Abstract
Cancer screening can enable early detection and improve survival but a focus on single cancers limits cost-effectiveness. Here we present OMAFound (carcinOMA Finder foundation), a foundation model capable of simultaneous multi-cancer screening at both organ level and patient level using widely accessible non-contrast computed tomography (CT). The model was developed and tested on 325,197 CT volumes from 151,386 patients across 10 Chinese and international datasets, achieving performance comparable to mammography-based approaches for breast cancer detection and matching existing lung-specific models for lung cancer detection. In a prospective multi-centre cohort of 21,601 patients undergoing low-dose CT screening, OMAFound demonstrated balanced accuracy of 82.2% for breast cancer and 88.0% for lung cancer in females, while attaining 86.1% balanced accuracy for lung cancer detection in males. When assisted by OMAFound, 7 generalist radiologists showed improvement in sensitivity (mean increases of 38.9% for breast cancer, 16.0% for lung cancer and 21.3% at patient level), without compromising specificity. These findings highlight the potential of OMAFound as a multi-cancer screening tool to offer robust preventive medicine strategies with minimal costs.
Similar content being viewed by others
Main
Cancer is a major global health challenge and remains one of the leading causes of mortality worldwide, with nearly 20 million new cases and 9.7 million deaths reported in 20221. This substantial cancer burden continues to escalate globally, driven by factors such as ageing populations and the prevalence of risk factors such as smoking, obesity and unhealthy diets2,3. Among the various types of cancer, lung cancer stands as the most commonly diagnosed malignancy and the leading cause of cancer-related deaths across populations, accounting for 12.4% of all new cases. Breast cancer follows closely behind as the second most prevalent form, constituting 11.6% of new cases and disproportionately affecting women. Despite cancer’s detrimental impact, the 5-year survival rate for early-stage cancer is notably higher than that of late-stage disease4, underscoring the urgent need for early detection.
Early detection of cancers through screening programmes in the vast asymptomatic population generally shows improved survival and outcomes5,6, especially for high-risk cases, compared with those diagnosed outside of surveillance programmes via standard clinical diagnostic workflows. For instance, low-dose computed tomography (CT) screening has resulted in a marked reduction in lung cancer mortality7,8, while mammography-based screening has been a universally recommended standard for breast cancer detection for over three decades9,10. However, medical image interpretation is a highly challenging task for radiologists owing to anatomical complexity and cognitive load, particularly with volumetric imaging11, leading to subjective characterization and persistent intra- and inter-observer variability12,13. Predictive artificial intelligence (AI), with its robust capability to extract representative features from medical images, has shown promising results in cancer screening, including lung14,15,16, breast17,18,19,20,21 and pancreatic22 cancers. This potential is further validated by pioneer population studies on real-world AI deployment, which demonstrate enhanced cancer detection rates without negatively affecting the recall rates23,24.
Despite the proven benefits of existing screening programmes, they remain constrained by the ‘single test for one cancer’ paradigm, where each imaging examination is optimized for detecting only one specific cancer type. This approach necessitates multiple separate screening examinations for comprehensive cancer detection, increasing both out-of-pocket costs for patients25 and cumulative ionizing radiation exposure risks26,27. Non-contrast CT28, particularly low-dose CT in physical examination centres, offers a low-cost and widely accessible imaging solution, even in low-resource regions. Its broad clinical applicability makes it an ideal candidate for implementing a ‘single test for multi-cancer’ screening approach. However, detecting multiple abnormalities across diverse regions from CT scans presents substantial challenges for conventional predictive AI models, which are typically designed for organ-specific analysis and show limited cross-organ generalizability.
Recent advances in self-supervised learning (SSL)29-based foundation models, leveraging task-agnostic representations from large-scale unlabelled data, has sparked a renaissance in the medical AI field30,31. Although existing CT-focused foundation models have shown great potential on multi-task scenarios32,33,34, such as imaging captioning, detection and segmentation, their potential for multi-cancer screening faces three critical challenges. First, cancer screening requires the sophisticated differentiation of malignancy from general positive findings, a task substantially more complex than basic abnormality detection. Second, CT is not currently a primary screening tool for many cancers, including breast cancer. Thus, its potential value for routine or opportunistic screening using AI remains unexplored. Lastly, previous AI studies have primarily focused on model performance alone, failing to validate real-world effectiveness through prospective studies and how AI can improve the screening outcomes at both organ and patient levels.
In this study, we present OMAFound (carcinOMA Finder foundation), a three-dimensional (3D) CT foundation model-driven AI framework designed for automated multi-cancer screening in asymptomatic populations with minimal costs (monetary, radiation and time). We benchmark OMAFound’s performance against mammography-based AI models for breast cancer prediction, and existing CT-based AI models for lung cancer prediction using large-scale nationwide and international datasets. To assess the generalizability for multi-cancer screening, particularly in low-dose CT settings, we validate the performance of OMAFound in a prospective real-world study across 4 medical centres with 21,601 participants involved. To further evaluate clinical applicability, we compare OMAFound’s predictions with those made by seven generalist radiologists and subsequently explore the potential benefits of AI-assisted radiological decision-making.
Results
Figure 1 outlines the overall study design of OMAFound. The pretraining stage of OMAFound is an SSL-based task-agnostic vision foundation model using the SwinUNETR-V235 architecture (Supplementary Fig. 1). This architecture integrates residual convolution and Swin transformer blocks, enabling efficient processing of 3D medical data while capturing both local and global contextual features. The pretraining was conducted using a large-scale unlabelled dataset from Site A-CTunlabeled and CT-RATE (associated with the CT-CLIP model32), comprising 209,461 CT scans from 58,811 patients, without labelling of clinical disease status. The effectiveness of OMAFound’s pretraining stage is validated on benchmark comparisons (Supplementary Tables 1 and 2) with state-of-the-art CT-focused foundation models, including MedVersa33, Merlin34 and CT-CLIP32, as well as 3D extensions of base models of DINO v236 and ResNet-5037.
A total of 209,461 CT scans from 58,811 patients, acquired over a 10-year span from 7 manufacturers across nationwide and international medical centres, were retrospectively collected to develop a task-agnostic SSL-based foundation model (Supplementary Fig. 1) capable of robust CT image feature representation. Task-specific cancer screening modules (Supplementary Fig. 2) were subsequently fine-tuned using labelled data (non-cancer, breast cancer or lung cancer) to enable organ-specific and patient-level cancer predictions. Different from low-dose CT for routine lung cancer screening, we additionally benchmark its feasibility for breast cancer screening compared with the standard mammography-based approach. OMAFound for multi-cancer screening was prospectively evaluated in four large-scale cohorts, with its performance compared with that of seven experienced generalist radiologists. An AI-assisted reader study was conducted to demonstrate the potential benefit of OMAFound in enhancing screening outcomes.
To enhance OMAFound’s performance on cancer screening, we further leverage labelled data to fine-tune task-specific downstream modules (Supplementary Fig. 2) via a weakly supervised learning adaptation stage. Labelled data in this study refers to patient-level ground-truth status, categorized as either non-cancer, breast cancer or lung cancer, determined by pathology-confirmed results or follow-up screenings, respectively. Table 1 and Extended Data Fig. 1 provide comprehensive details on CT dataset utilization and patient recruitment criteria. Extended Data Table 1 (Site A-MG, Site A-CTMG and Site G) lists the mammography datasets for comparison purposes.
Both screening (low-dose CT) and diagnostic (standard-dose CT) examinations were included for OMAFound model development for several strategic reasons. Previous research has demonstrated that including diagnostic examinations in the training process can improve model performance even when evaluating on screening examinations exclusively18. In addition, incorporating diagnostic examinations, particularly those with cancer cases, can alleviate the class imbalance encountered when training models solely on screening examinations. Moreover, given the historically low screening rates in China, most available retrospective nationwide datasets predominantly consist of diagnostic examinations, making their inclusion practically necessary for model training.
The organ-specific breast cancer screening
Owing to the non-standardized application of chest CT in breast cancer screening, we had to retrospectively collect patients who had opportunistically undergone CT scans with either pathologically confirmed breast diagnoses or remained cancer-free during follow-up observations to develop our task-specific breast module (Pbreast). Specifically, the breast module of OMAFound was developed using the fine-tuning cohort of Site A-CTbreast with 16,979 patients (6,257 breast cancer). In the internal test cohort of Site A-CTbreast containing 5,782 patients (497 breast cancer), the module showed a balanced accuracy of 74.0%, a sensitivity of 68.0% and a specificity of 79.9% (Extended Data Table 2). Subsequent assessment on an external test cohort from Site B, consisting of 1,716 patients (55 breast cancer), yielded a corresponding performance of 76.6%, 74.5% and 78.7%, respectively. The area under the receiver operating characteristic curve (AUC-ROC) for both test cohorts is illustrated in Fig. 2.
a–c, ROC curves of the CT-based OMAFound for breast cancer prediction (breast-specific module; a), lung cancer prediction (lung-specific module; b) and patient-level cancer prediction (fusion module; c). d–f, The feasibility of OMAFound for breast cancer screening compared with the standard mammography (MG) approach, assessed by the baseline of the mammography-based AI model (d), comparison between models on a paired CT-mammography dataset (e) and comparison on a subset of the paired CT–mammography dataset benchmarked against breast radiologists (f). All ROC curves are presented with a 95% confidence band.
Given that mammography remains the gold standard for breast cancer screening, we additionally developed a mammography-based AI model as a benchmark for comparison with the CT-based breast module. As shown in Supplementary Fig. 3, this model, a derivative of BMU-Net20, was initialized with its pre-trained weights and re-designed to detect patient-level breast cancer by incorporating both cranial–caudal and mediolateral oblique views of bilateral breasts, using 46,800 mammography images from 11,700 patients in Site A-MG. When evaluated on the internal test cohort of 6,329 patients (612 breast cancer) from Site A-MG, our mammography model achieved an AUC of 0.856 (95% confidence interval (CI), 0.837–0.875). This performance aligned with previous large-scale mammography AI studies17,21,38 (Supplementary Table 3) and was further validated on an external test cohort from Site G, yielding an AUC of 0.844 (95% CI, 0.807–0.880).
On the basis of the developed CT-based breast module and mammography-based AI model, we conducted a rigorous breast cancer screening comparative assessment in a new test cohort of Site A-CTMG corresponding to 1,131 patients (358 breast cancer) who underwent both imaging modalities (that is, paired CT–mammography data). The mammography-based AI model achieved a balanced accuracy of 78.4%, while the CT-based breast module presented a marginally lower balanced accuracy of 76.5% (Extended Data Table 2). Notably, the mammography-based AI model showed superior specificity (90.0%), consistent with established literature17,39. By contrast, the CT-based breast module showed enhanced sensitivity compared with the mammography-based AI model (73.2% versus 66.8%), suggesting the potential role of AI-enhanced chest CT in breast cancer detection.
To avoid the bias caused by AI models, we further conducted a mammography reader study involving 5 experienced breast radiologists (with an average of over 10 years’ experience) and the mammography-based AI model, using a subset (190 cases) from Site A-CTMG. The reader study demonstrated that our mammography-based AI model achieved non-inferior performance compared with that of experienced radiologists in breast cancer detection. This comparison served to validate the fairness of our previous model comparative analysis by establishing a human expert-based reference benchmark, as depicted in Fig. 2f. Supplementary Table 4 lists weighted F1 score, balanced accuracy, sensitivity and specificity for each reader’s mammography interpretation.
The organ-specific lung cancer screening
The task-specific lung module (Plung) was developed by fine-tuning OMAFound on a retrospective dataset of 21,680 CT scans (3,372 lung cancer) from 20,626 patients. On an internal test cohort of Site A-CTlung comprising 5,777 patients (300 lung cancer), our lung module achieved an AUC of 0.894 (95% CI, 0.881–0.906). Additional evaluation metrics and comparison with current state-of-the-art models in lung cancer screening are provided in Extended Data Table 2 and Supplementary Table 5, respectively. When evaluated on an external test cohort (PublicX), consisting of 169 patients (7 lung cancer) from the Lung Image Database Consortium (LIDC)40 dataset and 227 patients (227 lung cancer) from the LungCT41 dataset, the lung module achieved an AUC of 0.819 (95% CI, 0.778–0.861). The performance decline in the external test cohort may be attributed to the high prevalence of cancer cases within this non-screening diagnostic population.
Different from CT-based breast applications, low-dose CT is routinely implemented for lung cancer screening, resulting in the availability of public cohorts for model generalizability evaluation. In this study, the lung module is further evaluated by using the widely adopted National Lung Screening Trial (NLST)42 dataset. Leveraging the long-term follow-up screenings offered by the NLST dataset, we performed a lung cancer risk analysis that used a single low-dose CT scan to predict lung cancers occurring 1–6 years after a screen. As depicted in Supplementary Fig. 4, the lung module achieved a 1-year AUC of 0.738 (95% CI, 0.706–0.770), a 2-year AUC of 0.732 (95% CI, 0.695–0.768), a 3-year AUC of 0.726 (95% CI, 0.684–0.769), a 4-year AUC of 0.721 (95% CI, 0.668–0.773), a 5-year AUC of 0.710 (95% CI, 0.639–0.780), and a 6-year AUC of 0.703 (95% CI, 0.603–0.803).
Moreover, we assess the overall effectiveness of lung cancer risk prediction using the concordance index (C-index)43. The lung module, which was fine-tuned using only weakly supervised patient-level labels (lung cancer or non-cancer), achieved a C-index of 0.736. This performance is non-inferior to the Sybil model16, which reported a C-index of 0.75 and was developed with additional nodule annotations (strong supervision) by expert radiologists on the same NLST dataset, indicating the potential advantage of our lung module to some extent.
The patient-level cancer screening
When organ-specific screening programmes operate independently, false positives can accumulate at the patient level, leading to increased referrals and unnecessary invasive diagnostic procedures. For instance, organ-specific modules can predict cancer simultaneously (such as the breast module predicting breast cancer and the lung module predicting lung cancer), then the combined prediction suggests multiple concurrent cancers in the same patient. This contradicts clinical reality where a patient may be cancer free or have a single malignancy but rarely presents with multiple primary cancers. Therefore, implementing a patient-level cancer prediction at the initial screening stage would help mitigate the potential bias introduced by independent organ-specific predictive models.
We investigated three strategies for patient-level cancer screening. Strategy 1 uses a ‘noisy-or’ probabilistic equation 1 − (1 − Pbreast) × (1 − Plung) without requiring new AI model development. Strategy 2 involves developing a novel end-to-end fusion module (Pfusion) that builds on our previously established breast and lung modules for patient-level cancer screening (Supplementary Fig. 2c). Unlike single-window-based organ-specific modules, the fusion module integrates feature representations from multiple CT window settings (soft tissue window and lung window), enabling direct ‘cancer’ versus ‘non-cancer’ prediction at the patient level. Strategy 3, which is ultimately adopted in this study following comparative analyses, implements an integrated approach to combine results from Pbreast, Plung and Pfusion (Fig. 3a).
a, A three-phase stratification is applied to the screening participants. Given the rare occurrence of patients presenting with multiple primary cancers, a fusion module is implemented to further refine potentially incorrect predictions at the patient level. The combined results of the four medical centres are presented as male and female cohorts using ROC with a 95% confidence band. b–e, Performance of organ-level breast cancer prediction, female only (b), organ-level lung cancer prediction, female only (c), patient-level cancer prediction on female population (d), and patient-level cancer prediction on male population (e), which is identical to organ-level lung cancer prediction, male only. The error bars represent 95% CIs computed from 1,000 bootstrap resamples.
It is important to note that the incidence of breast cancer in males is extremely rare, thereby obviating the necessity to differentiate between organ-specific and patient-level cancer screening in this population. In other words, our patient-level strategy is applicable to only the female population. On the combined female-only test cohort from Site A-CTbreast and Site A-CTlung, strategy 3 achieved optimal performance balance (balanced accuracy, 78.7%; sensitivity, 87.2%; specificity, 70.1%), compared with strategy 1 (balanced accuracy, 54.2%; sensitivity, 99.5%; specificity, 8.9%) and strategy 2 (balanced accuracy, 74.2%; sensitivity, 77.1%; specificity, 71.3%).
Prospective multi-cancer screening on low-dose CT
Although the performance of OMAFound has been demonstrated in retrospective CT datasets, its clinical applicability to low-dose CT screening has not yet been explored, particularly in breast cancer screening. To address this knowledge gap, we conducted a prospective real-world multi-centre study involving 21,601 screening participants who underwent low-dose CT scans across 4 medical centres, resulting in cohorts of 10,680 patients (5,581 females) at Site C (15 breast cancer and 62 lung cancer), 1,214 patients (614 females) at Site D (12 breast cancer and 10 lung cancer), 5,181 patients (2,576 females) at Site E (14 breast cancer and 27 lung cancer), and 4,526 patients (1,911 females) at Site F (43 breast cancer and 57 lung cancer).
Figure 3a illustrates the three-phase screening flowchart, which implements a sex-stratified approach as the first step (phase 1), separating participants into male and female cohorts. This stratification reflects the epidemiological reality that the male population is typically excluded from breast cancer screening programmes. In phase 2 (organ-level cancer prediction), the male cohort undergoes analysis using the lung module (Plung), while the female cohort is evaluated using both breast and lung modules (Pbreast and Plung). For phase 3 (patient-level cancer prediction), patient-level and organ-level cancer screening are identical for the male cohort. The female cohort, however, uses the previous established integration approach (strategy 3) of Pbreast, Plung and Pfusion.
The OMAFound showed excellent performance for lung cancer prediction, with a mean balanced accuracy of 86.1% in the male cohorts. In the female cohorts, OMAFound achieved a mean balanced accuracy of 82.2% for breast cancer, 88.0% for lung cancer at organ-level prediction and a mean balanced accuracy of 82.9% at patient-level cancer prediction. Figure 3b–e and Extended Data Table 3 show the detailed results of AUC, weighted F1 score, balanced accuracy, sensitivity and specificity for each prospective cohort, respectively.
Clinical outcomes of solo radiologists versus AI-assisted radiologists
To investigate the potential clinical value of OMAFound in supporting radiologist’s decision-making, we designed a sequential CT reader study and an AI-assisted CT reader study, as shown in Fig. 4a. The test cases in the reader study were strategically sampled from prospective cohorts using differential sampling rates (higher for minority cancer cases, lower for majority non-cancer cases) to enhance the difficulty of the screening task and statistical power. As a result, the CT reader study contains 165 male patients (52 lung cancer) and 200 female patients (34 lung cancer and 59 breast cancer).
a, Workflow of the two-part CT reader study. The important sensitivity improvement for lung specific (male and female), breast specific (female only) and patient level (male and female) for each reader are presented. b–d, Improved performance for seven individual readers (R; red colour in R1–R7 indicates statistical significance P < 0.05), as measured by weighted F1 score, balanced accuracy, sensitivity and specificity, from their solo assessment (light blue) to those assisted by OMAFound (dark blue) are shown for lung specific (b), breast specific (c) and patient level (d). The dashed line represents the standalone OMAFound benchmark performance.
As shown in Fig. 4, we first compared the performance between OMAFound and seven generalist radiologists alone. It was observed that radiologists maintained high specificity (96.1% to 100.0% for lung (male and female), and 95.0% to 100.0% for breast (female)) across all cancer prediction tasks, moderate sensitivity in lung cancer screening (65.1% to 80.2% (male and female), except 39.5% for reader 6), but limited sensitivity in breast cancer screening (16.9% to 49.2% (female)) especially for junior radiologists. By contrast, OMAFound achieved high sensitivity for both lung (90.7% (male and female)) and breast (86.4% (female)), with overall non-inferior performance in lung cancer prediction and substantially superior performance in breast cancer prediction.
An AI-assisted CT reader study was subsequently performed to evaluate the benefits of AI assistance to radiologists (Extended Data Table 4). To achieve this, we used the original reader’s assessment as the baseline for each reader. In addition to the original low-dose CT scans, corresponding heatmaps and OMAFound predictions of malignancy risk probability were both presented to the same readers to help them understand the justification of the AI predictions. According to the reader’s feedback, OMAFound could potentially guide them to making a better clinical outcome at the organ level, with a mean sensitivity improvement of 38.9% in breast cancer detection and 16.0% in lung cancer detection, without sacrificing the specificity. In terms of patient-level cancer presence prediction, OMAFound attained an eclectic performance with a mean sensitivity improvement of 21.3%.
The interpretability of OMAFound
To understand the regions influencing cancer predictions, we compared five post hoc interpretability approaches, including four based on class activation mapping (CAM) and one attention-based algorithm (Methods). We requested experienced radiologists’ comments on the correlation between each interpretable heatmap (all slice heatmaps including one representative slice with highest ranked activation score are provided) and the anatomical locations of different cancer types and their origins (Fig. 5a). Finer-CAM was eventually adopted in this study based on the majority voting.
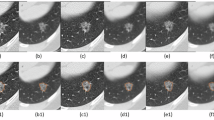
a, Heatmaps generated by five different post hoc interpretable approaches, including Grad-CAM, Grad-CAM++, Layer-CAM, Finer-CAM and attention-based GMAR. For breast cases, the right breast should match with the left side of the CT image due to anatomical opposite. b, Examples of non-cancer, lung cancer and breast cancer discrimination by OMAFound using preferable Finer-CAM, which were missed or partially missed by readers but well classified with the assistance of OMAFound. More examples including missed cases by OMAFound are shown in Extended Data Figs. 2 and 3.
We specifically analysed the attention made by OMAFound (Fig. 5b and Extended Data Figs. 2 and 3). For cancer cases, the focus of OMAFound concentrated primarily on the target organ and its immediate vicinity. In breast cancer cases, the highlighted regions predominantly included soft tissue areas in the lateral thorax, particularly the parenchyma. For lung cancer cases, the attention centred on the thoracic cavity, specifically focusing on nodular tissues. Given that chest CT is not the standard breast cancer screening modality, these interpretable heatmaps may offer valuable educational potential by helping clinicians identify breast cancer appearances in CT scans.
Both radiologists and AI models are susceptible to prediction errors, yet they exhibit distinct error profiles. Radiologists, with extensive training in radiological image interpretation, possess domain expertise in cancer appearances and origins. Their errors predominantly occur in missing cancer cases, especially small nodules and low-contrast lesions, resulting in lower sensitivity but preserved high specificity. Conversely, the data-driven OMAFound model makes errors in both cancer and non-cancer cases, demonstrating a balanced trade-off between sensitivity and specificity.
Discussion
Non-contrast CT, particularly low-dose CT, has been widely recommended for population-based cancer screening across many countries owing to its cost-effectiveness and reduced radiation exposure. However, current screening programmes follow a ‘single test for one cancer’ policy, failing to capitalize on the opportunity to maximize cancer detection from a single screening examination. In this study, we proposed OMAFound, an AI model that shifts towards a ‘single test for multi-cancer’ paradigm by leveraging all potential cancer biomarkers present within a single low-dose CT scan. Through large-scale real-world retrospective and prospective validation across multiple centres, OMAFound showed robust performance, highlighting the notable insights for enhancing existing screening programmes without incurring additional costs.
Conventional predictive AI models show limited cross-organ generalizability owing to organ-specific supervision and the resource-constrained nature of obtaining expert-annotated labelled data. To achieve cost-effective multi-cancer prediction, we developed a task-agnostic SSL-based foundation model that leverages large-scale unlabelled CT scans from diverse ethnic populations, varying dose levels and different scanner manufacturers. The superiority of OMAFound in extracting robust, generalizable CT feature representations has been validated through benchmark comparisons with state-of-the-art CT-focused foundation models such as MedVersa33, Merlin34 and CT-CLIP32, as well as 3D extensions of DINO v236 and ResNet 5037.
For organ-specific cancer screening, our downstream modules fine-tuned with weakly supervised patient-level labels showed good generalizability on large-scale representative CT test datasets. The lung module of OMAFound achieved AUCs of 0.819–0.955 across one standard-dose cohort (Site A-CTlung), four low-dose cohorts (Sites C–F) and two public cohorts (LIDC and LungCT), performing on par with established benchmark lung cancer screening models (AUCs 0.820–0.944). Similar generalizability was observed for the breast module of OMAFound across one external standard-dose cohort (Site B) and four low-dose cohorts (Sites C–F), with AUCs of 0.845–0.959. These results collectively underscore the clinical applicability of OMAFound for CT-based cancer screening.
Beyond organ-specific cancer screening, we evaluated OMAFound’s performance at the patient level via an integrated analytical approach. This integration strategy incorporated clinical knowledge (for instance, the rare occurrence of synchronous primary lung and breast cancers in clinical practice) to alleviate errors in predictive AI models, resulting in a higher cancer prediction accuracy than both the ‘noisy-or’ probabilistic equation and a simple end-to-end fusion module. The patient-level analysis proved particularly valuable for identifying high-risk individuals during initial screening, enabling efficient triage for targeted organ-specific cancer screening and diagnostic workup.
Given the non-standard role of chest CT in breast cancer screening, we specifically focused on breast performance analysis. Using paired CT–mammography data, we performed a systematic comparison between our CT-based breast module and mammography-based AI model, with the latter validated by five experienced mammography specialists. The mammography-based AI model achieved high performance (AUC 0.859), aligning with clinical expectations given mammography’s decades-long validation as the screening gold standard10. The CT-based breast module showed comparable performance (AUC 0.793), suggesting that existing imaging data, such as low-dose chest CT scans obtained during lung cancer screening in female individuals, could be leveraged for opportunistic breast cancer screening.
The multi-cancer screening capability of OMAFound has substantial clinical implications, offering robust preventive medicine strategies without incurring additional costs in terms of monetary, radiation or time. Although our current study focuses on chest CT scans for detecting the most prevalent cancers (lung and breast), future extensions of our model could potentially incorporate other types of lesion and neoplasm, moving towards comprehensive multi-cancer screening similar to liquid biopsy approaches44.
As clinical applicability is an important criterion for medical AI models, we evaluated OMAFound against experienced radiologists and investigated its advantages as a screening aid for multi-cancer prediction using low-dose CT. Our reader studies showed that OMAFound outperformed the majority of radiologists. Integration of OMAFound into the screening workflow yielded substantial improvements in the sensitivity of readers, particularly junior radiologists, with mean increases of 38.9% in breast (5 out of 7 with P < 0.05, remarkable opportunistic screening for breast cancer), 16.0% in lung (3 out of 7 with P < 0.05) and 21.3% at patient level (6 out of 7 with P < 0.05), without loss of specificity. Such a high sensitivity of OMAFound constitutes a substantial advantage for screening programmes in which minimizing missed cancer cases is a priority.
Transparent decision-making remains crucial in healthcare45. Current AI explainability approaches fall into two main categories: post hoc explanations for unconstrained black-box models and intrinsically interpretable models such as prototype-based46. Previous studies47,48,49 on unstructured image data analysis indicate that black-box models learning hierarchical representations from raw pixels generally achieve superior performance compared with intrinsically interpretable models, highlighting the fundamental trade-off between model accuracy and interpretability. In our comparative analysis of five post hoc explanation methods, we observed varying saliency patterns, making it difficult to attribute these discrepancies to the model or to the explanation methods (or to both)—an unresolved trustworthiness challenge in medical AI50,51. Finer-CAM is preferable in this study because it more closely aligns with radiologists’ interpretations and is an improved version of Grad-CAM, which has been widely used in large-scale medical studies19,20,22.
There are a few limitations to our study. First, although we implemented various post hoc interpretability approaches to enhance transparent decision-making, studies indicate that the qualitative heatmap visualization generally has biases compared with expert radiologists regardless of model classification accuracy51. More advanced interpretability approaches should be investigated in the future. Second, the single patient-level label (low semantic information) is insufficient to improve the model’s predictive power. Strong patch-level lesion annotations, such as segmentation masks or detection boxes, could both improve predictive accuracy and enable interpretable localization analyses. Finally, OMAFound was currently limited to predict current cancer risk from a single CT scan. Future research should investigate personalized screening intervals based on individual risk stratification (low, moderate or high risk).
To conclude, we have developed OMAFound for image-based multi-cancer screening with improved generalizability. OMAFound was prospectively evaluated on low-dose CT scans from four medical centres under the evaluation tasks of organ-specific cancer type and patient-level cancer presence predictions, demonstrating performance that can assist clinicians in improving screening outcomes. The ‘single test for multi-cancer’ capability represents a step towards improved screening programmes in clinical scenarios.
Methods
Ethics approval
All retrospective non-public datasets (Sites A, B and G) in this investigation were approved by the institutional review board (IRB) of the hospitals with a waiver granted for the requirement of informed consent. With respect to the prospective study pre-registered at www.chictr.org.cn (identifier ChiCTR2400081249), all participants signed an informed consent developed and approved by the IRB of Sites C, D, E and F. All datasets were de-identified before model development and test in this investigation.
Chest CT dataset
Our study incorporated ten distinct CT datasets, including six Chinese (Sites A to F) and four international public datasets (CT-RATE, NLST, LIDC and LungCT). These datasets represented diverse clinical settings (emergency rooms, physical examination centres, inpatient and outpatient departments) and included scans from seven manufacturers (GE, Philips, SIEMENS, TOSHIBA, MinFound, UIH and Neusoft). Site A, Site B and all public datasets were characterized as retrospective cohorts used for the development and testing of the OMAFound model, while the remaining datasets (Sites C to F) provided prospective low-dose CT scans from screening populations for real-world validation.
The datasets were categorized into two types based on clinical interpretation availability. The first type consisted of unlabelled data (Site A-CTunlabeled and CT-RATE), which provided large-scale datasets exclusively for task-agnostic foundation model pretraining. The second type was weakly supervised labelled data with patient-level ground-truth status, confirmed either by pathology (cancer or non-cancer) or at least 2 years (unless otherwise specified) follow-up for non-cancer status confirmation. Within the labelled data, two labelling patterns emerged: retrospective datasets (Site A-CTbreast, Site A-CTlung, Site B, NLST, LIDC and LungCT) contained a single label per patient (either breast or lung), while prospective datasets (Sites C to F) provided comprehensive dual labelling, including both breast and lung assessments for each patient.
For model training, all eligible examinations per patient were utilized, whereas only a single CT scan per patient was used for model testing. To prevent the risk of label leakage, anonymized patient IDs were used across all datasets, ensuring no patient overlaps between training and test cohorts (all scans from the same patient were assigned to the same cohort). Table 1 and Extended Data Fig. 1 provide comprehensive details on dataset utilization and patient assignment criteria. Additional dataset specifications are provided below.
Site A (The First Affiliated Hospital of Anhui Medical University). Data were retrospectively collected from multiple clinical settings (emergency rooms, inpatient and outpatient departments) between October 2015 and April 2024, which were subsequently divided into unlabelled and labelled datasets. The Site A-CTunlabeled dataset comprised 159,273 unlabelled CT scans from 37,507 patients. The labelled data were further categorized into Site A-CTbreast dataset, containing scans from 16,007 non-cancer patients and 6,754 patients with breast cancer, and Site A-CTlung dataset, consisting of scans from 23,785 non-cancer patients and 3,672 patients with lung cancer. For the organ-specific adaptation phase, labelled data were randomly and selectively allocated to the fine-tuning cohort (most cancer cases were used here to alleviate class imbalance issue on training) and the internal test cohort.
Site B (No.2 People’s Hospital of Fuyang City). Standard-dose CT scans were retrospectively collected from the outpatient department between February 2020 and May 2024, resulting in a total of 1,716 labelled CT from 1,716 patients (1,661 non-cancer patients and patients with 55 breast cancer). Site B was used solely for external testing of the breast module of OMAFound to assess generalizability.
Site C (physical examination centres affiliated to Site A). Low-dose CT scans were collected through a pre-registered prospective study. A total of 10,680 screening participants were enroled between January 2024 and December 2024. The cohort comprised 10,603 non-cancer cases, confirmed through 6–12 months of short-term follow-up. The remaining cases included 15 breast cancer cases and 62 lung cancer cases (24 from female and 38 from male), all confirmed by pathology results. Site C was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site D (Lu’an People’s Hospital). Low-dose CT scans were prospectively collected from 1,214 screening participants between January 2024 and July 2024. Disease statuses were determined through either 6–12 months of short-term follow-up or pathology confirmation, identifying 1,192 non-cancer cases and 22 cancer cases (12 breast cancer, 4 female lung cancer and 6 male lung cancer). Site D was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site E (Weifang Traditional Chinese Hospital). Between January 2024 and December 2024, a total of 5,181 low-dose CT scans were prospectively collected during annual physical examinations. These scans represented 5,140 non-cancer patients, 14 patients with breast cancer and 27 patients with lung cancer (14 from female and 13 from male). Site E was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site F (Xuancheng People’s Hospital). We prospectively enroled participants from a local screening population for low-dose CT scans. Following standardized prospective labelling criteria, 4,426 non-cancer patients, 43 patients with breast cancer and 57 patients with lung cancer (35 from female and 22 from male) were finally collected between January 2024 and December 2024. Site F was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
CT-RATE (non-contrast chest CT dataset32). This public dataset was collected at Istanbul Medipol University Mega Hospital between May 2015 and January 2023. It comprises 50,188 unlabelled CT data from 21,304 unique patients. CT-RATE was used solely for task-agnostic foundation model pretraining.
NLST (National Lung Screening Trial42). The NLST dataset was collected across 33 US medical institutions, with participants randomized to receive annual low-dose CT screenings between August 2002 and 2007. In total, 41,805 labelled CT scans from 19,698 patients (18,717 non-cancer patients and 981 patients with lung cancer) were included, with long-term follow-up data available. A random subset (12.7%) at the patient level was allocated to the internal test cohort, while the remaining scans were used for training. NLST was used solely for multi-year lung cancer risk prediction, where a single low-dose CT scan was used to predict lung cancer occurrence 1–6 years post-screening.
PublicX (combined LIDC40 and LungCT41 datasets). The LIDC dataset with a mix of standard-dose and low-dose scans were collected from five different institutions between 1998 and 2010. The LungCT dataset contains standard-dose CT scans acquired between July 2004 and June 2011. On the basis of the same inclusion criteria for the nationwide dataset, the PublicX dataset includes 396 labelled CT data from 396 patients (162 non-cancer patients and 234 patients with lung cancer). The PublicX dataset was used solely for external testing of the lung module of OMAFound to assess generalizability.
Mammography dataset
Given mammography’s status as the current gold standard for breast cancer screening, we developed a mammography-based AI model as a benchmark for comparison with the CT-based OMAFound. For this purpose, we retrospectively collected a dedicated mammography-only dataset, designated as Site A-MG to distinguish it from chest CT data of Site A, for the development and evaluation of this mammography-based AI model.
Specifically, Site A-MG includes 72,116 mammography images from 18,029 patients (bilateral cranial–caudal and mediolateral oblique views per patient), acquired between January 2014 and December 2023 from either a GE Senographe DS mammography system or Hologic Selenia Dimensions mammography system, covering both screening and diagnostic populations. To assess the generalizability of our mammography-based AI model, we assembled an external test cohort from Anhui No.2 Provincial People’s Hospital (Site G). This cohort contained 3,280 mammography images from 820 patients (158 cancer-positive cases), retrospectively collected between March 2023 and August 2024 using a GE Senographe DS mammography system.
The labels of these mammography datasets were confirmed either by pathology (cancer or non-cancer) or through a minimum follow-up period of 2 years for non-cancer status confirmation. Detailed patient characteristics and labels are provided in Extended Data Table 1.
Paired CT–mammography dataset
Recognizing that model performance can vary across different populations and clinical settings, we thus established a more equitable comparison between the mammography-based AI model and CT-based OMAFound for breast cancer screening. That is, we additionally collected 1,131 paired CT and mammography scans from 1,131 patients (Extended Data Table 1), namely, as Site A-CTMG. Importantly, Site A-CTMG data had no overlap with either Site A-CTbreast or Site A-MG datasets.
OMAFound model
Image preprocessing before OMAFound model development was performed using Torchvision (version 0.20.1) and SciPy (version 1.14.1). The multi-institutional CT dataset showed slice spacing variations from 0.625 mm to 5 mm. To harmonize the difference in slice thickness and spatial resolution, all CT scans were resampled to a uniform 1 × 1 × 1 mm before resizing to voxel dimensions of 128 × 128 × 128. Intensity distributions (Hounsfield units) were standardized using min–max normalization, and foreground regions of lung window and soft tissue window were extracted from each scan. In this study, the model development process did not incorporate any image annotations, such as lesion bounding boxes or segmentation masks.
The architecture of the SSL-based OMAFound model is detailed in Supplementary Fig. 1 and the task-specific downstream modules are shown in Supplementary Fig. 2. For the foundation model, we used the encoder from SwinUNETR-V235 as the backbone for feature extraction, integrating 3D stage-wise convolution and shifted window-based self-attention mechanisms. A residual convolution (ResConv) block was added at the beginning of each resolution level, followed by a Swin transformer block.
In the organ-specific breast and lung modules, a 3D adaptive average pooling layer was utilized to aggregate spatial features, followed by a fully connected layer and softmax activation for cancer risk prediction task. Specifically, the breast module and lung module of OMAFound were developed using the fine-tuning cohort of Site A-CTbreast and Site A-CTlung, respectively.
For the fusion module, the encoders for the breast and lung branches were initialized with weights from the corresponding organ-specific modules and kept frozen during fusion training. Each encoder produced a 768-dimensional feature vector, which was used to generate classification logits and uncertainty estimates. A learnable class token was concatenated with the two feature vectors and passed through a transformer encoder to capture cross-organ interactions. The final cancer prediction was derived from the updated class token, and the total loss was calculated as the sum of the fusion loss and organ-specific uncertainty losses. The fusion module was developed using combined fine-tuning datasets from both breast and lung modules and tested on merged internal test cohorts of Site A-CTbreast and Site A-CTlung.
OMAFound was implemented using the PyTorch framework (version 2.5.1), and training was conducted using two Intel Xeon central processing units and eight NVIDIA A100 80GB graphics processing units. Inspired by previous research52, the objective of the SSL module was to minimize a combination of rotation loss, reconstruction loss and contrastive loss. For downstream tasks, label smoothing loss was applied. Optimization was performed using the adaptive moment estimation (ADAMW) optimizer, with a batch size of 96 and an initial learning rate of 0.0001. A linear warm-up ratio of 0.1 was applied, followed by a cosine function learning rate schedule. Training was capped at 15 epochs, with early stopping triggered if no further loss improvement was observed.
To address class imbalance, weighted sampling was used to ensure balanced representation of all classes during training. Data augmentation included random affine transformations (translation and scaling within the bounds of (0.1, 0.1, 0.1), random rotations (up to 15°), contrast adjustment with a random factor between 0.8 and 1.2, and the addition of random noise with intensities ranging from 0.005 to 0.05. All augmentations were constrained to maintain pixel values within the [0, 1] range.
Mammography-based AI model
To compare chest CT with the standard mammography-based approach for breast cancer screening, we developed an individual mammography-based AI model using the dataset from Site A-MG. Mammography scans containing both cranial–caudal and mediolateral oblique views of the bilateral breast were included for model development.
Supplementary Fig. 3 illustrates the architecture of the mammography-based AI model. The model, a derivative of BMU-Net20, integrates a ResNet-18 backbone with a transformer encoder for multi-view breast cancer classification. The ResNet-18 backbone, initialized with weights transferred from the large-scale, pre-trained Mirai model21, was used to extract features from each individual view. These features were then augmented with positional embeddings and passed through the transformer encoder to capture contextual dependencies across views. Separate classifiers were applied to each view, and their outputs were weighted by learnable parameters specific to the left and right sides. The final logit was obtained by averaging the weighted outputs.
Reader study on mammography
We conducted a mammography reader study to compare the performance of the mammography-based AI model with that of experienced breast radiologists. To be specific, each reader independently reviewed the same set of cases and assigned a BI-RADS (Breast Imaging Reporting and Data System) 5th edition53 rating using the values 1, 2, 3, 4a, 4b, 4c and 5, simulating routine clinical interpretation. To convert BI-RADS assessments into binary classification for sensitivity and specificity calculations, BI-RADS 4a or higher were considered as test positive, and all others negative. The average reader sensitivity and specificity were computed by averaging the individual sensitivity and specificity values across all readers. All readers were blinded to each other’s assessments, the original clinical reports and the AI model outputs. The study included 5 board-certified radiologists specializing in mammography, each with over 10 years of clinical experience. A total of 190 examinations—randomly selected from the test cohort of the Site A-CTMG dataset—were presented to the readers in a randomized order.
Reader study on low-dose CT
To evaluate the clinical utility of OMAFound in assisting generalist radiologists with improved screening outcomes, we conducted a 2-part CT reader study involving 365 patients (220 non-cancer, 59 breast cancer, 34 female lung cancer and 52 male lung cancer). Cases were randomly and selectively sampled (higher for cancer cases and lower for non-cancer cases to enhance the difficulty of the screening task and statistical power) from the prospective cohorts of Sites C, D, E and F. Seven board-certified generalist radiologists participated in this study, with their clinical experience summarized in Extended Data Table 4.
The sequential reader study consisted of a first reading (solo) and a second reading (+OMAFound). Each reader was requested to finish three tasks, including organ-level breast cancer detection, organ-level lung cancer detection and patient-level cancer presence prediction. During the first reading, each reader independently reviewed the same set of testing cases without time limit and provided initial binary decisions for each task (‘Yes’ for cancer, ‘No’ for non-cancer). In the second reading, readers were provided with OMAFound-generated heatmaps and prediction scores as a decision support. They were allowed to update their initial assessments based on the AI assistance.
Interpretability of the OMAFound model
To assure trust from human experts, it is essential to make the model’s decision-making process interpretable. In this study, we implemented and analysed five post-hoc explanation approaches, including four CAM-based (Grad-CAM54, Grad-CAM + + 55, Layer-CAM56 and Finer-CAM57) and one attention-based gradient-driven multi-head attention rollout (GMAR58) mappings, to visualize the heatmap localization regions that could aid human experts to understand the justification of the AI system for the cancer risk predictions. All post hoc methods in this study were applied to the normalization layer of the final stage of the model for each test image.
Specifically, Grad-CAM++ enhances Grad-CAM by implementing pixel-wise weights instead of channel-wise weights, improving small object localization capability. Layer-CAM generates more reliable boundary definitions by utilizing pixel-level activation with positive gradients within and across layers. Finer-CAM extends Layer-CAM by incorporating progressive cross-layer refinement and denoising, achieving superior semantic alignment. GMAR is a novel method to quantify the importance of each attention head using gradient-based scores.
Statistical analysis
The performance of the OMAFound model and the mammography-based AI model was evaluated using weighted F1 score, balanced accuracy, sensitivity, specificity and the AUC. The 95% CIs of the weighted F1 score, balanced accuracy and specificity were computed using 1,000 non-parametric bootstrap resamples. A dynamic approach (Wilson CIs and bootstrap-based CIs) was used for sensitivity due to low cancer prevalence. The C-index43 was computed to evaluate the predictive performance of time-to-event models. AUC comparisons were conducted using Delong’s test. All comparisons were two-sided, with a P value <0.05 considered statistically significant. All statistical analyses were performed using SPSS (version 22.0), and relevant Python packages.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The CT, mammography and paired CT–mammography datasets from participating medical centres are protected due to patient privacy and IRB restrictions. However, a portion of the test data can be made available for academic purposes from the lead corresponding author X.Q. upon reasonable request, with the approval of the hospital IRBs and a signed data-use agreement. The public datasets can be accessed from: CT-RARE at https://huggingface.co/datasets/ibrahimhamamci/CT-RATE, NLST at https://biometry.nci.nih.gov/cdas/learn/nlst/images/, LIDC at https://cancerimagingarchive.net/collection/lidc-idri, and LungCT at https://cancerimagingarchive.net/collection/lung-pet-ct-dx. Source data are provided with this paper.
Code availability
The source codes of OMAFound are available via GitHub at https://github.com/Qian-IMMULab/OMAFound (ref. 59). Custom codes for the deployment of the AI system are available for research purposes from the lead corresponding author X.Q. upon reasonable request.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Malik, V. S., Willett, W. C. & Hu, F. B. Global obesity: trends, risk factors and policy implications. Nat. Rev. Endocrinol. 9, 13–27 (2013).
Islami, F., Siegel, R. L. & Jemal, A. The changing landscape of cancer in the USA—opportunities for advancing prevention and treatment. Nat. Rev. Clin. Oncol. 17, 631–649 (2020).
Allemani, C. et al. Global surveillance of cancer survival 1995–2009: analysis of individual data for 25 676 887 patients from 279 population-based registries in 67 countries (CONCORD-2). Lancet 385, 977–1010 (2015).
Smith, R. A. et al. American Cancer Society guidelines for the early detection of cancer. CA Cancer J. Clin. 52, 8–22 (2002).
Crosby, D. et al. Early detection of cancer. Science 375, eaay9040 (2022).
The National Lung Screening Trial Research Team. Reduced lung-cancer mortality with low-dose computed tomographic screening. N. Engl. J. Med. 365, 395–409 (2011).
Oudkerk, M., Liu, S., Heuvelmans, M. A., Walter, J. E. & Field, J. K. Lung cancer LDCT screening and mortality reduction—evidence, pitfalls and future perspectives. Nat. Rev. Clin. Oncol. 18, 135–151 (2021).
Berry, D. A. et al. Effect of screening and adjuvant therapy on mortality from breast cancer. N. Engl. J. Med. 353, 1784–1792 (2005).
Bleyer, A. & Welch, H. G. Effect of three decades of screening mammography on breast-cancer incidence. N. Engl. J. Med. 367, 1998–2005 (2012).
de Koning, H. J. et al. Reduced lung-cancer mortality with volume CT screening in a randomized trial. N. Engl. J. Med. 382, 503–513 (2020).
Bankier, A. A., Levine, D., Halpern, E. F. & Kressel, H. Y. Consensus interpretation in imaging research: is there a better way? Radiology 257, 14–17 (2010).
Benchoufi, M., Matzner-Lober, E., Molinari, N., Jannot, A.-S. & Soyer, P. Interobserver agreement issues in radiology. Diagn. Interv. Imaging 101, 639–641 (2020).
Ardila, D. et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 954–961 (2019).
Jacobs, C. et al. Deep learning for lung cancer detection on screening CT scans: results of a large-scale public competition and an observer study with 11 radiologists. Radiol. Artif. Intell. 3, e210027 (2021).
Mikhael, P. G. et al. Sybil: a validated deep learning model to predict future lung cancer risk from a single low-dose chest computed tomography. J. Clin. Oncol. 41, 2191 (2023).
McKinney, S. M. et al. International evaluation of an AI system for breast cancer screening. Nature 577, 89–94 (2020).
Lotter, W. et al. Robust breast cancer detection in mammography and digital breast tomosynthesis using an annotation-efficient deep learning approach. Nat. Med. 27, 244–249 (2021).
Qian, X. et al. Prospective assessment of breast cancer risk from multimodal multiview ultrasound images via clinically applicable deep learning. Nat. Biomed. Eng. 5, 522–532 (2021).
Qian, X. et al. A multimodal machine learning model for the stratification of breast cancer risk. Nat. Biomed. Eng. 9, 356–370 (2025).
Yala, A. et al. Toward robust mammography-based models for breast cancer risk. Sci. Transl. Med. 13, eaba4373 (2021).
Cao, K. et al. Large-scale pancreatic cancer detection via non-contrast CT and deep learning. Nat. Med. 29, 3033–3043 (2023).
Ng, A. Y. et al. Prospective implementation of AI-assisted screen reading to improve early detection of breast cancer. Nat. Med. 29, 3044–3049 (2023).
Eisemann, N. et al. Nationwide real-world implementation of AI for cancer detection in population-based mammography screening. Nat. Med. 31, 917–924 (2025).
Black, W. C. et al. Cost-effectiveness of CT screening in the National Lung Screening Trial. N. Engl. J. Med. 371, 1793–1802 (2014).
Sodickson, A. et al. Recurrent CT, cumulative radiation exposure, and associated radiation-induced cancer risks from CT of adults. Radiology 251, 175–184 (2009).
Smith-Bindman, R. et al. Projected lifetime cancer risks from current computed tomography imaging. JAMA Intern. Med. 185, 710–719 (2025).
Rubin, G. D. Computed tomography: revolutionizing the practice of medicine for 40 years. Radiology 273, S45–S74 (2014).
Krishnan, R., Rajpurkar, P. & Topol, E. J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 6, 1346–1352 (2022).
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Xu, H. et al. A whole-slide foundation model for digital pathology from real-world data. Nature 630, 181–188 (2024).
Hamamci, I. E. et al. Developing generalist foundation models from a multimodal dataset for 3D computed tomography. Preprint at https://arxiv.org/abs/2403.17834 (2024).
Zhou, H.-Y. et al. MedVersa: a generalist foundation model for medical image interpretation. Preprint at https://arxiv.org/abs/2405.07988 (2024).
Blankemeier, L. et al. Merlin: a vision language foundation model for 3D computed tomography. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-4546309/v1 (2024).
He, Y. et al. SwinUNETR-v2: stronger swin transformers with stagewise convolutions for 3D medical image segmentation. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2023 Lecture Notes in Computer Science Vol. 14223 (eds Greenspan, H. et al.) https://doi.org/10.1007/978-3-031-43901-8_40 (Springer, 2023).
Oquab, M. et al. Dinov2: learning robust visual features without supervision. Preprint at https://arxiv.org/abs/2304.07193 (2023).
Hara, K., Kataoka, H. & Satoh, Y. Learning spatio-temporal features with 3D residual networks for action recognition. In Proc. IEEE International Conference on Computer Vision Workshops 3154–3160 (IEEE, 2017).
Wu, N. et al. Deep neural networks improve radiologists’ performance in breast cancer screening. IEEE Trans. Med. Imaging 39, 1184–1194 (2019).
Lehman, C. D. et al. Diagnostic accuracy of digital screening mammography with and without computer-aided detection. JAMA Intern. Med. 175, 1828–1837 (2015).
Armato III, S. G. et al. The Lung Image Database Consortium (LIDC) and Image Database Resource Initiative (IDRI): a completed reference database of lung nodules on CT scans. Med. Phys. 38, 915–931 (2011).
Clark, K. et al. The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26, 1045–1057 (2013).
National Lung Screening Trial Research Team. The National Lung Screening Trial: overview and study design. Radiology 258, 243–253 (2011).
Uno, H., Cai, T., Pencina, M. J., D’Agostino, R. B. & Wei, L.-J. On the C-statistics for evaluating overall adequacy of risk prediction procedures with censored survival data. Stat. Med. 30, 1105–1117 (2011).
Wan, J. C., Sasieni, P. & Rosenfeld, N. Promises and pitfalls of multi-cancer early detection using liquid biopsy tests. Nat. Rev. Clin. Oncol. 22, 566–580 (2025).
Ng, M. Y., Kapur, S., Blizinsky, K. D. & Hernandez-Boussard, T. The AI life cycle: a holistic approach to creating ethical AI for health decisions. Nat. Med. 28, 2247–2249 (2022).
Chen, C. et al. This looks like that: deep learning for interpretable image recognition. In 33rd Conference on Neural Information Processing Systems (NeurIPS 2019) https://proceedings.neurips.cc/paper_files/paper/2019/file/adf7ee2dcf142b0e11888e72b43fcb75-Paper.pdf (2019).
London, A. J. Artificial intelligence and black-box medical decisions: accuracy versus explainability. Hastings Cent. Rep. 49, 15–21 (2019).
Imrie, F., Davis, R. & van der Schaar, M. Multiple stakeholders drive diverse interpretability requirements for machine learning in healthcare. Nat. Mach. Intell. 5, 824–829 (2023).
Yan, L. et al. A domain knowledge-based interpretable deep learning system for improving clinical breast ultrasound diagnosis. Commun. Med. 4, 90 (2024).
Arun, N. et al. Assessing the trustworthiness of saliency maps for localizing abnormalities in medical imaging. Radiol. Artif. Intell. 3, e200267 (2021).
Saporta, A. et al. Benchmarking saliency methods for chest X-ray interpretation. Nat. Mach. Intell. 4, 867–878 (2022).
Tang, Y. et al. Self-supervised pre-training of Swin transformers for 3D medical image analysis. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 20730–20740 (IEEE, 2022).
Radiology, A. C. o., D’Orsi, C. J., Sickles, E. A., Mendelson, E. B. & Morris, E. A. ACR BI-RADS Atlas: Breast Imaging Reporting and Data System; Mammography, Ultrasound, Magnetic Resonance Imaging, Follow-up and Outcome Monitoring, Data Dictionary (American College of Radiology, 2013).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. In Proc. IEEE International Conference on Computer Vision 618–626 (IEEE, 2017).
Chattopadhay, A., Sarkar, A., Howlader, P. & Balasubramanian, V. N. Grad-CAM++: generalized gradient-based visual explanations for deep convolutional networks. In IEEE Winter Conference on Applications of Computer Vision (WACV) 839–847 (IEEE, 2018).
Jiang, P.-T., Zhang, C.-B., Hou, Q., Cheng, M.-M. & Wei, Y. LayerCAM: exploring hierarchical class activation maps for localization. IEEE Trans. Image Process. 30, 5875–5888 (2021).
Zhang, Z. et al. Finer-CAM: spotting the difference reveals finer details for visual explanation. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 9611–9620 (2025).
Jo, S., Jang, G. & Park, H. GMAR: gradient-driven multi-head attention rollout for vision transformer interpretability. Preprint at https://arxiv.org/abs/2504.19414 (2025).
Qian, X. et al. OMAFound. GitHub https://github.com/Qian-IMMULab/OMAFound (2025).
Acknowledgements
We thank the radiologists for their participation in our reader study and heatmap analysis. We are also grateful to D. Yu, Y. Chen, Z. Zhou, Q. Song, P. Liu and X. Zhang for their contributions to image preprocessing. This study was supported by the National Natural Science Foundation of China (82371993 to X.Q.), the Anhui Province Health Scientific Research Project (AHWJ2023A20096 to J.P.), the First Affiliated Hospital of Anhui Medical University Clinical Research Initiation Plan Project (LCYJ2021YB008 to J.P.) and HPC Computing Platform of ShanghaiTech University.
Author information
Authors and Affiliations
Contributions
X.Q. conceived of, designed and supervised the project. J.P. provided clinical expertise and co-supervised for the study. Z.L., Y.W. and C. Hu executed the research and developed the deep learning framework and software tools necessary for the experiments. Q.N., Jinmei Wang and C. Han analysed and interpreted the data and defined the clinical labels. Q.L., B.Z., X.H., Zhaorui Wang, X.W., C.L., Y.Z., Jingjing Wang, Zikang Wang and Y.N. collected the raw CT, mammography, paired CT–mammography and patients’ pathology/follow-ups results in clinic. X.Q. conducted the literature search and wrote the paper. All authors contributed to the review and editing of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Health thanks Eleftherios Trivizakis, Matthew Warner-Smith and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ben Johnson, in collaboration with the Nature Health team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of the flowchart of patient’s recruitment and assignment.
All patients were selected based on pre-defined inclusion and exclusion criteria. Only non-contrast chest CT scans with complete anatomical coverage of both breast and lung regions were considered. The study population was derived from both retrospective and prospective data collections, with CT scans acquired between 2015 and 2024 across various institutions.
Extended Data Fig. 2 Female examples of radiologist’s decision with and without AI assistance.
Each case was first read independently by radiologists, then re-read with AI-generated prediction score and heatmaps overlaid on CT images. Heatmaps were derived from the normalization layer in the final stage to highlight regions contributing to the prediction. (a) A lung cancer case which was correctly identified by OMAFound. (b) lung cancer and (c) breast cancer cases which were overdiagnosed by OMAFound.
Extended Data Fig. 3 Male examples of radiologist’s decision with and without AI assistance.
Each case was first read independently by radiologists, then re-read with AI-generated prediction score and heatmaps overlaid on CT images. Heatmaps were derived from the normalization layer in the final stage to highlight regions contributing to the prediction. Lung cancer cases which were (a) identified and (b) missed by OMAFound. (c) A non-lung cancer case which was overdiagnosed by OMAFound.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–4 and Tables 1–5.
Source data
Source Data Fig. 2 (download XLSX )
Statistical data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical data for Fig. 3.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, Z., Niu, Q., Wang, J. et al. A foundation model for breast and lung cancer screening using non-contrast computed tomography. Nat. Health 1, 403–415 (2026). https://doi.org/10.1038/s44360-026-00055-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44360-026-00055-8