Abstract
Oligodendrocyte progenitor cells (OPCs) are highly dynamic, widely distributed glial cells of the central nervous system responsible for generating myelinating oligodendrocytes throughout life. However, the rates of OPC proliferation and differentiation decline dramatically with aging, which may impair homeostasis, remyelination and adaptive myelination during learning. To determine how aging influences OPCs, we generated a transgenic mouse line (Matn4-mEGFP) and performed single-cell RNA sequencing, providing enhanced resolution of transcriptional changes during key transitions from quiescence to proliferation and differentiation across the lifespan. We found that aging induces distinct transcriptomic changes in OPCs in different states, including enhanced activation of HIF-1α and WNT pathways. Pharmacological inhibition of these pathways in aged OPCs was sufficient to increase their ability to differentiate in vitro. Ultimately, Matn4-mEGFP mouse line and the sequencing dataset of cortical OPCs across ages will help to define the molecular changes guiding OPC behavior in various physiological and pathological contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All raw and preprocessed sequencing data generated for this study as well as the processed Monocle 3 cell_data_set (cds) object have been deposited in NCBI Gene Expression Omnibus (GEO) with accession code GSE249268. To promote open access to data, we deposited the annotated dataset onto the Chan Zuckerberg CELL by GENE Discover Platform (https://tinyurl.com/aging-opcs). Source data are provided with this paper. Any other data reported in this paper are available from the lead contact upon reasonable request.
Code availability
No new code for the analysis of data was created.
References
Salas, A. et al. JAK-STAT pathway targeting for the treatment of inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 17, 323–337 (2020).
Clarke, L. E. et al. Normal aging induces A1-like astrocyte reactivity. Proc. Natl Acad. Sci. USA 115, E1896–E1905 (2018).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Guttenplan, K. A. et al. Neurotoxic reactive astrocytes induce cell death via saturated lipids. Nature 599, 102–107 (2021).
Young, K. M. et al. Oligodendrocyte dynamics in the healthy adult CNS: evidence for myelin remodeling. Neuron 77, 873–885 (2013).
Hill, R. A., Li, A. M. & Grutzendler, J. Lifelong cortical myelin plasticity and age-related degeneration in the live mammalian brain. Nat. Neurosci. 21, 683–695 (2018).
Zonouzi, M. et al. Individual oligodendrocytes show bias for inhibitory axons in the neocortex. Cell Rep 27, 2799–2808.e3 (2019).
Bacmeister, C. M. et al. Motor learning promotes remyelination via new and surviving oligodendrocytes. Nat. Neurosci. 23, 819–831 (2020).
Hughes, E. G., Orthmann-Murphy, J. L., Langseth, A. J. & Bergles, D. E. Myelin remodeling through experience-dependent oligodendrogenesis in the adult somatosensory cortex. Nat. Neurosci. 21, 696–706 (2018).
Nishiyama, A., Shimizu, T., Sherafat, A. & Richardson, W. D. Life-long oligodendrocyte development and plasticity. Semin. Cell Dev. Biol. 116, 25–37 (2021).
Sim, F. J., Zhao, C., Penderis, J. & Franklin, R. J. M. The age-related decrease in CNS remyelination efficiency is attributable to an impairment of both oligodendrocyte progenitor recruitment and differentiation. J. Neurosci. 22, 2451–2459 (2002).
Neumann, B. et al. Metformin restores CNS remyelination capacity by rejuvenating aged stem cells. Cell Stem Cell 25, 473–485 (2019).
Kirby, L. et al. Oligodendrocyte precursor cells present antigen and are cytotoxic targets in inflammatory demyelination. Nat. Commun. 10, 3887 (2019).
Falcão, A. M. et al. Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat. Med. 24, 1837–1844 (2018).
Park, H. et al. Single-cell RNA-sequencing identifies disease-associated oligodendrocytes in male APP NL-G-F and 5XFAD mice. Nat. Commun. 14, 802 (2023).
Dawson, M. R. L., Polito, A., Levine, J. M. & Reynolds, R. NG2-expressing glial progenitor cells: an abundant and widespread population of cycling cells in the adult rat CNS. Mol. Cell. Neurosci. 24, 476–488 (2003).
Uckelmann, H. et al. Extracellular matrix protein Matrilin-4 regulates stress-induced HSC proliferation via CXCR4. J. Exp. Med. 213, 1961–1971 (2016).
Hrvatin, S. et al. Single-cell analysis of experience-dependent transcriptomic states in the mouse visual cortex. Nat. Neurosci. 21, 120–129 (2018).
Nishiyama, A., Chang, A. & Trapp, B. D. NG2+ glial cells: a novel glial cell population in the adult brain. J. Neuropathol. Exp. Neurol. 58, 1113–1124 (1999).
Daneman, R. et al. The mouse blood-brain barrier transcriptome: a new resource for understanding the development and function of brain endothelial cells. PLoS ONE 5, e13741 (2010).
Dani, N. et al. A cellular and spatial map of the choroid plexus across brain ventricles and ages. Cell 184, 3056–3074.e21 (2021).
Cembrowski, M. S., Wang, L., Sugino, K., Shields, B. C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
Sullivan, K. E. et al. Sharp cell-type-identity changes differentiate the retrosplenial cortex from the neocortex. Cell Rep. 42, 112206 (2023).
Hughes, E. G., Kang, S. H., Fukaya, M. & Bergles, D. E. Oligodendrocyte progenitors balance growth with self-repulsion to achieve homeostasis in the adult brain. Nat. Neurosci. 16, 668–676 (2013).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Fard, M. K. et al. BCAS1 expression defines a population of early myelinating oligodendrocytes in multiple sclerosis lesions. Sci. Transl. Med. 9, eaam7816 (2017).
Xiao, L. et al. Rapid production of new oligodendrocytes is required in the earliest stages of motor-skill learning. Nat. Neurosci. 19, 1210–1217 (2016).
He, D. et al. lncRNA functional networks in oligodendrocytes reveal stage-specific myelination control by an lncOL1/Suz12 complex in the CNS. Neuron 93, 362–378 (2017).
Adams, K. L. et al. Endothelin-1 signaling maintains glial progenitor proliferation in the postnatal subventricular zone. Nat. Commun. 11, 2138 (2020).
Beiter, R. M. et al. Evidence for oligodendrocyte progenitor cell heterogeneity in the adult mouse brain. Sci. Rep. 12, 12921 (2022).
Psachoulia, K., Jamen, F., Young, K. M. & Richardson, W. D. Cell cycle dynamics of NG2 cells in the postnatal and ageing brain. Neuron Glia Biol. 5, 57–67 (2009).
Kang, S. H., Fukaya, M., Yang, J. K., Rothstein, J. D. & Bergles, D. E. NG2+ CNS glial progenitors remain committed to the oligodendrocyte lineage in postnatal life and following neurodegeneration. Neuron 68, 668–681 (2010).
Spitzer, S. O. et al. Oligodendrocyte progenitor cells become regionally diverse and heterogeneous with age. Neuron 101, 459–471 (2019).
Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023).
Stein-O’Brien, G. L. et al. Decomposing cell identity for transfer learning across cellular measurements, platforms, tissues, and species. Cell Syst 8, 395–411.e8 (2019).
Sharma, G., Colantuoni, C., Goff, L. A., Fertig, E. J. & Stein-O’Brien, G. projectR: an R/Bioconductor package for transfer learning via PCA, NMF, correlation and clustering. Bioinformatics 36, 3592–3593 (2020).
CZI Cell Science Program. CZ CELLxGENE Discover: a single-cell data platform for scalable exploration, analysis and modeling of aggregated data. Nucleic Acids Res. 53, D886–D900 (2021).
Zheng, S. C. et al. Universal prediction of cell-cycle position using transfer learning. Genome Biol 23, 41 (2022).
Stein-O’Brien, G. L. et al. Enter the matrix: factorization uncovers knowledge from omics determining the dimensions of biology from omics data. Trends Genet. 34, 790–805 (2018).
Baruch, K. et al. Aging. Aging-induced type I interferon signaling at the choroid plexus negatively affects brain function. Science 346, 89 (2014).
Xu, H., Dzhashiashvili, Y., Ming, G.-L., He, C. & Correspondence, B. P. m6A mRNA methylation is essential for oligodendrocyte maturation and CNS myelination. Neuron 105, 293–309 (2020).
Kotter, M. R., Stadelmann, C. & Hartung, H.-P. Enhancing remyelination in disease—can we wrap it up? Brain 134, 1882–1900 (2011).
Alto, L. T. & Terman, J. R. Semaphorins and their signaling mechanisms. Methods Mol. Biol. 1493, 1 (2017).
Cohen, R. I., Rottkamp, D. M., Maric, D., Barker, J. L. & Hudson, L. D. A role for semaphorins and neuropilins in oligodendrocyte guidance. J. Neurochem. 85, 1262–1278 (2003).
Bernard, F. et al. Role of transmembrane semaphorin Sema6A in oligodendrocyte differentiation and myelination. Glia 60, 1590–1604 (2012).
Segel, M. et al. Niche stiffness underlies the ageing of central nervous system progenitor cells. Nature 573, 130–134 (2019).
Emery, B. et al. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell 138, 172–185 (2009).
Zezula, J. et al. p21cip1 is required for the differentiation of oligodendrocytes independently of cell cycle withdrawal. EMBO Rep. 2, 27–34 (2001).
Qi, Y. et al. Control of oligodendrocyte differentiation by the Nkx2.2 homeodomain transcription factor. Development 128, 2723–2733 (2001).
Emery, B. & Lu, Q. R. Transcriptional and epigenetic regulation of oligodendrocyte development and myelination in the central nervous system. Cold Spring Harb. Perspect. Biol. 7, a020461 (2015).
Iram, T. et al. Young CSF restores oligodendrogenesis and memory in aged mice via Fgf17. Nature 605, 509–515 (2022).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and -independent cellular responses in Alzheimer’s disease. Nat. Med. 26, 131 (2020).
Liu, J. Z. et al. BDH2 triggers ROS-induced cell death and autophagy by promoting Nrf2 ubiquitination in gastric cancer. J. Exp. Clin. Cancer Res. 39, 123 (2020).
Krämer, A., Green, J., Pollard, J. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Neumann, B. et al. Myc determines the functional age state of oligodendrocyte progenitor cells. Nat Aging 1, 826–837 (2021).
Tang, K. et al. HIF-1α stimulates the progression of oesophageal squamous cell carcinoma by activating the Wnt/β-catenin signalling pathway. Br. J. Cancer 127, 474–487 (2022).
Xu, W. et al. Hypoxia activates Wnt/β-catenin signaling by regulating the expression of BCL9 in human hepatocellular carcinoma. Sci. Rep. 7, 40446 (2017).
Fancy, S. P. J. et al. Dysregulation of the Wnt pathway inhibits timely myelination and remyelination in the mammalian CNS. Genes Dev. 23, 1571 (2009).
de la Fuente, A. G. et al. Changes in the oligodendrocyte progenitor cell proteome with ageing. Mol. Cell. Proteomics 19, 1281–1302 (2020).
Zhang, P. et al. Senolytic therapy alleviates Aβ-associated oligodendrocyte progenitor cell senescence and cognitive deficits in an Alzheimer’s disease model. Nat. Neurosci. 22, 719–728 (2019).
Chapman, T. W., Olveda, G. E., Bame, X., Pereira, E. & Hill, R. A. Oligodendrocyte death initiates synchronous remyelination to restore cortical myelin patterns in mice. Nat. Neurosci. 26, 555–569 (2023).
von Streitberg, A. et al. NG2-glia transiently overcome their homeostatic network and contribute to wound closure after brain injury. Front Cell Dev Biol 9, 662056 (2021).
Gibson, E. M. et al. Neuronal activity promotes oligodendrogenesis and adaptive myelination in the mammalian brain. Science 344, 1252304 (2014).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Marques, S. et al. Transcriptional convergence of oligodendrocyte lineage progenitors during development. Dev. Cell 46, 504–517.e7 (2018).
Kessaris, N. et al. Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat. Neurosci. 9, 173–179 (2006).
Tripathi, R. B. et al. Dorsally and ventrally derived oligodendrocytes have similar electrical properties but myelinate preferred tracts. J. Neurosci. 31, 6809–6819 (2011).
Vallstedt, A., Klos, J. M. & Ericson, J. Multiple dorsoventral origins of oligodendrocyte generation in the spinal cord and hindbrain. Neuron 45, 55–67 (2005).
Crawford, A. H., Tripathi, R. B., Richardson, W. D. & Franklin, R. J. M. Developmental origin of oligodendrocyte lineage cells determines response to demyelination and susceptibility to age-associated functional decline. Cell Rep. 15, 761–773 (2016).
Floriddia, E. M. et al. Distinct oligodendrocyte populations have spatial preference and different responses to spinal cord injury. Nat. Commun. 11, 5860 (2020).
Hilscher, M. M. et al. Spatial and temporal heterogeneity in the lineage progression of fine oligodendrocyte subtypes. BMC Biol. 20, 122 (2022).
Lin, J. P. et al. Transcriptomic architecture of nuclei in the marmoset CNS. Nat. Commun. 13, 5531 (2022).
Hill, R. A. et al. NG2 cells in white matter but not gray matter proliferate in response to PDGF. J. Neurosci. 33, 14558–14566 (2013).
Rivers, L. E. et al. PDGFRA/NG2 glia generate myelinating oligodendrocytes and piriform projection neurons in adult mice. Nat. Neurosci. 11, 1392–1401 (2008).
Gonsalvez, D. G. et al. The dynamics of oligodendrocyte generation: how distinct is the mouse from the human? Preprint at bioRxiv https://doi.org/10.1101/2019.12.23.887174 (2019).
Wang, S. et al. Ermin is a p116 RIP -interacting protein promoting oligodendroglial differentiation and myelin maintenance. Glia 68, 2264–2276 (2020).
Xing, Y. L. et al. High-efficiency pharmacogenetic ablation of oligodendrocyte progenitor cells in the adult mouse CNS. Cell Rep. Methods 3, 100414 (2023).
Buchanan, J. A. et al. Oligodendrocyte precursor cells ingest axons in the mouse neocortex. Proc. Natl Acad. Sci. USA 119, e2202580119 (2022).
Auguste, Y. S. S. et al. Oligodendrocyte precursor cells engulf synapses during circuit remodeling in mice. Nat. Neurosci. 25, 1273–1278 (2022).
Xiao, Y., Petrucco, L., Hoodless, L. J., Portugues, R. & Czopka, T. Oligodendrocyte precursor cells sculpt the visual system by regulating axonal remodeling. Nat. Neurosci. 25, 280–284 (2022).
Levine, J. M. Increased expression of the NG2 chondroitin-sulfate proteoglycan after brain injury. J. Neurosci. 14, 4716–4730 (1994).
Bradbury, E. J. et al. Chondroitinase ABC promotes functional recovery after spinal cord injury. Nature 416, 636–640 (2002).
Wei, H. et al. Glial progenitor heterogeneity and key regulators revealed by single-cell RNA sequencing provide insight to regeneration in spinal cord injury. Cell Rep 42, 112486 (2023).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Safaiyan, S. et al. White matter aging drives microglial diversity. Neuron 109, 1100–1117.e10 (2021).
Kaya, T. et al. CD8+ T cells induce interferon-responsive oligodendrocytes and microglia in white matter aging. Nat. Neurosci. 25, 1446–1457 (2022).
Harrington, E. P. et al. MHC class i and MHC class ii reporter mice enable analysis of immune oligodendroglia in mouse models of multiple sclerosis. eLife 12, e82938 (2023).
Nicaise, A. M. et al. Cellular senescence in progenitor cells contributes to diminished remyelination potential in progressive multiple sclerosis. Proc. Natl Acad. Sci. USA 116, 9030–9039 (2019).
Yuen, T. J. et al. Oligodendrocyte-encoded HIF function couples postnatal myelination and white matter angiogenesis. Cell 158, 383–396 (2014).
Allan, K. C. et al. Non-canonical targets of HIF1a impair oligodendrocyte progenitor cell function. Cell Stem Cell 28, 257–272.e11 (2021).
Kuhlmann, T. et al. Differentiation block of oligodendroglial progenitor cells as a cause for remyelination failure in chronic multiple sclerosis. Brain 131, 1749–1758 (2008).
Call, C. L. & Bergles, D. E. Cortical neurons exhibit diverse myelination patterns that scale between mouse brain regions and regenerate after demyelination. Nat. Commun. 12, 4767 (2021).
Melsted, P. et al. Modular, efficient and constant-memory single-cell RNA-seq preprocessing. Nat. Biotechnol. 39, 813–818 (2021).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Schwarzkopf, M. et al. Hybridization chain reaction enables a unified approach to multiplexed, quantitative, high-resolution immunohistochemistry and in situ hybridization. Development 148, dev199847 (2021).
Acknowledgements
We thank C. Hawkins at JHMI Transgenic Core Laboratory for performing CRISPR–Cas9 microinjections and assisting in the generation of the Matn4-mEGFP mouse line. We also thank M. Pucak and A. Smirnov at JHMI Neuroscience Imaging Center for their assistance with image acquisition and analysis. R. Kawaguchi at UCLA provided oversight for the sequencing of 10x scRNA-seq libraries through AMRF. We also thank our colleagues for their invaluable support throughout this study. This research was supported by grants from the NIH (AG072305, NS041435), the Goldman Foundation and the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (AMRF). D.H. and A.K. were supported by fellowships from the NIH (F31NS110204 and F30AG084193, respectively). Y.M. was supported by a fellowship from the National MS Society (FG-1708-28962).
Author information
Authors and Affiliations
Contributions
D.H., L.A.G. and D.E.B. conceived the project, designed the experiments, and wrote the manuscript with input from the other authors. A.A.K. and V.N.D. performed FISH validation experiments. B.N. and R.J.M.F. performed and analyzed in vitro pharmacological validation experiments. Y.K.T.X. and Y.A.M. performed in vivo imaging of OPC reporter mouse lines. D.H., J.S. and L.A.G. performed bioinformatical analyses of the scRNA-seq dataset. L.A.G., R.J.M.F. and D.E.B. provided funding for the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Matn4-mEGFP expression is restricted to OPCs and a subset of neurons.
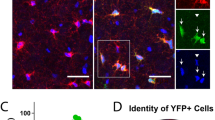
a. Matn4 expression is specific to OPCs and newly-formed oligodendrocytes (NFO) in 6-7 week-old mouse V1 cortex (reanalysis of a publicly available scRNA-seq dataset18). b. Genotyping result of Matn4-mEGFP mouse line. Wildtype (wt) band size is 178 bp whereas the mutant, knock-in band size is 356 bp. c. EGFP signal in the optic nerve of Matn4-mEGFP mouse line is restricted to NG2 + PDGFRα+ OPCs. d. Matn4-mEGFP is also expressed by hippocampal granule cells and neurons in the somatosensory cortex barrel field and retrosplenial cortex. e. Matn4-mEGFP signal is absent from Iba1+ microglia and GFAP+ astrocytes. f. In vivo imaging of GFP+ cells in Matn4-mEGFP, NG2-mEGFP, and Pdgfra-CreER; RCE mouse lines. None of the vascular cells (red arrowheads) express EGFP in the cortex of Matn4-mEGFP mice.
Extended Data Fig. 2 Preprocessing of the OPC scRNA-seq dataset.
a. Expression of oligodendrocyte lineage cell genes in the uncleaned dataset. Most cells in the dataset express Cspg4, Pdgfra, and Olig2 (OPCs) or Enpp6 and Olig2 (differentiating OPCs). b. Only a small group of cells that were FACS isolated from Matn4-mEGFP mouse line express non-oligodendrocyte lineage cell genes. c. UMAP plot of uncleaned dataset colorized by the percentage of mitochondrial-related genes (cutoff at 10%). Those cells with relatively high mitochondrial gene ratio (>5%) were removed for downstream analyses. d. Expression of classic oligodendrocyte lineage marker genes (oligodendrocyte lineage: Olig2, Sox10; OPC: Pdgfra, Cspg4, Matn4; differentiating OPC: Bcas1, Enpp6, 9630013A20Rik; oligodendrocyte: Mbp, Mobp) as well as the subtype marker genes identified in this study (Cycling OPC: Top2a, Mcm3, Mki67; Transitioning OPC: Gap43, Rplp0) in the cleaned, preprocessed, final dataset.
Extended Data Fig. 3 Fluorescent in situ hybridization (FISH) for Cycling and Differentiating OPC subtypes.
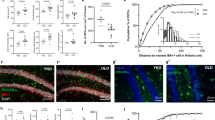
a. FISH for Top2a, Pdgfra, and Sox10 to identify Cycling OPC in situ in postnatal day 9 (P9) mouse brain. b. Comparison of the density of Cycling OPC in highly myelinated, somatosensory cortex (SS) and that in sparsely myelinated, temporal association cortex (TEA). c. Quantification of the density of Cycling OPC (Top2a+ Pdgfra + ) in SS and TEA. d. FISH for LncOL1 to identify Differentiating OPC in situ in P74 mouse brain. e. Quantification of the frequency of LncOL1+ Differentiating OPC in SS and TEA. The exact p-values are reported in Source Data file for Extended Data Fig. 3.
Extended Data Fig. 4 OPC subtype-specific gene patterns are shared between mouse and human cortical OPCs.
a. UMAP plot of the dataset colorized by their four age groups (blue: 29-yr old, red: 42-yr old, green: 50-yr old, and purple: 60-yr old). b. UMAP plot of 26,357 human cortical OPCs colorized by their identified subtypes (quiescent OPC, cycling OPC, differentiating OPC, and oligodendrocytes). c. Expression of classic oligodendrocyte lineage marker genes (oligodendrocyte lineage: OLIG2, SOX10; OPC: PDGFRA, CSPG4, MATN4; differentiating OPC: BCAS1, ENPP6; oligodendrocyte: MBP, MOBP) as well as the subtype marker genes identified in this study (Cycling OPC: TOP2A, MCM3, MKI67; Transitioning OPC: GAP43, RPLP0) in the human cortical OPC dataset. d. UMAP plots of mouse Cycling OPC gene pattern 5 projected on the mouse scRNA-seq dataset and on the human snRNA-seq dataset. e. UMAP plots of mouse Differentiating OPC gene pattern 25 projected on the mouse scRNA-seq dataset and on the human snRNA-seq dataset. f. Dot plot of HIF1A and CTNNB1 expression in quiescent OPCs in the human cortex across aging.
Extended Data Fig. 5 Expression changes in individual cycling genes and groups of genes in Cycling OPC 1 and directly anteceding Quiescent OPC.
a. Expression levels of known cycling genes enriched in cycling OPCs are comparable in Cycling OPC 1 throughout aging. b. NMF gene patterns that are associated with either aged (P180-720) or young (P30) OPCs.
Extended Data Fig. 6 OPCs upregulate C4b, Hif1a, and Ctnnb1 mRNA with aging.
a. FISH with immunofluorescence staining (IF) for Pdgfra and C4b in P35 and P315 Matn4-mEGFP mouse cortex. OPC cell body masks were created based on EGFP fluorescence and Pdgfra FISH signal and used to quantify C4b transcript puncta/OPC (Two-tailed, Mann-Whitney U test, n = 65, 49 cells, * p-value = 0.0177, error bars = SEM). b. FISH with IF staining for Pdgfra and Hif1a in the P35 and P315 Matn4-mEGFP mouse cortex. Hif1a transcript puncta/OPC was quantified as described above (Two-tailed, Mann-Whitney U test, n = 56, 71 cells, * p-value = 0.0385, error bars = SEM). c. FISH with IF staining for Pdgfra and Ctnnb1 in the P35 and P315 Matn4-mEGFP mouse cortex (Two-tailed, Mann-Whitney U test, n = 59, 55 cells, ** p-value = 0.0069, error bars = SEM). The exact p-values are reported in Source Data file for Extended Data Fig. 6.
Extended Data Fig. 7 Wnt signaling pathway is activated in aged OPCs and may contribute to their decreased differentiation potential.
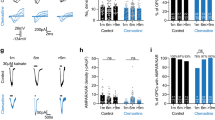
a. Dot plot of Ctnnb1 expression in Quiescent OPC from P180, P360, and P720 timepoints. b. Schematic of how two different Wnt inhibitors (IWP-2 and XAV939) differentially block Wnt signaling pathway. IWP-2 globally inhibits the Wnt pathway whereas XAV939 preferentially inhibits the canonical Wnt signaling pathway. Both non-canonical and canonical Wnt signaling pathways have been shown to regulate DNA damage response. c. Quantification of MBP+ differentiating OPC/Olig2+ oligodendrocyte proportions with and without Wnt inhibitor treatments in OPCs isolated from YA or AA (two-way ANOVA with Tukey’s multiple comparisons test, n = 6, 6, * p-value < 0.05, ** p-value < 0.01, error bars = SEM). The exact p-values are reported in Source Data file for Extended Data Fig. 7.
Extended Data Fig. 8 Matn4-mEGFP signal is restricted to OPCs even after a stab wound injury.
IHC against NG2 (red) and EGFP (green) was performed on the Matn4-mEGFP mouse following a stab wound injury to demonstrate the utility of the mouse line in studying OPC dynamics following injury and inflammation.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Tables 1–5. (download XLSX )
Table 1, List of marker genes used to define OPC subtypes. Table 2, List of genes corresponding to different gene modules associated with Quiescent, Transitioning and Differentiating OPC clusters. Table 3, Pseudotime differential gene expression (graph_test) of genes that encode transcription factors along the OPC differentiation trajectory. Table 4, Differential gene expression results of Quiescent OPCs in aging (P30 versus P180, P360 and P720). Table 5, Gene weights for Cycling and Differentiating OPC subtypes.
Supplementary Video 1 (download AVI )
1-h time-lapse imaging of OPCs in the motor cortex of Matn4-mEGFP.
Supplementary Video 2 (download AVI )
Z-stack image of the OPCs pseudocolored in green (at baseline of imaging) and magenta (50 min after baseline).
Source data
Source Data Fig. 1 (download XLSX )
Quantification for the graphs Fig. 1f–g.
Source Data Fig. 2 (download XLSX )
Raw cell counts for Fig. 2e.
Source Data Fig. 6 (download XLSX )
Quantification for the graphs Fig. 6c,e.
Source Data Extended Data Fig. 3 (download XLSX )
Quantification for the graphs Extended Data Fig. 3c,e.
Source Data Extended Data Fig. 6 (download XLSX )
Raw puncta quantification for the graphs Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Quantification for Fig. 7c.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Heo, D., Kim, A.A., Neumann, B. et al. Transcriptional profiles of mouse oligodendrocyte precursor cells across the lifespan. Nat Aging 5, 675–690 (2025). https://doi.org/10.1038/s43587-025-00840-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-00840-2
This article is cited by
-
The Power of Neuroglia in Driving Brain Function
Neurochemical Research (2025)