Abstract
Background
Systematic profiling of plasma proteins in population studies offers a complementary approach to discovery of novel risk factors and may provide new insights into the causes of coronary heart disease.
Methods
To explore relationships between the circulating proteome and coronary heart disease (CHD), we evaluated associations of 4780 plasma proteins with incident CHD in the Cardiovascular Health Study (CHS, N=2856, 575 CHD events) and replicated significant associations in the Atherosclerosis Risk in Communities Study (ARIC, N = 10456; 1375 events).
Results
We find that 11 proteins significantly associate with incident CHD after adjusting for risk factors; and eight significantly replicated in ARIC. Several proteins correlate with carotid intimal medial thickness and CHD associations are attenuated in participants without subclinical atherosclerosis. Macrophage metalloelastase (MMP12) is the strongest observed association (Hazard Ratio, 1.31; 95% Confidence Interval, 1.19-1.44). Mendelian randomization (MR) identifies a causal relationship between higher MMP12 and lower CHD (Odds Ratio, OR 0.94) and ischemic stroke (OR 0.90) risk, while reverse MR found that genetic propensity to CHD increased MMP12. Taken together, multivariable MR confirms a direct protective effect of higher plasma MMP12 on CHD risk and a genetic effect of atherosclerosis and CHD on elevating MMP12.
Conclusions
Proteomic analyses reveal associations with incident CHD and genomic evidence suggests that therapeutic MMP12 inhibition may confer adverse cardiovascular effects.
Plain Language Summary
Measuring many protein levels in the blood may provide new insights into the causes of heart disease (CHD). We evaluated thousands of plasma proteins in a study of older adults to determine whether any predicted risk of heart attacks or heart-related deaths and confirmed the findings in a second study. Eleven proteins showed evidence of association with CHD; often with stronger effects among those with pre-existing atherosclerosis. The strongest finding was for macrophage metalloelastase (MMP12), an endopeptidase expressed in vascular tissue and atherosclerotic plaque with roles in tissue repair, vascular remodeling, and reduction of inflammation. Our findings point to a complex role for this protein in response to underlying heart disease risk factors.
Similar content being viewed by others
Introduction
Despite the dramatic decline in morbidity and mortality associated with coronary heart disease (CHD) since the 1960s1, the rate of decline in mortality has slowed2,3 and CHD remains the leading cause of death in the US and in most of the developed world4,5. Large-scale genetic association studies have identified over 200 loci for CHD6,7,8, uncovering new biology and potential mechanisms of disease. However, many of these genetic loci lie in noncoding regions and have not been linked to individual proteins, limiting translation to new treatments.
Systematic profiling of plasma proteins in population studies offers a complementary approach to discovery and may provide new insights into the causes of CHD. Proteins are detectible in blood due to secretion or release during cell damage or turnover, and their quantification can provide a snapshot of the current state of human health9. In addition, the integration of proteomic and genomic data can help to identify proteins that may be causal factors and therefore potential therapeutic targets for CHD prevention10,11,12. An emerging literature has demonstrated the effectiveness of this approach, and novel candidate protein targets have been identified for stroke, dementia, and heart failure13,14,15. Such investigations in well-characterized individuals from population-based studies have yet to be conducted for CHD.
We first determine whether plasma proteins associate with incident CHD using an aptamer-based platform in the Cardiovascular Health Study (CHS), a prospective cohort study of cardiovascular disease in older adults. We then replicate our findings in additional community-based prospective cohort studies. Using genomic data and Mendelian randomization we next determine whether proteins associated with incident CHD are potential causal risk factors for CHD or ischemic stroke, a related cardiovascular outcome. Finally, we interpret how these findings inform our understanding of the biology of atherosclerotic cardiovascular disease by reconciling evidence from our observational study with predicted genetic levels of proteins using Mendelian randomization. In total, we identify eleven significant protein associations with CHD and leverage observational and genomic evidence to provide context for their roles in CHD biology.
Methods
Study design and participants
The Cardiovascular Health Study (CHS) is a prospective cohort study of risk factors for cardiovascular diseases in adults ages 65 years and older, who were recruited from four field centers in the United States (Sacramento, CA; Hagerstown, MD; Winston-Salem, NC; Pittsburgh, PA)16. Between June 1989 and June 1990, 5201 participants were recruited from random samples of Medicare eligibility lists, and an additional 687 Black participants were recruited between November 1992 and June 1993. Study participants were seen in the clinic annually between enrollment and 1998–1999 and were contacted by telephone at 6-month intervals through June 2023 to collect information about hospitalizations and cardiovascular events, which were adjudicated by committee through 201517.
This study was exempt from institutional review board (IRB) approval as only de-identified data was used, but the parent CHS study was approved by the institutional review boards of all participating sites (University of Washington, University of Vermont, University of Pittsburgh, Johns Hopkins University, Wake Forest University), and all study participants provided informed consent.
Plasma proteomic profiling
For this study, we set the 1992-3 clinic visit as the baseline (Supplemental Fig. 1). Blood samples were collected at each of the four field centers at baseline and processed using standard protocols. Among 5265 participants who attended, we used the SomaScan V4 assay (SomaLogic Inc, Boulder, CO) to measure relative protein concentrations from 4979 aptamers in a substudy of 3188 participants with previously unthawed EDTA-plasma collected at the 1992–1993 examination.
The SomaScan assay uses chemically modified single-stranded DNA oligonucleotides, called aptamers, that bind tightly and specifically at a ratio of 1:1 with a target protein. Aptamer levels are quantitated by a DNA oligo-array plate reader and expressed as relative fluorescent units, and these measurements reflect the concentrations of target proteins within a sample18,19,20,21. In CHS, the median intra-assay coefficient of variation (CV) from calibrator samples was 3.4% (10–90th percentiles: 1.6–7.6%) and the median inter-assay CV from quality control samples was 4.4% (10–90th percentiles: 2.6–10.1%)22.
Coronary heart disease
The outcome incident CHD included physician-adjudicated incident myocardial infarction (MI) or death from CHD. Participants with prevalent MI at the 1992-93 examination (analytic baseline) were excluded. CHS subjects were contacted every 6 months by either an in-person interview or by phone to assess risk factors and occurrence of events or hospitalizations. Discharge summaries and diagnoses were obtained for all hospitalizations, and all potential CHD events were reviewed and classified by an adjudication committee using an algorithm including cardiac symptoms as chest pain, abnormal cardiac enzyme concentrations, and serial ECG changes. Fatal CHD included deaths not meeting criteria for myocardial infarction if occurring within 72 h of chest pain or with previous history of ischemic heart disease23. Surveillance and adjudication methods in CHS were adapted from those developed in the Atherosclerosis Risk in Communities Study (ARIC), which serves as validation, and adopted by the Jackson Heart Study (JHS), which serves as an additional validation population, ensuring consistency of definitions across the three studies. Events follow up extended through June 30, 2015. Participants were censored at the end of events follow up or at the time of a non-CHD death.
Covariates
All covariates were measured at the 1992–1993 visit. Current cigarette smoking status was self-reported on a questionnaire in which participants described their smoking habits and categorized themselves as current, former, or never smokers. Prevalent diabetes mellitus was defined as a fasting blood glucose measurement \(\ge\)126 mg/dL, a non-fasting blood glucose measurement \(\ge\)200 mg/dL, or the use of glucose-lowering treatment, and treated hypertension was defined as a self-reported physician diagnosis of hypertension plus the use of an anti-hypertension medication24. Prescription medication use was assessed by the inventory method while non-prescription medication use was self-reported on a questionnaire16. Two seated, resting blood pressure measurements were obtained using a random-zero sphygmomanometer according to a standard protocol, and the average was recorded for analyses25. Body mass index (BMI) was calculated from measured height and weight. Fasting plasma concentrations of total cholesterol, high-density lipoprotein cholesterol, and triglycerides were performed on an Olympus Demand system (Olympus Corp., Lake Success, NY), and low-density lipoprotein cholesterol (LDL-C) was calculated according to the Friedewald equation16,26,27. Glomerular filtration rate (eGFR) was estimated from serum creatinine and cystatin C levels using the Chronic Kidney Disease Epidemiology Collaboration formula28. Concentrations of high-sensitivity cardiac troponin T (hs-cTnT) and N-terminal pro-B-type natriuretic peptide (NT-proBNP) were measured on an Elecsys 2010 analyzer (Roche Diagnostics, Indianapolis, IN)29,30. Serum concentrations of C-reactive protein (CRP) were measured by an enzyme-linked immunosorbent assay (ELISA) developed at the CHS central blood laboratory31, and serum concentrations of interleukin-6 (IL-6) were measured using the Quantikine HS Human IL-6 Immunoassay (R&D Systems, Minneapolis, MN)32. Ankle-brachial index and duplex ultrasonography of the carotid arteries were performed as previously described16.
Our analyses of incident CHD were restricted to those with complete covariate data after QC and no history of myocardial infarction at the time of the blood draw (N = 2856).
Protein associations with incident CHD
Plasma protein concentrations were log2 transformed and standardized to a mean of zero and a standard deviation of one, and associations of protein concentrations with time to incident CHD were evaluated using Cox proportional hazards regression. Each protein was modeled separately, adjusting for clinic site, age, sex, race/ethnicity, current smoking status, prevalent diabetes mellitus, treated hypertension, systolic blood pressure, diastolic blood pressure, BMI, plasma LDL-C concentration, and eGFR. Participants with missing values for diabetes (n = 5), treated hypertension (n = 1), systolic blood pressure (n = 1), diastolic blood pressure (n = 3), BMI (n = 18), or LDL-C (n = 57) were excluded from analyses. Analyses were repeated restricting follow-up time to ten years and to five years to evaluate protein associations with events occurring more temporally proximal to study baseline. A two-sided p-value of 2.1 × 10-5, Bonferroni-corrected for the effective number of independent tests, as defined by the number of principal components that explained 99% of the variance (n = 2377) in protein concentrations, was considered statistically significant. Significant proteins were carried forward to secondary analyses, replication, and genetic analyses.
In secondary analyses, we additionally adjusted for levels of established cardiac (hs-cTnT and NT-proBNP) and inflammatory (CRP and IL-6) biomarkers measured by conventional immunoassays to determine whether proteomic associations changed after accounting for clinical assays. We repeated primary analyses in sex and race subgroups. Using the ankle-brachial index, carotid artery stenosis, and carotid wall thickness to define subclinical atherosclerosis based on established CHS criteria33, we evaluated protein associations with incident CHD in subgroups defined by the presence of prevalent subclinical atherosclerosis at baseline. Since carotid intima media thickness (IMT) is a measure of atherosclerosis burden, we further evaluated cross-sectional associations of significant proteins with common carotid and internal carotid artery IMT measured by duplex ultrasonography.
Replication analyses in the Atherosclerosis Risk in Communities and the Jackson Heart Study
Atherosclerosis Risk in Communities (ARIC)
ARIC is an ongoing, community-based cohort study that enrolled 15,792 adults ages 45–64 between 1987–1989 from four communities in the United States (Washington County, MD; Forsyth County, NC; suburbs of Minneapolis, MN; Jackson, MS)34. Study participants completed a baseline clinic visit and six follow-up clinic visits through 2018–2019. Plasma protein concentrations for our analyses were measured using the SomaScan V4 assay. The ARIC study protocol was approved by institutional review boards at each participating institution: University of North Carolina at Chapel Hill, Chapel Hill, NC; Wake Forest University, Winston-Salem, NC; Johns Hopkins University, Baltimore, MD; University of Minnesota, Minneapolis, MN; and University of Mississippi Medical Center, Jackson, MS. All participants provided written informed consent at each study visit.
Incident CHD, a composite outcome of incident non-fatal or fatal MI or death from CHD, was determined by physician-adjudication, and plasma protein concentrations were log2 transformed and standardized to a mean of zero and a standard deviation of one.
We first replicated proteins found to be significantly associated with CHD in the CHS discovery analyses among ARIC participants with plasma protein measurements at the 1990–1992 clinic visit. Each analysis excluded participants with prevalent MI at the time of proteomic profiling, those whose samples failed proteomics QC, and those who were missing key covariates as well as non-Black and non-White participants as well as the black participants from Minneapolis and Washington County. After exclusions, 10,456 participants were included in the analyses, among whom 1375 experienced CHD during follow-up.
Cox proportional hazards regression models adjusted for clinic site, age, sex, race/ethnicity, current smoking status, prevalent diabetes mellitus, treated hypertension, systolic and diastolic blood pressure, BMI, LDL-C, and eGFR. A p value < 0.05 / number of proteins was considered statistically significant.
Jackson Heart Study (JHS)
JHS is an ongoing, community-based cohort study that recruited 5306 non-institutionalized African Americans in 2000-04 from the Jackson, MS metropolitan statistical area35. Using the SomaScan 1.3k assay (SomaLogic, Boulder, CO), plasma protein concentrations were measured in 2140 study participants from EDTA-plasma samples collected at the 2000-04 examination, and a subset of 570 participants also had plasma protein concentrations measured using the Olink Explore 1536 panel (Olink Proteomics AB, Uppsala, Sweden)36,37. The JHS study was approved by Jackson State University, Tougaloo College, and the University of Mississippi Medical Center IRBs, and all participants provided written informed consen.
Incident CHD (probable or definite non-fatal or fatal MI or death from CHD) and incident coronary revascularization were determined by physician adjudication. For these analyses, this composite outcome was determined using events ascertained through 2014. Plasma protein concentrations measured by either platform were log2 transformed and standardized to a mean of zero and a standard deviation of one.
We validated associations of significant proteins with time to incident CHD or incident coronary revascularization in two separate subgroups of participants: (1) those with with SomaScan or (2) those with Olink proteomic profiling using Cox proportional hazards regression. Participants with prevalent MI or prior revascularization at the 2000–2004 examination and those whose samples failed QC were excluded from analyses, and models adjusted for age, sex, current smoking status, prevalent diabetes mellitus, systolic and diastolic blood pressure, BMI, LDL-C, and eGFR. After QC and exclusion for missing covariates, 1772 participants (N = 118 CHD cases) were included in the SomaScan analysis and 491 (N = 95 CHD cases) were included in the Olink analysis. A p-value < 0.05 / number of proteins was considered statistically significant.
Mendelian randomization
We used Mendelian randomization to evaluate causal associations of top protein findings with CHD and ischemic stroke. We calculated F-statistics to determine the strength of the genetic instruments, and we classified instruments with F-statistics <10 as weak. For univariable MR, causal effects of each protein on outcomes were estimated by the debiased inverse variance weighted (dIVW) method38 using a three-sample design. When multiple independent cis-pQTLs were available for a protein, sensitivity analyses were conducted using the MR-Egger, weighted-median method, simple mode-based estimate, and weighted mode-based methods. All analyses were performed using the mr.divw (the GitHub version released on September 9, 2023) and TwoSampleMR (version 0.5.6) packages in R39, and a p-value < 0.05 Bonferroni-corrected for the number of tests performed in each set of analyses was considered statistically significant.
Forward Mendelian randomization
Specifically, for MR of protein concentrations (exposure) we first selected SNP cis-pQTLs among 7213 European ancestry participants of the ARIC study40 (associated with exposure at P\(\le 5\times {10}^{-8}\), independent as defined by LD \({r}^{2} > 0.001\) in 1000 G EUR population and distance > 10 mb). Weights for those SNPs were effect estimates from analyses of 35,559 Icelanders from the Icelandic Cancer Project and deCODE genetics41. Both studies measured relative protein concentrations using the SomaScan V4 assay. Genetic instruments were identified (F-statistics \(\ge \,\)95 for all proteins) for six of the eleven significant proteins: GDF15 (rs1058587, rs74953716); LEAP2 (rs60545802); MMP12 (rs2276109, rs72987587); NPPB (rs198379); REG3A (rs114874655, rs72913277); and VOPP1 (rs117042408).
For prevalent CHD (outcome), our primary analysis used summary GWAS meta-analysis results bringing together 181,522 cases among 1,165,690 participants of predominantly European ancestry participants from multiple studies, UK Biobank, and the CARDIoGRAMplusC4D consortium8. Mendelian randomization analyses of ischemic stroke used GWAS results from MEGAStroke42.
Reverse Mendelian randomization
We conducted reverse Mendelian randomization analyses to estimate effects of CHD (exposure) on proteins concentrations (outcome), using GWAS results for prevalent CHD from multiple studies, UK Biobank, and the CARDIoGRAMplusC4D consortium8 and pQTLs from a study of 35,559 Icelanders from the Icelandic Cancer Project and deCODE genetics41. The SNPs for CHD were selected based on the primarily European GWAS of CHD (associated with exposure at P\(\le 5\times {10}^{-8}\), independent as defined by LD \({r}^{2} > 0.001\) in 1000 G EUR population and distance > 10 mb, excluding SNPs within ±1 mb of each protein tested). Primary analyses estimated effects of CHD on proteins were estimated by the dIVW method. To evaluate the robustness of the findings additional methods were used, including the weighted median, weighted mode, simple mode, MR Egger and inverse variance weighted tests. (Error! Reference source not found).
Multivariable Mendelian randomization
The SNPs for cIMT were selected based on a European GWAS of maximum cIMT43 (associated with exposure at P\(\le 5\times {10}^{-8}\), independent as defined by LD \({r}^{2} > 0.001\) in 1000 G EUR population and distance > 10 mb and excluding SNPs within ±500 kb of the protein). Multivariable Mendelian randomization results were obtained from the multivariable dIVW approach using the mr.divw R package44.
Gene Expression, Protein Annotation
We examined gene expression of significant proteins in 54 non-diseased tissue sites across nearly 1000 individuals from the Genotype-Tissue Expression project45. Using previously identified cis-pQTLs37,40,41,46,47,48,49, we also used the OpenGWAS database to identify associations of cis-pQTLs with non-transcriptomic and non-proteomic phenotypes at a p-value < 5 × 10−850. The OpenGWAS database was queried on October 18, 2022 using the ieugwasr (version 0.1.5) package in R.
Statistics and Reproducibility
Primary analyses were conducted using Cox proportional hazards models in R. The discovery sample size was 2856 participants (575 of whom experienced the primary outcome of incident CHD). Significant associations were those with two-tailed p-values less than 2.1 × 10-5 was considered significant. Discovery associations were tested in a replication sample that included 10,456 participants (1375 CHD events); two-tailed p-values less than 0.05/number of tests were considered significant.
Results
In CHS we used the SomaScan V4 assay (SomaLogic Inc, Boulder, CO) to measure relative plasma protein concentrations (4985 aptamers, 4,780 unique proteins) at the 1992–1993 examination, which served as the baseline visit for this study. Our analyses were limited to the 2856 participants with proteomic data and complete covariates who had not experienced a prior myocardial infarction (MI) at the time of blood collection. In the analytic sample, the mean age was 74 years (age range: 65–98); 16% were Black, 62% were female, 10% were current smokers, and 38% had treated hypertension (Supplemental Table 1). Participants were seen in the clinic annually until 1998–1999 and were contacted by telephone at 6-month intervals through June 2023 to collect information about hospitalizations and cardiovascular events, which were adjudicated by committee through 201523. During a median follow-up of 12.5 years, 575 (20%) experienced a physician-adjudicated51 incident CHD event (366 incident MI, 209 CHD death).
Protein associations with CHD
In primary analyses, relative levels of ten proteins were significantly associated with incident CHD after adjustment for baseline characteristics and CHD risk factors (age, sex, self-described race/ethnicity, clinic site, self-reported current smoking, prevalent diabetes mellitus, treated hypertension, systolic blood pressure, diastolic blood pressure, BMI, plasma LDL-C concentration, and eGFR) and correcting for the effective number of proteins22 tested (p < 2.1 × 10-5) (Fig. 1A). When follow-up was restricted to 5 or 10 years, an eleventh protein, palmitoleoyl-protein carboxylesterase (NOTUM), reached statistical significance. (Fig. 1B, Supplemental Table 2). The most strongly associated protein was macrophage metalloelastase (MMP12); each SD higher protein concentration was associated with a higher hazard ratio (HR) of 1.31 (95% CI: 1.19–1.44) and associations were stronger for MMP12 in analyses of shorter follow-up time. Other significant proteins included known cardiac biomarkers NT-proBNP (NPPB, HR: 1.29; 95% CI: 1.18–1.42), cardiac troponin T (TNNT2, HR: 1.20; 95% CI: 1.12–1.29) and growth/differentiation factor 15 (GDF15; HR: 1.26; 95% CI: 1.14–1.40), as well as proteins including liver-expressed antimicrobial peptide 2 (LEAP2) and CREB-binding protein (CREBBP). All associated proteins passed QC and all relative levels of protein abundance observed in our study samples were above the median observed in buffer samples. Additional adjustment for established clinical protein biomarkers (high-sensitivity cardiac troponin T, N-terminal pro-B-type natriuretic peptide, C-reactive protein, and interleukin-6) measured using conventional immunoassays did not substantially attenuate associations for the other proteins (Fig. 1C, Supplemental Table 3). There was little evidence of effect modification by sex (Supplemental Table 4) or race/ethnicity (Supplemental Table 5).
Plots show associations from Cox regression analyses of incident CHD among 2856 CHS participants. (A) Volcano plot of hazard ratios and -log P values for associations of each protein with incident CHD, adjusted for clinic site, age, sex race/ethnicity, current smoking status, prevalent diabetes mellitus, treated hypertension, systolic blood pressure, body mass index, low-density lipoprotein cholesterol, and estimated glomerular filtration rate. The dotted red line is the threshold for statistical significance. Panels (B) – (D) show hazard ratios and 95% confidence intervals from secondary analyses. Panel B compares primary analyses to those restricting follow-up time to ten and five years (N = 2856). Panel C displays additional adjustment for established clinical assays (N = 2541) and inflammation biomarkers (N = 2291). Panel D compares the primary analyses to subsets of the cohort with (N = 1488) or without (N = 1233) subclinical atherosclerosis at baseline.
In contrast, we found that associations for several proteins differed by the presence of subclinical atherosclerosis at baseline, as defined using established CHS criteria33 on the basis of ankle-brachial index, carotid artery stenosis, and carotid intima media thickness (cIMT). For several proteins, associations were attenuated in the subgroup with no subclinical atherosclerosis. For instance, the HR for MMP12 was 1.43 (95% CI: 1.30–1.55) in the subgroup with subclinical atherosclerosis and was 1.02 (95% CI: 0.87–1.20) in the subgroup without subclinical atherosclerosis (p-value for interaction, 0.007; (Fig. 1D, Supplemental Table 6). Each of the attenuated proteins was also associated with common and internal cIMT, with direction of association consistent with the incident CHD findings. (Supplemental Fig. 2).
Replication in the Atherosclerosis Risk in Communities Study
We attempted to replicate significant protein associations in the Atherosclerosis Risk in Communities (ARIC) Study. Among 10,456 ARIC participants with plasma proteomics measurements from the 1990-92 visit (mean age 57 years, 58% female, 23% Black), 1,375 (13%) had an incident CHD event during a mean (SD) follow-up time of 21.3 (7.9) years (Supplemental Table 7). Among the eleven proteins significant in CHS (from the primary analysis or those with restricted follow-up), eight were significantly associated with incident CHD after correction for multiple comparisons, including MMP12 (HR: 1.35; 95% CI: 1.27–1.43) and GDF15 (HR: 1.34; 95% CI: 1.26–1.43) (Table 1).
Characterization of Proteomic Associations in the Jackson Heart Study
We further sought to characterize significantly-associated proteins in the Jackson Heart Study (JHS), a prospective cohort study of Black adults. SomaScan 1.3 proteomics were available for 1772 participants (of whom 118 experienced a CHD event). Another 491 participants (N = 95 CHD cases) had plasma protein measurements using the Olink Explore 1536 platform (Supplemental Table 8). Of the five significant CHS proteins that were measured in JHS, three (NPPB, GDF15, and REG3A) were significantly associated with the composite outcome of incident CHD (a composite of probable or definite non-fatal or fatal MI or death from CHD) or coronary revascularization (Supplemental Table 9).
Protein validation
We examined published correlations of protein levels measured among 1514 Icelandic individuals using both the SomaScan V4 and Olink Explore 3072 platforms52. Four significant proteins from the primary analysis were available on both platforms; among them the Spearman correlations were high, ranging from 0.81 (GDF15) to 0.90 (NPPB) (Supplementary Data 3). Next, we identified whether cis-pQTLs were identified from seven previous genome-wide association studies of SomaScan protein levels37,40,41,46,47,48,49. and identified 80 conditionally independent cis-pQTLs for eight of the eleven significant proteins (Supplementary Data 4).
Mendelian randomization
Mendelian randomization (MR) was used to evaluate potential causal associations of top protein findings with CHD using a “three sample” debiased Inverse Variance Weighting (dIVW) approach38. Genetic instruments for protein concentrations, available for six of eleven proteins, were selected from cis-pQTLs among 7213 European ancestry participants of the ARIC Study40 and weighted using effect estimates from analyses of 35,559 Icelanders from the Icelandic Cancer Project and deCODE genetics41. For the outcome of CHD, which were largely prevalent cases, we used summary GWAS results of 181,522 cases among 1,165,690 participants of predominantly European ancestry participants8. We found evidence of a causal relationship between higher levels of MMP12 and a lower risk of CHD (OR: 0.97; 95% CI: 0.95–0.99), which is the opposite direction of the observational association (Table 2). In a previous publication we found that higher plasma levels of MMP12 were associated a higher risk of incident ischemic stroke, another atherosclerotic cardiovascular disease53, therefore we also performed MR of MMP12 on this outcome using summary GWAS results from MEGAStroke42. As with CHD, we also found that higher levels of MMP-12 were causally related to a lower risk of ischemic stroke (OR: 0.89; 95% CI: 0.86–0.93) (Supplemental Table 10).
Reverse Mendelian randomization
We then conducted reverse MR analyses to estimate effects of CHD (exposure) on protein concentrations (outcome), using GWAS results for prevalent CHD from CARDIoGRAMplusC4D8 and pQTLs from a study of 35,559 Icelanders from the Icelandic Cancer Project and deCODE genetics41. (Supplemental Table 11) Using up to 159 independent genetic instruments for CHD, we found evidence for a causal association between CHD (exposure) and four of the significant proteins (outcomes). All four associations were directionally concordant with the observational results: higher genetically-proxied CHD risk was associated with higher levels of ARL5B (B = 0.052), MMP12 (B = 0.066), and NPPB (B = 0.056), and lower genetically-proxied CHD risk with lower levels of VOPP1 (B = -0.068) (Fig. 2, Supplemental Fig. 4). In other words, the genetic liability to CHD, which encompasses various biological processes that lead to this complex phenotype, results in upregulation of MMP12 and other proteins.
X-axis shows the observed hazard ratios for associations of each protein with incident CHD in CHS, adjusted for clinic site, age, sex race/ethnicity, current smoking status, prevalent diabetes mellitus, treated hypertension, systolic blood pressure, body mass index, low-density lipoprotein cholesterol, and estimated glomerular filtration rate; Y-axis shows the beta estimate from forward (left panel) or reverse (right panel) Mendelian randomization analysis of protein levels for cis-pQTLs.
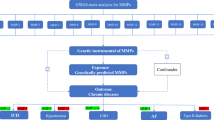
To attempt to reconcile the apparently conflicting evidence from observational and genomic data, we employed a combination of univariable and multivariable Mendelian randomization (MVMR)38 to estimate causal directions and effects between MMP12 levels and CHD. We also considered the causal effects of cIMT, a known marker of subclinical atherosclerosis that precedes CHD54. This technique used genetic instruments for MMP12 and cIMT to determine whether MMP12 has a direct effect on CHD, and whether MMP12 is a confounder or a mediator of the well-established causal cIMT-CHD relationship55,56. As combined in the MVMR framework, we showed that (1) MMP12 has a direct protective effect against CHD, independent of the development of subclinical atherosclerosis; and (2) subclinical atherosclerosis and clinical CHD causes elevated MMP12 plasma levels. (Fig. 3).
A causal diagram shows multivariable mendelian randomization (MVMR) estimates of associations of genetically proxied MMP12 levels with risk of CHD after incorporating effects of genetically proxied cIMT with MMP12 levels and CHD. B shows results from univariate and multivariable MR for each of these phenotypes. P-value is the two-sided value from the Wald test corresponding to the MR method used to estimate the causal effects.
Annotation of significant proteins
To better understand potential biological roles for the associated proteins we examined gene expression in 54 non-diseased tissue sites in the Genotype-Tissue Expression project45. We observed that genes encoding several proteins (DNJB9, CREBBP, and VOPP1) were highly expressed across most GTEX tissues, while known cardiac biomarkers NPPB and TNNT2 were selectively upregulated in the left atrial appendage and left ventricle (Supplemental Fig. 3). Using previously identified cis-pQTLs37,40,41,46,47,48,49, we also used the OpenGWAS database50 (queried October 18, 2022) to identify associations of cis-pQTLs with published phenotypes using ieugwasr (version 0.1.5) package in R. These phenome-wide association analyses identified associations between 17 cis-pQTLs in five proteins and 87 traits at P < 5 × 10−8 (Supplementary Data 5). Several proteins had cis-pQTLs that associated with potentially relevant cardiovascular traits: the cis-pQTL for NPPB was associated with systolic and diastolic blood pressure, hypertension, and anti-hypertension medication; the cis-pQTL for NOTUM with hemoglobin A1c and pulmonary function; the cis-pQTL for LEAP2 with triglyceride and insulin-like growth factor 1 levels; and cis-pQTLs for GDF15 with immune cell subsets and anthropometric measurements.
Discussion
We performed proteomic profiling in a cohort of Black and White older adults and found eleven proteins significantly associated with risk of incident CHD, independent of established CHD risk factors, eight of which replicated in an independent cohort. The most strongly associated protein was MMP12, replicating a finding observed in previous studies46,57,58,59,60,61. In contrast with the observational evidence, our MR analyses suggest that higher genetically-determined levels of MMP12 protect against the development of CHD through atherosclerosis-independent mechanisms, while atherosclerosis and CHD results in elevated plasma levels of MMP12. Taken together, proteomic profiling in observational cohorts and genomic evidence from external studies helped to inform our understanding of the biology of atherosclerotic cardiovascular disease.
MMP12, matrix metalloelastase (also known as matrix metallopeptidase 12), is an endopeptidase expressed in vascular tissue and atherosclerotic plaque that degrades the extracellular matrix and has roles in tissue repair, vascular remodeling, and reduction of inflammation. Our findings for MMP12 align with previous human observational studies, which found that MMP12 levels are elevated in the setting of carotid atherosclerosis and vulnerable plaques57, and associated with a greater risk of subclinical atherosclerosis58, peripheral artery disease62, stroke53, combined CVD63, recurrent cardiovascular events59, and heart failure15. We contribute to this growing literature by replicating prior findings46 derived from Somascan technology, as well as others that observed similar associations using Olink’s Proximity Extension Assay58,60,61,62,63.
We observed a stronger magnitude of association for MMP12 (and other associated proteins NPPB, and NOTUM) when analyses were restricted to a shorter follow-up period, highlighting their potentially short-term role in signaling the onset of clinical CHD. We then attempted to better understand the role of perturbations of the plasma proteome in the development of CHD by considering measures of subclinical atherosclerosis available in our cohort. Here we found that several significant proteins for CHD were mirrored in magnitude by associations with cIMT, a known risk factor in the development and progression of CHD. Further, we observed that associations of MMP12 (as well as NPPB and TNNT2) with CHD were attenuated among those without subclinical disease at baseline. Taken together, these findings raised the possibility that elevated levels of CHD-associated proteins might be a result of atherosclerosis rather than a direct cause of CHD.
Genomic data and MR supported this hypothesis, revealing that among six significant proteins from our observational analysis with valid cis-pQTLs, only MMP12 showed evidence of a causal effect on CHD risk. In contrast, reverse MR found that increases in plasma protein levels often result from genetic predisposition to CHD, and for all these proteins the directions of the reverse MR associations aligned with the observational associations. Whereas higher observed baseline MMP12 levels associated with higher incident CHD risk, higher genetically-predicted MMP12 levels associated with lower CHD risk. These findings mirror those from other studies which have also observed an inverse genomic association for cardiovascular disease46, ischemic stroke58,64, large artery ischemic stroke13, atrial fibrillation15, peripheral artery disease61, and heart failure15.
The biological mechanisms by which MMP-12 impacts the development of cardiovascular disease are not well understood. Some animal model studies suggest a protective role for MMP12 in the early stages of atherosclerosis65 and healing after ischemia in the post-MI setting66, but others have found potentially harmful, context-dependent roles in promoting atherosclerosis67. To advance our understanding of the function of MMP-12, we used univariable and multivariable MR to disentangle “indirect” causal effects of MMP-12 that lead to CHD through atherogenesis from “direct” causal effects that are atherosclerosis-independent. These analyses demonstrated that MMP12 has a protective role against the development of both CHD and ischemic stroke, likely independent of the development of subclinical atherosclerosis, and that the processes leading to atherosclerosis and CHD result in increased expression or release of MMP12 in the blood. This may represent a previously unrecognized negative feedback loop with MMP-12 and atherosclerosis, similar to the release of natriuretic peptides in the setting of cardiac dysfunction, which has beneficial effects on lowering blood pressure and intravascular volume68.
Other replicated associations included proteins in innate immunity pathways (REG3A; LEAP2), apoptosis (DNJB9, VOPP1), glucocorticoid receptor pathways (ARL5B), and cytokine response to cellular injury and overall stress responses (GDF15). We were able to further evaluate associations of five proteins among a cohort of African Americans where we found evidence that four (NPPB, TNNT2, REG3A, and GDF15) generalized as CHD risk markers.
While these proteins were not supported by MR as causal associations with CHD, annotation suggested potential roles in the development of CHD. Some genes encoding these proteins are highly expressed in cardiac and other relevant tissues, and phenome-wide analyses of cis-pQTLs for these proteins highlighted potential causal roles in relation to hyperglycemia, triglyceride levels, pulmonary function, and immune activation.
Although several of the proteins identified in this study are “druggable”, the MMP12 example suggests that proteomic findings alone are insufficient to guide drug development. The observation that MMPs are dysregulated in a variety of pathological conditions has led to interest in these proteins as therapeutic targets for cardiovascular and pulmonary diseases, as well as cancer treatment69,70, Early-stage trials of MMP12 inhibition have been conducted in the setting of chronic obstructive pulmonary disease, cystic fibrosis (AZD-1236), and asthma (FP-025, NCT03858686). The key finding from our study—that elevated MMP12 levels are a consequence rather than a cause of atherosclerosis, and that lowering levels of MMP12 may have a direct causal effect on the development of CHD—raise the possibility that long-term therapeutic inhibition of MMP12 may result in unintended adverse cardiovascular events. Because regulatory approval of therapeutics is increasingly based on trials with short-term follow up and sometimes only hundreds of patients exposed to an investigational drug, such adverse effects may not be observed until after many years of widespread use.
There are limitations to our study. Findings from the observational cohorts may not generalize to settings other than those studied (e.g., younger adults). Despite the prospective nature of our study and adjustment for participant characteristics, we cannot eliminate the possibility of selection bias and confounding. Our study conducted affinity-based proteomic profiling in EDTA plasma, so we were unable to investigate tissue-specific expression, isoforms, or post-translational modifications. To avoid false positive associations, we chose a stringent statistical significance threshold that corrected for the effective number of tests; as such, false negatives remain possible. In addition, the proteomic platform used assays only a fraction of the detectable plasma proteome, and many of the included protein targets have not been verified empirically or with orthogonal techniques. We attempted to interrogate this possibility by referencing correlations between proteins measured with orthogonal techniques and by investigating cis-pQTLs to focus our discussion on true protein differences, but these methods cannot fully substitute for direct laboratory validation. Our ability to investigate the causal role of several plasma proteins in CHD was limited by our ability to genetically proxy protein levels. Therefore, it is possible that some of the proteins studied other than MMP12 may be etiological or protective factors, and the lack of MR findings does not exclude this possibility. Lastly, the validity of the MR analyses is based on several assumptions, such as the absence of horizontal pleiotropy and epitope effects, which are difficult to test empirically. For MMP-12, the use of cis-pQTLs to instrument protein levels and the high degree of correlation in protein levels across different platforms increases confidence in our findings.
In summary, we identified and replicated several proteins that associated with incident CHD in a cohort of older adults, independent of known clinical and biomarker risk factors. Careful examination of our findings among our study population pointed to a role for several of these proteins as markers of the early stages of atherosclerosis and potential novel risk markers. Finally, use of Mendelian randomization points to a potentially causal protective role for MMP12 and helps to untangle seemingly contradictory findings between observational and genetically-proxied levels. In total, our findings point to new biology and provide a cautionary lesson about the interpretation of proteomic findings in drug development.
Cardiovascular Health Study
This research was supported by contracts HHSN268201200036C, HHSN268200800007C, HHSN268201800001C, N01HC35129, N01HC45133, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083, N01HC85084, N01HC85085, N01HC85086, 75N92021D00006, and grants U01HL080295,U01HL130114, R01HL105756, and R01HL172803 from the National Heart, Lung, and Blood Institute (NHLBI), with additional contribution from the National Institute of Neurological Disorders and Stroke (NINDS). Additional support was provided by R01AG023629 from the National Institute on Aging (NIA). A full list of principal CHS investigators and institutions can be found at CHS-NHLBI.org.
Atherosclerosis Risk in Communities Study
The Atherosclerosis Risk in Communities study has been funded in whole or in part with Federal funds from the National Heart, Lung, and Blood Institute, National Institutes of Health, Department of Health and Human Services, under Contract nos. (75N92022D00001, 75N92022D00002, 75N92022D00003, 75N92022D00004, 75N92022D00005). SomaLogic Inc. conducted the SomaScan assays in exchange for use of ARIC data. This work was supported in part by NIH/NHLBI grant R01HL134320. The authors thank the staff and participants of the ARIC study for their important contributions.
Jackson Heart Study
JHS acknowledgement- The Jackson Heart Study (JHS) is supported and conducted in collaboration with Jackson State University (HHSN268201800013I), Tougaloo College (HHSN268201800014I), the Mississippi State Department of Health (HHSN268201800015I) and the University of Mississippi Medical Center (HHSN268201800010I, HHSN268201800011I and HHSN268201800012I) contracts from the National Heart, Lung, and Blood Institute (NHLBI) and the National Institute on Minority Health and Health Disparities (NIMHD). The views expressed in this manuscript are those of the authors and do not necessarily represent the views of the National Heart, Lung, and Blood Institute; the National Institutes of Health; or the U.S. Department of Health and Human Services.
UK Biobank
The genetic and phenotypic data used in the GWAS of incident CHD were obtained from the UK Biobank Resource under application number 52569.
Authors
Authors were additionally supported by T32HL007828 (MPH), K08HL161445-01 (UAT), and R01HL144483 (REG, RPT, BMP); R01HL149706 (JSF); R01HL142599 (JSF) from the National Heart Lung and Blood Institute. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Data availability
The association estimates underlying Fig. 1 are provided as Supplemental Data 1. The association estimates underlying Fig. 2 are provided as Supplemental Data 2. Details on accessing the participant-level data used in this manuscript are available on the websites of each participating study: CHS, https://chs-nhlbi.org/CHS_DistribPolicy; ARIC, https://aric.cscc.unc.edu/aric9/researchers/Obtain_Submit_Data; JHS, https://biolincc.nhlbi.nih.gov/studies/jhs/;UK Biobank, https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access.
References
Mensah, G. A. et al. Decline in cardiovascular mortality: possible causes and implications. Circ. Res. 120, 366–380 (2017).
Sidney, S. et al. Recent trends in cardiovascular mortality in the United States and public health goals. JAMA Cardiol. 1, 594–599 (2016).
Manemann, S. M. et al. Recent trends in cardiovascular disease deaths: a state specific perspective. BMC Public Health 21, 1031 (2021).
Moran, A. E. et al. The global burden of ischemic heart disease in 1990 and 2010: the Global Burden of Disease 2010 study. Circulation 129, 1493–1501 (2014).
Collaborators, G.C.O.D Global, regional, and national age-sex-specific mortality for 282 causes of death in 195 countries and territories, 1980-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 392, 1736–1788 (2018).
Hartiala, J. A. et al. Genome-wide analysis identifies novel susceptibility loci for myocardial infarction. Eur. Heart J. 42, 919–933 (2021).
Tcheandjieu, C. et al. Large-scale genome-wide association study of coronary artery disease in genetically diverse populations. Nat. Med. 28, 1679–1692 (2022).
Aragam, K. G. et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Nat. Genet. 54, 1803–1815 (2022).
Suhre, K., McCarthy, M. I. & Schwenk, J. M. Genetics meets proteomics: perspectives for large population-based studies. Nat. Rev. Genet. 22, 19–37 (2021).
Emdin, C. A., Khera, A. V. & Kathiresan, S. Mendelian randomization. JAMA 318, 1925–1926 (2017).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Zheng, J. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat. Genet 52, 1122–1131 (2020).
Chong, M. et al. Novel drug targets for ischemic stroke identified through mendelian randomization analysis of the blood proteome. Circulation 140, 819–830 (2019).
Walker, K. A. et al. Large-scale plasma proteomic analysis identifies proteins and pathways associated with dementia risk. Nat. Aging 1, 473–489 (2021).
Henry, A. et al. Therapeutic targets for heart failure identified using proteomics and mendelian randomization. Circulation 145, 1205–1217 (2022).
Fried, L. P. et al. The cardiovascular health study: design and rationale. Ann. Epidemiol. 1, 263–276 (1991).
Psaty, B. M. et al. Study of cardiovascular health outcomes in the era of claims data: the cardiovascular health study. Circulation 133, 156–164 (2016).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One 5, e15004 (2010).
Candia, J. et al. Assessment of variability in the SOMAscan assay. Sci. Rep. 7, 14248 (2017).
Kim, C. H. et al. Stability and reproducibility of proteomic profiles measured with an aptamer-based platform. Sci. Rep. 8, 8382 (2018).
Rohloff, J. C. et al. Nucleic acid ligands with protein-like side chains: modified aptamers and their use as diagnostic and therapeutic agents. Mol. Ther. Nucleic Acids 3, e201 (2014).
Austin, T. R. et al. Proteomics and population biology in the cardiovascular health study (CHS): design of a study with mentored access and active data sharing. Eur. J. Epidemiol. 37, 755–765 (2022).
Ives, D. G. et al. Surveillance and ascertainment of cardiovascular events. The Cardiovascular Health Study. Ann. Epidemiol. 5, 278–285 (1995).
Mittelmark, M. B. et al. Prevalence of cardiovascular diseases among older adults. The Cardiovascular Health Study. Am. J. Epidemiol. 137, 311–317 (1993).
Tell, G. S. et al. Correlates of blood pressure in community-dwelling older adults. The Cardiovascular Health Study. Cardiovascular Health Study (CHS) Collaborative Research Group. Hypertension 23, 59–67 (1994).
Kalani, R. et al. Cholesterol variability and cranial magnetic resonance imaging findings in older adults: the cardiovascular health study. Stroke 51, 69–74 (2020).
Friedewald, W. T., Levy, R. I. & Fredrickson, D. S. Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin. Chem. 18, 499–502 (1972).
Inker, L. A. et al. Estimating glomerular filtration rate from serum creatinine and cystatin C. N. Engl. J. Med. 367, 20–29 (2012).
Tehrani, D. M. et al. Trends in blood pressure and high-sensitivity cardiac troponin-T with cardiovascular disease: the cardiovascular health study. Am. J. Hypertens. 32, 1013–1020 (2019).
deFilippi, C. R., Christenson, R. H., Gottdiener, J. S., Kop, W. J. & Seliger, S. L. Dynamic cardiovascular risk assessment in elderly people. The role of repeated N-terminal pro-B-type natriuretic peptide testing. J. Am. Coll. Cardiol. 55, 441–450 (2010).
Tracy, R. P. et al. Relationship of C-reactive protein to risk of cardiovascular disease in the elderly. Results from the cardiovascular health study and the rural health promotion project. Arterioscler Thromb. Vasc. Biol. 17, 1121–1127 (1997).
Carty, C. L. et al. Interaction between fibrinogen and IL-6 genetic variants and associations with cardiovascular disease risk in the Cardiovascular Health Study. Ann. Hum. Genet 74, 1–10 (2010).
Kuller, L. H. et al. Subclinical disease as an independent risk factor for cardiovascular disease. Circulation 92, 720–726 (1995).
Wright, J. D. et al. The ARIC (Atherosclerosis risk in communities) study: JACC Focus Seminar 3/8. J. Am. Coll. Cardiol. 77, 2939–2959 (2021).
Taylor, H. A., Jr, et al. Toward resolution of cardiovascular health disparities in African Americans: design and methods of the Jackson Heart Study. Ethn. Dis. 15, S6-4-17 (2005).
Katz, D. H. et al. Multiomic profiling in black and white populations reveals novel candidate pathways in left ventricular hypertrophy and incident heart failure specific to black adults. Circ. Genom. Precis. Med. 14, e003191 (2021).
Katz, D. H. et al. Whole genome sequence analysis of the plasma proteome in black adults provides novel insights into cardiovascular disease. Circulation 145, 357–370 (2022).
Ye, T., Shao, J. & Kang, H. Debiased inverse-variance weighted estimator in two-sample summary-data mendelian randomization. Ann. Stat. 49, 2079–2100 (2021).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Zhang, J. et al. Plasma proteome analyses in individuals of European and African ancestry identify cis-pQTLs and models for proteome-wide association studies. Nat. Genet. 54, 593–602 (2022).
Ferkingstad, E. et al. Large-scale integration of the plasma proteome with genetics and disease. Nat. Genet 53, 1712–1721 (2021).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
Yeung, M. W. et al. Twenty-five novel loci for carotid intima-media thickness: a genome-wide association study in >45000 individuals and meta-analysis of >100000 individuals. Arteriosclerosis Thrombosis Vasc. Biol. 42, 484–501 (2022).
Wu, Y., Kang, H., & Ye, T. A more robust approach to multivariable Mendelian randomization. Biometrika, 112, asaf053 (2025).
Consortium, G. The genotype-tissue expression (GTEx) project. Nat. Genet 45, 580–585 (2013).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Folkersen, L. et al. Genomic and drug target evaluation of 90 cardiovascular proteins in 30,931 individuals. Nat. Metab. 2, 1135–1148 (2020).
Pietzner, M. et al. Mapping the proteo-genomic convergence of human diseases. Science 374, eabj1541 (2021).
Gudjonsson, A. et al. A genome-wide association study of serum proteins reveals shared loci with common diseases. Nat. Commun. 13, 480 (2022).
Elsworth, B. et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv (2020).
Psaty, B. M. et al. The study of cardiovascular health outcomes in the era of claims data: the cardiovascular health study. Circulation 133, 156–164 (2015).
Eldjarn, G. H. et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 622, 348–358 (2023).
Kalani, R. et al. Plasma proteomic associations with incident ischemic stroke in older adults: the cardiovascular health study. Neurology 100, e2182–e2190 (2023).
Greenland, P. et al. 2010 ACCF/AHA guideline for assessment of cardiovascular risk in asymptomatic adults: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. J. Am. Coll. Cardiol. 56, e50–e103 (2010).
Simon, A., Megnien, J. L. & Chironi, G. The value of carotid intima-media thickness for predicting cardiovascular risk. Arteriosclerosis, Thrombosis Vasc. Biol. 30, 182–185 (2010).
Polak, J. F., Szklo, M. & O’Leary, D. H. Carotid intima-media thickness score, positive coronary artery calcium score, and incident coronary heart disease: the multi-ethnic study of atherosclerosis. J. Am. Heart Assoc. 6, e004612 (2017).
Hu, W. et al. Correlations of MMP-1, MMP-3, and MMP-12 with the degree of atherosclerosis, plaque stability and cardiovascular and cerebrovascular events. Exp. Therapeutic Med. 15, 1994–1998 (2018).
Lind, L. et al. Plasma protein profile of carotid artery atherosclerosis and atherosclerotic outcomes: meta-analyses and mendelian randomization analyses. Arteriosclerosis Thrombosis Vasc. Biol. 41, 1777–1788 (2021).
Ganz, P. et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA 315, 2532–2541 (2016).
Mahdessian, H. et al. Integrative studies implicate matrix metalloproteinase-12 as a culprit gene for large-artery atherosclerotic stroke. J. Intern. Med. 282, 429–444 (2017).
Titova, O. E. et al. Plasma proteome and incident myocardial infarction: sex-specific differences. Eur. Heart J. 45, 4647–4657 (2024).
Yuan, S. et al. Circulating proteins and peripheral artery disease risk: observational and Mendelian randomization analyses. Eur. Heart J. open 3, oead056 (2023).
Lind, L. et al. Plasma protein profiling of incident cardiovascular diseases: a multisample evaluation. Circul. Genom. Precis. Med. 16, e004233 (2023).
Carcel-Marquez, J. et al. Causal Effect of MMP-1 (Matrix Metalloproteinase-1), MMP-8, and MMP-12 levels on ischemic stroke: a mendelian randomization study. Stroke 52, e316–e320 (2021).
Luttun, A. et al. Loss of matrix metalloproteinase-9 or matrix metalloproteinase-12 protects apolipoprotein E-deficient mice against atherosclerotic media destruction but differentially affects plaque growth. Circulation 109, 1408–1414 (2004).
Iyer, R. P. et al. Early matrix metalloproteinase-12 inhibition worsens post-myocardial infarction cardiac dysfunction by delaying inflammation resolution. Int. J. Cardiol. 185, 198–208 (2015).
Johnson, J. L., George, S. J., Newby, A. C. & Jackson, C. L. Divergent effects of matrix metalloproteinases 3, 7, 9, and 12 on atherosclerotic plaque stability in mouse brachiocephalic arteries. Proc. Natl. Acad. Sci. USA 102, 15575–15580 (2005).
Ichiki, T., Jinno, A. & Tsuji, Y. Natriuretic peptide-based novel therapeutics: long journeys of drug developments optimized for disease states. Biology 11, 859 (2022).
Myasoedova, V. A., Chistiakov, D. A., Grechko, A. V. & Orekhov, A. N. Matrix metalloproteinases in pro-atherosclerotic arterial remodeling. J. Mol. Cell. Cardiol. 123, 159–167 (2018).
Abd-Elaziz, K., Jesenak, M., Vasakova, M. & Diamant, Z. Revisiting matrix metalloproteinase 12: its role in pathophysiology of asthma and related pulmonary diseases. Curr. Opin. Pulm. Med. 27, 54–60 (2021).
Acknowledgements
The authors thank the other investigators, the staff, and the participants of ARIC, CHS, JHS, and the UK Biobank for their valuable contributions.
Author information
Authors and Affiliations
Contributions
Conceptualization: (K.M., B.M.P., J.C.B., and J.S.F.); Data curation: (T.R.A.); Formal analysis: (M.P.H., J.A.B., U.A.T., J.C.h., and T.Y.); Funding acquisition: (B.M.P. and J.S.F.); Project administration: (M.P.H. and T.R.A.); Resources: (R.E.G. and R.P.T.); Software: (C.M.S.); Supervision: (K.M., R.E.G., and J.S.F.); Visualization: (M.P.H. and J.C.B.); Critical Revision: (J.A.B., C.M.S., T.R.A., U.A.T., S.D., D.M., J.C.h., K.M., C.M.B., J.C.o., R.E.G., R.P.T., T.Y., and B.M.P.); Drafting Manuscript: (M.P.H., J.C.B., and J.S.F.).
Corresponding author
Ethics declarations
Competing interests
Bruce M. Psaty serves on the Steering Committee of the Yale Open Data Access Project funded by Johnson & Johnson. All other authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Anders Mälarstig and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huber, M.P., Brody, J.A., Sitlani, C.M. et al. Plasma proteomics and incident coronary heart disease. Commun Med 6, 98 (2026). https://doi.org/10.1038/s43856-025-01363-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01363-y