in humans, independent of external temperature16. As chemogenetic modulation of SG1/Rxfp1+ neurons alters glycaemia in the context of a glucose load but not at baseline, it is possible that crosstalk with other metabolic organs is involved.
The ability to target SG1 projections to iBAT without affecting thermogenesis may be particularly beneficial in the context of diabetes, which is associated with impaired glucose uptake but normal thermogenesis in BAT6,48. In the long term, uncovering SNS targets that mediate the metabolically beneficial effects on glycaemia without activating thermogenic programmes that are subject to robust negative feedback could lead to novel approaches to treat metabolic diseases.
Methods
Ethics statement
All procedures were performed within the guidelines of the Institutional Animal Care and Use Committee at the Columbia University Health Science Division. All efforts were made to minimize animal suffering and to use the minimum number of animals required for statistical validity.
Animals
Mice were housed at 22 ± 1 °C on a 12 h light:12 h dark cycle in the Russ Berrie Medical Sciences Pavilion barrier facility with ad libitum access to chow (13.2% calories from fat; 5053, PicoLab Rodent Diet 20) and autoclaved water. C57BL/6J (000664), Ai949 (007909), Flp-dependent Tomato50 (032864) and NPY-GFP51 (006417) strains were purchased from the Jackson Laboratories, Th-Cre (B6.129×1-Tht1(cre)Te/Kieg) from the European Mouse Mutant Archives (RRID: IMSR_EM:00254) and Rxfp1-Cre (B6.Cg-Rxfp1tm1(cre)Zel), Rxfp1-Flp (B6.Cg-Rxfp1tm1(cre)Zel) and Vmat1-Cre (B6.Cg-Slc18a1tm1(cre)Zel) strains were generated by CRISPR-mediated T2A-nlsCre insertion at the Genetically Modified Mouse Model Shared Resource at Columbia University Irving Medical Center. Consistent with findings from other groups31, the Jackson Th-Cre line (008601) drove minimal Ai9 recombination. Mice receiving viral injections were single housed post-surgery.
Retrograde tracing with cholera toxin B
Under isoflurane anaesthesia, the upper back was shaved and sterilized with alternating betadine and 70% isopropyl alcohol. Buprenorphine (0.1 mg kg−1) was given for analgesia. Mice were placed on a heating pad under a sterile drape and a 1 cm incision was made over the interscapular region. The upper white adipose layer was dissected to expose iBAT, and CTB–Alexa Fluor 555 (Thermo Fisher, C22843; 1 μl of 1% in SAL) was injected at two sites per depot using a glass-pulled needle and mouth pipette. The fat pad was replaced and the incision closed with 4-0 PS-4 sutures (Ethilon, 1662G). CTB–Alexa Fluor 488 (C22841) was similarly injected at three sites per forelimb. Wounds were cleaned and mice monitored for distress. After 72 h, stellate ganglia were collected following euthanasia with Avertin (500 mg kg−1; Fisher, AC421430100), dissociated for analysis or post-fixed in 4% paraformaldehyde (PFA) for 1 h, washed and mounted in LIMPID52.
Virus injection
Mice were anaesthetized and prepared as described above. A horizontal incision was made ~2 cm below the iBAT to avoid scar interference with Doppler imaging. The fat pad was exposed, separated along the midline and flipped. Virus was injected at three to five evenly spaced sites per depot using a 25 µl Neuros syringe (Hamilton, 65460-11). Each mouse received a total dose of 1.3 × 10¹² gc, with the injection volume (20–50 µl) adjusted to viral titre (Supplementary Tables). The fat pad was repositioned, and the incision closed and sanitized. At the end of the study, SGs and DRGs were collected, post-fixed in 4% PFA for 1 h, washed and mounted in LIMPID.
Imaging of isolated ganglia from virus-injected mice
SGs and DRGs were incubated overnight at room temperature in the dark and imaged on a Zeiss LSM 710 confocal microscope (20×, tiled z stacks, 5 µm steps). Fluorescent reporter-expressing neurons were quantified using Stereo Investigator Cleared Tissue Edition software (MBF Bioscience).
Temperature and blood flow recordings
Mice were anaesthetized with Avertin (350 mg kg−1, i.p.) and the interscapular area shaved. Animals were maintained at 35 °C on a heating pad with rectal temperature feedback (Harvard Apparatus, 75-1500). A perfusion needle connected to a syringe pump (Medfusion 3500, ICUMedical) delivered Avertin continuously (190 mg kg−1 h−1) via the lateral vein to maintain anaesthesia. A small incision was made over the previous surgical site to expose iBAT and the fat pad was cut caudally to insert an implantable thermal probe (IT-21, Physitemp). A second probe was placed subcutaneously on the lower back and a third measured ambient temperature. Probes were connected to a thermometer (TC-2000, Sable Systems), and signals digitized via LabJack U3-LV for acquisition on a laptop. Blood flow over the interscapular region was recorded using a high-resolution laser Doppler imager (MOORLDI2-HIR, Moor Instruments). Baseline measurements were acquired for 10 min, followed by i.p. injections of SAL, CNO (10 µg g−1; Sigma, SML2304) and, when indicated, propranolol (5 µg g−1; Thermo Fisher, H26645.06) or phentolamine (20 µg g−1; Millipore-Sigma, P7561). Norepinephrine (0.5 mg kg−1) served as a positive control. All injections were volume-matched to 5 µl g−1. After recordings, mice were euthanized by anaesthetic overdose and ganglia were collected as described above.
Doppler data were processed with Moor LDI Review v6.0. The interscapular region was defined as the area showing maximal norepinephrine-induced activity, and this region of interest was applied to all frames to extract mean perfusion values. Thermal data were analysed in R using a custom script. For each injection, we computed the stepwise temperature difference between iBAT and lower back probes. Baseline (2 min pre-injection) and treated (5–10 min post-injection) averages were calculated and the difference (treated − baseline) was used for SAL, CNO and adrenergic blocker conditions. The same procedure was applied to relative blood flow values.
Oral glucose tolerance tests
Mice were transferred to clean cages without food for 4 h before testing. Baseline blood glucose was measured from tail vein pricks (Contour Next EZ, Ascensia). Mice then received an oral gavage of d-(+)-glucose (2 mg kg−1; Sigma, G8769, 1:2 in ddH2O, 20 µm filtered) and an i.p. injection of either DCZ (200 ng g−1) or SAL. Gavage and injection volumes were 9 and 5 µl g−1, respectively. Blood glucose was recorded at 15, 30, 45, 60, 90 and 120 min after gavage. Each mouse received DCZ and SAL in a balanced crossover design, with a 7-day washout between sessions.
Indirect calorimetry
Mice were single housed in a Promethion Core Metabolic System (Sable Systems) for 1 week under controlled ambient temperature (22 ± 1 °C), a 12 h light:12 h dark cycle and with ad libitum access to food and water from the same barrier facility. Mice were acclimated to the metabolic cages and daily i.p. SAL injections at 16:00 for 3 days. On days 4 and 5, mice received two i.p. SAL injections per day (10:00 and 16:00). During the final 2 days, mice received two daily injections of either DCZ (200 ng g−1) or SAL in a balanced crossover design. Specifically, on day 6, half of the mice received DCZ and half received SAL (10:00 and 16:00), and treatment groups were reversed on day 7. All injections were volume-matched to 5 µl g−1 body weight. Mice were returned to their home cages after each recording session.
Energy expenditure (heat) and RER (CO2 produced/O2 consumed) were quantified. For experiments comparing DCZ and CNO in WT mice, animals were acclimated for 2 days with a single daily i.p. SAL injection, followed by two daily SAL injections on days 3 and 4. Over the next 3 days, mice received two daily injections of either SAL, DCZ or CNO in a randomized and counterbalanced order to minimize carryover effects.
Statistical analyses were performed by comparing average values before and after injections for each mouse and parameter. For each parameter, the mean value during the hour following injection (excluding the first 9 min to account for gas equilibration and stress-induced hyperthermia) was calculated and the equivalent pre-injection mean was subtracted. Differences between groups were analysed using paired t-tests.
Bulk RNA sequencing
Three weeks after intra-iBAT viral injections of Th-Cre+/− mice (six males and six females) with MaCPNS1:CAG-DIO-hM3D-mCherry as described above, mice were anaesthetized with Avertin (350 mg kg−1, i.p.) and injected intraperitoneally with either DCZ (200 ng g−1, i.p.) or SAL. Mice were maintained on a heating pad at 35 °C for 30 min before euthanasia via thoracotomy. Transcardial perfusion with 5 ml of cold PBS (4 °C) was performed to remove residual blood, after which iBAT and SG were collected. Each iBAT was bisected along the midline, weighed and snap-frozen in liquid nitrogen. One lobe per mouse was used for RNA extraction. SG were post-fixed and washed as previously described, mounted in LIMPID mounting medium and imaged to confirm viral expression.
Total RNA was extracted from iBAT using the Tri-Isolate RNA Pure Kit (IBI Scientific, IB47631) following the manufacturer’s instructions. RNA integrity was assessed using RIN analysis, and sequencing was performed by the Genome Center Core at Columbia University Irving Medical Center. mRNA was enriched using poly(A) selection and sequencing libraries were prepared with Illumina TruSeq chemistry. The final PCR step was modified using the KAPA HiFi HotStart Ready Mix. Libraries were sequenced on an Element AVITI platform to generate paired-end 75 bp reads. Raw BCL files were converted to FASTQ format and adaptor-trimmed using bases2fastq (v1.4.0.833301531). Reads were pseudoaligned to a kallisto index generated from the Ensembl v96 mouse transcriptome (GRCm38.p6) using kallisto (v0.44.0).Gene count matrices were analysed using R53 with DESeq54, while GSEA and Gene Ontology (GO) over-representation analysis (ORA) were done with clusterProfiler55.
Lipidomics
Shotgun lipidomic analysis of murine plasma and BAT samples was performed as previously described56. Briefly, 25 µl of plasma or 3–5 mg of BAT samples were spiked with internal standards and mixed with MTBE/methanol/water 10:3:2.5 (v/v/v) as described57. Following homogenization, samples were centrifuged (4 °C, 10,000g, 10 min) and the lipid-containing upper phase was transferred into a new vial. Lipid extracts were concentrated by vacuum centrifugation and reconstituted in a mixture of dichloromethane (50): methanol (50)-containing 10 mM ammonium acetate. Direct infusion of lipid extracts into the mass spectrometer was conducted by an ultrahigh-pressure liquid chromatography system (Nexera X2, Shimadzu) using dichloromethane (50): methanol (50)-containing 10 mM ammonium acetate as eluent at a flowrate of 0.008 ml min−1. The separation and targeted profiling of lipid species was performed combining differential mobility spectrometry and a QTRAP system (QTRAP 5500, SCIEX) run in multiple reaction monitoring mode operated via Analyst (v1.6.8, SCIEX). The mass spectrometry-generated raw data were converted to mzML format using MSconvertGUI (v3.0.21245-5724be1). Data processing and lipid quantification were performed using the Shotgun Lipidomic Assistant software, a Python-based application according to ref. 56. Values are given as nmol ml−1 (plasma) or nmol mg−1 (BAT).
Dissociation of SG neurons
SG were dissociated following modified published protocols58,59. SGs were collected bilaterally from two 6–8-week-old C57BL/6J males (four SG per batch) in ice-chilled Eagle’s balanced salt solution equilibrated to 95% O2/5% CO2 for 1 h. After removing fat and connective tissue, SGs were minced into three to four pieces and transferred to low-bind tubes containing 1.67 ml pre-heated (37 °C) digestion solution for 1.5 h with agitation. The digestion mix contained papain (1,034 µl), collagenase/dispase (200 µl, 20 mg ml−1), d-trehalose (167 µl, 50%), AP-V (3 µl, 25 mM), kynurenic acid (13 µl, 100 mM) and DNase I (250 µl).
Digestion-stop solution consisted of 50% d-trehalose (1,050 µl), AP-V (11 µl, 25 mM), kynurenic acid (44 µl, 100 mM), DNase I (250 µl) and FBS (250 µl). The medium solution contained 50% d-trehalose (1,050 µl), AP-V (8 µl, 25 mM), kynurenic acid (19 µl, 100 mM) and FBS (107 µl) in 9.45 ml DMEM/F12.
Following digestion, half of the suspension was transferred to a new tube, mixed with digestion-stop solution, inverted and centrifuged (300g, 4 °C, 5 min). Pellets were resuspended in 500 µl digestion-stop solution, combined and triturated with BSA-coated fire-polished pipettes of decreasing diameter (400 µm to 150 µm). The cell suspension was washed twice with 1 ml medium, centrifuged (300g, 5 min) and resuspended in 200 µl medium. Cells were filtered through a 40 µm strainer, stained with Sytox Blue (1:1,000) and kept on ice before FACS. A detailed description of this protocol can be found on protocols.io (https://doi.org/10.17504/protocols.io.14egn79e6v5d/v1).
Cell sorting
Single cells were sorted into three to four 96-well plates using a FACSAria sorter equipped with a 130-µm nozzle (12 PSI). Approximately two-thirds of wells were filled with CTB–A555 or CTB–A488-positive neurons, and the remaining third with non-labelled neurons. Plates were centrifuged (200g, 2 min) and snap-frozen on dry ice. Three sorting sessions yielded a total of 1,039 cells across 11 plates from 6 mice.
Plate-based scRNA-seq of SG neurons
ScRNA-seq was conducted by the Single Cell Analysis Core at Columbia University Irving Medical Center as previously described60. Briefly, libraries were prepared using template-switching reverse transcription with adaptor-linked oligo(dT) primers containing cell- and molecule-specific barcodes. cDNA from each plate was pooled for PCR amplification and library construction, and 3′-end libraries were sequenced on an Illumina NextSeq 500/550.
Analysis of scRNA-seq datasets
Reads were aligned to the mouse genome (GRCm38, Gencode M13, Ensembl 88) using STAR v2.5.3a61 and assigned to genes with featureCounts v1.5.362. BAM files were sorted, duplicates removed and unique molecular identifier (UMI) count matrices generated with UMI-tools63, yielding 1,039 cells.
Downstream analysis was performed in Seurat v364. Quality control excluded cells with <1,000 genes, >10% mitochondrial content or no CTB dye assignment, retaining 879 cells. Expression matrices were log-normalized and scaled while regressing out batch, nUMI and mitochondrial fraction. Highly variable genes were identified, and dimensionality reduction was performed by principal component analysis. JackStraw identified 42 significant principal components, of which 30 were retained for clustering.
Cells were clustered using Seurat’s FindClusters function on a KNN graph, refined by Jaccard distance and modularity optimization. Cluster markers were identified using FindMarkers (Wilcoxon rank-sum test). To prevent glial gene interference, 136 genes expressed by satellite glial cells65 were removed from the highly variable set (3,555 genes retained). Reclustering across resolutions was visualized using clustree66, and a resolution of 0.6 was chosen. Cluster markers were defined as genes enriched in ≥40% of cells within each cluster. Expression was visualized via heat maps and violin plots. Analyses were conducted in R53.
Immunofluorescence staining and analysis
Mice injected with MaCPNS1 virus were perfused with PBS and 4% PFA at 4 °C. Vertebral columns were post-fixed overnight, washed in PBS and SGs and DRGs were extracted, embedded in OCT, frozen at −80 °C and sectioned at 10 µm.
Thawed sections were washed in PBST (0.1% Triton X-100) and circled with a hydrophobic pen. Slides were blocked in PBST + 5% normal donkey serum for 1 h at room temperature, then incubated overnight at 4 °C with primary antibodies (anti-NPY, anti-RFP, anti-GFP and anti-TH, 1:1,000 each). After washing, sections were incubated 1 h at room temperature with secondary antibodies and 4,6-diamidino-2-phenylindole (DAPI) (1:500), washed and mounted with ProLong Glass Antifade.
SGs were imaged on a Zeiss Axio Observer 7 (20×, tiled), DRGs on a Keyence BZ-8000 (20×, tiled). Neurons were manually counted in ImageJ70, using negative controls for background subtraction.
DRGs counts were imaged with a Keyence BZ-8000 fluorescent microscope at 20× as a tiled image. Neurons were manually counted using ImageJ67. Negative control slides (incubated without primary antibodies) were used for background subtraction.
Single-molecule in situ hybridization
SGs from WT mice and SCG/celiac ganglion from Npy-GFP mice were post-fixed in 4% PFA for 1 h, embedded in OCT, frozen at −80 °C, sectioned at 10 µm and mounted on slides. Multiplex smFISH was performed using the RNAscope Multiplex Fluorescent Reagent Kit following fixed-frozen tissue protocols. To preserve GFP fluorescence, a mild antigen retrieval was used (boiled buffer, 5 min at room temperature). After smFISH, sections were immunostained for GFP (Bio-Rad, 4745-1051, 1:250; Invitrogen, A-11015, 1:500) and imaged on a Zeiss LSM confocal microscope (40× and 63×).
Whole-tissue imaging in iBAT
Whole iBAT clearing was performed using a modified Fast 3D Clear protocol42. Mice were perfused with cold PBS followed by 4% PFA, and iBAT was post-fixed overnight at 4 °C. Tissues were washed in PBS (3× 2 h, 4 °C), incubated overnight in 25% Quadrol (Sigma, 122262) in PBS at 37 °C, and washed in double-distilled water. Samples were dehydrated in 50%, 70% and 90% tetrahydrofuran (Sigma, 186562) with pH adjusted to 9–9.5 using triethylamine, then rehydrated in reverse order. Clearing solution was prepared with 48 g Histodenz (Millipore-Sigma, D2158), 0.6 g of diatrizoic acid (Millipore-Sigma, D9268), 1 g of N-methyl-d-glucamine (Millipore-Sigma, M2004) and 10 g of ultrapure urea (Millipore-Sigma, 15505) in 13 ml water, incubated overnight at 37 °C. Samples were incubated in clearing solution overnight at 37 °C and imaged in the same medium on a Zeiss LSM 710 confocal microscope (20×).
Analysis of the distance between the nerves and blood vessels
Whole-tissue iBAT images were acquired as described above using a 2-µm z step. Each stack was divided into consecutive 20-µm substacks (10 images each) and converted into maximum intensity projections. Images were processed in ImageJ with custom Python scripts. For each channel (nerves and blood vessels), projections were converted to 8-bit greyscale, brightness/contrast were manually adjusted and binary masks were generated by manual thresholding.
A custom script calculated the Euclidean distance from each nerve pixel to the nearest vessel pixel, capped at 100 pixels, and converted to micrometres using microscope calibration (2.4089 pixels per µm). Distance distributions between genotypes were compared using the Kolmogorov–Smirnov test in R53.
iBAT innervated blood vessel diameter measurement
Whole-tissue images of iBAT were acquired as described above, using a 2-µm z step. Each image stack was visually inspected and split into two or more substacks to avoid overlapping between innervated blood vessels, then converted into maximum intensity projections. Blood vessels innervated by tdTomato-positive fibres were manually annotated in ImageJ and traced along their long axis. For each vessel, the diameter was measured manually at five evenly spaced points along the long axis. The average of these measurements was taken as the vessel diameter and converted from pixels to micrometres using the microscope calibration (2.4089 pixels per µm).
Whole-tissue staining and imaging of mouse trunk
Trunk samples were immunostained and cleared using a modified combination of PEGASOS68 and iDISCO69 protocols. Mice were transcardially perfused with ice-cold PBS (Thermo Fisher, 10010023) followed by 4% PFA (Electron Microscopy Sciences, 15710). Thoracic and lumbar vertebral columns were post-fixed overnight at 4 °C, washed in PBS and decalcified in 20% EDTA (Millipore-Sigma, E5134) for 4 days at 37 °C with daily solution changes. Samples were washed, incubated in 25% Quadrol (Sigma-Aldrich, 122262) for 48 h, washed again, treated with 3% ammonia (Sigma-Aldrich, 338818) for 6 h and washed overnight at 37 °C.
Tissues were delipidated and dehydrated through graded tert-butanol (tB, 30%, 50% and 70%; Sigma-Aldrich, 360538 with 3% quadrol; Sigma-Aldrich 122262) at 37 °C, rehydrated in reverse order and permeabilized and blocked for 48 h each as in iDISCO. Samples were incubated with rabbit anti-RFP (Rockland, 600-401-379, 1:10,000) for 6 days at 37 °C, washed for 24 h and then incubated with donkey anti-rabbit Alexa Fluor 568 (Invitrogen, A10042, 1:1,000) for 5 days at 37 °C, followed by a 24-h PBS wash. Samples were embedded in 1% agarose (Invitrogen, 16500500), dehydrated again in graded tert-butanol, incubated in tB-protected polyethylene glycol (tB-PEG, 75% tB + 22% PEG + 3% quadrol) for 48 h and cleared in benzyl benzoate PEG (BB-PEG, 75% tB + 22% PEG + 3% quadrol) for 48 h at 37 °C following the PEGASOS protocol. Imaging was performed with an Ultramicroscope II light-sheet microscope (Miltenyi Biotec) at 4× in BB-PEG clearing solution.
Statistical analyses and reproducibility
All statistical analysis were performed in R53 and Graphpad Prism v10.2.0 with the indicated statistical tests. Data distribution was assumed to be normal, but this was not formally tested. No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications32,70, and sample sizes for each experiment are reported in the figure legends. For metabolic data, the sample size was determined based on cage availability. For experiments involving viruses, the sample size was determined based on virus availability. No data were excluded, except mice with deteriorating health issues after surgery or during the experiment and mice with missed viral targeting assessed by histology. All in vivo experiments were performed at least twice or grouped from two independent cohorts with the same conclusion. Experiments were randomized across experimental days and daily blocks were chosen balancing sex and experimental conditions. The Investigators were not blinded to allocation during experiments and outcome assessment. For representative images, the numbers of experimental repetitions were as follows: Fig. 3a: over ten animals across six independent experiments; Fig. 3h,i: over six animals per genotype across four independent experiments; Fig. 3j–l: three animals per experiment; Extended Data Fig. 1a–h: over six animals per experiment, with the exception of Extended Data Fig. 1e, which was two animals per experiment; Extended Data Fig. 9b,c,e,f: over three animals per experiment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data were uploaded to the Gene Expression Omnibus (scRNA-seq of stellate ganglion: GSE310280; bulk RNA-seq of iBAT chemogenetics: GSE310279). Numerical source data and raw imaging files are published alongside this Letter, available via Figshare at https://doi.org/10.6084/m9.figshare.30491075 (ref. 71), temperature and Doppler recordings; https://doi.org/10.6084/m9.figshare.30524474 (ref. 72), lipidomics; and https://doi.org/10.6084/m9.figshare.30642500 (ref. 73), raw images and remaining source data. Source data are provided with this paper.
Code availability
Custom code for this Letter is available via GitHub at https://github.com/DanieleN90/Sympathetic-control-of-brown-fat.
References
Shimizu, Y., Nikami, H. & Saito, M. Sympathetic activation of glucose utilization in brown adipose tissue in rats. J. Biochem. 110, 688–692 (1991).
Engel, B. T., Sato, A. & Sato, Y. Responses of sympathetic nerves innervating blood vessels in interscapular, brown adipose tissue and skin during cold stimulation in anesthetized C57BL/6J mice. Jpn. J. Physiol. 42, 549–559 (1992).
Takahashi, A., Shimazu, T. & Maruyama, Y. Importance of sympathetic nerves for the stimulatory effect of cold exposure on glucose utilization in brown adipose tissue. Jpn. J. Physiol. 42, 653–664 (1992).
Soderlund, V., Larsson, S. A. & Jacobsson, H. Reduction of FDG uptake in brown adipose tissue in clinical patients by a single dose of propranolol. Eur. J. Nucl. Med. Mol. Imaging 34, 1018–1022 (2007).
Ulaner, G. A., Samstein, R., Cahlon, O., Weber, W. A. & Rimner, A. Unilateral suppression of brown fat on FDG PET/CT in Horner syndrome. Clin. Nucl. Med. 41, 797–798 (2016).
Blondin, D. P. et al. Selective impairment of glucose but not fatty acid or oxidative metabolism in brown adipose tissue of subjects with type 2 diabetes. Diabetes 64, 2388–2397 (2015).
Richard, G. et al. High-fructose feeding suppresses cold-stimulated brown adipose tissue glucose uptake independently of changes in thermogenesis and the gut microbiome. Cell Rep. Med. 3, 100742 (2022).
Cannon, B. et al. Neuropeptide tyrosine (NPY) is co-stored with noradrenaline in vascular but not in parenchymal sympathetic nerves of brown adipose tissue. Exp. Cell. Res. 164, 546–550 (1986).
De Matteis, R., Ricquier, D. & Cinti, S. TH-, NPY-, SP-, and CGRP-immunoreactive nerves in interscapular brown adipose tissue of adult rats acclimated at different temperatures: an immunohistochemical study. J. Neurocytol. 27, 877–886 (1998).
Huesing, C. et al. Organization of sympathetic innervation of interscapular brown adipose tissue in the mouse. J. Comp. Neurol. 530, 1363–1378 (2022).
Kumari, R. et al. Sympathetic NPY controls glucose homeostasis, cold tolerance, and cardiovascular functions in mice. Cell Rep. 43, 113674 (2024).
Zhu, Y. et al. Sympathetic neuropeptide Y protects from obesity by sustaining thermogenic fat. Nature 634, 243–250 (2024).
Hankir, M. K. et al. Dissociation between brown adipose tissue 18F-FDG uptake and thermogenesis in uncoupling protein 1-deficient mice. J. Nucl. Med. 58, 1100–1103 (2017).
Olsen, J. M. et al. β3-Adrenergically induced glucose uptake in brown adipose tissue is independent of UCP1 presence or activity: mediation through the mTOR pathway. Mol. Metab. 6, 611–619 (2017).
Fischer, A. W. et al. Brown adipose tissue lipoprotein and glucose disposal is not determined by thermogenesis in uncoupling protein 1-deficient mice. J. Lipid Res. 61, 1377–1389 (2020).
Becher, T. et al. Brown adipose tissue is associated with cardiometabolic health. Nat. Med. https://doi.org/10.1038/s41591-020-1126-7 (2021).
Cannon, B. & Nedergaard, J. Brown adipose tissue: function and physiological significance. Physiol. Rev. 84, 277–359 (2004).
Weir, G. et al. Substantial metabolic activity of human brown adipose tissue during warm conditions and cold-induced lipolysis of local triglycerides. Cell. Metab. 27, 1348–1355 e1344 (2018).
Roberts-Toler, C., O’Neill, B. T. & Cypess, A. M. Diet-induced obesity causes insulin resistance in mouse brown adipose tissue. Obesity 23, 1765–1770 (2015).
Heine, M. et al. Lipolysis triggers a systemic insulin response essential for efficient energy replenishment of activated brown adipose tissue in mice. Cell. Metab. 28, 644–655 e644 (2018).
Furness, J. B., Morris, J. L., Gibbins, I. L. & Costa, M. Chemical coding of neurons and plurichemical transmission. Annu. Rev. Pharmacol. Toxicol. 29, 289–306 (1989).
Li, C. & Horn, J. P. Physiological classification of sympathetic neurons in the rat superior cervical ganglion. J. Neurophysiol. 95, 187–195 (2006).
Lundberg, J. M. et al. Neuropeptide Y (NPY)-like immunoreactivity in peripheral noradrenergic neurons and effects of NPY on sympathetic function. Acta Physiol. Scand. 116, 477–480 (1982).
Pernow, J., Ohlen, A., Hokfelt, T., Nilsson, O. & Lundberg, J. M. Neuropeptide Y: presence in perivascular noradrenergic neurons and vasoconstrictor effects on skeletal muscle blood vessels in experimental animals and man. Regul. Pept. 19, 313–324 (1987).
Gibbins, I. L. Vasomotor, pilomotor and secretomotor neurons distinguished by size and neuropeptide content in superior cervical ganglia of mice. J. Auton. Nerv. Syst. 34, 171–183 (1991).
Zhang, L. et al. Stress- and diet-induced fat gain is controlled by NPY in catecholaminergic neurons. Mol. Metab. 3, 581–591 (2014).
Yan, C. et al. Peripheral-specific Y1 receptor antagonism increases thermogenesis and protects against diet-induced obesity. Nat. Commun. 12, 2622 (2021).
Wang, T., Teng, B., Yao, D. R., Gao, W. & Oka, Y. Organ-specific sympathetic innervation defines visceral functions. Nature https://doi.org/10.1038/s41586-024-08269-0 (2024).
Tervo, D. G. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Morrison, S. F., Ramamurthy, S. & Young, J. B. Reduced rearing temperature augments responses in sympathetic outflow to brown adipose tissue. J. Neurosci. 20, 9264–9271 (2000).
Francois, M. et al. Sympathetic innervation of the interscapular brown adipose tissue in mouse. Ann. N. Y. Acad. Sci. 1454, 3–13 (2019).
Wang, Y. et al. The role of somatosensory innervation of adipose tissues. Nature 609, 569–574 (2022).
Chen, X. et al. Engineered AAVs for non-invasive gene delivery to rodent and non-human primate nervous systems. Neuron 110, 2242–2257 e2246 (2022).
Liu, X., Perusse, F. & Bukowiecki, L. J. Mechanisms of the antidiabetic effects of the beta 3-adrenergic agonist CL-316243 in obese Zucker-ZDF rats. Am. J. Physiol. 274, R1212–R1219 (1998).
Xiao, C., Goldgof, M., Gavrilova, O. & Reitman, M. L. Anti-obesity and metabolic efficacy of the beta3-adrenergic agonist, CL316243, in mice at thermoneutrality compared to 22 °C. Obesity 23, 1450–1459 (2015).
Kong, X. et al. IRF4 is a key thermogenic transcriptional partner of PGC-1α. Cell 158, 69–83 (2014).
Wang, X. & Wahl, R. Responses of the insulin signaling pathways in the brown adipose tissue of rats following cold exposure. PLoS ONE 9, e99772 (2014).
Gutierrez, L. S. & Gutierrez, J. Thrombospondin 1 in metabolic diseases. Front. Endocrinol. 12, 638536 (2021).
Laduron, P. & Belpaire, F. Transport of noradrenaline and dopamine-β-hydroxylase in sympathetic nerves. Life Sci. 7, 1–7 (1968).
Furlan, A. et al. Visceral motor neuron diversity delineates a cellular basis for nipple- and pilo-erection muscle control. Nat. Neurosci. 19, 1331–1340 (2016).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 e1022 (2018).
Kosmidis, S., Negrean, A., Dranovsky, A., Losonczy, A. & Kandel, E. R. A fast, aqueous, reversible three-day tissue clearing method for adult and embryonic mouse brain and whole body. Cell Rep. Methods 1, 100090 (2021).
Hansson, S. R., Mezey, E. & Hoffman, B. J. Ontogeny of vesicular monoamine transporter mRNAs VMAT1 and VMAT2. II. Expression in neural crest derivatives and their target sites in the rat. Brain Res. Dev. Brain Res. 110, 159–174 (1998).
Schutz, B., Schafer, M. K., Eiden, L. E. & Weihe, E. Vesicular amine transporter expression and isoform selection in developing brain, peripheral nervous system and gut. Brain Res. Dev. Brain Res. 106, 181–204 (1998).
Jensen, M. D. Brown adipose tissue—not as hot as we thought. J. Physiol. 593, 489 (2015).
Shin, H. et al. Lipolysis in brown adipocytes is not essential for cold-induced thermogenesis in mice. Cell Metab. 26, 764–777 e765 (2017).
Qi, L. et al. A mouse DRG genetic toolkit reveals morphological and physiological diversity of somatosensory neuron subtypes. Cell 187, 1508–1526 e1516 (2024).
Matsen, M. E. et al. In uncontrolled diabetes, thyroid hormone and sympathetic activators induce thermogenesis without increasing glucose uptake in brown adipose tissue. Am. J. Physiol. Endocrinol. Metab. 304, E734–E746 (2013).
Madisen, L. et al. A toolbox of Cre-dependent optogenetic transgenic mice for light-induced activation and silencing. Nat. Neurosci. 15, 793–802 (2012).
Daigle, T. L. et al. A suite of transgenic driver and reporter mouse lines with enhanced brain-cell-type targeting and functionality. Cell 174, 465–480 e422 (2018).
van den Pol, A. N. et al. Neuromedin B and gastrin-releasing peptide excite arcuate nucleus neuropeptide Y neurons in a novel transgenic mouse expressing strong Renilla green fluorescent protein in NPY neurons. J. Neurosci. 29, 4622–4639 (2009).
Suh, J. et al. A simple and fast optical clearing method for whole-mount fluorescence in situ hybridization (FISH) imaging. J. Biophoton. https://doi.org/10.1002/jbio.202400258 (2024).
R: a language and environment for statistical computing. R Foundation for Statistical Computing v4.1.0 2021-05-18 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Su, B. et al. A DMS shotgun lipidomics workflow application to facilitate high-throughput, comprehensive lipidomics. J. Am. Soc. Mass Spectrom. 32, 2655–2663 (2021).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Saxena, A. et al. Trehalose-enhanced isolation of neuronal sub-types from adult mouse brain. Biotechniques 52, 381–385 (2012).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 20, 484–496 (2017).
Snyder, M. E. et al. Generation and persistence of human tissue-resident memory T cells in lung transplantation. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aav5581 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Bai, L. et al. Genetic identification of vagal sensory neurons that control feeding. Cell 179, 1129–1143 e1123 (2019).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. GigaScience https://doi.org/10.1093/gigascience/giy083 (2018).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Jing, D., Men, Y. & Zhao, H. Tissue clearing and 3-D visualization of vasculature with the PEGASOS method. Methods Mol. Biol. 2319, 1–13 (2021).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Wang, Y. et al. A key role of PIEZO2 mechanosensitive ion channel in adipose sensory innervation. Cell Metab. 37, 1001–1011 e1007 (2025).
Neri, D. et al. Temperature and Doppler recordings from anesthetized mice. figshare https://doi.org/10.6084/m9.figshare.30491075 (2025).
Neri, D. et al. Lipidomics data from anesthetized mice (Th-Cre mice injected with MaCPNS1:CAG–DIO–hm3Dq–mCherry in iBAT). figshare https://doi.org/10.6084/m9.figshare.30524474 (2025).
Neri, D. et al. Distinct sympathetic projections to brown fat regulate thermogenesis and glucose tolerance. figshare https://doi.org/10.6084/m9.figshare.30642500 (2025).
Acknowledgements
We thank J. Campbell for guidance in ganglia dissociation, G. Mentis for advice on the CTB retrograde labelling, Z. Knight for help with removing glial genes from scRNA-seq analyses, M. Jenkins for providing the LIMPID reagent, S. Kosmidis for help with Fast 3D Clear and S. Ho for generating the exported BioRender files. This research was funded by the NIH OT2OD023853 (to L.M.Z.), R01DK125094 (to L.M.Z.), P01AG032959 (to L.M.Z.), R56DK140355 (to L.M.Z.), the Chan Zuckerberg Initiative (to L.M.Z and J.H.), the American Diabetes Association 1-17-IBS-208 (to L.M.Z.), the Russell Berrie Foundation (to L.M.Z.), the DFG-funded research consortium SFB-Transregio 333, project-ID: 450149205 (to A.W. and J.H.) and a fellowship from the Kwanjeong Educational Foundation (to S.L.). This research was supported by several core facilities at Columbia University: NCI Cancer Center support grant P30CA013696 for the Confocal and Specialized Microscopy, Genetically Modified Mouse Models and Molecular Pathology shared resources of the Herbert Irving Comprehensive Cancer Center; CTSA grant UL1 TR001873 for the Single Cell Analysis Core, NIDDK Diabetes Research Center support grant P30DK063608 and S10OD034377 for the Advanced Tissue Pathology and Imaging Core. Microscopy was performed with support from the Zuckerman Institute’s Cellular Imaging Platform (grant no. S10OD023587). Zuckerman Institute’s Molecular Tools platform generated custom adeno-associated virus (AAV) transfer plasmids and packing for the inhibitory DREADD. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
D.N., S.L. and L.M.Z. conceived of and designed the experiments and interpreted the data. S.L. performed the scRNA-seq. D.N. and V.V.T. analysed the scRNA-seq data. S.L., A.J.L. and D.N. performed and imaged the smFISH experiments. X.C. and V.G. provided the MACPNS1 viruses. N.R.L., L.C.S. and D.N. performed the immunostaining and imaging of virus-injected ganglia. D.N. performed the chemogenetic experiments. D.B. maintained all mouse lines and performed glucose tolerance tests with D.N., with help from L.C.S. D.N., A.M.R.-L. and A.M.F. dissected the ganglia. G.M.N counted the labelled neurons in whole SG and performed correlation analyses with help from D.N. M.H., A.W. and J.H. performed and analysed the lipidomics experiment and made intellectual contributions to their interpretation. L.M.Z. and D.N. wrote the paper with discussions and contributions from S.L., V.V.T. and J.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Jake Willows and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Jean Nakhle, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Retrograde transport of Cre-dependent viruses targets SNS neurons projecting to iBAT without affecting sensory neurons.
Whole tissue imaging from mice after injection of viruses and CTB into iBAT. a, SG harvested from a WT mouse injected with CTB-555. b, SG harvested from a WT mouse injected with AAVrg:CAG-GFP. c, SG harvested from a WT mouse injected with MaCPNS1:CAG-GFP. d, SG harvested from a Th-Cre mouse injected with MaCPNS1:CAG-DIO-mCherry. e, Intact cleared trunk tissue containing the thoracic sympathetic ganglia from a Th-Cre mouse injected with MaCPNS1:CAG-DIO-mCherry in iBAT. f, SG and DRGs harvested from a Th-Cre mouse injected with MaCPNS1:CAG-DIO-hM3D-mCherry. g, SG and DRGs harvested from a Rxfp1-Cre mouse injected with MaCPNS1:CAG-DIO-hM3D-mCherry. h, SG and DRGs harvested from a Vmat1-Cre mouse injected with MaCPNS1:CAG-DIO-hM3D-mCherry. a-d and f-h, SGs are shown as the central section across the z-stack, DRGs are maximum projections of the z-stack. Scale bars are 200 µm. e, Maximum projection of the z-stack, scale bar is 700 µm.
Extended Data Fig. 2 Sensory neurons projecting to iBAT do not express tyrosine hydroxylase.
a, Representative picture of tyrosine hydroxylase (TH) and green fluorescent protein (GFP) immunofluorescent staining in the DRGs of WT mice injected with MaCPNS1:CAG-GFP in iBAT. b, Average number of neurons expressing GFP and/or TH per DRG in mice from a (161 sections from n = 3 mice). Data presented as the mean + s.e.m.
Extended Data Fig. 3 Effects of chemogenetic stimulation of SNS projections to iBAT are not proportional to the number of neurons infected.
a, Correlation between the increase in iBAT temperature following CNO injection and the average number of SG neurons expressing mCherry from mice shown in Fig. 1f and g (n = 16 mice, Spearman’s r = 0.3, P = 0.26). b, Correlation between the increase in iBAT blood flow following CNO injection and the average number of SG neurons expressing mCherry from mice shown in Fig. 1f and g (n = 16 mice, Spearman’s r = -0.18, P = 0.52).
Extended Data Fig. 4 Dissociation between iBAT thermogenesis and blood flow following chemogenetic activation of iBAT-projecting SNS in anesthetized animals.
Effects of treatment with adrenergic receptor antagonists on chemogenetically-induced changes in Th-Cre mice injected with a Cre-dependent hM3D-mCherry-expressing MaCPNS1 virus in iBAT. a,b, Average traces for thermogenesis (Δ iBAT-LB)(brown), lower back temperature (orange) and blood flow (blue) following injection of propranolol (n = 6 mice) (a) or phentolamine (n = 3 mice) (b). Data presented as mean (solid line) ± s.e.m. (shaded area). c, Changes in thermogenesis in the same animals from a,b, P = 0.0228. d, Changes in blood flow in the same animals from a and b, P = 0.713. Data presented as mean ± s.e.m. c,d unpaired Mann-Whitey test, two-tailed *P < 0.05.
Extended Data Fig. 5 DREADD ligands reduce RER but do not affect thermogenesis or glucose tolerance.
a, Schematic illustration of the random crossover experimental design to assess the independent impact of DCZ and CNO versus saline treatment in awake WT mice (data shown in b-f). b, Effects of CNO, DCZ, or saline delivered immediately before a glucose gavage on circulating glucose levels. Data presented as the mean ± s.e.m. (n = 7 mice). c, Within-subject effects (DCZ minus saline and CNO minus saline) on blood glucose levels from the same mice shown in b. d, Average traces of energy expenditure (kcal/h). Solid lines represent means, shaded areas represent ± s.e.m. (n = 7 mice). e, f, Changes in energy expenditure (e) and respiratory exchange ratio (RER, f) following saline, DCZ, or CNO injection (n = 7 mice). Dots represent average values, lines represent individual mice. f, Repeated measure two-way ANOVA (sex and treatment, P = 0.418 for sex and P = 0.043 for treatment), *P < 0.05 after Dunnett’s post-hoc test with saline as reference.
Extended Data Fig. 6 Chemogenetic stimulation of SNS projections to iBAT does not affect baseline glycemia.
Effects of chemogenetic stimulation of iBAT-projecting neurons in awake mice without glucose gavage. All Cre driver lines were injected with a Cre-dependent hM3D-mCherry-expressing MaCPNS1 virus in iBAT. a-c, Effect of DCZ or saline on circulating glucose levels in Th-Cre mice (n = 16 mice, each tested in both conditions) (b) and within-subject effects (DCZ minus saline) on blood glucose levels from the same mice (c). Data presented as the mean ± s.e.m. d-f, Effect of DCZ or saline on circulating glucose levels in Rxfp1-Cre mice. (n = 4 mice, each tested in both conditions) (e) and within-subject effects (DCZ minus saline) on blood glucose levels from the same mice (f). Data presented as the mean ± s.e.m. g-i, Effect of DCZ or saline on circulating glucose levels in Vmat1-Cre mice. (n = 6 mice, each tested in both conditions) (h) and within-subject effects (DCZ minus saline) on blood glucose levels from the same mice (i). Data presented as the mean ± s.e.m.
Extended Data Fig. 7 Strategy for scRNA-Seq in conjunction with retrograde tracing.
a, Mice were injected with CTB-A555 into iBAT and CTB-A488 into forelimb. b, SG were harvested and cells were dissociated for FACS. c, Large, viable neurons were gated based on the absence of Sytox blue staining and forward scatter. d, A555 + , A488+ and unlabeled (A000) cells were sorted into 96-well plates containing lysis buffer for scRNA-Seq.
Extended Data Fig. 8 The distribution of stellate ganglion cluster markers is shared with the superior cervical ganglion but not with the celiac ganglion.
a-b, Validation of SG cluster marker genes using smFISH in conjunction with the Npy-GFP reporter in n = 4 mice. SG1 markers assessed were Npy-GFP, Rfxp1, Fst and Sctr. Markers for other SG populations were Cckar (SG2-3), Colq (SG2-4) and Aqp1 (SG3). a, Superior cervical ganglion. b, Celiac ganglion. Scale bar is 20 µm.
Extended Data Fig. 9 The Rxfp1-Cre::tdTOM lineage trace is restricted to Rxfp1+ neurons, while the Vmat1-Cre::tdTOM lineage trace is broadly expressed in the SG.
a-c, Validating expression of the Rxfp1-Cre::tdTomato lineage trace. a, Schematic of breeding strategy used to generate Rxfp1-Cre::tdTomato mice. b, Whole tissue imaging of SG harvested from a Rxfp1-Cre::tdTomato mouse. c, smFISH for Rxfp1 and tdTomato (top), and for Colq and tdTomato (bottom) expression in SG cryosections. Scale bar is 10 µm. d-f, Validating expression of the Vmat1-Cre::tdTomato lineage trace. d, Schematic of breeding strategy used to generate Vmat1-Cre::tdTomato mice. e, Whole tissue imaging of SG harvested from a Vmat1-Cre::tdTomato mouse. f, smFISH for Rxfp1 and tdTomato expression in SG cryosections. Scale bar is 10 µm. SG in b and e are shown as maximum projections of the z-stack, scale bar is 200 µm. Schematics in a and d were generated with Biorender.
Extended Data Fig. 10 Nerves from Rxfp1+ sympathetic neurons are closely associated with blood vessels in mouse iBAT.
a, Diameter of iBAT blood vessels innervated by tdTomato-positive nerves in Rxfp1-Cre::tdTomato mice (2 mice, 3 regions analyzed per mouse, n = 24 unique innervated blood vessels from one mouse and n = 47 from the other). Data presented as box plots. Whiskers indicate min and max values, left and right borders are 25th and 75th percentile, center is the median. b, Proportion of distances between tdTomato-positive nerves and their closest blood vessel in iBAT of Rxfp1-Cre::tdTomato and Th-Cre::tdTomato mice, grouped into 1 µm bins (2 mice per genotype, 3 regions analyzed per mouse, P < 0.001). c, Same data from b, grouped in bins of either 0, 0.41-2, 2-15 or >15 µm, P < 0.001. b, Chi-squared test, two-tailed, P < 0.001. c, Kolmogorov-Smirnov test, two-sided, P < 0.001.
Supplementary information
Supplementary Table (download XLSX )
Complete lists of GO terms associated with Fig. 2b,d,e and information about the AAVs utilized in each experiment.
Supplementary Video 1 (download AVI )
Representative video of the distribution of Th-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Video 2 (download AVI )
Representative video of the distribution of Th-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Video 3 (download AVI )
Representative video of the distribution of Th-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Video 4 (download AVI )
Representative video of the distribution of Rxfp1-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Video 5 (download AVI )
Representative video of the distribution of Rxfp1-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Video 6 (download AVI )
Representative video of the distribution of Rxfp1-Cre::tdTOM fibres relative to blood vessels labelled with isolectin.
Supplementary Data 1 (download XLSX )
Source data for Supplementary Fig. 1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Neri, D., Lee, S., Fohn, A.M. et al. Distinct sympathetic projections to brown fat regulate thermogenesis and glucose tolerance. Nat Metab 8, 313–326 (2026). https://doi.org/10.1038/s42255-025-01429-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01429-0
This article is cited by
-
Heterogeneous sympathetic control of brown adipose tissue
Nature Metabolism (2026)