Abstract
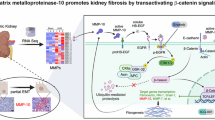
The fibrotic kidney microenvironment is shaped by cellular crosstalk, extracellular matrix (ECM) remodelling, metabolic reprogramming and spatial heterogeneity. While late-stage ECM changes dominate fibrosis, the role of early-activated matrix proteins remains unclear. Here we show that ECM1 is an early regulator of kidney remodelling. Global Ecm1 knockout mice develop spontaneous fibrosis and early death, whereas ECM1 levels markedly increase in biofluids during chronic kidney disease. Targeting Ecm1 through AAV9-mediated knockdown or fibroblast-specific deletion substantially reduces renal fibrosis. Mechanistically, Ecm1 deletion disrupts the integrin α2β1–RhoC axis, suppressing YAP activity. Reduced YAP nuclear translocation and diminished YAP–TEAD4 complex formation relieve TEAD4-mediated repression of Pgc1a, enhancing mitochondrial oxidative phosphorylation (OXPHOS) and promoting repair. Spatial transcriptomics and proteomics confirm this mechano-metabolic pathway, revealing mitochondrial reprogramming in tubules that counteracts fibrotic progression. Notably, YAP inactivation in fibroblasts limits aberrant activation without impairing their OXPHOS. This selective ECM–mitochondrial crosstalk uncovers a mechano-metabolic pathway in which mitochondrial shifts drive defence against kidney fibrosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All unique materials generated in this study are available from the corresponding author on completion of a materials transfer agreement. Raw mass spectrometry data have been deposited in MassIVE under the identifier MSV000100327 (global proteomics) and in the ProteomeXchange Consortium via the PRIDE partner repository under the dataset identifier PXD072386 (phosphoproteomics). Bulk RNA-sequencing data are available in the Gene Expression Omnibus (GEO) under accession number GSE226534. Spatial transcriptomics data are available in the GEO database under accession number GSE281998. Source data are provided with this paper.
References
Kovesdy, C. P. Epidemiology of chronic kidney disease: an update 2022. Kidney Int. Suppl. (2011) 12, 7–11 (2022).
Bello, A. K. et al. An update on the global disparities in kidney disease burden and care across world countries and regions. Lancet Glob. Health 12, e382–e395 (2024).
Time to sound the alarm about the hidden epidemic of kidney disease. Nature 628, 7–8 (2024).
Li, L., Fu, H. & Liu, Y. The fibrogenic niche in kidney fibrosis: components and mechanisms. Nat. Rev. Nephrol. 18, 545–557 (2022).
Zhang, H. et al. The extracellular matrix integrates mitochondrial homeostasis. Cell 187, 4289–4304 (2024).
Bülow, R. D. & Boor, P. Extracellular matrix in kidney fibrosis: more than just a scaffold. J. Histochem. Cytochem. 67, 643–661 (2019).
Wang, D., Sant, S., Lawless, C. & Ferrell, N. A kidney proximal tubule model to evaluate effects of basement membrane stiffening on renal tubular epithelial cells. Integr. Biol. (Camb.) 14, 171–183 (2022).
Abrahamson, D. R. & Leardkamolkarn, V. Development of kidney tubular basement membranes. Kidney Int. 39, 382–393 (1991).
Vink, H. & Duling, B. R. Identification of distinct luminal domains for macromolecules, erythrocytes, and leukocytes within mammalian capillaries. Circ. Res. 79, 581–589 (1996).
Miguel, V., Shaw, I. W. & Kramann, R. Metabolism at the crossroads of inflammation and fibrosis in chronic kidney disease. Nat. Rev. Nephrol. https://doi.org/10.1038/s41581-024-00889-z (2024).
Xu, M., Zhou, J., Yan, J. & Wang, J. Identification of a novel mutation of extracellular matrix protein 1 gene in a Chinese family with lipoid proteinosis. Clin. Cosmet. Investig. Dermatol. 16, 1515–1519 (2023).
Yin, H. et al. Extracellular matrix protein-1 secretory isoform promotes ovarian cancer through increasing alternative mRNA splicing and stemness. Nat. Commun. 12, 4230 (2021).
Xu, K. et al. Elevated extracellular matrix protein 1 in circulating extracellular vesicles supports breast cancer progression under obesity conditions. Nat. Commun. 15, 1685 (2024).
Meng, X.-Y., Liu, J., Lv, F., Liu, M.-Q. & Wan, J.-M. Study on the correlation between extracellular matrix protein-1 and the growth, metastasis and angiogenesis of laryngeal carcinoma. Asian Pac. J. Cancer Prev. 16, 2313–2316 (2015).
Hardy, S. A. et al. Novel role of extracellular matrix protein 1 (ECM1) in cardiac aging and myocardial infarction. PLoS ONE 14, e0212230 (2019).
Hardy, S. A. et al. Extracellular matrix protein-1 as a mediator of inflammation-induced fibrosis after myocardial infarction. JACC Basic Transl. Sci. 8, 1539–1554 (2023).
Feng, D. et al. Downregulation of extracellular matrix protein 1 effectively ameliorates osteoarthritis progression in vivo. Int. Immunopharmacol. 126, 111291 (2024).
Zhang, C. et al. ECM1 promotes migration and invasion in endometriosis. Reprod. Biol. 24, 100826 (2024).
Fan, W. et al. ECM1 prevents activation of transforming growth factor β, hepatic stellate cells, and fibrogenesis in mice. Gastroenterology 157, 1352–1367 (2019).
Su, P. et al. Novel function of extracellular matrix protein 1 in suppressing Th17 cell development in experimental autoimmune encephalomyelitis. J. Immunol. 197, 1054–1064 (2016).
He, L. et al. Extracellular matrix protein 1 promotes follicular helper T cell differentiation and antibody production. Proc. Natl Acad. Sci. USA 115, 8621–8626 (2018).
Kong, L. et al. Extracellular matrix protein 1, a direct targeting molecule of parathyroid hormone-related peptide, negatively regulates chondrogenesis and endochondral ossification via associating with progranulin growth factor. FASEB J. 30, 2741–2754 (2016).
Kong, L. et al. Interaction between cartilage oligomeric matrix protein and extracellular matrix protein 1 mediates endochondral bone growth. Matrix Biol. 29, 276–286 (2010).
Kirita, Y., Wu, H., Uchimura, K., Wilson, P. C. & Humphreys, B. D. Cell profiling of mouse acute kidney injury reveals conserved cellular responses to injury. Proc. Natl Acad. Sci. USA 117, 15874–15883 (2020).
Tabula Sapiens Consortium. The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Gui, Y. et al. Fibroblast expression of transmembrane protein smoothened governs microenvironment characteristics after acute kidney injury. J. Clin. Invest. 134, e165836 (2024).
Tran, M. T. et al. PGC1α drives NAD biosynthesis linking oxidative metabolism to renal protection. Nature 531, 528–532 (2016).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Gui, Y. et al. Yap/Taz mediates mTORC2-stimulated fibroblast activation and kidney fibrosis. J. Biol. Chem. 293, 16364–16375 (2018).
Balzer, M. S. et al. Single-cell analysis highlights differences in druggable pathways underlying adaptive or fibrotic kidney regeneration. Nat. Commun. 13, 4018 (2022).
Chen, J. et al. YAP activation in renal proximal tubule cells drives diabetic renal interstitial fibrogenesis. Diabetes 69, 2446–2457 (2020).
Xu, C. et al. Tubule-specific Mst1/2 deficiency induces CKD via YAP and non-YAP mechanisms. J. Am. Soc. Nephrol. 31, 946–961 (2020).
Tang, T. T. et al. Small molecule inhibitors of TEAD auto-palmitoylation selectively inhibit proliferation and tumor growth of NF2-deficient mesothelioma. Mol. Cancer Ther. 20, 986–998 (2021).
Smits, P. et al. The human extracellular matrix gene 1 (ECM1): genomic structure, cDNA cloning, expression pattern, and chromosomal localization. Genomics 45, 487–495 (1997).
Sercu, S. et al. Interaction of extracellular matrix protein 1 with extracellular matrix components: ECM1 is a basement membrane protein of the skin. J. Invest. Dermatol. 128, 1397–1408 (2008).
Hynes, R. O. The extracellular matrix: not just pretty fibrils. Science 326, 1216–1219 (2009).
Eckenstaler, R., Hauke, M. & Benndorf, R. A. A current overview of RhoA, RhoB, and RhoC functions in vascular biology and pathology. Biochem. Pharmacol. 206, 115321 (2022).
Acharya, B. R. et al. A mechanosensitive RhoA pathway that protects epithelia against acute tensile stress. Dev. Cell 47, 439–452 (2018).
Ohgushi, M., Minaguchi, M. & Sasai, Y. Rho-signaling-directed YAP/TAZ activity underlies the long-term survival and expansion of human embryonic stem cells. Cell Stem Cell 17, 448–461 (2015).
Höffken, V., Hermann, A., Pavenstädt, H. & Kremerskothen, J. WWC proteins: important regulators of Hippo signaling in cancer. Cancers (Basel) 13, 306 (2021).
Elbediwy, A. et al. Integrin signalling regulates YAP and TAZ to control skin homeostasis. Development 143, 1674–1687 (2016).
Koo, J. H. & Guan, K.-L. Interplay between YAP/TAZ and metabolism. Cell Metab. 28, 196–206 (2018).
Lee, K.-M. et al. ECM1 promotes the Warburg effect through EGF-mediated activation of PKM2. Cell. Signal. 27, 228–235 (2015).
Kashihara, T. & Sadoshima, J. Regulation of myocardial glucose metabolism by YAP/TAZ signaling. J. Cardiol. 83, 323–329 (2024).
White, S. M. et al. YAP/TAZ inhibition induces metabolic and signaling rewiring resulting in targetable vulnerabilities in NF2-deficient tumor cells. Dev. Cell 49, 425–443 (2019).
Dai, J.-Z. et al. YAP dictates mitochondrial redox homeostasis to facilitate obesity-associated breast cancer progression. Adv. Sci. (Weinh.) 9, e2103687 (2022).
Kami Reddy, K. R. et al. Mitochondrial reprogramming by activating OXPHOS via glutamine metabolism in African American patients with bladder cancer. JCI Insight 9, e172336 (2024).
Liu, Y. et al. Activation of the YAP/KLF5 transcriptional cascade in renal tubular cells aggravates kidney injury. Mol. Ther. 32, 1526–1539 (2024).
Wu, J. et al. Serum protein biomarkers of fibrosis aid in risk stratification of future stricturing complications in pediatric Crohn’s disease. Am. J. Gastroenterol. 114, 777–785 (2019).
Oyama, N. et al. Autoantibodies to extracellular matrix protein 1 in lichen sclerosus. Lancet 362, 118–123 (2003).
Martin, K. R. et al. Development of an efficient, effective, and economical technology for proteome analysis. Cell Rep. Methods 4, 100796 (2024).
Wu, C. et al. Global and site-specific effect of phosphorylation on protein turnover. Dev. Cell 56, 111–124 (2021).
Mehnert, M., Li, W., Wu, C., Salovska, B. & Liu, Y. Combining rapid data independent acquisition and CRISPR gene deletion for studying potential protein functions: a case of HMGN1. Proteomics 19, e1800438 (2019).
Li, W. et al. Assessing the relationship between mass window width and retention time scheduling on protein coverage for data-independent acquisition. J. Am. Soc. Mass Spectrom. 30, 1396–1405 (2019).
Li, W. et al. Turnover atlas of proteome and phosphoproteome across mouse tissues and brain regions. Cell 188, 2267–2287 (2025).
Tsou, C.-C. et al. DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat. Methods 12, 258–264 (2015).
Bekker-Jensen, D. B. et al. Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries. Nat. Commun. 11, 787 (2020).
Olsen, J. V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Zhou, D. et al. Fibroblast-specific β-catenin signaling dictates the outcome of AKI. J. Am. Soc. Nephrol. 29, 1257–1271 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Acknowledgements
We extend our gratitude to Vivace Therapeutics, Inc. (San Mateo, CA), for providing small-molecule inhibitors targeting the YAP signalling pathway. We gratefully acknowledge the contributions of the Single Cell Biology service and the cyberinfrastructure high-performance computing resources at The Jackson Laboratory for their expert assistance with this study. We thank S. Liu and L. Zhang at the First Affiliated Hospital of Nanjing University of Chinese Medicine for their invaluable help with the collection of human serum and urine samples. We are also grateful to A. Vella from the Department of Immunology at UConn Health for generously providing access to the Seahorse assay platform. Finally, we thank B. L. Cook, a science editor and illustrator at UConn Health, for his artwork and his careful reading and editing of the paper before submission. Some illustrations were created using BioRender.com. Funding: National Institutes of Health (NIH) grants DK116816 (D.Z.), DK128529 (D.Z.) and DK132059 (D.Z.); the University of Pittsburgh Center for Research Computing (S10OD028483); and the JAX Cancer Center (P30 CA034196). S.L. is supported by NIH grant GM159862. Y.G. is supported by the American Heart Association Career Development Award 25CDA1435759 and the University of Connecticut InCHIP Faculty Seed Grant.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.G., D.Z. Methodology: Y.G., W.L., Y.Y., H.F., T.T.T., S.L., Y.L., D.Z. Investigation: Y.G., Y.W., W.L., J.-J.L., C.J., R.B., S.M.M., H.W.S. Visualization: Y.G., D.Z. Funding acquisition: D.Z. Project administration: D.Z. Supervision: D.Z. Writing—original draft: Y.G., D.Z. Writing—review and editing: S.L., Y.L., D.Z.
Corresponding author
Ethics declarations
Competing interests
T.T.T. is the head of biology at Vivace Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Benjamin Humphreys, Katalin Susztak and Vishal Vaidya for their contribution to the peer review of this work. Primary Handling Editor: Revati Dewal, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ECM1 expression in human and animal models of CKD.
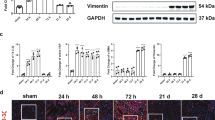
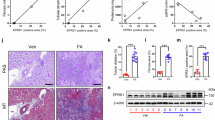
(a) Bulk RNA sequencing shows Ecm1 gene expression at days 0 (n = 4), 1 (n = 5), 3 (n = 5), and 10 (n = 3) after renal IRI. In the box plot, the minima, first quartile, median, third quartile, and maxima are represented by the lower whisker, lower hinge of the box, middle line, upper hinge of the box, and upper whisker, respectively. (b) Western blot analysis demonstrates ECM1 expression in the kidneys after IRI at 1 day and 3 months. Numbers indicate individual mouse in each group. (c) Quantitative data of ECM1 protein levels in fibrotic kidneys induced by IRI (n = 6), UUO (n = 5), and diabetes (16-week-old db/db mice, n = 5). (d) Sex-disaggregated data of urinary ECM1 levels in CKD patients. (e) Correlation between urinary ECM1 and eGFR levels in CKD patients (n = 66). (f, g) Sex-disaggregated data of correlations between urinary ECM1 and Scr or eGFR in CKD patients. (h) Serum ECM1 levels in non-diabetic CKD patients, with sex-disaggregated data shown on the right. (i) Serum (left panel) and urine (right panel) ECM1 levels in 16- and 28-week-old db/db mice (n = 11). (j) Genotyping of mice via PCR analysis of genomic DNA. Lane 1 represents Ecm1 knockout (KO) mice (Ecm1-/-), lane 2 indicates heterozygous controls (F/W), and lane 3 shows wild type (WT) mice (Ecm1+/+). (k) Quantitative data for Vimentin and α-SMA protein expression in Ecm1+/+ and Ecm1-/- mice kidneys at different ages (n = 3). IRI, ischemia-reperfusion injury; UUO, unilateral ureter obstruction; Scr, serum creatinine; eGFR, estimated glomerular filtration rate. Data are presented as means ± s.e.m. Differences among groups were analyzed using two-sided unpaired t-tests or one-way ANOVA followed by the Student-Newman-Keuls test. Correlation analyses were performed using Spearman’s test. r denotes the correlation coefficient.
Extended Data Fig. 2 Knockdown of Ecm1 alleviates kidney fibrosis induced by IRI and UUO.
(a) Double immunofluorescence staining of ECM1 with markers for mesengial cells (PDGFRβ), endothelial cells (CD31), and podocytes (PODXL) in diseased glomeruli 3 weeks after doxorubicin injection. (b) Western blot analyses confirming the efficiency of ECM1 knockdown in rat (left) or human (right) cell lines using three individual predesigned DsiRNA (1, 2, and 3). NC, negative control siRNA. (c) Immunofluorescence staining of markers for fibroblasts/pericytes (PDGFRβ), endothelial cells (CD31), proximal tubules (AQP1), distal tubules (AQP3), glomerular mesangial cells (PDGFRβ), and macrophages (F4/80) in ischemic kidneys two weeks after IRI, performed four weeks after GFP-tagged AAV-9 ShEcm1 vector injection. (d, e) Western blot analysis (d) and quantitative data (e) of ECM1 protein levels in AAV9-Scramble (Scram) and AAV9-ShEcm1 kidneys after UUO. Numbers indicate individual animals in each group (n = 6). (f) Quantitative data of COL1A1, FN, and α-SMA protein expression in AAV9-Scramble and AAV9-ShEcm1 kidneys after unilateral IRI combined with nephrectomy (n = 6). (g) Quantitative data of FN and α-SMA protein expression in AAV9-Scramble and AAV9-ShEcm1 kidneys after UUO (n = 5). (h) Quantitative data from immunohistochemical staining (α-SMA, COL1A1, FN, CD45) and Masson’s trichrome staining in AAV9-Scramble and AAV9-ShEcm1 kidneys after unilateral IRI combined with nephrectomy and UUO (Sham, n = 3; IRI, n = 5; UUO, n = 5). Each dot represents the score of the corresponding image. PDGFRβ, Platelet-derived growth factor receptor b; PODXL, Podocalyxin; AQP1, Aquaporin 1; AQP3, Aquaporin 3; α-SMA, α-Smooth Muscle Actin; COL1A1, Collagen 1α1; FN, fibronectin; Dsi, Dicer-substrate siRNA. Scale bar, 25 µm. Arrows indicate double-positive staining. Arrowheads indicate non-overlapping signals. Data are presented as mean ± s.e.m. Differences between groups were analyzed using two-sided unpaired t-tests or one-way ANOVA followed by the Student-Newman-Keuls test.
Extended Data Fig. 3 Global proteomics reveals knockdown of Ecm1 activates mitochondrial OXPHOS in fibrotic kidneys.
(a) Correlation of kidney proteome profiles between AAV9-Scram and AAV9-ShEcm1 mice after unilateral IRI combined with nephrectomy. (b-d) Gene Ontology (GO) enrichment analysis under biological process (b), cellular compartment (c), and molecular function (d) terms in each cluster of proteins are plotted with their names and significance. (e) Heatmap of differentially expressed proteins involved in fatty acid, Carbohydrate, and Amino acid metabolism in AAV9-Scram and AAV9-ShEcm1 mice fibrotic kidneys. (f) Quantitative data for protein expression of ATP5A, UQCRC2, MTCO1, SDHB, and NDUFB8 in AAV9-Scram and AAV9-ShEcm1 fibrotic kidneys (n = 6). (g) Quantitative data for protein expression of HK3, PGC1α, and PPARα in AAV9-Scram and AAV9-ShEcm1 fibrotic kidneys (n = 6). Data are presented as means ± s.e.m. Differences among groups were analyzed using two-sided unpaired t-tests.
Extended Data Fig. 4 Loss of Ecm1 in fibroblasts alleviates kidney fibrosis after IRI and UUO.
(a) PCR-based genotyping of mice genomic DNA. Lane 1: Col1a2+ fibroblast-specific Ecm1 conditional knockout (cKO) mice (genotype: Ecm1fl/fl, Cre), designated as Col1α2-Ecm1-/-. Lane 2: wild type (WT) mice (genotype: Ecm1fl/fl), designated as Col1α2-Ecm1+/+. (b) PCR genotyping of Pdgfrb+ fibroblast-specific Ecm1 knockout mice. Lane 1: Pdgfrb-Ecm1+/+ (WT) mice (genotype: Ecm1fl/fl). Lane 2: Pdgfrb-Ecm1-/- (cKO) mice (genotype: Ecm1fl/fl, Cre). (c) Scr levels in normal Col1α2-Ecm1+/+ (n = 5) versus Col1α2-Ecm1-/- mice (n = 5) (upper) and Pdgfrb-Ecm1+/+ (n = 5) versus Pdgfrb-Ecm1-/- mice (lower) (n = 4). Age- (8–10 weeks) and sex-matched littermates were used. (d) Kidney weight (KW) to body weight (BW) ratio in Col1α2-Ecm1+/+ (n = 5) versus Col1α2-Ecm1-/- (n = 5) mice (upper) and Pdgfrb-Ecm1+/+ (n = 5) versus Pdgfrb-Ecm1-/- (n = 4) mice (lower). (e) PAS staining of both strains of WT and cKO kidneys. (f) qPCR of Ecm1 mRNA in whole kidneys from both strains at 2 weeks after IRI. (g) Immunohistochemistry of kidney ECM1 expression in both strains after unilateral IRI at 2 weeks. (h) Quantitative data of fibronectin (FN) and α-SMA protein levels in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys following unilateral IRI (upper, n = 6) or UUO (lower, n = 5). (i) Quantitative data of MTS and immunohistochemistry (COL1A1, FN, CD45) in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys after unilateral IRI and UUO (Sham, n = 3; IRI, n = 5; UUO, n = 5). (j) Quantitative data of ATP5A, UQCRC2, MTCO1, SDHB, and NDUFB8 protein levels in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys after unilateral IRI (n = 6). (k) Western blot analysis of mitochondrial OXPHOS complexes I-V in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys after UUO. Numbers represent individual animals in each group. (l) Quantitative data of SDHB and NDUFB8 protein levels in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys after UUO (n = 5). (m) Quantitative data of FN and α-SMA protein levels in Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- kidneys after unilateral IRI (n = 5). (n, o) Representative micrographs of COL1A1 staining (n) and quantitative analysis (o) in Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- kidneys after unilateral IRI (Sham, n = 3; IRI, n = 5). (p, q) Quantitative data of MTS and CD45 staining in Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- kidneys after unilateral IRI (Sham, n = 3; IRI, n = 5). (r) Quantitative data of UQCRC2, MTCO1, and SDHB protein levels in Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- kidneys after IRI (n = 5). (s-v) Western blot analysis of FN in male (s) and female (t) Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys and male (u) and female (v) Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- kidneys after bilateral IRI at 28 days. Quantitative data is shown on the right of each panel (n = 6). Scale bar, 25 µm. Data are presented as mean ± s.e.m. Statistical differences were assessed using two-sided unpaired t-tests or one-way ANOVA followed by the Student-Newman-Keuls test.
Extended Data Fig. 5 Proteomics reveals altered signaling pathways after Ecm1 knockdown in fibrotic kidneys after IRI.
(a) Quantitative data for the protein expression of integrin (ITG) A2, B1, AV, RhoC, RhoA, and CDC42 in AAV9-Scramble (Sc) and AAV9-ShEcm1 kidneys after unilateral IRI combined with nephrectomy (n = 6). (b, c) Quantitative data for the protein expression of ITGA2, ITGB1, and RhoC in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- (b, n = 6) or Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- (c, n = 5) kidneys after unilateral IRI. (d) Structural representation of protein-protein interactions. The image shows the structure of ECM1 (purple), integrin β1 (blue), and integrin α2 (green). Key amino acid residues involved in hydrogen bonding and electrostatic interactions are labeled and depicted as sticks. Dashed lines indicate potential hydrogen bonds or salt bridges stabilizing the interaction interface. Data are presented as mean ± s.e.m. Differences between groups were analyzed using two-sided unpaired t tests.
Extended Data Fig. 6 Phosphoproteomics reveals energy metabolism changes are major events in fibrotic kidneys after knockdown of Ecm1.
(a) Poylogenetic tree of all protein kinases families detected in proteomics study. (b) Selected pathways of GO analysis for enriched up- and down-regulated phosphoproteins in the fibrotic kidneys after knockdown of Ecm1. (c) Quantitative data for the protein expression of LATS1, YAP, and TAZ in AAV9-Scramble (Sc) and AAV9-ShEcm1 kidneys after unilateral IRI combined with nephrectomy (n = 6). (d) Bioinformatics analysis showing TEAD4 is predicted to repress Pgc1a. (e) Human TEAD 4 motif. (f) Yap Dsi (rat) was transfected into NRK-52E cells. (g, h) Quantitative data for the protein expression of YAP and TAZ in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- (g, n = 6) or Pdgfrb-Ecm1+/+ and Pdgfrb-Ecm1-/- (h, n = 5) kidneys after unilateral IRI. Data are presented as mean ± s.e.m. Differences between groups were analyzed using two-sided unpaired t tests.
Extended Data Fig. 7 Spatial transcriptomic profiling of Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- fibrotic kidneys.
(a, b) Spatial feature plots showing total transcript counts (nCount_Spatial) and detected gene features (nFeature_Spatial) in kidneys from Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- mice (three biological pairs). (c) Clustering of the spots to reveal the spatial regions of kidney cortex and medulla. (d) Spatially resolved Ecm1 expression in Col1α2-Ecm1+/+ and Col1α2-Ecm1-/- kidneys. (e) Spatially resolved expression levels for fibrosis- and OXPHOS-related genes (Col1α1, Col1α2, Col3α1, Sdhb, Uqcrc1, Uqcrc2). (f) Violin plots showing normalized expression of Sdhb, Uqcrc1, Uqcrc2, and Slc34a1 between genotypes. (g) Spatially resolved expression of proximal tubular marker Slc34a1. (h) Deconvolution maps displaying spatial distribution of renal cell populations including distal convoluted tubule (DCT), endothelial cell (Endo), fibroblast (Fib), glomerular endothelial cell (GEC), intercalated cell (IC), immune cells (Immune), loop of Henle (LOH), podocyte (Podo), and proximal tubule (PT). (i) Dot plot summarizing marker gene expression across major renal cell types. (j) Spatial definition of fibrotic niches and their k nearest neighbors. For the violin plots, statistical significance was assessed using a two-sided Wilcoxon rank-sum test.
Extended Data Fig. 8 VT103 has no adverse effects on multiple organs.
(a) Experiment design. VT103 was administered orally to mice daily for 14 days. (b, c) Scr (b) and alanine aminotransferase (ALT) (c) levels in normal VT103-treated mice (n = 6). Age- (8–10 weeks) and sex-matched mice littermates were used. Each dot represents one mouse. (d) Representative PAS-stained sections of kidney, liver, intestine, lung, and heart reveal no morphological changes after VT103 treatment in mice. Scale bar, 25 µm. Graphs are presented as mean ± s.e.m. Differences between groups were analyzed using one-way ANOVA followed by the Student-Newman-Keuls test. Panel a created using BioRender.com.
Extended Data Fig. 9 Pharmacological inhibition of Yap signaling ameliorates kidney fibrosis and promotes mitochondrial OXPHOS.
(a) qPCR analysis showing Tead1, Tead2, Tead3, and Tead4 mRNA levels in vehicle (Veh.) and VT103-treated kidneys after IRI (n = 5). (b) Quantitative data for the protein expression of COL1A1, FN, and α-SMA in vehicle and VT103-treated kidneys after IRI (n = 5). (c) Quantitative data for MTS and immunohistochemical staining (α-SMA, COL1A1, and CD45) in vehicle and VT103-treated mice kidneys after IRI (Sham, n = 3; IRI, n = 5). (d) Quantitative data for the protein expression of SDHB and NDUFB8 in vehicle and VT103-treated mice kidneys after IRI (n = 5). Data are presented as mean ± s.e.m. Differences between groups were analyzed using two-sided unpaired t tests or one-way ANOVA followed by the Student-Newman-Keuls test.
Extended Data Fig. 10 Knockdown of Ecm1 enhances tubular mitochondrial function in vitro.
(a) Quantitative data of FN, a-SMA, and ITGA2 in NRK-49F cells after transfected with Ecm1 Dsi under TGFβ stress (n = 3). (b) Western blot assay showing reduced HK3 levels in NRK-52E cells seeded on Ecm1-deficient matrix scaffold isolated from Ecm1-knockdown fibroblasts. (c, d) qPCR analyses of Hk2, Hk3, or Plin2 in tubular cells seeded on Ecm1-deficient matrix scaffold (c) or treat with recombinant ECM1 protein (d) (n=3). (e) Western blot analyses confirming the efficiency of RhoC knockdown in NRK-52E using three individual predesigned DsiRNA (1, 2, and 3). CM. conditioned medium; Plin2, perilipin 2. Data are presented as mean ± s.e.m. Differences between groups were analyzed using two-sided unpaired t tests or one-way ANOVA followed by the Student-Newman-Keuls test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1. Primer sequence information. Supplementary Table 2. Demographic characteristics of human participants. Supplementary Table 3. Antibody information.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download PDF )
Unprocessed immunoblots.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download PDF )
Unprocessed immunoblots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed immunoblots.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed immunoblots.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed immunoblots.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed immunoblots.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed immunoblots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gui, Y., Li, W., Liu, JJ. et al. Early-activated extracellular matrix proteins shape the metabolic and spatial dynamics of the kidney fibrotic microenvironment. Nat Metab 8, 604–623 (2026). https://doi.org/10.1038/s42255-026-01458-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-026-01458-3