Abstract
From insects to mammals, essential brain functions, such as forming long-term memories (LTMs), increase metabolic activity in stimulated neurons to meet the energetic demand associated with brain activation. However, while impairing neuronal metabolism limits brain performance, whether expanding the metabolic capacity of neurons boosts brain function remains poorly understood. Here, we show that LTM formation of flies and mice can be enhanced by increasing mitochondrial metabolism in central memory circuits. By knocking down the mitochondrial Ca2+ exporter Letm1, we favour Ca2+ retention in the mitochondrial matrix of neurons due to reduction of mitochondrial H+/Ca2+ exchange. The resulting increase in mitochondrial Ca2+ over-activates mitochondrial metabolism in neurons of central memory circuits, leading to improved LTM storage in training paradigms in which wild-type counterparts of both species fail to remember. Our findings unveil an evolutionarily conserved mechanism that controls mitochondrial metabolism in neurons and indicate its involvement in shaping higher brain functions, such as LTM.
Similar content being viewed by others
Main
The energetic cost of brain functioning imposes substantial energetic demands to organisms. Transmission of information between neurons generates acute and local energy costs at synaptic sites, which amount to ~75% of the brain’s total energy expenditure1. Organismal metabolic states in which the brain experiences limited energy supply, such as hypoxia or hypoglycaemia, constrain the information processing capability of synapses and impair brain function2,3. As such, neurons must ensure that energy levels are preserved, even during energy-demanding neuronal computations associated with complex brain functions. Mitochondria are known to generate more than 90% of neuronal energy in the form of ATP via oxidative phosphorylation (OXPHOS)4 and are strategically located along the complex neuronal morphology to be in the ideal position to locally provide ATP on demand5,6,7,8,9,10,11,12,13. When neural circuits are activated, fuels are provided on demand to activated brain regions14,15. This activation is accompanied by transient increases in neuronal metabolic rates, which enables neurons to use these fuels and counterbalance energy usage associated with neurotransmission. Dynamic adjustments of neuronal mitochondrial metabolism are thus essential to enable a wide variety of brain computations, ranging from the formation of LTMs to the control of social behaviours in flies and mammals16,17,18,19. LTM formation, in particular, is well known to be an energetically demanding process20,21,22. A large body of research in Drosophila melanogaster has demonstrated that adaptations in neuronal metabolism enable the formation and maintenance of LTM, and that impaired mitochondrial function can hinder this process19,23. These results support the widely accepted idea in which energy is seen as an enabler of function, but not as a driving factor. This standpoint implies that once the energy needs of neural circuits are met, brain function should proceed optimally and additional energy will not necessarily enhance behavioural efficiency.
At the cellular level, axonal mitochondria sense neuronal activity and produce ATP on demand to preserve the metabolic integrity of presynaptic function24,25,26. During neurotransmission, Ca2+ invades presynaptic sites27,28, increasing axonal mitochondrial Ca2+29,30,31,32,33. While during high-frequency stimulation this buffering process may impact vesicle cycling31,32, in physiological conditions mitochondrial Ca2+ uptake serves as a feed-forward mechanism that transiently increases axonal mitochondrial metabolism to indirectly favour ATP production and compensate for energy usage10,34. A similar coordination between neuronal activity, mitochondrial Ca2+ uptake and increases in metabolism also occurs in somatodendritic compartments12,35, although depending on the experimental conditions, matrix Ca2+ increases have been shown not to be required for somatodendritic metabolic upregulation36,37,38. While cell-type-specific and compartment-specific differences likely exist, non-pathological Ca2+ increases in the axonal mitochondrial matrix are thought to speed up tricarboxylic acid (TCA) cycle rates in a reversible manner5,14,39,40 because Ca2+ transiently binds to several TCA cycle dehydrogenases to increase their enzymatic activity41,42. When neurons cease to fire, however, mitochondrial Ca2+ must be actively extruded from the matrix10,29,30,31,32, a process that should in theory decelerate mitochondrial metabolism back to baseline rates. The inability to do so would effectively increase the time Ca2+ stays in the mitochondrial matrix, over-activating mitochondrial metabolism beyond what would be necessary to counterbalance activity-driven ATP consumption, thus theoretically leading to ATP accumulation. However, while such perturbation could be ideal to boost metabolism in firing neurons, the molecular control of mitochondrial Ca2+ efflux remains poorly understood, limiting our understanding of how this process may serve as a control point in the adjustment of synaptic bioenergetics and the metabolic states of brain circuits and behaviour.
Two non-exclusive mitochondrial systems can potentially extrude Ca2+ from the matrix in cells: NCLX, a Na+/Ca2+/Li+ exchanger43,44, and Letm1 (Leucine zipper and EF-hand containing transmembrane protein 1), an H+/Ca2+ exchanger45,46,47,48. Here, we identify Letm1 as an activity-dependent mitochondrial Ca2+ extruder that controls efflux rates in neuronal mitochondria. We found that increased matrix Ca2+ retention over-activated mitochondrial metabolism in rodent primary neurons in vitro and in mushroom body (MB) neurons of D. melanogaster in vivo. This upregulated metabolic state relied on Ca2+ activation of mitochondrial metabolism, as the knockdown (KD) of Ca2+-activated pyruvate dehydrogenase phosphatase 1 (PDP1) abolished metabolic acceleration in both systems. Given that LTM is an energetically demanding brain function, we knocked down Letm1 in specific memory circuits of flies and rodents and found that both species presented robust long-term olfactory memories in conditions in which wild-type counterparts failed to remember. Together, our results reveal the importance of mitochondrial Ca2+ efflux in shaping neuronal metabolism and suggest that targeted metabolic modulations in neural circuits can considerably enhance specific memories across species.
Results
Letm1 controls mitochondrial Ca2+ efflux rates in firing neurons
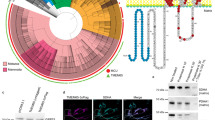
While NCLX contributes to mitochondrial Ca2+ extrusion in firing axons49,50,51, NCLX inactivation leads to pronounced increases in resting mitochondrial Ca2+ levels50,52,53, which causes neurodegeneration both in vitro and in vivo in rodent hippocampal neurons52,54. To circumvent this, we explored the contribution of Letm1 to the control of mitochondrial Ca2+ extrusion during activity (Fig. 1a). We first used primary dissociated rat hippocampal neurons, cultured within a layer of astrocytes (Methods and Extended Data Fig. 1a,b), as they allow high-resolution quantitative measurements of neuronal mitochondrial physiology during activity6,10,40,49,55. Resting free Ca2+ levels in neuronal mitochondria are higher than in the cytosol, yet remain in the hundreds of nanomolar range10,56,57,58, enabling sensors originally developed for cytosolic Ca2+ to report robust mitochondrial responses10,13. We thus co-expressed a mitochondria-targeted GCaMP6 sensor (Mito4x-GCaMP6f10) with a short hairpin RNA (shRNA) that depleted Letm1 levels by ∼70% (Methods and Extended Data Fig. 1c,d) and compared axonal mitochondrial Ca2+ fluxes in wild-type and Letm1 KD neurons using physiological firing paradigms of hippocampal neurons in vivo, such as firing at 20 Hz for 1 s59 (Fig. 1b). We found that impairing Letm1 expression caused a ~3-fold reduction in the rate of mitochondrial Ca2+ extrusion (Fig. 1b,c), indicating that Letm1 is involved in controlling activity-driven mitochondrial Ca2+ efflux in axonal mitochondria. We observed similar results when neurons were stimulated at a higher frequency (Extended Data Fig. 1e,f), suggesting that Letm1 is active in conditions with variable levels of Ca2+ entry. Repeating both these experiments in imaging media with 1.2 mM glucose, a physiological concentration found in the hippocampus60,61, yielded indistinguishable outcomes (Extended Data Fig. 1g,h), indicating that elevated glucose in the imaging media does not alter Letm1-mediated mitochondrial Ca2+ extrusion dynamics. As a control, we confirmed that Letm1 KD neurons did not present alterations in activity-driven mitochondrial Ca2+ uptake capacity (Fig. 1d and Extended Data Fig. 1e–h).
a, Letm1 is a protein located on the inner mitochondrial membrane where it transports one Ca2+ in exchange for two H+. It contains a Ca2+-binding EF-hand domain facing the mitochondrial matrix. OMM, outer mitochondrial membrane; IMM, inner mitochondrial membrane; MCU, mitochondrial Ca2+ uniporter. b, Axonal mitochondrial Ca2+ responses to stimulation (20 action potentials (APs), 20 Hz) in control and Letm1 KD neurons. The figure shows traces corresponding to the mean ± s.e.m. of Mito4x-GCaMP6f in axons. c, Rate of mitochondrial Ca2+ decay measured as half-time decay (t1/2) in axonal mitochondria following stimulation in control and Letm1 KD neurons. Statistical significance was assessed using a two-tailed Mann–Whitney U-test (****P < 0.0001; n = 27 for control neurons, n = 21 for Letm1 KD neurons; U = 72). d, Peak Mito4x-GCaMP6f responses (ΔF/F0) after stimulation. Non-significance (NS) was determined using a two-tailed Mann–Whitney U-test (P = 0.578; n = 27 for control neurons, n = 21 for Letm1 KD neurons; U = 256). e, Baseline mitochondrial Ca2+ concentration (nM) in control and Letm1 KD neurons. Non-significance was determined using a two-tailed unpaired t-test (NS, P = 0.6023, n = 16 control neurons, n = 11 Letm1 KD neurons, t25 = 0.5279). f, Half-time decay in axonal mitochondria following stimulation in control, Letm1 KD neurons, Letm1 KD neurons expressing rat Letm1 wild-type protein and Letm1 KD neurons expressing rat Letm1 protein with the EF-hand domain mutated. P values were determined by a Kruskal–Wallis test followed by Dunn’s multiple-comparisons test against the control group (****P < 0.0001; n = 19 for control, n = 13 for Letm1 KD, n = 7 for wild-type (WT) Letm1 rescue and n = 8 for EF-hand mutant Letm1. Post hoc P values; control versus Letm1 KD, P = 0.0001; control versus Letm1 KD + Letm1 rescue, P = 0.3126; Letm1 KD + Letm1ΔEF-hand rescue, P = 0.0022). g, Dendritic mitochondrial Ca2+ responses to a single glutamate uncaging pulse in a single spine in control neurons and Letm1 KD neurons. Traces correspond to the mean ± s.e.m. of Mito4x-GCaMP6f. h, Half-time decay in dendritic mitochondrial Ca2+ following stimulation in control and Letm1 KD neurons. P value was determined using a two-tailed Mann–Whitney U-test (***P < 0.0001, n = 63 dendrites for control, n = 47 dendrites for Letm1 KD, U = 809). i, Fold change in half-time decay in Letm1 KD neurons in axonal and dendritic compartments. P value was determined using a two-tailed unpaired t-test (****P < 0.0001, with n = 33 for axons and n = 47 for dendrites, t78 = 4.959). Data are represented as the mean ± s.e.m. See Extended Data Figs. 1 and 2.
Letm1 KD neurons did not present alterations in resting mitochondrial Ca2+ levels, estimated using two different methods (Fig. 1e, Extended Data Fig. 1i,j and Methods). Quantitative estimates revealed that mitochondrial Ca2+ levels rose to ~425 nM during 20-Hz stimulation and to ~530 nM at 100 Hz in both control and Letm1 KD neurons (Extended Data Fig. 1K), indicating that neuronal activity elicits moderate increases in mitochondrial Ca2+. Axonal responses to field stimulation were measured in the presence of the postsynaptic blockers 6-cyano-7-nitroquinoxaline-2,3-dione (CNQX) and d,l-2-amino-5-phospho-novaleric acid (AP5) to study presynaptic function without the concurrent incoming excitation from stimulating surrounding unlabelled neurons10,49,62. Re-expression of a shRNA-resistant variant of wild-type Letm1 in Letm1 KD neurons rescued mitochondrial Ca2+ efflux rates to a large extent, supporting that Letm1 controls activity-driven Ca2+ efflux (Fig. 1f). Letm1 presents a Ca2+-binding EF-hand domain oriented towards the mitochondrial matrix, which may drive its activation63,64 (Fig. 1a). We generated a shRNA-resistant variant of Letm1 in which the key conserved coordinating aspartates of the EF-hand motif were mutated to alanines (p.Asp676Ala and p.Asp680Ala, ΔEF-hand), reducing its Ca2+-binding capacity65. Co-expression of this mutant with Mito4x-GCaMP6f and Letm1 shRNA failed to rescue the slower Ca2+ efflux rates caused by the Letm1 KD (Fig. 1f), suggesting that the EF-hand domain of Letm1 is required for its export function. To confirm that overexpressed Letm1 localized to mitochondria, we fused it to mRFP1 and transfected it together with Mito4x-GCaMP6f, which confirmed that Letm1–mRFP1 localized correctly to axonal mitochondria (Extended Data Fig. 1l). Taken together, these results indicate that Letm1 acts as a mitochondrial Ca2+ exporter in neurons.
We examined any possible effect on cytosolic Ca2+ dynamics in axons by Letm1 KD using a cytosolic Ca2+ sensor, cytoGaMP8f66 (Extended Data Fig. 1m,n). We did not observe any differences in the amplitude of cytosolic Ca2+ responses (Extended Data Fig. 1n) or in its extrusion rates (Extended Data Fig. 1o). These results indicate that manipulating the slow dynamics of mitochondrial Ca2+ efflux, which occurs over tens of seconds, does not affect the rapid dynamics of cytosolic Ca2+ during firing, which takes place within a few seconds. Additionally, no differences were observed in resting cytosolic Ca2+ levels using established quantitative methods62 (Extended Data Fig. 1p). As Letm1 exchanges Ca2+ for H+, we examined whether removing it from axonal mitochondria could impact neuronal and mitochondrial pH physiology. Using optical pH sensors for the mitochondrial matrix (Mito4x-pHluorin10) and the cytosol (Cyto-pHluorin25), we first confirmed that resting axonal mitochondrial or cytosolic pH were not affected by Letm1 KD (Methods and Extended Data Fig. 2a–d). Next, we measured activity-driven mitochondrial and cytosolic pH dynamics and found them to be indistinguishable between wild-type and Letm1 KD neurons (Extended Data Fig. 2e). These results suggest that mitochondrial pH is not strongly affected by Letm1 KD. Collectively, these results show that Letm1 primarily acts as a mitochondrial Ca2+ exporter during neuronal activity without significantly impacting other aspects of presynaptic Ca2+ signalling.
Axonal and dendritic mitochondria present marked differences in their structure, function and regulation67,68. We next examined whether Letm1 was also involved in controlling mitochondrial Ca2+ efflux in dendritic mitochondria. During neurotransmission, dendritic mitochondria uptake Ca2+ to later release it back to the cytosol when transmission is over55. To study this process, we used two-photon glutamate uncaging to stimulate single spines once and quantified cytosolic and mitochondrial Ca2+ dynamics simultaneously in dendritic shafts of neurons in the presence or absence of Letm1 shRNA expression. We observed a ~70% reduction in dendritic mitochondrial Ca2+ efflux in Letm1 KD neurons compared to wild-type neurons (Fig. 1g,h). This effect, while present, appeared to be less pronounced than that observed in axons (Fig. 1i), although given the differences in biology and stimulation protocols between the two compartments it is difficult to draw conclusions about whether the rates of exclusion are actually different between axons and dendrites. We show normalized mitochondrial Ca2+ responses in dendrites to facilitate visualization of the Letm1-mediated slowdown effect (Fig. 1g,h), but the amplitude of dendritic mitochondrial Ca2+ uptake was reduced in Letm1 KD neurons (Extended Data Fig. 2f) despite cytosolic Ca2+ responses remaining unchanged (Extended Data Fig. 2g,h). Under these stimulation paradigms, the total amount of Ca2+ ions within the matrix of dendritic mitochondria measured as the area under the curve reported no difference, in contrast to axonal mitochondrial responses (Extended Data Fig. 2i). Given the established role of axonal mitochondrial Ca2+ handling in governing presynaptic energy levels10,35,69, we next focused on examining the importance of mitochondrial Ca2+ extrusion in controlling the metabolism of the presynapse.
Increased Ca2+ retention times in axonal mitochondria over-activate synapse metabolism
Mitochondrial Ca2+ uptake in firing axons transiently activates mitochondrial metabolism to produce ATP and sustain synaptic function locally4,26,27. We hypothesized that such transitory acceleration of mitochondrial metabolism should conclude when Ca2+ is exported back to the cytosol. Therefore, using ATP as a readout, we next studied the role of Letm1 in regulating presynaptic metabolism. We removed Letm1 sparsely in cultures of otherwise wild-type neurons, and allowed spontaneous activity to take place for 10 days. This paradigm challenges Letm1 KD neurons to produce and consume ATP repeatedly in response to spontaneous activity with average firing rates similar to hippocampal firing paradigms in vivo70,71, revealing the possible cumulative effect of unbalanced production and consumption over time under physiological firing conditions. We measured presynaptic ATP levels in Letm1 KD neurons in these conditions using either Syn-ATP25 or ATeam72,73, and found that Letm1 KD resulted in significant ATP accumulation in presynapses (Fig. 2a–c and Extended Data Fig. 3a–c). Calibrating Syn-ATP data reported that Letm1 KD neurons presented on average 2.64 mM ATP compared to 1.32 mM obtained from controls (Fig. 2a,b).
a, Representative images of ATP levels (green) measured at presynapses (red) with Syn-ATP in control, Letm1 KD and dual Letm1 KD + PDP1 KD neurons. Pseudocolour calibration bar indicates range of luminescence. Scale bars, 10 μm. b, Resting synaptic ATP levels in control, Letm1 KD and dual Letm1 KD + PDP1 KD neurons, together with control and Letm1 KD neurons with chronic TTX treatment. Left, P values were determined using a Kruskal–Wallis test (*P = 0.0142; n = 21 for control, n = 27 for Letm1 KD, n = 7 for Letm1/PDP1 KD, n = 8 for control + TTX, and n = 9 for Letm1 KD + TTX, H = 12.46, followed by Dunn’s multiple-comparisons test. Post hoc P values: control versus Letm1 KD, **P = 0.0058; control versus Letm1/PDP1 KD, NS, P > 0.9999; Letm1 KD versus Letm1/PDP1 KD, *P = 0.0370. Right, P value was determined using an unpaired two-tailed t-test. Control + TTX versus Letm1 KD + TTX, NS, t15 = 0.0026, n = 8, n = 9, P > 0.9980. c, Resting presynaptic ATP levels measured with the FRET ATP sensor ATeam 1.03. P value was determined using a two-tailed Mann–Whitney U-test (*P = 0.0357; n = 33 for control, n = 25 for Letm1 KD, U = 279). Experiments shown in a–c were done in 1.25 mM lactate and 1.25 mM pyruvate with no glucose. d, Model output 1: Computational simulation of 10,000 s of the evolution of ATP dynamics in control, Letm1 KD and dual Letm1 KD + PDP1 KD neurons with neuronal activity of ~2 Hz. y-axis units are arbitrary. e, Model output 2: fold change in ATP levels at the end of the 10,000-s simulation for control, Letm1 KD and dual Letm1 KD + PDP1 KD neurons firing with different mean firing rates (0 Hz, 1 Hz, 2 Hz, 5 Hz and 10 Hz). f, Representative traces of cytosolic pyruvate accumulation upon inhibition of mitochondrial metabolism with sodium azide in rat hippocampal neurons for control and Letm1 KD conditions. Pyruvate is measured using the FRET sensor Pyronic. g, Rate of pyruvate accumulation in control and Letm1 KD neurons. P value was determined using a two-tailed Mann–Whitney U-test (**P = 0.0021, n = 19 for control, n = 18 for Letm1 KD, U = 72). h, Model output 3: pyruvate consumption during 10,000 s for control, Letm1 KD and dual Letm1 KD + PDP1 KD neurons firing with different mean firing rates (0 Hz, 1 Hz, 2 Hz, 5 Hz and 10 Hz). Consumption is calculated as the change in pyruvate levels from start to end. Data are represented as the mean ± s.e.m. See also Extended Data Figs. 3 and 4. a.u., arbitrary units.
The most widely recognized effect of Ca2+ on mitochondrial metabolism is through its catalytic action on certain enzymes of the TCA cycle74,75,76. This regulatory effect includes the pyruvate dehydrogenase complex (PDHc), which converts pyruvate to acetyl-CoA at the entry point of the TCA cycle. PDHc is regulated by PDP, a Ca2+-activated enzyme that enhances PDHc activity through dephosphorylation77,78,79,80. PDHc will remain activated until pyruvate dehydrogenase kinase re-phosphorylates PDHc to inactivate it, stopping the transient activation initiated by PDP. Although two isoforms of PDP exist, PDP1 and PDP2, neurons predominantly express the Ca2+-sensitive isoform PDP1 (refs. 81,82). We thus reasoned that Letm1-mediated regulation of synaptic metabolism should require Ca2+ activation of mitochondrial PDP1. Using a single vector to knock down both Letm1 and PDP1 (Extended Data Fig. 3d–f), we measured presynaptic ATP and found that KD of PDP1 abolished Letm1-mediated accumulation of ATP (Fig. 2a,b and Extended Data Fig. 3b), suggesting that Letm1 controls mitochondrial metabolism at least in part by modulating Ca2+-mediated activation of the conversion of pyruvate to acetyl-CoA.
We reasoned that, given that electrical activity drives both ATP consumption and production, eliminating action potential firing using tetrodotoxin (TTX) should block any imbalance between the two processes and thus abolish the increased ATP levels in Letm1 KD neurons. We applied TTX for several days and observed ATP levels in Letm1 KD neurons were rescued to wild-type levels, indicating that Letm1-mediated control of synapse metabolism is activity driven (Fig. 2b and Extended Data Fig. 3c). To explore this further, we quantified ATP dynamics during activity in synapses of Letm1 KD neurons. We coupled electrophysiological stimulation to ATP imaging and quantified presynaptic ATP dynamics over time, removing glucose but providing lactate and pyruvate as mitochondrial fuels to reduce glycolytic contributions to these measurements10,83. Using a 10-Hz stimulation for 1 min as done before10,24,25,33,34,84, we found that while wild-type neurons preserved constant overall ATP levels over time10,24,25 (Extended Data Fig. 3g; black trace), Letm1 KD neurons presented an overshoot in ATP levels after stimulation (Extended Data Fig. 3g; pink trace), indicating that Letm1 KD caused a mismatch between consumption and production (Extended Data Fig. 3h). We confirmed that Letm1 KD did not alter the overall total abundance of ETC proteins, nor did it affect the expression of key presynaptic or postsynaptic markers (Extended Data Fig. 3i,j). Circularity of axonal mitochondria was indistinguishable between control and Letm1 KD neurons, indicating preserved overall shape (Extended Data Fig. 3k). A modest (~15%) increase in both mitochondrial size (Extended Data Fig. 3l) and mitochondrial number (Extended Data Fig. 3m) was observed, consistent with previous reports in non-neuronal cells85,86,87,88. Lastly, we quantified synapse number and size and found that Letm1 KD neurons displayed a higher synapse density along axons, while the estimated size of individual synapses was reduced (Extended Data Fig. 3n,o). These phenotypes were rescued when PDP1 was knocked down in Letm1 KD neurons, suggesting they arise from Ca2+ regulation of mitochondrial metabolism. Despite these structural differences, Letm1 KD synapses appeared equally functional, shown by measuring synaptic vesicle exocytosis during activity using the optical reporter vGlut-pH62,89 (Extended Data Fig. 3p). As a control, we estimated synaptic vesicle pH, which was identical in both conditions (Extended Data Fig. 3q). Together, these findings indicate that Letm1 modulates mitochondrial and synaptic structure and number, without significantly altering synaptic vesicle cycling.
We next built a computational model to better develop a theoretical framework that could help to represent and explore the theoretical space of possible behaviours of the multiple complex mitochondrial metabolic processes occurring during neuronal activity and the expected consequences of modulating some of them through the transient presence of Ca2+ as a boosting factor. Starting with a previous model of the mitochondrial TCA cycle and ATP production90, we added a fixed Ca2+ dependency for the corresponding enzymes of the TCA cycle91,92 (PDP1-PDHc, isocitrate dehydrogenase, alpha-ketoglutarate dehydrogenase) and ATP synthase (complex V92; Extended Data Fig. 4a). We included in the model a basal energy cost at rest, a fixed energy cost per spike (Extended Data Fig. 4b,c) and a fixed increase in mitochondrial Ca2+ per firing event (Extended Data Fig. 4d). The rates of Ca2+ efflux after spiking were modelled to follow a single exponential decay (control (t1/2) = 7 s, Letm1 KD (t1/2) = 20 s) based on experimental data (Fig. 1b,c). Using this model, we first simulated ATP dynamics during 1 min at 10 Hz, which showed increased ATP production capacity following neuronal activity in Letm1 KD (Extended Data Fig. 4e), in agreement with experimental data (Extended Data Fig. 3g). We next modelled the expected changes in ATP levels at different firing frequencies in both control and Letm1 KD neurons, exploring theoretical scenarios under which mitochondrial Ca2+ dynamics could modulate ATP levels. Although these simulations do not recapitulate the full complexity of biological systems, they provide insight into theoretical constraints that shape mitochondrial metabolism during activity. Our simulations showed that after a sufficient amount of time, different firing paradigms lead to ATP accumulation in Letm1 KD neurons (Fig. 2d,e), as observed experimentally during physiological firing in culture (Fig. 2a–c), suggesting that accumulation of ATP in Letm1 KD neurons is proportional to the history of activity experienced by the neuron. We next tested the theoretical role of PDP1 as an initiator of this metabolic acceleration by removing it from Letm1 KD neurons. In the absence of PDP1, Letm1 KD-mediated ATP accumulation was abolished (Fig. 2d,e), consistent with our experimental measurements (Fig. 2a–c). Together, these modelling and experimental results suggest that slowing down mitochondrial Ca2+ extrusion rates in neurons firing at physiological rates can be sufficient to increase their metabolism.
Mitochondrial metabolism generates ATP using pyruvate as the main carbon source. Increased presynaptic metabolism in Letm1 KD neurons, thus, should require corresponding increases in pyruvate import fluxes into mitochondria. Cytosol-to-mitochondria pyruvate flux can be measured using genetically encoded sensors for cytosolic pyruvate, such as the FRET sensor Pyronic93, in the presence of agents that acutely block mitochondrial pyruvate consumption. This paradigm generates cytosolic pyruvate accumulation at a pace that indirectly reflects pyruvate uptake into mitochondria19,93,94. We expressed Pyronic in wild-type and Letm1 KD primary neurons, blocked pyruvate import using sodium azide (a potent inhibitor of mitochondrial complex IV that stalls pyruvate import) and quantified pyruvate accumulation rates, showing that Letm1 KD neurons presented a significantly faster increase in the Pyronic ratio (Fig. 2f,g). We next simulated pyruvate consumption at different frequencies, which suggested that Letm1 KD neurons should present increased pyruvate consumption as a consequence of their increased metabolism (Fig. 2h). Together, these pyruvate flux measurements and simulations, although indirect, provide additional data suggesting that Letm1 KD neurons present increased metabolic rates.
Letm1 controls mitochondrial Ca2+ and neuronal metabolism in vivo
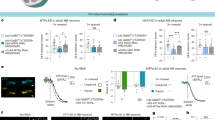
We next sought to establish in vivo the role of Letm1 in controlling neuronal metabolism in neurons and assess its putative impact on brain function and behaviour. Given that Letm1 is conserved across eukaryotes48 and is expressed in the fly brain, including MB neurons95, we first asked whether its function was conserved between rodents and flies. We expressed Mito4x-GCaMP6f in rat hippocampal neurons, knocked down endogenous Letm1 and re-expressed shRNA-resistant full-length Drosophila Letm1. We found that Drosophila Letm1 effectively rescued the impairment in mitochondrial Ca2+ efflux observed in rodent Letm1 KD neurons (Fig. 3a), suggesting functional equivalence between Drosophila Letm1 and its rat homologue.
a, Mitochondrial Ca2+ efflux (t1/2) in rat axonal mitochondria following stimulation (20 APs 20 Hz) in control, Letm1 KD neurons and Letm1 KD neurons expressing Drosophila Letm1. This experiment was performed in the same batch as Fig. 1f. P values were determined using a Kruskal–Wallis test (***P < 0.0003; n = 19 for control, n = 13 for Letm1 KD and n = 5 for Letm1 KD + fly Letm1, followed by Dunn’s multiple-comparisons test versus control. P values: Letm1 KD versus control, ***P = 0.0002; Letm1 KD + fly Letm1 versus control, NS, P > 0.9999). b, Mitochondrial Ca2+ levels were recorded in the vertical α-lobes of the MB of flies (tub-Gal80ts, c739-Gal4; UAS-mCarmine > UAS-Mito4x-LA-GCaMP6s) using two-photon microscopy as shown in the figure (white circle). These lobes consist of axonal projections of the MB neurons. Scale bar, 20 μm. c, Mitochondrial Ca2+ levels shown as the ratio of Mito4x-LA-GCaMP6s to mCarmine in control and Letm1 KD flies, measured 0.5–1.5 h after exposure to either 1× paired or unpaired protocol. P value for control flies was determined using an unpaired t-test (NS, P = 0.06; n = 16, 17, t31 = 1.955). P value for Letm1 KD flies was determined using an unpaired t-test (**P = 0.005; n = 16, 16; t30 = 3.038). Each group is shown normalized to the corresponding unpaired condition. d, Rates of cytosolic pyruvate accumulation after mitochondrial metabolism is blocked using sodium azide (black arrow) in control and Letm1 KD flies subjected to either 1× paired or unpaired protocol. Traces correspond to the mean ± s.e.m. of changes in CFP/YFP ratio (ΔR) normalized to the initial signal (R0) of Pyronic (ΔR/R0). e, Rate of pyruvate accumulation in MB neurons of control (tub-Gal80ts,c739;UAS-pyronic/+) and Letm1 KD flies (tub-Gal80ts,c739;UAS-pyronic > Letm1 RNAi no. 1) undergoing 1× paired or unpaired protocols. P value for control flies was determined using a two-tailed Mann–Whitney U-test (NS, P = 0.2973; n = 9,9; U = 28). P value for Letm1 KD flies was determined using a two-tailed Mann–Whitney U-test (**P = 0.0048; n = 11,10; U = 16). f,g, Letm1 RNAi flies were used to test memory. Genotypes include control 1 (dark grey, tub-Gal80ts,c739/+), control 2 (light grey, +/Letm1 RNAi no. 1) and Letm1 KD (tub-Gal80ts,c739>Letm1 RNAi no. 1). f, MTM tested 3 h after 1× conditioning in control and Letm1 KD flies. Group differences were assessed with a one-way analysis of variance (ANOVA; n = 12 per group). ANOVA: F(2,33) = 0.6233, P = 0.5424. g, LTM tested 24 h after 1× conditioning in control and Letm1 KD flies was increased in Letm1 RNAi flies (one-way ANOVA F(2,51) = 4.338, P = 0.0182, n = 18, 18, 18, Tukey’s post-test control 1 versus Letm1 KD, *P = 0.0387; control 2 versus Letm1 KD, *P = 0.0338, control 1 versus control 2, NS, P = 0.9983). h, LTM tested 24 h after 1× conditioning in control, Letm1 KD, PDP KD and Letm1;PDP dual KD flies. Increase in LTM by Letm1 KD (RNAi line no. 2, GD2208) was not observed if PDP RNAi is co-expressed. Genotypes include from left to right: tub-Gal80ts,c739/+; +/Letm1 RNAi no. 2, PDP RNAi; tub-Gal80ts,c739/Letm1 RNAi no. 2; tub-Gal80ts,c739/PDP RNAi and; tub-Gal80ts,c739/Letm1 RNAi no. 2, PDP RNAi. Group differences were assessed with a one-way ANOVA (n = 12 per group). ANOVA: F(4,55) = 5.141, P = 0.0014. Tukey’s post-test: tub-Gal80ts,c739/Letm1 RNAi no. 2 versus tub-Gal80ts,c739/+ *P = 0.0113; versus +/Letm1 RNAi no. 2, PDP RNAi **P = 0.0017; versus tub-Gal80ts,c739/PDP RNAi **P = 0.0059, versus tub-Gal80ts,c739/Letm1 RNAi no. 2, PDP RNAi *P = 0.0459. Data are represented as the mean ± s.e.m. See also Extended Data Figs. 5 and 6.
Memory formation following classical Pavlovian aversive olfactory conditioning in D. melanogaster96,97,98 has emerged as a key example of a cognitive function that is modulated by neuronal metabolism19,23,99. Given that LTM formation upregulates mitochondrial metabolism of MB neurons after associative training19,23,99, we next asked whether reducing Letm1 levels in α/β neurons could lead to increased mitochondrial Ca2+ retention and further enhance their metabolic state after training, thereby reshaping the dynamics of memory formation at the behavioural level. Following the paired delivery of an odour and an electric shock, flies form an avoidance memory that is encoded in neurons of the MB, a major integrative centre of insect brains that is considered functionally analogous to the mammalian hippocampus100,101. In wild-type flies, a single pairing of odour and shock (1×) forms a memory lasting only a few hours96. However, it is only when repeated sessions of odour and shock are presented spaced in time that LTM of the aversive olfactory stimulus is formed96. Olfactory conditioning activates mitochondrial metabolism in a specific group of neurons called α/β neurons19,102,103. However, this metabolic activation is complexly connected with how long the memory lasts. A short 1× training session causes a quick rise in pyruvate flux to mitochondria23 that is temporary and no longer detectable 3 h after conditioning99. The 5× spaced training, in contrast, leads to an extended period of increased mitochondrial pyruvate flux observable up to 8 h after conditioning99. This long-lasting upregulation of pyruvate usage in MB neurons reflects an early acceleration of mitochondrial metabolism that enables LTM19,99.
To explore the role of Letm1 in neuronal metabolism in vivo, we first studied how reducing Letm1 expression in MB neurons impacts mitochondrial Ca2+ retention after 1× training (1× paired training) or after an unpaired protocol in which the electric shock is temporally dissociated from the odour (1× unpaired training; Extended Data Fig. 5a and Methods). Previous work in Drosophila neurons showed that using Ca2+ sensors with low-micromolar affinity facilitates measuring mitochondrial Ca2+ dynamics104 despite this not being the case in cultured mammalian neurons13. To robustly measure mitochondrial Ca2+ in vivo in flies, we generated a low-affinity (LA) variant of GCaMP6s (LA-GCaMP6s) by introducing the single point mutation p.AspD362Ile that previously resulted in a low-micromolar-affinity shift for GCaMP3 (ref. 62). We measured the biophysical properties of LA-GCaMP6s in purified protein at room temperature and 37 °C and found that it presented an EC50 of ~6 μM in both cases while preserving the dynamic range of GCaMP6s (Supplementary Table 2 and Extended Data Fig. 5b,c). Mitochondrial expression of this variant (Mito4x-LA-GCaMP6s) enabled detecting activity-driven mitochondrial Ca2+ responses in axons of cultured neurons and the slower mitochondrial Ca2+ extrusion of Letm1 KD neurons (Extended Data Fig. 5d,e). We generated flies that co-expressed Mito4x-LA-GCaMP6s and the fluorescent protein mCarmine in α/β MB neurons under the control of the same GAL4 driver (Fig. 3b), and quantified mitochondrial Ca2+ levels in the α-lobes by measuring the fluorescence ratio of these two indicators. Training-induced changes in mitochondria Ca2+ levels were determined by comparing flies submitted to paired or unpaired protocols. In control flies, no learning-induced calcium elevation was detected either immediately or 3 h after conditioning (Fig. 3c and Extended Data Fig. 5f). However, Letm1 KD resulted in an increase in mitochondrial Ca2+ levels just after training (Fig. 3c). This effect was transient, as mitochondrial Ca2+ levels returned to baseline by 3 h after training (Extended Data Fig. 5f). These results show that Letm1 KD enhances mitochondrial Ca2+ retention in MB neurons immediately after associative training, in agreement with data in cultured neurons showing increased mitochondrial Ca2+ retention after neuronal activity.
To study whether transient changes in mitochondrial Ca2+ result in longer-lasting changes in metabolism in vivo, we next expressed Pyronic in α/β neurons of the adult Drosophila MB. Using two-photon microscopy, we first measured mitochondrial pyruvate usage in vivo in α/β neurons of control adult flies after 1× training or after an unpaired protocol, as previous work indicated that differences in experience-driven pyruvate accumulation reflected changes in mitochondrial metabolism specifically19. Similarly to rodent neurons in vitro (Fig. 2f,g), we quantified pyruvate accumulation rates in vivo following blocking mitochondrial pyruvate import, as a readout of cytosol-to-mitochondria pyruvate flux19 (Fig. 3d). We confirmed that 3 h after 1× training, mitochondrial pyruvate flux was not accelerated in α/β neurons of wild-type flies (Fig. 3d), as previously reported99. However, we found that Letm1 KD flies that were exposed to 1× paired conditioning presented a clear upregulation in cytosol-to-mitochondria pyruvate flux by ~75% when compared to Letm1 RNAi flies that received an unpaired protocol (Fig. 3d,e). These results suggest that reducing Letm1 expression in MB neurons increases neuronal metabolism in vivo after associative training.
Letm1 controls fly LTM through Ca2+-mediated upregulation of mitochondrial metabolism
A lack of sustained metabolic increases in MB neurons after associative olfactory training precludes the success of LTM formation19,99. This concept presents two major predictions with respect to Letm1 function: (1) Letm1 KD flies, which already present a marked increase in MB metabolism after 1× training (Fig. 3d,e), should be capable of forming LTM in these conditions; and (2) the absence of Letm1 should not modulate other shorter-lived types of memories that do not require long-lasting adaptations in metabolism, such as middle-term memory (MTM)19,99. To examine this hypothesis, we first measured MTM 3 h after 1× training and found no significant behavioural differences between wild-type and Letm1 KD flies, as expected (Fig. 3f). We validated this result using a second non-overlapping Letm1 RNAi fly line (Extended Data Fig. 6a). Using each of the Letm1 KD fly lines, we next examined LTM 24 h after 1× conditioning and found robust LTM formation exclusively when the expression of Letm1 was reduced (Fig. 3g and Extended Data Fig. 6b), indicating Letm1 has a functional role in modulating LTM. Using both RNAi lines, we confirmed that no increase in memory was observed when Letm1 RNAi was not induced (Extended Data Fig. 6c) and that expression of either of the two Letm1 RNAi constructs did not alter the flies’ innate avoidance responses to odour or electric shock (Extended Data Fig. 6d,e). When subjected to classical 5× space training protocols, Letm1 KD flies formed normal LTM as compared to genotypic controls (Extended Data Fig. 6f). Lastly, using quantitative PCR, we validated that both RNAi were able to reduce the expression of Letm1 in adult MB neurons (Extended Data Fig. 6g). Altogether, these results show that reducing the expression of Letm1 in adult MB α/β neurons facilitates associative LTM in conditions in which wild-type flies do not form LTM.
Since Letm1 KD controls both mitochondrial Ca2+ retention and neuronal metabolism, we examined whether the facilitation in LTM induced by Letm1 KD is driven by Ca2+-mediated upregulation of metabolism by PDP. To test this idea, we investigated 24-h memory formed after 1× training in flies expressing both PDP and Letm1 RNAi in MB neurons using a previously validated PDP RNAi19. Consistent with previous results, Letm1 RNAi flies presented robust memory performance 24 h after 1× conditioning, whereas the dual Letm1;PDP RNAi flies presented low memory scores indistinguishable from genotypic controls or PDP RNAi flies (Fig. 3h), showing that the absence of PDP hinders Letm1 KD-mediated LTM facilitation. These results indicate that disrupting the ability of mitochondria to modulate metabolism in response to Ca²⁺ eliminates the capacity of Letm1 to modulate LTM, thus suggesting that Letm1 KD allows LTM formation after 1× conditioning by regulating mitochondrial Ca2+ homeostasis.
Letm1 modulates LTM across species
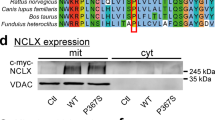
Given that the function of Letm1 as a mitochondrial Ca2+ exporter is conserved between flies and rodents (Fig. 3a), we next investigated whether Letm1 deficiency could control associative olfactory LTM in mice. To examine this hypothesis, we leveraged the miR-30 system105,106 to design viral constructs that express both mRFP1 and control or Letm1-specific shRNA sequences under the same promoter exclusively in excitatory principal neurons (Extended Data Fig. 7a). As the hippocampus acts as a major integrative centre of sensory information in rodents100,101, we bilaterally injected adeno-associated viruses (AAVs) carrying these constructs into the dorsal hippocampus of mice (Fig. 4a,b). We confirmed by western blot analysis of dissected hippocampi that this approach reduced Letm1 levels by ~80% (Fig. 4c). Notably, this genetic deletion did not impact locomotor activity as no significant differences were observed between Letm1 KD mice and controls (Extended Data Fig. 7b–e). However, lacking Letm1 for weeks should favour mitochondrial Ca2+ retention during the naturally occurring firing events in the hippocampus in vivo, favouring the over-activation of PDP1 and the dephosphorylation of PDHc in this population of neurons. Thus, we measured the phosphorylation state of PDH from dissected hippocampi of Letm1 KD mice and confirmed a strong reduction in PDH phosphorylation (Fig. 4d), indicating that Letm1 modulates the metabolic state of hippocampal neurons in vivo.
a,b, Visualization of the hippocampal brain region injected with the corresponding viral constructs, showing mRFP1 fluorescence in red and DAPI in blue. Scale bars, 500 μm. c, Left, representative western blot showing levels of Letm1 and β-actin proteins in lysates from dissected hippocampi of control and Letm1 KD mice. Right, quantification of Letm1 expression relative to controls. P value was determined using a two-tailed Wilcoxon signed-rank test (**P = 0.0078; n = 8; W = −36). d, Left, representative western blot showing levels of phosphorylated PDH (pPDH-S293) and total PDH in lysates from dissected hippocampi of control and Letm1 KD mice. Right, quantification of pPDH-S293 intensity relative to total PDH. P value was determined using a two-tailed Wilcoxon signed-rank test (**P = 0.0078; n = 8; W = 36). e, Schema showing the aversive olfactory conditioning paradigm in mice for MTM and LTM. f, MTM in control and Letm1 KD mice shows no difference between conditions. P value for MTM was determined using a two-tailed Mann–Whitney test (NS, P = 0.9395; n = 8 for control and n = 8 for Letm1 KD; U = 31). g, LTM in control and Letm1 KD mice shows that memory is only preserved in mice expressing Letm1 KD in the hippocampus. P value for LTM was determined using a two-tailed Mann–Whitney test (*P = 0.0303; n = 8 for control and n = 8 for Letm1 KD; U = 11.5). Data are represented as the mean ± s.e.m. See also Extended Data Fig. 7.
Following paired delivery of odour and aversive stimulus, the hippocampus is required for the consolidation of olfactory aversive memories107,108,109. We thus implemented an olfactory aversive conditioning protocol, outlined in Fig. 4e (also see Methods), to evaluate whether neuronal Letm1 can modulate olfactory LTM formation in rodents. After 2 days of habituation to water deprivation conditions, stereotaxically injected mice were exposed to two distinct pairing sessions: on day 3, mice were exposed to an odorized water bottle (CS+) paired with an injection of the unpleasant chemical lithium chloride (LiCl), which causes gastric malaise in rodents and generates aversion. On day 4, mice were exposed to a different odorized water bottle (CS−) coupled with a neutral saline (NaCl) injection. Lastly, on day 5 and day 15, two-choice tests were performed to assess the olfactory memory performance of both control and Letm1 KD mice at middle-term or long-term stages. We first measured MTM in mice 1 day after conditioning, and found that both wild-type and Letm1 KD mice presented robust MTM (Fig. 4f and Extended Data Fig. 7f). This result is consistent with findings in flies, which also exhibit similar levels of MTM following conditioning (Fig. 3f). However, when we evaluated LTM after 10 days, we found that memories were preserved exclusively in Letm1 KD mice (Fig. 4g and Extended Data Fig. 7g), indicating a functional role of Letm1 in regulating mouse olfactory LTM. We found no differences between control and Letm1 KD mice either in water consumption capacity (Extended Data Fig. 7h) or in the effectiveness of each odour as CS+ (Extended Data Fig. 7i,j). These results indicate that the loss of function of Letm1 in the mouse hippocampus increases neuronal mitochondrial metabolism and facilitates associative LTM. Collectively, our results observed across species suggest an evolutionarily conserved role for Letm1 in shaping experience-induced adjustments in neuronal metabolism within integrative associative memory centres, thereby controlling the formation of long-term olfactory memories.
Discussion
Expensive energy usage in neurons must be limited to avoid unnecessary overconsumption of fuels in the brain that could otherwise be useful for survival. During neuronal activity, synapses synthesize the exact levels of energy that are consumed during each firing event, without underproducing or overproducing ATP. While the work of several laboratories has identified how mitochondrial metabolism is upregulated on demand in activated neurons to preserve the metabolic integrity of synapses8,10,29,35,83,110,111, the importance and the molecular identity of mechanisms slowing down mitochondrial metabolism after firing have remained elusive.
In this study, we found that the efficiency of mitochondrial Ca2+ extrusion is actively tuned during neuronal activity to control mitochondrial metabolism and the metabolic state of neurons. Our observations indicate that Letm1, together with NCLX49,51, is essential in controlling this process. Our results suggest that Letm1 and NCLX may have differing modes of action, as indicated by two lines of evidence. First, ablating Letm1 does not alter resting mitochondrial Ca2+ levels, while impairing NCLX function increases mitochondrial Ca2+ at rest50,53, suggesting NCLX also controls constitutive Ca2+ efflux. This result likely reflects the higher affinity for Ca2+ of NCLX compared to Letm1 (refs. 47,112,113) and its mode of activation. While NCLX continuously exports Ca2+ through the thermodynamically favourable Na+ gradient and the membrane-potential exchange cycle53, the EF-hand domains in Letm1 enable its activation selectively during mitochondrial Ca2+ increases, such as those seen during neurotransmission, suggesting that Letm1 function in neurons is activity driven and it does not participate in mitochondrial Ca2+ efflux in resting conditions.
Despite that the expression of Letm1 is ubiquitous63 and its role as a Ca2+/H+ exchanger has been already shown in several different tissues45,46,48, the precise mechanism of Letm1-mediated mitochondrial Ca2+ exchange remains an active area of debate. Letm1 was initially proposed to drive 1Ca2+/1H+ electrogenic transport45. However, to drive electrogenic Ca2+ efflux, Letm1 would require either matrix Ca2+ in the high micromolar range or pronounced reductions in mitochondrial membrane potential114,115. Our results do not support such a 1Ca2+/1H+ stoichiometry because (1) during physiological activity mitochondrial Ca2+ increases up to ~425 nM, far from high micromolar levels, and (2) we observe robust axonal mitochondrial Ca2+ uptake, which suggests that axonal mitochondria are not depolarized as it would be required for electrogenic efflux. Alternatively, our observations support that Letm1 acts as an electroneutral Ca2+/H+ antiporter with a 1:2 exchange ratio, as proposed by work in reconstituted liposomes and non-neuronal cells45,46,47. Letm1 has also been proposed to exchange K+ for H+116,117. Although the affinity for K+ appears to be markedly less than for Ca2+47, future work should be aimed at clarifying the role of Letm1 in axonal mitochondrial K+ dynamics and dissecting whether it could have any role in modulating metabolism. On the other hand, by acting as a Ca2+/H+ exchanger, Letm1 should also transport H+ ions into the mitochondrial matrix. Letm1 KD neurons present accumulation of presynaptic ATP and increased pyruvate usage rates, and it is likely that the actions of both Ca2+ and H+ together contribute to such activity-dependent regulation of mitochondrial metabolism in neurons.
While previous work showed that mitochondrial Ca2+ enhances ATP synthesis only within a limited range39, we estimate that neuronal activity increases mitochondrial Ca2+ moderately up to 400–500 nM, far from the high mitochondrial Ca2+ levels observed to inhibit OXPHOS39. Such increases are also in the right range to activate PDP1 efficiently, as in vitro measurements of rat PDP1 activation show a half-maximal effective concentration (EC50) for Ca2+ of 400 nM118, supporting that the coupling between cytosolic and mitochondrial Ca2+ can increase mitochondrial metabolism. We used computational modelling to explore the possible theoretical relationships between mitochondrial Ca2+ transients and the synergistic modulation of the multiple complex metabolic processes occurring during neuronal activity. This approach suggested that a decrease in mitochondrial Ca2+ influx rates could be sufficient to modulate mitochondrial metabolism, and that the first step of acceleration by increasing PDH activity is theoretically well positioned to gate the acceleration of the subsequent reactions. Interestingly, our results in vivo in flies show that increases in mitochondrial Ca2+ after training are relatively brief, implying that PDP1 activation should be short-lived and conclude within 3 h after training. However, metabolic changes in MB neurons persist for a longer duration, as demonstrated by elevated pyruvate utilization several hours after training. Although PDP1 activation itself may be transient, the resulting PDHc dephosphorylation is likely to persist until actively reversed by pyruvate dehydrogenase kinase. Thus, we propose that brief mitochondrial Ca2+ elevations enable sustained increases in mitochondrial metabolism, effectively integrating a brief aversive experience lasting only seconds into metabolic responses maintained for hours. Such prolonged metabolic activation is well positioned to support the long-lasting metabolic demands of LTM formation.
Electrical and chemical signalling within and between neurons impose considerable metabolic challenges that, if not properly met, lead to a decline in cognitive performance119,120,121. Conversely, experimental increases in brain fuel availability markedly improve the cognitive abilities of rodents and humans22,122,123,124. These observations, combined with our data demonstrating that removal of Letm1 from memory centres facilitates LTM across species, suggest that the metabolic state of neurons could act as a master modulator of circuit function by enabling or limiting energy expenditure. However, while a robust facilitation in memory formation could be seen as beneficial, there are two main drawbacks that probably counteracted the selection of this mechanism during evolution: (1) forming memories based on associations occurring a single time does not necessarily lead to increased survival, as the aversive and innocuous stimuli may simply coincide in a random fashion, thus providing no useful information for future aversive behaviours; and (2) encoding memories comes with a high metabolic cost and should only occur in conditions in which it is necessary20. Thus, evolution has likely unified the cognitive and energetic constraints that are imposed on memory, limiting both energetic costs and useless memory associations unless they are ecologically relevant, as for example when the associative event has occurred several times.
In our study, we use mitochondrial ATP production and pyruvate consumption as parameters for measuring the metabolic state of neurons. However, these two specific metabolic factors alone are unlikely to fully account for the observed changes in memory performance. Indeed, increases in mitochondrial metabolism should influence a myriad of neuronal and metabolic pathways, including reactive oxygen species signalling125,126, neuronal excitability13,127, metabolite levels128, protein synthesis and/or gene expression128. These changes may occur as a consequence of alterations in post-translational modifications linked to metabolism, such as acetylation, which controls chromatin plasticity and memory129,130,131. Moreover, it was recently shown that calcium/calmodulin-dependent kinase II (CamKII), a key player in learning and memory132, regulates the fidelity and frequency of mitochondrial Ca2+ transients in response to neuronal activity69. CamKII, by shaping mitochondrial Ca2+ entry, could be a contributing factor to the enhanced memory observed in Letm1 KD animals. In our study, we used KD rather than knockout approaches, meaning that residual Letm1 protein remains. Future investigations should assess whether the observed memory benefits depend on the degree of Letm1 depletion, perhaps using conditional Letm1 mice. While off-target effects of KD cannot be fully excluded, we mitigated this risk by using two independent RNAi constructs and performing rescue experiments. Future investigations, including omics approaches, will aim to unravel these complex molecular mechanisms and explore their role in modulating neuronal function across various levels, from synaptic interactions and cell-type-specific responses to broader circuit dynamics and behavioural outcomes.
In the future, further understanding the role of mitochondrial calcium dynamics in neuronal metabolism may be particularly relevant in the context of memory disorders. Bioenergetic dysfunction is a prominent feature in early stages of memory disorders such as Alzheimer’s disease, in which dysfunctional mitochondria lead to debilitated metabolic states and consequently neurodegeneration133,134,135. In particular, as impaired mitochondrial metabolism alters both presynapse function10,32,136,137,138 and memory19,22,23,99,137, future experiments could explore whether reducing mitochondrial Ca2+ efflux could be used to rescue bioenergetic defects in early Alzheimer’s disease animal models, as Letm1 KD neurons present larger numbers of synapses and mitochondria, and facilitate memory formation. Overall, this work provides a theoretical and experimental framework to better understand the importance of the tight coupling between mitochondrial metabolism and neuronal function in health and disease, and define new molecular mechanisms controlling bioenergetics of neurotransmission, circuit physiology and behaviour across species.
Methods
Animals
Rodents
The rats used in the study to prepare primary cultures were either male or female, and were of the Sprague–Dawley strain Crl:CD(SD), which are bred worldwide by Charles River Laboratories according to the International Genetic Standardization programme. The experiments conducted in the study at the Paris Brain Institute were conducted at the PHENO-ICMice facility and adhered strictly to the guidelines outlined in the European Directive 2010/63/EU and the French Decree n° 2013-118 for the protection of animals used for scientific purposes. The glutamate uncaging experiments using rat pups (Fig. 1g,h and Extended Data Fig. 1l–o) were performed in accordance with the Max Planck Florida Institute for Neuroscience IACUC regulations (protocol number 22-005). Rats were fed with a standard rat breeding diet (V132400, SSNIFF). C57BL/6J male mice (Charles River, France) of 7–8 weeks of age were used for behavioural studies to facilitate direct comparison of our results with our previous work in hippocampal memory that have predominantly used male mice139,140. Animals were grouped housed and maintained in an environment in which both temperature (20–24 °C) and humidity (40–70%) were controlled. Mice were maintained under a 12-h light–dark cycle with food and water available ad libitum. All the experiments were performed during the dark phase of the light–dark cycle by a trained observer who was blind to experimental conditions. Animal procedures were conducted at the Parque de Investigación Biomédica de Barcelona and were in accordance with the standard guidelines of the European Directive on the protection of animals used for scientific purposes (2010/62/EU) and approved by the Animal Ethics Committee of the Parque de Investigación Biomédica de Barcelona. Mice were fed with the complete breeding vegetal diet for rats, mice and hamsters (SDS RM3 (P), Fibers for Life JRS).
Fly strains
Flies (D. Melanogaster) were maintained on a standard medium consisting of yeast, cornmeal and agar at 18 °C and 60% humidity under a 12-h–12-h light–dark cycle. The UAS-LETM1-RNAi lines correspond to HMS01644 (RNAi no. 1) from the Bloomington Drosophila Stock Center and GD2208 (RNAi no. 2) from the Vienna Drosophila Resource Center; the UAS-PDP-RNAi corresponds to GD31661 from the Vienna Drosophila Resource Center. The double RNAi line Letm1 no. 2, PDP was generated in this study using the above-mentioned RNAi lines. The tub-Gal80ts, c739-gal4; UAS-Pyronic was generated in this study from the previously described lines: UAS-Pyronic in Plaçais et al.19 and tub-Gal80ts, c739-gal4 in Turrel et al.141. To limit UAS/GAL4-mediated expression exclusively to the adult stage, the TARGET system was used142. The GAL4 activity was inhibited at 18 °C by a thermosensitive version of GAL80 ubiquitously expressed under the control of the tubulin promoter (tubulin-GAL80ts), as previously reported19. The GAL4 activity was released by transferring adult flies to 30 °C for 2–3 days allowing the expression of the UAS-transgene (RNAi and/or Pyronic FRET sensor). For the generation of the UAS-Mito4x-LA-GCaMP6s Drosophila line, the CMV-Mito4x-LA-GCaMP6s (Addgene, 243810) was cloned into a pJFRC-MUH plasmid (Addgene, 26213)143 using Gibson Cloning. The resulting construct was verified by sequencing. For the generation of the UAS-mCarmine Drosophila line, the mCarmine pcDNA3 plasmid (Addgene, 109486)144 was digested by EcoRI and BamHI. The resulting 746-bp fragment was purified by electrophoresis and cloned into a pJFRC-MUH plasmid (Addgene, 26213)143. The resulting construct was verified by sequencing. The molecular cloning was outsourced to RD-Biotech, France. Transgenic fly strains were obtained by site-specific embryonic injection of the resulting vector in the VK00005 landing site (third chromosome), which was outsourced to Rainbow Transgenic Flies. The tub-Gal80ts, c739-gal4; UAS-mCarmine was generated in this study from the above-described UAS-mCarmine line and tub-Gal80ts, c739-gal4 in Turrel et al.141.
Primary rat co-culture of postnatal neurons and astrocytes
All imaging experiments were performed in primary co-cultures of neurons and astrocytes obtained from the rat hippocampus. Postnatal day 0 (P0) to P2 rats of mixed gender were euthanized and their brains were dissected in a cold HBSS-FBS (1× HBSS, 20% FBS) solution to isolate the hippocampus, excluding the dentate gyrus. Hippocampi were washed with HBSS (Thermo Fisher Scientific, 14185045) and digested in a trypsin-digestion solution containing DNase I (Merck, D5025) for 5 min. Trypsin (Merck, T1005) was neutralized by the addition of HBSS-FBS solution, following which the tissue was washed several times with HBSS solution. The tissue was then transferred to a dissociation (1× HBSS, 5.85 mM MgSO4) solution and was dissociated into single cells by gentle and repeated triturations. Next, as a washing step, the cells were pelleted by centrifugation and resuspended in HBSS solution. The cells were then pelleted again and resuspended in warmed plating media composed of MEM (Thermo Fisher Scientific, 51200038) supplemented with 20 mM glucose, 0.1 mg ml−1 transferrin (Merck, 616420), 1% Glutamax (Thermo Fisher Scientific, 35050061), 24 μg ml−1 insulin (Merck, I6634), 10% FBS (Thermo Fisher Scientific, 10082147) and 2% N-21 (Bio-Techne, AR008). Finally, the cells were counted, and 38,000 cells were plated into 4.7-mm-diameter cloning cylinders attached onto coverslips coated with poly-ornithine (Merck, P3655). Once the supporting glial cell layer was established 2–4 days after plating, the cells were shifted to ‘feeding media’ composed of MEM supplemented with 20 mM glucose, 0.1 mg ml−1 transferrin, 1% Glutamax, 24 μg ml−1 insulin, 5% FBS, 2% N-21 and 4 μM cytosine β-d-arabinofuranoside (Merck, C6645). Cultures were incubated at 37 °C in a 95% air/5% CO2 humidified incubator. At 5–8 days in vitro (DIV), neurons were transfected using the calcium phosphate method based on a previously published protocol145. In brief, transfection was initiated by changing the medium to basal Advanced DMEM (Thermo Fisher Scientific, 12634-010) without any supplements. The cells were then returned to the incubator for 1 h to equilibrate in the new media. During this time, the DNA–calcium (Ca2+)–phosphate (PO4) mixture was prepared according to a previously published recipe146. The mixture was incubated for 30 min to allow formation of the DNA–Ca2+–PO4 precipitate, which was then added to the cells ~1 h after media change. The cells were incubated with the precipitate for 1 h after which the medium was changed back to feeding medium. The cells were maintained in the incubator up to 14–21 DIV before imaging. These mixed cultures are composed of 12.4% neurons, 75% astrocytes and 12.6% of cells that could be identified as neither neurons nor astrocytes (Extended Data Fig. 1a).
For cultures used in glutamate uncaging experiments shown in Fig. 1g,h and Extended Data Fig. 1l–o, conditions were as follows: hippocampal regions were dissected in artificial cerebrospinal fluid containing: 124 mM NaCl, 5 mM KCl, 1.3 mM MgSO4:7H2O, 1.25 mM NaH2PO4:H2O, 2 mM CaCl2, 26 mM NaHCO3 and 11 mM glucose (stored at 4 °C) and stored in hibernate E buffer (BrainBits, stored at 4 °C). Dissected hippocampi were dissociated using the Papain Dissociation System (Worthington Biochemical Corporation, stored at 4 °C) with a modified manufacturer’s protocol. In brief, hippocampi were digested in papain solution (20 units of papain per ml in 1 mM l-cysteine with 0.5 mM EDTA) supplemented with DNase I (final concentration 95 units per ml) and shaken for 30–60 min at 37 °C at 900 rpm. Digested tissue was triturated and set for 3 min, following which the supernatant devoid of tissue chunks was collected. The supernatant was centrifuged at 300g for 5 min and the pellet was resuspended in resuspension buffer (1 mg of ovomucoid inhibitor, 1 mg of albumin and 95 units of DNase I per ml in EBSS). The cells were forced to pass through a discontinuous density gradient formed by the resuspension buffer and the Ovomucoid protease inhibitor (10 mg per ml) with bovine serum albumin (BSA; 10 mg per ml) by centrifuging at 70g for 6 min. The final cell pellet devoid of membrane fragments was resuspended in Neurobasal-A medium (Gibco, stored at 4 °C) supplemented with Glutamax (Gibco, stored at −20 °C) and B27 (Gibco, stored at −20 °C). Cells were plated on poly-d-lysine-coated coverslips mounted on MatTek dishes at a density of 30,000–50,000 cells per cm2. Cultures were maintained at 37 °C and 5% CO2 with feeding every 3 days using the same medium until transfection. Transfections were performed 12 days after plating by magnetofection using Combimag (OZ biosciences, stored at 4 °C) and Lipofectamine 2000 (Invitrogen, stored at 4 °C) according to the manufacturer’s instructions.
Gene constructs
Constructs to specifically knock down Letm1 expression in primary cultures of rat neurons were designed using the Genetic Perturbation Platform (Broad Institute) and cloned into various versions of the pLKO cloning vector as indicated in Supplementary Table 1. For most imaging experiments the BFP expression version of the Letm1 KD plasmid construct was used to confirm double transfection through fluorescence. In the case of Pyronic experiments, the miRFPnano version was used to avoid spectral overlap with the sensor. The target sequence used for shRNA KD of Letm1 in rats (pLKO-U6-sh1-Letm1(rat)-hPGK-mTagBFP2; Addgene, 212664) was 5′-CCTTCCAGAAATTGTGGCAAA-3′. Rat Letm1 presents three very similar isoforms (Canonical, ENSRNOT00000099794.2, Letm1-202, 760 amino acids long; ENSRNOT00000022540.8 Letm1-201, 739 amino acids long; ENSRNOT00000147310.1, Letm1-203, 734 amino acids long). Our target sequence is located in exon 8, which is identical in all three isoforms. The target sequence against rat Pdp1 for shRNA KD was 5′-ATTATTGCCTACAGCATGGCG-3′, which was expressed in the same pLKO vector under the H1 promoter together with the Letm1 shRNA under the U6 promoter. For the rescue experiments, Mito4x-GCaMP6f was cloned into a plasmid with a CaMKII promoter and an IRES2 sequence to express two coding sequences. This was done by the Gibson cloning method using NEBuilder HiFi DNA Assembly Master Mix (E2621, NEB). Individual rat Letm1 and Drosophila Letm1 protein-coding sequences were obtained from the Ensembl genome browser, and plasmids containing these sequences were synthesized using Invitrogen GeneArt Gene Synthesis. Site-directed mutagenesis using the Q5 Site-Directed Mutagenesis Kit (NEB) was performed to inactivate the Ca2+ coordinating amino acids in the EF-hand of Letm1. The aspartate (D) amino acids at positions 276 and 280 were replaced by alanine. We designed the Letm1–mRFP1 construct to express rat Letm1, followed by a short linker (RPVVAV) and mRFP1 and synthesized this construct at GeneArt (Thermo Fisher Scientific), where it was cloned into pcDNA3.1 to be expressed under the CMV promoter. In the multiple cloning site located after the IRES2 sequence, the synthesized shRNA-resistant Letm1 coding sequences were cloned using restriction enzyme cloning, generating the following constructs: CamKII-Mito4x-GCaMP6f-IRES2-MCS, CamKII-Mito4x-GCaMP6f-IRES2-rat-Letm1, CamKII-Mito4x-GCaMP6f-IRES2-rat-Letm1-ΔEF-hand and CamKII-Mito4x-GCaMP6f-Drosophila-Letm1. For the mouse behaviour experiments, the microRNA encoding plasmids and AAVs—pAAV-Camk2a(short)-mRFP1.Letm1[miR-30-shRNA]-WPRE, pAAV-Camk2a(short)-mRFP1.Scramble[miR-30-shRNA]-WPRE and GCaMP6s p.AspD362Ile—were designed in the lab and generated by VectorBuilder (Supplementary Table 1). Mouse Letm1 has only one major protein-coding isoform (ENSMUST00000005431.6 Letm1-201, 738 amino acids long). Other annotated mouse transcripts are truncated or non-coding. Therefore, the target sequence is present in the main mouse isoform, and a single western blot band corresponding to this isoform is expected.
Synthetic DNA oligonucleotides were purchased from Integrated DNA technologies. Q5 high fidelity DNA polymerase (New England Biolabs, NEB) was used for all PCR amplifications. Isothermal assembly reactions were performed with a NEBuilder HiFi kit (NEB). Small-scale DNA isolation was performed with QIAprep Spin Miniprep Kit (Qiagen). The pRSET vector backbone was acquired from Life Technologies. Inserts and vector backbones were amplified by PCR amplification. Vector backbones and inserts were assembled by isothermal assembly with 10–30-base-pair overlap, and sequence verified by Sanger sequencing (Azenta Life Sciences) or by nanopore full-plasmid sequencing (Plasmidsaurus). pRSET plasmids encoding GCaMP6s147 and GCaMP6-150 (ref. 62) were used from an in-house source (Janelia Research Campus). Other plasmids used in this study, which have been previously described, are listed in Supplementary Table 1.
Protein expression and purification of GCaMP6s, GCaMP-150 and GCaMP6s p.AspD362Ile
For expression and purification of proteins, T7 express cells (NEB) were transformed with pRSET plasmids encoding the protein of interest. The bacteria were grown in auto-induction media using the Studier method148 with antibiotics at 30 °C for 48 h with shaking at 200 rpm. Cell pellets were collected by centrifugation, lysed in Tris-buffered saline (TBS; 19.98 mM Tris, 136 mM NaCl, pH 8.0), with n-octyl-β-d-thioglucopyranoside (5 g l−1)). Aggregations were disrupted by sonication and the lysate cleared by centrifugation. Protein purification was performed on an N-terminal poly-histidine (His6) tag using HisPur Ni-NTA resin (Thermo Fisher Scientific), according to the manufacturer’s recommendations. Purified proteins were buffer exchanged into TBS using Amicon concentration filters (Merck). Protein aliquots were stored at 4 °C.
Calcium titrations in purified protein
To determine the calcium affinity and cooperativity of the calcium indicators, calcium titrations were performed in a buffer system made from ethylene glycol tetraacetic acid (EGTA) and Ca2+/EGTA from the Calcium Calibration Buffer Kit no. 1 (Invitrogen) or nitrilotriacetic acid (NTA) and Ca2+/NTA prepared using the pH titration method described by Tsien and Pozzan149. The solutions were mixed in specific ratios to generate known free calcium concentrations. The free Ca2+ concentration was calculated assuming the dissociation constant of EGTA for Ca2+ to be 150 nM at 22 °C and pH 7.2, and assuming the dissociation constant of NTA for Ca2+ to be 67 µM. In total, 2 µl of purified protein at around 20 µM was diluted into 98 µl of a pre-mixed solution of Ca2+/EGTA or Ca2+/NTA in black 96-well plates. Fluorescence intensities were read on a plate reader (Tecan Spark 20 M). Fluorescence intensity was measured at 26 °C and 37 °C. The excitation was set to 488 nm and emission was 525 nm. All bandwidths were set to 10 nm. Changes in fluorescence after addition of Ca2+ were calculated in Microsoft Excel. The fluorescence (y) was plotted against the free calcium concentration (x) and a four-parameter dose–response curve (variable slope) using GraphPad Prism software was fit where a is the value of fluorescence at the bottom of the curve, b is the value of fluorescence at the top of the curve, EC50 is the concentration of agonist that gives a response halfway between bottom and the top, and (n) is the hill or cooperative coefficient according to equation (1):
Lentivirus production
HEK 293T cells were purchased from the American Type Culture Collection (CRL-3216). They were transfected with the pLKO shRNA vector plasmid along with third-generation packaging, transfer and envelope plasmids, using the vesicular stomatitis virus G glycoprotein as the envelope protein with transient transfection in a medium containing chloroquine (Merck). The medium was replaced after 6 h and the supernatant was collected after 36 h. The supernatant was treated with DNase I (Roche) and then ultracentrifugation was carried out at 60,000g for 90 min. The resulting pellet was resuspended in 0.1 M PBS, aliquoted and frozen at −80 °C until use. Lentivirus was produced at the iVector facility at the Paris Brain Institute in BSL2 facilities. The lentivirus production presented a titre of 4.01 × 109 viral particles per μl, measured by ELISA using the p24 ZeptoMetrix kit (Merck).
Primary culture of embryonic rat neurons for western blotting
To assess the efficiency of Letm1-targeted shRNA specifically in neurons, we used primary cultures of rat embryonic neurons, which do not present astrocytes, allowing the assessment of Letm1 levels only from neurons. Pregnant rats (embryonic day 18) were euthanized by CO2 asphyxiation, and the embryos were then isolated onto sterile ice-cold HBSS solution, followed by the dissection of the cortex and the hippocampus. After removal of meninges, the tissue was digested with papain (Worthington Biochemical, LK003178) to isolate single cells. The dissociated cells were plated onto six-well plates coated with poly-d-lysine (Merck, P2636). Around 0.5 M cells were plated per well in plating media (prepared according to a previously published recipe150). At 5 DIV, half of the medium was replaced with maintenance media composed of BrainPhys Neuronal Medium supplemented with 2% (vol/vol) SM1 (STEMCELL Technologies, 05792) and 12.5 mM d-(+)-glucose (Merck, G8270) in addition to 10 μM 5’-fluoro-2’-deoxyuridine (Fisher Scientific, 10144760). Media replacement was carried out every 4–5 days. At 8 DIV, for assessing the different KD conditions, neurons were transduced with lentiviruses expressing the plKO-Letm1 shRNA or plKO-PDP1-shRNA-Letm1 shRNA constructs at a multiplicity of infection of 50.
Immunocytochemistry
Immunocytochemistry was performed on co-cultures at 17 DIV. Briefly, cells were fixed in 4% paraformaldehyde solution in PBS, washed, permeabilized using PBS 0.2% Triton X-100 (PBS-T) and blocked in PBS-T containing 3% BSA. Primary antibodies used were anti-MAP-2 (AB183830, Abcam) and anti-GFAP (173008, Synaptic Systems), diluted at 1:200 in blocking solution overnight at 4 °C. Secondary antibodies used were Goat anti-mouse IgG coupled to Alexa Fluor Plus 555 (A32727, Invitrogen) and Goat anti-rabbit IgG coupled to Alexa Fluor 488 (A11034, Invitrogen), diluted at 1:200 in blocking solution at room temperature for 2 h. Cells were washed in PBS three times after each antibody incubation. DAPI was included in the final wash. Cells were mounted in slides with fluorescence mounting media (F4680, Sigma-Aldrich). Images were acquired in a widefield Apotome Zeiss microscope at ×10 magnification covering the whole area containing cells. Analysis was performed manually using ImageJ by counting the total number of nuclei, the number of nuclei corresponding with the neuronal marker MAP-2 and the number of nuclei corresponding with the astrocytic marker GFAP. The percentage of neurons, astrocytes and unidentified cells was determined for every image and averaged for all images for each coverslip.
Western blotting
For analysis of protein levels of neurons in culture, lysates of embryonic neurons were prepared using RIPA buffer supplemented with 1× protease inhibitor cocktail (from 100×; Merck, P8849) and 1 mM phenylmethylsulfonyl fluoride (Roche, 10837091001) at 18 DIV. Lysates with 30 μg of protein were loaded onto SDS–PAGE gels and transferred onto nitrocellulose membranes after separation. The blots were probed with anti-Letm1 (514136, Santa Cruz Biotechnology), anti-synaptophysin 1 (101011, Synaptic Systems), anti-Total OXPHOS (110413, Abcam) and anti-PDP1 (84612-3-RR, Proteintech), and β-actin (PA5-85271, Thermo Fisher Scientific) was used as the loading control.
For analysis of protein levels of mouse hippocampus, mice were stereotaxically injected as described below with control virus (pAAV[mir-30]-CamK2(short)>mRFP1: scramble) or shLETM1 virus (pAAV[mir-30]-CamK2-mir-30-shRNA no. 1]: WPRE). Mice were euthanized and hippocampi were obtained. Given that mRFP1 has a naturally visible red colour due to its chromophore, we dissected the dorsal region of the hippocampus that appeared red by eye and prepared lysates using RIPA buffer supplemented with 1× protease inhibitor cocktail (from 100×; Merck, P8849) and 1 mM phenylmethylsulfonyl fluoride (Roche, 10837091001). Lysates with 30 μg of protein were loaded onto SDS–PAGE gels and transferred onto nitrocellulose membranes after separation. The blots were probed with anti-Letm1 (514136, Santa Cruz Biotechnology), anti-PDH E1 Alpha (Proteintech, 18068-1-AP), anti phospho-293 PDH (Proteintech, 84612-3-RR) and β-actin (PA5-85271, Thermo Fisher Scientific). Chemiluminescence images of the blots were obtained using Clarity Max ECL Western Blotting Substrate (Bio-Rad, 1705062) and the Chemi-doc Touch imaging system (Bio-Rad) following which the blots were quantified using the Image Lab (Bio-Rad) software.
All primary antibodies were prepared in 10% skimmed milk (Sigma, 70166) dissolved in TBS-Tween 0.2%, except for the anti-Total OXPHOS, which was prepared in 5% BSA in TBS-Tween. The primary antibodies and dilutions used in this study were anti-Letm1 (1:1,000 dilution), anti-PDH E1 Alpha (1:2,000 dilution), anti phospho-293 PDH (1:1,000 dilution), anti-synaptophysin1 (1:2,000 dilution), anti-Total OXPHOS (1:2,000 dilution), anti-PDP1 (1:1,000 dilution) and anti-β-actin (1:4,000 dilution). As secondary antibodies, we used Goat anti-Rabbit or Goat anti-Mouse IgG (Bio-Rad, 1706516) using a 1:5,000 dilution in 10% milk.
Western blot molecular weight assignment
Each blot was run with a pre-stained protein molecular weight ladder (PageRuler 10–180 kDa, Thermo Fisher, 26616). Immediately after transfer and before blocking, the PVDF membrane was photographed in colour to record the ladder positions. After chemiluminescence detection, the ECL exposure was registered to the colour image in Fiji/ImageJ, enabling assignment of apparent molecular weights by interpolation from molecular weight versus relative migration standard curve. For all targets, the detected bands matched the expected sizes from UniProt.
Live imaging of primary neurons
Unless otherwise noted, primary hippocampal neurons were transfected using Ca2+ phosphate at 7 DIV as described above and in previous work89, and were imaged from 14 to 21 DIV. Experiments using an shRNA against Letm1 were always performed at least 10 days after transfection to ensure protein turnover at mitochondria4. Imaging experiments were performed using a custom-built laser-illuminated epifluorescence microscope (Zeiss Axio Observer 3) coupled to an Andor iXon Ultra camera (model no. DU-897U-CSO-#BV), whose chip temperature is cooled down to −90 °C to reduce noise in the measurements using the Oasis UC160 Cooling System. Illumination using fibre-coupled lasers of wavelengths 488 (Coherent OBIS 488 nm LX 30 mW) and 561 (Coherent OBIS 561 nm LS 80 mW) was combined through using the Coherent Galaxy Beam Combiner, and laser illumination was controlled using a custom Arduino-based circuit coupling imaging and illumination. Primary neuron–astrocyte cultures were grown on poly-ornithine-coated coverslips (D = 0.17 mm, Warner instruments), mounted onto an RC-21BRFS imaging chamber for field stimulation (Warner Instruments) and imaged through a ×40 Zeiss oil objective ‘Plan-Neofluar’ with an NA of 1.30 (WD = 0.21 mm). Imaging frequencies used in experiments were 5 Hz for Mito4x-GCaMP6f, 100 Hz for cytosolic GCaMP8f and 2 Hz for all the others. The temperature of all experiments was clamped at 36.5 °C and was kept constant by heating the chamber through a platform (PH-2, warner instruments) together with an in-line solution heater (SHM-6, Warner Instruments), through which solutions flowed at 0.35 ml min−1. The temperature was kept constant using a feedback loop temperature controller (TC-344C, Warner Instruments). External Ca2+ in the medium was maintained constant at a level of 2 mM across all experiments. We identified axons as neuronal projections extending beyond ~500 μm from the cell body, while dendrites were identified by being thick proximal projections presenting spines. In axons, mitochondria are also easily recognized relative to dendrites because they are typically smaller and more rounded49,51.
In the case of glutamate uncaging experiments, live cell imaging was conducted between 18 and 19 days after plating. Experiments were performed at 37 °C and in a modified E4 imaging buffer containing 120 mM NaCl, 3 mM KCl, 10 mM HEPES (buffered to pH 7.4), 4 mM CaCl2 and 10 mM glucose. Imaging during glutamate uncaging was performed using a custom-built inverted spinning-disk confocal microscope (3i Imaging Systems, CSU-W1) with an Andor iXon Life 888 for confocal fluorescence imaging. Image acquisition was controlled by SlideBook 2023 software. Images were acquired with a Plan-Apochromat ×63/1.4-NA oil objective, M27 with DIC III prism, using a CSU-W1 Dichroic for 488/561-nm excitation with a Quad emitter and individual emitters. During imaging, the temperature was maintained at 37 °C using an Okolab stage top incubator with temperature control.
All experiments in cultured cells were performed in continuously flowing Tyrode’s solution containing 2 mM Ca2+. Unless otherwise noted, Tyrode’s solution contained the following components: 119 mM NaCl, 2.5 mM KCl, 2 mM CaCl2, 2 mM MgCl2, 20 mM glucose, 10 μM CNQX and 50 μM AP5, buffered to pH 7.4 at 37 °C using 25 mM HEPES. However, key experiments were replicated in the same Tyrode’s solution but containing 1.2 mM glucose and 43.8 mM HEPES to avoid possible contributions of high glucose to results. Experiments using 1.2 mM glucose are shown in Fig. 1e and Extended Data Figs. 1g–l, 2a–d and 5d,e. Experiments in Fig. 2 and Extended Data Fig. 3a,b,g,h were done in a Tyrode’s solution containing no glucose but lactate and pyruvate as fuel, with the following composition: 119 mM NaCl, 2.5 mM KCl, 2 mM CaCl2, 2 mM MgCl2, 0 mM glucose, 1.25 mM lactate, 1.25 mM pyruvate, 10 μM CNQX and 50 μM AP5, buffered to pH 7.4 at 37 °C using 42.5 mM HEPES. None of the Tyrode’s solutions contain glutamine.
NH4Cl solutions for calibrating pHluorin measurements had a similar composition as Tyrode’s buffer except they contained 100 mM NH4Cl and 19 mM NaCl for a pH of 7.4, pH of 7.8 and pH of 8.2 at 37 °C for mitochondrial estimates, and 50 mM NH4Cl and 69 mM NaCl for a pH of 7.4 at 37 °C for synaptic vesicle pH estimates. Chronic incubation with TTX (Tocris) was performed by adding a 1 µM final concentration of TTX in the culture media one day after transfection, and it was refreshed 5 days after the initial addition and maintained until synaptic ATP levels were measured using Syn-ATP. Luminescence imaging of the presynaptic ATP reporter Syn-ATP was performed as previously described10,25. We did not observe significant differences in pH changes when comparing wild-type neurons and Letm1 KD neurons (Extended Data Fig. 2d); therefore, we did not correct for pH changes in ATP measurements. vGlut-pHluorin signals during electrical stimulation are reported as a percentage of the total vesicle pool, whose fluorescence is obtained by perfusion of a Tyrode’s solution containing 50 mM NH4Cl buffered at pH 7.4 using 25 mM HEPES.
Live FRET imaging of pyruvate or ATP using Pyronic or ATeam 1.03, respectively, was carried out using a widefield inverted Zeiss Axio Observer 7 microscope with an incubation chamber maintained at 37 °C. Cells were imaged in Tyrode’s buffer without AP5 and CNQX. The ×40 LD Plan-Neofluar objective with 0.75 NA was used for image acquisition with excitation at 430 nm and emissions recorded at 480 nm ± 20 nm (mTFP or CFP) and 535 nm ± 15 nm (Venus or YFP). A HXP 120-V metal halide lamp (Leistungselektronik Jena) was used as the illumination source. The inverse FRET ratio (R = mTFP/Venus) was calculated as a measure of pyruvate levels while the Venus/eCFP ratio was calculated as a measure of ATP levels. In the case of Pyronic imaging, the slope was determined automatically using the ‘statelevels’, ‘risetime’ and ‘slewrate’ functions in the MATLAB signal processing toolbox to analyse the ΔR/R0 traces.
Single spine stimulation experiments and analysis
Neurons were transfected with RCaMP1.07 and Mito4x-GCaMP6f10 plasmid constructs, along with the Letm1 shRNA when specified. Transfected neurons were identified by changes in RCaMP1.07 fluorescence in dendrites corresponding to calcium transients. Before glutamate uncaging, neurons were replaced with 1 μM TTX, 2 mM 4-methoxy-7-itroindolinyl-caged-l-glutamate (MNI caged glutamate; Tocris Bioscience, 100 mM stock made in modified E4 buffer) in modified E4 buffer lacking Mg2+ (see above). Glutamate uncaging was performed using a multiphoton laser at 720 nm (Chameleon, Coherent) and a Pockels cell (Conoptics) to control the uncaging pulses. To test a spine’s response to an uncaging pulse, an uncaging spot (2 μm2) close to a spine head was selected and two to three uncaging pulses, each at 10-ms pulse duration per pixel at 5.9–9.6-mW power, were given. Only spines with pulse-specific calcium transients were selected for the following experiments. A single uncaging pulse was given, followed by acquisition.
Image analysis for in vitro experiments
We used the ImageJ plugin Time Series Analyzer V3 for imaging analysis. This involved the selection of 150–250 regions of interest (ROIs) for synaptic boutons, or 10–150 ROIs for responding boutons, and measuring the fluorescence over time. Mitochondrial and cytosolic Ca2+ signals in response to electrical activity (ΔF) were normalized to the resting fluorescence (F0), unless otherwise mentioned in the text.
Mitochondrial calcium measurements
For activity-driven mitochondrial Ca2+ measurements using Mito4x-GCaMP6f, data were obtained from imaging axonal mitochondria responses to electrical stimulation. As previously reported10, in some cases a small fraction of Mito4x-GCaMP6f appeared mislocalized in the cytosol, which would contaminate the quantification of peak mitochondrial responses. Leveraging the kinetic differences of cytosolic and mitochondrial Ca2+ responses, we quantified exclusively mitochondrial Ca2+ by choosing a 1-s delay after the stimulus as the peak response. At this point cytosolic Ca2+ has returned to baseline, allowing a clean estimate of mitochondrial responses without any contribution from possible mislocalized probes. Neurons with apparent cytosolic mislocalization of Mito4x-GCaMP6f were excluded. Based on this criterion, 3 of 51 neurons were excluded from the analysis.
For measuring the rate of efflux, we did not fit our data to any model because mitochondrial Ca2+ decays differently in different conditions. Therefore, we used the time to reach half the peak value that is, half-time decay (t1/2), as a comparable indicator of the rate of Ca2+ efflux across conditions.
Mitochondrial free Ca2+ concentration estimates, one-point calibration protocol
In this method, to estimate free Ca2+ concentration in axonal mitochondria expressing Mito4x-GCaMP6f, we measured fluorescence at saturating [Ca2+] in mitochondria (Fmax). This was obtained by applying Tyrode’s solution containing 500 μM ionomycin, 4 mM CaCl2 and 0 mM MgCl2 at pH 6.9 buffered with 25 mM HEPES, as done previously10,62. Knowing the parameters of purified GCaMP6f147, baseline mitochondrial [Ca2+], [Ca2+]r is calculated from Fmax using equation (2):
EC50 is the affinity constant of the indicator, Fr is the measured fluorescence at rest, Rf is the dynamic range (Fsat/Fapo) and n is the Hill coefficient. The values for EC50, Rf and n were obtained from those calculated in a previously published paper on GCaMP6f147. Ionomycin application does not produce a change in mitochondrial matrix pH10.
Mitochondrial free Ca2+ concentration estimates, two-point calibration protocol
To avoid relying on the in vitro-determined dynamic range of the indicator, we also performed a two-point calibration that estimates this value in situ. In this method, both the minimum (Fmin) and maximum (Fmax) fluorescence of the sensor were determined directly in axonal mitochondria. Neurons were imaged in basal conditions first using a Tyrode’s solution containing 1.2 mM glucose and 2 mM Ca2+. Before measuring Fmin, neurons were washed with the same Tyrode’s solution but containing no Ca2+. Next, we obtained Fmin by incubating neurons for 8 min with Tyrode’s 0 mM Ca2+, 1.2 mM glucose, 50 μM CCCP and 10 μM BAPTA-AM. Finally, to obtain Fmax, neurons were incubated with Tyrode’s 4 mM Ca2+, 1.2 mM glucose and 50 μM ionomycin, and imaged after a quick increase in fluorescence. Resting fluorescence (F0) was converted into fractional sensor saturation (S) by normalizing between the Fmin and Fmax according to equation (3):
Under the assumption of a Hill binding model (with dissociation constant EC50 and Hill coefficient n), the fractional saturation is (equation (4)):
This equation can be algebraically inverted to estimate resting free Ca2+ concentration according to equation (5):
Equation 5 was used to calculate the resting mitochondrial calcium values. The values for EC50, Rf and n were obtained from those calculated in a previously published paper on GCaMP8f66.
Mitochondrial pH measurements
Mitochondrial pH measurements were obtained by measuring Mito4x-pHluorin fluorescence in axonal mitochondria. Neurons were perfused sequentially with three different Tyrode’s solutions containing 100 mM NH4Cl but buffered at pH 7.4, pH 7.8 and pH 8.2 (Extended Data Fig. 2a). After an initial overshoot of fluorescence in the presence of 100 mM NH4Cl, which has been previously described to occur when using endoplasmic reticulum pHluorin62, signals stabilized to equilibrated mitochondrial pH corresponding with those of the Tyrode’s solution. These three fluorescence values in these conditions were used for estimating resting mitochondrial pH using the modified Henderson–Hasselbalch equation according to equation (6):
Using the biophysical properties described for pHluorin151, pKa is the pKa of pHluorin, 7.2, and n is the hill coefficient of pHluorin responsiveness, 1.9. pHNH4Cl,i is the pH of the 100 mM NH4Cl calibration buffer (i = 7.8 or 8.2), F0 is the fluorescence of Mito4x-pHluorin measured before NH4Cl perfusion, FNH4Cl is the fluorescence of Mito4x-pHluorin measured after NH4Cl perfusion when signal is stable in each buffer (i = 7.8 or 8.2). To obtain the final estimates of mitochondrial pH, we averaged results from changes obtained in pH 7.8 and pH 8.2, as these are clearly distinguishable from baseline fluorescence and thus present less noise when measured.
Synaptic vesicle pH measurements
Synaptic vesicle pH measurements were obtained by measuring vGlut-pHluorin fluorescence in axons. Neurons were perfused with a Tyrode’s solutions containing 50 mM NH4Cl buffered at pH 7.4, causing a stable peak in fluorescence corresponding to pH 7.4, as previously described. Synaptic vesicle pH was estimated as described for mitochondria but using the values obtained at pH 7.4.
Cytosolic pH measurements
Cytosolic pH measurements were obtained by measuring cytosolic pHluorin fluorescence in axons. Neurons were perfused sequentially with three different modified Tyrode’s solutions buffered to pH 7.4, 7.8 and 8.2. These solutions lack Na+ and Ca2+ but contain 1.2 mM glucose, 2 mM MgCl2, 123.5 mM KCl, 53.8 mM HEPES, 10 mM CNQX, 50 mM APV and 20 µM nigericin. Fluorescence was stabilized after each perfusion in around 30 s, allowing the measurement of the change in fluorescence in each of the cases. Cytosolic pH was estimated as described for mitochondria, averaging results from fluorescence changes obtained in pH 7.8 and pH 8.2.
Permeabilization and ATP estimates
To enable quantification of ATP from experimental L/F measurements, Syn-ATP was calibrated by determining its response in the presence of 0 mM ATP (L/Fmin) and 5 mM ATP (L/Fmax) under imaging conditions identical to those used in the experiments. Calibration was achieved by permeabilizing neurons by incubation with 1,000 U ml−1 sreptolysin-O (Sigma, 98072-47-0) for 2 min in a permeabilization buffer containing 139 mM KCl, 20 mM PIPES (buffered to pH 7.0 at 37 °C), 5.22 mM MgCl2, 0.186 mM CaCl2 and 0.91 mM EGTA. This experiment was performed four independent times, which allowed obtaining L/Fmin and L/Fmax, defining the dynamic range of the sensor and allowing for normalization of all experimental L/F values as shown in equation (7):
The normalized L/F values were then related to ATP concentration using a Michaelis–Menten equation with the apparent affinity (Km) fixed at the published value of 2.3 mM25. The ATP concentration corresponding to each measurement was obtained by solving equation (8):
Only normalized L/F values within the calibration range were converted. Values above 1 were excluded being above calibration range (1 of 73 measurements).
Estimates of synapse number per axonal distance
Neurons were transfected to express the synaptic marker synapsin1-mRuby and scramble or Letm1 shRNAs. Both shRNA vectors also expressed cytosolic mTagBFP2, which we used as a filler to identify axons. Using ImageJ, we manually drew segmented lines in axons using the BFP channel and generated plot profiles for the corresponding synapsin1-mRuby signals. Synapses were counted as those whose mRuby fluorescence was 3.5 times the fluorescence of the inter-synapse space corresponding to the axon, divided by the distance of the axon drawn and multiplied by 100 to estimate the number of synapses per 100 μm.
Estimates of mitochondrial shape
Different morphological aspects of mitochondria were analysed from the images taken in the mitochondrial calcium measurement experiments in which neurons expressed Mito4x-GCaMP6f and scramble or Letm1 shRNAs. Images were analysed using the ImageJ plugin Trainable Weka Segmentation following the criteria described previously152,153. For cultured neurons, in addition to the global background of the image not containing any cell type, pixels containing astrocytes were also selected as a secondary background, which increased specificity in the mitochondrial shaping measurements performed by the Trainable Weka Segmentation plugin. The morphological aspects of mitochondria analysed for neurons were area, perimeter, circularity and aspect ratio. We averaged the results per neuron for statistical comparison.
Dendritic responses during glutamate uncaging
Image analysis was performed using ImageJ. ROIs of ~2 μm2 were drawn at the base of the stimulated spine to measure the dendritic calcium signal in the 561-nm channel and the mitochondrial matrix signal in the 488-nm channel. For spine calcium measurement, ROIs of ~0.8 μm2 were drawn on the spine and measured in the 561-nm channel. Only dendrites with apparent mitochondrial matrix response were analysed. The average intensity of each ROI was measured, and the background was subtracted using the intensity measured from an adjacent background area. F0 was defined as the average of the intensities measured at the five time points before spine stimulation. For each successive time point during and after stimulation, the normalized intensity (Fnorm) was calculated using the equation: Fnorm = (F − F0)/F0. For normalization to maximum analysis, F/F0 corresponding to each time point was divided by the maximum Fnorm value (Fmax) of the entire time trace. Traces with ectopic peaks following the stimulation pulse or offshoots of Mito4x-GCaMP6f10 fluorescence were excluded from the analysis.
Computational model for neuronal activity and mitochondrial metabolism
We reproduced and adapted a previously published mitochondrial metabolism model90 that simulates some essential components of the TCA and ATP production with eight differential equations using mass action kinetics and irreversible reactions (henceforth referred to as the Nazareth model). The Nazareth model is a minimalist metabolism model that starts with pyruvate and includes acetyl-coenzyme A, citrate, alpha-ketoglutarate, oxaloacetate, nicotinamide adenine dinucleotide (NAD+), ATP and the intrinsic mitochondrial membrane potential (ΔΨ). The levels of these metabolic intermediates are set by the rate at which ATP in the mitochondria is exchanged with the cytosolic ADP, and in the model is controlled by a variable (KANT) that sets the rate constant of the adenine nucleotide translocator.
To simulate the metabolic expense of spiking in neurons, we modulated this variable (KANT) as a function of time. We separated the costs of a neuron into two components, the first of which accounted for the non-spike-related metabolic expenses of a neuron and this remained the same in all our simulations (100/ks). The second component was a per-spike metabolic expense, which was an additional transient increase in this KANT after each spike (0 to 10/ks, followed by a slow delay to 0 with a 100-ms decay constant). Effectively, this guaranteed that the metabolic expenses incurred by a neuron were dependent on the spiking rate of the neuron.
Spikes in our model
To simulate the effect of spiking, we assumed spikes occur either regularly (Fig. 2f), or as a Poisson point process in case of random spikes (Fig. 2g–i). Each spike is merely a point process with no specific details of ionic currents. This serves as a proxy for a detailed neuronal model that is sufficient to explore the phenomenon we wish to observe.
Mitochondrial free Ca2+ and its influence on TCA and ETC reactions
In addition to the ATP expense incurred due to each spike, each spike also added a unitless value of 0.1 to an excess free Ca2+ variable (ca_mito). This variable decayed to 0 over time depending on if it was the case of control (decay constant of 7 s) or if it was the case of Letm1 KD (decay constant of 20 s). In our implementation of the Nazareth model, the excess free Ca2+ variable increased the rate of reactions for the equivalent enzymes of pyruvate, isocitrate and alpha-ketoglutarate dehydrogenases and the complex V according to equation (9):
Where ‘frac’ is the fractional increase of the rate constants due to the Ca2+ binding, and ‘ca_mito’ is the excess free Ca2+ in the mitochondria.
In vivo imaging in D. melanogaster
Crosses for imaging experiments were raised at 23 °C and fly progeny were induced for 3 days at 30.5 °C to drive sufficient expression of the probe (and the desired RNAi) for use in imaging. All in vivo imaging was performed on female flies, which are preferred as their larger size facilitates surgery. Around 30 min to 1 h (for mitochondrial Ca2+ imaging) and 3 h (for pyruvate or mitochondrial Ca2+ imaging) after 1× paired or 1× unpaired olfactory conditioning, flies were gently handled by aspiration without anaesthesia and glued on their dorsal side to a plastic coverslip coated with a thin transparent plastic sheet. The coverslip was then placed on a recording chamber. Surgery was performed to obtain an imaging window on the fly head by removing the cuticle, trachea and fat bodies, thereby exposing the underlying MB neurons. During the procedure, the head capsule is bathed in a drop of artificial haemolymph: 130 mM NaCl (Merck, S9625), 5 mM KCl (Merck, P3911), 2 mM MgCl (Merck, M9272), CaCl2 2 mM (Merck, C3881), 5 mM d-trehalose (Merck, 9531), 30 mM sucrose (Merck, S9378) and 5 mM HEPES hemisodium salt (Merck, H7637). At the end of the procedure, any remaining solution was absorbed and a fresh 90-µl droplet was applied on the preparation.
For measuring mitochondrial Ca2+ levels in vivo, confocal excitation was achieved using dual excitation at 488 nm (Mito4x-LA-GCaMP6s) and 633 nm (mCarmine). Images (512 × 250) were acquired at a rate of one image every 1.5 s. Calcium levels were measured by quantifying Mito4x-LA-GCaMP6s and normalizing it by the mCarmine signal as both are driven by the same GAL4-UAS system. The Mito4x-LA-GCaMP6s and mCarmine signals were recorded for 5 min to calculate the average Mito4x-LA-GCaMP6s/mCarmine ratio over this period. The ROIs were delimited by hand around each visible MB vertical lobe. The final ratios were normalized for each experiment using the average ratio of the unpaired dataset for each genotype. The final normalized ratios were pooled together. The indicated ‘n’ is the number of animals that were assayed in each condition.
For measuring pyruvate fluxes in vivo, two-photon imaging was performed using a Leica TCS-SP5 upright microscope equipped with a ×25, 0.95-NA water immersion objective. Two-photon excitation was achieved using a Mai Tai DeepSee laser tuned to 825 nm. Images (512 × 150) were acquired at a rate of two images per second. Measurements of pyruvate consumption were performed according to a previously well-characterized protocol19. After 1 min of baseline acquisition, 10 μl of a 50 mM sodium azide solution (Merck, 71289; prepared in the same artificial haemolymph solution) were injected into the 90-μl droplet bathing the fly’s brain, bringing sodium azide to a final concentration of 5 mM. Image analysis was performed as previously described19. ROIs were delimited by hand around each visible MB vertical lobe, and the average intensity of the mTFP and Venus channels over each ROI was calculated over time after background subtraction. The Pyronic sensor was designed so that FRET from mTFP to Venus decreases when the pyruvate concentration increases. To obtain a signal that positively correlates with pyruvate concentration, the inverse FRET ratio was computed as mTFP intensity divided by Venus intensity. This ratio was normalized by a baseline value calculated over the 1-min preceding drug injection. The slope was calculated between 10% and 70% of the plateau. The indicated ‘n’ is the number of animals that were assayed in each condition.
Aversive olfactory conditioning and memory test in D. melanogaster
The behavioural experiments, including sample sizes, were conducted similarly to previous studies from our research group19,154. For all experiments, training and testing were performed in a soundproof and odour-proof room at 25 °C and 80% humidity. Experimental flies (male and female) were transferred to fresh bottles containing standard medium on the day before conditioning for the non-induced condition. For the induced condition, flies were transferred 2 days before the experiment at 30.5 °C to allow RNAi expression.
Conditioning
Flies were conditioned by exposure to one odour paired with electric shocks and subsequent exposure to a second odour in the absence of shock. A barrel-type machine was used for simultaneous automated conditioning of six groups of 40–50 flies each. Throughout the conditioning protocol, each barrel was attached to a constant air flow at 2 l min−1. The odorants 3-octanol and 4-methylcyclohexanol, diluted in paraffin oil at 0.360 mM and 0.325 mM, respectively, were alternately used as conditioned stimuli (CS+). For a single cycle of associative training, flies were first exposed to an odorant (the CS+) for 1 min while 12 pulses of 5-s long, 60-V electric shocks were delivered; flies were then exposed 45 s later to a second odorant without shocks (the CS−) for 1 min. Here, the groups of flies were subjected to one of the following olfactory conditioning protocols: 1 cycle training (1×) or five associative cycles spaced by 15-min intertrial intervals (5× spaced conditioning). Non-associative control protocols (unpaired protocols) were also used for in vivo imaging experiments. During unpaired conditioning, the odour and shock stimuli were delivered separately in time, with shocks occurring 3 min before the first odorant. After training and until memory testing, flies were kept on regular food at 25 °C (for 3 h memory test) or at 18 °C (for 24 h memory test).
Memory test
The memory test was performed either 3 h after 1× conditioning or 24 h after 1× or 5× spaced conditioning in a T-maze apparatus comprising a central elevator to transfer the flies to the centre of the maze arms. During the test, flies were exposed simultaneously to both odours (the same concentration as during conditioning) in the T-maze. After 1 min of odorant exposure in the dark, flies were trapped in either T-maze arm, retrieved and counted. A memory score was calculated as follows according to equation (10):
where PI is the performance index, #CS− is the number of flies avoiding the conditioned odour, and #CS+ is the number of flies preferring the conditioned odour. A single memory score is the average of two scores obtained from two groups of genotypically identical flies conditioned in two reciprocal experiments, using either odorant (3-octanol or 4-methylcyclohexanol) as the CS+. The indicated ‘n’ is the number of independent memory score values for each genotype.
Odour perception test
The olfactory acuity of flies was tested after conditioning with the CS+, as electric shocks modify their olfactory perceptions. Flies were then immediately tested in a T-maze, where they had to choose between the CS− or its solvent (paraffin oil). Odour concentrations used in this assay were the same as for the memory assays. At these concentrations, both odorants are innately repulsive. The odour-interlaced side was alternated for successively tested groups. After 1 min, flies were counted, and naive odour avoidance was calculated as for the memory test.
Electric shock perception test
During the test, flies must choose between two barrels: one delivering the electric shocks, and one that is neutral. The compartment where the electric shocks are delivered was alternated between two consecutive groups. After 1 min, flies were counted, and shock avoidance was calculated as for the memory test.
Quantitative PCR on D. melanogaster brains
To assess the efficiency of Letm1 RNAi lines to knock down Letm1 in MB neurons, female flies carrying the tubulin-Gal80ts;VT30559-Gal4 MB neuron driver were either crossed with UAS-Letm1 RNAi no. 1 or UAS-Letm1 RNAi no. 2 males, or with CS males for controls. In flies, Letm1 is encoded by three very similar transcripts (FBtr0072348 Letm1-RB, FBtr0072349 Letm1-RC, FBtr0072347 Letm1-RA), all producing the same 1,013 amino acid protein isoform (UniProt P91927, Ensembl Canonical, APPRIS P1). Thus, both RNAi used target the single Letm1 protein isoform expressed in Drosophila. Fly progeny was raised at 23 °C throughout development. Newly hatched flies were transferred to fresh food vials at 30.5 °C for 4 days of induction before RNA extraction, as previously reported155. RNA extraction and cDNA synthesis were performed using the RNeasy Plant Mini Kit (Qiagen), RNA MinElute Cleanup Kit (Qiagen), oligo(dT)20 primers and the SuperScript III First-Strand kit (Thermo Fisher Invitrogen). Amplification was performed using a LightCycler 480 (Roche) and the SYBR Green I Master mix (Roche). Specific primers used for Letm1 cDNA are forward 5′-GCCCAGAATGTCTACAAGCG-3′ and reverse 5′-GCGGGTTCATCTGTGACGTT-3′ and for the reference αTub84B (Tub, CG1913) cDNA: forward 5′- TTGTCGCGTGTGAAACACTTC-3′ and reverse 5′ CTGGACACCAGCCTGACCAAC-3′. The level of Letm1 cDNA was compared against the level of the αTub84B reference cDNA. Each reaction was performed in triplicate. The specificity and size of amplification products were assessed by melting curve analyses. Expression relative to the reference was presented as the fold change compared to the calibrator (relative quantification RQ = 2−ΔΔCt, where Ct is the cycle threshold).
Aversive olfactory conditioning and memory test (Mus musculus)
Drugs
LiCl (Merck, 203637) was dissolved in saline (0.9% NaCl solution) to obtain a working solution of 0.3 M concentration. For the conditioned odour aversion protocol, a banana odour (isoamyl acetate solution at 0.05% in water) and an almond odour (benzaldehyde solution at 0.01% in water) were used.
Stereotaxic surgeries
C57BL/6J male mice of 7–8 weeks of age were anaesthetized with a mixture of ketamine–medetomidine (75:1 mg per kg body weight) and placed in a stereotaxic apparatus. Animals were infused bilaterally in the dorsal hippocampus (ML ± 1.5; AP −2; DV −1.5) with 500 nl of control virus (pAAV[mir-30]-CamK2(short)>mRFP1: scramble) or shLETM1 virus (pAAV[mir-30]-CamK2-mir-30-shRNA no. 1]: WPRE) with a flow rate of 1 nl s−1. At the end of all behavioural tests, animals were anaesthetized with a cocktail mixture of ketamine–xylazine (50:20 mg per kg body weight) and perfused with 4% paraformaldehyde in 0.1 M phosphate buffer. Next, brains were removed to confirm the expression of the viral infection by fluorescence microscopy.
Conditioned odour aversion
Four weeks after the stereotaxic surgeries and before assessing the conditioned odour aversion protocol, mice were placed in a behavioural chamber (25 × 15 cm) to evaluate locomotor activity for 20 min. Subsequent analysis was done by combining DeepLabCut156 and the BehaviorFlow package157, which enables automated behavioural tracking and classification. Next, the conditioned odour aversion was assessed in mice using two odours and LiCl, based on a modified protocol described previously. Briefly, the principle of this task is that if we devalue one of the odours by coupling it with an injection of LiCl (unconditioned stimulus), mice will prefer the non-conditioned odour (conditioned stimulus, CS−) over the devalued one (aversive conditioned stimulus, CS+). During the protocol, mice were individualized only during the 1-h access to the drinking bottles and then returned to grouped housing. Before starting the first day of the protocol, mice were water deprived for 24 h, and this water deprivation lasted for 5 consecutive days of the protocol. During the first 2 days of habituation, individualized mice had access to the two water drinking bottles for 1 h each day. On the third day, mice received two identical bottles of water but containing the banana or almond odours (randomized) and after 1 h, one odour was devalued (CS+) with an intraperitoneal injection of LiCl (0.3 M concentration; 10 ml per kg body weight). LiCl causes gastric malaise that causes a reduction in locomotor activity due to the sickness. On the fourth day, mice had 1 h of access to bottles of water along with the second odour (CS−) and at the end of this session mice were injected intraperitoneally with an innocuous physiological solution (0.9% NaCl). To assess MTM, on the fifth day a two-bottle choice test was conducted, offering access to both water-based odours for 1 h. Preferences were assayed by quantifying consumption of each bottle (Extended Data Fig. 4a). The bottle positions were alternated to avoid bias. To assess LTM, the two-bottle test was repeated after 10 days, following a 24-h water deprivation period and preference was quantified using the same method (Extended Data Fig. 4b). The time points selected to study MTM (1 day) and LTM (10 days) in this study were chosen in mice to be approximately proportional to those tested in Drosophila by relativizing them to species lifespan.
Statistics and reproducibility
Statistical analyses were performed in GraphPad Prism v8 for Windows. For each dataset, distributional assumptions were checked (Shapiro–Wilk test and inspection of residuals) and, based on this, either parametric or non-parametric tests were used appropriately. Exact tests used, sample sizes (n) and exact P values are reported in the figure legends. Data are shown as the mean ± s.e.m. For experiments with more than two groups, analyses included ANOVA (or Kruskal–Wallis for non‑parametric data) with appropriate post hoc multiple‑comparison corrections as specified in the figure legends.
In each figure, n denotes independent biological replicates. For single-neuron measurements, each biological replicate is considered a single neuron. For fly memory tests, each biological replicate is a measurement involving several flies, as indicated in each case. For in vivo fly imaging, each fly used is a biological replicate. For in vivo mouse work, each mouse is a biological replicate. Multiple measurements made from the same biological sample (for example, multiple fields of a single axon or repeated runs from the same lysate) were treated as technical replicates; technical replicates were averaged within each biological replicate and were not used to increase n. All key findings were reproduced in at least three independent experiments unless otherwise indicated. Uncropped western blots corresponding to the cropped panels shown in the figures are provided in the Supplementary Information.
No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications10,99,140. No data were excluded from the analyses. Phenotype studies were randomized to minimize potential bias. For each experimental replicate, the order in which phenotypes were analysed was alternated systematically. In cases involving two groups, such as A and B, we ensured that the sequence alternated (for example, A, B, B, A, B, A, A, B) to account for any order effects. The investigators were not blinded to allocation during experiments and outcome assessment. No formal randomization was used. To minimize batch and time‑dependent effects, experiments were blocked by culture preparation/day and data acquisition was interleaved across groups.
Significance is denoted by *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Materials availability
Plasmids generated in this study are available at Addgene (https://www.addgene.org/), as indicated in Supplementary Table 1. Any plasmid not deposited in Addgene is available upon request to the corresponding author. The Mito4x-LA-GCaMP6s and the mCarmine fly lines are available from the Energy & Memory group (contact pierre-yves.placais@espci.fr or alice.pavlowsky@espci.fr) upon request.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data reported in this paper will be shared by the corresponding author upon request. Source data are provided with this paper. Any additional information required to reanalyse the data reported in this paper is available from the corresponding author upon request.
Code availability
We have generated new code for modelling mitochondrial metabolism in firing neurons. The source code of the simulations, installation instructions and software dependencies are provided with an open-source licence (GPL-3.0) via https://github.com/ccluri/letm1/.
References
Faria-Pereira, A. & Morais, V. A. Synapses: the brain’s energy-demanding sites. Int. J. Mol. Sci. 23, 3627 (2022).
Jang, S. et al. Glycolytic enzymes localize to synapses under energy stress to support synaptic function. Neuron 90, 278–291 (2016).
Dienel, G. A. Brain glucose metabolism: integration of energetics with function. Physiol. Rev. 99, 949–1045 (2019).
Yellen, G. Fueling thought: management of glycolysis and oxidative phosphorylation in neuronal metabolism. J. Cell Biol. 217, 2235–2246 (2018).
Devine, M. J. & Kittler, J. T. Mitochondria at the neuronal presynapse in health and disease. Nat. Rev. Neurosci. 19, 63–80 (2018).
Rangaraju, V., Lauterbach, M. & Schuman, E. M. Spatially stable mitochondrial compartments fuel local translation during plasticity. Cell 176, 73–84 (2019).
Stoler, O. et al. Frequency- and spike-timing-dependent mitochondrial Ca2+ signaling regulates the metabolic rate and synaptic efficacy in cortical neurons. eLife 11, e74606 (2022).
Díaz-García, C. M. et al. The distinct roles of calcium in rapid control of neuronal glycolysis and the tricarboxylic acid cycle. eLife 10, e64821 (2021).
Lujan, B. J., Singh, M., Singh, A. & Renden, R. B. Developmental shift to mitochondrial respiration for energetic support of sustained transmission during maturation at the calyx of Held. J. Neurophysiol. 126, 976–996 (2021).
Ashrafi, G., de Juan-Sanz, J., Farrell, R. J. & Ryan, T. A. Molecular tuning of the axonal mitochondrial Ca2+ uniporter ensures metabolic flexibility of neurotransmission. Neuron 105, 678–687 (2020).
Bapat, O. et al. VAP spatially stabilizes dendritic mitochondria to locally support synaptic plasticity. Nat. Commun. 15, 205 (2024).
Ghosh, I., Fan, R., Shah, M., Bapat, O. & Rangaraju, V. Synapses drive local mitochondrial ATP synthesis to fuel plasticity. Preprint at bioRxiv https://doi.org/10.1101/2025.04.09.648032 (2025).
Groten, C. J. & MacVicar, B. A. Mitochondrial Ca2+ uptake by the MCU facilitates pyramidal neuron excitability and metabolism during action potential firing. Commun. Biol. 5, 900 (2022).
Li, S. & Sheng, Z. -H. Energy matters: presynaptic metabolism and the maintenance of synaptic transmission. Nat. Rev. Neurosci. 23, 4–22 (2022).
Kaplan, L., Chow, B. W. & Gu, C. Neuronal regulation of the blood–brain barrier and neurovascular coupling. Nat. Rev. Neurosci. 21, 416–432 (2020).
Filiou, M. D. & Sandi, C. Anxiety and brain mitochondria: a bidirectional crosstalk. Trends Neurosci. 42, 573–588 (2019).
Hollis, F. et al. Mitochondrial function in the brain links anxiety with social subordination. Proc. Natl Acad. Sci. USA 112, 15486–15491 (2015).
Kanellopoulos, A. K. et al. Aralar sequesters GABA into hyperactive mitochondria, causing social behavior deficits. Cell 180, 1178–1197 (2020).
Plaçais, P. -Y. et al. Upregulated energy metabolism in the Drosophila mushroom body is the trigger for long-term memory. Nat. Commun. 8, 15510 (2017).
Plaçais, P. -Y. & Preat, T. To favor survival under food shortage, the brain disables costly memory. Science 339, 440–442 (2013).
Sgammeglia, N. & Sprecher, S. G. Interplay between metabolic energy regulation and memory pathways in Drosophila. Trends Neurosci. 45, 539–549 (2022).
Suzuki, A. et al. Astrocyte-neuron lactate transport is required for long-term memory formation. Cell 144, 810–823 (2011).
Rabah, Y. et al. Glycolysis-derived alanine from glia fuels neuronal mitochondria for memory in Drosophila. Nat. Metab. 5, 2002–2019 (2023).
Pathak, D. et al. The role of mitochondrially derived ATP in synaptic vesicle recycling. J. Biol. Chem. 290, 22325–22336 (2015).
Rangaraju, V., Calloway, N. & Ryan, T. A. Activity-driven local ATP synthesis is required for synaptic function. Cell 156, 825–835 (2014).
Sobieski, C., Fitzpatrick, M. J. & Mennerick, S. J. Differential presynaptic ATP supply for basal and high-demand transmission. J. Neurosci. 37, 1888–1899 (2017).
Schneggenburger, R. & Rosenmund, C. Molecular mechanisms governing Ca2+ regulation of evoked and spontaneous release. Nat. Neurosci. 18, 935–941 (2015).
Schneggenburger, R. & Neher, E. Intracellular calcium dependence of transmitter release rates at a fast central synapse. Nature 406, 889–893 (2000).
Devine, M. J. et al. Mitochondrial Ca2+ uniporter haploinsufficiency enhances long-term potentiation at hippocampal mossy fibre synapses. J. Cell Sci. 135, jcs259823 (2022).
Gazit, N. et al. IGF-1 receptor differentially regulates spontaneous and evoked transmission via mitochondria at hippocampal synapses. Neuron 89, 583–597 (2016).
Kwon, S. -K. et al. LKB1 regulates mitochondria-dependent presynaptic calcium clearance and neurotransmitter release properties at excitatory synapses along cortical axons. PLoS Biol. 14, e1002516 (2016).
Vaccaro, V., Devine, M. J., Higgs, N. F. & Kittler, J. T. Miro1-dependent mitochondrial positioning drives the rescaling of presynaptic Ca2+ signals during homeostatic plasticity. EMBO Rep. 18, 231–240 (2017).
Bredvik, K. & Ryan, T. A. Differential control of inhibitory and excitatory nerve terminal function by mitochondria. Preprint at bioRxiv https://doi.org/10.1101/2024.05.19.594864 (2024).
Bredvik, K., Liu, C. & Ryan, T. A. Characterization of β-hydroxybutyrate as a cell autonomous fuel for active excitatory and inhibitory neurons. Preprint at bioRxiv https://doi.org/10.1101/2024.06.08.598077 (2024).
Zampese, E. et al. Ca2+ channels couple spiking to mitochondrial metabolism in substantia nigra dopaminergic neurons. Sci. Adv. 8, eabp8701 (2022).
Del Arco, A. et al. Regulation of neuronal energy metabolism by calcium: role of MCU and Aralar/malate–aspartate shuttle. Biochim. Biophys. Acta Mol. Cell. Res. 1870, 119468 (2023).
Llorente-Folch, I. et al. Calcium-regulation of mitochondrial respiration maintains ATP homeostasis and requires ARALAR/AGC1-malate aspartate shuttle in intact cortical neurons. J. Neurosci. 33, 13957–13971 (2013).
Baeza-Lehnert, F. et al. Non-canonical control of neuronal energy status by the Na+ pump. Cell Metab. 29, 668–680 (2019).
Vilas-Boas, E. A., Cabral-Costa, J. V., Ramos, V. M., Caldeira da Silva, C. C. & Kowaltowski, A. J. Goldilocks calcium concentrations and the regulation of oxidative phosphorylation: too much, too little, or just right. J. Biol. Chem. 299, 102904 (2023).
Li, S., Xiong, G. -J., Huang, N. & Sheng, Z. -H. The cross-talk of energy sensing and mitochondrial anchoring sustains synaptic efficacy by maintaining presynaptic metabolism. Nat. Metab. 2, 1077–1095 (2020).
Denton, R. M. Regulation of mitochondrial dehydrogenases by calcium ions. Biochim. Biophys. Acta 1787, 1309–1316 (2009).
Glancy, B. & Balaban, R. S. Role of mitochondrial Ca2+ in the regulation of cellular energetics. Biochemistry 51, 2959–2973 (2012).
Palty, R. et al. Lithium–calcium exchange is mediated by a distinct potassium-independent sodium-calcium exchanger. J. Biol. Chem. 279, 25234–25240 (2004).
Palty, R. et al. NCLX is an essential component of mitochondrial Na+/Ca2+ exchange. Proc. Natl Acad. Sci. USA 107, 436–441 (2010).
Jiang, D., Zhao, L. & Clapham, D. E. Genome-wide RNAi screen identifies Letm1 as a mitochondrial Ca2+/H+ antiporter. Science 326, 144–147 (2009).
Shao, J. et al. Leucine zipper-EF-hand containing transmembrane protein 1 (LETM1) forms a Ca2+/H+ antiporter. Sci. Rep. 6, 34174 (2016).
Tsai, M. -F., Jiang, D., Zhao, L., Clapham, D. & Miller, C. Functional reconstitution of the mitochondrial Ca2+/H+ antiporter Letm1. J. Gen. Physiol. 143, 67–73 (2013).
Austin, S. & Nowikovsky, K. LETM1: essential for mitochondrial biology and cation homeostasis?. Trends Biochem. Sci. 44, 648–658 (2019).
Lewis, T. L., Kwon, S. -K., Lee, A., Shaw, R. & Polleux, F. MFF-dependent mitochondrial fission regulates presynaptic release and axon branching by limiting axonal mitochondria size. Nat. Commun. 9, 5008 (2018).
Stavsky, A. et al. Aberrant activity of mitochondrial NCLX is linked to impaired synaptic transmission and is associated with mental retardation. Commun. Biol. 4, 666 (2021).
Jang, D. C. et al. Asymmetric distribution of mitochondrial Ca2+ regulators specifies compartment-specific mitochondrial function and neuronal development. Preprint at bioRxiv https://doi.org/10.1101/2025.06.23.660978 (2025).
Hagenston, A. M. et al. Disrupted expression of mitochondrial NCLX sensitizes neuroglial networks to excitotoxic stimuli and renders synaptic activity toxic. J. Biol. Chem. 298, 101508 (2022).
Ruiz, A., Alberdi, E. & Matute, C. CGP37157, an inhibitor of the mitochondrial Na+/Ca2+ exchanger, protects neurons from excitotoxicity by blocking voltage-gated Ca2+ channels. Cell Death Dis. 5, e1156–e1156 (2014).
Qiu, J. et al. Mitochondrial calcium uniporter Mcu controls excitotoxicity and is transcriptionally repressed by neuroprotective nuclear calcium signals. Nat. Commun. 4, 2034 (2013).
Hirabayashi, Y. et al. ER-mitochondria tethering by PDZD8 regulates Ca2+ dynamics in mammalian neurons. Science 358, 623–630 (2017).
Zhou, Z., Matlib, M. A. & Bers, D. M. Cytosolic and mitochondrial Ca2+ signals in patch clamped mammalian ventricular myocytes. J. Physiol. 507, 379–403 (1998).
Di Lisa, F., Gambassi, G., Spurgeon, H. & Hansford, R. G. Intramitochondrial free calcium in cardiac myocytes in relation to dehydrogenase activation. Cardiovasc. Res. 27, 1840–1844 (1993).
Miyata, H. et al. Measurement of mitochondrial free Ca2+ concentration in living single rat cardiac myocytes. Am. J. Physiol. 261, H1123–H1134 (1991).
Grieves, R. M., Wood, E. R. & Dudchenko, P. A. Place cells on a maze encode routes rather than destinations. eLife 5, e15986 (2016).
McNay, E. C. & Gold, P. E. Extracellular glucose concentrations in the rat hippocampus measured by zero-net-flux: effects of microdialysis flow rate, strain, and age. J. Neurochem. 72, 785–790 (1999).
McNay, E. C., Fries, T. M. & Gold, P. E. Decreases in rat extracellular hippocampal glucose concentration associated with cognitive demand during a spatial task. Proc. Natl Acad. Sci. USA 97, 2881–2885 (2000).
Juan-Sanz et al. Axonal endoplasmic reticulum Ca2+ content controls release probability in CNS nerve terminals. Neuron 93, 867–881 (2017).
Endele, S., Fuhry, M., Pak, S. -J., Zabel, B. U. & Winterpacht, A. LETM1, a novel gene encoding a putative EF-hand Ca2+-binding protein, flanks the Wolf–Hirschhorn Syndrome (WHS) critical region and is deleted in most WHS patients. Genomics 60, 218–225 (1999).
Lin, Q. -T., Lee, R., Feng, A. L., Kim, M. S. & Stathopulos, P. B. The leucine zipper EF-hand containing transmembrane protein-1 EF-hand is a tripartite calcium, temperature, and pH sensor. Protein Sci. 30, 855–872 (2021).
Gifford, J. L., Walsh, M. P. & Vogel, H. J. Structures and metal-ion-binding properties of the Ca2+-binding helix–loop–helix EF-hand motifs. Biochem. J 405, 199–221 (2007).
Zhang, Y. et al. Fast and sensitive GCaMP calcium indicators for imaging neural populations. Nature 615, 884–891 (2023).
Delgado, T. et al. Comparing 3D ultrastructure of presynaptic and postsynaptic mitochondria. Biol. Open 8, bio044834 (2019).
Faitg, J. et al. 3D neuronal mitochondrial morphology in axons, dendrites, and somata of the aging mouse hippocampus. Cell Rep. 36, 109509 (2021).
Lin, Y. et al. Brain activity regulates loose coupling between mitochondrial and cytosolic Ca2+ transients. Nat. Commun. 10, 5277 (2019).
Belle, A. M. et al. Evaluation of in vitro neuronal platforms as surrogates for in vivo whole brain systems. Sci. Rep. 8, 10820 (2018).
Cadena, J. et al. Modeling the temporal network dynamics of neuronal cultures. PLoS Comput. Biol. 16, e1007834 (2020).
Kotera, I., Iwasaki, T., Imamura, H., Noji, H. & Nagai, T. Reversible dimerization of aequorea victoria fluorescent proteins increases the dynamic range of FRET-based indicators. ACS Chem. Biol. 5, 215–222 (2010).
Imamura, H. et al. Visualization of ATP levels inside single living cells with fluorescence resonance energy transfer-based genetically encoded indicators. Proc. Natl Acad. Sci. USA 106, 15651–15656 (2009).
Finkel, T. et al. The ins and outs of mitochondrial calcium. Circ. Res. 116, 1810–1819 (2015).
Williams, G. S. B., Boyman, L. & Lederer, W. J. Mitochondrial calcium and the regulation of metabolism in the heart. J. Mol. Cell. Cardiol. 78, 35–45 (2015).
Llorente-Folch, I. et al. The regulation of neuronal mitochondrial metabolism by calcium. J. Physiol. 593, 3447–3462 (2015).
Chen, J. et al. Identification of novel modulators of mitochondrial function by a genome-wide RNAi screen in Drosophila melanogaster. Genome Res. 18, 123–136 (2008).
Dung, V. M. et al. Neuron-specific knockdown of Drosophila PDHB induces reduction of lifespan, deficient locomotive ability, abnormal morphology of motor neuron terminals and photoreceptor axon targeting. Exp. Cell. Res. 366, 92–102 (2018).
Harris, R. A., Bowker-Kinley, M. M., Huang, B. & Wu, P. Regulation of the activity of the pyruvate dehydrogenase complex. Adv. Enzyme Regul. 42, 249–259 (2002).
Lander, N. et al. Calcium-sensitive pyruvate dehydrogenase phosphatase is required for energy metabolism, growth, differentiation, and infectivity of Trypanosoma cruzi. J. Biol. Chem. 293, 17402–17417 (2018).
Kumar, V. & Greenberg, M. L. Emerging roles of pyruvate dehydrogenase phosphatase 1: a key player in metabolic health. Front. Physiol. 16, 1596636 (2025).
Yang, W. et al. Rheb mediates neuronal-activity-induced mitochondrial energetics through mTORC1-independent PDH activation. Dev. Cell 56, 811–825 (2021).
Tiwari, A. et al. Sirtuin3 ensures the metabolic plasticity of neurotransmission during glucose deprivation. Preprint at bioRxiv https://doi.org/10.1101/2023.03.08.531724 (2023).
Koshenov, Z., Kokotos, A. C., Benedetti, L., Schwartz, J. L. & Ryan, T. A. A genetic driver of epileptic encephalopathy impairs gating of synaptic glycolysis. Preprint at bioRxiv https://doi.org/10.1101/2025.06.17.660213 (2025).
Durigon, R. et al. LETM1 couples mitochondrial DNA metabolism and nutrient preference. EMBO Mol. Med. 10, e8550 (2018).
Hasegawa, A. & van der Bliek, A. M. Inverse correlation between expression of the Wolfs Hirschhorn candidate gene Letm1 and mitochondrial volume in C. elegans and in mammalian cells. Hum. Mol. Genet. 16, 2061–2071 (2007).
Shi, Y. et al. Aberrant LETM1 elevation dysregulates mitochondrial functions and energy metabolism and promotes lung metastasis in osteosarcoma. Genes Dis. 11, 100988 (2024).
Piao, L. et al. Association of LETM1 and MRPL36 contributes to the regulation of mitochondrial ATP production and necrotic cell death. Cancer Res. 69, 3397–3404 (2009).
Sankaranarayanan, S., De Angelis, D., Rothman, J. E. & Ryan, T. A. The use of phluorins for optical measurements of presynaptic activity. Biophys. J. 79, 2199–2208 (2000).
Nazaret, C., Heiske, M., Thurley, K. & Mazat, J. -P. Mitochondrial energetic metabolism: a simplified model of TCA cycle with ATP production. J. Theor. Biol. 258, 455–464 (2009).
Denton, R. M. & McCormack, J. G. On the role of the calcium transport cycle in heart and other mammalian mitochondria. FEBS Lett. 119, 1–8 (1980).
Territo, P. R., Mootha, V. K., French, S. A. & Balaban, R. S. Ca2+ activation of heart mitochondrial oxidative phosphorylation: role of the F0/F1-ATPase. Am. J. Physiol. 278, C423–C435 (2000).
Martín, A. S. et al. Imaging mitochondrial flux in single cells with a FRET sensor for pyruvate. PLoS ONE 9, e85780 (2014).
Fernández-Moncada, I. et al. Neuronal control of astrocytic respiration through a variant of the Crabtree effect. Proc. Natl Acad. Sci. USA 115, 1623–1628 (2018).
Davie, K. et al. A single-cell transcriptome atlas of the aging Drosophila brain. Cell 174, 982–998 (2018).
Tully, T., Preat, T., Boynton, S. C. & Del Vecchio, M. Genetic dissection of consolidated memory in Drosophila. Cell 79, 35–47 (1994).
Tully, T. & Quinn, W. G. Classical conditioning and retention in normal and mutant Drosophila melanogaster. J. Comp. Physiol. A 157, 263–277 (1985).
Bouzaiane, E., Trannoy, S., Scheunemann, L., Plaçais, P. -Y. & Preat, T. Two independent mushroom body output circuits retrieve the six discrete components of Drosophila aversive memory. Cell Rep. 11, 1280–1292 (2015).
Comyn, T., Preat, T., Pavlowsky, A. & Plaçais, P. -Y. PKCδ is an activator of neuronal mitochondrial metabolism that mediates the spacing effect on memory consolidation. eLife 13, RP92085 (2024).
Wolff, G. H. & Strausfeld, N. J. Genealogical correspondence of mushroom bodies across invertebrate phyla. Curr. Biol. 25, 38–44 (2015).
Wolff, G. H. & Strausfeld, N. J. Genealogical correspondence of a forebrain centre implies an executive brain in the protostome–deuterostome bilaterian ancestor. Philos. Trans. R. Soc. B Biol. Sci. 371, 20150055 (2016).
Pascual, A. & Préat, T. Localization of long-term memory within the Drosophila mushroom body. Science 294, 1115–1117 (2001).
Yu, D., Akalal, D. -B. G. & Davis, R. L. Drosophila α/β mushroom body neurons form a branch-specific, long-term cellular memory trace after spaced olfactory conditioning. Neuron 52, 845–855 (2006).
Pérez-Moreno, J. J. et al. Drosophila SPG12 ortholog, reticulon-like 1, governs presynaptic ER organization and Ca2+ dynamics. J. Cell Biol. 222, e202112101 (2023).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Chang, K., Marran, K., Valentine, A. & Hannon, G. J. Creating an miR30-Based shRNA vector. Cold Spring Harb. Protoc. 2013, 631–635 (2013).
Aqrabawi, A. J. & Kim, J. C. Hippocampal projections to the anterior olfactory nucleus differentially convey spatiotemporal information during episodic odour memory. Nat. Commun. 9, 2735 (2018).
Tanaka, K. Z. et al. Cortical representations are reinstated by the hippocampus during memory retrieval. Neuron 84, 347–354 (2014).
Grella, S. L., Fortin, A. H., McKissick, O., Leblanc, H. & Ramirez, S. Odor modulates the temporal dynamics of fear memory consolidation. Learn. Mem. 27, 150–163 (2020).
Yu, S. B. et al. Neuronal activity-driven O-GlcNAcylation promotes mitochondrial plasticity. Dev. Cell 59, 2143–2157.e9 (2024).
Kumar, M. et al. Triglycerides are an important fuel reserve for synapse function in the brain. Nat. Metab. 7, 1392–1403 (2025).
Paucek, P. & Jabůrek, M. Kinetics and ion specificity of Na+/Ca2+ exchange mediated by the reconstituted beef heart mitochondrial Na+/Ca2+ antiporter. Biochim. Biophys. Acta 1659, 83–91 (2004).
Vercesi, A. E. et al. Mitochondrial calcium transport and the redox nature of the calcium-induced membrane permeability transition. Free Radic. Biol. Med. 129, 1–24 (2018).
Garbincius, J. F. & Elrod, J. W. Mitochondrial calcium exchange in physiology and disease. Physiol. Rev. 102, 893–992 (2022).
Santo-Domingo, J. & Demaurex, N. Calcium uptake mechanisms of mitochondria. Biochim. Biophys. Acta 1797, 907–912 (2010).
Austin, S. et al. LETM1-mediated K+ and Na+ homeostasis regulates mitochondrial Ca2+ efflux. Front. Physiol. 8, 839 (2017).
Froschauer, E., Nowikovsky, K. & Schweyen, R. J. Electroneutral K+/H+ exchange in mitochondrial membrane vesicles involves Yol027/Letm1 proteins. Biochim. Biophys. Acta 1711, 41–48 (2005).
Midgley, P. J., Rutter, G. A., Thomas, A. P. & Denton, R. M. Effects of Ca2+ and Mg2+ on the activity of pyruvate dehydrogenase phosphate phosphatase within toluene-permeabilized mitochondria. Biochem. J. 241, 371–377 (1987).
Fernandez, A. et al. Mitochondrial dysfunction leads to cortical under-connectivity and cognitive impairment. Neuron 102, 1127–1142 (2019).
Kapogiannis, D. & Mattson, M. P. Disrupted energy metabolism and neuronal circuit dysfunction in cognitive impairment and Alzheimer’s disease. Lancet Neurol. 10, 187–198 (2011).
Sharma, C., Kim, S., Nam, Y., Jung, U. J. & Kim, S. R. Mitochondrial dysfunction as a driver of cognitive impairment in Alzheimer’s disease. Int. J. Mol. Sci. 22, 4850 (2021).
Messier, C. Glucose improvement of memory: a review. Eur. J. Pharmacol. 490, 33–57 (2004).
Gold, P. Role of glucose in regulating the brain and cognition. Am. J. Clin. Nutr. 61, 987S–995S (1995).
Messier, C. Object recognition in mice: improvement of memory by glucose. Neurobiol. Learn. Mem. 67, 172–175 (1997).
Papa, S. & Skulachev, V. P. Reactive oxygen species, mitochondria, apoptosis and aging. Mol. Cell. Biochem. 174, 305–319 (1997).
Lopez-Fabuel, I. et al. Complex I assembly into supercomplexes determines differential mitochondrial ROS production in neurons and astrocytes. Proc. Natl Acad. Sci. USA 113, 13063–13068 (2016).
Jakkamsetti, V. et al. Brain metabolism modulates neuronal excitability in a mouse model of pyruvate dehydrogenase deficiency. Sci. Transl. Med. 11, eaan0457 (2019).
Martínez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Alexander, D. C. et al. Targeting acetyl-CoA metabolism attenuates the formation of fear memories through reduced activity-dependent histone acetylation. Proc. Natl Acad. Sci. USA 119, e2114758119 (2022).
Currais, A. et al. Elevating acetyl-CoA levels reduces aspects of brain aging. eLife 8, e47866 (2019).
Santoni, G. et al. Chromatin plasticity predetermines neuronal eligibility for memory trace formation. Science 385, eadg9982 (2024).
Bayer, K. U. & Giese, K. P. A revised view of the role of CaMKII in learning and memory. Nat. Neurosci. 28, 24–34 (2025).
Wang, W. et al. Damaged mitochondria coincide with presynaptic vesicle loss and abnormalities in alzheimer’s disease brain. Acta Neuropathol. Commun. 11, 54 (2023).
Du, H. et al. Early deficits in synaptic mitochondria in an Alzheimer’s disease mouse model. Proc. Natl Acad. Sci. USA 107, 18670–18675 (2010).
Swerdlow, R. H. The neurodegenerative mitochondriopathies. J. Alzheimers Dis. 17, 737–751 (2009).
Shil, S. K. et al. Ndufs4 ablation decreases synaptophysin expression in hippocampus. Sci. Rep. 11, 10969 (2021).
Drago, I. & Davis, R. L. Inhibiting the mitochondrial calcium uniporter during development impairs memory in adult Drosophila. Cell Rep. 16, 2763–2776 (2016).
Tiwari, A. et al. Mitochondrial pyruvate transport regulates presynaptic metabolism and neurotransmission. Sci. Adv. 10, eadp7423 (2024).
Cruz et al. Specific hippocampal interneurons shape consolidation of recognition memory. Cell Rep. 32, 108046 (2020).
Pinho, J. S., Ramon-Duaso, C., Manzanares-Sierra, I. & Busquets-García, A. Dorsal hippocampus mediates light-tone sensory preconditioning task in mice. eLife 14, RP105863 (2025).
Turrel, O., Goguel, V. & Preat, T. Amnesiac is required in the adult mushroom body for memory formation. J. Neurosci. 38, 9202–9214 (2018).
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003).
Pfeiffer, B. D. et al. Refinement of tools for targeted gene expression in Drosophila. Genetics 186, 735–755 (2010).
Fabritius, A. et al. Imaging-based screening platform assists protein engineering. Cell Chem. Biol. 25, 1554–1561 (2018).
Sun, M., Bernard, L. P., DiBona, V. L., Wu, Q. & Zhang, H. Calcium phosphate transfection of primary hippocampal neurons. J. Vis. Exp. https://doi.org/10.3791/50808 (2013).
Dudek, H., Ghosh, A. & Greenberg, M. E. Calcium phosphate transfection of DNA into neurons in primary culture. Curr. Protoc. Neurosci. 3, 3.11.1–3.11.6 (1998).
Chen, T. -W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Studier, F. W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Tsien, R. & Pozzan, T. Measurement of cytosolic free Ca2+ with quin2. Methods Enzymol. 172, 230–262 (1989).
Loh, K. H. et al. Proteomic analysis of unbounded cellular compartments: synaptic clefts. Cell 166, 1295–1307 (2016).
Shen, Y., Rosendale, M., Campbell, R. E. & Perrais, D. pHuji, a pH-sensitive red fluorescent protein for imaging of exo- and endocytosis. J. Cell Biol. 207, 419–432 (2014).
Merrill, R. A., Flippo, K. H. & Strack, S. in Techniques to Investigate Mitochondrial Function in Neurons (eds. Strack, S. et al.) 31–48 (Springer, 2017).
Mohylyak, I. et al. Temporal transcriptional regulation of mitochondrial morphology primes activity-dependent circuit connectivity. Nat. Commun. 16, 8173 (2025).
Scheunemann, L., Plaçais, P. -Y., Dromard, Y., Schwärzel, M. & Preat, T. Dunce phosphodiesterase acts as a checkpoint for Drosophila long-term memory in a pair of serotonergic neurons. Neuron 98, 350–365 (2018).
Silva, B. et al. Glia fuel neurons with locally synthesized ketone bodies to sustain memory under starvation. Nat. Metab. 4, 213–224 (2022).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
von Ziegler, L. M. et al. Analysis of behavioral flow resolves latent phenotypes. Nat. Methods 21, 2376–2387 (2024).
Acknowledgements
We thank all members of the laboratory of J.d.J.-S. for insightful discussions and comments. We thank S. Perez for technical assistance. This work was made possible by the Paris Brain Institute Diane Barriere Chair in Synaptic Bioenergetics awarded to J.d.J.-S., who is also supported by an ERC Starting Grant (SynaptoEnergy, European Research Council; ERC-StG-852873), 2019 ATIP-Avenir Grant (CNRS, Inserm), a Big Brain Theory Grant (ICM Foundation) and a Kavli Exploratory Award (Kavli Foundation). This work was also supported by an ERC Advanced Grant (EnergyMeMo; ERC-AdG-741550) to T.P. and grants from the Agence Nationale de la Recherche to P.Y.P. (ANR-20-CE92-0047-01), T.P. (ANR-23-CE16-0029-01), A.P. and J.d.J.-S. (ANR-22-CE16-0020) and J.d.J.-S. (ANR-24-CE16-0221). T.P., P.Y.P. and J.d.J.-S. are permanent CNRS researchers. A.P. is a permanent ESPCI associate professor. T.C. was funded by the French Ministry of Research and the Fondation pour la Recherche Médicale. V.R. was funded by the Max Planck Society, the Chan Zuckerberg Initiative DAF, an advised fund of the Silicon Valley Community Foundation grant number 2024-349543 and the NIH Director’s New Innovator Award (DP2 MH140148). A.B.-G. and C.R.-D. received funding from an ERC Starting Grant (HighMemory; ERC-StG-948217), the Ministry of Economy and Competitiveness (PID2021-122795OB-I00) and the Departament d’Economia i Coneixement de la Generalitat de Catalunya (SGR 00022). T.P.V. was funded by the Wellcome Trust and a Royal Society Sir Henry Dale Research Fellowship (WT100000) and a Wellcome Trust Senior Research Fellowship (214316/Z/18/Z). K.G. was supported by the DIM C-BRAINS, funded by the Conseil Régional d’Ile-de-France. The contributions of H.F. and E.R.S. were supported by the Howard Hughes Medical Institute. The PHENO-ICMice animal Core at ICM is supported by two ‘Investissements d’avenir’ (ANR-10- IAIHU-06 and ANR-11-INBS-0011-NeurATRIS) and the Fondation pour la Recherche Médicale.
Author information
Authors and Affiliations
Contributions
Conceptualization, A.A.V. and J.d.J.-S.; methodology, A.A.V., T.C., R.G.M., C.B., C.P-C., M.F., K.B., C.C., C.R.-D., R.F., K.G., H.F., R.S., T.P., T.P.V., V.R., A.B.-G., P.-Y.P., A.P. and J.d.J.-S.; investigation, A.A.V., T.C., R.G.M., C.B., C.P-C., M.F., K.B., C.C., C.R.-D., R.F., K.G., H.F., J.-P.B., E.R.S., T.P., T.P.V., V.R., A.B.-G., P.-Y.P., A.P. and J.d.J.-S.; project administration and supervision, J.d.J.-S.; writing—original draft, A.A.V. and J.d.J.-S.; writing—review and editing, A.A.V., T.C., R.G.M., C.B., C.P-C., M.F., K.B., C.C., C.R.-D., R.F., K.G., H.F., J.-P.B., E.R.S., T.P., T.P.V., V.R., A.B.-G., P.-Y.P., A.P. and J.d.J.-S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks L. Felipe Barros and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Gimenez-Cassina, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Related to Fig. 1. Validation of the effects of Letm1 KD in primary hippocampal rat neurons.
(A) Representative immunofluorescence images of co-cultured neurons and astrocytes stained for MAP2 (neuronal marker, red), GFAP (astrocyte marker, green), and DAPI (nuclei, blue). Scale bar: 100 µm. This experiment was performed in two independent experiments with 5 technical replicates. (B) Quantification of cell-type composition within the cultures. Pie chart shows the percentage of neurons (MAP2+), astrocytes (GFAP+) and undefined cells (DAPI+/MAP2−/GFAP−). Data indicate neurons constitute 12.4% of the total cell population. (C) Representative western blot showing levels of Letm1 and β-actin proteins in lysates from pure neuronal cultures of control and Letm1 KD neurons. (D) Densitometric quantification of Letm1 levels in control and Letm1 KD neurons. P value was determined using the two-tailed Mann–Whitney U-test (**P = 0.0022; n = 6 neurons for each group; U = 0). (E) Traces represent the mean ± SEM of Mito4x-GCaMP6f in axons stimulated with 100AP at 100 Hz in 20 mM glucose. (F) Quantification of peak (left) and half-time decay (t1/2, right) of Mito4x-GCaMP6f responses (ΔF/F0) after 100AP 100 Hz stimulation in high glucose (20 mM) Tyrode’s solution. P value for peaks was determined using the two-tailed Mann–Whitney U-test (n.s., P = 0.8518; n = 6 neurons for control and n = 8 for Letm1 KD; U = 22). P value for t1/2 was determined using a two-tailed unpaired t‑test (**P = 0.008; n = 6 neurons for control and n = 8 for Letm1 KD; U = 4). (G) Quantification of peak (left) and half-time decay (t1/2, right) of Mito4x-GCaMP6f responses (ΔF/F0) after 100AP 100 Hz stimulation in physiological glucose (1.2 mM) Tyrode’s solution. P value for peaks was determined using the two-tailed Mann–Whitney U-test (n.s., P = 0.0501; n = 31 neurons for control and n = 28 for Letm1 KD; U = 305). P value for t1/2 was determined using a two-tailed Mann-Whitney U test (**P = 0.0025; n = 31 neurons for control and n = 28 for Letm1 KD; U = 237.5). (H) Quantification of peak (left) and of Mito4x-GCaMP6f responses (ΔF/F0) after 20AP 20 Hz stimulation in physiological glucose (1.2 mM) Tyrode’s solution. P value for peaks was determined using a two-tailed Mann–Whitney U test (n.s., P = 0.4609 n = 29 neurons for control and n = 28 for Letm1 KD; U = 359). P value for t1/2 was determined using a two-tailed Mann–Whitney U-test (**P = 0.0026; n = 29 neurons for control and n = 28 for Letm1 KD; U = 0.0026). (I) Estimates of mitochondrial Ca2+ at rest using two-point or one-point calibration methods. P values were determined using a Kruskal-Wallis Test (P = 0.0065; n = 16 for control and n = 11 for Letm1 KD in two-point calibrations, and n = 13 for control and n = 12 for Letm1 KD in one-point calibrations, followed by Dunn’s multiple comparisons test. Post hoc P values: Two-point, Control vs Letm1 KD, n.s. P > 0.9999; One-point, Control vs Letm1 KD, n.s. P > 0.9999; Control, two-point vs one-point, n.s. P = 0.105; Letm1 KD, two-point vs one-point, n.s. P = 0.0623). (J) Representative images of two-point calibration of Mito4x-GCaMP6f fluorescence to obtain minimum fluorescence (Fmin) and saturation of the sensor (Fmax). Pseudocolor calibration bar indicates range of fluorescence. Scale bar 8 µm. (K) Estimated matrix Ca2+ peak during 20AP 20 Hz or 100AP 100 Hz (nM) in physiological glucose (1.2 mM) Tyrode’s solution. P value for 20 Hz was determined using a two-tailed Mann–Whitney U-test (n.s., P = 0.4269; n = 28 neurons for control and n = 27 for Letm1 KD; U = 330). P value for 100 Hz was determined using a two-tailed unpaired t-test (n.s., P = 0.0820; n = 19 neurons for control and n = 22 for Letm1 KD; t39 = 1.785). (L) Representative image of an axon of a neuron transfected with Mito4x-GCaMP6f and Letm1-mRFP1, showing co-localization in mitochondria. The scale bar is 4 μm. This pattern was observed in 12 independently transfected neurons. (M) Traces represent the mean ± SEM of cytoGCaMP8f responses in axonal varicosities following stimulation of 20AP 20 Hz. (N) Peak cytoGCaMP8f responses (ΔF/F0) during stimulation in control and Letm1 KD neurons. P value was determined using a two-tailed unpaired t‑test (n.s., P = 0.7374; n = 32 neurons for control and n = 36 for Letm1 KD; t66 = 0.3367). (O) Rate of cytosolic Ca2+ decay (t1/2) in presynaptic varicosities following 20AP 20 Hz stimulation in control and Letm1 KD neurons. P value was determined using the two-tailed Mann–Whitney U-test (n.s., P = 0.37; n = 32 neurons for control and n = 36 for Letm1 KD; U = 502.5). (P) Resting levels of cytosolic Ca2+ in control and Letm1 KD neurons. P value was determined using a two-tailed Unpaired t-test (n.s. P = 0.3634; n = 10 neurons for control and Letm1 KD; t18 = 0.9326).
Extended Data Fig. 2 Related to Fig. 1. Axonal pH calibrations in cytosol and mitochondria and dendrite Ca2+ responses in cytosol and mitochondria.
(A) Representative responses of Mito4x-pHluorin to NH4Cl-mediated mitochondrial pH clamping used to estimate resting mitochondrial pH. Tyrode’s solutions at pH 7.4, 7.8, and 8.2 were sequentially perfused, each containing 100 mM NH4Cl. (B) Resting mitochondrial pH was estimated using fluorescence values before and after NH4Cl perfusion at pH 7.8 or pH 8.2, or using the average of both values. P values were determined using two-tailed Mann-Whitney U-tests (pH 7.8; n.s., P = 0.777; n = 17 for Control, n = 12 for Letm1 KD neurons; U = 95; pH 8.2; n.s., P = 0.527; n = 17 for Control, n = 12 for Letm1 KD neurons; U = 87; average from 7.8 and 8.2; n.s., P = 0.7438; n = 17 for Control, n = 12 for Letm1 KD neurons; U = 94). (C) Representative responses of Cyto-pHluorin to nigericin-mediated cytosolic pH clamping used to estimate resting cytosolic pH. Tyrode’s solutions at pH 7.4, 7.8, and 8.2 were sequentially perfused, each containing 20 µM nigericin. (D) Resting cytosolic pH was estimated using fluorescence values before and after nigericin perfusion at pH 7.8 or pH 8.2, or using the average of both values. P values were determined using two-tailed Mann-Whitney U-tests (pH 7.8; n.s., P = 0.1802; n = 12 for Control, n = 10 for Letm1 KD neurons; U = 39) (pH 8.2; n.s., P = 0.107; n = 12 for Control, n = 10 for Letm1 KD neurons; U = 35) (average from 7.8 and 8.2; n.s., P = 0.159; n = 12 for Control, n = 10 for Letm1 KD neurons; U = 38). (E) (left) Peak Mito4x-pHluorin responses (ΔF/F0) after 20AP 20 Hz stimulation in control and Letm1 KD neurons. P value was determined using a two-tailed Mann–Whitney U-test (n.s., P = 0.0971; n = 18 neurons per group; U = 109). (right) Peak Cyto-pHluorin responses (ΔF/F0) in axons of control and Letm1 KD neurons stimulated with 600APs at 10 Hz. P value was determined using a two-tailed unpaired t-test (n.s., P = 0.4223; n = 6 neurons per group; t10 = 0.8366). (F) Peak dendritic mitochondrial calcium responses in control and Letm1 KD neurons. P value was determined using a two-tailed Mann–Whitney U-test (***P = 0.0006; n = 63 neurons for control and n = 47 for Letm1 KD; U = 916). (G) Traces represent the mean ± SEM of cytosolic calcium responses in neuronal dendrites measured using RCaMP1.07 following stimulation with a single pulse of glutamate uncaging. (H) Peak RCaMP1.07 responses (ΔF/F0) following stimulation in control and Letm1 KD neurons. P value was determined using a two-tailed Mann–Whitney U-test (n.s., P = 0.5512; n = 63 dendrites for control and n = 47 dendrites for Letm1 KD; U = 1381). (I) Area under the curves (AUC) for stimulation-induced dendritic and axonal mitochondrial Ca2+ responses. When comparing dendrite responses, P value was determined using a two-tailed Mann–Whitney U-test (n.s., P = 0.3504; n = 63 neurons for control and n = 47 for Letm1 KD; U = 1325). When comparing axonal responses, P value was determined using a two-tailed Mann–Whitney U-test (**P = 0.0016; n = 46 neurons for control and n = 34 for Letm1 KD; U = 462). Data are represented as mean ± SEM.
Extended Data Fig. 3 Related to Fig. 2. Metabolic and structural phenotypes of Letm1 KD neurons.
(A) Representative images of neurons expressing Syn-ATP. From left to right, neurons are 1) live, 2) permeabilized and with 0 mM ATP and 3) permeabilized and with 5 mM ATP (see methods). (B-C) Non-calibrated values corresponding to main Fig. 2b, shown as ratio of luminescence by fluorescence (L/F). (B) P values were determined using a Kruskal-Wallis test followed by Dunn’s multiple comparisons test (P = 0.018; n = 30 for control, n = 28 for Letm1 KD, n = 7 for Letm1/PDP1 KD. Post hoc P values: Control vs Letm1 KD, **P = 0.0064; Control vs Letm1/PDP1 KD, n.s., P > 0.9999; Letm1 KD vs Letm1/PDP1 KD, *P = 0.0214. (C) ATP levels in the presence of TTX. P value was determined using a two-tailed unpaired two-tailed t-test. (n.s., P > 0.9208; n = 8 for Control + TTX, n = 9 for Letm1 KD + TTX; t15 = 0.1011). Experiments shown in A-C were done in 1.25 mM lactate and 1.25 pyruvate with no glucose. (D) Strategy used to simultaneously KD PDP1 and Letm1. H1 and U6 are RNA Pol III promoters. PGK is a promoter to express mTagBFP2 to confirm transfection. (E) Western blot of pure neuronal cultures to knock-down Letm1 or Letm1 and PDP1 using lentivirus. (F) Quantification of KD for Letm1 and PDP1 using the dual vector strategy. P values were determined using a one-sample Wilcoxon signed rank test (**P = 0.078 for Letm1 KD, *P = 0.0156 for PDP1 KD; n = 8 for Letm1 KD and n = 7 for PDP1 KD; W = -36, -28). (G) Synaptic ATP levels in Control and Letm1 KD neurons upon stimulation with 600APs at 10 Hz. The dotted line indicates baseline ATP levels before stimulation. L/F values are normalized to the baseline. Data are shown as mean±SEM (n = 11). (H) Stimulation induced changes in synaptic ATP levels (ΔL/F) in control and Letm1 KD neurons. P value was determined using a two-tailed unpaired two-tailed t-test (*P = 0.0137; n = 11 for both control and Letm1 KD neurons; t20 = 2.703). (I) Representative Western blot of Letm1 KD pure neurons, showing Letm1, Syphy1 (synaptophysin 1), PSD95, CI-NDUFB8, CII-SDHB, CIII-UQCRC2, COXIV, CV-ATP5A and β-actin. (J) Protein levels are normalized to control. P values were calculated using two-tailed one-sample t-tests: Letm1 (**** P < 0.0001, t₃ = 32.30), Syphy1 (ns P = 0.5755, t₃ = 0.6263), PSD95 (ns P = 0.0659, t₃ = 2.835), Complex I (ns P = 0.6821, t₃ = 0.4518), Complex II (n.s., P = 0.5133, t₃ = 0.7394), Complex III (n.s., P = 0.7529, t₃ = 0.3450), Complex IV (n.s., P = 0.2027, t₃ = 1.625), Complex V (n.s., P = 0.4177, t₃ = 0.9375). (K-M) Axonal mitochondrial shape properties measured using Mito4x-GCaMP6f. Circularity is measured in arbitrary units (a.u.), 1 = circular; 0 = non-circular. P values were determined using two-tailed Mann–Whitney U-tests for circularity (n.s., P = 0.0925; n = 29 neurons for control, n = 22 for Letm1 KD; U = 230) and size (*P = 0.0160; n = 29 neurons for control, n = 22 for Letm1 KD; U = 193), and using a two-tailed unpaired t-test for mitochondria number (*P = 0.0115, n = 8 for both conditions; t14 = 2.906). (N) Number of synapses per 100 µm measured from synaptophysin-mCherry signal from SynATP. P values were determined using a ordinary one-way ANOVA (**P = 0.0019; F(2,24) = 0.2812; n = 9 for control, n = 11 for Letm1 KD, n = 7 for Letm1/PDP1 KD, followed by Dunn’s multiple comparisons test. Post hoc P values: Control vs Letm1 KD, *P = 0.0135; Control vs Letm1/PDP1 KD, n.s., P = 0.7126; Letm1 KD vs Letm1/PDP1 KD, **P = 0.0032). (O) Estimated synapse size, shown as area in µm2. P values were determined using a one-way ANOVA with n = 23 for control, n = 22 for Letm1 KD, n = 7 for Letm1/PDP1 KD, followed by Tukey’s multiple comparisons test. Post hoc P values: Control vs Letm1 KD, **P = 0.0031; Control vs Letm1/PDP1 KD, n.s., P > 0.9999; Letm1 KD vs Letm1/PDP1 KD, *P = 0.0469. (P) Quantification of synaptic vesicle cycling using vGlut-pHluorin. The figure shows traces corresponding to the mean ± SEM of responses normalized to the total synaptic vesicle pool, obtained by applying NH4Cl at pH 7.4. P value was determined using a two-tailed Mann Whitney U-test (n.s. P = 0.6126; n = 8 for control neurons and n = 7 Letm1 KD; U = 23). (Q) Comparison of synaptic vesicle pH estimates using vGlut-pH. P value was determined using a two-tailed unpaired two-tailed t-test (n.s. P = 0.66; n = 8 for both control and Letm1 KD neurons; t14 = 0.4389).
Extended Data Fig. 4 Related to Fig. 2. Parameters and outputs of neuronal metabolic modeling.
(A) Schematic of the computational model showing how activity-dependent Ca2+ influx elevates cytosolic and mitochondrial Ca2+, increasing ATP consumption and mitochondrial metabolism. Mitochondrial Ca2+ export is controlled by NCLX and Letm1 in control neurons, and mainly by NCLX in Letm1 KD. Ca2+ activates TCA cycle reactions, including PDP1-mediated boosting of PDHc activity. (B) Simulation of the metabolic expense of spiking. KANT is the change in the rate of adenine nucleotide translocator (ANT) caused by a neuronal spike. ANT exchanges cytosolic ADP for mitochondrial ATP. Ks = 1/1000 seconds. Spike-associated (dark red) and spike-independent (light red) metabolic cost are depicted. (C) Change in the mitochondrial fraction of ATP and ADP due a single neuronal spike. (D) Decay in free mitochondrial Ca2+ following 1AP (top) or 20AP at 20 Hz (bottom), modeled to be slower in Letm1 KD. (E) Simulation of activity-driven ATP dynamics in control and Letm1 KD in arbitrary units (a.u.).
Extended Data Fig. 5 Related to Fig. 3. Mito4x-LA-GCaMP6s properties and additional experiments.
(A) Aversive olfactory conditioning was used as the behavioral training paradigm to elicit neural circuit activation of the drosophila MB. Flies subjected to unpaired training do not form associative memory. Mitochondrial Ca2+ imaging was performed 0.5-1.5 hours after training while pyruvate imaging and mitochondrial Ca2+ imaging were done 3 hours after training. Retrieval at 3 hours corresponds to middle-term memory, while retrieval at 24 hours tests long-term memory. In the unpaired protocol, the lack of temporal association between shock and odor prevents memory formation and imaging at corresponding time points serves as a non-associative control. (B-C) Ca2+ titrations in purified protein for GCaMP6s (grey), LA-GCaMP6s (GCaMP6s-D362I) and GCaMP6-150 at room temperature (B) or at 37 °C (C) with n = 2 for each measurement. (D) Axonal mitochondrial Ca2+ responses to stimulation (100 AP, 100 Hz) in control and Letm1 KD neurons using Mito4x-LA-GCaMP6s. The figure shows traces corresponding to the mean ± SEM of Mito4x-LA-GCaMP6f in axons. (E) Rate of mitochondrial Ca2+ decay measured as half-time decay (t1/2) in axonal mitochondria following stimulation in control and Letm1 KD neurons. P value was determined using a two-tailed Mann-Whitney U-test (****P < 0.0001; n = 18 for control and n = 15 for Letm1 KD; U = 30.50). (F) Mitochondrial Ca2+ levels shown as the ratio of Mito4x-LA-GCaMP6s to mCarmine in control and Letm1 KD flies, measured 3 hours after exposure to either 1x paired or unpaired protocol. P value for control flies was determined using a two-tailed unpaired t-test (n.s., P = 0.7206; n = 13,13; t24 = 0.362). P value for Letm1 KD flies was determined using a two-tailed unpaired t-test (n.s., P = 0.3031); n = 14,14; t26 = 1.051).
Extended Data Fig. 6 Related to Fig. 3. Control experiments for the effect of Letm1 KD on fly olfactory memory.
(A-B) Letm1 RNAi #2 flies were used to test memory. (A) MTM tested 3 h after 1x conditioning in control and Letm1 RNAi #2 flies. Group differences were assessed with a one-way ANOVA (n = 12 per group). ANOVA: F(2,24) = 0.5172, P = 0.6027 (B) LTM tested 24 h after 1x conditioning in control and Letm1 RNAi #2 flies, is increased LTM in Letm1 KD flies. (one-way ANOVA F(2,33) = 6.565, P = 0.0040, n = 12,12,12, Tukey’s post-test Control 1 vs Letm1 KD, **P = 0.0045; Control 2 vs Letm1 KD, *P = 0.0271. Control 1 vs Control 2, n.s., P = 0.7579). (C) No RNAi expression was induced and LTM was tested 24 h after 1x conditioning in control and Letm1 RNAi #1 or #2 lines and no significant differences were detected. P values were determined using one-way ANOVA (left: Letm1 RNAi #1: F(2,51) = 0.2267, P = 0.7670, n = 18,18,18; right Letm1 RNAi #2: F(2,33) = 0.1892, P = 0.8285, n = 12,12,12) (D) Avoidance of control and Letm1 RNAi #1 or #2 lines to the odors used for olfactory conditioning shows no difference in odor avoidance. P values were determined using one-way ANOVA (top panel: Letm1 RNAi #1: Octanol: F(2,33) = 0.8653, P = 0.4303, n = 12,12,12; Methyl-cyclohexanol: F(2,33) = 2.749, P = 0.0787, n = 12,12,12; bottom panel: Letm1 RNAi #2: Octanol: F(2,33) = 2.859, P = 0.0716, n = 12,12,12; Methyl-cyclohexanol: F(2,33) = 0.2520, P = 0.7787, n = 12,12,12) (E) Avoidance of control and Letm1 RNAi #1 or #2 lines to the shock stimulus shows no difference in shock avoidance. P values were determined using one-way ANOVA (left: Letm1 RNAi #1: F(2,33) = 0.5629, P = 0.5749, n = 12,12,12; right Letm1 RNAi #2: F(2,33) = 0.7137, P = 0.4972, n = 12,12,12) (F) Testing for LTM formation 24 hours after a 5x spaced training in control and Letm1 RNAi #1 or #2 flies show normal LTM formation in all conditions. P values were determined using one-way ANOVA (left: Letm1 RNAi #1: F(2,51) = 0.2275, P = 0.7973, n = 18,18,18; right Letm1 RNAi #2: F(2,33) = 0.9020, P = 0.4155, n = 12,12,12) For graph, A-F: Data are represented as mean ± SEM and genotypes are shown in the top legend. (G) Letm1 RNA levels measured by qPCR from fly brains, normalized to α-Tub84B as reference gene. Note that knock down is underestimated as RNAi is only expressed in MB neurons and not in the entire brain. P values were determined using a one-way ANOVA followed by Tukey’s multiple comparison test versus control with n = 4 for control, n = 4 for Letm1 RNAi #1 and n = 5 for Letm1 RNAi #2, one-way ANOVA: F(2,10) = 8.838, P = 0.0.0062, Tukey’s multiple comparison P values: Letm1 RNAi #1 vs control, *P = 0.0084; Letm1 RNAi #2 vs control, *P = 0.0147. Data are represented as mean ± SEM.
Extended Data Fig. 7 Related to Fig. 4. Control experiments for the effect of Letm1 KD on rodent olfactory memory.
(A) Genetic design of viral constructs injected into the mouse hippocampus. (B-E) Analysis of motor behavior in mice before being subjected to memory tests. No differences were found between control and Letm1 KD mice in (B) distance travelled or (C) speed. For (B), P value was determined using a two-tailed unpaired t-test (n.s., P = 0.7321; n = 8 per group; t14 = 0.3492). For (C), P value was determined using a two-tailed unpaired t-test (n.s., P = 0.8231; n = 8 per group; t14 = 0.2278). (D, E) Representative trajectory plots of locomotor activity in control (left) and Letm1 KD (right) mice, respectively. Grey rectangles indicate the base of the arena while lines outside these markings represent rearing events. (F-J) CS+ corresponds to the odor previously paired with aversive LiCl injection and CS- corresponds to the innocuous odor. Control and Letm1 KD refer to mice that have received viral injections in the dorsal hippocampus of control or Letm1 miRNA-based shRNAs, respectively. (F) Water consumption from CS+ and CS- drinking bottles in control and Letm1 KD mice on day 5. These raw data are used for quantifying memory scores in Fig. 4f. P values were determined using two-way ANOVA (n = 8 per group), followed by Tukey’s multiple comparisons test. For control: CS- vs CS + , ****P < 0.0001. For Letm1 KD: CS- vs CS + , ****P < 0.0001. (G) Water consumption from CS+ and CS- drinking bottles in control and Letm1 KD mice on day 15. These raw data are used for quantifying memory scores in Fig. 4g. P values were determined using two-way ANOVA (n = 8 per group), followed by Tukey’s multiple comparisons test. For control: CS- vs CS + , n.s., P = 0.9704. For Letm1 KD: CS- vs CS + , **P < 0.0023. (H) Water consumption during habituation sessions in control and Letm1 KD mice. P values were determined using a two-tailed Mann–Whitney U-tests session #1 (n.s., P = 0.9818; n = 8 per group; U = 31.5) and session #2 (n.s.; P = n = 8 per group; U = 23.5). No significant differences were found. (Iand J) Water consumption in control and Letm1 KD mice when either banana or almond were devalued with LiCl (CS + ). These data are obtained from the main pool of raw data shown in (F) by analyzing separately paired responses to each odor. P values were determined using two-way ANOVA, followed by Tukey’s multiple comparisons test. For panel (I): Control, Banana vs Almond, *P = 0.0161; Letm1 KD, Banana vs Almond, **P = 0.0083. For panel (J): Control, Almond vs Banana, **P = 0.0038; Letm1 KD, Almond vs Banana, **P = 0.0081.
Supplementary information
Supplementary Table 1 (download XLSX )
The table shows details of organisms/strains, antibodies, recombinant DNA, chemicals, critical commercial assays/kits and software used in this work. This table contains the addgene references for plasmids generated in this work.
Supplementary Table 2 (download XLSX )
An EGTA/Ca2+ buffer system was used to determine indicator responses from 0 to 3 µM and used for the fit of GCaMP6s. An NTA/ Ca2+ buffer system was used from 0 to 140 µM and used for the fit of LA-GCaMP6s (GCaMP6s-D362I) and GCaMP6-150. Mean ± SEM are shown, n = 3 replicates.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3s and 4 (download PDF )
Source data for Extended Data Figs. 3b,f and 4c–e.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Amrapali Vishwanath, A., Comyn, T., Mira, R.G. et al. Mitochondrial Ca2+ efflux controls neuronal metabolism and long-term memory across species. Nat Metab 8, 467–488 (2026). https://doi.org/10.1038/s42255-026-01451-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-026-01451-w