Abstract
Cancer immunotherapies have revolutionized cancer treatment, yet many patients fail to respond. Activating innate immunity offers a promising approach to enhance therapeutic efficacy, but the signaling kinases directly regulating this process to boost antitumor responses remain elusive. Here we conduct an in vivo kinome CRISPR screen and identify CDK10 as a key suppressor of tumor immune surveillance. Mechanistically, CDK10 phosphorylates DNMT1 and RAP80 to reduce the accumulation of double-stranded RNA and R-loops, which alleviates the activation of innate immune pathways mediated by MDA5 and cGAS. Kinase inhibitor screens identify NVP-AST487 and ponatinib as selective CDK10 inhibitors. Both genetic and pharmacological inhibition of CDK10 activates MDA5 and cGAS pathways, fostering an immunoactive tumor microenvironment that enhances cancer immunotherapy in multiple mouse tumor models. Clinically, low CDK10 expression in tumors correlates with better immunotherapy responses. These findings establish CDK10 as a pivotal modulator of tumor immunity and a potential therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The scRNA-seq, CRISPR screening and RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession codes GSE287052, GSE287053 and GSE287054. The TMT, IP–MS proteomics and TMT phosphoproteomics data have been deposited in ProteomeXchange under primary accession code PXD059695. Data in Fig. 1e were derived from https://cancergenome.nih.gov/. Data in Figs. 1f and 6s–v and Extended Data Fig. 8p–r were derived from http://tide.dfci.harvard.edu and https://cide.ccr.cancer.gov/. Data in Fig. 6n were derived from https://cistrome.shinyapps.io/timer/. The remaining data are available within the Article, Supplementary Information and Source Data files, or available from the corresponding author on request. Source data are provided with this paper.
References
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv324 (2016).
Sharma, P. et al. Immune checkpoint therapy-current perspectives and future directions. Cell 186, 1652–1669 (2023).
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Carroll, S. L., Pasare, C. & Barton, G. M. Control of adaptive immunity by pattern recognition receptors. Immunity 57, 632–648 (2024).
Demaria, O. et al. Harnessing innate immunity in cancer therapy. Nature 574, 45–56 (2019).
Cao, L. L. & Kagan, J. C. Targeting innate immune pathways for cancer immunotherapy. Immunity 56, 2206–2217 (2023).
Lynch, C., Pitroda, S. P. & Weichselbaum, R. R. Radiotherapy, immunity, and immune checkpoint inhibitors. Lancet Oncol. 25, e352–e362 (2024).
Kornepati, A. V. R., Rogers, C. M., Sung, P. T. & Curiel, T. J. The complementarity of DDR, nucleic acids and anti-tumour immunity. Nature 619, 475–486 (2023).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563 e519 (2018).
Maxwell, M. B. et al. ARID1A suppresses R-loop-mediated STING-type I interferon pathway activation of anti-tumor immunity. Cell 187, 3390–3408 e3319 (2024).
Zhang, Z. D. & Zhong, B. Regulation and function of the cGAS-MITA/STING axis in health and disease. Cell Insight 1, 100001 (2022).
Jiang, X. et al. Ubiquitin-induced oligomerization of the RNA sensors RIG-I and MDA5 activates antiviral innate immune response. Immunity 36, 959–973 (2012).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Deng, J. et al. CDK4/6 inhibition augments antitumor immunity by enhancing T-cell activation. Cancer Discov. 8, 216–233 (2018).
Fassl, A., Geng, Y. & Sicinski, P. CDK4 and CDK6 kinases: from basic science to cancer therapy. Science 375, eabc1495 (2022).
Alvarez-Fernandez, M. & Malumbres, M. Mechanisms of sensitivity and resistance to CDK4/6 Inhibition. Cancer Cell 37, 514–529 (2020).
Dietrich, C. et al. INX-315, a selective CDK2 inhibitor, induces cell cycle arrest and senescence in solid tumors. Cancer Discov. 14, 446–467 (2024).
Arora, M. et al. Rapid adaptation to CDK2 inhibition exposes intrinsic cell-cycle plasticity. Cell 186, 2628–2643 e2621 (2023).
Cidado, J. et al. AZD4573 is a highly selective CDK9 inhibitor that suppresses MCL-1 and induces apoptosis in hematologic cancer cells. Clin. Cancer Res. 26, 922–934 (2020).
Parua, P. K. et al. A Cdk9-PP1 switch regulates the elongation-termination transition of RNA polymerase II. Nature 558, 460–464 (2018).
Hluchy, M. et al. CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1. Nature 609, 829–834 (2022).
Dubbury, S. J., Boutz, P. L. & Sharp, P. A. CDK12 regulates DNA repair genes by suppressing intronic polyadenylation. Nature 564, 141–145 (2018).
Iniguez, A. B. et al. EWS/FLI confers tumor cell synthetic lethality to CDK12 inhibition in Ewing sarcoma. Cancer Cell 33, 202–216 e206 (2018).
Quereda, V. et al. Therapeutic targeting of CDK12/CDK13 in triple-negative breast cancer. Cancer Cell 36, 545–558 e547 (2019).
Insco, M. L. et al. Oncogenic CDK13 mutations impede nuclear RNA surveillance. Science 380, eabn7625 (2023).
Guen, V. J. et al. CDK10/cyclin M is a protein kinase that controls ETS2 degradation and is deficient in STAR syndrome. Proc. Natl Acad. Sci. USA 110, 19525–19530 (2013).
Iorns, E. et al. Identification of CDK10 as an important determinant of resistance to endocrine therapy for breast cancer. Cancer Cell 13, 91–104 (2008).
You, Y. et al. Downregulated CDK10 expression in gastric cancer: association with tumor progression and poor prognosis. Mol. Med. Rep. 17, 6812–6818 (2018).
Li, H., You, Y. & Liu, J. Cyclin‑dependent kinase 10 prevents glioma metastasis via modulation of Snail expression. Mol. Med. Rep. 18, 1165–1170 (2018).
Weiswald, L. B. et al. Inactivation of the kinase domain of CDK10 prevents tumor growth in a preclinical model of colorectal cancer, and is accompanied by downregulation of Bcl-2. Mol. Cancer Ther. 16, 2292–2303 (2017).
Windpassinger, C. et al. CDK10 mutations in humans and mice cause severe growth retardation, spine malformations, and developmental delays. Am. J. Hum. Genet. 101, 391–403 (2017).
Guen, V. J. et al. A homozygous deleterious CDK10 mutation in a patient with agenesis of corpus callosum, retinopathy, and deafness. Am. J. Med. Genet. A 176, 92–98 (2018).
Gu, S. S. et al. Therapeutically increasing MHC-I expression potentiates immune checkpoint blockade. Cancer Discov. 11, 1524–1541 (2021).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Belnap, L. P., Cleveland, P. H., Colmerauer, M. E., Barone, R. M. & Pilch, Y. H. Immunogenicity of chemically induced murine colon cancers. Cancer Res. 39, 1174–1179 (1979).
Wang, X. et al. In vivo CRISPR screens identify the E3 ligase Cop1 as a modulator of macrophage infiltration and cancer immunotherapy target. Cell 184, 5357–5374 e5322 (2021).
Guo, E. et al. WEE1 inhibition induces anti-tumor immunity by activating ERV and the dsRNA pathway. J. Exp. Med. https://doi.org/10.1084/jem.20210789 (2022).
Bell, H. N. & Zou, W. Beyond the barrier: unraveling the mechanisms of immunotherapy resistance. Annu. Rev. Immunol. 42, 521–550 (2024).
Della Corte, C. M. & Byers, L. A. Evading the STING: LKB1 loss leads to STING silencing and immune escape in KRAS-mutant lung cancers. Cancer Discov. 9, 16–18 (2019).
Zulato, E. et al. LKB1 loss is associated with glutathione deficiency under oxidative stress and sensitivity of cancer cells to cytotoxic drugs and γ-irradiation. Biochem. Pharmacol. 156, 479–490 (2018).
Skoulidis, F. et al. CTLA4 blockade abrogates KEAP1/STK11-related resistance to PD-(L)1 inhibitors. Nature 635, 462–471 (2024).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459 e429 (2020).
Dexter, D. L. et al. Heterogeneity of cancer cells from a single human colon carcinoma. Am. J. Med. 71, 949–956 (1981).
van den Heuvel, S. & Harlow, E. Distinct roles for cyclin-dependent kinases in cell cycle control. Science 262, 2050–2054 (1993).
Robert, T. et al. Development of a CDK10/CycM in vitro kinase screening assay and identification of first small-molecule inhibitors. Front. Chem. 8, 147 (2020).
Brown, N. R., Noble, M. E., Endicott, J. A. & Johnson, L. N. The structural basis for specificity of substrate and recruitment peptides for cyclin-dependent kinases. Nat. Cell Biol. 1, 438–443 (1999).
Zhou, X. et al. Pharmacologic activation of p53 triggers viral mimicry response thereby abolishing tumor immune evasion and promoting antitumor immunity. Cancer Discov. 11, 3090–3105 (2021).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Crossley, M. P. et al. R-loop-derived cytoplasmic RNA-DNA hybrids activate an immune response. Nature 613, 187–194 (2023).
Yasuhara, T. et al. RAP80 suppresses the vulnerability of R-loops during DNA double-strand break repair. Cell Rep. 38, 110335 (2022).
Valk, E., Ord, M., Faustova, I. & Loog, M. CDK signaling via nonconventional CDK phosphorylation sites. Mol. Biol. Cell 34, pe5 (2023).
Kusubata, M. et al. cdc2 kinase phosphorylation of desmin at three serine/threonine residues in the amino-terminal head domain. Biochem. Biophys. Res. Commun. 190, 927–934 (1993).
Li, Z. et al. Synthesis and structure-activity relationships of cyclin-dependent kinase 11 inhibitors based on a diaminothiazole scaffold. Eur. J. Med. Chem. 238, 114433 (2022).
Duster, R., Ji, Y., Pan, K. T., Urlaub, H. & Geyer, M. Functional characterization of the human Cdk10/Cyclin Q complex. Open Biol. 12, 210381 (2022).
Sachs, R. E., Ginsburg, P. B. & Goldman, D. P. Encouraging new uses for old drugs. JAMA 318, 2421–2422 (2017).
Weisberg, E. et al. Antileukemic effects of the novel, mutant FLT3 inhibitor NVP-AST487: effects on PKC412-sensitive and -resistant FLT3-expressing cells. Blood 112, 5161–5170 (2008).
Kantarjian, H. M. et al. ponatinib-review of historical development, current status, and future research. Am. J. Hematol. 99, 1576–1585 (2024).
Eide, C. A. et al. Overcoming clinical BCR-ABL1 compound mutant resistance with combined ponatinib and asciminib therapy. Cancer Cell 42, 1486–1488 (2024).
Li, H. et al. USP8-governed GPX4 homeostasis orchestrates ferroptosis and cancer immunotherapy. Proc. Natl Acad. Sci. USA 121, e2315541121 (2024).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Dai, P. et al. USP2 inhibition unleashes CD47-restrained phagocytosis and enhances anti-tumor immunity. Nat. Commun. 16, 4564 (2025).
Crossley, M. P. et al. Catalytically inactive, purified RNase H1: a specific and sensitive probe for RNA-DNA hybrid imaging. J. Cell Biol. https://doi.org/10.1083/jcb.202101092 (2021).
Zhang, J. et al. Cyclin D-CDK4 kinase destabilizes PD-L1 via cullin 3-SPOP to control cancer immune surveillance. Nature 553, 91–95 (2018).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Zhao, J. et al. Immune and genomic correlates of response to anti-PD-1 immunotherapy in glioblastoma. Nat. Med. 25, 462–469 (2019).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Nathanson, T. et al. Somatic mutations and neoepitope homology in melanomas treated with CTLA-4 blockade. Cancer Immunol. Res. 5, 84–91 (2017).
Gong, L. et al. Cancer immunology data engine reveals secreted AOAH as a potential immunotherapy. Cell 188, 5062–5080 e5032 (2025).
Patil, N. S. et al. Intratumoral plasma cells predict outcomes to PD-L1 blockade in non-small cell lung cancer. Cancer Cell 40, 289–300 e284 (2022).
Acknowledgements
We thank P. Sicinski (Dana-Farber Cancer Institute) for the discussion and advice. This work was supported by grants from the National Key Research and Development Program of China (2023YFC3402100 and 2022YFC3401500 to J.Z.), the National Natural Science Foundation of China (82273062 to J.Z.; 22193073 and 92253305 to X.L.; 82504056 to C.H.; 82503353 to H.L.; 82403052 to W.X.; 22407012 to F.G.; and 82503802 to B.C.), the Fundamental Research Funds for the Central Universities (2042022dx0003 to J.Z.), Natural Science Foundation of Wuhan (2024040701010031 to J.Z.), the open Research Fund of the National Center for Protein Sciences at Peking University in Beijing (KF-202504 to J.Z.), the Open Projects of Hubei Key Laboratory of Tumor Biological Behavior (220172107 to J.Z.), Translational Medicine and Interdisciplinary Research Joint Fund of Zhongnan Hospital of Wuhan University (ZNJC202312 to J.Z.), Postdoctoral Fellowship Program of CPSF (GZC20250976 to C.H. and GZC20241276 to W.X.), China Postdoctoral Science Foundation (2022M722462 to H.L.), China Postdoctoral Science Foundation (2024M762491 to C.H.), Postdoctoral Project of Hubei Province (2004HBBHJD057 to C.H.; 2004HBBHCXA069 to W.X.), Beijing National Laboratory for Molecular Sciences (BNLMS- CXX-202106 to X.L.) and the New Cornerstone Science Foundation through the XPLORER PRIZE to X.L. We also thank the staff at the core facility of the Medical Research Institute at Wuhan University for their technical support. The funders had no role in study design, data collection, analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.Z., G.X., X.L. and F.G. conceived of and designed the study. G.X., F.G., C.H. and X.W. conducted most of the experiments, analyzed the data and prepared the figures with help from B.X., L.F., B.C., J.P., Y.S., J.S., X.X., Y.Y., P.D., H.L. and W.X. J.P. and R.X. analyzed RNA sequencing data. L.F., B.C. and C.J. helped analyze the clinical data. F.G., B.J. and X.L. helped with high-throughput kinase library screening; J.Z. and X.L. obtained funding and supervised the study. J.Z., X.L., G.X. and F.G. wrote the manuscript. H.L. and G.Q. provided the intellectual support and discussed the research. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Daniel Schramek and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CRISPR screen identifies CDK10 as a driver of cancer immune evasion.
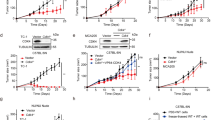
a–c, The sgRNA coverage (a), gini index (b), and pearson correlation (c) in cells and tumors from immunocompetent BALB/c WT and immunodeficient BALB/c nude mice. d–f, Cell Counting Kit-8 (CCK8) assay (d) and colony formation assay (e, f) for sgControl and sgCdk10 CT26 cells. g, h, Tumor growth curve (g) and end point tumor weight (h) of sgControl or sgCdk10 CT26 tumors in immunodeficient NSG mice. n = 6 mice per group. i–k, CCK8 assay (i) and colony formation assay (j, k) for sgControl and sgCdk10 MC38 cells. l, Immunoblot (IB) analysis of whole cell lysates (WCL) derived from sgControl and sgCdk10 B16F10 cells. m–o, CCK8 assay (m) and colony formation assay (n, o) for sgControl and sgCdk10 B16F10 cells. p–s, Tumor growth curve and end point tumor weight of sgControl and sgCdk10 B16F10 tumors in immunodeficient C57BL/6J Rag1−/− mice (p, q) or immunocompetent C57BL/6J WT mice (r, s). n = 6 mice for sgControl group and n = 7 mice for sgCdk10 group in C57BL/6J Rag1−/− mice, n = 10 mice for sgControl group and n = 12 mice for sgCdk10 group in C57BL/6J WT mice. t, IB analysis of WCL derived from CT26 cells with indicated treatments. u, v, Tumor growth curve (u) and end point tumor weight (v) of indicated CT26 tumors in immunocompetent BALB/c WT mice. n = 8 mice for sgControl group and sgCdk10 group, n = 9 mice for sgCdk10 group + CDK10 WT group and sgCdk10 group + CDK10 D181A group. Data are presented as mean ± SEM. For d, g, i, m, p, r, and u, analyzed by two-way ANOVA with Tukey’s multiple-comparisons test; for f, h, k, o, q, s, and v, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For d–f, i–k, m–o, n = 3 biologically independent samples per group, the graph is representative of three independent experiments with similar results. For l, t, three independent experiments were performed for each image with similar results.
Extended Data Fig. 2 Clustering analysis of various cell populations in sgControl and sgCdk10 CT26 tumors.
a, UMAP of total cells from sgControl and sgCdk10 CT26 tumors in immunocompetent BALB/c WT mice, colored by cluster. b–g, Feature plots displaying the expression of conventional marker genes across different cell types: Mcm6 (b), Mcm7 (c), Cd3g (d), Itgam (e), Col1a1 (f), and Hbb-bs (g). h, UMAP of lymphoid cells from sgControl and sgCdk10 CT26 tumors in immunocompetent BALB/c WT mice, colored by cluster. i–o, Feature plots showing the expression of lymphoid-specific marker genes across different cell types: Tcf7 (i), Cd8a (j), Ncr1 (k), Foxp3 (l), Cd79a (m), Sirpa (n) and Cd40 (o). p, The representative fluorescence-activated cell sorting (FACS) gating strategy for analyzing tumor-infiltrating immune cells was designed to comprehensively assess the leukocyte population. Gating commenced with two sequential steps to exclude doublets, followed by the selection of live cells. For immune cell analysis, a size gate and subsequent CD45 gating were applied to identify the following populations: CD8+ T cells (CD45+CD3+CD8+, as shown in Figs. 2g, q, 6d, 8e,m and Extended Data Fig. 2w,x), CD4+ T cells (CD45+CD3+CD4+, as shown in Extended Data Fig. 2q, u, v), NK cells (CD45+ CD3−NK1.1+, as shown in Fig. 2k,t), MDSC (CD45+CD11b+Gr1+, as shown in Extended Data Fig. 3n), and DCs (CD45+CD11b−CD11c+MHCⅡ+, as shown in Extended Data Fig. 3o). CD8⁺ T cells and CD4+ T cells were further assessed for frequency of IFNγ+, TNF+, and GZMB+, as shown in Figs. 2h–j, 6e–g, 8f–h, n–p and Extended Data Fig. 2r–t. NK cells, natural killer cells; MDSC, Myeloid-derived suppressor cells; DCs, Dendritic cells. q, The number of CD4+ T cells normalized to tumor weight per gram in sgControl and sgCdk10 CT26 tumors. n = 5 mice/group. r–t, The percentage of IFNγ+ (r), TNF+ (s), and GZMB+ (t) in CD4+ T cells from sgControl and sgCdk10 CT26 tumors. n = 5 mice/group. IFNγ+/CD4+ (%), TNF+/CD4+ (%), GZMB+/CD4+ (%), represent the percentages of IFNγ+, TNF+, and GZMB+ cells among total CD4+ T cells within the tumor tissue, respectively. u, v, The percentage of CD4+ T cells in CD3+ T cells (u) and the number of CD4+ T cells (v) in draining lymph nodes (dLNs) of sgControl and sgCdk10 CT26 tumors. n = 5 mice/group. w, x, The percentage of CD8+ T cells in CD3+ T cells (w) and the number of CD8+ T cells (x) in dLNs of sgControl and sgCdk10 CT26 tumors. n = 5 mice/group. CD4+/CD3+ (%) represent the percentages of CD4+ cells among total CD3+ T cells. For a–o, n = 2 biologically independent samples per group, with each sample comprising a pool of three tumors. For q–x, data are presented as mean ± SEM, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test.
Extended Data Fig. 3 Analysis of the tumor-infiltrating myeloid cells from sgControl and sgCdk10 CT26 tumors.
a, UMAP of myeloid cells from sgControl and sgCdk10 CT26 tumors in immunocompetent BALB/c WT mice, colored by cluster. b, Dot plots showing expression patterns of conventional myeloid cell marker genes across different myeloid cell types in subcutaneous sgControl and sgCdk10 CT26 tumors from immunocompetent BALB/c WT mice. TAM, Tumor-associated macrophage. c–h, Feature plots displaying the expression of conventional myeloid marker genes across different cell types: C1qa (c), Spp1 (d), Plac8 (e), Vcan (f), H2-Aa (g), and S100a8 (h). i, j, UMAP of seven distinct myeloid cell types (i) and the percentage of each myeloid cell type (j) in sgControl and sgCdk10 CT26 tumors from immunocompetent BALB/c mice. k–m, Representative IHC images (k) and quantification (l, m) of C1qc and Spp1 from sgControl and sgCdk10 CT26 tumors in immunocompetent BALB/c WT mice. Scale bar, 50 μm. n = 5 mice/group. n, o, Flow cytometry analysis showing the percentage of MDSC+ (n) and DC+ (o) cells in CD45+ cells from sgControl and sgCdk10 CT26 tumors in immunocompetent BALB/c mice. n = 5 mice/group. p, Representative multiplex IHC (mIHC) images showing tumor cells (GFP, green), macrophages (F4/80, red), and nuclei (DAPI, blue) from GFP-overexpressing sgControl and sgCdk10 CT26 tumors. Yellow signals (indicated by arrows), representing the colocalization of GFP (green) and F4/80 (red), indicate macrophage-mediated phagocytosis of tumor cells. Scale bar: 50 μm. q, Quantification of yellow colocalized cells was performed in high-power fields. Each data point represents the average number of double-positive cells across three high-power fields. n = 4 mice/group. For a–j, n = 2 biologically independent samples per group, with each sample comprising a pool of three tumors. Data are presented as mean ± SEM. For l, m, and q, analyzed by two-tailed unpaired t-test, for n, o, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test.
Extended Data Fig. 4 Loss of Cdk10 triggers innate immune activation.
a, b, Heatmap displaying differentially expressed genes in sgControl and sgCdk10 CT26 cells, based on transcriptomic (a) and proteomic (b) sequencing data. c, Reverse transcription quantitative PCR (RT-qPCR) analysis of the indicated genes in sgControl and sgCdk10 B16F10 cells. d, IB analysis of WCL derived from sgControl and sgCdk10 B16F10 cells. e, IB analysis of WCL derived from sgControl and sgCdk10 NCM460 cells. f, RT-qPCR analysis of the indicated genes in sgControl and sgCdk10 NCM460 cells. g, Western blot analysis of CDK10 in normal colon cells (NCM460) and various CRC cell lines. Vinculin served as a loading control. h, IB analysis of WCL derived from CT26 cells with indicated treatments. i, RT-qPCR analysis of the indicated genes. j, IB of WCL from human colorectal cancer (CRC) tumor samples, n=11 tumor samples. k–u, Correlations between the CDK10 expression and pS172-TBK1 (k), pS396-IRF3 (l), IFIT3 (m), OAS2 (n), BST2 (o), IRGM (p), GBP2 (q), OAS1 (r), STAT1 (s), IFI44 (t) and ISG15 (u) in colorectal tumor samples, n = 11 tumor samples. Data are presented as mean ± SEM, for c, f, and i, n = 3 biologically independent samples, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For k–u, correlation coefficients were calculated using the Pearson two-taided test. For d, e, g, and h, three independent experiments were performed for each image with similar results.
Extended Data Fig. 5 Cdk10 suppresses nucleic acid sensors-mediated innate immune responses.
a, IB analysis of WCL derived from CT26 cells with indicated sgRNA treatments. b, c, pS172-TBK1 level (b) and pS396-IRF3 level (c) was quantified by ImageJ for a, which was normalized to vinculin. d, IB analysis of WCL derived from CT26 cells with indicated sgRNA treatments. e, f, pS172-TBK1 level (e) and pS396-IRF3 level (f) was quantified by ImageJ for d, which was normalized to vinculin. g, RT-qPCR analysis of the indicated genes. h, i, IB analysis of WCL derived from sgControl and sgCdk10 CT26 cells treated with Ru.521 (10 µM, 36 h) (h) or H-151 (1 µM, 36 h) (i). j, RT-qPCR analysis of the indicated genes derived from sgControl and sgCdk10 CT26 cells treated with Ru.521 (10 µM, 36 h) or H-151 (1 µM, 36 h). k, IB analysis of WCL derived from CT26 cells with indicated sgRNA treatments. l, RT-qPCR analysis of the indicated genes. m, n, Tumor growth curve (m) and end point tumor weight (n) of indicated CT26 tumors in immunocompetent BALB/c WT mice. n = 7 mice for sgControl group and sgCdk10 group, n = 6 mice for sgCdk10 + sgcGas group, sgCdk10 + sgMda5 group and sgCdk10 + sgcGas + sgMda5 group. For g, j, and l, n = 3 biologically independent samples. Data are presented as mean ± SEM, for b, c, e, f, g, j, l, and n, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test; for m, analyzed by two-way ANOVA with Tukey’s multiple-comparisons test. For a, d, h, i, and k, three independent experiments were performed for each image with similar results.
Extended Data Fig. 6 CDK10 interacts with and phosphorylates DNMT1, thereby suppressing dsRNA production.
a, Heatmap showing differentially expressed genes in sgControl and sgCdk10 CT26 cells based on phosphoproteomics sequencing. b, RT-qPCR analysis of Dnmt1 mRNA levels in sgControl and sgCdk10 CT26 cells. c, IB analysis of WCL and anti-DNMT1 immunoprecipitation (IP) from DLD1 cells. d, IB analysis of glutathione S-transferase (GST) pull-down assays from WCL of HEK293T cells expressing Flag-DNMT1, incubated with bacterially purified recombinant GST or GST-CDK10 protein. e, Schematic illustration of DNMT1, showing its various domains and truncated mutants. f, g, Immunofluorescence (IF) images showing anti-dsRNA (J2) staining and quantification of cytoplasmic dsRNA mean fluorescence intensity (MFI) in sgControl and sgCDK10 DLD1 cells. Scale bar, 20 μm. n = 20 cell/group. h, RT-qPCR analysis of the indicated genes in sgControl and sgCdk10 CT26 cells. i, IB analysis of WCL derived from CT26 cells with indicated treatments. For b and h, n = 3 biologically independent samples, data are represented as mean ± SEM, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For g, the box represents the interquartile range (IQR); the central line represents the median, while the upper and lower edges of the box correspond to the first and third quartiles, respectively; the whiskers extend to the minimum and maximum values; all individual data points are shown, analyzed by two-tailed unpaired t-test. For c, d, f, and i, three independent experiments were performed for each image with similar results.
Extended Data Fig. 7 CDK10 interacts with and phosphorylates RAP80 at serine 379, thereby suppressing R-loop production.
a, IB analysis of GST pull-down assays from WCL of HEK293T cells expressing HA-CDK10, incubated with bacterially purified recombinant GST or GST-hRAP80 protein. b, A schematic presentation of putative CDK10 phosphorylation motif around S379 site in RAP80. c, in vitro kinase assays using bacterially purified recombinant GST-mRap80-WT and S379A mutant proteins, incubated with or without active CDK10/cyclin M kinase. d, IB analysis of WCL and anti-HA IPs from HEK293T cells transfected with indicted mRap80 WT or S379A mutant. e, f, Representative IF images of γH2AX (red) and corresponding foci quantification. Images show γH2AX staining in sgControl and sgCdk10 CT26 cells (e). Scale bar, 20 µm. The graph quantifies the number of γH2AX foci per nucleus (f); n = 30 cells/group. g, h, IF images of S9.6 staining and quantification of nuclear and cytoplasmic R-loop (MFI) in sgControl and sgCDK10 DLD1 cells. Scale bars, 20 µm. n = 20 cell/group. i, j, IF images of GFP-dRNASEH1 staining and quantification of nuclear R-loop and cytoplasmic hybrid MFI in sgControl and sgCdk10 CT26 cells with or without treatment of RNase H (37 °C, 4 h). Scale bar, 20 μm. n = 20 cell/group. k, l, IF images of GFP-dRNASEH1 staining and quantification of nuclear R-loop and cytoplasmic hybrid MFI in indicated cells. Scale bar, 20 μm. n = 20 cell/group. For f, h, j, and l, the box represents the IQR; the central line represents the median, while the upper and lower edges of the box correspond to the first and third quartiles, respectively; the whiskers extend to the minimum and maximum values; all individual data points are shown, for f, h, analyzed by two-tailed unpaired t-test; for j, l,analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For a, c–e, g, i, and k, three independent experiments were performed for each image with similar results.
Extended Data Fig. 8 Phenotypes of mice with Cdk10 conditional knockout (cKO) in intestinal epithelial cells.
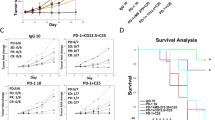
a, Schematic diagram illustrating the cKO strategy for Cdk10 in intestinal epithelial cells of C57BL/6J mice, achieved using CRISPR/Cas9 technology to edit the Cdk10 gene. b, IB of WCL from intestinal epithelial cells (IECs) sorted from WT and Cdk10 cKO mice. n = 3 mice/group. c, d, Mendelian inheritance ratios from the interbreeding of Cdk10fl/+ and Cdk10fl/+; Vil-Cre/+ mice (c). Number and sex ratio of mice were analyzed (d). e, Body weight of female and male 8-week-old WT and Cdk10 cKO mice. For males, n = 5 mice/group; For females n = 7 mice/group. f, General appearance of representative 8-week-old WT and Cdk10 cKO mice. n = 3 mice/group. g, Representative images of the kidneys and spleens from 8-week-old WT and Cdk10 cKO mice. n = 3 mice/group. h, IB of WCL from IECs sorted from WT and Cdk10 cKO mice. n = 3 mice/group. i, RT-qPCR analysis of mRNA levels of indicated genes in IECs derived from WT and Cdk10 cKO mice. n = 3 mice/group. j, Schematic of the AOM/DSS-induced colorectal cancer model. k–m, Representative images of tumor-bearing colon and rectum (k), tumor count per mouse (l), and representative H&E staining (m) of colonic tissues from WT and Cdk10 cKO mice after AOM/DSS treatment. Tumor count was assessed at the study end point. Scale bars, 1 mm (m). n = 6 mice/group (l). n, IB of WCL from colorectal tumors sorted from WT and Cdk10 cKO mice. n = 3 mice/group. o, RT-qPCR analysis of mRNA levels of indicated genes in colorectal tumors derived from WT and Cdk10 cKO mice. n = 3 mice/group. p–r, Clinical studies showing that high CDK10 expression is associated with poor survival in patients treated with anti-PD-1 (p), anti-PD-L1 (q), and anti-CTLA-4 (r) therapy. A positive Z-value indicates that higher CDK10 expression is associated with worse survival, whereas a negative value suggests a protective effect. For e, i, l, and o, data are presented as mean ± SEM, analyzed by two-tailed unpaired t-test. For p–r, a Cox proportional hazards (CoxPH) model was applied to compute a P value reflecting the strength of the correlation between gene expression and OS.
Extended Data Fig. 9 Ponatinib and NVP-AST487 induce CDK10-dependent activation of innate immune signaling.
a, Chemical structures of NVP-AST487 and ponatinib. b, Relative activity of CDK10/cyclin M, CDK2/cyclin E1, CDK4/cyclin D1 and CDK6/cyclin D1 following treatment with ponatinib and NVP-AST487, assessed by the NanoBret assay. c, d, IB analysis of WCL derived from sgControl and sgCdk10 CT26 cells treated with ponatinib (c) or NVP-AST487 (d) for 48 h. e, IF images showing anti-dsRNA (J2) staining and quantification of cytoplasmic dsRNA signal intensity in sgControl and sgCdk10 CT26 cells treated with ponatinib (0.25 µM, 24 h), NVP-AST487 (0.25 µM, 24 h), or DMSO. Scale bars, 20 µm. n = 20 cell/group. f, IF images of GFP-dRNASEH1 staining and quantification of nuclear R-loops and cytoplasmic hybrids in sgControl and sgCdk10 CT26 cells treated with ponatinib (0.25 µM, 24 h), NVP-AST487 (0.25 µM, 24 h), or DMSO. Scale bars, 20 µm. n = 20 cell/group. g, h, mRNA expression levels of the indicated genes in sgControl and sgCdk10 CT26 cells treated with ponatinib (0.25 μM, 48 h) (g), NVP-AST487 (0.25 µM, 48 h) (h) or DMSO. For e, f, the box represents the IQR; the central line represents the median, while the upper and lower edges of the box correspond to the first and third quartiles, respectively; the whiskers extend to the minimum and maximum values, all individual data points are shown, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For g, h, n = 3 biologically independent samples, data are presented as mean ± SEM, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For b–f, three independent experiments were performed with similar results, and a representative experiment is shown.
Extended Data Fig. 10 Ponatinib and NVP-AST487 exhibit no notable toxicity in mice and demonstrate superior antitumor efficacy compared to palbociclib in the CT26 murine model.
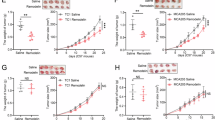
a, b, Tumor growth curve (a) and end point tumor weight (b) of sgControl and sgCdk10 CT26 tumors in immunodeficient BALB/c nude mice, treated with ponatinib (30 mg/kg), NVP-AST487 (30 mg/kg) or Vehicle. n = 9 mice for sgControl + Vehicle group, sgControl + ponatinib group and sgControl + NVP-AST487 group, n = 10 mice/group for sgCdk10 + Vehicle group, sgCdk10 + ponatinib group and sgCdk10 + NVP-AST487 group. c, Time-course monitoring of body weight in immunocompetent BALB/c WT mice during the indicated treatment. n = 7 mice/group. d–f, RBC (d), WBC (e), and PLT (f) counts were measured in immunocompetent BALB/c WT mice with the indicated treatments. n = 7 mice/group. g, h, Liver function was evaluated by measuring AST (g) and ALT (h) levels in immunocompetent BALB/c WT mice following the indicated treatments. n = 7 mice/group. i, H&E staining for the slices of the several organs (liver, spleen, lung and colon) from immunocompetent BALB/c WT mice. Scale bar, 50 μm. j, k, Growth curve (j) and Kaplan–Meier (k) survival curves of subcutaneous WT CT26 tumors in immunocompetent BALB/c WT mice following the indicated treatments. n = 12 mice/group. Anti-PD-1 antibody treatment was given by i.p. injection of 200 µg per mouse every three days, for a total of seven injections. l, A schematic model of CDK10-mediated regulation of tumor immunity. Schematic model of CDK10-mediated regulation of tumor immunity. CDK10 plays a critical role in suppressing the accumulation of dsRNA and R-loops by phosphorylating DNMT1 and RAP80. This phosphorylation prevents the activation of the MDA5 and cGAS-STING innate immune pathways. Consequently, genetic or pharmacological inhibition of CDK10 triggers the activation of both the MDA5 and cGAS-STING pathways, thereby enhancing antitumor immunity and improving the efficacy of immune-checkpoint blockade (ICB) therapy. Data are represented as mean ± SEM, for a, c and j, analyzed by two-way ANOVA with Tukey’s multiple-comparisons test; for b, d–h, analyzed by one-way ANOVA with Tukey’s multiple-comparisons test. For k, Kaplan–Meier survival analysis and two-tailed Gehan–Breslow-Wilcoxon test were performed.
Supplementary information
Supplementary Tables 1–10 (download XLSX )
Supplementary Tables 1–10.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data (download PDF )
Unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, G., Guo, F., He, C. et al. CDK10 suppresses nucleic acid sensors-mediated antitumor immunity. Nat Cancer 7, 283–303 (2026). https://doi.org/10.1038/s43018-025-01100-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01100-3
This article is cited by
-
CDK10 inhibits innate sensing to constrain antitumor immunity
Nature Cancer (2026)