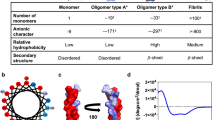
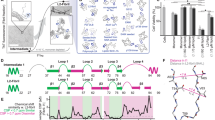
Abstract
Parkinson’s disease involves the accumulation of aggregates of ɑ-synuclein (ɑ-Syn), both as intracellular fibrils and as cytotoxic soluble oligomeric species (ɑSOs). No available nanobodies show exclusive preference for the oligomeric state of ɑ-Syn. Here, we describe two nanobodies NB1 and NB2, obtained by immunizing a llama with αSOs, which bind ɑSOs with nM affinity and do not show any measurable affinity for monomeric ɑ-Syn or ɑ-Syn fibrils. While the nanobodies were not useful for high-throughput screening for therapeutic compounds or high-resolution cryoEM, they retained their ability to discriminate against ɑ-Syn monomers in brain tissue and were able to detect ɑ-Syn aggregates in diseased tissue. In addition, αSO binding affinity was improved by DNA-scaffold-mediated NB1 dimerization compared to scaffolded monomeric NB1. The nanobodies promote the uptake of ɑSOs into HEK93 cells via the Sortilin receptor pathway. Their absolute specificity for oligomeric ɑ-Syn makes them promising reagents to detect oligomeric ɑ-Syn in patient samples.
Similar content being viewed by others
Introduction
Nanobodies are small (~13.5 kDa) protein binders derived from the variable domain of camelid heavy-chain antibodies which show antibody (Abs)-level specificity toward a broad range of targets1. This occurs via an architecture similar to Abs’ variable heavy chain, based on three highly variable loops (H1–H3). Further, thanks to their small size and convex shape, nanobodies can bind to epitopes sometimes inaccessible to conventional Abs. They show particular structural nvariation in the H3 loop1 which vary greatly in length. Long H3 loops can form finger-like protrusions to access epitope cavities and can also help shield the surface that is exposed due to the absence of the light chain, reducing aggregation propensity2. Nanobodies are reasonably stable, also in vivo, and are attractive for both therapy and diagnostics for several reasons. Unlike full-length Abs, they are generally unproblematic to express recombinantly in E. coli in large quantities and show superior abilities to penetrate tissue and cross the blood-brain barrier3 while maintaining low immunogenicity4 and avoiding Abs-dependent cytotoxicity. They are also straightforward to obtain by immunizing camelids such as alpacas and llamas, extraction of lymphocytes from blood samples, cloning into phagemids, and panning against the desired target1. However, their small size also leads to a significantly faster clearance from the blood, with half-lives in the order of a few hours, as opposed to several weeks for Abs3.
Nanobodies have also been investigated for both therapeutic and diagnostic purposes in Parkinson’s Disease, which is characterized by the accumulation of aggregates of the natively unfolded protein ɑ-synuclein (ɑ-Syn). These aggregates typically manifest themselves as intracellular inclusions known as Lewy Bodies which take up an amyloid or fibrillar structure5. In addition, ɑ-Syn forms soluble oligomeric species (ɑSOs) which are at least as cytotoxic as the fibrillar species6. ɑSOs can form in a pathway distinct from fibrillation7 and show a remarkable ability to permeabilize cell membranes8,9, making them important targets for both diagnostics and therapy. Their N- and C-terminal regions are highly dynamic10, making it difficult to obtain high-resolution structures despite intense efforts over the past decade11,12, which includes the enlistment of peptides such as PSMα3 binding to the ɑSO with high affinity12. Nevertheless, nanobodies have been developed which recognize both fibrillar and oligomeric species of ɑ-Syn. The two nanobodies NbSyn2 and NbSyn87, obtained by immunization with monomeric ɑ-Syn, both bind to the unstructured C-terminal tail of ɑ-Syn13 which is generally highly exposed and also remains unstructured in oligomers and fibrils. These two nanobodies inhibit ɑ-Syn fibrillation and reduce ɑSO cell toxicity by oligomer remodeling14. NbSyn87 is sensitive to phosphorylation of Tyr125 which decreases binding affinity ca. 400-fold15. Another set of nanobodies, sdAbs 2D10 and 2D816, also raised by immunizing with monomeric ɑ-Syn, is specific for ɑ-Syn’s NAC region (ca. residues 60–100) which form the core of both oligomers10 and fibrils17 of α-Syn. These nanobodies bind to rodent brains to an extent which correlates with the level of α-Syn pathology16. Furthermore, these two nanobodies were potentiated further by coupling them to thalidomide, an E3 ligase ligand, which allows them to enter the proteasomal pathway to clear both insoluble and soluble α-Syn18, both in primary neuron cultures and in mouse models of synucleinopathy. There are other variations on the theme of small Abs against α-Syn aggregates. Thus, disulfide-free nanobodies (able to remain folded within the reducing environment of the cytosol), selected against α-Syn fibrils in a yeast library, dissociate fibrils in vitro and reduce spread of pathology in murine neurons19, while a single-chain Ab fragment (scFv) called D5, obtained from a phage library selected against immobilized αSOs, was able to block α-Syn aggregation in vitro and reduce αSO cell toxicity in both SHSY5Y20 and HEK29321 cells. Finally, the nanobody Nb α-syn01, obtained from a camel immunized with monomeric α-Syn, binding to the stretch 42–56 in α-Syn, preferentially (but not exclusively) binds to fibrillar α-Syn in an ingenious mechanism where the peptide displaces the initial 8 N-terminal residues of the nanobody via residues 48–5222.
Among these numerous nanobodies, none show an exclusive preference for oligomeric state or even just the aggregated (fibrillar or oligomeric) state of ɑ-Syn, although this would be very useful for both therapeutic and diagnostic purposes. We reasoned that immunization with pre-formed αSOs would increase the likelihood of obtaining conformationally specific nanobodies. Here, we developed two nanobodies NB1 and NB2 which we have obtained by llama immunization and phage-display selection with αSOs. The αSO we use forms spontaneously upon incubation of ɑ-Syn under fibrillating conditions and can be obtained by size-exclusion chromatography as a stable well-defined species consisting of ~30 ɑ-Syn subunits7. Unlike the large collection of monoclonal Abs (mAbs) which we raised against a variety of different types of ɑ-Syn oligomers23, NB1 and NB2 do not show any measurable affinity for monomeric or fibrillar ɑ-Syn in vitro but exclusively bind to oligomers with nM affinity. They retain the ability to discriminate against monomeric ɑ-Syn in spiked brain tissue lysate and detect ɑ-Syn aggregates in diseased tissue. While we were unable to determine the structure of the ɑSO:NB complexes by the NabFab approach, NB1 and NB2 both promote uptake of ɑSOs into HEK293 cells via the Sortilin receptor pathway. Their absolute specificity for aggregated ɑ-Syn and preference for αSOs makes them promising reagents for the diagnosis of aggregated ɑ-Syn.
Results
Generation of two nanobodies against the α-Syn oligomer
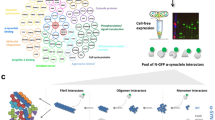
To obtain nanobodies specific for the α-Syn oligomer (αSO), a llama was immunized with 0.5 mg of αSO made from human α-Syn, recombinantly expressed in E. coli. This αSO forms spontaneously upon incubation of monomeric α-Syn while shaking at high concentrations (8–12 mg/ml) at 37 °C, and is sufficiently stable that it can be purified by size-exclusion chromatography and concentrated to ~1 mg/ml. We have previously characterized this αSO extensively in terms of structure, dynamics, stability, and functionality in membrane permeabilization and neuronal assays (summarized in refs. 6,24). Its ease of preparation and clear biological activity make it a physiologically relevant and experimentally convenient representative of α-Syn aggregate toxicity. We have used similar αSO preparation to raise mAbs against αSO in mice23. After harvesting lymphocytes from a blood sample taken from the llama, a cDNA library was obtained, from which nanobody-coding sequences were amplified by PCR and cloned into phagemids for selection by phage display against immobilized αSO. αSO specificity was enhanced by including negative selection against monomeric and fibrillar α-Syn in the phage display. The 10 phagemids obtained from the panning of this phage display library were sequenced, revealing only two unique sequences (three of one type and seven of the other), deviating from each other at seven positions (5.6% deviation) (Table 1). Most of these mutations were conservative. These two unique NB sequences, subsequently referred to as NB1 and NB2, were transferred into a BL21d vector for bacterial expression and purification by a C-terminal His-tag. Expression levels were 5–10 mg per L culture. Both proteins showed high and very similar stability, unfolding without stable intermediates with a midpoint around 65 °C and 1.95 M guanidinium chloride as measured by chemical and thermal denaturation (Fig. S1 and associated text, results summarized in Table 2).
FIDA analysis shows specific binding of NBs to αSOs
Since NB1 and NB2 were generated by immunizing llamas with αSOs prepared from recombinant wildtype α-Syn, we decided to investigate the binding of the two NBs to αSOs. Due to the dynamic nature of different regions of the αSOs (particularly the C-terminal region, residues 95–14010), we sought to avoid immobilization of the αSOs. In addition, the relatively low yields of αSOs (typically only 1–5% of monomeric α-Syn converts to oligomer7) limited us to techniques requiring low amounts of αSOs. Accordingly, we turned to a microcapillary technique in the form of Flow-induced Dispersion Analysis (FIDA), which detects changes in the hydrodynamic radius (RH) of fluorescently labeled species (e.g., through binding to non-labeled partners) through their diffusive properties25. We generated binding curves for the NB:αSO and NB:monomeric α-Syn complexes for both NB1 and NB2 by titrating either αSO or monomeric α-Syn (the analyte) against fluorescently labeled nanobodies (indicators) using a capillary mixing mode in which the nanobodies are bathed in a solution of α-Syn, allowing equilibrium to be reached. Since only the NBs are fluorescently labeled (using AlexaFluor 488), RH of free NBs could be determined from the initial measurement with no α-Syn in each experiment25. Representative examples of raw FIDA profiles are shown in Fig. S2a, b.
Increasing concentrations of αSO led to an increase in RH for both NB1 and NB2, from around 2 nm to 9.8 nm (Fig. 1a). This fits well with the size of αSO determined by FIDA (9.74 nm), transmission electron microscopy (EM) (~10 nm), and atomic force microscopy (AFM) (~10 nm)7,26. To determine binding affinity, a 1:1 model in the FIDA software was used to fit the data. The KD for NB1 was estimated at 31.3 nM, while NB2 had a KD of 41.2 nM (correlation coefficients R of 0.984 and 0.992, respectively), indicating that NB1 has a slightly higher affinity. A 1:2 binding model with 2 NB molecules per oligomer led to a R value of 0.98 while the 1:3 model scored even worse with a R value of 0.97. Therefore, we restrict ourselves to a simple 1:1 binding model. Gratifyingly, increasing concentrations of monomeric α-Syn did not lead to any change in RH when titrated into the capillary (Fig. 1a), indicating no binding between either NB1 or NB2 and monomeric α-Syn. This is consistent with our observation that the two NBs did not show any binding to α-Syn peptides in a peptide array displaying 14-mer peptides spanning the whole α-Syn sequence (Fig. S3a). Furthermore, we performed ELISA assays in 96-well plates in which α-Syn was deposited on the well bottom, followed by the addition of NB1 or NB2 and detection of bound nanobody using anti-His Abs; remarkably, only αSOs led to a significant signal, and neither monomer nor fibrillated α-Syn led to any significant binding (Fig. S3b). Thus the NBs are absolutely specific for aggregated α-Syn, in contrast to the mAbs we recently reported to favor the aggregated state but also to bind with measurable affinity to the monomeric species and to peptides from the α-Syn sequence23. The results are summarized in Table 2.
a FIDA analysis reveals a high affinity for αSOs but no detectable binding to monomeric α-Syn. Errors based on triplicates. SPR of binding of b NB1 and c NB2 to immobilized αSOs (tethered to the monoclonal antibody 18-3A5-H2, which in turn was bound to protein G on the chip surface). d Left: Western blots of binding of NB1 and NB2 to different types of αSOs and (right) Coomassie-Blue stained gels of the same αSOs. e Calcein release assays in which 1 µg/ml αSO was incubated with NB1 or NB2 in different molar ratios, followed by exposure to calcein-filled vesicles. Data normalized to complete calcein release by detergent lysis. Errors based on triplicates.
Our FIDA-binding data were further validated using SPR studies. To determine kinetic parameters, we developed a double capture assay in which the αSO was tethered on the surface via a mAb (to avoid unspecific surface interactions) and then exposed to NBs. Specifically, we used a protein G-coupled chip to capture the αSO-binding mAb 18-3A5-H223. A titration series of NBs at increasing concentration was then injected over the surfaces, and the obtained sensorgrams were fitted with a 1:1 binding model (Fig. 1b, c), resulting in KD values of 3.2 ± 0.1 nM and 31 ± 1.2 nM for NB1 and NB2, respectively. Given that FIDA measures in solution and SPR uses immobilized αSOs, it is satisfactory to see reasonably modest deviations between the two sets of results from FIDA and SPR.
Further confirmation of the specificity for this type of oligomer was obtained by Western blots, in which we used oligomers of α-Syn prepared by incubation with oxononenal (ONE), EGCG, and nitration with tetranitromethane (see “Methods”) in addition to our spontaneously formed αSOs. In all cases, only the original αSO was detected (Fig. 1d and original gel data in Fig. S13), along with a very faint band for the oligomer formed by nitration of α-Syn.
Surprisingly, neither NB1 nor NB2 had any significant effect on the ability of the αSOs to permeabilize calcein-encapsulated vesicles. There was a small decline in calcein release between 1 and 10-fold molar excess of nanobody, but the release increased at higher concentrations (Fig. 1e). Similarly, neither nanobody had any significant effect on α-Syn fibrillation monitored by ThT fluorescence (Fig. S4a, b) in terms of the rapidity of the reaction, since the half-time of fibrillation (i.e., the time needed to reach 50% of the final fluorescence level) only showed a modest decline (Fig. S4c), and this was largely due to the decline in plateau level (Fig. S4d) since the onset of fibrillation (tlag, determined as the intercept of the growth phase extrapolated to meet the x axis) did not change significantly (Fig. S4e).
The nanobodies are able to detect αSOs in brain lysates
To test how well our nanobodies could detect α-Syn under physiological conditions, we prepared brain homogenates from healthy mice brains and spiked them with different concentrations of monomeric, oligomeric, and fibrillar α-Syn. The mixtures were used to coat microwells in a 96-well plate and incubated with NB1 or NB2, after which nanobodies were detected with an anti-His antibody. Both nanobodies detected αSO very robustly but showed no signal with monomeric α-Syn (Fig. 2a, b). They showed intermediate signals with equivalent amounts of fibrillar α-Syn, although controls with pure samples of α-Syn without brain lysate showed complete restriction to αSOs (Fig. 2c). We cannot exclude that this reflects structural rearrangement of the fibrils in vitro. The nanobodies also detected α-Syn in brain lysates from mice transgenic for human A53T α-Syn (Fig. 2d) at levels only slightly lower than the commercial MJF14 antibody.
The ability of nanobodies a NB1 and b NB2 to detect different species of α-Syn spiked into mouse brain lysates. c Control experiments. Different α-Syn species were used to coat wells, after which NB1 or NB2 was added. The signal was detected using an anti-His antibody (detects nanobody) or the anti α-Syn antibody 14-9E7 A1. d Brain tissues from 4 PD mice were used to coat wells in 1:1 or 1:5 dilution, after which the anti-α-Syn antibody MJF14 or nanobody was added. Background level (0.14 units) subtracted. Errors based on triplicates.
Dimerization of NB1 using DNA scaffolds improves binding to αSO
Inspired by antibodies’ multivalency, we investigated the potential of using DNA scaffolds to dimerize NB1 and improve its binding affinity towards αSO. NB1 was conjugated onto two different DNA linkers (L1 and L2) using click chemistry and which subsequently were dimerized using scaffold strands complementary to the two linkers (Fig. 3a). To further examine whether the distance between two NB1 molecules influences binding, scaffold strands were engineered with either three (3 T) or nine thymidines (9 T), spacing the NB1-DNA linker hybridization sites ~2 nm or 5 nm apart, respectively (Fig. 3a). BioLayer Interferometry (BLI) was used to assess the specificity of scaffolded NB1 and to evaluate the avidity effect in αSO binding (Fig. 3b). The scaffolded constructs with two nanobodies exhibited higher BLI signal intensity, indicating increased binding affinity. The scaffolded NB1 constructs maintained their specificity for αSO, evidenced by the lack of binding to α-Syn monomer and fibrils. In addition, we observed 2.2–4.6-fold increase in binding affinity when comparing the scaffolds carrying two nanobodies versus one (Fig. 3c–h). The BLI-derived KD values spanned 255–581 nM for the constructs containing one NB1 and 90.5–173 nM for constructs containing two NB1. The increased affinity derived from the dimerization of NB1 was corroborated by FIDA (Fig. 3i, j), which showed KD values ranging between 128 and 254 nM for the constructs with one NB1 and 57.6–110 nM for the dimerized constructs. This represents a 1.2–4.3-fold increase in binding affinity upon dimerization. Both BLI and FIDA data validated the 3 T DNA scaffold as a more effective candidate, though it should be noted that even the dimerized NB showed lower affinity to αSO than non-conjugated NB1 (KD of 31.3 nM).
a Schematic of NB1 conjugation and hybridization. NB1 alone (Nanobody), NB1 conjugated to DNA linkers (L1 and L2) alone and hybridized to different DNA scaffold strands (T + L1, T + L2 and T + L1 + L2), containing either three thymidine (3 T) or nine thymidine (9 T) spacers. The scaffold strands were labeled (green dot) and contained either biotin for BLI or ATTO488 for FIDA applications. b BLI binding screening of NB1 against αSO (500 nM), α-Syn monomer (20 µM), and fibrils (2 mg/mL), with or without DNA conjugation. c–h BLI analysis of scaffolded and multimerised NB1 showing improved binding against αSO (0–500 nM) when dimerized. i, j FIDA analysis validating improved binding against αSO (0.500 mM) of the dimerized NB1. Data are fitted with a 1:1 binding curve, and the dissociation constant KD is provided. Errors bars based on triplicates.
Formation of αSO:NB:NabFab complexes
To aid the structural analysis of the αSOs, we decided to construct complexes between the αSOs, NBs and a synthetic Fab fragment (NabFab) binding to an epitope on the NB scaffold distant from the Complementarity Determining Regions (CDRs). This approach has been used to obtain high-resolution cryoEM structures of otherwise intractable or dynamic targets such as membrane proteins by trapping them in specific conformational states27. Similarly, Ventura and coworkers have used the αSO-binding peptide PSMα3 to obtain better-resolved structures of the αSO12. Accordingly, we constructed NabFab-binding nanobody variants of NB1 and NB2, in which the CDRs of our two NBs were grafted onto a NB scaffold that is able to bind to NabFab. This grafting procedure required us to introduce 9 (NB1) or 10 (NB2) individual mutations (Table 1) to make NBg1 and NBg2, respectively. We start by determining their affinity for NabFab. The NabFab domain showed high affinities for both nanobodies measured by FIDA, with values of 28.4 nM and 32.5 nM for NBg1 and NBg2, respectively (Fig. 4a, b). Both nanobodies exhibited slightly larger RH values than expected, both for the NB alone and in complex with NabFab (Table 3), which we attribute to slight baseline drift. A nanobody with a Mwt of ~13.5 kDa is expected to have an RH of ~2.1 nm, but we observed RH values of 2.37 nm for NBg1 and 2.45 nm for NBg2. The expected RH value for the 50 kDa complex formed with NabFab is 3.6 nm, but the measured complex sizes were 3.17 nm and 3.03 nm for NBg1 and NBg2, respectively. Nonetheless, the stable plateau observed suggests that full complex formation was achieved (see Fig. 4a, b).
Next, we tested whether the grafted nanobodies retained an affinity for αSOs, as previously observed with the original nanobodies NB1 and NB2 (Fig. 1). The KD values for complex formation with αSOs were somewhat higher for both grafted nanobodies (Fig. 4c, d). For NBg1, we measured an affinity constant of 132.5 nM, which is >fourfold weaker than the 31.3 nM for the original NB1; the difference is likely due to the grafting of the binding regions to a different NB. However, the expected size of NBg1 was confirmed at 2.1 nm, while the αSO:NBg1 complex was estimated by FIDA to be 13.6 nm, which is larger than the previously observed ~10 nm. However, there is some uncertainty in the complex size measurements, as titration plateaus were not reached due to the limited solubility of αSO. Similarly, the affinity of NBg2 for αSOs was measured at 120.7 nM, about 3-fold higher than the 41.2 nM KD of the original NB2. The size of NBg2 alone was the expected 2.0 nm, while the αSO:NBg2 complex size was the expected 10.3 nm. These results indicate that while the grafting process slightly reduces affinity, the resulting KD values are within an order of magnitude of the original NBs. Data are summarized in Table 3.
a NabFab binding to NBg1. b NabFab binding to NBg2. c NBg1 binding to αSOs. d NBg2 binding to αSOs. e NBg1:NabFab binding to αSOs. f NBg2:NabFab binding to αSOs. g SPR data of binding of NBg1:NabFab to αSOs (immobilized as in Fig. 2b, c). In all cases, data are fitted with a 1:1 binding curve and the dissociation constant KD is provided. Errors based on triplicates.
Finally, we measured the affinity of the αSOs for the two NabFab:NBg complexes. We premixed 100 nM of either NBg with 1000 nM NabFab to ensure the full complex formation and titrated this complex against αSO. For the NBg1 complex, we observed a KDapp of 350 nM (Fig. 4e), which is higher than the KD of αSO for NBg1 alone (132.5 nM). The size of the indicator molecule (NBg1:NabFab) was 3.4 nm, and the complex with αSO was 14.5 nm, larger than our estimation of the size of the complex formed by NBg1 alone. Similar results were observed for the NBg2 complex, with a KDapp of 551.5 nM (Fig. 4f), significantly higher than αSO’s affinity for NBg2 alone (120.7 nM). The size of the NBg2 complex was 3.4 nm, and the complex with αSO estimated to 14.8 nm. As with previous titrations of the NBg’s alone, titration plateaus were not achieved due to the low solubility of αSO. These results indicate that the association of the NBg’s with the NabFab domain reduces their affinity for αSO, resulting in KD values 3–4-fold greater than those of the original nanobodies. Alternatively, there could be steric competition between αSO and NabFab for binding to the nanobodies. This is not unreasonable. Although there is a distinct separation of binding sites on the nanobodies for binding to the αSO (in the CDR loops) and the NabFab (in the constant domain), the large size of αSO compared to both NB and NabFab could lead to a sphere of sterical perturbation that affects the access to other parts of NB. In this case, there would be multiple species present with distinct (but not FIDA distinguishable) RH values and with resulting KD values whose values reflect the extent of structural interference between the different sites but with affinities that can be reduced compared to the expected values, as modeled theoretically28. We investigated whether NabFab can outcompete NB for binding to αSO by titrating NabFab into a pre-formed NB: αSO complex. At the highest NabFab concentration (8 µM), we observed a modest reduction in RH from ~9.2 nm to ~8.6 nm. However, we did not detect the formation of previously observed complex sizes corresponding to NBg1 and NBg2 in the complex with NabFab (3.17 nm and 3.03 nm, respectively) (Fig. S5). These findings support the proposal that different epitopes exhibit varying affinities for NabFab.
To further investigate the binding between NBg’s and αSO, we turned to SPR. For oriented capture of NBg’s, we initially prepared a chip with immobilized NabFab. Although this surface showed a high binding capacity for NBg’s, no subsequent binding of αSO was observed, indicating that NabFab and αSO compete for the same binding site on NBg’s (not shown) which is consistent with our conclusions in the previous section. Using this chip, we instead developed an assay based on competitive binding to measure affinity of NBg1 to αSO. NBg1, at a constant concentration of 1 µg/ml, was preincubated with a titration series of αSO and injected over the NabFab surface as described in Materials and Methods. In Fig. 4g, the binding of NBg1: αSO relative to NBg1 alone is plotted as a function of the logarithm of the αSO concentration. Almost complete inhibition of NBg1 binding to NabFab is observed at 10 µM αSO. Fitting resulted in an inhibition constant (Ki) of 390 nM, i.e. the affinity of the interaction between NBg1 and αSO, determined by this assay, is 390 nM, which is only threefold greater than the FIDA-determined value of 132.5 nM.
Mass photometry suggests complex formation between αSOs, NB, and NabFab but is not accurate enough to deduce stoichiometry
To establish the stoichiometry of binding between αSO, grafted nanobodies, and NabFab, we turned to mass photometry, which uses interferometric scattering to determine molecular masses of individual particles29. Mass photometry measurements of the αSOs led to a mass of 457 kDa, which is close to the expected size of ~ 432 kDa (Fig. 5a). However, we also observed a lower mass of 92 kDa, which is possibly a buffer artifact made more prominent by the low number of sample counts. This artifact is found in all the measurements, although it somewhat varies in molecular size. The addition of the NBg1-NabFab complex to αSO led to particles with average masses of either 514 kDa or 404 kDa, when mixed in 1:3 or 1:10 ratio, respectively (Fig. 5b, c). Notably, the errors (σ) from the multi-Gaussian fittings are high, so the differences are not significant. Similarly, mixing with the NBg2-NabFab complex results in 516 kDa and 552 kDa in 1:3 and 1:10 ratios, respectively (Fig. 5d, e). Although the overlapping errors prevent us from concluding on the exact stoichiometries of binding, we can clearly see the effect of binding of the NBg’s in complex with NabFab to the αSO molecules (combined in Fig. 5f). Attempts to obtain high-resolution structures of αSO:NB:NabFab complexes were however unsuccessful (described in Fig. 6 and in Supplementary Information with Figs. S6 and S7 and associated text).
a αSOs alone followed by αSOs in complex with NBg1:NabFab at b 1:3 and c 1:10 molar ratio. d, e As in panels (b, c) with NBg2. f Superposition of the five curves in (a–e).
Left: αSOs alone show the characteristic hollow or doughnut-like structure with a central cavity. Complexes of αSOs with NBg1:NabFab (centre) and NBg2:NabFab (right) show the same general structure but lose resolution, likely due to the formation of heterogeneous populations of species. Arrows point to individual αSOs also shown in close-up (upper right corner). More details can be found in Fig. S7.
Besides these efforts, we carried out a major effort to develop a high-throughput assay to screen for compounds binding to αSOs. This was based on a sandwich strategy in which NB1, immobilized in a microplate well, captured αSOs as detected by the mAb 14-9E7-A1. This binding could potentially be disrupted by small molecules, and we screened 50,000 such compounds. The 23 best hits showed low-nM IC50 values in this assay, but their binding could unfortunately not be validated in orthogonal calcein release assays and MST measurements. This is described in more detail in Supplementary Information along with Figs. S8–11.
Uptake of nanobodies into HEK293 cells
Finally, to investigate whether our nanobodies were able to target αSOs for intracellular degradation, we turned to a HEK293 cell model for Sortilin receptor targeting. Here HEK293 cells are transfected to express the Sortilin receptor, which in turn recognizes biomolecules tagged with the tetrapeptide RQLL and internalizes them for endosome-lysosome pathway transport30,31,32. When fluorescently tagged αSOs bind to NBs tagged with a C-terminal RQLL sequence (prepared recombinantly as described in “Methods”), the RQLL sequence promotes a ternary complex with the Sortilin receptor displayed on the membrane of HEK293 cells, and this in turn mediates transport into the cells32. We quantified cellular uptake with microscopy through the fluorescence of the Alexa488-labeled αSOs. Uptake is followed by plotting cell fluorescence versus NB concentration, which ideally should present a bell-shaped profile32,33. The decline in uptake at higher concentrations, known as the hook effect, occurs through competition between free and bound NBs. That is, higher concentrations of the RQLL-tagged NBs lead to the binding of free NBs (not bound to αSOs) to Sortilin, thus blocking the uptake of the αSO-bound RQLL-NBs.
Accordingly, HEK293 cells were exposed to a constant concentration of αSOs preincubated with increasing concentrations of RQLL-NBs. While both NB1 and NB2 lead to the uptake of αSOs into the cells (Fig. 7a), there were notable differences. NB1 led to slightly higher fluorescence intensity than NB2 but without completing the bell-shaped profile at the available NB concentrations. Furthermore, both NBs showed significantly lower levels of fluorescence uptake than mAbs raised against αSOs (Fig. 7b). The mAb shown in Fig. 7b shows the highest uptake around 10−7.1 M (8 × 10−8 M) mAb, while the corresponding number for NB2 is one magnitude higher (8 × 10−7 M). Given that mAbs are ~12 times larger than NBs, the two ligands are comparable in efficiency on a per-mass basis, but the mAb also shows a ~ 12-fold higher fluorescence intensity, suggesting that the mAb is better able to target αSOs for intracellular degradation.
a NB1 and NB2 both facilitate uptake of a certain amount of αSOs into cells but b the effect is about 12-fold weaker in terms of internalized fluorescence signal compared to a monoclonal antibody binding to αSOs. Errors based on triplicate measurements.
Discussion
Nanobodies raised against the α-Syn oligomer are specific for the oligomer
We undertook this study to raise nanobodies specifically against the oligomeric species of α-synuclein, which to the best of our knowledge had not been used before to immunize llamas and generate VHH antibodies from which nanobodies could be obtained. We only obtained two different nanobody sequences; a similarly low number of hits has also been observed in other studies in which monomeric α-Syn was used as antigen13,16. Both our nanobodies showed respectable levels of expression and thermodynamic stability and were therefore subjected to additional analysis. The most striking and attractive feature of the nanobodies is their utter specificity for oligomeric α-Syn and their inability to bind either the monomeric species, the fibrillated state, or individual peptides corresponding to parts of the α-Syn primary structure. This indicates that binding is not a simple avidity effect (i.e., the binding to multiple copies of the same linear epitope in close proximity in homo-oligomeric aggregates). Avidity is in any case not expected to play a significant role in nanobodies binding to their antigens, due to the presence of a single binding site on each nanobody, in contrast to the two arms found on a conventional mAb (as well as Fab and ScFv constructs). We note, however, that other nanobodies raised against monomeric α-Syn showed affinity for specific peptides in the α-Syn sequence; thus 2D10 and 2D8 target peptides 71–85 and 81–9516. While the other NBs Nbsyn87 and NbSyn2 show nM affinity for monomeric α-Syn (KD values of 14 and 130 nM, respectively13), their affinity for αSOs was not demonstrated. We believe that our measurements are the first direct reports on nanobodies that are specific for αSOs and demonstrate the superior qualities of FIDA as a sample-economic approach that uses in-solution techniques to obtain binding affinities under true equilibrium conditions. Our results were corroborated by SPR values which agree well with FIDA values for NB2 and show a 3-fold difference in affinity versus FIDA for NB1, probably due to the difference in presentation (Ab-tethered versus in-solution).
To increase the nanobodies’ avidity, we designed different DNA scaffolds to dimerize NB1 and enhance its binding affinity. Previous multimerization using nucleic acid scaffolds has successfully increased the valency and avidity of different binders34. While the binding affinity of scaffolded NB1 was lower than that of NB1 alone, multimerization clearly showed enhanced binding. The lower affinities found for the scaffolded constructs suggest a negative contribution of the scaffold and the DNA linkers in the nanobody binding. We hypothesize that this effect may be due to steric hindrance or electrostatic repulsion between DNA scaffolds and αSO. Both BLI and FIDA data validated the 3 T DNA scaffold with 2 nm spacing between nanobodies as the most effective candidate. Overall, our data strongly suggests that the αSO has at least two binding sites for NB1. DNA scaffolding of the nanobodies can enhance valence and avidity, and the use of alternative scaffolds that do not hinder αSO binding holds potential for the design of more potent NB1 multimers in the future.
Prospects for using our nanobodies for diagnostic purposes
The nanobodies retained an ability to distinguish between α-Syn monomers and aggregates in the more complex environment of murine brain homogenates, although they also recognized fibrils as well as oligomers (though to a slightly lower extent). Whether this reflects a rearrangement of the fibrils caused by factors found in the lysate or reflects other intermolecular interactions in the lysate remains unclear. However, it is encouraging that the monomeric state is completely ignored and that the nanobodies also elicit signals at a level comparable to the commercial antibody MJF14 in brain lysates from PD mice.
The nanobodies did not show strong effects on α-Syn fibrillation, which is not unexpected given their inability to bind to immobilized α-Syn fibrils and their remarkable preference for the oligomeric state, which is largely an off-pathway species that does not directly contribute to the fibrillation process7. Nevertheless, the inability of the NBs to block αSO permeabilization of DOPG vesicles loaded with calcein contrasts with the performance of our mAbs raised against different forms of oligomeric α-Syn; several of these mAbs showed very significant efficacy in blocking calcein release23. We attribute these effects to the larger sizes of the mAbs which effectively envelop the αSOs, curtailing contacts with other potential binding partners. The calcein release assay is in our experience a good proxy for bona fide cell assays35, so we do not expect the nanobodies to perform better in actual cytotoxicity measurements. In addition, they showed inferior performance compared to mAbs in inducing the uptake of extracellular αSOs into HEK293 cells via the Sortilin system for endosomal degradation. They may be more successful when targeting other receptors. Nanobodies raised against monomeric α-Syn (but without reported affinities for α-Syn) have been coupled to thalidomide (an E3 ligase ligand) and enter the brain16, clearing both soluble and insoluble α-Syn18.
In conclusion, while the scope of NB1 and NB2 for therapy currently is limited, it remains of potential interest to apply them for diagnostic purposes (particularly given their superior BBB crossing abilities compared to mAbs). In this context, it should be noted that the best way for the nanobodies to discriminate between the different states of α-Syn involve binding of the α-Syn species (monomer, oligomer, or fibril) to the surface of a 96-well plate, followed by transfer of nanobodies and their detection by anti-His antibodies. This indicates that the optimal operational flow to analyze patient samples for the presence of αSOs is to first immobilize them in plates, followed by the addition of the nanobodies. Efforts are currently underway to investigate this further.
Methods
Materials
α-Syn monomer, oligomer, and fibrils were produced as described23. For high-throughput screening, commercial reagents used included DMSO (99% biograde, Cas: 67-68-5, VWR, Cat: ICNA0219481983), PBS (ThermoFisher, cat: 10010015), Tween20 (Fisher Scientific, Cat: 11851465), Horseradish peroxidase (Jackson, cat. 115-035-146), TMB One (Kem-en-tec, Cat: 4380A) and H2SO4 (95%, Cas: 7664-93-9, Fisher Scientific, Cat: 10294300).
Immunization of llamas and phage display
For nanobody production, a llama (Llama glama) was maintained in pasture and immunized by Capralogics Inc (Gilbertville, MA). All animal procedures described in this study were approved by the Institutional Animal Care and Use Committee of Capralogics, Inc., a USDA-regulated research facility. All experiments were performed in accordance with the guidelines and regulations for animal care. All procedures related to animal care were performed by Capralogics, Inc., and have been through scientific peer review at Aarhus University. The llama was immunized four times with 500 μg of purified αSOs. Peripheral blood lymphocytes were isolated from a blood sample, and total RNA was isolated using QIAshredder and RNeasy Plus Mini Kit (Qiagen). From this, total cDNA was generated using the Superscript IV First-Strand Kit (Invitrogen) with oligo(dT) and random hexamer primers. Coding regions of the nanobodies (NBs) were amplified by PCR, restriction digested, and gel purified before it was ligated into a phagemid vector backbone where the NBs were C-terminally fused to an E-tag followed by the pIII coat protein. To generate the final M13 phage display NB library, the phagemid vector was transformed into TG1 electroporation-competent cells (Agilent), infected with VCSM13 helper phages.
The selection was carried out as described36 with small modifications. Briefly, an ELISA plate (High Bind; F, Sarstedt) well was coated with either 2% BSA or 0.1 μg of purified αSOs and was blocked with 2% BSA in PBS. 2.5 × 1012 M13 phage particles in PBS with 2% BSA were first negatively selected against BSA with 1 h incubation followed by 1 h incubation with αSO. The well containing αSO was washed 15 times with 200 μl PBS with 0.1% Tween20 and 15 times with 200 μl PBS. Phages were eluted with 100 μl 0.2 M glycine pH 2.2 and were neutralized with 15 μl 1 M Tris pH 9.1. The eluted phage particles were amplified in ER2738 cells and subsequently used for a second round of phage display. This time the negative selection was done against BSA, α-Syn monomer and fibril followed by selection against αSO. 2% BSA or 0.1 μg of purified antigen was used to coat the ELISA plate and 2.5 × 1010 M13 phage particles from the first round were added. The phages were first incubated with BSA followed by α-Syn monomer, α-Syn fibril and finally αSO. The incubation time for each step was 1 h. The phages were eluted from the αSO well, incubated with ER2738 and then plated out.
Single colonies were picked and grown in LB medium in a 96-well plate format for 4 h before NB expression was induced with 1.0 mM IPTG. An ELISA test was performed to evaluate the Nbs’ ability to bind BSA, α-Syn monomer, fibril, and αSOs. This was done by coating an ELISA plate with 0.1 μg of antigen, followed by blocking with 2% BSA in PBS with 0.1% Tween20 for 1 h at 37 °C. The plate was washed in 0.05% Tween20 in PBS three times on a Intelispeed Washer IW-8 (BioSan, Riga, Latvia). Growth media from single colonies were added to the ELISA plate, incubated for 1 h, and washed three times with PBS with 0.05% Tween20 before anti-E-tag-HPR antibody (Bethyl) was added at a 1:10,000 dilution. The plate was incubated for 1 h, washed, and developed with 3,3’,5,5’-tetramethylbenzidine. The reaction was quenched with 0.5 M H2SO4, and the absorbance was measured at 450 and 620 nm on a HiPo Microplate Photometer MPP-96 (BioSan, Riga, Latvia).
Sequencing, expression, and purification of nanobodies
Selected clones from the phage display were isolated and sequenced using the Eurofins Mix2Seq service with the primer KRA242-VHH-1stSeqf (TCCGGCTCGTATGTTGTGTGGAAT). Sequence similarities were analyzed using CLC Workbench. The two unique NB sequences were amplified from the phagemids using PCR amplification using the primers Llama Nanobody Fwd (AATGCAGGUGCAGCTCGTGGAGTCTGG) and Llama Nanobody Rev (ATGGTGUGAGGAGACGGTGACCTGG). The amplified NB genes (NB1 and NB2) were subcloned into a pET11d expression vector with Uracil-Specific Excision Reagent (USER) cloning and transformed into chemically competent XL1-Blue cells. The transformed cells were plated out on two separate LB-agar plates containing 100 µg/ml ampicillin and incubated overnight at 37 °C. Five potential clones were harvested from each plate (10 clones in total) and transferred to separate tubes of 10 ml LB-media containing ampicillin and incubated in a shaking incubator at 37 °C overnight. Plasmids from all ten clones were purified using a ThermoFisher GeneJET Plasmid Miniprep Kit and sequenced with Eurofins Mix2Seq service using the primer pET11d Seq Rev (CTTTCAGCAAAAAACCCCT). The sequences were analyzed, and a successful NB1 and NB2 clone were selected for further expression and purification. The NB constructs contained a C-terminal His6-tag for purification and were transformed into BL21(DE3) cells, which were incubated for 2 h at 37 °C in LB medium until reaching an OD600 of 0.8 and were then induced with 1 mM IPTG. Cells were harvested after 4 h incubation at 37 °C by centrifugation at 4 °C. The pellet was resuspended in lysis buffer (50 mM Tris pH 8, 150 mM NaCl) containing 10 µg/ml DNase and one protease inhibitor tab (EDTA-free cOmplete ULTRA tablet, Roche) per 50 mL buffer. The solution was sonicated for 6 × 20 s at 4 °C and centrifuged at 10,000 × g at 4 °C to collect the supernatant containing NBs. A HisTrap HP 1 mL column was equilibrated in lysis buffer, and the NB solution was loaded on the column and washed in lysis buffer. A 0–100% lysis buffer with 0.5 M imidazole was used to elute the NB. Subsequently, the NB was transferred to PBS buffer using a PD-10 column equilibrated in PBS.
NabFab production
NabFab protein was produced as described in ref. 26 with some modifications. E. coli C43(DE3) cells were transformed with pRH2.2 NabFab plasmid and cultured at 37 °C in lysogeny broth supplemented 100 μg/mL ampicillin. NabFab expression was induced by the addition of 0.4 mM IPTG at OD600 = 0.6, and cultures were incubated overnight at 18 °C. E. coli were harvested by centrifugation and lysed by sonication. The lysate was incubated at 60 °C for 30 min, and the precipitate was removed by centrifugation. The supernatant was loaded onto a 1 mL HiTrap protein L column and washed extensively in 50 mM Tris-HCl pH 8, 500 mM NaCl before NabFab was eluted in 0.1 M acetic acid. Elution fractions were neutralized in 50 mM Tris-HCl pH 8 and dialyzed against 50 mM Tris-HCl pH 8, 200 mM NaCl.
Denaturant-induced unfolding
Unfolding of NB1 and NB2 in the presence of denaturants was monitored using a LS 55 Luminescence Spectrometer (Perkin Elmer, USA). Fluorescence measurements were performed with an excitation wavelength (λexc) of 280 nm and an emission wavelength (λem) range of 300–440 nm, with a scanning rate of 100 nm/min. NBs were diluted to ca. 80 µM in Tris buffer (20 mM Tris, 300 mM NaCl, pH 7.4) and then diluted 20-fold to 4 µM in 200 µL in 0–9.5 M urea. An analogous procedure was followed to make solutions in 0–6 M guanidinium chloride (GdmCl). The samples were incubated for 1 h at 25 °C before being transferred to a 10 mm quartz cuvette for measurement. Each measurement was recorded as an average of three scans.
Differential scanning fluorimetry (DSF)
Thermal unfolding of nanobodies was monitored using a Tycho NT.6 instrument (Nanotemper Technologies, Germany). NB samples were prepared at three different concentrations (1 mg/mL, 0.2 mg/mL, and 0.04 mg/mL, i.e., 3–75 µM) and loaded into Tycho NT.6 capillaries (Nanotemper Technologies, Germany). Fluorescence was detected at 330 nm and 350 nm as the temperature was increased from 35 °C to 95 °C at a scan rate of 30 °C/min. The ratio of the fluorescence emission at these two wavelengths was plotted versus temperature using the built-in software of the Tycho NT.6 instrument to determine the unfolding temperature tm.
Nanobody labeling with AlexaFluor™ 488
Nanobodies NB1 and NB2 were labeled with AlexaFluor™ 488 amine (ThermoFisher, A20000) following the NHS ester labeling protocol outlined by Fluidic Analytics (https://fluidic.com/wp-content/uploads/2024/07/NHS-ester-labeling-guide_UG-002.pdf) to achieve a 3:1 molar ratio of dye to protein. Briefly, 72.3 µL of 1 M sodium bicarbonate (NaHCO3) buffer (Merck, 1.06329.1000) was added to 200 µL of NB1 solution (850 µg/mL), and 59.5 µL of 1 M NaHCO3 buffer was added to 200 µL of NB2 solution (810 µg/mL). Subsequently, 5.0 µL of a 7.8 mM AlexaFluor™ 488 stock solution was added to NB1, and 4.8 µL of the same stock solution was added to NB2, with the dye volumes calculated using the Fluidic Analytics labeling calculator (https://fluidic.com/labeling-calculator/). The labeling mixtures were gently mixed by pipetting and incubated at 25 °C for 3 h in the dark. Post-incubation, unreacted dye was removed, and the samples were desalted using PD-10 desalting columns (Cytiva) equilibrated with phosphate-buffered saline (PBS, pH 7.4). The labeled nanobodies were maintained in PBS for subsequent analyses.
Nanobody conjugation onto DNA-linkers
Nanobody NB1 was expressed with the unnatural amino acid 4-azido-L-phenylalanine (pAzF) as described for other nanobodies37, allowing the incorporation of an azide group for conjugation. The NB1 was conjugated to either amine-conjugated DNA Linker 1 (5’-5AmMC6/TTCCCGACACCCGCGGAT-3’) or DNA Linker 2 (5’-CGACGCCCTCTTGCCTCCCTT/3AmMO-3’) in a two-step reaction. Firstly, DNA linkers and 25-fold excess of NHS-DBCO were incubated in 100 mM HEPES, pH 8 and 62% DMSO for 4 h at 30 °C, 1000 rpm, in the dark and spiked with 25 excess NHS-DBCO at the 2 h point. DBCO-DNA linkers were precipitated in 0.3 M NaOAc, pH 5.2, and 5–10 excess 96% EtOH (for NaOAc) for 1 h at −80 °C and centrifuged for 30 min at 4 °C, 12,700 rpm. Then redissolved in nuclease-free water and purified using RP-HPLC on a Kinetex C18 column (Phenomenex) using the 1260 Infinity II HPLC system (Agilent). RP-HPLC program used: 2 min at 5% MeCN, 13 min gradient from 5% to 95% MeCN, 2 min at 95% MeCN. Flow rate: 0.4 mL/min. In total, 50 mM TEAA was present at all times during purification. Purified DBCO-DNA linkers were lyophilized, redissolved in nuclease-free water and stored at −20 °C after concentration measurements using DeNovix DS-11 FX+ Spectrophotometer. NB1 with pAzF was conjugated to DBCO-DNA linkers using strain-promoted alkyne-azide cycloaddition (SPAAC) click chemistry. Threefold molar excess of NB1 was incubated with DBCO-DNA linkers 100 mM HEPES, pH 7.4, and 40% DMSO overnight at RT, 600 rpm. NB1-DNA linkers were purified using BioRad FPLC on ion exchange Enrich column 5/50.
NB1-DNA Linker 1 and 2 were hybridized onto biotinylated or ATTO488-labeled DNA scaffold 3 T (5’-5ATTO488N or 5BiosG/TTTATCCGCGGGTGTCGGGTTTGGGAGGCAAGAGGGCGTCG-3’) or DNA scaffold 9 T (5’-5ATTO488N or 5BiosG/TTTATCCGCGGGTGTCGGGTTTTTTTTTGGGAGGCAAGAGGGCGTCG-3’) in 1× PBS for 2 min at 70 °C and 2 °C/min to 20 °C.
Biolayer interferometry (BLI) of dimerized NB1
BLI experiments were carried out using an Octet®RED96e System (Pall®FortéBio) applying Octet®Streptavidin (SA) biosensors. Two hundred microlitres 500 nM biotinylated scaffold 3 T and 9 T, NB-DNA Linker 1 and 2 and 0–500 nM αSO, 20 µM α-Syn monomer, and 2 mg/mL α-Syn fibrils were prepared in 1× PBS containing 0.02% Tween20 and 0.1 mg/mL BSA. Hybridization was executed stepwise, by first immobilizing the biotinylated 3 T or 9 T DNA scaffold to a threshold of 0.4 and then hybridizing one and the second NB1-DNA linkers until saturation. The tip was regenerated by applying 500 mM phosphoric acid and NB1-DNA linkers were re-hybridized between each measurement.
Flow-induced dispersion analysis (FIDA)
Binding of nanobodies to αSO
A range of concentrations (0.1–1024 nM) of monomeric α-Syn and αSO was prepared along with a 125 nM sample of the labeled nanobodies. The FIDA assay was performed on a FIDA1 instrument (Fidabio, Denmark) with a 480 nm detector and a standard Fida capillary (75 µm inner diameter). The instrument was configured to flow Milli-Q water at 3500 mbar for 45 s (wash), followed by PBS at 3500 mbar for 75 s (buffer equilibration), analyte (αSO) at 3500 mbar for 20 s (analyte injection), and the labeled nanobody at 50 mbar for 10 s (indicator injection), concluding with analyte (αSO) at 350 mbar for 205 s (second analyte injection). Notably, the last analyte injection results in at least tenfold dilution of the indicator at the given pressure. This setup facilitated the binding of the indicator and analyte within the capillary column. Fluorescence detection was performed using the instrument’s 488 nm detector. All measurements were conducted at 25 °C, with the capillaries washed with NaOH before use. The same protocol was applied for monomeric α-Syn interactions. Each measurement was performed in triplicate to ensure reproducibility. Data were fitted with the following 1:1 binding model:
Here [A] is the αSO concentration, RI is the hydrodynamic radius for the indicator (nanobody) and RIA is the hydrodynamic radius for the indicator:analyte (nanobody:αSO) complex.
Binding of grafted NBs to Nab-Fab
NBg1 and NBg2 were labeled by the same approach as NB1 and NB2, however, as they were stored in a 5% glycerol, we carried out buffer exchange into PBS using a PD-10 column prior to the labeling reaction. To measure the affinity between the two NBg’s and NabFab by FIDA, NabFab (0–8 µM) in PBS + 0.05% pluronic acid was titrated against 100 nM of either NBg, using in-capillary mixing and a standard injection method at 400 mbar. The affinity between the NBg or NBg:NabFab complex and αSOs was measured by titrating 0–0.97 µM αSO in PBS + 0.05% pluronic acid, using the same in-capillary mixing and injection method. When NBg:NabFab complex was used as the ligand in ternary complex formation with αSO, 100 nM of either NBg1 or NBg2 were mixed and incubated with 1000 nM NabFab for 15 min before titration. All measurements were fitted using the single species fit in the FIDA Software.
Binding of dimerized nanobodies
Either one, two or both of the NB1-DNA linkers were hybridized onto the 3 T and 9 T scaffold as described above to concentrations of 200 nM. αSO (0–500 nM) was prepared in 1× PBS and the experiments were run as described above for the unscaffolded nanobodies.
Surface plasmon resonance (SPR)
SPR experiments were carried out on a Biacore 3000 (Cytiva) using 10 mM NaH2PO4, pH 7.5 + 150 mM NaCl + 0.05% Tween20 as running buffer. Binding analysis was performed at 25 °C, and data were collected at a rate of 1 Hz. Kinetics of nanobody binding to αSOs was performed on an HC200M chip (XanTec bioanalytics). Using standard amine chemistry, in accordance with the instructions of the manufacturer, flow cells (FCs) 1 and 2 were coated with protein G (08062, Merck). Next, the αSO-binding mAb 18-3A5-H2 (which shows ~100-fold greater affinity for αSOs over monomeric α-Syn23) was captured in both FCs, followed by capture of αSOs in FC2 only. Titration series of nanobodies were then injected over both surfaces for 120 s, at a flow rate of 30 µl/min, followed by a 300 s dissociation phase. At the end of each round, surfaces were regenerated by a 30 s injection of 10 mM glycine pH 1.5. Using the BIA evaluation 4.1.1 software (Cytiva), recorded signals from FC2 were double referenced, i.e., the signal from the in-line reference flow cell, FC1, was subtracted as was the signal from a blank run (0 nM analyte).
To evaluate the competitive binding of NabFab and αSOs to Nbg1, NabFab was immobilized in FC1 and FC2 on a CM5 chip (Cytiva), using standard amine chemistry. At a flow rate of 10 µl/min, Nbg1 was sequentially injected for 120 s without (FC1) or with (FC2) a preincubated titration series of αSOs. Regeneration between rounds was done by a 120 s injection of 10 mM glycine pH 2.2. Assuming only free nanobody will bind to NabFab, relative binding (FC1/FC2) was calculated from the initial association rates (the linear part of the association curve immediately following the start of injection, where free binding sites are not limiting). Data analysis was performed using GraphPad Prism 10. Relative binding was plotted as a function of log (competitor) and fitted with the built-in model “one site fit Ki”. The FIDA-determined KD of 28.4 nM for the interaction between NAbFab and NBg1 was used for the fitting.
Western blotting
To prepare oligomeric species of α-Syn by nitration, 1 ml α-Syn at 3.5 mg/ml (242 µM) was incubated with a fourfold excess of tetranitromethane (Sigma-Aldrich, 10% stock solution in ethanol (0.827 M)) for 3 h at 30 °C, followed by desalting on a PD Miditrap G-25 column to collect the protein fraction based on absorbance at 280 nm. The oligomeric fraction was separated from monomer and dimer fractions on a Superose 6 column and concentrated to ca. 1 mg/ml with an Amicon spin 10 K cut-off spin filter. Oligomers of α-Syn made by incubation with EGCG or ONE were prepared, as described in ref. 23. The different oligomers were run on SDS-PAGE, electroblotted onto a nitrocellulose membrane, blocked with 2% BSA, and incubated with 1 µg/ml NB1 or NB2 in PBS buffer for 2 h at room temperature, after which the solution was developed with 1:10,000-fold diluted anti-His-HRP antibody (Sigma-Aldrich) for 1 h at room temperature. The membrane was washed with PBS and 0.05% Tween20 between all steps. Color was developed using TMB blotting from Kementec.
Calcein release assays
These were performed as described38 using DOPG vesicles filled with 50 mM calcein. Dye release in the presence of αSOs was normalized to full release in the presence of 0.1% Triton-X-100. 6 µg/ml αSO was preincubated with different concentrations of nanobodies (molar ratios NB:αSO from 0.03 to 320), after which they were mixed with DOPG vesicles, and the reaction followed in a 96-well plate until the fluorescence signal had stabilized at a plateau.
Thioflavin T fluorescence assays
These were performed at 37 oC while shaking in the presence of a glass bead as described in ref. 23 using 1 mg/ml α-Syn and 4–1000 µg/ml nanobody.
Mass photometry
αSO at a final concentration of 1.62 µM (oligomer units) was mixed separately with either NBg1-NabFab or NBg2-NabFab complexes in 1:3 or 1:10 ratio. To make the NBg-NabFab complexes, NBg1 and NBg2 were mixed with NabFab and incubated for 10 min at ambient temperature prior to the addition of αSO. The samples were frozen and stored for later measurements. The molecular mass of the complexes was measured on a TwoMP mass photometer (Refeyn, Oxford, England). The system was calibrated with BSA (66 and 132 kDa for monomer and dimer, respectively) and thyroglobulin (670 kDa). The αSO complex samples were first diluted 4-fold. Then, using the drop-dilution method, an 18 µl drop was used to establish a baseline signal. After that, 2 µl of the αSO samples were added, resulting in a final 40-fold dilution to 40.5 nM αSO. The samples were recorded over 60 s, and the resulting ratiometric contrast distributions were converted to mass using the calibration curve.
TEM analysis of αSO:NB:NabFab complexes
NabFab was prepared as described in ref. 27. The αSO was prepared by incubation of 8 mg/ml monomeric α-Syn at 37 °C while shaking at 900 rpm for 3.5–5 h. In a separate reaction, the NB:NabFab complex was prepared by incubating 18 µM NB with 18 µM NabFab (final concentrations after mixing with αSO, see below) for 1 h at room temperature. Subsequently the complex was diluted 3.75-fold to 5 µM of each component by being mixed 1:3 (αSO:complex molar ratio expressed in α-Syn monomer units based on a yield of 1% αSO in the α-Syn incubation mixture, i.e., 10:1 αSO:complex when expressed in αSO units) with αSO for 1 h at room temperature, after which the mixture was run over a Superose 6 prep grade SEC column23. The first peak after the void volume was collected and visualized by TEM, as described in ref. 39.
High-throughput screening assay
This was based on an ELISA sandwich screening assay previously described38. Briefly, 60 μL of 5 μg/mL NB1 per well was used to coat a 96-well High Bind ELISA plate, after which 2 μg/mL αSO and 25 µL compound (typically 10 µM concentration) was added, followed by the addition of 25 μL of 1 μg/mL of the primary anti-αSO mAb 14-9E7-A1, 50 μL secondary Jackson GAM-HRP antibody and addition of 50 μL of HRP substrate TMB One (3,3´,5,5´-tetramethylbenzidine). The α-Syn assay was miniaturized and adapted to 384-well plate format by dividing the volumes used in this procedure by four. Based on this, volumes, concentrations, and conditions were optimized in order to screen the full library in a high-throughput format. The screen was carried out at DTUSCore laboratory automation system using a 384-well plate format (clear, flat-bottom, NUNC plates, Corning Inc., Corning, NY, USA), and absorbance (450 nm) was measured by Tecan Spark 600 (Tecan, Männedorf, Switzerland). The compounds screened were from the in-house chemicals library at DK-OPENSCREEN, Kongens Lyngby, Denmark. The entire screening campaign was done in a single dose with a final compound concentration at 10 µM and 0.1% DMSO concentration. All compound stock solutions were dissolved in DMSO (99% biograde, 67-68-5, VWR, cat: ICNA0219481983), resulting in a 10 mM concentration. The outer circle of wells was excluded from the assay to avoid edge effects but was still filled with reagents. Control wells (two rows) were either treated with no compounds but same amount of DMSO (0.1% v/v) to get maximum signal, or with no α-Synuclein oligomer to measure background signal (layout shown in Fig. S12).
The optimized assay for HTS campaign was performed in a two-day experiment as follows.
Day 1
In total, 15 µL of a 5 µg/µL of NB1 in PBS (ThermoFisher, cat: 10010015) were dispensed using a Multiflo dispenser (Agilent, Santa Clara, California, United States) and incubated overnight at 4 °C.
Day 2
All wells were emptied using the EL406 dispenser/washer (Agilent, Santa Clara, CA, USA). Then 20 µL of 1% of blocking agent BSA (9048-46-8, Merck, cat A2153) in PBS were dispensed by EL406 and left to incubate for 30 min at 37 °C.
Washing
All wells were emptied and washed three times by EL406 with 1× PBS and 0.005% TWEEN®20 (Fisher Scientific, Cat: 11851465), and emptied.
Addition of oligomer
Overall, 13 µL of αSO were added to all wells except one entire row (background signal) using the Multiflo and left for 60 min at 37 °C after which all wells were washed three times by the EL406.
Addition of compounds
In total, 12 nL of 10 mM stock solution of compounds were dispensed by an ECHO550 (Labcyte/Beckman, Brea, CA, USA) to all wells, except two rows where 12 nL of DMSO were added (background and maximum signal), The wells were topped off with 6 µL PBS with 0.005% TWEEN®20 with the help of the Felix pipetting robot (Analytik Jena GmbH+Co. KG, Jena, Germany) and incubated for 30 min at 37 °C.
Addition of primary antibody
In all, 6 µL of 1 µg/µL of primary mAb 14-9E7-A123 in PBS and 0.005% TWEEN®20 was added to all wells using the Multiflo and left to incubate for 60 min at 37 °C before washing three times with the Multiflo.
Addition of secondary antibody
Commercial secondary goat-anti-mouse Ab combined with Horse-peroxide-radish (1:20,000 in PBS and 0.005% TWEEN®20 was dispensed in a volume of 13 µL with the Multiflo. Another 60 min of incubation at 37 °C was followed by the washing procedure described above.
Development of signal
Overall, 13 µL of TMB ONE reagent kit (Kem-en-tec cat: 4380A) were added to each well with the EL406 to start the reaction with horseradish peroxidase-reaction. After incubation for 30 min at 37 °C the reaction was stopped by the addition of 25 µL of 0.5 M H2SO4 (95%, CAS: 7664-93-9, Fisher Scientific, cat: 10294300), and absorbance at 450 nm was immediately measured by the Tecan Spark plate reader.
Based on the HTS campaign, 56 compounds were chosen for validation. The selected hits were confirmed by ten points concentration dose–response, with 90 µM, 30 µM, 10 µM, 4 µM, 1 µM, 0.375 µM, 0,125 µM, 41 nM, 10 nM, and 4.6 nM concentration, in triplicates, also using the above described automated assay in 384-well format. In total, 23 compounds showed below 30% remaining activity at 10 µM.
Screening data analysis
All data from the plate reader was analyzed by MolFormatic, a Python program developed at DTU Chemistry that handles data from compounds libraries and activity data generated from screenings. The data generated after each screening plate readout was first imported into MolFormatic, followed by a background correction and normalization by subtracting the background (the row with no αSO). Hits were determined by comparing the normalized value with our predetermined threshold (<30% of binding compared to positive control). For each plate, we obtained the average, sample standard deviation of data, population standard deviation of data, population variance of data, sample variance of data, and standard deviation % for minimum and maximum controls, samples, and empty wells. For the original data, the signal/baseline (S/B) was calculated based on the normalized data. Finally, one histogram per plate was prepared and checked for a normal distribution, otherwise the data was discarded. A heat map for the normalized data was created to visualize the results.
Dose response
Dose–response is calculated using MolFormatic. The calculations are based on normalized data to provide the Hill constant, slope, R2, and EC50. The Brent Equation was used to draw the dose–response curve, due to it being more robust for an HTS screen:
Response = upper + (lower - upper)/(1 + (x/EC50) ** -hillslope)
Background correction: wellorg – minAvg.
R2:
Dose response:
Peptide array studies
Peptide arrays were produced, as described40. Each peptide array was blocked overnight in 5 mL 2% BSA in PBS at 4 °C. After blocking, the arrays were washed 3 times in a block buffer (PBS pH 7.5, 0.1% Tween20) and then incubated for 1 h at 37 °C on a rolling board at room temperature with 3 mL mAb or NB at 1.0 µg/mL. The array was then incubated 1 h on a rolling board at room temperature with 3 mL anti-His-tag (Merck, cat. SAB2702218) at 1:1000 dilution (1 µg/mL). The arrays were washed in buffer (PBS pH 7.5, 0.1% Tween20) 3 times followed by 1 h incubation with the secondary antibody GAM-HRP, washed 3 times with (PBS pH 7.5, 0.1% Tween20) and developed with TMB blotting (Kementec) and imaged with GELDOCGo (Biorad).
ELISA measurements
To measure the binding of NB1 and NB2 to immobilized α-Syn (monomer, oligomer, and fibrils prepared as described in ref. 23), a 96-well plate was coated with 5 µg/ml of the α-Syn sample, after which it was blocked with 2% BSA, washed and incubated with twofold dilution series of NB1 and NB2 from 20 to 0.75 µg/ml. As primary Ab, we used anti-His SAB2702218 (1 µg/mL) and as secondary Ab GAM-HRP diluted 1:20000 (Jackson). HRP signal was developed as described23 using absorption at 450 nm.
Preparation of brain homogenate
Healthy male (wildtype) Swiss mice (n = 3) were housed with ad libitum access to food and water, in a climate-controlled facility under a 12 h/12 h night/daylight cycle. All experiments were conducted in accordance with the ethical commitments for the use of laboratory animals at Aarhus University and approved by the Danish Supervisory Authority on Animal Testing. One-month-old mice were terminated by cervical dislocation, the brain was extracted, and each hemisphere was cut into three coronal pieces (cut approx. −1.22 mm and −3.8 mm A–P from Bregma) and snap frozen on dry ice. The middle of the left hemisphere was homogenized as previously described41, with some modifications. Briefly, the brain was dounce-homogenized 1:10 in PBS buffer (PBS, 5 mM NaF, 22.5 mM β-glycerolphosphate, 1 mM Na3VO4, 1% Triton-X-100) containing protease and phosphatase inhibitors (cOmplete Mini EDTA-free inhibitor cocktail tablet, Merck), 20 bumps with rotor set at 700 rpm. The homogenates were sonicated (4 × 10 strokes using a Branson Ultrasonics Sonifer S250 A, duty cycle 80, and output control 6). Homogenates from three mice were diluted eight times from a protein concentration of approx. 3 mg/ml (measured using a DS-11 spectrophotometer (Denovix, Wilmington, DE, assuming 1 OD280 corresponds to 1 mg/ml), spiked with 10 µg/ml of either monomeric, fibrillar, or oligomeric α-syn and incubated ≥10 min on ice. Subsequently, samples were subjected to a twofold dilution series in PBS (from 3 to 0.0015 mg/ml), and indirect ELISA assays with 5 µg/ml NB1 and NB2 were carried out as described above. Subsequent to incubation with NB1 or NB2, we quantified the amount of bound nanobody using the anti-His antibody SAB2702218, while α-Syn was quantified using the monoclonal antibody 14-9E7 A1 as described23. To analyze α-syn aggregates in PD mice, we used a human A53T transgenic M83 line as described23 and extracted brain homogenates as described above. Homogenates were diluted 8 times and then diluted again either 1:1 or 1:5, after which they were added to microwell plates and incubated with either 1 µ/ml MJF14 antibody or 5 µg/ml NB1 or NB2. The signal was developed with either Goat Anti-Rabbit (MJF14) or Goat Anti-Mouse HRP-linked antibody as described23.
Microscale thermophoresis
To measure the binding of small molecule hits to αSOs, a 12-point titration series of small molecules was prepared in TTP LVDS 384-well plates (SPT Labtech) in 5 µL in a PBS buffer containing 4% DMSO (v/v) and 0.05% Tween20 (v/v). The plate was spun down before the addition of 5 µL solution containing 4.45 nM αSO labeled with Alexa647 in 2× PBS (Fisher BioReagents) and 0.05% Tween20 (v/v) to each well. At this point, the plate was sealed with adhesive strip and placed on an Eppendorf thermomixer to shake at 300 rpm for 5 min before spinning down the plate and further incubation 1 h at 19 °C. MST was measured on a Monolith NT Automated (Nanotemper) in standard-treated capillaries at 10% LED-power and high MST-power. Dose–response was extracted from the raw thermographs at 10 sec hot-time and the dissociation constant was estimated by sigmoidal curve fitting in MO Affinity analysis program (Nanotemper). Experiments were run in duplicates. To measure the binding of small molecules to NB1 and mAb 14-9E7-A1, a 24-point titration series of small molecule was prepared as above, using instead of αSO either 4 nM mAb-NHS-RED (labeling according to manufactures protocol, degree of labeling (DOL) = 0.6) or 100 nM NB1-NHS-RED (labeling according to manufactures protocol, DOL = 0.23).
α-Syn uptake in HEK293 cells in the presence of nanobodies
For these experiments, NB1 and NB2 were N-terminally tagged with a Met-Gly-His6 sequence immediately preceding the QVQ… N-terminus, while the C-terminus contained the linker sequence SS(GGGGS)3GG followed by RQLL (negative controls omitted the RQLL sequence). The sequences were cloned into pET11d vectors by Genscript and expressed as described for NB1 and NB2 above. The HEK293 cells were transfected to express the Sortilin receptor42 with zeocin used for selection. The cells were grown in Dulbecco’s Modified Eagle Medium with 10% fetal bovine serum, 1% penicillin/streptomycin, and 1% Glutamax either with or without zeocin (Med-Zeo and Med, respectively). In all, 2 mio. cells in 15 mL Med-Zeo were seeded in 55 mL cell culture bottle and incubated for about 75 h. Subsequently, old media was removed, and cells were washed using PBS. Trypsin was added to the cells to dissociate cells from the cell culture bottle. Fresh Med-Zeo was added, and the cells were transferred to a 15-mL tube which was spun down for 3 min at 400 × g. Media was removed from the cell pellet, and the pellet was resuspended in fresh Med-Zeo. Cells were then seeded in a 96-well plate (coated with poly-L-Lys and preincubated for 30 min at 37 °C with 5% CO2, after which it was washed twice with 100 µL demineralized water per well) with 50 µL/well in a dilution of 800,000 cells/mL (40,000 cells/well) and incubated at 37 °C with 5% CO2 for 24 h.
A dilution series of NB1 and NB2 was prepared (both with RQLL-OH and RQLL-NH2 C-terminal extension) using Med in a twofold 9-step dilution series from 5 µM nM to 20 nM and a 0 nM control. Med with 200 nM αSO (monomer units) labeled with Alexa488 (αSO488) and 160 nM leupeptin was prepared by adding 17.9 µL 56 µM αSO-488 and 16 µL 50 µM leupeptin to 5 mL Med and diluted 1:1 with the different samples from the NB dilution series. 50 µL of each dilution sample was then added to the wells of the 96-well plate in duplicates, after which the cells were incubated for 24 h at 37 °C with 5% CO2. Old media was removed, and cells were washed three times in 3 × 50 µL cold PBS. After the last wash, PBS was left in the wells. Fluorescence from the αSO488 (taken up by the cells) was measured with a CLARIOstar microplate reader (BMG Labtech) using excitation at 488 ( + /−8) nm and emission at 535 ( + /−15) nm. Monomers and fibrils were tested using the same assay using 100 nM each of monomer and fibrils (fibril concentration in monomer units).
Data availability
All data are provided in the manuscript and will be made available upon request.
Abbreviations
- α-Syn:
-
α-synuclein
- αSO:
-
α-synuclein oligomer
- FIDA:
-
flow-induced dispersion analysis
- NabFab:
-
synthetic Fab fragment
References
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu Rev. Biochem. 82, 775–797 (2013).
Kunz, P. et al. The structural basis of nanobody unfolding reversibility and thermoresistance. Sci. Rep. 8, 7934 (2018).
Messer, A. & Butler, D. C. Optimizing intracellular antibodies (intrabodies/nanobodies) to treat neurodegenerative disorders. Neurobiol. Dis. 134, 104619 (2020).
Siontorou, C. G. Nanobodies as novel agents for disease diagnosis and therapy. Int. J. Nanomed. 8, 4215–4227 (2013).
Yang, Y. et al. Structures of α-synuclein filaments from human brains with Lewy pathology. Nature 610, 791–795 (2022).
Alam, P., Bousset, L., Melki, R. & Otzen, D. E. α-synuclein oligomers and fibrils: a spectrum of species, a spectrum of toxicities. J. Neurochem. 150, 522–534 (2019).
Lorenzen, N. et al. The role of stable α-synuclein oligomers in the molecular events underlying amyloid formation. J. Am. Chem. Soc. 136, 3859–3868 (2014).
Lorenzen, N. et al. How epigallogatechin gallate can inhibit α-synuclein oligomer toxicity in vitro. J. Biol. Chem. 289, 21299–21310 (2014).
Volles, M. J. et al. Vesicle permeabilization by protofibrillar α-synuclein: implications for the pathogenesis and treatment of Parkinson’s disease. Biochemistry 40, 7812–7819 (2001).
Paslawski, W., Mysling, S., Thomsen, K., Jørgensen, T. J. D. & Otzen, D. E. Co-existence of two different α-synuclein oligomers with different core structures determined by hydrogen/deuterium exchange mass spectrometry. Angew. Chem. Int Ed. Engl. 53, 7560–7563 (2014).
Fusco, G. et al. Structural basis of membrane disruption and cellular toxicity by α-synuclein oligomers. Science 358, 1440–1443 (2017).
Santos, J. et al. Targetable N-terminal motif orchestrates α-synuclein oligomer-to-fibril conversion. J. Am. Chem. Soc. 146, 12702–12711 (2024).
Guilliams, T. et al. Nanobodies raised against monomeric α-synuclein distinguish between fibrils at different maturation stages. J. Mol. Biol. 425, 2397–2411 (2013).
Iljina, M. et al. Nanobodies raised against monomeric ɑ-synuclein inhibit fibril formation and destabilize toxic oligomeric species. BMC Biol. 15, 57 (2017).
El Turk, F. et al. Exploring the role of post-translational modifications in regulating α-synuclein interactions by studying the effects of phosphorylation on nanobody binding. Protein Sci. 27, 1262–1274 (2018).
Jiang, Y. et al. Single-domain antibody-based noninvasive in vivo imaging of α-synuclein or tau pathology. Sci. Adv. 9, eadf3775 (2023).
Tuttle, M. D. et al. Solid-state NMR structure of a pathogenic fibril of full-length human α-synuclein. Nat. Struct. Mol. Biol. 23, 409–415 (2016).
Jiang, Y. et al. Single-domain antibody-based protein degrader for synucleinopathies. Mol. Neurodegeneration 19, 44 (2024).
Butler, Y. R. et al. α-Synuclein fibril-specific nanobody reduces prion-like α-synuclein spreading in mice. Nat. Commun. 13, 4060 (2022).
Emadi, S., Barkhordarian, H., Wang, M. S., Schulz, P. & Sierks, M. R. Isolation of a human single chain antibody fragment against oligomeric α-synuclein that inhibits aggregation and prevents α-synuclein-induced toxicity. J. Mol. Biol. 368, 1132–1144 (2007).
Yuan, B. & Sierks, M. R. Intracellular targeting and clearance of oligomeric alpha-synuclein alleviates toxicity in mammalian cells. Neurosci. Lett. 459, 16–18 (2009).
Islam, Z., Vaikath, N. N., Hmila, I., El-Agnaf, O. M. A. & Kolatkar, P. R. Structural insights into the unique recognition module between α-synuclein peptide and nanobody. Protein Sci. 33, e4875 (2024).
Nielsen, J. et al. Monoclonal antibodies targeting cytotoxic α-synuclein oligomers: molecular properties and diagnostic potential. NPJ Parkinson’s. Dis. 10, 139 (2024).
Otzen, D. E. Antibodies and α-synuclein: what to target against Parkinson’s disease?. Biochim Biophys. Acta Proteins Proteom. 1872, 140943 (2023).
Otzen, D. E., Buell, A. K. & Jensen, H. Microfluidics and the quantification of biomolecular interactions. Curr. Opin. Struct. Biol. 70, 8–15 (2021).
Paslawski, W., Lorenzen, N. & Otzen, D. E. Formation and characterization of α-synuclein oligomers. Methods Mol. Biol. 1345, 133–150 (2016).
Bloch, J. S. et al. Development of a universal nanobody-binding Fab module for fiducial-assisted cryo-EM studies of membrane proteins. Proc. Natl. Acad. Sci. USA 118, e2115435118 (2021).
Hlavacek, W. S., Posner, R. G. & Perelson, A. S. Steric effects on multivalent ligand-receptor binding: exclusion of ligand sites by bound cell surface receptors. Biophys. J. 76, 3031–3043 (1999).
Young, G. et al. Quantitative mass imaging of single biological macromolecules. Science 360, 423–427 (2018).
Zheng, Y., Brady, O. A., Meng, P. S., Mao, Y. & Hu, F. C-terminus of progranulin interacts with the beta-propeller region of sortilin to regulate progranulin trafficking. PLoS ONE 6, e21023 (2011).
Petersen, C. M. et al. Molecular identification of a novel candidate sorting receptor purified from human brain by receptor-associated protein affinity chromatography. J. Biol. Chem. 272, 3599–3605 (1997).
Gustafsen, C. et al. Reshaping the progranulin/sortilin interaction for targeted degradation of extracellular proteins. Preprint at http://www.biorxiv.org/content/10.1101/2025.03.03.641244v1.full.pdf (2025).
Han, B. A suite of mathematical solutions to describe ternary complex formation and their application to targeted protein degradation by heterobifunctional ligands. J. Biol. Chem. 295, 15280–15291 (2020).
Valero, J. et al. A serum-stable RNA aptamer specific for SARS-CoV-2 neutralizes viral entry. Proc. Natl. Acad. Sci. USA 118, e2112942118 (2021).
Farzadfard, A. et al. The C-terminal tail of α-synuclein protects against aggregate replication but is critical for oligomerization. Commun. Biol. 5, 123 (2022).
Hansen, S. B., Laursen, N. S., Andersen, G. R. & Andersen, K. R. Introducing site-specific cysteines into nanobodies for mercury labelling allows de novo phasing of their crystal structures. Acta Crystallogr D. Struct. Biol. 73, 804–813 (2017).
Laura, T. et al. Jørgen, site-specific nanobody-oligonucleotide conjugation for super-resolution imaging. JBM 9, 1 (2022).
Somavarapu, A. K. et al. Drug repurposing screens identify compounds that inhibit α-synuclein oligomers’ membrane disruption and block antibody interactions. Chem. Sci. 14, 3030–3047 (2023).
Sønderby, T. V. et al. Otzen, sequence-targeted peptides divert functional bacterial amyloid towards destabilized aggregates and reduce biofilm formation. J. Mol. Biol. 435, 168039 (2023).
Bleem, A. et al. Protein engineering reveals mechanisms of functional amyloid formation in Pseudomonas aeruginosa biofilms. J. Mol. Biol. 430, 3751–3763 (2018).
Nielsen, J. et al. Molecular properties and diagnostic potential of monoclonal antibodies targeting cytotoxic α-synuclein oligomers. NPJ Parkinsons Dis. 10, 139 (2024).
Petersen, C. M. et al. Propeptide cleavage conditions sortilin/neurotensin receptor-3 for ligand binding. EMBO J. 18, 595-604 (1999).
Acknowledgements
This work has generously been funded by the Lundbeck Foundation (grant no. R276-2018-671) as well as instrument grants from the Carlsberg Foundation (grants no. CF19-0337 and CF20-0610). DK-OPENSCREEN is supported by the Ministry of Higher Education and Science (grant case no. 5072-00019B), Aarhus University, the Technical University of Denmark, and other contributing universities. The DTU Screening Core is supported by the Novo Nordisk Foundation (NNF19OC0055818), the Carlsberg Foundation (CF19-0072), and DTU. Work related to developing nanobodies and NabFab was supported by the Novo Nordisk Foundation (NNF0086732) and the Danish Council for Independent Research (3103-00137B). M.G.M. was funded by Lundbeck Foundation grant no. R380-2021-1393. We warmly thank Dr. Henrik Jensen from FIDAbio for his constant and constructive input regarding FIDA analysis. We are grateful to Dr. Lasse Reimer from the PHJ group at DANDRITE, Aarhus University for technical support, guidance and inputs regarding the processing of mice brain tissue. J.K. and K.F.M. are funded by the Danish National Research Foundation to Center for Cellular Signal Patterns (CellPAT, Grant DNRF135). J.V. is supported by the Lundbeck Foundation (grant no. R346-2020-1999).
Author information
Authors and Affiliations
Contributions
D.E.O. wrote the main manuscript text and prepared the figures with input from J.N., J.S.N., F.N., B.B.B, J.H.C., and A.S. In addition, J.N., J.N.P., G.K., J.S.N., F.N., C.J., E.S., B.B.B., N.M.L., B.V.B., J.H.C., A.S., W.P.O., B.W.S., J.H.M., V.S., M.G.M., M.K., A.B., S.B., G.C., S.B.H., and P.S.M. carried out investigations. T.B., K.S., C.G., S.G., K.R.A., M.H.C., and D.E.O. supervised. D.E.O. conceived and coordinated the study and was in charge of overall fundraising and project management. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.G., P.M., C.G., A.S., M.K. are all employees of Draupnir Bio, a biotech company with an interest in targeted protein degradation. Furthermore, Draupnir Bio has a pending patent application related to present study entitled “Bifunctional molecules that selectively induce degradation of extracellular targets in lysosomes” with the number WO2023247754A1. The remaining authors have no competing interests related to the present study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nielsen, J., Pedersen, J.N., Kleijwegt, G. et al. Nanobodies raised against the cytotoxic α-synuclein oligomer are oligomer-specific and promote its cellular uptake. npj Biosensing 2, 23 (2025). https://doi.org/10.1038/s44328-025-00042-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44328-025-00042-1