Abstract
Acinar-to-ductal metaplasia (ADM) is a reversible cell state that facilitates pancreas repair following injury. Oncogenic KRAS mutations can progress ADM to pancreatic intraepithelial neoplasia (PanIN) and pancreatic ductal adenocarcinoma (PDAC). However, the metabolic alterations in these precancerous lesions are understudied. Here, we identify global changes in central carbon metabolism genes and metabolites during ADM formation. In particular, NRF2-target genes are significantly induced in ADM. Among these, we focus on genes encoding NADPH-producing enzymes glucose-6-phosphate dehydrogenase (G6PD) and malic enzyme 1 (ME1), which participate in the regulation of oxidative stress. In mouse models of pancreatic tumourigenesis, G6PD deficiency or Me1 loss increases reactive oxygen species and lipid peroxidation, which is accompanied by accelerated formation of ADM and PanIN lesions. Notably, Me1 loss, but not G6PD deficiency, promotes faster PDAC progression. We demonstrate that oxidative stress is required for ADM, as pharmacological antioxidant treatment attenuates ADM progression in vivo and ex vivo. Conversely, depleting the antioxidant glutathione promotes precancerous lesions in primary human acinar cells and in mice. Together, our findings shed light on metabolic reprogramming in the precancerous pancreas.
Similar content being viewed by others
Main
Oncogenic KRAS mutations are initiating events in PDAC1. PDAC has a low 5-year survival rate owing to few early detection methods, a paucity of effective therapies and late-stage diagnosis2. PDAC tumourigenesis progresses stepwise, beginning with ADM, followed by PanIN lesions. ADM is a reversible, healing response after pancreatic injury or inflammation, in which acinar cells transdifferentiate into ductal progenitor-like cells, proliferate and restore damaged tissue3,4,5. Upon healing, ADM cells redifferentiate into acinar cells; however, oncogenic KRAS blocks this process, leading to persistent ADM and PanIN formation. PanIN lesions are common in otherwise healthy tissue6,7, underscoring the need to define mechanisms that control precancer formation and progression8.
Pancreatic cancer cells require biosynthetic precursors and energy to sustain growth9,10,11. Although metabolic reprogramming in PDAC is well characterized, the role of metabolism in driving precancerous lesions remains underexplored. Requirements for acetyl-CoA, cholesterol metabolism and tight regulation of reactive oxygen species (ROS) have been demonstrated in precancer formation and progression12,13,14. MYC and NFE2L2 (NRF2) are key transcription factors facilitating metabolic changes during cancer initiation15,16,17,18.
NADPH supports biosynthesis (fatty acids, cholesterol, deoxyribonucleotides) and antioxidant systems that control ROS19. The primary source of NADPH is G6PD, the first and rate-limiting enzyme in the oxidative pentose phosphate pathway (PPP). G6PD deficiency is an X-linked disorder and the most common human enzyme deficiency20, often causing haemolytic anaemia21. We previously showed that oncogenic KRAS activates the nonoxidative branch of the PPP by enhanced glycolysis and transcriptional upregulation of Rpia and Rpe22,23,24. As such, pancreatic cancer cells rely on NADPH produced by ME1 (refs. 23,25). Loss of TIGAR, which supports oxidative PPP activity, delays PanIN formation but enhances metastasis14, highlighting complex regulation of PPP flux, NADPH production and ROS control during cancer initiation and progression.
Here, we use KrasG12D-driven mouse models to define metabolic pathways supporting precancer development. G6PD deficiency accelerates ADM and PanIN formation, while pancreas-specific Me1 loss accelerates early lesions and promotes PDAC progression. Loss of these NADPH-generating enzymes increases ROS and lipid peroxidation; antioxidant supplementation rescues accelerated lesion formation in mice and primary cultures. Inhibition of glutathione biosynthesis similarly promotes ADM in healthy human acinar cells and mice. Integrated RNA sequencing (RNA-seq) and metabolomics analyses further define metabolic changes during cancer initiation and establish the energetic demands of pancreatic precancerous lesions.
Metabolic changes accompany ADM
To investigate KrasG12D-driven metabolic changes during pancreatic cancer initiation, we performed RNA-seq and targeted liquid chromatography–tandem mass spectrometry (LC–MS/MS)-based metabolomics on primary, ex vivo cultured acinar cells undergoing ADM. We isolated acinar cells from LSL-KrasG12D; Ptf1aCreERTM mice and treated them with vehicle or 4-hydroxytamoxifen (4-OHT) to induce mutant KrasG12D (Fig. 1a). Cells were collected 1 day after vehicle treatment and 1, 2 and 3 days after 4-OHT treatment (days 1, 2 and 3). Multidimensional scaling of RNA-seq data demonstrated temporal transcriptional changes (Extended Data Fig. 1a). No differential gene expression was observed at day 1 of 4-OHT treatment compared to vehicle (Extended Data Fig. 1b), but a significant increase in upregulated and downregulated genes were observed on days 2 and 3 (Fig. 1b). Across the time course, a decrease in acinar genes with a concurrent increase in ductal genes was detected (Extended Data Fig. 1c). Principal component analysis of intracellular metabolites paralleled the transcriptomics changes (Extended Data Fig. 2a). Notably, the abundance of eight metabolites was significantly different at day 1 (Extended Data Fig. 2b), with a larger set of metabolites altered at days 2 and 3 (Fig. 1c). Metabolomics on culture media during the time course revealed complementary findings, with pronounced metabolic changes at days 2 and 3 (Extended Data Fig. 2c–f).
a, Scheme of the experimental model. Pancreata were dissected from LSL-KrasG12D; Ptf1aCreERTM mice (n = 3) and acinar clusters were isolated for primary, ex vivo culture. Acinar clusters were grown free-floating in media and were treated with vehicle or 2 μM 4-OHT to induce mutant Kras. Cells were collected after 1, 2 or 3 days in culture. For each timepoint, cells from one well were divided for RNA extraction and metabolite analysis. b, Volcano plots of differentially expressed genes at day 2 and day 3 following 4-OHT treatment compared to day 1 vehicle. Significantly upregulated genes are red; significantly downregulated genes are blue. False discovery rate (FDR) < 0.05 was considered significant, log2(fold change (FC)) > 0.5 was considered upregulated andlog2(FC) < –0.5 was considered downregulated. Differential gene expression was estimated using the quasi-likelihood negative binomial generalized log-linear approach in edgeR. c, Volcano plots of differentially abundant metabolites when comparing day 2 and day 3 following 4-OHT treatment compared to day 1 vehicle. Significantly increased intracellular metabolites are red; decreased metabolites are blue. P < 0.1 was considered significant, log2(FC) > 0.5 was considered increased and log2(FC) < –0.5 was considered decreased. P values were calculated using a Student’s t-test (unpaired, two-tailed). d, Significant metabolism-related pathways at day 2, identified by gene set enrichment analysis (GSEA) from KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways. Blue bars highlight pathways of interest. e, Heatmap of detected metabolites in the TCA cycle. Scale represents log2(FC) of median normalized abundance relative to vehicle. f, Violin plot of the leading-edge genes in the TCA cycle KEGG pathway. The y axis represents log(FC) relative to vehicle. Lines inside the plots represent the quartiles and median. g, Violin plot of the leading-edge genes in the oxidative phosphorylation (OxPhos) KEGG pathway. The y axis represents log(FC) relative to vehicle. Lines inside the plots represent the quartiles and median. h, Heatmap of metabolites related to glycolysis and the PPP. Scale represents log2(FC) of median normalized abundance relative to vehicle. i, Violin plot of the leading-edge genes in the glycolysis and gluconeogenesis KEGG pathway. The y axis represents log(FC) relative to vehicle. Lines inside the plots represent the quartiles and median. j, Violin plot the leading-edge genes in the PPP KEGG pathways. The y axis represents log(FC) relative to vehicle. Lines inside the plots represent the quartiles and median. k, Heatmap of metabolites related to glutathione metabolism. Scale represents log2(FC) of median normalized abundance relative to vehicle. GSSG, oxidized glutathione. l, Violin plot of leading-edge genes in the glutathione metabolism KEGG pathway. The y axis represents log(FC) relative to vehicle. Lines inside the plots represent the quartiles and median. m, Transcription factor enrichment analysis generated using Enrichr from ENCODE and ChEA consensus transcription factors. The top 500 upregulated genes from day 2 were analysed. The blue bar highlights a transcription factor of interest. Enrichr was used to identify differentially enriched pathways and calculate P values with Fisher’s exact test. n, Heatmap showing log(FC) of significantly differentially expressed NRF2-target genes. Genes under investigation are indicated with blue text.
To better understand transcriptional changes during ADM, we performed gene set enrichment analysis (Fig. 1d and Supplementary Figs. 1 and 2). Integration of RNA-seq and metabolomics data identified dynamic metabolic reprogramming during ADM. A decrease in tricarboxylic acid (TCA) cycle metabolites was observed at day 1 (Fig. 1e), consistent with a shift away from oxidative phosphorylation at the onset of ADM18. A sharp increase in alpha-ketoglutarate and succinate abundance was detected at day 2, remaining high at day 3 (Fig. 1e). Concurrently, at days 2 and 3, we saw a stepwise increase in expression of leading-edge genes included in the KEGG TCA cycle and oxidative phosphorylation gene sets (Fig. 1f,g and Extended Data Fig. 3a–d). Glycolysis and the PPP were broadly upregulated during ADM. However, ribose 5-phosphate, arabinose-5-phosphate and uridine diphosphate N-acetylglucosamine (UDP-GlcNAc) levels decreased in days 1–3 compared to vehicle (Fig. 1h). RNA-seq showed a stepwise increase in leading-edge genes from the PPP and glycolysis (Fig. 1i,j and Extended Data Fig. 3e–h). Extracellular metabolite profiling supported these findings, with a significant increase in lactate detected in the media (Extended Data Fig. 2f), suggesting increased glycolysis. Glutathione metabolism exhibited distinct temporal dynamics. We noted that reduced glutathione (GSH) peaked at day 1 with a reduction at days 2 and 3, whereas oxidized glutathione and other metabolites involved in glutathione synthesis increased by day 3 (Fig. 1k). There was a complementary, stepwise increase in leading-edge genes for glutathione metabolism during the time course (Fig. 1l and Extended Data Fig. 3i,j). Additionally, we observed a general upregulation of purine metabolism gene expression but an overall decrease in purine metabolites across timepoints (Extended Data Fig. 4a–d). There was variability in the levels of metabolites and expression of genes related to pyrimidine metabolism, and we observed a progressive increase in uracil, uridine and cytidine, with a decrease in uridine diphosphate, cytidine monophosphate and deoxycytidine monophosphate (Extended Data Fig. 4e,f). There was a cumulative increase in genes involved in fatty acid degradation (Extended Data Fig. 4g). Amino acid abundance exhibited a biphasic trend, generally low at days 1 and 3 but higher at day 2 (Extended Data Fig. 4h). Many genes involved in branched-chain amino acid metabolism were upregulated as ADM progressed (Extended Data Fig. 4i).
An NRF2 pathway signature is present in mutant Kras-driven ADM
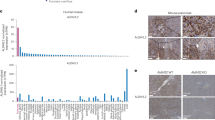
We performed transcription factor enrichment analysis using the Enrichr web tool26,27,28 (Fig. 1m and Extended Data Fig. 5). Consensus sites for NRF2 transcription factor binding were significantly enriched among the top 500 upregulated genes at day 2 (Fig. 1m and Extended Data Fig. 5a). NRF2 is known to have a key role in pancreatic tumourigenesis, progression and metastasis, in which it regulates expression of genes involved in ROS detoxification, cell metabolism, proliferation, invasion and migration12,14,15,29,30. We assayed our dataset for NRF2-target genes and found that several genes coding for NADPH-producing enzymes, such as G6pdx and Me1, increased during ADM formation (Fig. 1n and Supplementary Fig. 3a). We also observed protein expression from NRF2 targets G6PD, ME1 and NQO1 in mouse pancreas (Supplementary Fig. 3b–e).
To assess whether 4-OHT treatment was nonspecifically affecting acinar cells, we cultured wild-type cells for 3 days with transforming growth factor-α (TGFα) to induce ADM, and administered 4-OHT or vehicle (Extended Data Fig. 6a). We observed that 4-OHT treatment alongside TGFα still reduced expression of Amy (acinar gene coding for amylase) and increased expression of Spink1 (ductal gene coding for a trypsin inhibitor) (Extended Data Fig. 6b,c). We then checked whether an alternative ADM culture system yielded the same candidate genes. In this culture model, acinar cells from LSL-KrasG12D mice were isolated and treated with adenovirus expressing green fluorescent protein (GFP) for 1 day or adenovirus expressing Cre recombinase for 1, 2 and 3 days (Extended Data Fig. 6d), after which cells were collected for RNA-seq31. We observed a concurrent decrease in acinar gene expression and an increase in ductal gene expression over the 3 days (Extended Data Fig. 6e), supporting the formation of ADM. We then assayed for the NRF2-target genes from Fig. 1n and observed the same patterns reproduced in this additional dataset (Extended Data Fig. 6f). Specifically, we saw a stepwise increase in G6pdx and Me1 (Extended Data Fig. 6g,h), encoding NADPH-generating enzymes from the two major NADPH pathways in the cell. Collectively, data on ADM metabolism from these ex vivo profiling methods suggested an important role for NADPH regulatory enzymes G6PD and ME1.
G6PD deficiency decreases oxidative PPP flux in Kras G12D-expressing acinar cells
The PPP branches from glycolysis and consists of two arms: oxidative and nonoxidative (Fig. 2a). Flux through the oxidative arm, where G6PD resides, yields NADPH and ribose for nucleotides. Given the significant increase of G6pdx in ADM (Fig. 1n, Extended Data Fig. 6f,g and Supplementary Fig. 3a) and its contrast with activation of the nonoxidative PPP in PDAC, we sought to determine the role of oxidative PPP metabolism in ADM. A mouse model of human G6PD deficiency demonstrates enzyme activity of 15–20% in hemizygous mice (male G6pdmut/y), 50–60% in heterozygous mice (female G6pdmut/+) and 15–20% in homozygous mutant mice (female G6pdmut/mut)32,33. To determine how G6PD deficiency and decreased oxidative PPP flux alters pancreatic tumourigenesis, we generated LSL-KrasG12D; Ptf1aCre; G6pdmut mice (KC;G6pdmut) (breeding scheme in Extended Data Fig. 7a).
a, Schematic of the PPP. In the first step of glycolysis, glucose is converted into glucose 6-phosphate (glucose 6P). G6PD, the first and rate-limiting enzyme of the oxidative PPP (oxPPP; blue), converts glucose 6P into 6-phosphogluconolactone and NADPH. Another NADPH-producing enzyme in the oxPPP, 6-phosphogluconate dehydrogenase (6PGD), generates ribulose 5-phosphate (ribulose 5P). Ribulose 5P can be used in the nonoxidative PPP (nonoxPPP; green) by the enzymes ribulose-5-phosphate 3-epimerase (RPE) and ribose-5-phosphate isomerase (RPI) to generate intermediate metabolites that enter glycolysis or nucleic acid biosynthesis, including ribose 5-phosphate (ribose 5P). b, H&E staining and immunostaining for amylase (AMY; acinar cells), cytokeratin 19 (CK19; ductal, metaplastic, neoplastic cells) and Ki67 (proliferation) in pancreata from 16-week-old KC;G6pdwt and KC;G6pdmut mice. Scale bars, 50 μm. c, Percentage of AMY-positive area in pancreata as quantified from male (closed circles) and female (open circles) KC;G6pdwt and KC;G6pdmut mice at 16 weeks; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a Student’s t-test (unpaired, two-tailed). d,e, Percentage of CK19+ cells (d) and Ki67+ cells (e) in the pancreas as quantified from male (closed circles) and female (open circles) KC;G6pdwt and KC;G6pdmut mice at 16 weeks; n = 5 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a Student’s t-test (unpaired, two-tailed). f, H&E staining, Alcian Blue staining (for PanIN-produced mucins) and CK19 (ductal, metaplastic, neoplastic cells) immunostaining in 26-week-old KC;G6pdwt and KC;G6pdmut mice. Scale bars, 100 μm. g, Pathological grading of pancreas tissues from 26-week-old KC;G6pdwt and KC;G6pdmut mice, representing the percentage of total tissue area with acinar cells, ADM and PanIN lesions; n = 5 KC;G6pdwt mice; n = 7 KC;G6pdmut mice. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with Tukey’s post hoc test. h, H&E staining in 1-year-old KC;G6pdwt and KC;G6pdmut mice. Scale bars, 100 μm. i, Pathological grading of pancreas tissues from 1-year-old KC;G6pdwt and KC;G6pdmut mice, representing the percentage of total tissue area with acinar cells, ADM and PanIN lesions; n = 5 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with Tukey’s post hoc test.
To determine whether oxidative PPP activity was decreased in G6pdmut pancreas, we traced radioactive carbon incorporation into carbon dioxide (CO2) produced from oxidative decarboxylation in the PPP and TCA cycle. [1-14C]glucose labels CO2 derived from both oxidative PPP and the TCA cycle, while [6-14C]glucose specifically traces TCA-derived CO2. By following the tracers in tandem, we can determine oxidative PPP activity (Extended Data Fig. 7b). As expected, KC;G6pdmut acinar cells have decreased oxidative PPP flux, as measured by [1-14C]O2, compared to G6pdwt cells (Extended Data Fig. 7c). TCA cycle flux was not affected by G6pd status, as measured by [6-14C]O2 (Extended Data Fig. 7c). These data suggest that G6pdmut acinar cells have less oxidative PPP activity, establishing a physiologically relevant model to study how G6PD deficiency affects pancreatic tumourigenesis.
G6PD deficiency accelerates ADM and PanIN lesions in mouse models
ROS generated during pancreatic tumourigenesis is countered by flux into the PPP to provide NADPH13,14,22. We reasoned that decreasing oxidative PPP activity through G6PD deficiency would promote KrasG12D-driven ADM and tumourigenesis owing to increased oxidative stress. To test this hypothesis, we aged KC;G6pdmut and KC;G6pdwt mice to 8 weeks, when ADM is present within the pancreas of KC mice. We observed ADM in KC;G6pdwt mice as noted by haematoxylin and eosin (H&E)-stained tissue sections, decreased amylase (AMY), increased cytokeratin 19 (CK19) and increased proliferation (Ki67) within ADM lesions (Extended Data Fig. 7d). In KC;G6pdmut mice, there was a trend for less acinar cell area and significantly increased proliferation (Extended Data Fig. 7e–h). At 16 weeks, when ADM and early PanIN lesions are present in KC mice, KC;G6pdmut mice showed more ADM and PanIN lesions, indicated by a significant decrease in AMY and significantly more proliferation compared to KC;G6pdwt mice (Fig. 2b–e and Extended Data Fig. 7h). We did not find a difference in pancreas weight to body weight ratios in KC;G6pdmut mice, an indirect measure of tissue transformation (Extended Data Fig. 7i).
To determine the impact of G6PD deficiency on PanIN formation, we aged mice to 26 weeks, when low-grade PanINs are present in KC models (Fig. 2f). There was no difference in pancreas weight to body weight ratios in 26-week-old mice (Extended Data Fig. 7i). We observed PanINs present in pancreata from both KC;G6pdwt and KC;G6pdmut mice, as assessed by CK19 expression and Alcian Blue stain for PanIN-specific mucins (Fig. 2f and Extended Data Fig. 7h). Pathological grading of H&E-stained tissues demonstrated more transformed area in KC;G6pdmut mice (Fig. 2g and Extended Data Fig. 7j). Additionally, tissues from KC;G6pdmut mice contained more low-grade PanIN lesions (PanIN 1A, 1B, 2) (Fig. 2g and Extended Data Fig. 7j). Interestingly, two KC;G6pdmut mice displayed high-grade lesions (PanIN 3), which were absent in KC;G6pdwt mice. Pancreata from KC;G6pdmut mice also exhibited less acinar cell area and more ADM. We aged mice to 1 year to determine whether the high-grade PanIN lesions present in KC;G6pdmut mice progress to PDAC (Fig. 2h,i and Extended Data Fig. 7k). However, we saw no significant difference in pancreata between KC;G6pdwt and KC;G6pdmut mice after pathological grading of H&E-stained tissues (Fig. 2i and Extended Data Fig. 7k).
Overall, these data illustrate that G6PD deficiency in KrasG12D-driven mouse models increased the number and rate of precancerous lesion formation in 16-week-old and 26-week-old animals. However, after 1 year, our results suggest that G6PD deficiency does not affect PanIN to PDAC progression.
G6PD deficiency does not affect survival in the PDAC mouse model
We next assessed the impact on pancreatic cancer progression and survival using the more rapid KPC mouse model (LSL-KrasG12D; LSL-Trp53R172H; Ptf1aCre)34. KPC mice were crossed to G6PD-deficient mice to determine whether accelerated ADM and PanIN lesions enhanced invasive PDAC and decreased survival (breeding scheme in Supplementary Fig. 4a). There was no significant difference in overall survival between KPC;G6pdmut and KPC;G6pdwt mice (Supplementary Fig. 4b). We assessed histology from pancreas of 90-day-old mice and observed that both genotypes had neoplasia, with significantly increased proliferation in pancreata from KPC;G6pdmut mice (Supplementary Fig. 4c–e). We did not see evidence of metastasis to the liver (Supplementary Fig. 4f), and there was no difference in pancreas weight to body weight ratios (Supplementary Fig. 4g). There was also no significant difference in pathological tissue grading of pancreas from 90-day-old KPC;G6pdmut and KPC;G6pdwt mice (Supplementary Fig. 4h). Although G6PD deficiency promotes precancerous lesions in KC mice, these data indicate that G6PD deficiency does not affect progression to cancer or survival in KPC mice, highlighting a difference in the metabolic pathways that underlie precancer and established cancer.
Me1 loss accelerates ADM, PanIN and PDAC in mouse models
We next sought to determine the role of ME1 in Kras-driven pancreatic carcinogenesis. ME1 is a cytoplasmic enzyme that catalyses the reversible oxidative decarboxylation of malate to pyruvate, generating NADPH (schematic in Fig. 3a). Similar to G6pdx, Me1 was increased in a temporal manner in the primary acinar cultures (Fig. 1n and Extended Data Fig. 6f–h). To assess the necessity of Me1 in pancreatic tumourigenesis, we crossed Me1flox/flox mice35 into the KC model, generating KC;Me1flox/flox mice, in which oncogenic KrasG12D and Me1 loss are induced in the pancreas by Ptf1aCre (breeding scheme in Extended Data Fig. 8a). ME1 protein expression was reduced in pancreata from KC;Me1flox/flox mice (Extended Data Fig. 8b), demonstrating knockout efficiency in the pancreas.
a, Schematic of malic enzyme 1 (ME1) function and related metabolic pathways. ME1 is a cytosolic enzyme that converts malate to pyruvate, generating NADPH in the process. Malate can come from the cytosol or be transported across the mitochondrial membrane from the TCA cycle. The pyruvate generated by ME1 can enter the TCA cycle. Mitochondria (Mito) presented in pink. b, H&E staining and immunostaining for AMY (acinar cells), CK19 (ductal, metaplastic, neoplastic cells) and Ki67 (proliferation) in pancreas of 16-week-old KC;Me1+/+ and KC;Me1flox/flox mice. Scale bars, 50 μm. c, Percentage of AMY-positive area in pancreas as quantified from male (closed circles) and female (open circles) KC;Me1+/+ and KC;Me1flox/flox mice at 16 weeks; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a Student’s t-test (unpaired, two-tailed). d,e, Percentage of CK19+ cells (d) and Ki67+ cells (e) in the pancreas as quantified from male (closed circles) and female (open circles) KC;Me1+/+ and KC;Me1flox/flox mice at 16 weeks; n = 6 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a Student’s t-test (unpaired, two-tailed). f, H&E staining, Alcian Blue staining (for PanIN-produced mucins) and CK19 (ductal, metaplastic, neoplastic cells) immunostaining in 26-week-old KC;Me1+/+ and KC;Me1flox/flox mice. Scale bars, 100 μm. g, Pathological grading of pancreas tissues from 26-week-old KC;Me1+/+ and KC;Me1flox/flox mice, representing the percentage of total tissue area with acinar cells, ADM and PanIN lesions; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with Tukey’s post hoc test. h, H&E staining in 1-year-old KC;Me1+/+ and KC;Me1flox/flox mice. Scale bar, 100 μm. i, Pathological grading of pancreas tissues from 1-year-old KC;Me1+/+ and KC;Me1flox/flox mice, representing the percentage of total tissue area with acinar cells and PDAC; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with Tukey’s post hoc test.
At 8-weeks-old, there was no significant difference in ADM lesions present in KC;Me1flox/flox mice compared to KC;Me1+/+ controls (Extended Data Fig. 8c–g). However, at 16 weeks, more ADM and PanIN lesions were present in KC;Me1flox/flox mice, as indicated by significantly decreased AMY, increased CK19 and increased Ki67 (Fig. 3b–e). No significant difference in pancreas weight to body weight ratios were observed at 8 weeks and 16 weeks (Extended Data Fig. 8h). Precancerous lesions are found in pancreata of 26-week-old KC;Me1+/+ and KC;Me1flox/flox mice, as observed by H&E, Alcian blue and CK19 staining (Fig. 3f and Extended Data Fig. 8g). Yet there was no significant difference in pathological grading (Fig. 3g and Extended Data Fig. 8i). We also did not find significant differences in pancreas weight to body weight ratios (Extended Data Fig. 8h). To determine whether Me1 loss influences progression to PDAC, we aged mice to 1 year and assessed pancreas histology by H&E staining and pathological grading (Fig. 3h,i). Surprisingly, all KC;Me1flox/flox mice developed PDAC (Fig. 3i), while none of the KC;Me1+/+ controls progressed past PanIN 2 lesions (Extended Data Fig. 8j). Additionally, KC;Me1flox/flox pancreata had a smaller acinar cell area (Fig. 3i).
Overall, ADM and PanIN lesions are more abundant and occur sooner (at 16 weeks) in pancreata of KC;Me1flox/flox mice than in KC;Me1+/+ controls. These data suggest that Me1 loss, consistent with G6PD deficiency, promotes precancerous lesions in KrasG12D-driven mouse models. However, we also observe no significant effect of Me1 loss at 26 weeks but a unique promotion of PDAC at 1 year. Together, these results reveal common and diverging roles of NADPH-generating enzymes at distinct stages of tumourigenesis and progression.
G6PD deficiency and Me1 loss do not alter pancreatic endocrine and exocrine function
We sought to determine whether impaired function of NADPH-producing enzymes disrupts normal pancreas physiology, which could predispose to precancerous lesion formation36,37. We performed glucose tolerance tests on Me1+/+, Me1−/−, wild-type, KC;G6pdwt and KC;G6pdmut mice after an overnight fast (Supplementary Fig. 5a–h). There were no significant differences in glucose tolerance across genotypes, although previous studies show that hemizygous G6PD-deficient male mice have smaller islets and impaired glucose tolerance38. The only significant change we detected was a reduction in fasted body weight in Me1−/− mice compared to Me1+/+ controls (Supplementary Fig. 5c). We also tested exocrine pancreas function through serum amylase activity tests, using cerulein treatment as a positive control (Supplementary Fig. 5i). We did not observe significant differences in serum amylase activity across groups compared to wild-type mice. Finally, we checked for premature carboxypeptidase A1 (CPA1) enzyme activation (Supplementary Fig. 5j). Cerulein-treated positive controls showed cleavage of the CPA1 pro-peptide39, but this was not observed in the other mice.
Increased ROS is present with G6PD deficiency and Me1 loss
We used mass spectrometry to assess NADPH, NADP+ and glutathione levels in 16-week-old pancreas tissue. The NADPH/NADP+ ratios trended toward less cellular reducing capacity in pancreata of KC;G6pdmut and KC;Me1flox/flox mice relative to controls but did not reach statistical significance, and there were no significant differences in glutathione levels (Supplementary Fig. 6a–l). These results demonstrate the critical requirement for tissues to maintain NADPH and antioxidants, and sufficient compensation by other metabolic pathways.
We next asked whether reduced NADPH-producing enzyme function led to increased oxidative stress in the pancreas. To assess lipid peroxidation, we performed immunoblotting on pancreas lysates for hyperoxidized peroxiredoxin 3 (HO-PRDX3), malondialdehyde (MDA) and 4-hydroxynonenal (4-HNE) (Fig. 4a,b). Pancreata from KC;G6pdmut and KC;Me1flox/flox mice exhibited increased lipid peroxidation compared with their littermate controls (Fig. 4a and Extended Data Fig. 9a). There was trend toward increased MDA immunostaining in KC;G6pdmut pancreas (compared to KC;G6pdwt) and significantly more MDA in KC;Me1flox/flox pancreas (compared to KC;Me1+/+) (Extended Data Fig. 9b–d). Pancreata from KC;G6pdmut mice also had higher levels of 4-HNE than KC;G6pdwt controls (Extended Data Fig. 9e,f).
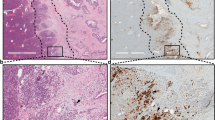
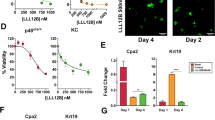
Western blots for lipid peroxidation markers malondialdehyde (MDA), hyperoxidized peroxiredoxin 3 (HO-PRDX3) and 4-hydroxynonenal (4-HNE) normalized to Vinculin loading control. Samples are pancreas lysates (n = 2 per genotype). Western blot was repeated two independent times. b, H&E staining of pancreas from 11–12-week-old KC;G6pdwt and KC;G6pdmut mice after administration of control or N-acetylcysteine (NAC; 30 mM)-containing water for 5–6 weeks. Scale bars, 50 μm. c, Pathological grading of pancreas tissues from KC;G6pdwt and KC;G6pdmut mice from b representing the percentage of total tissue area with acinar cells, ADM and PanIN; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with multiple comparisons. Where P < 0.0001, the exact value is 1.0 × 10−5. d, Immunostaining for CK19 (ductal, metaplastic, neoplastic cells) in pancrease of 11–12-week-old KC;G6pdwt and KC;G6pdmut mice after administration of untreated or N-acetylcysteine water for 5–6 weeks. Scale bars, 50 μm. e, Percentage of CK19+ cells in the pancreas as quantified from male (closed circles) and female (open circles) KC;G6pdwt and KC;G6pdmut mice in d; n = 4 mice for each genotype. Bars represent the mean values; error bars, s.d. P values were calculated using ordinary one-way ANOVA with Tukey’s multiple comparisons test. f, Schematic of de novo glutathione synthesis. Intracellular cysteine (Cys) is converted to glutathione (GSH). The rate-limiting enzyme glutamate cysteine ligase (GCL) is made up of a catalytic subunit (GCLC) and a regulatory, modifier subunit (GCLM). GCL catalyses the condensation of cysteine and glutamate (Glu) to the dipeptide precursor γ-glutamylcysteine (γ-Glu-Cys). Glutathione synthetase (GSS) catalyses the condensation of γ-Glu-Cys and glycine (Gly) to GSH. Buthionine sulfoximine (BSO) inhibits GCL. g, Brightfield images of primary acinar cell cultures isolated from human donor pancreata at day 0 and day 6. Cells were treated with vehicle (DMSO), TGFα (50 ng ml−1) or BSO (100 μM). Scale bars, 100 μm. h, Quantification of acinar cells undergoing ADM in primary cultures at day 6 in vehicle-treated, TGFα-treated and BSO-treated cells. Each symbol shape represents technical duplicates from a unique donor (n = 4 donor pancreata). Bars represent the mean values; error bars, s.d. P values were calculated using repeated measures one-way ANOVA with Dunnett’s multiple comparisons test. Where P < 0.0001, the exact value is 6.1 × 10−5. i, Representative immunostaining for G6PD and ME1 from human ADM primary cultures at day 6. Scale bars, 25 μm. j, H&E staining in pancreata from 5–6-week-old KC mice administered control or BSO-containing water for 5 weeks. Scale bars, 50 μm. k, Pathological grading of pancreata from 5–6-week-old KC mice administered control or BSO-containing water for 5 weeks. Datapoints represent the percentage of total tissue area with acinar cells, ADM and PanIN lesions; n = 4 mice for each treatment. Bars represent the mean values; error bars, s.d. P values were calculated using a two-way ANOVA with Tukey’s multiple comparisons test.
Antioxidant treatment rescues accelerated ADM formation in primary acinar cultures
Given the established role of ROS in ADM formation13, we hypothesized that accelerated ADM could be rescued by antioxidant treatment. Acinar cells from KC;G6pdmut and KC;G6pdwt mice were treated with vehicle or the cell-permeable glutathione analogue glutathione ethyl ester (GSHee). Consistent with our in vivo findings, KC;G6pdmut acinar cells underwent ADM more rapidly in ex vivo culture compared with KC;G6pdwt cells, which was restored by GSHee treatment (Extended Data Fig. 9g–k). We performed similar experiments using acinar cells isolated from Me1+/+ and Me1−/− mice (Supplementary Fig. 7a–c). Acinar cells were 3D-embedded in Matrigel and cultured for 3 days with TGFα to induce ADM. Cells were treated with vehicle, GSHee or N-acetylcysteine (NAC) antioxidants. We observed a significant increase in ADM formation in Me1−/− cells compared to controls, which could be rescued with NAC treatment (Supplementary Fig. 7a–c). As with G6PD deficiency, the persistence of enhanced ADM in isolated Me1−/− acinar cells indicates that this phenotype is redox regulated. Finally, we wanted to determine how ADM formation rates compared between G6pd-mutant, Me1-null and Nrf2-null acinar cells. In collagen cultures, both KC;G6pdmut and Me1−/− cells had significantly higher ADM formation rates than the respective controls (Supplementary Fig. 7d–g). Surprisingly, Nrf2−/− cells failed to form ADM in both collagen cultures and Matrigel cultures (Supplementary Fig. 7f–h). Taken together, our experiments suggest that ROS induce NRF2-signalling to promote ADM.
Antioxidant treatment rescues accelerated lesion formation in KC;G6pdmut mice
To test whether antioxidants can rescue lesion formation in vivo, we provided KC;G6pdwt and KC;G6pdmut mice drinking water with or without NAC. No significant differences in pancreas to body weight ratios were observed between mice given control and NAC water (Extended Data Fig. 9l). H&E staining and histological grading showed that pancreas from KC;G6pdmut mice had significantly more ADM and PanIN lesions, coincident with a decrease in per cent acinar area, compared to pancreas from KC;G6pdwt mice (Fig. 4b–c and Extended Data Fig. 9m). NAC treatment in KC;G6pdmut mice reduced lesions back to levels found in KC;G6pdwt mice and increased per cent acinar area (Fig. 4b,c). This finding was also supported by a significant increase in the percentage of CK19+ cells in pancreas from KC;G6pdmut mice compared to KC;G6pdwt controls, which could be rescued by NAC treatment (Fig. 4d,e).
Inhibiting glutathione biosynthesis promotes ADM in human acinar cells and PanIN formation in KC mice
We analysed a previously published human ADM RNA-seq dataset40 and observed a significant upregulation of both G6PD and ME1 at day 6 following TGFα treatment (Extended Data Fig. 10a). We obtained human donor pancreas through a research partnership with the Gift of Life Michigan organ and tissue donation programme6. Human acinar cells from four independent donors were isolated and 3D-cultured in Matrigel for 6 days (Extended Data Fig. 10b,c). To induce ADM, cells were treated with TGFα (positive control)41. To decrease antioxidant capacity, we depleted cellular GSH pools with buthionine sulfoximine (BSO), an inhibitor of de novo glutathione synthesis (schematic in Fig. 4f). Compared to vehicle, TGFα and BSO treatment significantly increased ADM formation (Fig. 4g,h) and significantly increased the diameter of ADM spheres (Extended Data Fig. 10d). We also saw elevated G6PD and ME1 protein expression in human ADM by immunostaining (Fig. 4i).
Based on the experiments in human cultures, we reasoned that decreasing glutathione would be sufficient to drive more metaplastic and neoplastic lesions in vivo. We provided KC mice control drinking water or water that was supplemented with BSO for 5 weeks. BSO treatment significantly reduced glutathione levels in the pancreas (Extended Data Fig. 10e–g) and did not significantly change pancreas weight to body weight ratios (Extended Data Fig. 10h). Histological grading of H&E-stained pancreas tissue showed that BSO treatment significantly increased ADM and PanIN lesions, concurrent with a decrease in acinar cell area (Fig. 4j,k and Extended Data Fig. 10i). This finding was also supported by a significant increase of CK19+ cells in BSO-treated KC mice (Extended Data Fig. 10j,k). These data suggest that inhibiting glutathione synthesis is sufficient to accelerate lesion formation in KC mice, overall pointing to redox regulation as a key factor in pancreatic precancer formation.
Discussion
Metabolic dependencies in advanced cancers have been extensively studied and targeted therapeutically, but metabolic alterations in precancer are not well characterized. Previous work has demonstrated that KrasG12D drives ROS production and antioxidant pathways during ADM and pancreatic tumourigenesis12,13,14,42,43,44. Yet the function of ROS in tumourigenesis is complex, with studies showing pro-tumour and anti-tumour roles. This duality stems, in part, from ROS-dependent cell toxicity versus mitogenic signalling. Our study and others show that antioxidant treatment dampens ADM formation ex vivo and tumourigenesis in vivo by lowering ROS levels13,45. By contrast, abrogating antioxidant defences through genetic deletion of Nrf2, a primary antioxidant regulator, results in high levels of ROS that drive cellular senescence and reduce PanIN formation12. NRF2-target genes also have dynamic roles in tumourigenesis, as seen in the lung, where Gsr loss impairs lung tumour initiation, while Txnrd1 loss impairs tumour progression17. Our work complements these studies and shows that NRF2-target genes have distinct roles during each step in cancer initiation and progression, as demonstrated by the promotion of early lesions with G6PD deficiency or Me1 loss, but the promotion of PDAC only with Me1 loss.
We interrogated the role of NADPH-producing enzymes G6PD and ME1 in pancreatic precancer formation. G6PD deficiency accelerated the rate of KrasG12D-driven ADM in primary, ex vivo acinar cultures and increased the rate and number of ADM and PanIN lesions in KC mice. Similarly, Me1 loss accelerated ADM in cultures and increased the rate and number of ADM and PanIN lesions in KC mice. Our data suggest that this acceleration is a result of increased oxidative stress from decreased NADPH production, as antioxidant treatment can rescue metaplasia and neoplasia acceleration in cultures and in vivo. Additionally, depleting antioxidant levels by glutathione synthesis inhibition promotes ADM and PanIN formation in KC mice. These findings are consistent with a recent study showing increased precancerous lesions with the loss of Aldh1l2, a gene that encodes a mitochondrial NADPH-producing enzyme46. Together, this highlights the interconnection of NADPH metabolism and how it functions to maintain cellular reducing potential as a barrier against precancer formation.
Although both G6PD deficiency and Me1 loss promote early tissue transformation, Me1 loss uniquely accelerates malignant progression to PDAC. This highlights clear differences in the necessity of these metabolic enzymes in precancer and advanced disease. Outside of their shared roles in NADPH production, G6PD and ME1 appear to impose distinct metabolic constraints on evolving lesions, which may only become limiting as cells have more demands in malignant progression. Our data suggest a conserved function of NADPH regulation during early tissue transformation, but NADPH-independent metabolic activities for PanIN to PDAC transformation.
Recent studies show a surprisingly high frequency of precancerous lesions present in the healthy human pancreas6,7. These findings raise several questions on how lesions form, which lesions progress to cancer and what pathways underlie formation and progression. Changes in cell and systemic metabolism during lesion development and progression pose attractive targets for biomarkers and dietary interventions. Indeed, new research suggests that circulating formate levels may serve as a biomarker for PDAC progression46. Furthermore, this supports a role of oxidative stress in regulating cancer initiation in the pancreas12,13,14,18,29. However, a careful and comprehensive understanding of precancer metabolic changes, especially in antioxidant pathways, is required before translation to humans. As a cautionary example, clinical trials of antioxidant supplementation in individuals who were at high risk for lung cancer increased cancer incidence and mortality47,48. Therefore, additional studies in precancer across tissues and in diverse patient populations are necessary to improve human health outcomes.
Methods
Animal use
Mouse experiments were conducted under the guidelines of the Office of Laboratory Animal Welfare and approved by the Institutional Animal Care and Use Committees at the University of Michigan under protocol PRO00010606. Animal experiments complied with all relevant ethical regulations. Mice were housed in ventilated racks in a pathogen-free animal facility with a 12 h light, 12 h dark cycle, 30–70% humidity and 20–23 °C temperatures maintained in the animal facility at Rogel Cancer, University of Michigan. Mice were overseen by the Unit for Laboratory Animal Medicine. Mice were fed ad libitum (5L0D, PicoLab Laboratory Rodent Diet). Both male and female mice were used in experiments unless specifically indicated. Tumour monitoring was performed to ensure that the maximal tumour size and burden was not exceeded. In line with ethical guidelines, a tumour diameter of 2 cm in any single dimension required that the mice be killed. All mice in this study were killed at the experimental endpoint, and the maximum tumour size was never exceeded.
Mouse strains
LSL-KrasG12D mice (Jackson Laboratory, strain no. 008179) were crossed with Ptf1aCreERTM mice49 to generate LSL-KrasG12D; Ptf1aCreERTM mice and maintained on a C57BL/6J background. G6PD-deficient mice32,33,50 were originally generated in the C3H strain. They have been crossed to other strains, including C57BL/6J. G6PD-deficient mice were maintained on a mixed background and compared to age-matched littermate controls. LSL-KrasG12D; Ptf1aCre; G6pdmut or G6pdwt mice were maintained on a mixed background and were confirmed Nnt wild-type by genotyping. LSL-KrasG12D; Ptf1aCre mice51 were crossed to Me1flox/flox mice35 to generate LSL-KrasG12D; Ptf1aCre/+; Me1flox/flox mice and maintained on a C57BL/6J background. All the LSL-KrasG12D; Ptf1aCre; Me1flox/flox mice presented in these studies were Nnt-null. LSL-KrasG12D; Ptf1aCre; Me1flox/flox mice in these studies were compared to LSL-KrasG12D; Ptf1aCre; Me1+/+ littermates and LSL-KrasG12D; Ptf1aCre age-matched, historic controls. Germline, whole-body Me1-null mice (Me1−/−)35 were maintained on a C57BL/6J background and compared to wild-type littermates for experiments. Nrf2−/− mice52 (Nfe2l2tm1Ywk; Jackson Laboratory, strain no. 017009) were maintained on a C57BL/6J background.
Genotyping
Genotyping was performed through a commercial vendor (Transnetyx) or in house. Genotyping performed by Transnetyx used real-time PCR with specific probes designed for each target. In-house genotyping was performed using DNA extracted from tail or ear tissue using DirectPCR Lysis Reagent (Viagen Biotech). Genotyping primer sequences are included in Supplementary Table 1. Thermal cycling was performed using GoTaq Green Master Mix (Promega). Gel electrophoresis was performed on 2% agarose gel with 10,000X SYBR Safe DNA gel stain (Invitrogen). DNA Ladders (1 kb and 100 bp) (New England Biolabs) were used to measure amplicon size. Gels were run at 140 V for 50 min and imaged using the ChemiDoc Imaging System (Bio-Rad). G6pd mouse PCR primers were made between exon 1 and intron 1, around the reported mutation site. The PCR product was digested using the DdeI restriction enzyme and run on a 3% agarose gel. G6pd wild-type mice have two bands (214 and 55 bp), G6pd-mutant homozygotes have one band (269 bp) and G6pd-mutant heterozygotes have three bands (269, 214 and 55 bp).
Treatment procedures in mice
NAC
NAC (N-acetyl-L-cysteine; Cayman Chemical, 20261) was added to drinking water at a concentration of 30 mM with pH adjusted to 7, matching control water. Water was changed weekly. Male and female LSL-KrasG12D; Ptf1aCre; G6pdmut or G6pdwt mice aged 5–6 weeks were randomly selected to receive NAC water or normal water for 5–6 weeks.
BSO
BSO (L-Buthionine-(S,R)-sulfoximine; Cayman Chemical, 14484) was added to drinking water at a concentration of 20 mM53 and changed weekly. Male and female LSL-KrasG12D; Ptf1aCreERTM mice aged 5–6 weeks were randomly selected to receive BSO water or normal water for 5 weeks.
Cerulein
Serum and pancreas from cerulein-treated wild-type mice (C57BL/6J) were used as positive controls for serum amylase tests and anti-CPA1 western blots, respectively. Two male and two female mice were administered 75 µg kg−1 cerulein (dissolved in sterile PBS) by intraperitoneal injection once every hour for 8 h, then administered two additional hourly injections the following day before blood and tissue collection.
Acinar cell isolation from mice
Pancreata from 8–10 week-old mice were dissected and rinsed twice in 5 ml cold Hanks’ Balanced Salt Solution (HBSS; Gibco). Tissue was minced with sterile scissors into 1–5 mm-sized pieces, then centrifuged for 2 min at 300g and 4 °C. Media was aspirated, and minced tissue was digested with 5 mg of Collagenase P (Roche, Sigma, 11213857001) dissolved in 5 ml cold HBSS. Tissue was digested at 37 °C for 15–20 min with gentle agitation on an orbital shaker. Collagenase P was inhibited by the addition of 5 ml cold 5% FBS (Corning, 35-010-CV) in HBSS. Cells were centrifuged for 2 min at 300g and 4 °C, then washed with 5 ml cold 5% FBS in HBSS. This process was repeated twice. Cells were passed through a 500 μm strainer (Pluriselect, Fisher, NC0822591), followed by a 100 μm strainer (Fisherbrand, 22363549) and then pelleted through 10 ml of 30% FBS gradient. Cells were resuspended in media and incubated at 37 °C for at least 4 h before plating. Cells were cultured in 1× Waymouth’s media (Gibco, 11220035) supplemented with 0.4 mg ml−1 soybean trypsin inhibitor (Gibco, 17075-029), 1 μg ml−1 dexamethasone (Sigma-Aldrich, D4902) and 0.5% gentamicin (Lonza, 17-519L). All media was sterilized through a 0.22 μm PVDF membrane (Millipore, Stericup Filter Unit, SCGVU01RE; Steriflip Filter Unit, SE1M179M6).
3D acinar cell culture
For measuring transdifferentiation rates, acinar cells were embedded in Matrigel (Corning, 354234) domes or Collagen gel (Gibco, A1048301; as described in previous work54) and plated in 48-well plates. Culture media was added on top of the solidified matrix and changed on day 2 after plating. Cells were treated with vehicle (water), 1 mM GSHee (Cayman Chemical, 14953) or 500 µM NAC (Cayman Chemical, 20261; pH adjusted to 7) dissolved in water. Cells were imaged using the EVOS FL Auto Imaging System (ThermoFisher Scientific). Isolated acinar cells were seeded in triplicate, and per cent ADM quantifications were counted from 4× optical fields per well. The diameter of ADM spheres was measured using the line tool in ImageJ (v.1.51n) from imported 4× images. Statistics for per cent ADM and diameter were calculated using GraphPad Prism 10 and analysed using ordinary one-way ANOVA with Dunnett’s multiple comparisons test.
For KrasG12D induction in acinar cell cultures, acinar cells were isolated from three independent male LSL-KrasG12D; Ptf1aCreERTM littermates at 11 weeks old. Cells were treated with either vehicle (ethanol) or 2 μM 4-OHT (Cayman Chemical, 19519) to induce expression of mutant KrasG12D. RNA and metabolites were isolated 24 h after plating vehicle-treated acinar cells (vehicle) and 24, 48 and 72 h after plating 4-OHT-treated cells (days 1, 2 and 3, respectively).
RNA isolation and RNA-seq from acinar cell cultures
Acinar cells were grown free-floating in non-tissue culture-treated six-well plates for RNA isolation. Primary acinar cell cultures were collected by centrifugation for 2 min at 300g. Half of the cell pellet was lysed in RLT buffer containing 1% β-mercaptoethanol (Sigma-Aldrich, M6250), flash-frozen in liquid nitrogen and stored at −80 °C for brief storage. Samples were quickly thawed and passed through a Qiashredder column (Qiagen, 79654). RNA was purified using a RNeasy Plus Mini Kit (Qiagen, 74136) and analysed on a Nanodrop 2000c (Thermo Scientific) spectrophotometer for quantification. RNA was further assessed with a Qubit Assay to determine concentration and quality assessment with an Agilent TapeStation through the University of Michigan Advanced Genomics Core. All samples for sequencing had an RNA integrity number greater than 7.9 and a DV200 greater than 90. Samples were submitted to GENEWIZ from Azenta Life Sciences for standard RNA-seq using Illumina NovaSeq X Plus with 20 million reads per sample and ribosomal RNA removal using PolyA selection for messenger RNA (mRNA) species. ERCC RNA Spike-In Mix (ThermoFisher Scientific, 4456740) was added to normalized total RNA before library preparation using the manufacturer’s protocol. RNA-seq libraries were prepared using the NEBNext Ultra II RNA Library Prep Kit for Illumina following the manufacturer’s instructions (NEB). In brief, mRNAs were initially enriched with Oligod(T) beads. Enriched mRNAs were fragmented for 15 min at 94 °C. cDNA fragments were end-repaired and adenylated at 3′ ends, and universal adaptors were ligated to cDNA fragments, followed by index addition and library enrichment by PCR with limited cycles. The sequencing library was validated on the Agilent TapeStation (Agilent Technologies) and quantified by a Qubit 3.0 Fluorometer (Invitrogen) and qPCR (KAPA Biosystems). Data are publicly available from NCBI’s Gene Expression Omnibus (GEO) database (GSE313699).
RNA-seq data analysis
RNA-seq fastq reads were transferred to the University of Michigan Great Lakes computational cluster for pseudoalignment to the mouse genome (Gencode version M35) using Salmon (v.1.9.0)55 to generate a counts per sample matrix. RNA-seq count matrices were analysed in edgeR to identify differentially expressed genes between experimental conditions. The expression data were filtered to remove genes with low counts using the edgeR filterByExpr() function with default settings. Normalization factors and dispersion parameters were then calculated before generating a log2-transformed counts per million matrix for analysis. Overall gene expression patterns per sample were first visualized with a multidimensional scaling plot. Differential gene expression between each timepoint (days 1, 2 and 3) and vehicle control was estimated using the quasi-likelihood negative binomial generalized log-linear approach in edgeR. Genes were considered differentially expressed between a treatment and control at a false discovery rate-adjusted P value of <0.05. Enriched differentially expressed genes per treatment were identified using gene set enrichment analysis in FGSEA56. To identify enriched transcription factor binding, the top 500 overexpressed or underexpressed genes within a condition were uploaded to the Enrichr web tool26,27,28 to quantify enrichment for ENCODE and ChEA consensus transcription factor targets. We visualized gene expression patterns using pheatmap() in R or using GraphPad Prism. Analyses were conducted using R software (v.4.2.2).
14C glucose incorporation into CO2
Cells were treated with 1 μCi 1-14C (Perkin Elmer, NEC043X050UC) or 6-14C glucose (Perkin Elmer, NEC045X050UC) and incubated at 37 °C for 4 h. To release 14CO2, 150 µl of 3 M perchloric acid (Sigma-Aldrich, 244252) was added to each well and immediately covered with Whatman paper saturated with phenylethylamine (Sigma-Aldrich, P6513) and then incubated at room temperature (25 °C) overnight. The Whatman paper was then analysed by scintillation counting (Beckman, LS6500) and normalized to surrogate protein quantification.
Real-time qPCR
RNA was extracted from acinar cell cultures using the RNeasy Mini Kit (Qiagen, 74104) according to the manufacturer’s instructions. RQ1 RNase-Free DNase (Promega, M6101) was added to samples, and then cDNA was generated using iScript Reverse Transcription Supermix (Bio-Rad, 1708840). Quantitative PCR was performed using Fast SYBR Green Master Mix (Applied Biosystems, 4385612) on a QuantStudio 5 PCR System (Applied Biosystems). Gene expression levels were normalized to the expression of the endogenous control Mrpl. Relative expression was calculated using the 2−ΔΔCT method57. Experiments were performed in technical triplicates. Primer sequences are included in Supplementary Table 2.
Sample processing for histology
Mice were killed by CO2 asphyxiation or isoflurane overdose, followed by cervical dislocation. Tissue was quickly dissected and fixed overnight at room temperature with zinc formalin fixative (Z-Fix; Anatech). Fixed tissues were transferred to 70% ethanol and embedded in paraffin. Primary acinar cells cultured in Matrigel were rinsed in cold PBS, pelleted and fixed in 10% formalin at 4 °C for 1 h. Cells were rinsed in PBS, moved to 70% ethanol, pelleted, mounted in 2% agar (Fisher, BP1423) and embedded in paraffin. Samples were processed using a Leica ASP300S Tissue Processor (Leica Microsystems) and cut into 5 μm-thick sections.
Immunohistochemistry
Tissue sections were deparaffinized with Histo-Clear (National Diagnostics) and rehydrated with graded ethanol and water. Samples were quenched with 1.5% hydrogen peroxide in 100% methanol for 15 min at room temperature. Antigen retrieval was performed in sodium citrate buffer (2.94 g sodium citrate, 500 μl Tween 20, pH 6.0) using a pressure cooker on high or by maintaining slides at a rolling boil for 20 min. Slides were blocked in PBS containing 2.5% BSA (Sigma-Aldrich) and 0.2% Triton X-100 for 45 min to 1 h at room temperature. The slides were then incubated overnight at 4 °C in primary antibodies, with the exception of the anti-malondialdehyde primary antibody, which was incubated for 1 h at room temperature. After rinsing with PBS, slides were incubated with secondary antibodies for 1 h at room temperature. Blocking of endogenous biotin, biotin receptors and avidin binding sites was performed using the Avidin/Biotin Blocking Kit (Vector Laboratories) per the manufacturer’s protocol. Substrate reaction and detection was performed using DAB Peroxidase (HRP) Substrate Kit (With Nickel), 3,3′-diaminobenzidine (Vector Laboratories, SK4100) as detailed per the manufacturer’s protocol. Slides were counterstained, mounted in Permount Mounting Medium (Fisher Scientific), coverslipped and allowed to dry overnight before imaging. Antibody information is included in Supplementary Table 3.
Counterstains and special stains
H&E staining was performed using Mayer’s haematoxylin solution and Eosin Y (Thermo Fisher Scientific). Blue and Nuclear Fast Red staining was performed using the Alcian Blue (pH 2.5) Stain Kit (Vector Laboratories, H-3501) per the manufacturer’s protocol.
Imaging and quantification of immunostaining
Imaging was performed automatically using a whole-slide scanner (Vectra Polaris, Akoya Biosciences) or manually using an Olympus BX53F microscope. Whole-slide scanning was performed by the University of Michigan Tissue and Molecular Pathology Shared Resource. To quantify staining from slide scans, three to five regions of interest at ×10 magnification were randomly selected using QuPath58 (v.0.6.0) and the positive DAB signal was quantified. Manual imaging was performed on an Olympus BX53F microscope fitted with an Olympus DP80 digital camera (Olympus Life Science), using CellSens Standard software. To quantify staining from manual images, three to five images were taken per slide at ×10 magnification. Images were anonymized and randomized, and a positive DAB signal was quantified using QuPath58 (v.0.6.0). For tissue grading, at least three ×20 fields were reviewed by a pathologist and graded in a blinded manner.
Polar metabolite isolation
From primary ex vivo acinar cell cultures
Floating cells from primary cultures were collected by centrifugation for 2 min at 300g. Half of the cell pellet was used to isolate intracellular metabolites by lysis in dry ice-cold 80% methanol with repeated vortexing for 10 min. Media from cultures was used to isolate extracellular metabolites by adding 0.2 ml of media to 0.8 ml dry ice-cold 100% methanol. The solution was repeatedly vortexed and incubated on dry ice for 10 min. Both intracellular and extracellular samples were centrifuged at 15,000g for 10 min at 4 °C to pellet insoluble material. The metabolite supernatant was transferred to a fresh tube and dried by vacuum centrifugation. The dried pellet was resuspended in a 50:50 methanol:water mixture for metabolomics.
From pancreas tissue
Upon collection, pancreas tissue was flash-frozen in liquid nitrogen and stored at −80 °C until metabolite isolation. Frozen tissue samples were weighed and added to 2 ml Eppendorf tubes with 1 ml of ice-cold 80% methanol (diluted in 20% ultrapure water). Then, samples were immediately disrupted on a Qiagen TissueLyser II with metallic beads (Qiagen, 69989) at 30 s intervals until fully homogenized. Samples were centrifuged at 15,000g for 10 min at 4 °C to pellet insoluble material. Metabolite supernatant volume was normalized by pancreas tissue weight, transferred to a fresh tube and dried by nitrogen purging or vacuum centrifugation. The dried pellet was resuspended in a 50:50 methanol:water mixture. To detect reduced glutathione in samples, pancreas tissue was dissociated in ice-cold 80% methanol (diluted in 20% ultrapure water) containing 25 mM N-ethylmaleimide (Cayman Chemical, 19938, CAS 128-53-0). After homogenization, samples were incubated at room temperature for 30 min, allowing for derivatization of thiols. The remainder of sample preparation followed the same procedure described above.
Metabolomics methods
General LC–MS-grade chemicals and solutions used included acetonitrile (Fisher, A955-4), methanol (Sigma-Aldrich, A456-4), isopropanol (Fisher, A461-4), ammonia acetate (Sigma-Aldrich, 73594-25G-F), ammonia solution (Sigma-Aldrich, 5438300100) and formic acid (Fisher, A117-50). The water was purified to 18 MΩ·cm using a Millipore filtration system.
Targeted metabolomics from primary acinar cell cultures
Steady state LC–MS/MS-based targeted metabolomics was performed according to published methods59,60 and described here. Samples were run on an Agilent 1290 Infinity II LC system coupled to a 6470 Triple Quadrupole (QqQ) mass spectrometer. The LC–MS/MS platform comprised a 1290 Infinity II LC Flexible Pump (Quaternary Pump), a 1290 Infinity II Multisampler, a 1290 Infinity II Multicolumn Thermostat with six-port valves and the 6470 triple quad mass spectrometer. Compound optimization, calibration and data acquisition were performed using the Agilent MassHunter Workstation Software LC–MS Data Acquisition for 6400 Series Triple Quadrupole MS (software version 10). A total of 145 metabolites were detected in the samples from a 242-metabolite reference library.
The QqQ data were pre-processed with Agilent MassHunter Workstation QqQ Quantitative Analysis Software (B0700). Raw metabolite abundance levels from day 1, day 2 and day 3 samples were divided by the median of raw metabolite abundance levels in vehicle samples to allow for metabolite comparison across days, dependent on their change compared to vehicle. Values were then log2 transformed for visualization in volcano plots and heatmaps. Median normalized values were analysed by a two-tailed t-test to determine statistical significance, with a P < 0.1 cutoff to indicate metabolites of interest. Heatmaps and volcano plots were generated using GraphPad Prism 10. Principal component analysis plots were generated using MetaboAnalyst (https://www.metaboanalyst.ca). Metabolomics source data for this experiment are included in Supplementary Tables 4–6 (intracellular metabolites) and Supplementary Tables 7–9 (extracellular metabolites).
NADP+ and NADPH detection in pancreas tissue
Samples were analysed on an Agilent 1290 Infinity II LC system coupled to an Agilent 6470 triple quadrupole (QqQ) mass spectrometer, according to previously published methods59,60 and described here. An additional method for the quantification of NAD, NADH, NADP and NADPH was performed on the same hardware with the following parameters.
Solvent A comprised 97% water and 3% methanol, with 15 mM acetic acid and 10 mM tributylamine at a pH of 5. Solvent C consisted of 15 mM acetic acid and 10 mM tributylamine in methanol. Solvent D (acetonitrile) was used as the wash solvent. The LC system seal wash consisted of 90% water and 10% isopropanol, and the needle wash solvent was 75% methanol and 25% water. Tributylamine (GC-grade, 99%; ACROS Organics), acetic acid (LC–MS-grade Optima; Fisher Chemical), InfinityLab Deactivator additive and ESI–L Low Concentration Tuning mix (Agilent Technologies) and LC–MS-grade water, acetonitrile, methanol (Millipore) and isopropanol (Fisher Chemical) were used.
An Agilent ZORBAX RRHD Eclipse Plus C18 column (2.1 × 50 mm, 1.8 µm particle size) was used. The gradient profile was 90% solvent A, 10% solvent B for 0–0.8 min; 80% solvent A, 20% solvent B for 0.8–2.5 min; 1% solvent A, 99% solvent B for 2.5–6.5 min; 1% solvent A, 99% solvent B for 6.5–7.5 min; 90% solvent A, 10% solvent B for 7.5–7.6 min; and 90% solvent A, 10% solvent B for 7.6–8.5 min. The flow rate was set at 0.5 ml min−1 with an injection volume of 3 µl at a temperature of 35 °C.
Analytical standards were used to optimize compounds, and a dynamic multiple reaction monitoring (dMRM) strategy was used. Samples were also run alongside standards for NADPH (Cayman Chemical, 9000743, CAS 2646-71-1) and NADP+ (Cayman Chemical, 10004675, CAS 698999-85-8) dissolved in 50% methanol. Data were pre-processed in Agilent MassHunter Quantitative Analysis Software (B0700). Metabolomics source data for this experiment are included in Supplementary Table 10.
Glutathione detection in pancreas tissue
The Thermo Scientific Orbitrap IQ-X LC–MS system was used to detect glutathione. The system consisted of two Vanquish Horizon UHPLC binary pumps, a dual Vanquish autosampler, two Vanquish column compartments with a switch valve for a two-column setup configuration, a Thermo Multiwavelength Detector (CG) and an Orbitrap IQ-X high-resolution mass spectrometer. Data acquisition, instrument tuning, calibration and optimization were performed using the Thermo Scientific Xcalibur data system (v.4.6.67.17) on a Dell Opex-EX3 computer running Windows 10 Professional.
For targeted and untargeted applications, the Waters Acquity UPLC BEH amide column (2.1 × 100 mm, 1.7 µm particle size) with a Waters UPLC BEH Amide VanGuard pre-column (1.7 µm) was used for high-throughput acquisition. Solvent A consisted of 10 mM ammonium acetate in water with 10 mM ammonia (pH 8.0). Solvent B consisted of acetonitrile. The pump seal wash and autosampler wash comprised 50% isopropanol and 50% water with 0.1% formic acid, and the needle wash solvent consisted of 80% acetonitrile and 20% water. The LC profile was a 0.2 ml per minute flow rate of of 85% B for 0–0.5 min, 15% B for 0.5–15 min, 15% B for 15–17 min, 85% B for 17.1 min and 85% B for 22 min. The column compartment temperature was maintained at 30 °C, and the autosampler was held at 5 °C; the injection volume was 3 µl.
The Thermo Scientific Orbitrap IQ-X mass spectrometer was calibrated weekly using Pierce FlexMix Calibration Solution (A39329), delivered by infusion into the micro-ESI probe at 5 µl per minute, with 2 ppm accuracy for all calibrations. Source parameters included H-ESI positive static voltage 4,000 V; negative static voltage, 3,500 V; sheath gas, 40 arbitrary units (a.u.); auxiliary gas, 5 a.u.; sweep gas, 1 a.u.; ion transfer tube temperature, 300 °C and vapourizer temperature, 350 °C. Orbitrap resolution was 60 K, and the mass scanning range was 50–1,000 Da. The data type was centroid, using polarity-switching data acquisition mode.
Data processing was performed by Skyline (v.12.1) and Thermo Compound Discoverer (v.3.4).
MS/MS analysis was performed using the Thermo AcqX acquisition method, applying the same LC method on both the MS1 scan and MS/MS, with collision energies of 20, 30 and 50 eV in positive and negative modes separately. One blank, one scan and five MS/MS injections were used from the pooled quality control samples. All MS1 and MS2 datasets were combined to build a data workflow, using Compound Discoverer (v.3.4) for the final output.
Metabolomics source data for this experiment are included in Supplementary Table 11.
Donor sample procurement and processing
The process of acquiring donor pancreas tissue for research purposes through the Gift of Life Michigan has been previously published6 and is described in brief here. Donor pancreata that were ineligible for transplant or for which there were no eligible recipients were collected at the Gift of Life Michigan Donor Care Center and transported to the University of Michigan. Written consent from family members for the use of tissues for research was obtained by the Gift of Life Michigan. Upon arrival, the pancreas was dissected into head, body and tail regions by a pancreatobiliary surgeon. Tissue was used for primary, ex vivo acinar cultures. An adjacent region was fixed overnight in Z-fix (Anatech), rinsed with PBS and 70% ethanol and then paraffin-embedded for histology. Acquisition and use of donor pancreas for research purposes was approved by the Gift of Life Michigan research review group and the University of Michigan Institutional Review Board (HUM00025339). We obtained informed consent, and the studies complied with all relevant ethical regulations.
Primary acinar cell culture from donor pancreas
Donor mouse pancreata were prepared for primary, ex vivo acinar culture using a similar procedure as described above. In brief, tissue was minced with sterile scissors in cold HBSS (Gibco) into 1–5 mm-sized pieces, centrifuged, rinsed and then digested with Collagenase P (Roche, Sigma, 11213857001) dissolved in HBSS. Tissue was digested at 37 °C for 20–30 min with gentle agitation on an orbital shaker. HBSS containing 5% FBS (Corning, 35-010-CV) was added, and cells were centrifuged and rinsed with 5 ml cold 5% FBS in HBSS three times. Cells were passed through a 500 μm strainer (Pluriselect, Fisher, NC0822591), then through a 100 μm strainer (Fisherbrand, 22363549) and then pelleted through a 30% FBS gradient. Cells were resuspended in 1× Waymouth’s media (Gibco, 11220035) supplemented with 0.4 mg ml−1 soybean trypsin inhibitor (Gibco, 17075-029), 1 μg ml−1 dexamethasone (Sigma-Aldrich, D4902) and 0.5% gentamicin (Lonza, 17-519 L). Cells were cultured at 37 °C overnight in a non-tissue culture-treated six-well plate. The following day, floating cells were pelleted, embedded in Matrigel (Corning, 354234) and plated in 24-well plates for measuring ADM transdifferentiation rates. Culture media was added on top of the solidified matrix and changed on days 1, 3 and 5 after plating. Cells were with vehicle (PBS), 50 ng ml−1 TGFα (R&D Systems, 239A100) or BSO (Cayman Chemical, 14484; dissolved in PBS). Cells were imaged daily using the EVOS FL Auto Imaging System (ThermoFisher Scientific). Isolated acinar cells were seeded in triplicates, and per cent ADM quantifications were counted from 4× optical fields per well. The diameter of ADM spheres was measured using the line tool in ImageJ (v.1.51n) from imported 4× images.
Protein detection by western blot
Pancreas tissue was flash-frozen in liquid nitrogen upon collection and stored at −80 °C. Frozen pancreas tissue was added to RIPA buffer (Sigma-Aldrich, R0278) containing protease (Roche, 04693132001) and phosphatase (Sigma-Aldrich, P5726) inhibitors. Then, samples were immediately homogenized on a Retsch TissueLyser II (129251128) with metallic beads at intervals of 30 s. Samples were centrifuged at 4 °C, and the supernatant was collected to measure protein concentration using the Pierce BCA Protein Assay Kit (ThermoFisher, 23227), according to the manufacturer’s protocol. Equal amounts of protein (10–20 µg) were subjected to separation at 135 V for 85 min on SDS–PAGE and then transferred to a methanol-activated PVDF membrane at 20 V for 60 min. Membranes were blocked with 5% milk in TBST (Tris-buffered saline containing 0.1% Tween 20) followed by incubation with primary antibodies diluted in 5% milk at 4 °C overnight. Following incubation, the membranes were washed three times with TBST and incubated with the species-appropriate secondary antibody conjugated to horseradish peroxidase for 1 h at room temperature. Membranes were then washed three times for 5 min each with TBST. Chemiluminescence was detected using Clarity ECL substrate (Bio-Rad, 170-5060), and the signal was captured with a Bio-Rad ChemiDoc Imaging System (Image Lab Touch Software v.2.4.0.03). Densitometric analysis of western blot bands was carried out using ImageJ (v.1.51n). The resulting values were normalized to the intensity of the loading control from the same membrane. Additional proteins of interest were probed on the same membrane after incubation in stripping buffer (ThermoFisher, 46430) and re-blocking.
Glucose tolerance tests
Female mice aged 12–15 weeks were fasted for 15 h, and fasting blood glucose was measured using a Contour Next EZ Glucometer (Ascensia Diabetes Care, 90007941) with Contour Next Test strips (Ascensia Diabetes Care, 85701231). Afterwards, animals were given an intraperitoneal injection of 1.5 g kg−1 glucose (Sigma-Aldrich, G7021-100G) in sterile saline solution (Braun, R5201-01). Blood glucose was measured 30, 60, 90 and 120 min after injection by tail vein bleeding. Animals were restrained for less than 1 min each time blood was collected. Basal blood glucose measurements were taken from the same mice 10 days after glucose tolerance tests were performed.
Serum amylase activity
Blood was collected from 16–18-week-old mice, and serum was separated using BD Diagnostic Solutions Serum Separator Tubes (Fisher Scientific, 02-675-185). Serum was frozen and stored until testing. Quantitative kinetic determination of α-amylase activity in serum was measured using Pointe Scientific Amylase Reagent (Fisher 23-666-111) following the manufacturer’s instructions. In brief, 10 μl of serum was added to 400 μl of reagent and read on a plate reader at 409 nm three times over 2 min. To calculate amylase activity in U l−1 , the average absorbance per minute was multiplied by the assay volume, then by 1,000. This product was divided by the millimolar absorptivity of 2-chloro-p-nitrophenol times the sample volume times the light path.
Statistics and reproducibility
Statistics were calculated using GraphPad Prism 10. A two-tailed Student’s t-test was used when comparing two groups; an unpaired test was used for independent groups and a paired test was performed for matched or repeated measurements. An ordinary one-way ANOVA was used to compare the means of groups with more than 2 single-variable and unpaired or unmatched measurements. A repeated measures one-way ANOVA was used to compare the means of more than 2 groups with matched or paired measurements. A two-way ANOVA was used to compare groups with more than two variables. Tukey’s multiple comparisons test was performed to compare every mean with every other mean. Dunnett’s multiple comparison test was performed to compare every mean to a control mean. Data distribution was assumed to be normal, but this was not formally tested. Survival analysis was performed using the log-rank (Mantel–Cox) test. No statistical methods were used to pre-determine sample sizes, but our sample sizes are similar to those reported in previous publications12,13,14. Sample sizes were selected based on standard experimental group sizes to achieve acceptable power, taking into account the increased variability of animal models (three to four replicates for ex vivo experiments; four to 12 replicates for in vivo experiments). No animals were excluded from analyses. No datapoints were excluded from analyses, except in Extended Data Fig. 10d, in which two outlier datapoints were removed. Outliers were identified by analysing the raw dataset in GraphPad Prism using the ROUT (Robust Regression and Outlier Removal) method, with Q = 1%. Experiments were repeated at least once and results were replicated, except for the RNA-seq and metabolomics experiments, which were performed once with independent biological replicates. Samples from animals were collected on several different days, from different litters and handled by multiple investigators. Staining was performed by at least two independent investigators and replicated on separate days. RNA-seq and metabolomics experiments were performed once using samples prepared from three biological replicates. Where applicable, the sample size (n) of biological replicates and technical replicates is indicated in the figure legends. Animals with the relevant genotypes were randomly placed into groups to collect tissue at each timepoint, with representation from approximately equal numbers of each sex. Staining, analysis and quantification were performed on samples from each group at random. For staining from mouse tissues, an investigator imaged samples and the genotypes were known. The images were then de-identified and randomized, and the positive signal was quantified using QuPath software by a separate investigator in a blinded manner. For tissue grading, slides were de-identified, imaged and graded by a pathologist in a blinded manner.
Availability of biological materials
All unique materials used will be promptly available from companies/suppliers or from the authors upon reasonable request, without undue qualifications. Antibodies can be purchased from corresponding suppliers (Developmental Studies Hybridoma Bank, Abcam, Sigma-Aldrich and Vector Laboratories). Mouse strains can be purchased from Jackson Laboratories (where applicable) or provided by the authors.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data from this study, referenced in Fig. 1, are publicly available from NCBI’s Gene Expression Omnibus (GEO) database (GSE313699). RNA-seq data referenced in Extended Data Fig. 6 are publicly available from the University of Michigan’s Deep Blue Data Repository (10.7302/nd65-zg69)31. Human ADM RNA-seq data referenced in Extended Data Fig. 10 were obtained from GEO (GSE179248)40. All metabolomics datasets are available in Supplementary Tables 4–11. Source data are provided with this paper.
References
Shi, C. et al. KRAS2 mutations in human pancreatic acinar-ductal metaplastic lesions are limited to those with PanIN: implications for the human pancreatic cancer cell of origin. Mol. Cancer Res. 7, 230–236 (2009).
Halbrook, C. J., Lyssiotis, C. A., Pasca di Magliano, M. & Maitra, A. Pancreatic cancer: advances and challenges. Cell 186, 1729–1754 (2023).
Storz, P. Acinar cell plasticity and development of pancreatic ductal adenocarcinoma. Nat. Rev. Gastroenterol. Hepatol. 14, 296–304 (2017).
Burclaff, J. & Mills, J. C. Plasticity of differentiated cells in wound repair and tumorigenesis, part I: stomach and pancreas. Dis. Model. Mech. 11, dmm033373 (2018).
Mills, J. C. & Sansom, O. J. Reserve stem cells: differentiated cells reprogram to fuel repair, metaplasia, and neoplasia in the adult gastrointestinal tract. Sci. Signal. 8, re8 (2015).
Carpenter, E. S. et al. Analysis of donor pancreata defines the transcriptomic signature and microenvironment of early neoplastic lesions. Cancer Discov. 13, 1324–1345 (2023).
Braxton, A. M. et al. 3D genomic mapping reveals multifocality of human pancreatic precancers. Nature 629, 679–687 (2024).
Stangis, M. M. et al. The hallmarks of precancer. Cancer Discov. 14, 683–689 (2024).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Halbrook, C. J. & Lyssiotis, C. A. Employing metabolism to improve the diagnosis and treatment of pancreatic cancer. Cancer Cell 31, 5–19 (2017).
Kerk, S. A., Papagiannakopoulos, T., Shah, Y. M. & Lyssiotis, C. A. Metabolic networks in mutant KRAS-driven tumours: tissue specificities and the microenvironment. Nat. Rev. Cancer 21, 510–525 (2021).
DeNicola, G. M. et al. Oncogene-induced Nrf2 transcription promotes ROS detoxification and tumorigenesis. Nature 475, 106–109 (2011).
Liou, G. Y. et al. Mutant KRas-induced mitochondrial oxidative stress in acinar cells upregulates EGFR signaling to drive formation of pancreatic precancerous lesions. Cell Rep. 14, 2325–2336 (2016).
Cheung, E. C. et al. Dynamic ROS control by TIGAR regulates the initiation and progression of pancreatic cancer. Cancer Cell 37, 168–182.e4 (2020).
Antonucci, L. et al. Self-amplifying NRF2–EZH2 epigenetic loop converts KRAS-initiated progenitors to invasive pancreatic cancer. Nat. Cancer 6, 1263–1282 (2025).
DeBlasi, J. M. et al. Distinct Nrf2 signaling thresholds mediate lung tumor initiation and progression. Cancer Res. 83, 1953–1967 (2023).
Sherwood, A. M., Yasseen, B. A., DeBlasi, J. M., Caldwell, S. & DeNicola, G. M. Distinct roles for the thioredoxin and glutathione antioxidant systems in Nrf2-mediated lung tumor initiation and progression. Redox Biol. 83, 103653 (2025).
Neuss, T. et al. Metabolic reprogramming is an initial step in pancreatic carcinogenesis that can be targeted to inhibit acinar-to-ductal metaplasia. Cancer Res. 84, 2297–2312 (2024).
TeSlaa, T., Ralser, M., Fan, J. & Rabinowitz, J. D. The pentose phosphate pathway in health and disease. Nat. Metab. 5, 1275–1289 (2023).
Beutler, E. Glucose-6-phosphate dehydrogenase deficiency: a historical perspective. Blood 111, 16–24 (2008).
Mak, G.K. & Shah, M. Glucose-6-phosphate dehydrogenase deficiency. StatPearls https://www.ncbi.nlm.nih.gov/books/NBK470315 (2026).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Santana-Codina, N. et al. Oncogenic KRAS supports pancreatic cancer through regulation of nucleotide synthesis. Nat. Commun. 9, 4945 (2018).
Nelson, B. S. et al. Tissue of origin dictates GOT1 dependence and confers synthetic lethality to radiotherapy. Cancer Metab. 8, 1 (2020).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Chio, I. I. C. et al. NRF2 promotes tumor maintenance by modulating mRNA translation in pancreatic cancer. Cell 166, 963–976 (2016).
Todoric, J. et al. Stress-activated NRF2–MDM2 cascade controls neoplastic progression in pancreas. Cancer Cell 32, 824–839.e8 (2017).
Lyssiotis, C. Dataset of gene counts and RPKM from RNA-sequencing of KrasG12D-induced acinar-to-ductal metaplasia in primary mouse pancreas cells [Data set]. Deep Blue Data https://doi.org/10.7302/nd65-zg69 (2025).
Pretsch, W., Charles, D. J. & Merkle, S. X-linked glucose-6-phosphate dehydrogenase deficiency in Mus musculus. Biochem. Genet. 26, 89–103 (1988).
Nicol, C. J., Zielenski, J., Tsui, L. C. & Wells, P. G. An embryoprotective role for glucose-6-phosphate dehydrogenase in developmental oxidative stress and chemical teratogenesis. FASEB J. 14, 111–127 (2000).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Alektiar, J. M. et al. Malic enzyme 1 knockout has no deleterious phenotype and is favored in the male germline under standard laboratory conditions. PLoS ONE 19, e0303577 (2024).
Zhang, A. M. Y. et al. Endogenous hyperinsulinemia contributes to pancreatic cancer development. Cell Metab. 30, 403–404 (2019).
Zhang, A. M. Y. et al. Hyperinsulinemia acts via acinar insulin receptors to initiate pancreatic cancer by increasing digestive enzyme production and inflammation. Cell Metab. 35, 2119–2135.e5 (2023).
Zhang, Z. et al. High glucose inhibits glucose-6-phosphate dehydrogenase, leading to increased oxidative stress and beta-cell apoptosis. FASEB J. 24, 1497–1505 (2010).
Szmola, R. et al. Chymotrypsin C is a co-activator of human pancreatic procarboxypeptidases A1 and A2. J. Biol. Chem. 286, 1819–1827 (2011).
Jiang, J. et al. Transcriptional profile of human pancreatic acinar ductal metaplasia. Gastro Hep Adv. 2, 532–543 (2023).
Means, A. L. et al. Pancreatic epithelial plasticity mediated by acinar cell transdifferentiation and generation of nestin-positive intermediates. Development 132, 3767–3776 (2005).
Doppler, H. R., Liou, G. Y. & Storz, P. Generation of hydrogen peroxide and downstream protein kinase D1 signaling is a common feature of inducers of pancreatic acinar-to-ductal metaplasia. Antioxidants (Basel) 11, 137 (2022).
Durand, N. & Storz, P. Targeting reactive oxygen species in development and progression of pancreatic cancer. Expert Rev. Anticancer Ther. 17, 19–31 (2017).
Fleming Martinez, A. K. et al. Downregulation of Sod2 increases atypical flat lesions and dysplasia to advance pancreatic ductal adenocarcinoma. Mol. Cancer 24, 300 (2025).
Al Saati, T. et al. Oxidative stress induced by inactivation of TP53INP1 cooperates with KrasG12D to initiate and promote pancreatic carcinogenesis in the murine pancreas. Am. J. Pathol. 182, 1996–2004 (2013).
Hennequart, M. et al. ALDH1L2 regulates reactive oxygen species and acinar-to-ductal metaplasia in the pancreas. Nat. Metab. https://doi.org/10.1038/s42255-026-01456-5 (2026).
Albanes, D. et al. α-Tocopherol and β-carotene supplements and lung cancer incidence in the Alpha-Tocopherol, Beta-Carotene Cancer Prevention Study: effects of base-line characteristics and study compliance. J. Natl Cancer Inst. 88, 1560–1570 (1996).
Goodman, G. E. et al. The Beta-Carotene and Retinol Efficacy Trial: incidence of lung cancer and cardiovascular disease mortality during 6-year follow-up after stopping β-carotene and retinol supplements. J. Natl Cancer Inst. 96, 1743–1750 (2004).
Kopinke, D. et al. Ongoing Notch signaling maintains phenotypic fidelity in the adult exocrine pancreas. Dev. Biol. 362, 57–64 (2012).
Stincone, A. et al. The return of metabolism: biochemistry and physiology of the pentose phosphate pathway. Biol. Rev. Camb. Philos. Soc. 90, 927–963 (2015).
Hingorani, S. R. et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 4, 437–450 (2003).
Chan, K., Lu, R., Chang, J. C. & Kan, Y. W. NRF2, a member of the NFE2 family of transcription factors, is not essential for murine erythropoiesis, growth, and development. Proc. Natl Acad. Sci. USA 93, 13943–13948 (1996).
Watanabe, T. et al. A novel model of continuous depletion of glutathione in mice treated with L-buthionine (S,R)-sulfoximine. J. Toxicol. Sci. 28, 455–469 (2003).
Fleming Martinez, A. K. & Storz, P. Mimicking and manipulating pancreatic acinar-to-ductal metaplasia in 3-dimensional cell culture. J. Vis. Exp. 144, e59096 (2019).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Korotkevich, G et al. Fast gene set enrichment analysis. Preprint at https://doi.org/10.1101/060012 (2021).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Lee, H. J., Kremer, D. M., Sajjakulnukit, P., Zhang, L. & Lyssiotis, C. A. A large-scale analysis of targeted metabolomics data from heterogeneous biological samples provides insights into metabolite dynamics. Metabolomics 15, 103 (2019).
Kerk, S. A. et al. Metabolic requirement for GOT2 in pancreatic cancer depends on environmental context. Elife 11, e73245 (2022).
Acknowledgements
We thank the members of the Lyssiotis, Crawford and Shah laboratories and the Rogel and Blondy Center for Pancreatic Cancer at the Rogel Cancer Center for their valuable comments and input. We would also like to thank C. Strayhorn, M. Mattea and D. Long for providing histopathology support. This study was funded by T32HD007505 (M.D.R.), F32CA275283 (M.D.R.), K99GM159354 (M.D.R.), T32DK094775 (B.S.N.), T32CA009676 (B.S.N.), the Sky Foundation (M.T.R.), R37CA283575 (C.J.H.), the Tower Cancer Research Foundation Career Development Award (C.J.H.), P30CA062203 (C.J.H.), F32CA247492 (M.S.), the Department of Molecular & Integrative Physiology Summer Undergraduate Research Fellowship (B.L.L.), T32CA009676 (M.D.P., K.B. and P.S.), T32CA140044 (M.D.P. and P.S.), CA271510 (F.B.), the Association of Academic Surgery (F.B.), the Association of VA Surgeons (F.B.), the American Surgical Association Foundation (F.B.), R01CA268426 (T.L.F.), R01DK128102 (T.L.F.), U01CA274154 (T.L.F.), VA BLR&D Merit Award 5I01BX005777 (T.L.F.), U54CA274371 (M.P.), R01CA271510 (M.P.), R01CA275182 (M.P.), R01CA268426 (M.P.), R01ES028802 (J.A.C.), R21CA273646 (J.A.C.) and P30ES017885 (J.A.C.). Support was also provided by National Institutes of Health grants R01CA148828 (Y.M.S.), R01CA245546 (Y.M.S.), R01DK095201 (Y.M.S.), R01CA247516 (H.C.C.), U01CA224145 (H.C.C.) and U01CA274154 (H.C.C.), as well as the Sky Foundation (H.C.C.), R37CA237421 (C.A.L.), U54CA274371 (C.A.L.), U01CA274154 (C.A.L.), the UMCCC Core Grant P30CA046592 (C.A.L.), the DOD Idea Development Award PA240250 (C.A.L.), R01CA248160 (C.A.L.), the Stephenson Global Pancreatic Cancer Research Institute (C.A.L.), the Emerald Foundation (C.A.L.), the Rogel and Blondy Center for Pancreatic Cancer (C.A.L.), the Pancreatic Cancer Action Network/AACR Pathway to Leadership award 13-70-25-LYSS (C.A.L.), the V Foundation for Cancer Research V2016-009 (C.A.L.), the Sidney Kimmel Foundation for Cancer Research SKF-16-005 (C.A.L.) and the AACR NextGen Grant 17-20-01-LYSS (C.A.L.). We acknowledge use of the Tissue and Molecular Pathology Shared Resource, supported by P30CA046592. The funders had no role in study design, data collection, data analysis, decision to publish or preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
M.D.R., B.N.S., Y.M.S., H.C.C. and C.A.L. conceptualized the study. M.D.R., Y.M.S. and C.A.L. wrote the manuscript. M.D.R. B.S.N., M.T.R., B.L.L., Y.M.S. and C.A.L. revised the manuscript. C.J.H., J.M.A., L.S., G.C., F.B., T.L.F., M.P., J.A.C. and H.C.C. provided feedback on the manuscript. M.D.R., B.S.N., M.T.R., C.J.H., M.S., J.M.A., B.L.L., L.S., M.D.P., K.B., A.W., H.S.H., P.S. and L.Z. performed and analysed the experiments. W.Y. provided expert pathological evaluation. J.A.C. performed RNA-seq analysis and interpretation. G.C. provided the G6PD-deficient mice. F.B., T.L.F., M.P., Y.M.S., H.C.C. and C.A.L. provided conceptual input on the study, helped interpret data and provided resources. Y.M.S., H.C.C. and C.A.L. provided funding. All authors reviewed and approved the study.
Corresponding authors
Ethics declarations
Competing interests
In the past 3 years, C.A.L. has consulted for Astellas Pharmaceuticals, Odyssey Therapeutics, Third Rock Ventures and T-Knife Therapeutics and is an inventor on patents pertaining to Kras-regulated metabolic pathways, redox control pathways in pancreatic cancer and targeting the GOT1–ME1 pathway as a therapeutic approach (US Patent No. 2015126580-A1, 05/07/2015; US Patent No. 20190136238, 05/09/2019; International Patent No. WO2013177426-A2, 04/23/2015). G.C. was the scientific founder of Decibel Therapeutics, had equity interest in the company and received compensation for consulting. The company was not involved in this study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Jean Nakhle, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Continued summary of RNA-sequencing data.
a. Multidimensional scaling plot of RNA-sequencing data from Vehicle (Control), Day 1, Day 2, and Day 3 cells treated with 4-OHT. Cells from each mouse are identified by color. b. Volcano plot of genes expressed in 4-OHT-treated Day 1 cells compared to Vehicle. There are no significantly differentially expressed genes. c. Heatmap of acinar and ductal gene expression. Scale represents log2FC in expression relative to Vehicle.
Extended Data Fig. 2 Continued summary of metabolomics data.
a. Principal component analysis (PCA) plot of intracellular metabolites collected from Vehicle, or 4-OHT-treated Day 1, Day 2, and Day 3 cells generated using MetaboAnalyst. Cells from each mouse are identified by color. b. Volcano plots of differentially abundant intracellular metabolites when comparing Day 1 (4-OHT treated) to Vehicle. c. PCA plot of extracellular metabolites collected from Vehicle, Day 1, Day 2, or Day 3 4-OHT-treated cells generated using MetaboAnalyst. Cells from each mouse are identified by color. d. Volcano plots of differentially abundant extracellular metabolites when comparing Day 1 (4-OHT treated) to Vehicle. e. Volcano plots of differentially abundant extracellular metabolites when comparing Day 2 (4-OHT treated) to Vehicle. f. Volcano plots of differentially abundant extracellular metabolites when comparing Day 3 (4-OHT treated) to Vehicle. P-values for b, d, e, and f were calculated using a Student’s t-test (unpaired, two-tailed). P-value < 0.1 was considered significant, log2FC > 0.5 was considered increased, and log2FC < -0.5 was considered decreased. Significantly increased metabolites are labeled in red; decreased metabolites are labeled in blue.
Extended Data Fig. 3 Heatmaps of leading edge genes and volcano plots of all the genes detected in respective pathways.
a. Heatmap of the leading edge genes in the Oxidative Phosphorylation (OxPhos) KEGG pathway. Scale represents log2FC relative to Vehicle. b. Violin plot of the 106 detected genes in the OxPhos KEGG pathway. Y-axis represents log2FC relative to Vehicle. c. Heatmap of the leading edge genes in the tricarboxylic acid (TCA) cycle KEGG pathway. Scale represents log2FC relative to Vehicle. d. Violin plot of the 28 detected genes in the TCA cycle KEGG pathway. Y-axis represents log2FC relative to Vehicle. e. Heatmap of the leading edge genes in the Pentose Phosphate Pathway (PPP) KEGG pathway. Scale represents log2FC relative to Vehicle. f. Violin plot of the 27 detected genes in the PPP KEGG pathway. Y-axis represents log2FC relative to Vehicle. g. Violin plot of the 49 detected genes in the Glycolysis & Gluconeogenesis KEGG pathway. Y-axis represents log2FC relative to Vehicle. h. Heatmap of the edge genes in the Glycolysis & Gluconeogenesis KEGG pathway. Scale represents log2FC relative to Vehicle. i. Heatmap of the leading edge genes in the Glutathione Metabolism KEGG pathway. Scale represents log2FC relative to Vehicle. j. Violin plot of the 57 detected genes in the Glutathione Metabolism KEGG pathway. Y-axis represents log2FC relative to Vehicle.
Extended Data Fig. 4 Other pathways of interest identified from RNAseq & Metabolomics.
a. Heatmap of the leading edge genes in the Purine Metabolism KEGG pathway. Scale represents log2FC relative to Vehicle. b. Violin plot of the 40 leading edge genes in the Purine Metabolism KEGG pathway. Y-axis represents log2FC relative to Vehicle. c. Violin plot of the 93 detected genes in the Purine Metabolism KEGG pathway. Y-axis represents log2FC relative to Vehicle. d. Heatmap of detected purine-related metabolites. Scale represents log2FC of median normalized abundance relative to Vehicle. e. Heatmap of detected pyrimidine-related metabolites. Scale represents log2FC of median normalized abundance relative to Vehicle. f. Heatmap of genes that encode core enzymes in pyrimidine metabolism. Scale represents log2FC relative to Vehicle. g. Violin plot of the 38 detected genes in the Fatty Acid Degradation KEGG pathway. Y-axis represents log2FC relative to Vehicle. h. Heatmap of detected amino acid abundances. Scale represents log2FC of median normalized abundance relative to Vehicle. i. Heatmap of genes that encode core enzymes in branched-chain amino acid (BCAA) metabolism. Scale represents log2FC relative to Vehicle.
Extended Data Fig. 5 Enrichr transcription factor enrichment analysis.
Transcription Factor Enrichment Analyses generated using Enrichr from ENCODE and ChEA Consensus Transcription Factors. a. Analysis from the top 500 upregulated genes at Day 2. b. Analysis from the top 500 downregulated genes at Day 2. c. Analysis from the top 500 upregulated genes at Day 3. d. Analysis from the top 500 downregulated genes at Day 3. Enrichr was used to identify differentially enriched pathways and calculate p-values with Fisher’s exact test.
Extended Data Fig. 6 RNA analysis using additional ADM culture approaches.
a. Scheme of the experimental model. Pancreata were dissected from Wildtype mice and acinar clusters were isolated for primary, ex vivo culture. Cells were collected after dissociation (Day 0) or grown free-floating in media and treated with TGFα (50 ng/mL), with or without 4-hydroxytamoxifen (4-OHT; 2 μM) for 3 days. Relative expression of acinar gene, Amy (b.) and ductal gene, Spink1 (c.) using real-time quantitative PCR. Bars represent the mean of n = 3 biological replicates for Day 0, n = 2 biological replicates for Day 3 and Day 3 + 4-OHT. d. Scheme of the experimental model. Pancreata were dissected from LSL-KrasG12D mice (n = 3) and acinar clusters were isolated for primary, ex vivo culture. Acinar clusters were grown free-floating in media and were treated with adenovirus expressing GFP (ad-GFP) or adenovirus expressing Cre recombinase (ad-Cre) to induce mutant Kras. Cells were collected after 1, 2, or 3 days in culture. e. Heatmap of acinar and ductal gene expression. Scale represents log2FC in expression relative to ad-GFP. f. Heatmap showing log2FC of key NRF2-target genes, as referenced from Fig. 1n. Individual datapoints for log2FC of gene expression of G6pdx (g.) and Me1 (h.) from ADM cultures from each mouse (n = 3) after ad-Cre for 1, 2, or 3 days (relative to ad-GFP). Bars represent the mean with standard deviation. P-values were calculated using ordinary one-way ANOVA with Tukey’s multiple comparisons test.
Extended Data Fig. 7 Additional description of and analyses in KC;G6pdwt and KC;G6pdmut mice.
a. Mice with mutant G6pd, that mimics human G6PD-deficiency, were bred into the KC (LSL-KrasG12D; Ptf1aCre) line. Cre was always maintained in female breeders. Experiments used both male and female G6pd mutant mice, where females were homozygous for mutant G6pd (G6pdmut/mut) and males were hemizygous for mutant G6pd (G6pdmut/y), as G6pd is an X-linked gene. G6pd wildtype mice used in experiments were obtained in the same colony and age matched littermates of G6pd mutant mice when possible. Female G6pd wildtype mice have two wildtype copies of G6pd (G6pdwt/wt) and males have one wildtype copy (their only copy) of G6pd (G6pdwt/y). In the schematics and labelling, “y” in male mice refers to the y chromosome, which does not contain a copy of G6pd. b. Schematic of 14C-labeling experiment. [1-14C]glucose (blue) can be used in both the oxidative pentose phosphate pathway (ox PPP) and the TCA cycle. [6-14C]glucose (red) can be used in the TCA cycle. Released CO2 represented as clouds; G6PD mutation represented as star. c. Graph depicting the relative amount of [14C]-labeled CO2 derived from glucose tracing in G6pdmut mice relative to G6pdwt mice (n = 2 biological replicates). Blue bar shows the relative [14C]-labeled CO2 derived from [1-14C]glucose, which is generated from either the oxidative pentose phosphate pathway (oxPPP) or the TCA cycle. Red bar shows the relative amount of [14C]-labeled CO2 derived from [6-14C]glucose, which is only generated from the TCA cycle. Grey bar shows the ratio of [14C]-labeled CO2 derived from [1-14C]O2 to [6-14C]O2 representing flux from the oxPPP. Bars represent the biological replicates’ mean. d. Hematoxylin & eosin (H&E) staining and immunostaining for amylase (AMY; acinar cells), cytokeratin 19 (CK19; ductal, metaplastic, neoplastic cells), and KI67 (proliferation) in pancreata from 8-week-old KC;G6pdwt and KC;G6pdmut mice. Scale bar = 50μm. e. Percent of Amylase+ area in pancreas as quantified from male (closed circle) and female (open circle) mice in KC;G6pdwt (grey bar) and KC;G6pdmut (blue bar) mice at 8 weeks. n = 4 mice for each genotype. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). f. Percent of CK19+ cells and g. Ki67+ cells in the pancreas as quantified from male (closed circle) and female (open circle) KC;G6pdwt (grey bar) and KC;G6pdmut (blue bar) mice at 8 weeks. n = 5 mice for each genotype. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). h. Immunostaining for Amylase (AMY), in 8-week and 16-week-old KC;G6pdwt and KC;G6pdmut pancreas. Alcian blue (PanIN-produced mucin) & nuclear fast red counterstain in pancreas of 26-week-old KC;G6pdwt and KC;G6pdmut mice. Scale bar = 500μm. i. Pancreas weight (PW) to body weight (BW) ratios in 8-, 16-, and 26-week-old KC;G6pdwt and KC;G6pdmut mice. Bars represent the mean with standard deviation. Ratios are not significantly different, as calculated using a Student’s t-test (unpaired, two-tailed) for each age. Sample sizes for each age are as follows: 8 week, n = 11 KC;G6pdwt and n = 8 KC;G6pdmut; 16 week, n = 9 KC;G6pdwt and n = 8 KC;G6pdmut; 26 week, n = 8 KC;G6pdwt and n = 8 KC;G6pdmut. j. Pathological grading of 26-week-old KC;G6pdwt and KC;G6pdmut pancreas tissues representing the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 5 KC;G6pdwt mice; n = 7 KC;G6pdmut mice. Bars represent the mean with standard deviation. P-values were calculated using a two-way ANOVA with Tukey’s multiple comparisons test. k. Pathological grading of 1-year-old KC;G6pdwt and KC;G6pdmut pancreas tissues representing the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 5 mice per genotype. Bars represent the mean with standard deviation. P-values were calculated using a two-way ANOVA with Tukey’s multiple comparisons test.
Extended Data Fig. 8 Additional description of and analyses in KC;Me1+/+ and KC;Me1flox/flox mice.
a. Mice with Me1flox/flox alleles were bred into the KC (LSL-KrasG12D; Ptf1aCre) line. Cre was always maintained in female breeders. Experiments used both male and female mice that were homozygous for the Me1flox/flox allele and mice that were Me1+/+ (wildtype). b. Immunostaining for ME1 in 16-week-old KC;Me1+/+ and KC;Me1flox/flox pancreas. Scale bar = 50μm. c. Hematoxylin & eosin (H&E) staining and immunostaining for amylase (AMY; acinar cells), cytokeratin 19 (CK19; ductal, metaplastic, neoplastic cells), and Ki67 (proliferation) in pancreas of 8-week-old KC;Me1+/+ and KC;Me1flox/flox mice. Scale bar = 50μm. d. Percent of Amylase+ area in pancreas as quantified from male (closed circle) and female (open circle) mice in KC;Me1+/+ (grey bar) and KC;Me1flox/flox (blue bar) mice at 8 weeks. n = 4 mice for each genotype. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). e. Percent of CK19+ cells and f. Ki67+ cells in the pancreas as quantified from male (closed circle) and female (open circle) KC;Me1+/+ (grey bar) and KC;Me1flox/flox (blue bar) mice at 8 weeks. In CK19 plot: n = 5 KC;Me1+/+ and n = 6 KC;Me1flox/flox mice. In Ki67 plot: n = 6 mice for each genotype. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). g. Immunostaining for Amylase (AMY), in 8-week and 16-week-old KC;Me1+/+ and KC;Me1flox/flox pancreas. Alcian blue (PanIN-produced mucin) & nuclear fast red counterstain in pancreas of 26-week- KC;Me1+/+ and KC;Me1flox/flox mice. Scale bar = 500μm. h. Pancreas weight (PW) to body weight (BW) ratios in 8-, 16-, and 26-week-old KC;Me1+/+ and KC;Me1flox/flox mice. Bars represent the mean with standard deviation. Ratios are not significantly different as calculated using a Student’s t-test (unpaired, two-tailed) for each age. Sample sizes for each age are as follows: 8 week, n = 6 KC;Me1+/+ and n = 8 KC;Me1flox/flox; 16 week, n = 6 KC;Me1+/+ and n = 8 KC;Me1flox/flox; 26 week, n = 5 KC;Me1+/+ and n = 7 KC;Me1flox/flox. i. Pathological grading of 26-week-old KC;Me1+/+ and KC;Me1flox/flox pancreas tissues representing the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 4 mice per genotype. Bars represent the mean with standard deviation. P-values were calculated using a two-way ANOVA with Tukey’s multiple comparisons test. j. Pathological grading of 1-year-old KC;Me1+/+ pancreas tissues representing the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 4 mice. Bars represent the mean with standard deviation.
Extended Data Fig. 9 ROS markers in pancreas tissue and acinar cell cultures with antioxidant treatment.
a. Quantification of protein levels from Fig. 4a with samples normalized to Vinculin and relative to Wildtype expression. n = 2 per genotype. The experiment was repeated twice. MDA = Malondialdehyde; HO-PRDX3 = Hyperoxidized Peroxiredoxin 3; 4-HNE = 4-hydroxynonenal. b. Immunostaining for MDA in pancreas of 16-week-old KC;G6pdwt, KC;G6pdmut, KC;Me1+/+, and KC;Me1flox/flox mice, presented for acinar and lesion regions. Scale bar = 50μm. c-d. Quantification of mean optical density from MDA immunostaining in mouse pancreas separated by expression in acinar or lesion regions of the tissue. Each datapoint represents one mouse. n = 3 mice per genotype. Bars represent the mean with standard deviation. P-values were calculated using repeated measures one-way ANOVA with Tukey’s multiple comparisons test. e. Immunostaining for 4-HNE in pancreas of 16-week-old KC;G6pdwt and KC;G6pdmut mice Scale bar = 50μm. f. Quantification of mean optical density from 4-HNE immunostaining. Each datapoint represents one mouse. n = 3 mice per genotype. Bars represent the mean with standard deviation. P-values were calculated using repeated measures Student’s t-test (unpaired, two-tailed). g. Brightfield images of primary, ex vivo acinar cells from KC;G6pdwt and KC;G6pdmut mice grown in Matrigel for 0, 2, and 3 days. Cells were treated with vehicle (water) or glutathione ethyl ester (GSHee; 1 mM). Scale bars = 25μm. h-i. Quantification of acinar cells undergoing ADM in primary cultures at Day 2 (h.) and Day 3 (i). Grey bars are KC;G6pdwt cells treated with vehicle (light) or GSHee (dark), blue bars are KC;G6pdmut cells treated with vehicle (light) or GSHee (dark). Each datapoint represents one mouse. n = 3 mice per genotype. Bars represent the mean with standard deviation. P-values were calculated using ordinary one-way ANOVA with Tukey’s multiple comparisons test. j-k. Violin plots showing quantification of Diameter (µm) in ADM primary cultures at Day 2 (J.) and Day 3 (k). Grey plots are KC;G6pdwt cells treated with vehicle (light) or GSHee (dark), blue plots are KC;G6pdmut cells treated with vehicle (light) or GSHee (dark). Plots combine 3 mice per genotype. Lines inside of the plots represent the quartiles and median. P-values were calculated using ordinary one-way ANOVA with Tukey’s multiple comparisons test. Exact p-values are as follows: KC;G6pdwt vs. KC;G6pdmut = 1.54×10−13; KC;G6pdmut vs. KC;G6pdmut+GSHee = 2.54×10−7; KC;G6pdwt+GSHee vs. KC;G6pdmut+GSHee = 6.96×10−7. l. Pancreas to body weight ratios from mice of the indicated genotypes after receiving control drinking water or drinking water supplemented with NAC for 6 weeks. n = 4 KC;G6pdwt; n = 4 KC;G6pdwt + NAC; n = 6 KC;G6pdmut; n = 6 KC;G6pdmut + NAC. Bars represent the mean with standard deviation. Ratios are not significantly different, as calculated by using ordinary one-way ANOVA with Tukey’s multiple comparisons test. m. Pathological grading of pancreas tissue of the indicated genotypes after receiving control drinking water or drinking water supplemented with NAC for 6 weeks. Datapoints represent the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 4 mice for each treatment. Bars represent the mean with standard deviation.
Extended Data Fig. 10 Representative cultures from donor pancreas and additional analyses in BSO-treatment experiments.
a. Transcripts per million (TPM)-normalized expression of G6PD and ME1 from RNA-sequencing of human acinar cells treated with TGFα for 6 days (n = 11). Dataset from40 and accessed through NCBI GEO. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). Exact p-value for G6PD = 1.04×10−8; for ME1 = 1.06×10−7. b. Bright field images of ex vivo acinar cell cultures isolated from human pancreata (obtained from 4 independent tissue donors) at Day 0 and Day 6. Cells were treated with Vehicle (DMSO), TGFα (50 ng/mL), or buthionine sulfoximine (BSO; 100 μM). Scale bar = 100μm. c. Hematoxylin & eosin (H&E)-stained human donor pancreas. Tissue was adjacent to the region selected for acinar cell isolation and culture. Scale bar = 100μm. d. Violin plots showing quantification of Diameter (μm) in ADM primary cultures at Day 6. Grey plots are vehicle-treated cells, blue plots are TGFα-treated cells, green plots are BSO-treated. Plots combine counts from cells isolated from all 4 donors. Lines inside of the plots represent quartiles and median. P-values were calculated using two-way ANOVA with Dunnett’s multiple comparisons test. Metabolite peak areas of GSH (e.) and GSSG (f.) from pancreas tissue as determined by mass spectrometry. g. Ratio of GSH to GSSG peak areas. Data in e-g. represent male and female KC mice that were administered control or buthionine sulfoximine (BSO; 20 mM) water for 5 weeks. n = 4 mice per treatment. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). h. Pancreas to body weight ratios from mice that received control or BSO-containing drinking water for 5 weeks. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed). i. Pathological grading of pancreas tissue. Datapoints represent the % of total tissue area with acinar cells, ADM, and PanIN lesions. n = 4 mice for each treatment. Bars represent the mean with standard deviation. P-values were calculated using a two-way ANOVA with Tukey’s multiple comparisons test. j. Immunostaining for cytokeratin 19 (CK19; ductal, metaplastic, neoplastic cells) in pancreata from 5-6 week old KC mice administered control or BSO-containing water for 5 weeks. Scale bar = 50μm. k. Percent of CK19+ cells in the pancreas as quantified from KC mice administered control or BSO-containing water for 5 weeks. n = 4 mice for each genotype. Bars represent the mean with standard deviation. P-values were calculated using a Student’s t-test (unpaired, two-tailed).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7; Source Data Blot for Supplementary Fig. 5j
Supplementary Tables 1–3 (download XLSX )
Supplementary Tables 1–3 referenced in the Methods section
Supplementary Tables 4–11 (download XLSX )
Metabolomics datasets
Supplementary Data 1–7 (download XLSX )
Statistical Source Data for Supplementary Figs. 1–7
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data
Source Data Fig. 2 (download XLSX )
Statistical Source Data
Source Data Fig. 3 (download XLSX )
Statistical Source Data
Source Data Fig. 4 (download XLSX )
Statistical Source Data
Source Data Fig. 4 (download PDF )
Unprocessed western blots
Source Data Extended Data Fig./Table 1 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 2 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 3 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 4 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 5 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 6 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 7 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 8 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 9 (download XLSX )
Statistical Source Data
Source Data Extended Data Fig./Table 10 (download XLSX )
Statistical Source Data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Radyk, M.D., Nelson, B.S., Ruckert, M.T. et al. NADPH-producing enzymes restrict the formation of pancreatic precancerous lesions. Nat Metab 8, 824–839 (2026). https://doi.org/10.1038/s42255-026-01496-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-026-01496-x
This article is cited by
-
Good things come in twos
Nature Metabolism (2026)